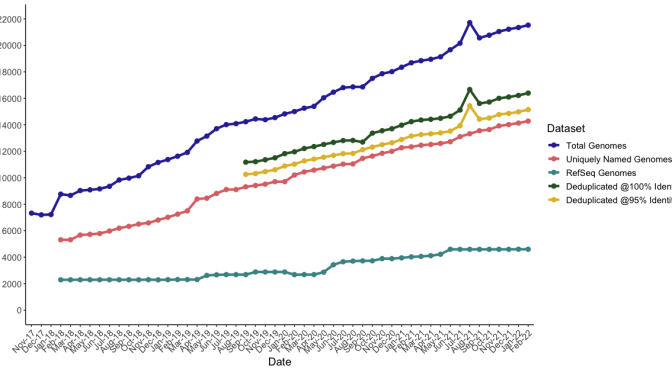
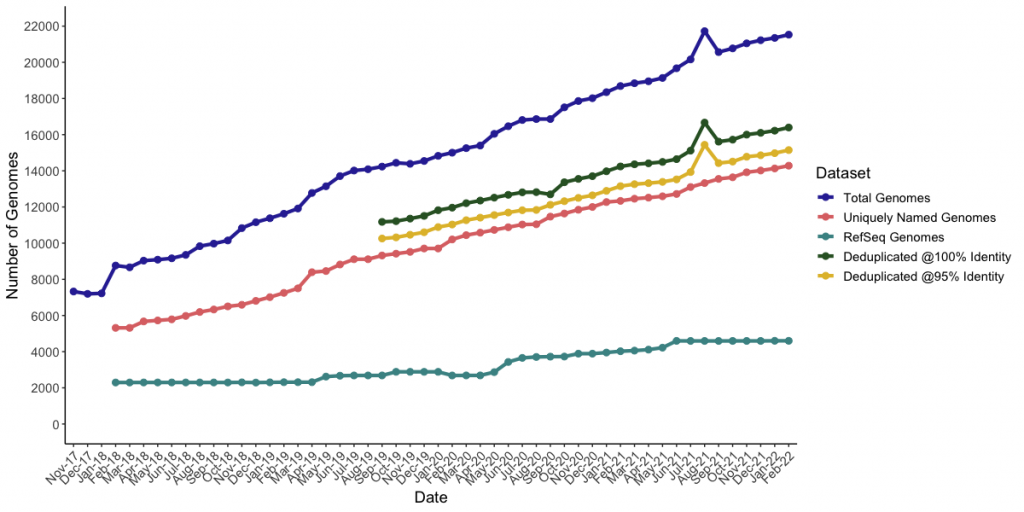
See our recent publication in PHAGE to read about how this dataset is produced and some of our analyses of it. Please consider citing this paper if you are using this database of information on this webpage. You can also generate an up-to-date version of the database, with useful files for vConTACT2, MASH, and IToL using our Perl script available on Github. Updates to the script this month include a new column to the tsv outputs which include anything identified as “host” or “lab_host” within the original Genbank files. However, these values may be inconsistent or downright bizarre (so please use them with caution).
We also recently added annotations using PHROGs (more details available here), and you can download the updated annotations from HERE (please note that we won’t be re-uploading the updated annotations on a monthly basis, as the file is huge. Having the first ~19,000 already annotated will save users a lot of time when using the Perl script themselves).
If you don’t want to run the script yourself, please download all of the files ready-made from below:
- 1Mar2022_data_excluding_refseq.tsv
- 1Mar2022_data.tsv
- 1Mar2022_genomes.db
- 1Mar2022_genomes_excluding_refseq.fa
- 1Mar2022_genomes.fa
- 1Mar2022_genomes.fa.msh
- 1Mar2022_itol_family_annotations.txt
- 1Mar2022_itol_genus_annotations.txt
- 1Mar2022_itol_host_annotations.txt
- 1Mar2022_itol_length_annotations.txt
- 1Mar2022_itol_lowest_taxa_annotations.txt
- 1Mar2022_itol_node_label_annotations.txt
- 1Mar2022_itol_subfamily_annotations.txt
- 1Mar2022_phages_downloaded_from_genbank.gb
- 1Mar2022_refseq_genomes.fa
- 1Mar2022_vConTACT2_family_annotations.tsv
- 1Mar2022_vConTACT2_gene_to_genome.csv
- 1Mar2022_vConTACT2_genus_annotations.tsv
- 1Mar2022_vConTACT2_host_annotations.tsv
- 1Mar2022_vConTACT2_lowest_taxa_annotations.tsv
- 1Mar2022_vConTACT2_proteins.faa
- 1Mar2022_vConTACT2_subfamily_annotations.tsv
- PHROGs HMMs for consistent annotation of genomes (see the new –PHROG optional flag)
- GenomesDB Directory (please note that this doesn’t get updated each month, it’s just here as a time-saver if you run the script yourself. This version is for 13/Dec/2021)
| Accession | Description | Classification | Genome Length (bp) | molGC (%) | Genus | Sub-family | Family | Host |
|---|---|---|---|---|---|---|---|---|
| LR756511 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 595163 | 42.11 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756508 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 484177 | 39.553 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756504 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 636363 | 26.393 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756503 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 735411 | 32.225 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756502 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 642428 | 31.472 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756501 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 735411 | 32.225 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR756500 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 462018 | 24.326 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR745208 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 485034 | 55.114 | Unclassified | Unclassified | Unclassified | Unspecified |
| LR745206 | uncultured phage | uncultured phage environmental samples unclassified bacterial viruses Viruses | 634780 | 29.181 | Unclassified | Unclassified | Unclassified | Unspecified |
| MK250029 | Prevotella phage Lak-C1 | Prevotella phage Lak-C1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 540217 | 25.796 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250028 | Prevotella phage Lak-B9 | Prevotella phage Lak-B9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 550053 | 26.012 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250027 | Prevotella phage Lak-B8 | Prevotella phage Lak-B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 551627 | 26.022 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250026 | Prevotella phage Lak-B7 | Prevotella phage Lak-B7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 550702 | 26.02 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250025 | Prevotella phage Lak-B6 | Prevotella phage Lak-B6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 546689 | 26.029 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250024 | Prevotella phage Lak-B5 | Prevotella phage Lak-B5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 543529 | 26.053 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250023 | Prevotella phage Lak-B4 | Prevotella phage Lak-B4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 550552 | 26.019 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250022 | Prevotella phage Lak-B3 | Prevotella phage Lak-B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 546746 | 26.032 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250021 | Prevotella phage Lak-B2 | Prevotella phage Lak-B2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 549839 | 25.986 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250020 | Prevotella phage Lak-B1 | Prevotella phage Lak-B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 547991 | 26.031 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250019 | Prevotella phage Lak-A2 | Prevotella phage Lak-A2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 541299 | 25.968 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250018 | Prevotella phage Lak-A1 | Prevotella phage Lak-A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 541644 | 25.878 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250017 | Prevotella phage Lak-A1 | Prevotella phage Lak-A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 541724 | 25.88 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250016 | Prevotella phage Lak-A1 | Prevotella phage Lak-A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 541643 | 25.879 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK250015 | Prevotella phage Lak-A1 | Prevotella phage Lak-A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 541664 | 25.877 | Unclassified | Unclassified | Myoviridae | Prevotella |
| MK796797 | Shewanella phage X14 | Shewanella phage X14 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36396 | 44.104 | Unclassified | Unclassified | Podoviridae | Shewanella |
| OL770074 | Escherichia phage vB_EcoM-RPN187 | Escherichia phage vB_EcoM-RPN187 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167158 | 37.773 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| OL770073 | Escherichia phage vB_EcoM-RPN226 | Escherichia phage vB_EcoM-RPN226 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167390 | 35.552 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ326858 | Stenotrophomonas phage Pepon | Stenotrophomonas phage Pepon unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42532 | 60.039 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| NC_041920 | Escherichia phage phiEC1 | Escherichia phage phiEC1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170254 | 37.56 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_041919 | Escherichia phage HP3 | Escherichia phage HP3 Escherichia virus HP3 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168188 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JF912400 | Cronobacter phage ESP2949-1 | Cronobacter phage ESP2949-1 Kyungwonvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49116 | 50.092 | Kyungwonvirus | Unclassified | Drexlerviridae | Cronobacter |
| JF704107 | Mycobacterium phage George | Mycobacterium phage George Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51578 | 60.954 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704115 | Mycobacterium phage Yoshi | Mycobacterium phage Yoshi Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58714 | 61.023 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704110 | Mycobacterium phage PackMan | Mycobacterium phage PackMan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51339 | 62.576 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704108 | Mycobacterium phage JoeDirt | Mycobacterium phage JoeDirt Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74914 | 58.781 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704117 | Mycobacterium phage RockyHorror | Mycobacterium phage RockyHorror Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56719 | 61.128 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704101 | Mycobacterium phage Microwolf | Mycobacterium phage Microwolf Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50864 | 64.041 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704098 | Mycobacterium phage JHC117 | Mycobacterium phage JHC117 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 63.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704097 | Mycobacterium phage Gladiator | Mycobacterium phage Gladiator Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52213 | 61.427 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| HM246724 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24608 | 27.329 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| JF744988 | Mycobacterium phage Faith1 | Mycobacterium phage Faith1 Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75960 | 58.918 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| HM246723 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14701 | 26.427 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| HM246722 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27985 | 28.426 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| HM246720 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53237 | 27.265 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| HM755814 | Mycobacterium phage Wonder | Mycobacterium phage Wonder Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48491 | 63.95 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| HM624080 | Pseudomonas phage PA1/KOR/2010 | Pseudomonas phage PA1/KOR/2010 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34553 | 64.767 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ326859 | Stenotrophomonas phage Siara | Stenotrophomonas phage Siara Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61427 | 56.5 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| OM339528 | Enterobacteria phage SV76 | Enterobacteria phage SV76 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163826 | 35.316 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| OM033136 | Aeromonas phage phiA047 | Aeromonas phage phiA047 unclassified bacterial viruses Viruses | 121865 | 35.446 | Unclassified | Unclassified | Unclassified | Aeromonas |
| OM033135 | Aeromonas phage phiA019 | Aeromonas phage phiA019 unclassified bacterial viruses Viruses | 115393 | 35.99 | Unclassified | Unclassified | Unclassified | Aeromonas |
| OM033134 | Aeromonas phage phiA009 | Aeromonas phage phiA009 unclassified bacterial viruses Viruses | 115276 | 35.998 | Unclassified | Unclassified | Unclassified | Aeromonas |
| OM033133 | Aeromonas phage phiA005 | Aeromonas phage phiA005 unclassified bacterial viruses Viruses | 48663 | 49.448 | Unclassified | Unclassified | Unclassified | Aeromonas |
| OM475435 | Escherichia phage E26 | Escherichia phage E26 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48850 | 49.785 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| OM475433 | Escherichia phage 10W | Escherichia phage 10W unclassified bacterial viruses Viruses | 32279 | 51.888 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475434 | Escherichia phage 12W | Escherichia phage 12W unclassified bacterial viruses Viruses | 31347 | 51.903 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475432 | Escherichia phage 4W | Escherichia phage 4W unclassified bacterial viruses Viruses | 48634 | 49.84 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475431 | Escherichia phage 9W | Escherichia phage 9W unclassified bacterial viruses Viruses | 31622 | 52.106 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475430 | Escherichia phage 3W | Escherichia phage 3W unclassified bacterial viruses Viruses | 49277 | 49.776 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475429 | Escherichia phage 11W | Escherichia phage 11W unclassified bacterial viruses Viruses | 31564 | 51.384 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475428 | Escherichia phage 1W | Escherichia phage 1W unclassified bacterial viruses Viruses | 49662 | 50.016 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475427 | Escherichia phage D2 | Escherichia phage D2 unclassified bacterial viruses Viruses | 49197 | 50.007 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM475426 | Escherichia phage D8 | Escherichia phage D8 unclassified bacterial viruses Viruses | 31345 | 51.951 | Unclassified | Unclassified | Unclassified | Escherichia |
| OL964948 | Acinetobacter phage vB_AbaP_PE21 | Acinetobacter phage vB_AbaP_PE21 unclassified bacterial viruses Viruses | 41655 | 39.287 | Unclassified | Unclassified | Unclassified | Acinetobacter |
| OM030343 | Staphylococcus phage vB_SaS_GE1 | Staphylococcus phage vB_SaS_GE1 unclassified bacterial viruses Viruses | 138106 | 30.403 | Unclassified | Unclassified | Unclassified | Staphylococcus |
| OM287554 | Alteromonas phage vB_AmaS-R9Y1 | Alteromonas phage vB_AmaS-R9Y1 unclassified bacterial viruses Viruses | 40851 | 55.061 | Unclassified | Unclassified | Unclassified | Alteromonas |
| OM141125 | Pseudomonas phage PP9W2 | Pseudomonas phage PP9W2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54311 | 59.122 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| OM350013 | Bacillus phage MrBubbles | Bacillus phage MrBubbles Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161581 | 38.68 | Unclassified | Unclassified | Myoviridae | Bacillus |
| OM350012 | Bacillus phage Darren | Bacillus phage Darren Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160916 | 38.723 | Unclassified | Unclassified | Myoviridae | Bacillus |
| OM350011 | Bacillus phage BillyBob | Bacillus phage BillyBob Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165429 | 37.772 | Unclassified | Unclassified | Myoviridae | Bacillus |
| OM355495 | Escherichia phage ZH4 | Escherichia phage ZH4 unclassified bacterial viruses Viruses | 39496 | 49.711 | Unclassified | Unclassified | Unclassified | Escherichia |
| OM456492 | Salmonella phage SaP7 | Salmonella phage SaP7 unclassified bacterial viruses Viruses | 240613 | 48.418 | Unclassified | Unclassified | Unclassified | Salmonella |
| OL770276 | Salmonella phage vB_SalS-S10 | Salmonella phage vB_SalS-S10 unclassified bacterial viruses Viruses | 43324 | 49.481 | Unclassified | Unclassified | Unclassified | Salmonella |
| MW526260 | Pseudomonas phage PhL_UNISO_PA-DSM_ph0041x | Pseudomonas phage PhL_UNISO_PA-DSM_ph0041x unclassified bacterial viruses Viruses | 44477 | 52.108 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| MW512573 | Clostridioides phage phiCD418 | Clostridioides phage phiCD418 unclassified bacterial viruses Viruses | 53311 | 29.056 | Unclassified | Unclassified | Unclassified | Clostridioides |
| MW512572 | Clostridioides phage phiCD08011 | Clostridioides phage phiCD08011 unclassified bacterial viruses Viruses | 31394 | 29.805 | Unclassified | Unclassified | Unclassified | Clostridioides |
| MW512571 | Clostridioides phage CD2301 | Clostridioides phage CD2301 unclassified bacterial viruses Viruses | 38695 | 29.472 | Unclassified | Unclassified | Unclassified | Clostridioides |
| MW512570 | Clostridioides phage CD1801 | Clostridioides phage CD1801 unclassified bacterial viruses Viruses | 44363 | 28.781 | Unclassified | Unclassified | Unclassified | Clostridioides |
| NC_053005 | Enterobacter phage KNP7 | Enterobacter phage KNP7 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58058 | 56.48 | Chivirus | Unclassified | Siphoviridae | Enterobacter |
| NC_052992 | Salmonella phage KNP6 | Salmonella phage KNP6 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59230 | 55.756 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052980 | Proteus phage ASh-2020a | Proteus phage ASh-2020a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59078 | 46.72 | Unclassified | Unclassified | Siphoviridae | Proteus |
| HM152767 | Mycobacterium phage CrimD | Mycobacterium phage CrimD Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59798 | 66.877 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| HM035024 | Shigella phage Shfl1 | Shigella phage Shfl1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50661 | 45.408 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| GU459069 | Aeromonas phage 65 | Aeromonas phage 65 Ishigurovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235229 | 37.196 | Ishigurovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| HM152763 | Mycobacterium phage LeBron | Mycobacterium phage LeBron Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73453 | 58.835 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| GU477322 | Staphylococcus phage SAP-26 | Staphylococcus phage SAP-26 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41207 | 34.016 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| HM114315 | Acinetobacter phage 133 | Acinetobacter phage 133 Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159801 | 39.672 | Unclassified | Tevenvirinae | Myoviridae | Acinetobacter |
| GQ403789 | Thermus phage P23-77 | Thermus phage P23-77 Gammasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 17036 | 67.95 | Gammasphaerolipovirus | Unclassified | Sphaerolipoviridae | Thermus |
| FJ848881 | Lactococcus phage SL4 | Lactococcus phage SL4 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28144 | 34.967 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| GQ303265 | Mycobacterium phage Pumpkin | Mycobacterium phage Pumpkin Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74491 | 62.958 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| GQ303263 | Mycobacterium phage Peaches | Mycobacterium phage Peaches Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.872 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ236310 | Streptococcus phage Abc2 | Streptococcus phage Abc2 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34882 | 38.986 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| FJ226752 | Streptococcus phage ALQ13.2 | Streptococcus phage ALQ13.2 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35525 | 39.395 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| OL828291 | Enterobacter phage vB-EclM_KMB20 | Enterobacter phage vB-EclM_KMB20 unclassified bacterial viruses Viruses | 174428 | 39.718 | Unclassified | Unclassified | Unclassified | Enterobacter |
| OL828290 | Enterobacter phage vB-EclM_KMB19 | Enterobacter phage vB-EclM_KMB19 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172697 | 39.792 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| OM319461 | Vibrio phage C-ZP2022 | Vibrio phage C-ZP2022 unclassified bacterial viruses Viruses | 287943 | 43.352 | Unclassified | Unclassified | Unclassified | Vibrio |
| OK169616 | Salmonella phage SGPC | Salmonella phage SGPC unclassified bacterial viruses Viruses | 42345 | 50.058 | Unclassified | Unclassified | Unclassified | Salmonella |
| NC_008722 | Staphylococcus phage CNPH82 | Staphylococcus phage CNPH82 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43420 | 34.675 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ358387 | Shigella virus Moo19 | Shigella virus Moo19 Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72458 | 44.64 | Unclassified | Enquatrovirinae | Schitoviridae | Shigella |
| OV877511 | Escherichia phage UGJNEcP2 | Escherichia phage UGJNEcP2 unclassified bacterial viruses Viruses | 169542 | 37.621 | Unclassified | Unclassified | Unclassified | Escherichia |
| OV877387 | Klebsiella phage UGKSKpnP1 | Klebsiella phage UGKSKpnP1 unclassified bacterial viruses Viruses | 51620 | 51.59 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OV877339 | Klebsiella phage UGKSKpnP2 | Klebsiella phage UGKSKpnP2 unclassified bacterial viruses Viruses | 51622 | 51.585 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OV877085 | Escherichia phage UGKSEcP1 | Escherichia phage UGKSEcP1 unclassified bacterial viruses Viruses | 167433 | 35.513 | Unclassified | Unclassified | Unclassified | Escherichia |
| OV877766 | Klebsiella phage UGKSKpnP3 | Klebsiella phage UGKSKpnP3 unclassified bacterial viruses Viruses | 51622 | 51.575 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OV877765 | Escherichia phage UGJNEcP4 | Escherichia phage UGJNEcP4 unclassified bacterial viruses Viruses | 170590 | 39.504 | Unclassified | Unclassified | Unclassified | Escherichia |
| OV876907 | Escherichia phage UGJNEcP3 | Escherichia phage UGJNEcP3 unclassified bacterial viruses Viruses | 170590 | 39.504 | Unclassified | Unclassified | Unclassified | Escherichia |
| OV876900 | Escherichia phage UGKSEcP2 | Escherichia phage UGKSEcP2 unclassified bacterial viruses Viruses | 167221 | 35.5 | Unclassified | Unclassified | Unclassified | Escherichia |
| OL800706 | Escherichia phage CLB_P3 | Escherichia phage CLB_P3 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49689 | 45.889 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MZ602148 | Klebsiella phage vB_KpnM_FRZ284 | Klebsiella phage vB_KpnM_FRZ284 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166376 | 39.635 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MZ602147 | Klebsiella phage vB_KpnM_VIK251 | Klebsiella phage vB_KpnM_VIK251 unclassified Mydovirus Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141994 | 44.615 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| MZ602146 | Klebsiella phage vB_KpnP_NER40 | Klebsiella phage vB_KpnP_NER40 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42674 | 54.272 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OM293948 | Levilactobacillus phage ENFP1 | Levilactobacillus phage ENFP1 unclassified Tybeckvirus Tybeckvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138676 | 32.484 | Tybeckvirus | Unclassified | Herelleviridae | Levilactobacillus |
| OL742560 | Arthrobacter phage Lilmac1015 | Arthrobacter phage Lilmac1015 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49978 | 69.433 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| OL964752 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43428 | 45.892 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| OL964751 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43528 | 45.913 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| OL964745 | Escherichia phage T4 | Escherichia phage T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159752 | 35.359 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL964735 | Escherichia phage T4 | Escherichia phage T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167548 | 35.281 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL964734 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43626 | 45.931 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| LC648442 | Staphylococcus phage vB_SsapH-Golestan-105-M | Staphylococcus phage vB_SsapH-Golestan-105-M unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142192 | 30.579 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| LC647030 | Staphylococcus phage vB_SsapH-Golestan-100 | Staphylococcus phage vB_SsapH-Golestan-100 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136433 | 33.748 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| LC557520 | Staphylococcus phage vB_SsapH-Golestan101-M | Staphylococcus phage vB_SsapH-Golestan101-M unclassified Baoshanvirus Baoshanvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144081 | 29.576 | Baoshanvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KM067278 | Pseudomonas phage vB_PaeP_PPA-ABTNL | Pseudomonas phage vB_PaeP_PPA-ABTNL Pseudomonas virus ABTNL Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43227 | 62.442 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| CP091911 | Enterococcus phage DPAraya-2022a | Enterococcus phage DPAraya-2022a Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28901 | 32.504 | Unclassified | Unclassified | Podoviridae | Enterococcus |
| OM291373 | Salmonella phage PBSE191 | Salmonella phage PBSE191 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41332 | 49.903 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OL976437 | Klebsiella phage Kpn BHU3 | Klebsiella phage Kpn BHU3 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43437 | 54.065 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OM249958 | Bacillus phage TBA3 | Bacillus phage TBA3 unclassified Salasvirus Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19105 | 39.901 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| OM317910 | Klebsiella phage P-K7R | Klebsiella phage P-K7R unclassified Drexlerviridae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52111 | 49.126 | Unclassified | Unclassified | Drexlerviridae | Klebsiella |
| OM022251 | Escherichia phage HZ41 | Escherichia phage HZ41 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38952 | 47.184 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| OM339718 | Aeromonas phage P19 | Aeromonas phage P19 unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 228236 | 38.799 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| OM025231 | Salmonella phage STM2 BHU | Salmonella phage STM2 BHU unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58828 | 54.426 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| OM421595 | Pseudomonas phage vB_PaeM_VL12 | Pseudomonas phage vB_PaeM_VL12 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88451 | 54.753 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| OM304382 | Burkholderia phage PhiBP82.2 | Burkholderia phage PhiBP82.2 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36280 | 64.766 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| OM401812 | Escherichia phage vB_EcoM_PD211 | Escherichia phage vB_EcoM_PD211 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149506 | 37.652 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| OM234793 | Pseudomonas phage PA28 | Pseudomonas phage PA28 unclassified Novosibovirus Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42910 | 41.384 | Novosibovirus | Slopekvirinae | Autographiviridae | Pseudomonas |
| OM234792 | Pseudomonas phage PA15 | Pseudomonas phage PA15 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72330 | 54.864 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| OM234791 | Pseudomonas phage PA02 | Pseudomonas phage PA02 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42880 | 53.862 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| OM439675 | Staphylococcus phage vB_SauS_Mh4 | Staphylococcus phage vB_SauS_Mh4 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44199 | 34.648 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439674 | Staphylococcus phage vB_SauS_Mh15 | Staphylococcus phage vB_SauS_Mh15 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43090 | 35.259 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439673 | Staphylococcus phage vB_SauS_Mh1 | Staphylococcus phage vB_SauS_Mh1 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43800 | 35.091 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439672 | Staphylococcus phage vB_SauS_I73 | Staphylococcus phage vB_SauS_I73 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43575 | 35.094 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439671 | Staphylococcus phage vB_SauS_832 | Staphylococcus phage vB_SauS_832 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43499 | 34.07 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439670 | Staphylococcus phage vB_SauS_775 | Staphylococcus phage vB_SauS_775 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43321 | 33.91 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439669 | Staphylococcus phage vB_SauS_760 | Staphylococcus phage vB_SauS_760 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42788 | 35.351 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439668 | Staphylococcus phage vB_SauS_713 | Staphylococcus phage vB_SauS_713 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42961 | 34.257 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439667 | Staphylococcus phage vB_SauS_690 | Staphylococcus phage vB_SauS_690 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45771 | 33.274 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| OM439666 | Staphylococcus phage vB_SauS_321c | Staphylococcus phage vB_SauS_321c unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42397 | 34.08 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439665 | Staphylococcus phage vB_SauS_320 | Staphylococcus phage vB_SauS_320 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45809 | 33.544 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| OM439664 | Staphylococcus phage vB_SauS_308 | Staphylococcus phage vB_SauS_308 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39968 | 34.112 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| OM439663 | Staphylococcus phage vB_SauS_287 | Staphylococcus phage vB_SauS_287 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43134 | 34.745 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM439662 | Staphylococcus phage vB_SauS_277g | Staphylococcus phage vB_SauS_277g unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44016 | 34.063 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| OM373202 | Cylindrospermopsis phage Cr-LKS3 | Cylindrospermopsis phage Cr-LKS3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46251 | 66.275 | Unclassified | Unclassified | Siphoviridae | Cylindrospermopsis |
| OL870612 | Enterococcus phage UTI-EfS7 | Enterococcus phage UTI-EfS7 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56144 | 39.935 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| OL870611 | Enterococcus phage UTI-EfS3 | Enterococcus phage UTI-EfS3 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150393 | 37.098 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| OM362897 | Escherichia phage SKA64 | Escherichia phage SKA64 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152401 | 39.245 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| OM334891 | Acinetobacter phage 4316 | Acinetobacter phage 4316 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41332 | 41.06 | Unclassified | Unclassified | Podoviridae | Acinetobacter |
| OM032871 | Klebsiella phage vB_Kpn-VAC110 | Klebsiella phage vB_Kpn-VAC110 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45195 | 50.437 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OM140653 | Yersinia phage vB_YenM_31.17 | Yersinia phage vB_YenM_31.17 unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32949 | 48.973 | Unclassified | Peduovirinae | Myoviridae | Yersinia |
| OM095401 | Yersinia phage vB_YenM_324 | Yersinia phage vB_YenM_324 unclassified Seongnamvirus Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29180 | 49.575 | Seongnamvirus | Peduovirinae | Myoviridae | Yersinia |
| OM112210 | Bacillus phage PK2 | Bacillus phage PK2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59399 | 42.513 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OM112209 | Bacillus phage PK1 | Bacillus phage PK1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41431 | 39.393 | Unclassified | Unclassified | Podoviridae | Bacillus |
| OM135608 | Serratia phage vB_SmaS_Bonzee | Serratia phage vB_SmaS_Bonzee Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42045 | 46.108 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OM284015 | Elizabethkingia phage EKP1 | Elizabethkingia phage EKP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38890 | 51.342 | Unclassified | Unclassified | Myoviridae | Elizabethkingia |
| OL944604 | Erwinia phage Gungnir39 | Erwinia phage Gungnir39 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48577 | 48.027 | Unclassified | Unclassified | Siphoviridae | Erwinia |
| OM158235 | Stenotrophomonas phage C121 | Stenotrophomonas phage C121 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73045 | 49.776 | Unclassified | Unclassified | Schitoviridae | Stenotrophomonas |
| OM131411 | Pseudomonas phage Pf17397_F_PD1 | Pseudomonas phage Pf17397_F_PD1 Pijolavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40763 | 56.392 | Pijolavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MW596414 | Bacillus phage BCP18 | Bacillus phage BCP18 unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159145 | 38.766 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| CP050061 | Lactococcus phage Bacchus | Lactococcus phage Bacchus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59331 | 35.368 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| OL964753 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122696 | 44.651 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964750 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37455 | 48.394 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964749 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37063 | 48.434 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964748 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37088 | 48.46 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964747 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122191 | 44.685 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964746 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123974 | 44.702 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964744 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41854 | 46.082 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| OL964743 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42512 | 46.067 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| OL964742 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41794 | 46.081 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| OL964741 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37534 | 48.511 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964740 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38646 | 48.45 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964739 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38846 | 48.471 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| OL964738 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139061 | 44.578 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964737 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143581 | 44.537 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964736 | Salmonella phage selz | Salmonella phage selz unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134470 | 44.661 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| OL964756 | Aeromonas phage pAEv1810 | Aeromonas phage pAEv1810 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235066 | 38.433 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| OL964755 | Aeromonas phage pAEv1818 | Aeromonas phage pAEv1818 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40808 | 55.239 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| OL964754 | Aeromonas phage pAEv1812 | Aeromonas phage pAEv1812 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61461 | 61.413 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MZ375347 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37892 | 46.345 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375232 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45807 | 45.927 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375358 | Salmonella phage SenALZ1 | Salmonella phage SenALZ1 Salmonella virus SenALZ1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118558 | 44.63 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MZ375357 | Salmonella phage SenALZ1 | Salmonella phage SenALZ1 Salmonella virus SenALZ1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121813 | 44.63 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MZ375356 | Escherichia phage T4 | Escherichia phage T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151790 | 35.49 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ375355 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36655 | 46.335 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375354 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36991 | 46.319 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375353 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37788 | 46.356 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375352 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35993 | 46.409 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375351 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36536 | 46.412 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375350 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37704 | 46.335 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375349 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36975 | 46.407 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375348 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37845 | 46.342 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375346 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37571 | 46.376 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375345 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37385 | 46.321 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375344 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38365 | 46.323 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375343 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38024 | 46.344 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375342 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37980 | 46.314 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375341 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37969 | 46.319 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375340 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37966 | 46.328 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375339 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38108 | 46.329 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375338 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37730 | 46.308 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375337 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38358 | 46.34 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375336 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40965 | 46.32 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375335 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38033 | 46.302 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375334 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39171 | 46.182 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375333 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38195 | 46.315 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375332 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39898 | 46.16 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375331 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40874 | 46.232 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375330 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37889 | 46.235 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375329 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38968 | 46.194 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375328 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38112 | 46.214 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375327 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39511 | 46.174 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375326 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38984 | 46.204 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375325 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37727 | 46.259 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375324 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40462 | 46.159 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375323 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40035 | 46.247 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375322 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39996 | 46.155 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375321 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39941 | 46.233 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375320 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45900 | 45.93 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375319 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40903 | 46.136 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375318 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40916 | 46.224 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375317 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40621 | 46.208 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375316 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40876 | 46.166 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375315 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40680 | 46.084 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375314 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42961 | 46.07 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375313 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45853 | 45.921 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375312 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41868 | 45.995 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375311 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41749 | 45.912 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375310 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42007 | 46.073 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375309 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42032 | 45.963 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375308 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42032 | 45.963 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375307 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42912 | 46.066 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375306 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43917 | 45.98 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375305 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43337 | 45.908 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375304 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44906 | 45.896 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375303 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44462 | 45.934 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375302 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45155 | 45.924 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375301 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43557 | 45.999 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375300 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44940 | 45.915 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375299 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44704 | 45.875 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375298 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45533 | 45.929 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375297 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45558 | 45.95 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375296 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45946 | 45.919 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375295 | Salmonella phage seszw | Salmonella phage seszw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45900 | 45.932 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ375294 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36575 | 48.577 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375293 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38241 | 48.469 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375292 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36844 | 48.589 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375291 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37262 | 48.527 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375290 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36858 | 48.529 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375289 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37254 | 48.534 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375288 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37388 | 48.513 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375287 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37125 | 48.544 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375286 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36542 | 48.561 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375285 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37137 | 48.534 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375284 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37337 | 48.507 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375283 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37228 | 48.504 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375282 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36619 | 48.57 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375281 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37125 | 48.531 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375280 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37320 | 48.499 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375279 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38030 | 48.446 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375278 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36783 | 48.552 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375277 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37330 | 48.489 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375276 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36532 | 48.557 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375275 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37757 | 48.494 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375274 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36786 | 48.546 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375273 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37757 | 48.489 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375272 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37478 | 48.506 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375271 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37825 | 48.481 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375270 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37751 | 48.468 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375269 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37730 | 48.481 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375268 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37210 | 48.498 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375267 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37743 | 48.481 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375266 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37732 | 48.489 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375265 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37690 | 48.485 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375264 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37892 | 48.498 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375263 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37712 | 48.478 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375262 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38168 | 48.48 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375261 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37533 | 48.501 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375260 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37759 | 48.489 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375259 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37825 | 48.489 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375258 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38167 | 48.448 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375257 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37825 | 48.486 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375256 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37111 | 48.476 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375255 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37752 | 48.487 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375254 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37682 | 48.445 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375253 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37861 | 48.504 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375252 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37779 | 48.514 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375251 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37861 | 48.501 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375250 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38163 | 48.461 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375249 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38166 | 48.452 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375248 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37861 | 48.432 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375247 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38050 | 48.439 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375246 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37681 | 48.454 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375245 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38697 | 48.456 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375244 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38718 | 48.414 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375243 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38728 | 48.427 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375242 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38916 | 48.425 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375241 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38926 | 48.451 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375240 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38831 | 48.405 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375239 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39105 | 48.434 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375238 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38112 | 48.434 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375237 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39218 | 48.434 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375236 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38457 | 48.428 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375235 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38457 | 48.423 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375234 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39938 | 48.4 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ375233 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39938 | 48.395 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN871446 | Klebsiella phage KpLz-2_45 | Klebsiella phage KpLz-2_45 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 296707 | 45.695 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| OL856139 | Rahnella phage Sarma103 | Rahnella phage Sarma103 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40678 | 49.378 | Unclassified | Studiervirinae | Autographiviridae | Rahnella |
| OL849997 | Enterobacter phage vB-EclM_KMB17 | Enterobacter phage vB-EclM_KMB17 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172607 | 39.843 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| OL825705 | Escherichia phage vB_EcoM_DE7 | Escherichia phage vB_EcoM_DE7 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86130 | 38.933 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| OL802211 | Pseudomonas phage vB_Pae_TR | Pseudomonas phage vB_Pae_TR unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43354 | 54.846 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| OL802210 | Pseudomonas phage vB_Pae_S1 | Pseudomonas phage vB_Pae_S1 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43058 | 62.223 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| OL791268 | Phage zth2-2 | Phage zth2-2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40470 | 53.729 | Unclassified | Unclassified | Podoviridae | Unspecified |
| OL791267 | Lentisphaerae phage WC36-1 | Lentisphaerae phage WC36-1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 16080 | 64.789 | Unclassified | Unclassified | Inoviridae | Lentisphaerae |
| OL791266 | Phage WC36-2 | Phage WC36-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28346 | 59.585 | Unclassified | Unclassified | Myoviridae | Unspecified |
| OL770365 | Aeromonas phage BUCT696 | Aeromonas phage BUCT696 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53372 | 51.13 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| OL754589 | Pseudomonas phage L5 | Pseudomonas phage L5 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42925 | 62.155 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| OL754588 | Pseudomonas phage PAP8 | Pseudomonas phage PAP8 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65466 | 55.569 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| OL762410 | Vibrio phage vB_ValP_FGH | Vibrio phage vB_ValP_FGH Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36470 | 48.544 | Unclassified | Unclassified | Podoviridae | Vibrio |
| OL742563 | Streptomyces phage Snorlax | Streptomyces phage Snorlax unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51592 | 65.652 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| OL742562 | Microbacterium phage Katzastrophic | Microbacterium phage Katzastrophic unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41712 | 67.851 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| OL742561 | Mycobacterium phage Duplicity | Mycobacterium phage Duplicity unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42938 | 66.265 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| OL742559 | Mycobacterium phage GoldenSpark | Mycobacterium phage GoldenSpark unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75132 | 62.955 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MW584228 | Bacillus phage BCPG3 | Bacillus phage BCPG3 unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159145 | 38.766 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| LR881104 | Klebsiella phage vB_KvM-Eowyn | Klebsiella phage vB_KvM-Eowyn Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265389 | 45.206 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| LR881103 | Escherichia phage vB_KppS-Ant | Escherichia phage vB_KppS-Ant unclassified Henuseptimavirus Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16548 | 49.958 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Escherichia |
| OV696614 | Escherichia phage Cyrano | Escherichia phage Cyrano Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108379 | 46.169 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| OV696612 | Escherichia phage Perceval | Escherichia phage Perceval Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50143 | 49.644 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| OV696611 | Escherichia phage Cartapus | Escherichia phage Cartapus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33368 | 50.971 | Unclassified | Unclassified | Myoviridae | Escherichia |
| OV696610 | Escherichia phage Tritos | Escherichia phage Tritos Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46360 | 48.296 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| OV696608 | Escherichia phage Gally | Escherichia phage Gally Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38682 | 47.043 | Unclassified | Unclassified | Podoviridae | Escherichia |
| OV754623 | Escherichia phage UGJNEcP1 | Escherichia phage UGJNEcP1 unclassified bacterial viruses Viruses | 169555 | 37.608 | Unclassified | Unclassified | Unclassified | Escherichia |
| MZ326866 | Stenotrophomonas phage Suso | Stenotrophomonas phage Suso Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44659 | 67.411 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ326862 | Stenotrophomonas phage Summit | Stenotrophomonas phage Summit unclassified Delepquintavirus Delepquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95728 | 58.473 | Delepquintavirus | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ326855 | Stenotrophomonas phage Suzuki | Stenotrophomonas phage Suzuki unclassified Sanovirus Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56042 | 62.607 | Sanovirus | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ326867 | Stenotrophomonas phage Silvanus | Stenotrophomonas phage Silvanus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45678 | 58.361 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ326865 | Enterococcus phage Sigurd | Enterococcus phage Sigurd unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41811 | 34.457 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MZ326861 | Stenotrophomonas phage Philippe | Stenotrophomonas phage Philippe Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74717 | 54.33 | Unclassified | Unclassified | Schitoviridae | Stenotrophomonas |
| MZ326860 | Stenotrophomonas phage Sonora | Stenotrophomonas phage Sonora Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63825 | 63.041 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ326856 | Stenotrophomonas phage Paxi | Stenotrophomonas phage Paxi unclassified Pokkenvirus Pokkenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74962 | 54.612 | Pokkenvirus | Unclassified | Schitoviridae | Stenotrophomonas |
| MZ326854 | Stenotrophomonas phage Ptah | Stenotrophomonas phage Ptah unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42593 | 59.928 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| LR991625 | Enterococcus phage dArtagnan | Enterococcus phage dArtagnan Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42884 | 35.335 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| LR990835 | Enterococcus phage Porthos | Enterococcus phage Porthos unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153193 | 37.106 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| LR990834 | Enterococcus phage Athos | Enterococcus phage Athos unclassified Picovirinae Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18345 | 33.824 | Unclassified | Picovirinae | Salasmaviridae | Enterococcus |
| LR990833 | Enterococcus phage Aramis | Enterococcus phage Aramis Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42697 | 35.091 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| LC680885 | Staphylococcus phage S6 | Staphylococcus phage S6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 267055 | 25.09 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| LC680884 | Bacillus phage PBS1 | Bacillus phage PBS1 Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252136 | 27.707 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| OK310514 | Escherichia phage ZCEC10 | Escherichia phage ZCEC10 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44776 | 54.686 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK310513 | Escherichia phage ZCEC12 | Escherichia phage ZCEC12 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44776 | 54.706 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK310512 | Escherichia phage ZCEC11 | Escherichia phage ZCEC11 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44776 | 54.701 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ361827 | Phage vB_LmoS_P7 | Phage vB_LmoS_P7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42553 | 36.646 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MZ361826 | Hydrogenophilus phage vB_LmoS_C996 | Hydrogenophilus phage vB_LmoS_C996 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40770 | 36.914 | Unclassified | Unclassified | Siphoviridae | Hydrogenophilus |
| MZ361825 | Listeria phage vB_LmoS_P9 | Listeria phage vB_LmoS_P9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36798 | 36.673 | Unclassified | Unclassified | Siphoviridae | Listeria |
| OL455900 | Arthrobacter phage BaileyBlu | Arthrobacter phage BaileyBlu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40447 | 65.837 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| OL455899 | Mycobacterium phage Vetrix | Mycobacterium phage Vetrix unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76520 | 58.939 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL455898 | Mycobacterium phage Abbyshoes | Mycobacterium phage Abbyshoes unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51324 | 63.744 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL455897 | Streptomyces phage Treat | Streptomyces phage Treat unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46094 | 59.596 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| OL455896 | Arthrobacter phage Reedo | Arthrobacter phage Reedo unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41823 | 65.304 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| OL455895 | Gordonia phage Jalebi | Gordonia phage Jalebi unclassified Lambovirus Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68501 | 58.364 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| OL455894 | Streptomyces phage Unstoppable | Streptomyces phage Unstoppable unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49556 | 66.184 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| OL455893 | Mycobacterium phage PenguinLover67 | Mycobacterium phage PenguinLover67 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70299 | 69.734 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| OL455892 | Microbacterium phage DesireeRose | Microbacterium phage DesireeRose unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 15488 | 54.997 | Unclassified | Unclassified | Tectiviridae | Microbacterium |
| OL455891 | Streptomyces phage SunsetPointe | Streptomyces phage SunsetPointe unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49957 | 66.307 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| OL455890 | Gordonia phage Wojtek | Gordonia phage Wojtek unclassified Lambovirus Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67754 | 58.196 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| OL455889 | Mycobacterium phage Bonamassa | Mycobacterium phage Bonamassa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51174 | 60.996 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL455888 | Gordonia phage GiKK | Gordonia phage GiKK Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47537 | 62.286 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| OL455887 | Gordonia phage Zany | Gordonia phage Zany unclassified Lambovirus Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68241 | 58.477 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| OL455886 | Streptomyces phage Piccadilly | Streptomyces phage Piccadilly Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37544 | 71.202 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| OK181862 | Escherichia phage 24-2-1 | Escherichia phage 24-2-1 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54117 | 39.592 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK181861 | Escherichia phage 24-2-1 | Escherichia phage 24-2-1 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79534 | 40.232 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK181860 | Escherichia phage 19-1-2 | Escherichia phage 19-1-2 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53997 | 39.591 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK181858 | Escherichia phage 19-1-2 | Escherichia phage 19-1-2 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77392 | 40.155 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OL979479 | Klebsiella phage Kpn BHU2 | Klebsiella phage Kpn BHU2 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42251 | 52.124 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OL979478 | Klebsiella phage Kpn BHU1 | Klebsiella phage Kpn BHU1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40410 | 52.833 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OL840290 | Staphylococcus phage Rih21 | Staphylococcus phage Rih21 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44789 | 33.258 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| OL799250 | Streptococcus phage MissF | Streptococcus phage MissF Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41035 | 41.048 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL739525 | Escherichia phage dw-ec | Escherichia phage dw-ec unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151825 | 39.069 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| OL699877 | Salmonella phage STM1 BHU | Salmonella phage STM1 BHU unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59328 | 53.968 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| OL702938 | Klebsiella phage pK8 | Klebsiella phage pK8 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49534 | 51.141 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OL743187 | Acinetobacter phage vB_AbM_WUPSU | Acinetobacter phage vB_AbM_WUPSU unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44241 | 37.23 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| OL741057 | Staphylococcus phage vB_Sau_P68 | Staphylococcus phage vB_Sau_P68 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139409 | 31.001 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| OL865411 | Klebsiella phage vB_1086 | Klebsiella phage vB_1086 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49473 | 50.476 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ456352 | Vibrio phage HNL01 | Vibrio phage HNL01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111792 | 38.047 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW478292 | Clostridium phage vB_CpeS-1181 | Clostridium phage vB_CpeS-1181 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48164 | 28.258 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MW478291 | Clostridium phage phiCp-D | Clostridium phage phiCp-D Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54735 | 30.098 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MW478290 | Clostridium phage phiCp-B | Clostridium phage phiCp-B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33985 | 28.083 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MW478289 | Clostridium phage phiCp-A | Clostridium phage phiCp-A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40619 | 28.221 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_028913 | Staphylococcus phage phinm2 | Staphylococcus phage phinm2 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43145 | 34.586 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028864 | Staphylococcus phage phinm4 | Staphylococcus phage phinm4 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43189 | 34.738 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_023739 | Mycobacterium phage BillKnuckles | Mycobacterium phage BillKnuckles Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51821 | 63.436 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023716 | Mycobacterium phage Alma | Mycobacterium phage Alma Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53177 | 62.461 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023592 | Haloarcula hispanica pleomorphic virus 2 | Haloarcula hispanica pleomorphic virus 2 Alphapleolipovirus HHPV2 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8176 | 53.694 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| NC_017090 | Halogeometricum pleomorphic virus 1 | Halogeometricum pleomorphic virus 1 Betapleolipovirus HGPV1 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9694 | 61.605 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halogeometricum |
| NC_017089 | Halorubrum pleomorphic virus 6 | Halorubrum pleomorphic virus 6 Alphapleolipovirus HRPV6 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8549 | 62.674 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_017088 | Halorubrum pleomorphic virus 3 | Halorubrum pleomorphic virus 3 Betapleolipovirus HRPV3 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8770 | 58.335 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_017087 | Halorubrum pleomorphic virus 2 | Halorubrum pleomorphic virus 2 Alphapleolipovirus HRPV2 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 10656 | 63.701 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_006882 | Prochlorococcus phage P-SSP7 | Prochlorococcus phage P-SSP7 Tiamatvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45176 | 38.824 | Tiamatvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_014458 | Mycobacterium phage Angelica | Mycobacterium phage Angelica Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59598 | 66.388 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_013936 | Mycobacterium phage Ardmore | Mycobacterium phage Ardmore Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52141 | 61.491 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| EU861005 | Staphylococcus phage phiSauS-IPLA35 | Staphylococcus phage phiSauS-IPLA35 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45344 | 33.246 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| FJ174693 | Mycobacterium phage Ramsey | Mycobacterium phage Ramsey Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58578 | 61.219 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ174692 | Mycobacterium phage Pacc40 | Mycobacterium phage Pacc40 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58554 | 61.263 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ174690 | Mycobacterium phage Fruitloop | Mycobacterium phage Fruitloop Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58471 | 61.773 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011054 | Mycobacterium phage Boomer | Mycobacterium phage Boomer Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58037 | 61.128 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| EU816591 | Mycobacterium phage Kostya | Mycobacterium phage Kostya Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75811 | 62.917 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| EU816588 | Mycobacterium phage Porky | Mycobacterium phage Porky Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76312 | 62.841 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| EU744251 | Mycobacterium phage Jasper | Mycobacterium phage Jasper Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50968 | 63.713 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU744250 | Mycobacterium phage Pukovnik | Mycobacterium phage Pukovnik Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52892 | 63.263 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU744249 | Mycobacterium phage Lockley | Mycobacterium phage Lockley Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51478 | 63.425 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU744248 | Mycobacterium phage KBG | Mycobacterium phage KBG Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53572 | 63.572 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU272037 | Pseudomonas phage MP38 | Pseudomonas phage MP38 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36885 | 64.514 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| EU272036 | Pseudomonas phage MP29 | Pseudomonas phage MP29 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36632 | 64.329 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| EU399241 | Halomonas phage phiHAP-1 | Halomonas phage phiHAP-1 Hapunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39245 | 58.978 | Hapunavirus | Unclassified | Myoviridae | Halomonas |
| DQ674738 | Aeromonas phage phiO18P | Aeromonas phage phiO18P Bielevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33985 | 61.745 | Bielevirus | Peduovirinae | Myoviridae | Aeromonas |
| EU100884 | Thermus phage P74-26 | Thermus phage P74-26 Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83319 | 57.759 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| EU100883 | Thermus phage P23-45 | Thermus phage P23-45 Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84201 | 57.762 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| EF529515 | Streptococcus phage 858 | Streptococcus phage 858 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35543 | 39.786 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| EF537008 | Phormidium phage Pf-WMP3 | Phormidium phage Pf-WMP3 Wumptrevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43249 | 46.48 | Wumptrevirus | Unclassified | Podoviridae | Phormidium |
| EF372997 | Synechococcus phage Syn5 | Synechococcus phage Syn5 Voetvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46214 | 54.97 | Voetvirus | Unclassified | Autographiviridae | Synechococcus |
| EF057797 | Vibrio phage VP882 | Vibrio phage VP882 Hapunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38197 | 56.952 | Hapunavirus | Unclassified | Myoviridae | Vibrio |
| DQ834250 | Staphylococcus phage PH15 | Staphylococcus phage PH15 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44041 | 34.908 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| DQ831957 | Staphylococcus phage CNPH82 | Staphylococcus phage CNPH82 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43420 | 34.675 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_008583 | Staphylococcus phage phinm1 | Staphylococcus phage phinm1 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43128 | 34.156 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ777876 | Xanthomonas phage Xop411 | Xanthomonas phage Xop411 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44520 | 51.903 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| DQ908929 | Staphylococcus phage 80 | Staphylococcus phage 80 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42140 | 35.565 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ875742 | Phormidium phage Pf-WMP4 | Phormidium phage Pf-WMP4 Wumpquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40938 | 51.803 | Wumpquatrovirus | Unclassified | Podoviridae | Phormidium |
| DQ873690 | Pseudomonas phage MP22 | Pseudomonas phage MP22 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36409 | 64.19 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| DQ631426 | Pseudomonas phage DMS3 | Pseudomonas phage DMS3 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36415 | 64.265 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| DQ535032 | Lactococcus phage KSY1 | Lactococcus phage KSY1 Chopinvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79232 | 35.111 | Chopinvirus | Unclassified | Podoviridae | Lactococcus |
| NC_007047 | Staphylococcus phage 187 | Staphylococcus phage 187 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39620 | 34.25 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_002371 | Salmonella phage P22 | Salmonella phage P22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41724 | 47.09 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_005135 | Aeromonas phage 44RR2.8t | Aeromonas phage 44RR2.8t Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173591 | 43.881 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| NC_003907 | Vibrio phage VpV262 | Vibrio phage VpV262 Vipivirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46012 | 49.52 | Vipivirus | Unclassified | Zobellviridae | Vibrio |
| OK143508 | Salmonella phage CKT1 | Salmonella phage CKT1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40923 | 49.825 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OL451946 | Salmonella phage BD13 | Salmonella phage BD13 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121769 | 40.045 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MZ286294 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4806 | 36.891 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL770309 | Erwinia phage Virsaitis27 | Erwinia phage Virsaitis27 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171269 | 38.796 | Winklervirus | Unclassified | Myoviridae | Erwinia |
| MZ611865 | Stenotrophomonas phage SM171 | Stenotrophomonas phage SM171 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44514 | 67.32 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MW526259 | Pseudomonas phage PhL_UNISO_PA-DSM_ph0034 | Pseudomonas phage PhL_UNISO_PA-DSM_ph0034 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106949 | 49.457 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW526258 | Pseudomonas phage PhL_UNISO_PA-DSM_ph0031 | Pseudomonas phage PhL_UNISO_PA-DSM_ph0031 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62522 | 55.459 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW760841 | Erwinia phage pEa_SNUABM_8 | Erwinia phage pEa_SNUABM_8 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 249194 | 51.362 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW349133 | Acinetobacter phage Liucustia | Acinetobacter phage Liucustia unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100533 | 37.111 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_060137 | Gordonia phage Syleon | Gordonia phage Syleon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75606 | 58.735 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060136 | Gordonia phage Kudefre | Gordonia phage Kudefre Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76731 | 58.619 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060135 | Gordonia Phage Sephiroth | Gordonia Phage Sephiroth Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76058 | 58.642 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060134 | Gordonia phage Octobien14 | Gordonia phage Octobien14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76358 | 58.733 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060133 | Gordonia phage Rabbitrun | Gordonia phage Rabbitrun Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76821 | 58.79 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060132 | Gordonia phage Trax | Gordonia phage Trax Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75943 | 58.706 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_060131 | Gordonia phage Neville | Gordonia phage Neville Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75813 | 58.752 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| OK570186 | Pantoea phage Nafs113 | Pantoea phage Nafs113 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75899 | 54.067 | Unclassified | Unclassified | Myoviridae | Pantoea |
| OK570185 | Pantoea phage Nufs112 | Pantoea phage Nufs112 unclassified Gajwadongvirus Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45951 | 47.701 | Gajwadongvirus | Unclassified | Autographiviridae | Pantoea |
| OK570184 | Pantoea phage Nifs112 | Pantoea phage Nifs112 unclassified Eracentumvirus Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46202 | 50.214 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Pantoea |
| OL964058 | Bacillus phage vB_BtM_BMBsp2 | Bacillus phage vB_BtM_BMBsp2 unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156575 | 39.923 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| OL774877 | Streptococcus phage MissG1 | Streptococcus phage MissG1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34071 | 40.774 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774867 | Streptococcus phage MismyG | Streptococcus phage MismyG Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42739 | 39.65 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| OL763420 | Acinetobacter phage PhaR5 | Acinetobacter phage PhaR5 unclassified Lazarusvirus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164549 | 36.742 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| OL396571 | Pantoea phage PdC23 | Pantoea phage PdC23 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44715 | 49.661 | Unclassified | Unclassified | Unclassified | Pantoea |
| MW557853 | Haloarcula virus Hardyhisp2 | Haloarcula virus Hardyhisp2 unclassified Pleolipoviridae Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16133 | 39.819 | Unclassified | Unclassified | Pleolipoviridae | Haloarcula |
| MW344767 | Haloarcula virus Harhisp1 | Haloarcula virus Harhisp1 unclassified Pleolipoviridae Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 19481 | 53.236 | Unclassified | Unclassified | Pleolipoviridae | Haloarcula |
| MW344766 | Haloferax virus Halfgib1 | Haloferax virus Halfgib1 unclassified Pleolipoviridae Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16280 | 56.486 | Unclassified | Unclassified | Pleolipoviridae | Haloferax |
| MW344764 | Halorubrum virus Humcor2 | Halorubrum virus Humcor2 unclassified Pleolipoviridae Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 11758 | 58.386 | Unclassified | Unclassified | Pleolipoviridae | Halorubrum |
| MT745956 | Bacillus phage z1a | Bacillus phage z1a unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39399 | 35.062 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| MT745954 | Bacillus phage F16Ba | Bacillus phage F16Ba unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38554 | 34.878 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| MT745955 | Bacillus phage J5a | Bacillus phage J5a unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40353 | 35.207 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| MG550113 | Halorubrum pleomorphic virus 11 | Halorubrum pleomorphic virus 11 Betapleolipovirus HRPV11 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9368 | 55.209 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| MG550111 | Halorubrum pleomorphic virus 10 | Halorubrum pleomorphic virus 10 Betapleolipovirus HRPV10 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9296 | 55.669 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| MG550110 | Halorubrum pleomorphic virus 12 | Halorubrum pleomorphic virus 12 Betapleolipovirus HRPV12 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9944 | 55.31 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| KY965934 | Halorubrum pleomorphic virus 9 | Halorubrum pleomorphic virus 9 Betapleolipovirus HRPV9 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16159 | 60.326 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| KY264020 | Haloarcula hispanica pleomorphic virus 4 | Haloarcula hispanica pleomorphic virus 4 Betapleolipovirus HHPV4 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 15010 | 54.63 | Betapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| KX344510 | Haloarcula hispanica pleomorphic virus 3 | Haloarcula hispanica pleomorphic virus 3 Betapleolipovirus HHPV3 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 11648 | 56.104 | Betapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| KF056323 | Haloarcula hispanica pleomorphic virus 2 | Haloarcula hispanica pleomorphic virus 2 Alphapleolipovirus HHPV2 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8176 | 53.694 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| JN882267 | Halogeometricum pleomorphic virus 1 | Halogeometricum pleomorphic virus 1 Betapleolipovirus HGPV1 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9694 | 61.605 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halogeometricum |
| JN882266 | Halorubrum pleomorphic virus 6 | Halorubrum pleomorphic virus 6 Alphapleolipovirus HRPV6 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8549 | 62.674 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| JN882265 | Halorubrum pleomorphic virus 3 | Halorubrum pleomorphic virus 3 Betapleolipovirus HRPV3 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8770 | 58.335 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| JN882264 | Halorubrum pleomorphic virus 2 | Halorubrum pleomorphic virus 2 Alphapleolipovirus HRPV2 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 10656 | 63.701 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| GU321093 | Haloarcula hispanica pleomorphic virus 1 | Haloarcula hispanica pleomorphic virus 1 Alphapleolipovirus HHPV1 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8082 | 55.778 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| FJ685651 | Halorubrum pleomorphic virus 1 | Halorubrum pleomorphic virus 1 Alphapleolipovirus HRPV1 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 7048 | 54.186 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| DQ398042 | Mycobacterium phage Halo | Mycobacterium phage Halo Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42289 | 66.722 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ530362 | Staphylococcus phage phinm4 | Staphylococcus phage phinm4 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43189 | 34.738 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ530360 | Staphylococcus phage phinm2 | Staphylococcus phage phinm2 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43145 | 34.586 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ530359 | Staphylococcus phage phinm1 | Staphylococcus phage phinm1 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43128 | 34.156 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ529280 | Aeromonas phage 25 | Aeromonas phage 25 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161475 | 41.04 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| DQ517338 | Staphylococcus phage 80alpha | Staphylococcus phage 80alpha Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43864 | 34.103 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| DQ398050 | Mycobacterium phage PMC | Mycobacterium phage PMC Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56692 | 61.418 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ398043 | Mycobacterium phage Che12 | Mycobacterium phage Che12 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52047 | 62.882 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AF191797 | His2 virus | His2 virus Gammapleolipovirus His2 Gammapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16067 | 40.368 | Gammapleolipovirus | Unclassified | Pleolipoviridae | Unspecified |
| DQ163916 | Pseudomonas phage M6 | Pseudomonas phage M6 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59446 | 64.464 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| DQ163914 | Pseudomonas phage 119X | Pseudomonas phage 119X Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43365 | 44.893 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| DQ114220 | Pasteurella phage F108 | Pasteurella phage F108 Irtavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30505 | 42.105 | Irtavirus | Peduovirinae | Myoviridae | Pasteurella |
| OL960030 | Acinetobacter phage vB_AbaM_P1 | Acinetobacter phage vB_AbaM_P1 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 105384 | 37.697 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| OM105886 | Gordonia phage Ligma | Gordonia phage Ligma unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61714 | 70.185 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| OM105885 | Mycobacterium phage Madiba | Mycobacterium phage Madiba unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56056 | 61.405 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| OM105884 | Mycobacterium phage Dussy | Mycobacterium phage Dussy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51344 | 63.737 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL800606 | Salmonella phage vB_SAg-RPN213 | Salmonella phage vB_SAg-RPN213 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86124 | 38.719 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| OL800604 | Salmonella phage vB_STy-RN29 | Salmonella phage vB_STy-RN29 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110591 | 39.595 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| OL800605 | Salmonella phage vB_SAg-RPN15 | Salmonella phage vB_SAg-RPN15 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38505 | 50.425 | Teetrevirus | Studiervirinae | Autographiviridae | Salmonella |
| OL800603 | Salmonella phage vB_STy-RN5i1 | Salmonella phage vB_STy-RN5i1 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39433 | 48.416 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| OM203162 | Mycobacterium phage Eyeball | Mycobacterium phage Eyeball unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52231 | 63.864 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OM203161 | Mycobacterium phage SororFago | Mycobacterium phage SororFago unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53516 | 62.116 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OM203160 | Gordonia phage MrBates | Gordonia phage MrBates unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45413 | 66.606 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| OM203159 | Mycobacterium phage HarryHoudini | Mycobacterium phage HarryHoudini unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52722 | 63.429 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OM203158 | Mycobacterium phage Zimmer | Mycobacterium phage Zimmer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53048 | 63.124 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OM203157 | Microbacterium phage AluminumJesus | Microbacterium phage AluminumJesus unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61926 | 66.835 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| OM203156 | Gordonia phage Niagara | Gordonia phage Niagara unclassified Demosthenesvirus Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75306 | 59.488 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| OL774876 | Streptococcus phage MissB2 | Streptococcus phage MissB2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12916 | 39.285 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774875 | Streptococcus phage MissE2 | Streptococcus phage MissE2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36665 | 39.864 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| OL774874 | Streptococcus phage MissE1 | Streptococcus phage MissE1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41671 | 41.41 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774873 | Streptococcus phage MissD | Streptococcus phage MissD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38511 | 38.584 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774872 | Streptococcus phage MissC | Streptococcus phage MissC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36804 | 41.145 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774871 | Streptococcus phage MissB1 | Streptococcus phage MissB1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36522 | 41.701 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774870 | Streptococcus phage OlisA3 | Streptococcus phage OlisA3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38516 | 38.537 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774869 | Streptococcus phage OlisA2 | Streptococcus phage OlisA2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30592 | 39.612 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774868 | Streptococcus phage OlisA1 | Streptococcus phage OlisA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30210 | 39.844 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| OL774866 | Streptococcus phage MissG2 | Streptococcus phage MissG2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21387 | 40.557 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MZ634351 | Klebsiella phage PWKp20 | Klebsiella phage PWKp20 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51939 | 51.73 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ634350 | Klebsiella phage PWKp19 | Klebsiella phage PWKp19 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51583 | 51.575 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ634349 | Klebsiella phage PWKp18 | Klebsiella phage PWKp18 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170712 | 40.852 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MZ634348 | Klebsiella phage PWKp17 | Klebsiella phage PWKp17 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51637 | 51.771 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ634347 | Klebsiella phage PWKp16 | Klebsiella phage PWKp16 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170706 | 40.759 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MZ634346 | Klebsiella phage PWKp15 | Klebsiella phage PWKp15 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50457 | 50.95 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ634345 | Klebsiella phage PWKp14 | Klebsiella phage PWKp14 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49365 | 51.028 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ634344 | Klebsiella phage PWKp9S | Klebsiella phage PWKp9S unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109179 | 45.13 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MZ634343 | Klebsiella phage PWKp9B | Klebsiella phage PWKp9B unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44881 | 53.501 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MZ634342 | Klebsiella phage PWKp7 | Klebsiella phage PWKp7 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44892 | 53.542 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MZ634341 | Klebsiella phage PWKp5 | Klebsiella phage PWKp5 unclassified Taipeivirus Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157530 | 46.416 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MZ634340 | Klebsiella phage PWKp3 | Klebsiella phage PWKp3 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44626 | 53.464 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MZ634339 | Klebsiella phage PWKp2 | Klebsiella phage PWKp2 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44612 | 53.557 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MZ634338 | Klebsiella phage PWKp1 | Klebsiella phage PWKp1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44602 | 53.538 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| CAKLQH020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 328530 | 39.357 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000002 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 254532 | 39.191 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000003 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240243 | 39.391 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000004 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 232245 | 38.362 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000005 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219710 | 40.09 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000006 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175221 | 38.45 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000007 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133721 | 39.74 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000008 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133321 | 39.508 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000009 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127735 | 39.457 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000010 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115414 | 40.081 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000011 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107676 | 37.463 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000012 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 105080 | 38.15 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000013 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94258 | 39.969 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000014 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92187 | 39.138 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000015 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91601 | 33.232 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000016 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89985 | 38.214 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000017 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82114 | 40.225 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000018 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74593 | 37.394 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000019 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69182 | 34.542 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000020 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64448 | 36.982 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000021 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57626 | 38.542 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000022 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57482 | 38.762 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000023 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57154 | 40.48 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000024 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56806 | 38.183 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000025 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50320 | 39.829 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000026 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50292 | 38.503 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000027 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50196 | 39.035 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000028 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49140 | 37.527 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000029 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48729 | 37.819 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000030 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45871 | 40.126 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000031 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44378 | 37.163 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000032 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43206 | 38.212 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000033 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42402 | 37.852 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000034 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39898 | 39.35 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000035 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39068 | 39.687 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000036 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37765 | 38.504 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000037 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35863 | 38.65 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000038 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34509 | 39.726 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000039 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33015 | 36.708 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000040 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28937 | 38.66 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000041 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28158 | 37.236 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000042 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25096 | 38.835 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000043 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24440 | 38.727 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000044 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24187 | 40.1 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000045 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22967 | 42.97 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000046 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21002 | 39.572 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000047 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20467 | 41.76 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000048 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18663 | 35.064 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000049 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17981 | 36.522 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000050 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17562 | 37.547 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000051 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17145 | 35.532 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000052 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16216 | 39.103 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000053 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15069 | 39.133 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000054 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14122 | 40.887 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000055 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13355 | 35.387 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000056 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13055 | 36.982 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000057 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12626 | 37.866 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000058 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12515 | 38.506 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000059 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12173 | 37.476 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000060 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11959 | 39.443 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000061 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11891 | 39.088 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000062 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11844 | 42.131 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQH020000063 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10014 | 39.754 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 378590 | 39.131 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000002 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 305545 | 38.638 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000003 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 254538 | 39.192 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000004 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240332 | 39.386 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000005 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219934 | 40.091 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000006 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 207647 | 38.131 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000007 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174261 | 38.464 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000008 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155736 | 38.658 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000009 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153613 | 37.252 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000010 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144066 | 37.39 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000011 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139490 | 39.771 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000012 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133390 | 39.513 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000013 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127828 | 39.463 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000014 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115472 | 40.076 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000015 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112182 | 38.208 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000016 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94268 | 39.969 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000017 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92259 | 39.129 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000018 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89980 | 38.216 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000019 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86531 | 38.597 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000020 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82134 | 40.225 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000021 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71601 | 39.593 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000022 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69750 | 39.702 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000023 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69202 | 34.545 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000024 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64471 | 36.981 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000025 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57492 | 38.758 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000026 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57183 | 40.481 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000027 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54152 | 37.45 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000028 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51762 | 39.465 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000029 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50641 | 38.437 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000030 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48771 | 37.811 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000031 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39146 | 39.672 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000032 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35987 | 38.683 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000033 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26725 | 39.394 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000034 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23215 | 39.418 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000035 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23017 | 42.964 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000036 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17248 | 39.57 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000037 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17181 | 35.568 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000038 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12142 | 37.449 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQF020000039 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10017 | 36.208 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQC020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165462 | 36.663 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQD020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165297 | 36.652 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQA020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50310 | 38.605 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQA020000002 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47665 | 37.73 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQA020000003 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37468 | 40.467 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQE020000001 | Acinetobacter phage MD-2021a | Acinetobacter phage MD-2021a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91606 | 33.219 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| CAKLQB020000001 | Acinetobacter phage MD-2021b | Acinetobacter phage MD-2021b Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42541 | 39.334 | Unclassified | Unclassified | Podoviridae | Acinetobacter |
| CAKLQB020000002 | Acinetobacter phage MD-2021b | Acinetobacter phage MD-2021b Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15209 | 39.917 | Unclassified | Unclassified | Podoviridae | Acinetobacter |
| CAKLQG020000001 | Acinetobacter phage MD-2021b | Acinetobacter phage MD-2021b Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42573 | 39.422 | Unclassified | Unclassified | Podoviridae | Acinetobacter |
| OL551674 | Enterobacter phage vB_EcRAM-01 | Enterobacter phage vB_EcRAM-01 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178477 | 44.985 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| OK288022 | Salmonella phage vB_SenS_PYC03 | Salmonella phage vB_SenS_PYC03 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114770 | 40.261 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| OL622073 | Pseudomonas phage Pa BHU-17 | Pseudomonas phage Pa BHU-17 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91079 | 51.89 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| OL549192 | Arthrobacter phage Lizalica | Arthrobacter phage Lizalica unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43055 | 67.391 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| OL549191 | Arthrobacter phage Amyev | Arthrobacter phage Amyev unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44284 | 67.934 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| OL549190 | Mycobacterium phage Marge | Mycobacterium phage Marge unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51268 | 63.855 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL549189 | Arthrobacter phage Crewmate | Arthrobacter phage Crewmate unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44354 | 68.844 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| OL549188 | Arthrobacter phage Warda | Arthrobacter phage Warda unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43792 | 67.62 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| OL840900 | Aeromonas phage AhMtk13a | Aeromonas phage AhMtk13a unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163879 | 41.207 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| OL676770 | Vibrio virus VPMCC5 | Vibrio virus VPMCC5 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48938 | 40.741 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MZ443773 | Erwinia phage pEa_SNUABM_31 | Erwinia phage pEa_SNUABM_31 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265846 | 49.516 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| OL606627 | Klebsiella phage vB_KaeS_Phraden | Klebsiella phage vB_KaeS_Phraden unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59052 | 56.55 | Chivirus | Unclassified | Siphoviridae | Klebsiella |
| OL799327 | Aeromonas phage BUCT695 | Aeromonas phage BUCT695 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86289 | 42.249 | Unclassified | Unclassified | Podoviridae | Aeromonas |
| OL674249 | Klebsiella phage KpS4 | Klebsiella phage KpS4 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49674 | 50.161 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OL617037 | Flyfo microvirus Tbat2_171 | Flyfo microvirus Tbat2_171 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4275 | 60.561 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617036 | Flyfo microvirus Tbat2_163 | Flyfo microvirus Tbat2_163 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4325 | 44.301 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617030 | Flyfo microvirus Tbat2_144 | Flyfo microvirus Tbat2_144 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4580 | 55.437 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617042 | Flyfo siphovirus Tbat2_3 | Flyfo siphovirus Tbat2_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50667 | 48.483 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| OL617041 | Flyfo siphovirus Tbat1_6 | Flyfo siphovirus Tbat1_6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48082 | 47.735 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| OL741059 | Escherichia phage SKA49 | Escherichia phage SKA49 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51548 | 44.128 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MZ923504 | Microcystis phage MinS1 | Microcystis phage MinS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49966 | 71.671 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| OM135583 | Escherichia phage EP01 | Escherichia phage EP01 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165577 | 35.436 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL960582 | Escherichia phage vB_EcoS_SA12KD | Escherichia phage vB_EcoS_SA12KD unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50709 | 45.333 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| OL960581 | Escherichia phage vB_EcoM_SA20RB | Escherichia phage vB_EcoM_SA20RB unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166337 | 35.458 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL960580 | Escherichia phage vB_EcoM_SA21RB | Escherichia phage vB_EcoM_SA21RB unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167177 | 35.641 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL960579 | Escherichia phage vB_EcoS_SA30RD | Escherichia phage vB_EcoS_SA30RD unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50465 | 45.529 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| OL960578 | Escherichia phage vB_EcoS_SA32RD | Escherichia phage vB_EcoS_SA32RD unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48980 | 45.553 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| OL960577 | Escherichia phage vB_EcoM_SA35RD | Escherichia phage vB_EcoM_SA35RD unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169615 | 37.638 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| OL960576 | Escherichia phage vB_EcoM_SA79RD | Escherichia phage vB_EcoM_SA79RD unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169615 | 37.638 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| OL960575 | Escherichia phage vB_EcoS_SA80RD | Escherichia phage vB_EcoS_SA80RD unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59778 | 44.098 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| OL960574 | Escherichia phage vB_EcoM_SA91KD | Escherichia phage vB_EcoM_SA91KD unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53673 | 44.058 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| OL960573 | Escherichia phage vB_EcoS_SA126VB | Escherichia phage vB_EcoS_SA126VB unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61753 | 44.236 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ995503 | Pasteurella phage vB_PmuP_PS07 | Pasteurella phage vB_PmuP_PS07 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38038 | 40.772 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| OL770310 | Escherichia phage PSa2 | Escherichia phage PSa2 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110539 | 39.885 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| OK597184 | Lactobacillus phage LPhi1ADMT15 | Lactobacillus phage LPhi1ADMT15 unclassified Hopescreekvirus Hopescreekvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16494 | 39.602 | Hopescreekvirus | Unclassified | Herelleviridae | Lactobacillus |
| MZ436628 | Microcystis phage Mwe-Yong1112-1 | Microcystis phage Mwe-Yong1112-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39679 | 66.599 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| LC597490 | Bacillus phage vB_BceM_WH1 | Bacillus phage vB_BceM_WH1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229829 | 37.193 | Unclassified | Unclassified | Myoviridae | Bacillus |
| AF083977 | Escherichia phage Mu | Escherichia phage Mu Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36717 | 52.055 | Muvirus | Unclassified | Myoviridae | Escherichia |
| OL614104 | Morganella phage Mecenats66 | Morganella phage Mecenats66 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86193 | 49.065 | Unclassified | Unclassified | Myoviridae | Morganella |
| OL310488 | Escherichia phage EC101 | Escherichia phage EC101 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167718 | 35.398 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL774570 | Aeromonas phage phiA008 | Aeromonas phage phiA008 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65284 | 52.901 | Unclassified | Unclassified | Unclassified | Aeromonas |
| OK665835 | Escherichia phage EC100 | Escherichia phage EC100 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108723 | 39.014 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ995506 | Pasteurella phage vB_PmuP_PS30 | Pasteurella phage vB_PmuP_PS30 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38176 | 40.774 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| MZ995505 | Clostridium phage vB_CpeP_PMQ04 | Clostridium phage vB_CpeP_PMQ04 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38947 | 30.613 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MZ995504 | Riemerella phage vB_RanS_PT03 | Riemerella phage vB_RanS_PT03 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44001 | 34.813 | Unclassified | Unclassified | Siphoviridae | Riemerella |
| OL960029 | Pseudoxanthomonas phage PW916 | Pseudoxanthomonas phage PW916 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47760 | 62.634 | Unclassified | Unclassified | Siphoviridae | Pseudoxanthomonas |
| OK254198 | Escherichia phage PSD2001 | Escherichia phage PSD2001 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172175 | 40.287 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| OK254197 | Escherichia phage PNJ1902 | Escherichia phage PNJ1902 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107506 | 38.996 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| OL744112 | Bacillus phage Ademby | Bacillus phage Ademby unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24162 | 30.527 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| OL744111 | Bacillus phage Arbo1 | Bacillus phage Arbo1 unclassified Salasvirus Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19362 | 39.696 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| OL989991 | Enterobacter phage vB_EclM_Q7622 | Enterobacter phage vB_EclM_Q7622 unclassified Kanagawavirus Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173871 | 40.022 | Kanagawavirus | Unclassified | Myoviridae | Enterobacter |
| OL959999 | Edwardsiella phage vB_EpP_ZHX | Edwardsiella phage vB_EpP_ZHX unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41777 | 48.11 | Kafunavirus | Unclassified | Podoviridae | Edwardsiella |
| OL773677 | Salmonella phage PRF-SP5 | Salmonella phage PRF-SP5 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39926 | 50.281 | Kayfunavirus | Studiervirinae | Autographiviridae | Salmonella |
| OL773676 | Salmonella phage PRF-SP4 | Salmonella phage PRF-SP4 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39976 | 50.243 | Kayfunavirus | Studiervirinae | Autographiviridae | Salmonella |
| OL580764 | Bacillus phage vB_BsuS-Goe14 | Bacillus phage vB_BsuS-Goe14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125490 | 35.541 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OL955261 | Erythrobacter phage vB_EliS-L02 | Erythrobacter phage vB_EliS-L02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150063 | 59.428 | Unclassified | Unclassified | Siphoviridae | Erythrobacter |
| OL856099 | Shigella phage CWP003 | Shigella phage CWP003 unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68721 | 46.14 | Wifcevirus | Unclassified | Myoviridae | Shigella |
| OL674541 | Stenotrophomonas phage vB_SmaS-AXL_1 | Stenotrophomonas phage vB_SmaS-AXL_1 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63962 | 67.277 | Pamexvirus | Unclassified | Siphoviridae | Stenotrophomonas |
| OL963731 | Citromicrobium phage vB_CbaS-RXM | Citromicrobium phage vB_CbaS-RXM Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104206 | 61.641 | Unclassified | Unclassified | Siphoviridae | Citromicrobium |
| OL956808 | Escherichia phage vB_EcoM-S1P5QW | Escherichia phage vB_EcoM-S1P5QW unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166102 | 35.451 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ516527 | Hafnia phage yong2 | Hafnia phage yong2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39546 | 49.906 | Unclassified | Unclassified | Myoviridae | Hafnia |
| MZ892993 | Pelagibacter phage HTVC033P | Pelagibacter phage HTVC033P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53075 | 33.78 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MZ892992 | Roseobacter phage CRP-738 | Roseobacter phage CRP-738 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53826 | 45.64 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ892991 | Roseobacter phage CRP-603 | Roseobacter phage CRP-603 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54551 | 43.119 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ892990 | Roseobacter phage CRP-345 | Roseobacter phage CRP-345 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54718 | 42.154 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ892989 | Roseobacter phage CRP-235 | Roseobacter phage CRP-235 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52729 | 45.688 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ892988 | Roseobacter phage CRP-212 | Roseobacter phage CRP-212 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54748 | 48.581 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ892987 | Roseobacter phage CRP-207 | Roseobacter phage CRP-207 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54895 | 46.203 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MZ318367 | Klebsiella phage BUCT610 | Klebsiella phage BUCT610 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46774 | 47.986 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MW205203 | Enterobacter phage BUCT554 | Enterobacter phage BUCT554 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40280 | 53.339 | Unclassified | Unclassified | Podoviridae | Enterobacter |
| NC_048198 | Erwinia phage vB_EhrS_59 | Erwinia phage vB_EhrS_59 Erwinia virus Eho59 Feofaniavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47116 | 50.405 | Feofaniavirus | Unclassified | Siphoviridae | Erwinia |
| OL505085 | Enterococcus phage EFap05-1 | Enterococcus phage EFap05-1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56564 | 39.987 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| OL505084 | Enterococcus phage EFap02 | Enterococcus phage EFap02 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39766 | 34.874 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| OL617038 | Flyfo microvirus Tbat2_185 | Flyfo microvirus Tbat2_185 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3911 | 41.37 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617035 | Flyfo microvirus Tbat2_160 | Flyfo microvirus Tbat2_160 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4362 | 48.304 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617034 | Flyfo microvirus Tbat2_158 | Flyfo microvirus Tbat2_158 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4380 | 37.489 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617033 | Flyfo microvirus Tbat2_156 | Flyfo microvirus Tbat2_156 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 47.665 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617032 | Flyfo microvirus Tbat2_153 | Flyfo microvirus Tbat2_153 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4405 | 44.427 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617031 | Flyfo microvirus Tbat2_151 | Flyfo microvirus Tbat2_151 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4440 | 47.635 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617029 | Flyfo microvirus Tbat2_130 | Flyfo microvirus Tbat2_130 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4969 | 56.631 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617028 | Flyfo microvirus Tbat2_116 | Flyfo microvirus Tbat2_116 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5648 | 40.74 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617027 | Flyfo microvirus Tbat2_112 | Flyfo microvirus Tbat2_112 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5767 | 56.841 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617026 | Flyfo microvirus Tbat2_110 | Flyfo microvirus Tbat2_110 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5849 | 56.386 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617025 | Flyfo microvirus Tbat2_108 | Flyfo microvirus Tbat2_108 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5857 | 57.965 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617024 | Flyfo microvirus Tbat2_105 | Flyfo microvirus Tbat2_105 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5949 | 55.522 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617023 | Flyfo microvirus Tbat2_103 | Flyfo microvirus Tbat2_103 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5964 | 38.146 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617022 | Flyfo microvirus Tbat2_98 | Flyfo microvirus Tbat2_98 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6062 | 57.473 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617021 | Flyfo microvirus Tbat2_95 | Flyfo microvirus Tbat2_95 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6096 | 55.43 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617020 | Flyfo microvirus Tbat2_93 | Flyfo microvirus Tbat2_93 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6111 | 59.368 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617019 | Flyfo microvirus Tbat2_91 | Flyfo microvirus Tbat2_91 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6153 | 53.876 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617018 | Flyfo microvirus Tbat2_88 | Flyfo microvirus Tbat2_88 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6260 | 55.304 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617017 | Flyfo microvirus Tbat2_83 | Flyfo microvirus Tbat2_83 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6376 | 54.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617016 | Flyfo microvirus Tbat2_43 | Flyfo microvirus Tbat2_43 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6049 | 54.422 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617015 | Flyfo microvirus Tbat1_66 | Flyfo microvirus Tbat1_66 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4179 | 55.157 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617014 | Flyfo microvirus Tbat1_59 | Flyfo microvirus Tbat1_59 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4315 | 36.848 | Unclassified | Unclassified | Microviridae | Unspecified |
| OL617040 | Flyfo podovirus Tbat2_2 | Flyfo podovirus Tbat2_2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51802 | 41.749 | Unclassified | Unclassified | Podoviridae | Unspecified |
| OL617039 | Flyfo myovirus Tbat2_7 | Flyfo myovirus Tbat2_7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31988 | 35.551 | Unclassified | Unclassified | Myoviridae | Unspecified |
| OK499971 | Enterobacter phage vB_EclS_CobraSix | Enterobacter phage vB_EclS_CobraSix Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47816 | 52.704 | Unclassified | Unclassified | Siphoviridae | Enterobacter |
| MZ447863 | Microcystis phage Mea-Yong924-1 | Microcystis phage Mea-Yong924-1 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40325 | 48.317 | Unclassified | Unclassified | Unclassified | Microcystis |
| MK075002 | Staphylococcus phage phiSP38-1 | Staphylococcus phage phiSP38-1 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43851 | 36.02 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AY078382 | Pseudomonas phage PaP3 | Pseudomonas phage PaP3 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45503 | 52.227 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| AF547987 | Shigella phage Sf6 | Shigella phage Sf6 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39043 | 47.468 | Lederbergvirus | Unclassified | Podoviridae | Shigella |
| AY052766 | Salmonella phage ST64T | Salmonella phage ST64T Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40679 | 47.523 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AF424782 | Staphylococcus phage phi 12 | Staphylococcus phage phi 12 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44970 | 33.347 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| AF189021 | Roseobacter phage SIO1 | Roseobacter phage SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39898 | 46.16 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| AF246223 | Mycoplasma phage P1 | Mycoplasma phage P1 Delislevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11660 | 26.792 | Delislevirus | Unclassified | Podoviridae | Mycoplasma |
| OL656110 | Escherichia phage vB_EcoM-RPN242 | Escherichia phage vB_EcoM-RPN242 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154840 | 48.831 | Unclassified | Aglimvirinae | Ackermannviridae | Escherichia |
| MZ666938 | Desulfovibrio phage ProddE | Desulfovibrio phage ProddE unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42637 | 51.387 | Unclassified | Unclassified | Autographiviridae | Desulfovibrio |
| MH399771 | Mycobacterium phage ByChance | Mycobacterium phage ByChance unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53123 | 61.314 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_008206 | Mycobacterium phage Wildcat | Mycobacterium phage Wildcat Wildcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78296 | 56.852 | Wildcatvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_025438 | Mycobacterium phage Soto | Mycobacterium phage Soto Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67744 | 66.558 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_025433 | Mycobacterium phage Taj | Mycobacterium phage Taj Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58550 | 61.863 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| AY939843 | Prochlorococcus phage P-SSP7 | Prochlorococcus phage P-SSP7 Tiamatvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45176 | 38.824 | Tiamatvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_009993 | Mycobacterium phage Giles | Mycobacterium phage Giles Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.447 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_015584 | Mycobacterium phage Faith1 | Mycobacterium phage Faith1 Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75960 | 58.918 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_014901 | Mycobacterium phage Wee | Mycobacterium phage Wee Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59230 | 61.786 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_014461 | Mycobacterium phage LeBron | Mycobacterium phage LeBron Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73453 | 58.835 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_013694 | Mycobacterium phage Peaches | Mycobacterium phage Peaches Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.872 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_013650 | Mycobacterium phage ET08 | Mycobacterium phage ET08 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155445 | 64.603 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_012788 | Mycobacterium phage Angel | Mycobacterium phage Angel unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41441 | 66.69 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011267 | Mycobacterium phage Solon | Mycobacterium phage Solon Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49487 | 63.843 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011055 | Mycobacterium phage Porky | Mycobacterium phage Porky Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76312 | 62.841 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011057 | Mycobacterium phage Phaedrus | Mycobacterium phage Phaedrus unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68090 | 67.566 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_011056 | Mycobacterium phage Kostya | Mycobacterium phage Kostya Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75811 | 62.917 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ003638 | Listeria phage A511 | Listeria phage A511 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137619 | 35.929 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_011023 | Mycobacterium phage Pukovnik | Mycobacterium phage Pukovnik Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52892 | 63.263 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011021 | Mycobacterium phage Lockley | Mycobacterium phage Lockley Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51478 | 63.425 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011019 | Mycobacterium phage KBG | Mycobacterium phage KBG Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53572 | 63.572 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011020 | Mycobacterium phage Jasper | Mycobacterium phage Jasper Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50968 | 63.713 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011022 | Mycobacterium phage DD5 | Mycobacterium phage DD5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51621 | 63.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_009877 | Mycobacterium phage U2 | Mycobacterium phage U2 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 63.699 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ054536 | Lactococcus phage P008 | Lactococcus phage P008 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28538 | 34.68 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| DQ004855 | Listeria phage P100 | Listeria phage P100 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131384 | 36.039 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| AY954970 | Staphylococcus phage Twort | Staphylococcus phage Twort Twortvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130706 | 30.261 | Twortvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| AY954969 | Staphylococcus phage G1 | Staphylococcus phage G1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138715 | 30.388 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| AY954968 | Staphylococcus phage X2 | Staphylococcus phage X2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43440 | 35.958 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954967 | Staphylococcus phage 92 | Staphylococcus phage 92 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42431 | 35.722 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954966 | Staphylococcus phage 88 | Staphylococcus phage 88 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43231 | 35.47 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954965 | Staphylococcus phage 52A | Staphylococcus phage 52A Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41690 | 35.534 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954955 | Staphylococcus phage 42E | Staphylococcus phage 42E Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45861 | 33.73 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| AY954953 | Staphylococcus phage 85 | Staphylococcus phage 85 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44283 | 34.587 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954952 | Staphylococcus phage 53 | Staphylococcus phage 53 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43883 | 34.077 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954951 | Staphylococcus phage 69 | Staphylococcus phage 69 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42732 | 34.302 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954950 | Staphylococcus phage 187 | Staphylococcus phage 187 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39620 | 34.25 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY699705 | Streptococcus phage 2972 | Streptococcus phage 2972 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34704 | 40.151 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| AY625898 | Pseudomonas phage F116 | Pseudomonas phage F116 Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65195 | 63.172 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| AY394005 | Pseudomonas phage D3112 | Pseudomonas phage D3112 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37611 | 64.34 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| AY375531 | Aeromonas phage 44RR2.8t | Aeromonas phage 44RR2.8t Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173591 | 43.881 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| AY288927 | Salmonella phage SP6 | Salmonella phage SP6 Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43769 | 47.227 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| AY299121 | Xanthomonas phage Xp10 | Xanthomonas phage Xp10 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44373 | 52.038 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| AY129330 | Mycobacterium phage Che8 | Mycobacterium phage Che8 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59471 | 61.304 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| OL840901 | Aeromonas phage AhMtk13b | Aeromonas phage AhMtk13b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61446 | 62.069 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| OL690335 | Staphylococcus phage 2PHSA1 | Staphylococcus phage 2PHSA1 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59779 | 32.871 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| OK539913 | Citrobacter phage vB_CbrM_HP1 | Citrobacter phage vB_CbrM_HP1 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89335 | 38.901 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| OL451225 | Escherichia phage P896 | Escherichia phage P896 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170656 | 40.425 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| OL691920 | Klebsiella phage vB8388 | Klebsiella phage vB8388 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39750 | 50.657 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| OL615011 | Shigella phage vB_SboS_Gloob | Shigella phage vB_SboS_Gloob unclassified Agtrevirus Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158676 | 50.359 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| OL770278 | Escherichia phage vB_EcoS_Opt212 | Escherichia phage vB_EcoS_Opt212 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44488 | 54.412 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| AY129332 | Mycobacterium phage Bxz2 | Mycobacterium phage Bxz2 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50913 | 64.184 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AY129336 | Mycobacterium phage Che9d | Mycobacterium phage Che9d Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 60.901 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| AY135486 | Salmonella phage PSP3 | Salmonella phage PSP3 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30636 | 52.827 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| AY212251 | Streptococcus phage C1 | Streptococcus phage C1 Fischettivirus Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16687 | 34.626 | Fischettivirus | Unclassified | Rountreeviridae | Streptococcus |
| AY129338 | Mycobacterium phage Omega | Mycobacterium phage Omega Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110865 | 61.375 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| AY129335 | Mycobacterium phage Corndog | Mycobacterium phage Corndog Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69777 | 65.384 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| AY129333 | Mycobacterium phage Che9c | Mycobacterium phage Che9c Chenonavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57050 | 65.378 | Chenonavirus | Unclassified | Siphoviridae | Mycobacterium |
| AY129331 | Mycobacterium phage Cjw1 | Mycobacterium phage Cjw1 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75931 | 63.115 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| AY095314 | Vibrio phage VpV262 | Vibrio phage VpV262 Vipivirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46012 | 49.52 | Vipivirus | Unclassified | Zobellviridae | Vibrio |
| NC_001909 | Lactococcus phage bIL170 | Lactococcus phage bIL170 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31754 | 34.345 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN685191 | Ralstonia phage vRsoP-WR2 | Ralstonia phage vRsoP-WR2 unclassified Gyeongsanvirus Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41158 | 59.041 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MN685190 | Ralstonia phage vRsoP-WM2 | Ralstonia phage vRsoP-WM2 unclassified Gyeongsanvirus Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40688 | 59.096 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MN685189 | Ralstonia phage vRsoP-WF2 | Ralstonia phage vRsoP-WF2 unclassified Gyeongsanvirus Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40690 | 59.071 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MZ666130 | Enterococcus phage Bp29 | Enterococcus phage Bp29 unclassified bacterial viruses Viruses | 41014 | 34.988 | Unclassified | Unclassified | Unclassified | Enterococcus |
| MZ506873 | Escherichia phage KW1E_UTAR | Escherichia phage KW1E_UTAR unclassified bacterial viruses Viruses | 45161 | 55.183 | Unclassified | Unclassified | Unclassified | Escherichia |
| MW286254 | Staphylococcus phage TSP | Staphylococcus phage TSP unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17987 | 29.427 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MW217515 | Vibrio phage VpJYP1 | Vibrio phage VpJYP1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50342 | 41.439 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW201959 | Salmonella phage SLMP1 | Salmonella phage SLMP1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40781 | 49.739 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT630408 | Enterobacteria phage phiC600P9 | Enterobacteria phage phiC600P9 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164691 | 35.621 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MT366943 | Yersinia phage vB_YenM_646 | Yersinia phage vB_YenM_646 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.8 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366942 | Yersinia phage vB_YenM_636 | Yersinia phage vB_YenM_636 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.798 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366941 | Yersinia phage vB_YenM_574 | Yersinia phage vB_YenM_574 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.798 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366940 | Yersinia phage vB_YenM_534 | Yersinia phage vB_YenM_534 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50480 | 46.791 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366939 | Yersinia phage vB_YenM_531 | Yersinia phage vB_YenM_531 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50482 | 46.817 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366938 | Yersinia phage vB_YenM_526-2 | Yersinia phage vB_YenM_526-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.795 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366937 | Yersinia phage vB_YenM_526-1 | Yersinia phage vB_YenM_526-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.791 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366936 | Yersinia phage vB_YenM_126 | Yersinia phage vB_YenM_126 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.796 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366935 | Yersinia phage vB_YenM_39 | Yersinia phage vB_YenM_39 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.796 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366934 | Yersinia phage vB_YenM_27 | Yersinia phage vB_YenM_27 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.8 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366933 | Yersinia phage vB_YenM_25 | Yersinia phage vB_YenM_25 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.798 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366932 | Yersinia phage vB_YenM_22 | Yersinia phage vB_YenM_22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.802 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MT366931 | Yersinia phage vB_YenM_12 | Yersinia phage vB_YenM_12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50476 | 46.798 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MN812694 | Pectobacterium phage Bobbio | Pectobacterium phage Bobbio unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98539 | 48.681 | Otagovirus | Unclassified | Myoviridae | Pectobacterium |
| MN812693 | Pectobacterium phage Cornix | Pectobacterium phage Cornix unclassified Ningirsuvirus Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40416 | 50.544 | Ningirsuvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MN812692 | Pectobacterium phage Rambam | Pectobacterium phage Rambam Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43928 | 49.083 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MN812690 | Dickeya phage Irene | Dickeya phage Irene Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44368 | 51.715 | Kotilavirus | Corkvirinae | Autographiviridae | Dickeya |
| MN812689 | Dickeya phage Setonix | Dickeya phage Setonix Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41545 | 46.537 | Unclassified | Unclassified | Siphoviridae | Dickeya |
| MN812688 | Dickeya phage Okapia | Dickeya phage Okapia Suwonvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53337 | 44.939 | Suwonvirus | Cleopatravirinae | Chaseviridae | Dickeya |
| MN812686 | Dickeya phage Monomax | Dickeya phage Monomax unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38777 | 50.7 | Kayfunavirus | Studiervirinae | Autographiviridae | Dickeya |
| MN812685 | Pectobacterium phage Arno152 | Pectobacterium phage Arno152 unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73119 | 48.522 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MZ547449 | Acidovorax phage ACF1 | Acidovorax phage ACF1 unclassified bacterial viruses Viruses | 59377 | 64.493 | Unclassified | Unclassified | Unclassified | Acidovorax |
| LC667452 | Burkholderia phage FLC10 | Burkholderia phage FLC10 unclassified Kisquattuordecimvirus Kisquattuordecimvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32867 | 61.286 | Kisquattuordecimvirus | Peduovirinae | Myoviridae | Burkholderia |
| LC667451 | Burkholderia phage FLC9 | Burkholderia phage FLC9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 321833 | 55.966 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| LC667450 | Burkholderia phage FLC8 | Burkholderia phage FLC8 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225545 | 52.051 | Chiangmaivirus | Unclassified | Myoviridae | Burkholderia |
| OL519843 | Salmonella phage vB_SenS_TUMS_E19 | Salmonella phage vB_SenS_TUMS_E19 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42813 | 49.751 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OK490474 | Klebsiella phage Kp4887-3 | Klebsiella phage Kp4887-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40133 | 50.353 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490473 | Klebsiella phage Kp4887-2 | Klebsiella phage Kp4887-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35131 | 51.052 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490472 | Klebsiella phage Kp4887-1 | Klebsiella phage Kp4887-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 50.636 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490471 | Klebsiella phage Kp4886-3 | Klebsiella phage Kp4886-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40133 | 50.353 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490470 | Klebsiella phage Kp4886-2 | Klebsiella phage Kp4886-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35131 | 51.052 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490469 | Klebsiella phage Kp4886-1 | Klebsiella phage Kp4886-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 50.636 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490468 | Klebsiella phage Kp4885-3 | Klebsiella phage Kp4885-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40133 | 50.353 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490467 | Klebsiella phage Kp4885-2 | Klebsiella phage Kp4885-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35131 | 51.052 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490466 | Klebsiella phage Kp4885-1 | Klebsiella phage Kp4885-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 50.636 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490465 | Klebsiella phage Kp4884-3 | Klebsiella phage Kp4884-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40133 | 50.353 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490464 | Klebsiella phage Kp4884-2 | Klebsiella phage Kp4884-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35131 | 51.052 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490463 | Klebsiella phage Kp4884-1 | Klebsiella phage Kp4884-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 50.636 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490462 | Klebsiella phage Kp4882-3 | Klebsiella phage Kp4882-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40133 | 50.353 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490461 | Klebsiella phage Kp4882-2 | Klebsiella phage Kp4882-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35131 | 51.052 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490460 | Klebsiella phage Kp4882-1 | Klebsiella phage Kp4882-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 50.636 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490459 | Klebsiella phage Kp4880-3 | Klebsiella phage Kp4880-3 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490458 | Klebsiella phage Kp4880-2 | Klebsiella phage Kp4880-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490457 | Klebsiella phage Kp4880-1 | Klebsiella phage Kp4880-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490456 | Klebsiella phage Kp4879-3 | Klebsiella phage Kp4879-3 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490455 | Klebsiella phage Kp4879-2 | Klebsiella phage Kp4879-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490454 | Klebsiella phage Kp4879-1 | Klebsiella phage Kp4879-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490453 | Klebsiella phage Kp4878-3 | Klebsiella phage Kp4878-3 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490452 | Klebsiella phage Kp4878-2 | Klebsiella phage Kp4878-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490451 | Klebsiella phage Kp4878-1 | Klebsiella phage Kp4878-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490450 | Klebsiella phage Kp4877-4 | Klebsiella phage Kp4877-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490449 | Klebsiella phage Kp4877-2 | Klebsiella phage Kp4877-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490448 | Klebsiella phage Kp4877-1 | Klebsiella phage Kp4877-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490447 | Klebsiella phage Kp4876-4 | Klebsiella phage Kp4876-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490446 | Klebsiella phage Kp4876-2 | Klebsiella phage Kp4876-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490445 | Klebsiella phage Kp4876-1 | Klebsiella phage Kp4876-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490444 | Klebsiella phage Kp4875-4 | Klebsiella phage Kp4875-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490443 | Klebsiella phage Kp4875-2 | Klebsiella phage Kp4875-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490442 | Klebsiella phage Kp4875-1 | Klebsiella phage Kp4875-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490441 | Klebsiella phage Kp4874-4 | Klebsiella phage Kp4874-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490440 | Klebsiella phage Kp4874-2 | Klebsiella phage Kp4874-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490439 | Klebsiella phage Kp4874-1 | Klebsiella phage Kp4874-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490438 | Klebsiella phage Kp4873-4 | Klebsiella phage Kp4873-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490437 | Klebsiella phage Kp4873-2 | Klebsiella phage Kp4873-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490436 | Klebsiella phage Kp4873-1 | Klebsiella phage Kp4873-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490435 | Klebsiella phage Kp4872-3 | Klebsiella phage Kp4872-3 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490434 | Klebsiella phage Kp4872-2 | Klebsiella phage Kp4872-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490433 | Klebsiella phage Kp4872-1 | Klebsiella phage Kp4872-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490432 | Klebsiella phage Kp4871-3 | Klebsiella phage Kp4871-3 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.431 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490431 | Klebsiella phage Kp4871-2 | Klebsiella phage Kp4871-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.422 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490430 | Klebsiella phage Kp4871-1 | Klebsiella phage Kp4871-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 50.466 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490429 | Klebsiella phage Kp4870-2 | Klebsiella phage Kp4870-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.089 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490428 | Klebsiella phage Kp4870-1 | Klebsiella phage Kp4870-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490427 | Klebsiella phage Kp4869-2 | Klebsiella phage Kp4869-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.089 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490426 | Klebsiella phage Kp4869-1 | Klebsiella phage Kp4869-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490425 | Klebsiella phage Kp4868-2 | Klebsiella phage Kp4868-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.089 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490424 | Klebsiella phage Kp4868-1 | Klebsiella phage Kp4868-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490423 | Klebsiella phage Kp4867-2 | Klebsiella phage Kp4867-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.089 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490422 | Klebsiella phage Kp4867-1 | Klebsiella phage Kp4867-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490421 | Klebsiella phage Kp4866-6 | Klebsiella phage Kp4866-6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45500 | 49.765 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490420 | Klebsiella phage Kp4866-3 | Klebsiella phage Kp4866-3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33090 | 54.998 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| OK490419 | Klebsiella phage Kp4865-4 | Klebsiella phage Kp4865-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45913 | 50.241 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490418 | Klebsiella phage Kp4865-2 | Klebsiella phage Kp4865-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46667 | 48.285 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490417 | Klebsiella phage Kp4865-1 | Klebsiella phage Kp4865-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40882 | 51.981 | Unclassified | Unclassified | Podoviridae | Klebsiella |
| OK490416 | Klebsiella phage Kp4864-1 | Klebsiella phage Kp4864-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46765 | 51.515 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490415 | Klebsiella phage Kp4862-1 | Klebsiella phage Kp4862-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48629 | 52.129 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490414 | Klebsiella phage Kp4861-1 | Klebsiella phage Kp4861-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37041 | 49.91 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490413 | Klebsiella phage Kp4860-7 | Klebsiella phage Kp4860-7 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33394 | 50.425 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490412 | Klebsiella phage Kp4860-2 | Klebsiella phage Kp4860-2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35006 | 54.434 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490411 | Klebsiella phage Kp4857-1 | Klebsiella phage Kp4857-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490410 | Klebsiella phage Kp4856-1 | Klebsiella phage Kp4856-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48629 | 52.129 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490409 | Klebsiella phage Kp4852-3 | Klebsiella phage Kp4852-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35168 | 54.021 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| OK490408 | Klebsiella phage Kp4852-2 | Klebsiella phage Kp4852-2 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34109 | 50.984 | Eganvirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490407 | Klebsiella phage Kp4851-2 | Klebsiella phage Kp4851-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490406 | Klebsiella phage Kp4851-1 | Klebsiella phage Kp4851-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490405 | Klebsiella phage Kp4850-2 | Klebsiella phage Kp4850-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490404 | Klebsiella phage Kp4850-1 | Klebsiella phage Kp4850-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490403 | Klebsiella phage Kp4849-2 | Klebsiella phage Kp4849-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.089 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490402 | Klebsiella phage Kp4849-1 | Klebsiella phage Kp4849-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490401 | Klebsiella phage Kp4848-2 | Klebsiella phage Kp4848-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490400 | Klebsiella phage Kp4848-1 | Klebsiella phage Kp4848-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490399 | Klebsiella phage Kp4847-2 | Klebsiella phage Kp4847-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490398 | Klebsiella phage Kp4847-1 | Klebsiella phage Kp4847-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490397 | Klebsiella phage Kp4846-2 | Klebsiella phage Kp4846-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490396 | Klebsiella phage Kp4846-1 | Klebsiella phage Kp4846-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490395 | Klebsiella phage Kp4845-2 | Klebsiella phage Kp4845-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35561 | 50.086 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| OK490394 | Klebsiella phage Kp4845-1 | Klebsiella phage Kp4845-1 unclassified Reipivirus Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36687 | 50.209 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| MW392801 | Bacillus phage BCP6 | Bacillus phage BCP6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81528 | 33.894 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MW392803 | Bacillus phage BCPG1 | Bacillus phage BCPG1 unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158861 | 38.753 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MW392802 | Bacillus phage BCPST | Bacillus phage BCPST Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81528 | 33.893 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| LC528882 | Burkholderia phage FLC5 | Burkholderia phage FLC5 unclassified Kisquattuordecimvirus Kisquattuordecimvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32090 | 61.792 | Kisquattuordecimvirus | Peduovirinae | Myoviridae | Burkholderia |
| MT338525 | Staphylococcus phage SAM1 | Staphylococcus phage SAM1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147588 | 30.209 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT226657 | Staphylococcus phage SAM2 | Staphylococcus phage SAM2 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137617 | 30.422 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ573924 | Salmonella phage vB_SenAt-pSL2 | Salmonella phage vB_SenAt-pSL2 unclassified Zindervirus Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44304 | 47.388 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| OK338687 | Vibrio phage vB_VpP_WS1 | Vibrio phage vB_VpP_WS1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5564 | 41.804 | Unclassified | Unclassified | Microviridae | Vibrio |
| OK129343 | Erwinia phage Fifi451 | Erwinia phage Fifi451 unclassified Loessnervirus Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53219 | 43.99 | Loessnervirus | Cleopatravirinae | Chaseviridae | Erwinia |
| OK073976 | Erwinia phage Fifi44 | Erwinia phage Fifi44 unclassified Loessnervirus Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53559 | 43.952 | Loessnervirus | Cleopatravirinae | Chaseviridae | Erwinia |
| MZ773939 | Pseudomonas phage PP9W | Pseudomonas phage PP9W Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47985 | 61.744 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW147603 | Yersinia phage PYps49T | Yersinia phage PYps49T unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39459 | 48.772 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| MW147602 | Yersinia phage PYps16T | Yersinia phage PYps16T unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52855 | 43.812 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Yersinia |
| MW147601 | Yersinia phage PYps16N | Yersinia phage PYps16N unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52376 | 44.02 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Yersinia |
| MW147600 | Yersinia phage PYps4T | Yersinia phage PYps4T unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52855 | 43.816 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Yersinia |
| MW147599 | Yersinia phage PYps3T | Yersinia phage PYps3T unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52854 | 43.813 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Yersinia |
| MW147598 | Yersinia phage PYps23T | Yersinia phage PYps23T unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54869 | 43.874 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Yersinia |
| MT515757 | Yersinia phage PYps50T | Yersinia phage PYps50T unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39655 | 48.738 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| MT828553 | Yersinia phage PYps10T | Yersinia phage PYps10T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169493 | 35.404 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT828552 | Yersinia phage PYps5T | Yersinia phage PYps5T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169375 | 35.355 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT828551 | Yersinia phage PYPS2T | Yersinia phage PYPS2T Yersinia virus PYPS2T Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169486 | 35.405 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515756 | Yersinia phage PYps55T | Yersinia phage PYps55T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167710 | 35.353 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515755 | Yersinia phage PYps47T | Yersinia phage PYps47T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165778 | 35.458 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515754 | Yersinia phage PYps35T | Yersinia phage PYps35T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166974 | 35.397 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515753 | Yersinia phage PYps32T | Yersinia phage PYps32T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165733 | 35.457 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515752 | Yersinia phage PYps15T | Yersinia phage PYps15T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166925 | 35.36 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT515751 | Yersinia phage PYps11T | Yersinia phage PYps11T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169490 | 35.406 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MT526905 | Yersinia phage PYps14T | Yersinia phage PYps14T unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166935 | 35.362 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| OL539467 | Escherichia phage vB_EcoD_Sadiya | Escherichia phage vB_EcoD_Sadiya unclassified Rogunavirinae Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48303 | 44.683 | Unclassified | Rogunavirinae | Drexlerviridae | Escherichia |
| OL539447 | Escherichia phage vB_ EcoD_Phleasolo | Escherichia phage vB_ EcoD_Phleasolo unclassified Rogunavirinae Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48301 | 44.689 | Unclassified | Rogunavirinae | Drexlerviridae | Escherichia |
| OL539472 | Klebsiella phage vB_KpnD_Chell | Klebsiella phage vB_KpnD_Chell unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51779 | 51.577 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OL539466 | Citrobacter phage vB_CfrD_SorkZaugg | Citrobacter phage vB_CfrD_SorkZaugg unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50023 | 42.75 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OL539460 | Citrobacter phage vB_CfrD_FairDinkum | Citrobacter phage vB_CfrD_FairDinkum unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49111 | 50.677 | Webervirus | Unclassified | Drexlerviridae | Citrobacter |
| OL539457 | Klebsiella phage vB_KaeD_HazelMika | Klebsiella phage vB_KaeD_HazelMika Vilniusvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51552 | 49.573 | Vilniusvirus | Unclassified | Drexlerviridae | Klebsiella |
| OL539451 | Escherichia phage vB_EcoD_Opt-719 | Escherichia phage vB_EcoD_Opt-719 unclassified Rogunavirinae Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48302 | 44.686 | Unclassified | Rogunavirinae | Drexlerviridae | Escherichia |
| OL539450 | Klebsiella phage vB_KpnD_Opt-817 | Klebsiella phage vB_KpnD_Opt-817 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48988 | 51.484 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OL539446 | Escherichia phage vB_EcoD_Phunderstruck | Escherichia phage vB_EcoD_Phunderstruck unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51274 | 44.635 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OL539445 | Escherichia phage vB_EcoD_Poky | Escherichia phage vB_EcoD_Poky unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 44.941 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OL539440 | Klebsiella phage vB_KaeS_Diencephalon | Klebsiella phage vB_KaeS_Diencephalon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47263 | 56.164 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ955866 | Salmonella phage vB_SenS_TUMS_E4 | Salmonella phage vB_SenS_TUMS_E4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43018 | 49.742 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MZ401009 | Clostridium phage CPQ10 | Clostridium phage CPQ10 unclassified bacterial viruses Viruses | 18254 | 34.683 | Unclassified | Unclassified | Unclassified | Clostridium |
| MZ401007 | Clostridium phage CPQ7 | Clostridium phage CPQ7 unclassified bacterial viruses Viruses | 17980 | 34.778 | Unclassified | Unclassified | Unclassified | Clostridium |
| MZ401006 | Clostridium phage CPQ4 | Clostridium phage CPQ4 unclassified bacterial viruses Viruses | 40182 | 30.307 | Unclassified | Unclassified | Unclassified | Clostridium |
| MZ401008 | Clostridium phage CPQ9 | Clostridium phage CPQ9 unclassified bacterial viruses Viruses | 18270 | 34.702 | Unclassified | Unclassified | Unclassified | Clostridium |
| MZ401005 | Clostridium phage CPQ3 | Clostridium phage CPQ3 unclassified bacterial viruses Viruses | 18246 | 34.72 | Unclassified | Unclassified | Unclassified | Clostridium |
| AF022214 | Mycobacterium phage D29 | Mycobacterium phage D29 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49127 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AF125163 | Bacteriophage K139 | Bacteriophage K139 Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33106 | 48.897 | Longwoodvirus | Peduovirinae | Myoviridae | Unspecified |
| AF145054 | Streptococcus phage 7201 | Streptococcus phage 7201 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35466 | 38.727 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| AF157835 | Hamiltonella phage APSE-1 | Hamiltonella phage APSE-1 Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36524 | 43.886 | Sendosyvirus | Unclassified | Podoviridae | Hamiltonella |
| AF115103 | Streptococcus phage Sfi21 | Streptococcus phage Sfi21 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40739 | 37.566 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| AF115102 | Streptococcus phage Sfi19 | Streptococcus phage Sfi19 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37370 | 38.274 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| AF155037 | Pseudoalteromonas phage PM2 | Pseudoalteromonas phage PM2 Corticovirus Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10079 | 42.167 | Corticovirus | Unclassified | Corticoviridae | Pseudoalteromonas |
| AF063097 | Bacteriophage P2 | Bacteriophage P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33593 | 50.168 | Peduovirus | Peduovirinae | Myoviridae | Unspecified |
| AF011378 | Lactococcus phage SK1 | Lactococcus phage SK1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28451 | 34.512 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JF937091 | Mycobacterium phage Bask21 | Mycobacterium phage Bask21 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74997 | 62.857 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JX307702 | Mycobacterium phage Arturo | Mycobacterium phage Arturo Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51500 | 64.052 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX015524 | Mycobacterium phage Astro | Mycobacterium phage Astro Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52494 | 61.38 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699005 | Mycobacterium phage Alma | Mycobacterium phage Alma Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53177 | 62.461 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699000 | Mycobacterium phage BillKnuckles | Mycobacterium phage BillKnuckles Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51821 | 63.436 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937090 | Mycobacterium phage Baka | Mycobacterium phage Baka Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111688 | 60.732 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704106 | Mycobacterium phage Anaya | Mycobacterium phage Anaya Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60835 | 66.394 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704093 | Mycobacterium phage Backyardigan | Mycobacterium phage Backyardigan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51308 | 63.688 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN083852 | Mycobacterium phage Benedict | Mycobacterium phage Benedict Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 59.797 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF957057 | Mycobacterium phage BPBiebs31 | Mycobacterium phage BPBiebs31 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53171 | 63.448 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| HM152764 | Mycobacterium phage Angelica | Mycobacterium phage Angelica Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59598 | 66.388 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| GU060500 | Mycobacterium phage Ardmore | Mycobacterium phage Ardmore Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52141 | 61.491 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ168659 | Mycobacterium phage Brujita | Mycobacterium phage Brujita Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47057 | 66.772 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| EU816590 | Mycobacterium phage Boomer | Mycobacterium phage Boomer Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58037 | 61.128 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ398041 | Mycobacterium phage 244 | Mycobacterium phage 244 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74483 | 62.937 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| AY500153 | Mycobacterium phage Bethlehem | Mycobacterium phage Bethlehem Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52250 | 63.246 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AF271693 | Mycobacterium phage Bxb1 | Mycobacterium phage Bxb1 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50550 | 63.65 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OL505078 | Escherichia phage BI-EHEC | Escherichia phage BI-EHEC unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151425 | 39.004 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| OL512804 | Hafnia phage Pocitis76 | Hafnia phage Pocitis76 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44148 | 46.29 | Unclassified | Unclassified | Siphoviridae | Hafnia |
| OL539470 | Cronobacter phage vB_CsaM_Invicta | Cronobacter phage vB_CsaM_Invicta unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179053 | 44.746 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| OL539459 | Escherichia phage vB_EcoD_Fulano1 | Escherichia phage vB_EcoD_Fulano1 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51759 | 43.95 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| OL449681 | Phage vB_EcoM-p111 | Phage vB_EcoM-p111 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50241 | 46.329 | Nouzillyvirus | Unclassified | Drexlerviridae | Unspecified |
| OL539727 | Escherichia virus fEg-Eco19 | Escherichia virus fEg-Eco19 unclassified bacterial viruses Viruses | 45805 | 50.23 | Unclassified | Unclassified | Unclassified | Escherichia |
| OL474141 | Salmonella phage ph2-2 | Salmonella phage ph2-2 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85944 | 38.803 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| OL473897 | Pseudomonas phage Pa BHU-15 | Pseudomonas phage Pa BHU-15 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91015 | 52.108 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| OK665488 | Pseudomonas phage VB_PaeS_VL1 | Pseudomonas phage VB_PaeS_VL1 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73308 | 54.731 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MZ156570 | Salmonella phage Liang dang 541 | Salmonella phage Liang dang 541 unclassified bacterial viruses Viruses | 157697 | 44.492 | Unclassified | Unclassified | Unclassified | Salmonella |
| MZ156569 | Salmonella phage SM86 | Salmonella phage SM86 unclassified bacterial viruses Viruses | 43508 | 49.492 | Unclassified | Unclassified | Unclassified | Salmonella |
| MZ156568 | Salmonella phage 95YS | Salmonella phage 95YS unclassified bacterial viruses Viruses | 43511 | 49.491 | Unclassified | Unclassified | Unclassified | Salmonella |
| MZ156567 | Salmonella phage 26YS | Salmonella phage 26YS unclassified bacterial viruses Viruses | 42161 | 49.57 | Unclassified | Unclassified | Unclassified | Salmonella |
| OV101294 | Escherichia phage vB_Eco_Alma | Escherichia phage vB_Eco_Alma unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136980 | 43.542 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| OV049961 | Escherichia phage vB_Eco_SPSP | Escherichia phage vB_Eco_SPSP unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70979 | 41.36 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| OV049960 | Escherichia phage vB_Eco_PATM | Escherichia phage vB_Eco_PATM unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40010 | 47.008 | Unclassified | Studiervirinae | Autographiviridae | Escherichia |
| OU734268 | Escherichia phage vB_Eco_Sip | Escherichia phage vB_Eco_Sip unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50809 | 44.638 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LR999871 | Escherichia phage vB_Eco_Jura | Escherichia phage vB_Eco_Jura unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71488 | 41.338 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| LR999870 | Escherichia phage vB_Eco_CS16 | Escherichia phage vB_Eco_CS16 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42656 | 54.471 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LR990704 | Escherichia phage vB_Eco_NR1 | Escherichia phage vB_Eco_NR1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167153 | 35.381 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LR990702 | Escherichia phage vB_Eco_Maverick | Escherichia phage vB_Eco_Maverick unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42624 | 54.512 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LR880803 | Escherichia phage Barry | Escherichia phage Barry unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87693 | 38.947 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| LR865361 | Escherichia phage Paula | Escherichia phage Paula unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146554 | 37.476 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| OV192280 | Escherichia phage P2_AC1 | Escherichia phage P2_AC1 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33349 | 50.655 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| OK272490 | Escherichia phage JLBYU32 | Escherichia phage JLBYU32 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86143 | 39.043 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| OK272489 | Escherichia phage JLBYU19 | Escherichia phage JLBYU19 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49897 | 44.732 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272488 | Escherichia phage JLBYU37 | Escherichia phage JLBYU37 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45011 | 54.431 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK272484 | Escherichia phage JLBYU22 | Escherichia phage JLBYU22 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166300 | 35.494 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OK272483 | Escherichia phage JLBYU26 | Escherichia phage JLBYU26 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51253 | 45.01 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272482 | Escherichia phage JLBYU08 | Escherichia phage JLBYU08 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136893 | 43.676 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK272481 | Escherichia phage JLBYU38 | Escherichia phage JLBYU38 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49995 | 44.744 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272480 | Escherichia phage JLBYU40 | Escherichia phage JLBYU40 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109776 | 39.062 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| OK272479 | Escherichia phage JLBYU41 | Escherichia phage JLBYU41 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 44.404 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272478 | Escherichia phage JLBYU01 | Escherichia phage JLBYU01 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51210 | 44.569 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272477 | Escherichia phage JLBYU43 | Escherichia phage JLBYU43 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111814 | 38.824 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| OK272476 | Escherichia phage JLBYU24 | Escherichia phage JLBYU24 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168125 | 35.525 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OK272475 | Escherichia phage JLBYU27 | Escherichia phage JLBYU27 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16547 | 54.427 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK272474 | Escherichia phage JLBYU60 | Escherichia phage JLBYU60 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44804 | 54.531 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK272473 | Escherichia phage JLBYU31 | Escherichia phage JLBYU31 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166825 | 35.419 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OK272472 | Escherichia phage JLBYU28 | Escherichia phage JLBYU28 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86784 | 38.968 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| OK272471 | Escherichia phage JLBYU16 | Escherichia phage JLBYU16 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49995 | 44.744 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OL539471 | Cronobacter phage vB_CsaD_Banach | Cronobacter phage vB_CsaD_Banach unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178878 | 44.686 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| OL539469 | Proteus phage vB_PmiP_Brookers | Proteus phage vB_PmiP_Brookers Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40152 | 40.698 | Lederbergvirus | Unclassified | Podoviridae | Proteus |
| OL539468 | Serratia phage vB_SmaM_Haymo | Serratia phage vB_SmaM_Haymo unclassified Moabitevirus Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 274604 | 46.807 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| OL539465 | Citrobacter phage vB_CfrD_Reinasaurus | Citrobacter phage vB_CfrD_Reinasaurus unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48103 | 42.889 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OL539464 | Serratia phage vB_SmaS_Stoker | Serratia phage vB_SmaS_Stoker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41797 | 46.377 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539462 | Citrobacter phage vB_CfrD_BlueShadow | Citrobacter phage vB_CfrD_BlueShadow unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49619 | 42.828 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OL539461 | Citrobacter phage_vB_CfrD_Devorator | Citrobacter phage_vB_CfrD_Devorator unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49712 | 42.869 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OL539458 | Klebsiella phage vB_KaeM_Nispero | Klebsiella phage vB_KaeM_Nispero unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166929 | 39.354 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OL539456 | Serratia phage vB_SmaS_LittleDog | Serratia phage vB_SmaS_LittleDog Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41738 | 45.795 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539455 | Serratia phage vB_SmaS_Niamh | Serratia phage vB_SmaS_Niamh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42053 | 45.956 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539454 | Escherichia phage vB_EcoS_OddieOddie | Escherichia phage vB_EcoS_OddieOddie unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59985 | 44.539 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| OL539453 | Klebsiella phage vB_KpnD_Opt-79 | Klebsiella phage vB_KpnD_Opt-79 unclassified Rogunavirinae Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48301 | 44.689 | Unclassified | Rogunavirinae | Drexlerviridae | Klebsiella |
| OL539452 | Serratia phage vB_SmaS_Opt-155 | Serratia phage vB_SmaS_Opt-155 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42792 | 51.94 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539449 | Enterococcus phage vB_EfaS_Paulomi | Enterococcus phage vB_EfaS_Paulomi unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41921 | 34.629 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| OL539448 | Klebsiella phage vB_KpnD_PeteCarol | Klebsiella phage vB_KpnD_PeteCarol unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49054 | 50.938 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OL539444 | Escherichia phage_vB_EcoD_Punny | Escherichia phage_vB_EcoD_Punny unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49765 | 42.829 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OL539443 | Citrobacter phage vB_CfrD_Brooksby | Citrobacter phage vB_CfrD_Brooksby unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50071 | 42.779 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OL539442 | Serratia phage vB_SmaS_Ulliraptor | Serratia phage vB_SmaS_Ulliraptor Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42052 | 45.974 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539441 | Serratia phage vB_SmaS_PhooPhighters | Serratia phage vB_SmaS_PhooPhighters Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39188 | 42.163 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539439 | Serratia phage vB_SmaS_Carrot | Serratia phage vB_SmaS_Carrot Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41293 | 45.739 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OL539438 | Serratia phage vB_SmaS_Swain | Serratia phage vB_SmaS_Swain Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41292 | 45.733 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OK500002 | Bacillus phage vB_BanS_Athena | Bacillus phage vB_BanS_Athena Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37369 | 35.267 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK500000 | Proteus phage vB_PmiS_DoubleBarrel | Proteus phage vB_PmiS_DoubleBarrel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59089 | 46.818 | Unclassified | Unclassified | Siphoviridae | Proteus |
| OK499999 | Baciilus phage vB_BanS_Booya | Baciilus phage vB_BanS_Booya unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39660 | 35.343 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| OK499998 | Proteus phage vB_PmiS_DanisaurMW | Proteus phage vB_PmiS_DanisaurMW Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58538 | 46.858 | Unclassified | Unclassified | Siphoviridae | Proteus |
| OK499995 | Serratia phage vB_SmaS_Tlacuache | Serratia phage vB_SmaS_Tlacuache Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42679 | 51.637 | Unclassified | Unclassified | Siphoviridae | Serratia |
| OK499994 | Bacillus phage vB_BanS_Skywalker | Bacillus phage vB_BanS_Skywalker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160928 | 34.274 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK499993 | Escherichia phage vB_EcoS_Teewinot | Escherichia phage vB_EcoS_Teewinot unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41800 | 54.493 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK499992 | Bacillus phage vB_BanS_Nate | Bacillus phage vB_BanS_Nate Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166879 | 33.601 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK499991 | Bacillus phage vB_BanS_Sophrita | Bacillus phage vB_BanS_Sophrita Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167995 | 32.359 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK499990 | Bacillus phage vB_BanH_RonSwanson | Bacillus phage vB_BanH_RonSwanson unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161711 | 37.814 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| OK499989 | Morganella phage vB_MmoM_Rgz1 | Morganella phage vB_MmoM_Rgz1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165808 | 34.712 | Unclassified | Tevenvirinae | Myoviridae | Morganella |
| OK499988 | Escherichia phage vB_EcoD_Pubbukkers | Escherichia phage vB_EcoD_Pubbukkers unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44476 | 54.571 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK499986 | Proteus phage vB_PmiS_NotEvenPhaged | Proteus phage vB_PmiS_NotEvenPhaged Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58566 | 46.928 | Unclassified | Unclassified | Siphoviridae | Proteus |
| OK499985 | Escherichia phage vB_EcoS_Over9000 | Escherichia phage vB_EcoS_Over9000 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44583 | 54.604 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK499984 | Escherichia phage vB_EcoD_Mishu | Escherichia phage vB_EcoD_Mishu unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49713 | 42.868 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK499981 | Klebsiella phage vB_KaeM_Merci | Klebsiella phage vB_KaeM_Merci unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165052 | 39.306 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OK499979 | Bacillus Phage vB_BanS_McSteamy | Bacillus Phage vB_BanS_McSteamy unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39748 | 34.933 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| OK499978 | Klebsiella phage vB_KpnM_JustaPhage | Klebsiella phage vB_KpnM_JustaPhage unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48129 | 48.594 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| OK499977 | Bacillus phage vB_BanH_Emiliahah | Bacillus phage vB_BanH_Emiliahah unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158566 | 39.914 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| OK499976 | Bacillus phage vB_BanH_JarJar | Bacillus phage vB_BanH_JarJar unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161711 | 37.811 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| OK499975 | Proteus_phage_vB_PmiS_Jing313 | Proteus_phage_vB_PmiS_Jing313 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58534 | 46.877 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| OK499973 | Citrobacter phage vB_CfrD_ElisaCorrea | Citrobacter phage vB_CfrD_ElisaCorrea unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49523 | 42.877 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| OK499997 | Kelbsiella phage vB_KaeM_Boboto | Kelbsiella phage vB_KaeM_Boboto unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166665 | 39.52 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OK499996 | Klebsiella phage vB_KaeM_Shinkou | Klebsiella phage vB_KaeM_Shinkou unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162839 | 39.56 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OK499987 | Bacillus phage vB_BanS_MrDarsey | Bacillus phage vB_BanS_MrDarsey Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164998 | 34.227 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK499983 | Bacillus phage vB_BanH_McCartney | Bacillus phage vB_BanH_McCartney unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158334 | 39.893 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| OK499982 | Morganella phage vB_MmoP_Lilpapawes | Morganella phage vB_MmoP_Lilpapawes Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39168 | 47.064 | Minipunavirus | Studiervirinae | Autographiviridae | Morganella |
| OK499980 | Klebsiella phage vB_KaeM_LilPanda | Klebsiella phage vB_KaeM_LilPanda unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164872 | 39.548 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OL615013 | Shigella phage vB_SboM_ChubbyThor | Shigella phage vB_SboM_ChubbyThor unclassified Agtrevirus Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159319 | 50.096 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| OL615012 | Shigella phage vB_SboM_Phaginator | Shigella phage vB_SboM_Phaginator unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166647 | 35.456 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| OL615010 | Shigella phage vB_SboS_StarDew | Shigella phage vB_SboS_StarDew unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44715 | 54.682 | Dhillonvirus | Unclassified | Siphoviridae | Shigella |
| OK500001 | Bacillus phage vB_BanH_Abinadi | Bacillus phage vB_BanH_Abinadi unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158201 | 38.724 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| OK499972 | Bacillus phage vB_BanS_Chewbecca | Bacillus phage vB_BanS_Chewbecca Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161151 | 34.293 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| OK499974 | Proteus phage vB_PmiS_Inception | Proteus phage vB_PmiS_Inception Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58589 | 46.811 | Unclassified | Unclassified | Siphoviridae | Proteus |
| OK310506 | Microbacterium phage Inventa | Microbacterium phage Inventa unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41802 | 63.552 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| OK310505 | Mycobacterium phage Boiiii | Mycobacterium phage Boiiii unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59907 | 66.551 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| OK310504 | Microbacterium phage Josuke | Microbacterium phage Josuke Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17453 | 68.722 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK310503 | Arthrobacter phage Tokki | Arthrobacter phage Tokki unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57652 | 50.33 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| OK310502 | Streptomyces phage Bartholomune | Streptomyces phage Bartholomune unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131879 | 50.201 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| OK275495 | Xanthomonas phage R3-22-T1 | Xanthomonas phage R3-22-T1 unclassified Pradovirus Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45281 | 61.578 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| OK275494 | Xanthomonas phage MYK3 | Xanthomonas phage MYK3 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47500 | 62.907 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| OK275493 | Xanthomonas phage MET13-T1 | Xanthomonas phage MET13-T1 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63593 | 68.193 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| OK275492 | Xanthomonas phage FMYAK-P1 | Xanthomonas phage FMYAK-P1 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64088 | 67.776 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| OK275491 | Pseudomonas phage REC1 | Pseudomonas phage REC1 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97884 | 49.001 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| OK272487 | Escherichia phage JLBYU09 | Escherichia phage JLBYU09 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48988 | 44.107 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272486 | Escherichia phage JLBYU10 | Escherichia phage JLBYU10 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50987 | 44.545 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272485 | Escherichia phage JLBYU07 | Escherichia phage JLBYU07 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50290 | 44.601 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| OK272470 | Escherichia phage JLBYU50 | Escherichia phage JLBYU50 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141597 | 37.458 | Unclassified | Unclassified | Myoviridae | Escherichia |
| OK999981 | Microbacterium phage Skylord | Microbacterium phage Skylord unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39843 | 67.088 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| OK999980 | Arthrobacter phage EastWest | Arthrobacter phage EastWest Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50397 | 61.541 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| OK999979 | Microbacterium phage Franklin22 | Microbacterium phage Franklin22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40567 | 66.096 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK999978 | Mycobacterium phage Grub | Mycobacterium phage Grub Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50841 | 64.033 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| OK539826 | Pseudomonas phage phipa10 | Pseudomonas phage phipa10 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92393 | 49.488 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| OK539825 | Pseudomonas phage phipa4 | Pseudomonas phage phipa4 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42943 | 53.734 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| OK539824 | Pseudomonas phage phipa2 | Pseudomonas phage phipa2 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43291 | 62.249 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| OK258140 | Xanthomonas phage DMF5-T1 | Xanthomonas phage DMF5-T1 Eisenstarkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45054 | 55.955 | Eisenstarkvirus | Unclassified | Siphoviridae | Xanthomonas |
| OK258139 | Pseudomonas phage COT4 | Pseudomonas phage COT4 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97989 | 49.05 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| OK258138 | Xanthomonas phage MUD8-T1 | Xanthomonas phage MUD8-T1 unclassified Pradovirus Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44322 | 61.62 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| OL581612 | Pseudomonas phage Eisa9 | Pseudomonas phage Eisa9 Pollyceevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40730 | 59.236 | Pollyceevirus | Unclassified | Autographiviridae | Pseudomonas |
| OL581611 | Pseudomonas phage Eir4 | Pseudomonas phage Eir4 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40177 | 57.284 | Unclassified | Unclassified | Autographiviridae | Pseudomonas |
| OL519844 | Pseudomonas phage vB_PaeS_TUMS_P81 | Pseudomonas phage vB_PaeS_TUMS_P81 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73167 | 54.955 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| OL519842 | Pseudomonas phage vB_PaeS_TUMS_P6 | Pseudomonas phage vB_PaeS_TUMS_P6 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73885 | 53.487 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MW598459 | Escherichia phage vB_EcoM_RZ | Escherichia phage vB_EcoM_RZ unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170318 | 40.147 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| OK340822 | Flavobacterium phage FpV4-1.3 | Flavobacterium phage FpV4-1.3 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88308 | 27.216 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| OK340821 | Flavobacterium phage FpV4-1.2 | Flavobacterium phage FpV4-1.2 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88308 | 27.216 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| OK340820 | Flavobacterium phage FpV4-1.1 | Flavobacterium phage FpV4-1.1 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88308 | 27.217 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| OK340819 | Flavobacterium phage FpV9-2.1 | Flavobacterium phage FpV9-2.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.012 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK340818 | Flavobacterium phage FpV9-1.3 | Flavobacterium phage FpV9-1.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.01 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK340817 | Flavobacterium phage FpV9-1.2 | Flavobacterium phage FpV9-1.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.005 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268427 | Flavobacterium phage FPSV-S20A3 | Flavobacterium phage FPSV-S20A3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42755 | 31.975 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268426 | Flavobacterium phage FPSV-S20A2 | Flavobacterium phage FPSV-S20A2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.977 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268425 | Flavobacterium phage FPSV-S20A1 | Flavobacterium phage FPSV-S20A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.977 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268424 | Flavobacterium phage FPSV-S20C | Flavobacterium phage FPSV-S20C Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.979 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268423 | Flavobacterium phage FPSV-S20B | Flavobacterium phage FPSV-S20B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.979 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OK268422 | Flavobacterium phage FPSV-S20A | Flavobacterium phage FPSV-S20A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.977 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| OL658619 | Oceanospirillum phage vB_OsaM_PD0307 | Oceanospirillum phage vB_OsaM_PD0307 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44421 | 57.131 | Unclassified | Unclassified | Myoviridae | Oceanospirillum |
| OL656105 | Rhodococcus phage P19 | Rhodococcus phage P19 unclassified bacterial viruses Viruses | 17065 | 55.429 | Unclassified | Unclassified | Unclassified | Rhodococcus |
| OL652658 | Salmonella phage JD02 | Salmonella phage JD02 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43955 | 49.532 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OL436139 | Salmonella phage LP31 | Salmonella phage LP31 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42600 | 49.667 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OL362041 | Escherichia phage vB_Eco_OMNI12 | Escherichia phage vB_Eco_OMNI12 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169246 | 35.324 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OL409034 | Klebsiella phage vB_KpnP_ZK2 | Klebsiella phage vB_KpnP_ZK2 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40946 | 52.271 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| OL348188 | Aeromonas phage vB_AsaM_LPM4 | Aeromonas phage vB_AsaM_LPM4 unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37312 | 58.134 | Unclassified | Peduovirinae | Myoviridae | Aeromonas |
| OL343755 | Pseudomonas phage BHU-1 | Pseudomonas phage BHU-1 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91108 | 51.925 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ560702 | Klebsiella phage vB_KpnM_TU02 | Klebsiella phage vB_KpnM_TU02 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166230 | 38.341 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN101229 | Klebsiella phage KMI13 | Klebsiella phage KMI13 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158756 | 39.644 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN101226 | Klebsiella phage KMI12 | Klebsiella phage KMI12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122724 | 39.137 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MN101225 | Klebsiella phage KMI11 | Klebsiella phage KMI11 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98975 | 39.667 | Unclassified | Tevenvirinae | Myoviridae | Klebsiella |
| OL539729 | Salmonella phage PRF-SP3 | Salmonella phage PRF-SP3 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39907 | 50.282 | Kayfunavirus | Studiervirinae | Autographiviridae | Salmonella |
| OL474125 | Klebsiella phage BUCT86 | Klebsiella phage BUCT86 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44542 | 54.328 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MZ427930 | Staphylococcus phage SAP23 | Staphylococcus phage SAP23 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151618 | 30.377 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| OK274152 | Escherichia phage APTC-EC-2A | Escherichia phage APTC-EC-2A unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166608 | 35.343 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OK330456 | Pseudomonas phage TehO | Pseudomonas phage TehO unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43015 | 53.684 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| OK330455 | Pseudomonas phage Kopi | Pseudomonas phage Kopi unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42820 | 53.475 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ836210 | Klebsiella phage BUCT541 | Klebsiella phage BUCT541 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46100 | 48.156 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ927746 | Pseudomonas phage Guyu | Pseudomonas phage Guyu unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43141 | 54.839 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ927745 | Pseudomonas phage Kaya | Pseudomonas phage Kaya unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43067 | 54.292 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ504995 | Klebsiella phage Geezett | Klebsiella phage Geezett unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50707 | 48.133 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MZ345003 | Xanthomonas phage M29 | Xanthomonas phage M29 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42891 | 59.644 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| NC_055030 | Paenibacillus phage Scottie | Paenibacillus phage Scottie Paenibacillus virus Scottie Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55990 | 48.519 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_055029 | Paenibacillus phage Halcyone | Paenibacillus phage Halcyone Paenibacillus virus Halcyone Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55560 | 48.56 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MW505928 | Propionibacterium phage TCUCAP1 | Propionibacterium phage TCUCAP1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29547 | 53.826 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MW021767 | Serratia phage vB_SmaS_Opt-169 | Serratia phage vB_SmaS_Opt-169 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38609 | 42.257 | Unclassified | Unclassified | Myoviridae | Serratia |
| NC_048688 | Paenibacillus phage BN12 | Paenibacillus phage BN12 unclassified Sitaravirus Sitaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39485 | 42.56 | Sitaravirus | Unclassified | Siphoviridae | Paenibacillus |
| MT409176 | Salmonella phage SapYZU01 | Salmonella phage SapYZU01 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88462 | 38.866 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MN905160 | Pseudomonas phage PE09 | Pseudomonas phage PE09 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99455 | 48.141 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| OK583892 | Klebsiella phage KPPK108.1 | Klebsiella phage KPPK108.1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43755 | 53.603 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OV121131 | Ralstonia phage UAM5 | Ralstonia phage UAM5 unclassified Cimandefvirus Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59830 | 65.026 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| LC645451 | Stx2a-converting phage Stx2_IB14005 | Stx2a-converting phage Stx2_IB14005 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63073 | 50.132 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645450 | Escherichia phage PPwrbA_EC3734 | Escherichia phage PPwrbA_EC3734 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62966 | 50.573 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| LC645449 | Stx2a-converting phage Stx2_EC3734 | Stx2a-converting phage Stx2_EC3734 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62983 | 50.275 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645448 | Stx2a-converting phage Stx2_140166 | Stx2a-converting phage Stx2_140166 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62048 | 50.335 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645447 | Stx2a-converting phage Stx2_KIH15-140 | Stx2a-converting phage Stx2_KIH15-140 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62282 | 50.294 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645446 | Stx2a-converting phage Stx2_H27V05_wrbA | Stx2a-converting phage Stx2_H27V05_wrbA unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66479 | 50.037 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645445 | Stx2a-converting phage Stx2_H27V05_argW | Stx2a-converting phage Stx2_H27V05_argW unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64958 | 50.105 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645444 | Stx2a-converting phage Stx2_Ech14022 | Stx2a-converting phage Stx2_Ech14022 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62996 | 50.179 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645443 | Stx2a-converting phage Stx2_EH2011 | Stx2a-converting phage Stx2_EH2011 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63180 | 50.003 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645442 | Stx2a-converting phage Stx2_112991 | Stx2a-converting phage Stx2_112991 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62305 | 49.996 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645441 | Stx2a-converting phage Stx2_EH2085 | Stx2a-converting phage Stx2_EH2085 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65600 | 49.227 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC645440 | Stx2a-converting phage Stx2_EH2086 | Stx2a-converting phage Stx2_EH2086 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64232 | 49.187 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC645439 | Stx2a-converting phage Stx2_16002 | Stx2a-converting phage Stx2_16002 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60591 | 48.865 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC645438 | Stx2a-converting phage Stx2_09E025 | Stx2a-converting phage Stx2_09E025 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62398 | 50.003 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645437 | Stx1a-converting phage Stx1_12188 | Stx1a-converting phage Stx1_12188 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51057 | 51.256 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645436 | Stx1a-converting phage Stx1_EH2197 | Stx1a-converting phage Stx1_EH2197 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51047 | 51.249 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645435 | Stx1a-converting phage Stx1_112808 | Stx1a-converting phage Stx1_112808 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51061 | 51.284 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645434 | Stx1a-converting phage Stx1_120412 | Stx1a-converting phage Stx1_120412 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50991 | 51.299 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645433 | Stx1a-converting phage Stx1_1380 | Stx1a-converting phage Stx1_1380 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51055 | 51.26 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645432 | Stx1a-converting phage Stx1_1649 | Stx1a-converting phage Stx1_1649 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51055 | 51.249 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645431 | Stx1a-converting phage Stx1_8100 | Stx1a-converting phage Stx1_8100 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51054 | 51.287 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645430 | Stx1a-converting phage Stx1_131719 | Stx1a-converting phage Stx1_131719 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51056 | 51.244 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645429 | Stx1a-converting phage Stx1_111698 | Stx1a-converting phage Stx1_111698 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51055 | 51.251 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645428 | Stx1a-converting phage Stx1_9793 | Stx1a-converting phage Stx1_9793 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52371 | 51.317 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645427 | Escherichia phage PPwrbA_EH2201 | Escherichia phage PPwrbA_EH2201 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61247 | 50.368 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| LC645426 | Stx2c-converting phage Stx2_5044 | Stx2c-converting phage Stx2_5044 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61198 | 51.044 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645425 | Stx1a-converting phage Stx1_121427 | Stx1a-converting phage Stx1_121427 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57335 | 50.531 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC645424 | Stx2a-converting phage Stx2_12E109 | Stx2a-converting phage Stx2_12E109 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63021 | 50.179 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645423 | Stx2a-converting phage Stx2_PV11-35 | Stx2a-converting phage Stx2_PV11-35 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63014 | 50.171 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645422 | Stx2a-converting phage Stx2_12E070 | Stx2a-converting phage Stx2_12E070 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64332 | 50.249 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645421 | Stx2a-converting phage Stx2_09E126 | Stx2a-converting phage Stx2_09E126 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64344 | 50.249 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645420 | Stx2a-converting phage Stx2_120517 | Stx2a-converting phage Stx2_120517 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64328 | 50.249 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645419 | Stx2a-converting phage Stx2_112716 | Stx2a-converting phage Stx2_112716 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62974 | 50.21 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645418 | Stx2a-converting phage Stx2_112312 | Stx2a-converting phage Stx2_112312 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62982 | 50.205 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645417 | Stx2a-converting phage Stx2_110509 | Stx2a-converting phage Stx2_110509 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64341 | 50.279 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645416 | Stx2a-converting phage Stx2_131713 | Stx2a-converting phage Stx2_131713 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63012 | 50.173 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645415 | Stx2a-converting phage Stx2_122200 | Stx2a-converting phage Stx2_122200 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64324 | 50.25 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645414 | Stx2a-converting phage Stx2_130296 | Stx2a-converting phage Stx2_130296 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63015 | 50.171 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645413 | Stx2a-converting phage Stx2_131990 | Stx2a-converting phage Stx2_131990 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65627 | 50.287 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645412 | Stx2a-converting phage Stx2_PV11-59 | Stx2a-converting phage Stx2_PV11-59 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63008 | 50.176 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645411 | Stx2a-converting phage Stx2_111430 | Stx2a-converting phage Stx2_111430 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63000 | 50.173 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645410 | Stx2a-converting phage Stx2_PV11-77 | Stx2a-converting phage Stx2_PV11-77 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62735 | 50.187 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645409 | Stx2a-converting phage Stx2_EC3283 | Stx2a-converting phage Stx2_EC3283 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63049 | 50.177 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| LC645408 | Stx2c-converting phage Stx2_EH1904 | Stx2c-converting phage Stx2_EH1904 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64967 | 51.374 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| OK149171 | Alcaligenes phage vB_AfaP_QDWS595 | Alcaligenes phage vB_AfaP_QDWS595 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70466 | 41.116 | Unclassified | Unclassified | Schitoviridae | Alcaligenes |
| OK649958 | Hardygib1 virus | Hardygib1 virus Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73193 | 56.277 | Unclassified | Unclassified | Unclassified | Unspecified |
| MZ398243 | Escherichia phage IME267 | Escherichia phage IME267 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76631 | 41.983 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MW927519 | Paenibacillus phage Neutron | Paenibacillus phage Neutron unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35988 | 41.85 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MZ171381 | Mycobacterium phage Neos5 | Mycobacterium phage Neos5 unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68886 | 67.49 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ171377 | Mycobacterium phage Jolene | Mycobacterium phage Jolene unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42305 | 66.692 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ171375 | Mycobacterium phage Yinz | Mycobacterium phage Yinz unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68093 | 67.572 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ171374 | Gordonia phage AlumE | Gordonia phage AlumE unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52698 | 66.784 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MZ171371 | Gordonia phage Sahara | Gordonia phage Sahara unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45072 | 66.678 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MZ171370 | Gordonia phage Roney | Gordonia phage Roney unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59907 | 51.044 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ348424 | Kosakonia phage Kc304 | Kosakonia phage Kc304 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171207 | 38.819 | Winklervirus | Unclassified | Myoviridae | Kosakonia |
| MZ348423 | Kosakonia phage 305 | Kosakonia phage 305 unclassified Kanagawavirus Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174783 | 40.019 | Kanagawavirus | Unclassified | Myoviridae | Kosakonia |
| MZ348422 | Kosakonia phage Kc263 | Kosakonia phage Kc263 Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253740 | 45.455 | Petsuvirus | Unclassified | Myoviridae | Kosakonia |
| MZ348421 | Kosakonia phage Kc283 | Kosakonia phage Kc283 unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76232 | 48.272 | Cbunavirus | Unclassified | Schitoviridae | Kosakonia |
| MZ388556 | Gordonia phage Nithya | Gordonia phage Nithya unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61092 | 51.326 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ388555 | Mycobacterium phage Leperchaun | Mycobacterium phage Leperchaun unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58194 | 61.58 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ388552 | Gordonia phage Norvs | Gordonia phage Norvs unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92405 | 61.974 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MZ388551 | Mycobacterium phage Delylah | Mycobacterium phage Delylah unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155626 | 64.757 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MZ274309 | Arthrobacter phage CastorTray | Arthrobacter phage CastorTray unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58377 | 50.006 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ274308 | Arthrobacter phage Popper | Arthrobacter phage Popper Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41822 | 65.238 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MZ274307 | Microbacterium phage Leafy | Microbacterium phage Leafy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17407 | 68.668 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ274306 | Mycobacterium phage Zizzle | Mycobacterium phage Zizzle unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59381 | 61.53 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005683 | Gordonia phage Runhaar | Gordonia phage Runhaar unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60229 | 51.553 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ005682 | Arthrobacter phage Kaylissa | Arthrobacter phage Kaylissa unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44124 | 67.616 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ005680 | Gordonia phage Fugax | Gordonia phage Fugax unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59506 | 67.828 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MZ005679 | Mycobacterium phage Chaser | Mycobacterium phage Chaser unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76685 | 58.539 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005678 | Microbacterium phage BrokMonster | Microbacterium phage BrokMonster Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17433 | 68.944 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ005677 | Gordonia phage Bock | Gordonia phage Bock unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50220 | 67.228 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MZ005676 | Microbacterium phage Aries55 | Microbacterium phage Aries55 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17452 | 68.726 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ005675 | Gordonia phage AlainaMarie | Gordonia phage AlainaMarie unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61315 | 51.293 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ005672 | Mycobacterium phage Hail | Mycobacterium phage Hail unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.451 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005671 | Gordonia phage DumpTruck | Gordonia phage DumpTruck unclassified Lambovirus Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67754 | 57.949 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MZ209304 | Microbacterium phage Trireme | Microbacterium phage Trireme Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17030 | 68.996 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ209303 | Arthrobacter phage DevitoJr | Arthrobacter phage DevitoJr unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58044 | 49.955 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ209301 | Gordonia phage Figliar | Gordonia phage Figliar unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61147 | 51.497 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ322027 | Microbacterium phage BoomRoasted | Microbacterium phage BoomRoasted Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17455 | 68.645 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ322026 | Microbacterium phage Phrancesco | Microbacterium phage Phrancesco unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53255 | 68.889 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ322025 | Mycobacterium phage Ulysses | Mycobacterium phage Ulysses unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51057 | 64.013 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322020 | Gordonia phage DoctorFroggo | Gordonia phage DoctorFroggo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61476 | 67.425 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322019 | Mycobacterium Phage MiculUcigas | Mycobacterium Phage MiculUcigas unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52760 | 63.525 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322018 | Microbacterium phage Blab | Microbacterium phage Blab unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62150 | 66.771 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ322017 | Mycobacterium phage Eaglepride | Mycobacterium phage Eaglepride unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50926 | 64.945 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322015 | Gordonia phage Fitzgerald | Gordonia phage Fitzgerald unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58988 | 67.785 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MZ322014 | Microbacterium phage Octopus | Microbacterium phage Octopus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17368 | 68.511 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ322013 | Gordonia phage AikoCarson | Gordonia phage AikoCarson unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45241 | 61.164 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MZ322012 | Microbacterium phage Abigail | Microbacterium phage Abigail unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41003 | 66.463 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ322011 | Microbacterium phage Jehoshaphat | Microbacterium phage Jehoshaphat unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62294 | 67.013 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ322009 | Gordonia phage Hans | Gordonia phage Hans Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59258 | 68.284 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322008 | Microbacterium phage Aztec | Microbacterium phage Aztec Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17368 | 68.494 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ322007 | Mycobacterium Phage Cookiedough | Mycobacterium Phage Cookiedough unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52469 | 61.535 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322006 | Microbacterium phage ManAs | Microbacterium phage ManAs Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.471 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ322004 | Mycobacterium Phage Sunshine924 | Mycobacterium Phage Sunshine924 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51349 | 63.861 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ150793 | Gordonia phage MScarn | Gordonia phage MScarn unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45677 | 60.335 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MZ150792 | Microbacterium phage Selwyn23 | Microbacterium phage Selwyn23 unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53205 | 68.898 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ150791 | Gordonia phage YungMoney | Gordonia phage YungMoney unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60495 | 67.265 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MZ150788 | Gordonia phage Yeet412 | Gordonia phage Yeet412 unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46684 | 66.768 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MZ150786 | Mycobacterium phage Knocker | Mycobacterium phage Knocker unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71459 | 69.732 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ150784 | Arthrobacter phage Zaheer | Arthrobacter phage Zaheer Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42874 | 65.037 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MZ150783 | Microbacterium phage A3Wally | Microbacterium phage A3Wally Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194724 | 60.073 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW862993 | Mycobacterium phage Freya | Mycobacterium phage Freya unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68035 | 66.503 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW862992 | Arthrobacter phage SilentRX | Arthrobacter phage SilentRX unclassified Tankvirus Tankvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65232 | 67.116 | Tankvirus | Unclassified | Siphoviridae | Arthrobacter |
| MW862989 | Mycobacterium phage Sassafras | Mycobacterium phage Sassafras unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57051 | 61.375 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW862988 | Gordonia phage Tolls | Gordonia phage Tolls unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44786 | 59.809 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MW862987 | Gordonia phage Aflac | Gordonia phage Aflac unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61288 | 51.446 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MW862983 | Mycobacterium phage Marsha | Mycobacterium phage Marsha unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54459 | 63.909 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW862982 | Arthrobacter phage Sicarius2 | Arthrobacter phage Sicarius2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49221 | 63.538 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| MW862981 | Microbacterium phage Honk | Microbacterium phage Honk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50154 | 70.301 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW862980 | Mycobacterium phage True | Mycobacterium phage True unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68922 | 66.436 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW712737 | Mycobacterium phage Winget | Mycobacterium phage Winget unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71544 | 65.361 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712735 | Arthrobacter phage Rizwana | Arthrobacter phage Rizwana unclassified Tankvirus Tankvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65823 | 63.612 | Tankvirus | Unclassified | Siphoviridae | Arthrobacter |
| MW712732 | Mycobacterium phage Burton | Mycobacterium phage Burton unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52607 | 63.644 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712731 | Mycobacterium phage Lars | Mycobacterium phage Lars unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67468 | 68.923 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW712729 | Mycobacterium phage BananaFence | Mycobacterium phage BananaFence unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155947 | 64.678 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MW712728 | Microbacterium phage Quammi | Microbacterium phage Quammi unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61903 | 66.943 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW712727 | Microbacterium phage Teehee | Microbacterium phage Teehee unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62291 | 66.957 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW712726 | Mycobacterium phage Cher | Mycobacterium phage Cher unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68855 | 66.441 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW712725 | Gordonia phage EdmundFerry | Gordonia phage EdmundFerry Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55873 | 67.594 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW712724 | Microbacterium phage StrawberryJamm | Microbacterium phage StrawberryJamm unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61953 | 66.956 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW712721 | Microbacterium phage CroZenni | Microbacterium phage CroZenni unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41471 | 66.591 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MW712720 | Mycobacterium phage Burr | Mycobacterium phage Burr unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68721 | 66.496 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW822149 | Mycobacterium phage Rose5 | Mycobacterium phage Rose5 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73949 | 58.719 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW822148 | Mycobacterium virus I3 | Mycobacterium virus I3 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156127 | 64.775 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MW822147 | Gordonia phage TuertoX | Gordonia phage TuertoX unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.632 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MW822146 | Gordonia phage Sedona | Gordonia phage Sedona unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59571 | 67.412 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MW822145 | Streptomyces phage TunaTartare | Streptomyces phage TunaTartare unclassified Annadreamyvirus Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130929 | 46.948 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MW822144 | Streptomyces phage KimJongPhill | Streptomyces phage KimJongPhill Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82521 | 63.991 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MW822143 | Mycobacterium phage Indra | Mycobacterium phage Indra unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52493 | 61.46 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ355728 | Mycobacterium Phage Leviathan | Mycobacterium Phage Leviathan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53239 | 63.249 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ355724 | Microbacterium phage Necrophoxinus | Microbacterium phage Necrophoxinus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64175 | 61.655 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ355722 | Mycobacterium phage Bexan | Mycobacterium phage Bexan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52078 | 63.843 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ028628 | Microbacterium phage Concrete | Microbacterium phage Concrete Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17432 | 68.592 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ028627 | Gordonia phage VanLee | Gordonia phage VanLee Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84560 | 67.773 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ028626 | Mycobacterium phage Latch | Mycobacterium phage Latch unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154550 | 64.745 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MW927525 | Paenibacillus phage Sneaky | Paenibacillus phage Sneaky unclassified Sitaravirus Sitaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39668 | 42.034 | Sitaravirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927524 | Paenibacillus phage Riker | Paenibacillus phage Riker unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39482 | 41.444 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927523 | Paenibacillus phage Picard | Paenibacillus phage Picard unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39416 | 41.788 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927522 | Paenibacillus phage Pahemo | Paenibacillus phage Pahemo unclassified Sitaravirus Sitaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42027 | 42.311 | Sitaravirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927521 | Paenibacillus phage Norbert | Paenibacillus phage Norbert unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39482 | 41.442 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927520 | Paenibacillus phage Newport | Paenibacillus phage Newport unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 41.533 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927518 | Paenibacillus phage Mock2 | Paenibacillus phage Mock2 Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38724 | 41.253 | Unclassified | Gochnauervirinae | Siphoviridae | Paenibacillus |
| MW927517 | Paenibacillus phage JackPlaque | Paenibacillus phage JackPlaque unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37997 | 41.774 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927516 | Paenibacillus phage Hobie | Paenibacillus phage Hobie unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39417 | 41.723 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927515 | Paenibacillus phage Gohan | Paenibacillus phage Gohan unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42083 | 41.501 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927514 | Paenibacillus phage Fitz | Paenibacillus phage Fitz unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 41.533 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927513 | Paenibacillus phage Bohemia | Paenibacillus phage Bohemia unclassified Sitaravirus Sitaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39670 | 42.014 | Sitaravirus | Unclassified | Siphoviridae | Paenibacillus |
| MW927512 | Paenibacillus phage Bert | Paenibacillus phage Bert Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38735 | 41.203 | Unclassified | Gochnauervirinae | Siphoviridae | Paenibacillus |
| MW927511 | Paenibacillus phage AlexiD | Paenibacillus phage AlexiD unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37997 | 41.764 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MW924649 | Mycobacterium phage Akhila | Mycobacterium phage Akhila unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56251 | 62.056 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW924647 | Microbacterium phage Stoor | Microbacterium phage Stoor unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41814 | 69.111 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MW924646 | Mycobacterium phage Lakes | Mycobacterium phage Lakes unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49127 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW924645 | Gordonia phage Sanjuju | Gordonia phage Sanjuju unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59075 | 67.731 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MW924644 | Mycobacterium phage GlenHope | Mycobacterium phage GlenHope unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68705 | 67.503 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW924643 | Mycobacterium phage Smeagan | Mycobacterium phage Smeagan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53311 | 63.328 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW924642 | Microbacterium phage BarBear | Microbacterium phage BarBear Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.747 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW924641 | Microbacterium phage SadLad | Microbacterium phage SadLad Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62801 | 68.959 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW924640 | Microbacterium phage BelmontSKP | Microbacterium phage BelmontSKP unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41665 | 66.581 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MW924639 | Mycobacterium phage Luna22 | Mycobacterium phage Luna22 unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53730 | 67.454 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW924638 | Microbacterium phage Fizzles | Microbacterium phage Fizzles Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62078 | 68.161 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW601225 | Gordonia phage ZiggyZoo | Gordonia phage ZiggyZoo Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50992 | 67.24 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MW601224 | Microbacterium phage Albright | Microbacterium phage Albright unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40883 | 66.262 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MW601223 | Arthrobacter phage Prairie | Arthrobacter phage Prairie Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49392 | 69.507 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| MW601221 | Microbacterium phage BabyYoda | Microbacterium phage BabyYoda unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41557 | 68.732 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MW601219 | Gordonia phage BeeGee | Gordonia phage BeeGee Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51776 | 66.243 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW601217 | Microbacterium phage Pulchra | Microbacterium phage Pulchra unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53312 | 68.844 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MW601216 | Mycobacterium phage TarsusIV | Mycobacterium phage TarsusIV unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52704 | 62.955 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW578837 | Microbacterium phage NoodlelyBoi | Microbacterium phage NoodlelyBoi unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53614 | 68.87 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MW578836 | Mycobacterium phage Nanosmite | Mycobacterium phage Nanosmite Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82676 | 61.655 | Unclassified | Mclasvirinae | Siphoviridae | Mycobacterium |
| MW534381 | Mycobacterium phage Bluephacebaby | Mycobacterium phage Bluephacebaby unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68899 | 66.496 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW534380 | Mycobacterium phage Anderson | Mycobacterium phage Anderson unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68726 | 66.49 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW534379 | Mycobacterium phage Lulwa | Mycobacterium phage Lulwa unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67689 | 66.5 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW534377 | Mycobacterium phage Aelin | Mycobacterium phage Aelin unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68702 | 66.452 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW534376 | Gordonia phage Ayotoya | Gordonia phage Ayotoya unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50854 | 67.049 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MW534375 | Gordonia phage ODay | Gordonia phage ODay Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58747 | 62.996 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW534374 | Gordonia phage Austin | Gordonia phage Austin Getseptimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73898 | 56.6 | Getseptimavirus | Unclassified | Siphoviridae | Gordonia |
| MW507136 | Streptomyces phage Cross | Streptomyces phage Cross unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132807 | 50.067 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MW507135 | Gordonia phage Dre3 | Gordonia phage Dre3 unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45810 | 60.539 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MW507134 | Streptomyces phage Battuta | Streptomyces phage Battuta unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130734 | 49.507 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MW507133 | Gordonia phage RoyalG | Gordonia phage RoyalG unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75686 | 59.291 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MW507131 | Arthrobacter phage Leona | Arthrobacter phage Leona Andrewvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38596 | 65.973 | Andrewvirus | Unclassified | Siphoviridae | Arthrobacter |
| MW507130 | Gordonia Phage Whiteclaw | Gordonia Phage Whiteclaw unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.632 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MW507127 | Microbacterium phage Wolfpack | Microbacterium phage Wolfpack Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17455 | 68.542 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW507125 | Gordonia phage Phlop | Gordonia phage Phlop unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58358 | 67.944 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MW507123 | Mycobacterium phage Rubeus | Mycobacterium phage Rubeus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49040 | 63.723 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW435858 | Mycobacterium phage Gandalf20 | Mycobacterium phage Gandalf20 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51735 | 63.609 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW435857 | Mycobacterium phage Hydro | Mycobacterium phage Hydro unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71159 | 68.878 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW435856 | Streptomyces phage Werner | Streptomyces phage Werner unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51566 | 65.698 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MW435855 | Mycobacterium phage OhShagHennessy | Mycobacterium phage OhShagHennessy unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72623 | 58.875 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW435853 | Streptomyces phage MeganTheeKilla | Streptomyces phage MeganTheeKilla unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128152 | 47.419 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MW365953 | Streptomyces phage Dwayne | Streptomyces phage Dwayne unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51582 | 65.721 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MW365952 | Streptomyces phage Belfort | Streptomyces phage Belfort unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128959 | 47.029 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MW365951 | Streptomyces phage Hippo | Streptomyces phage Hippo unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51566 | 65.65 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MW365950 | Microbacterium phage Phabia | Microbacterium phage Phabia unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61802 | 66.919 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW365949 | Microbacterium phage Namago | Microbacterium phage Namago unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61793 | 66.898 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW365946 | Microbacterium phage Rudy | Microbacterium phage Rudy unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61769 | 66.953 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MW132714 | Mycobacterium phage Pound | Mycobacterium phage Pound unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109901 | 60.85 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MW055913 | Arthrobacter phage Wollypog | Arthrobacter phage Wollypog Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63364 | 59.158 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MW055910 | Arthrobacter phage Tbone | Arthrobacter phage Tbone unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44111 | 67.641 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MW055909 | Microbacterium phage Namsahir | Microbacterium phage Namsahir Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17176 | 68.852 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW055908 | Gordonia phage Lamberg | Gordonia phage Lamberg unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44730 | 66.642 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MW055907 | Microbacterium phage JoBros | Microbacterium phage JoBros Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17363 | 68.49 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW055906 | Mycobacterium phage Indlovu | Mycobacterium phage Indlovu unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71061 | 69.442 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW055904 | Mycobacterium phage Flaverint | Mycobacterium phage Flaverint unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52593 | 63.746 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW055903 | Microbacterium phage DannyDe | Microbacterium phage DannyDe unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56087 | 63.598 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MW055902 | Mycobacterium phage Camster | Mycobacterium phage Camster unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47149 | 67.229 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MW314850 | Gordonia phage Lilbeanie | Gordonia phage Lilbeanie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59210 | 67.715 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW314849 | Gordonia phage Dexdert | Gordonia phage Dexdert Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55006 | 67.504 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW314847 | Gordonia phage Delrey21 | Gordonia phage Delrey21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61476 | 67.423 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW314845 | Mycobacterium phage TelAviv | Mycobacterium phage TelAviv unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71455 | 65.434 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW321655 | Microbacterium phage Lunatic | Microbacterium phage Lunatic Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16915 | 68.803 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW291027 | Rhodococcus phage PhailMary | Rhodococcus phage PhailMary unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45614 | 58.857 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MW291025 | Microbacterium phage Haunter | Microbacterium phage Haunter unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56083 | 63.645 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MW291024 | Mycobacterium phage Jubie | Mycobacterium phage Jubie unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75782 | 59.283 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW291022 | Gordonia phage Schwartz33 | Gordonia phage Schwartz33 unclassified Secretariatvirus Secretariatvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59457 | 54.005 | Secretariatvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MW291021 | Streptomyces phage EhyElimayoE | Streptomyces phage EhyElimayoE Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186386 | 66.707 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MW291020 | Microbacterium phage Jerky | Microbacterium phage Jerky Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17527 | 68.723 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW291018 | Microbacterium phage Atraxi | Microbacterium phage Atraxi unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53904 | 59.125 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW291017 | Streptomyces phage TurkishDelight | Streptomyces phage TurkishDelight Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93606 | 72.587 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MW291015 | Gordonia phage NancyRae | Gordonia phage NancyRae unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49775 | 67.275 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MW291014 | Streptomyces phage MindFlayer | Streptomyces phage MindFlayer unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130258 | 49.467 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MW291013 | Gordonia phage Mocha12 | Gordonia phage Mocha12 unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.628 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MW291012 | Mycobacterium phage StephanieG | Mycobacterium phage StephanieG unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156080 | 64.734 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT986029 | Gordonia phage Linetti | Gordonia phage Linetti unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78164 | 58.677 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MT986028 | Arthrobacter phage Thunderclap | Arthrobacter phage Thunderclap unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59150 | 53.011 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MT889395 | Mycobacterium phage Shifa | Mycobacterium phage Shifa unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154859 | 64.586 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT889394 | Arthrobacter phage Odyssey395 | Arthrobacter phage Odyssey395 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68939 | 65.609 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT889392 | Microbacterium phage Lahqtemish | Microbacterium phage Lahqtemish unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42174 | 66.259 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT889391 | Mycobacterium phage MiniMac | Mycobacterium phage MiniMac unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70064 | 59.166 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889389 | Microbacterium phage DickRichards | Microbacterium phage DickRichards unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41340 | 66.538 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT889388 | Arthrobacter phage Pippa | Arthrobacter phage Pippa unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50783 | 61.058 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MT889384 | Mycobacterium phage STLscum | Mycobacterium phage STLscum unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52265 | 63.674 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889383 | Mycobacterium phage MiniLon | Mycobacterium phage MiniLon unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75793 | 59.309 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889381 | Mycobacterium phage Suigeneris | Mycobacterium phage Suigeneris unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70079 | 68.443 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT889379 | Gordonia phage Diabla | Gordonia phage Diabla unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77117 | 59.156 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MT889378 | Gordonia phage Burnsey | Gordonia phage Burnsey unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46185 | 60.262 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MT889376 | Arthrobacter phage Phives | Arthrobacter phage Phives unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44204 | 67.256 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT889375 | Arthrobacter phage Adumb2043 | Arthrobacter phage Adumb2043 unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43100 | 66.981 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT889373 | Arthrobacter phage Brynnie | Arthrobacter phage Brynnie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38594 | 67.731 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT889372 | Microbacterium phage Avocadoman | Microbacterium phage Avocadoman unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41200 | 66.425 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT889371 | Microbacterium phage DelaGarza | Microbacterium phage DelaGarza Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39631 | 70.001 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT889370 | Gordonia phage GourdThymes | Gordonia phage GourdThymes unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75807 | 58.915 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MT889369 | Mycobacterium phage Inchworm | Mycobacterium phage Inchworm unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68929 | 66.342 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT889368 | Gordonia phage Sam12 | Gordonia phage Sam12 unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75280 | 58.94 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MT889367 | Gordonia phage Orla | Gordonia phage Orla Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47354 | 63.268 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT889366 | Arthrobacter phage London | Arthrobacter phage London unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43599 | 66.582 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT889365 | Microbacterium phage Quenya | Microbacterium phage Quenya unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41876 | 69.675 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT952854 | Mycobacterium phage Miniwave | Mycobacterium phage Miniwave unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74961 | 62.985 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MT952853 | Arthrobacter phage Orcanus | Arthrobacter phage Orcanus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39404 | 66.998 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT952852 | Streptomyces phage Beuffert | Streptomyces phage Beuffert unclassified Annadreamyvirus Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129935 | 47.742 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MT952851 | Mycobacterium phage Gervas | Mycobacterium phage Gervas unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69130 | 67.483 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT952850 | Microbacterium phage Sippinontea | Microbacterium phage Sippinontea Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17473 | 68.563 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT952849 | Mycobacterium phage Windsor | Mycobacterium phage Windsor unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68621 | 66.469 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT952848 | Microbacterium phage MuffinTheCat | Microbacterium phage MuffinTheCat unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 15494 | 55.054 | Unclassified | Unclassified | Tectiviridae | Microbacterium |
| MT952847 | Mycobacterium phage Topgun | Mycobacterium phage Topgun unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50977 | 63.915 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT952845 | Microbacterium phage GardenState | Microbacterium phage GardenState Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45424 | 70.016 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT952843 | Gordonia phage Twonlo | Gordonia phage Twonlo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56530 | 67.667 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT818427 | Microbacterium phage AnnaLie | Microbacterium phage AnnaLie unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41665 | 66.593 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT818426 | Microbacterium phage Lifes | Microbacterium phage Lifes Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59253 | 58.459 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT818424 | Gordonia phage GemG | Gordonia phage GemG unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46225 | 66.611 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MT818423 | Mycobacterium phage Janiyra | Mycobacterium phage Janiyra unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156653 | 64.621 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT818422 | Mycobacterium phage Periodt | Mycobacterium phage Periodt unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42498 | 66.657 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MT818421 | Mycobacterium phage Luke | Mycobacterium phage Luke unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53701 | 67.433 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT818419 | Mycobacterium phage Lolalove | Mycobacterium phage Lolalove unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71111 | 68.929 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT818416 | Mycobacterium phage BackyardAgain | Mycobacterium phage BackyardAgain unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153731 | 64.754 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT818415 | Microbacterium phage Scumberland | Microbacterium phage Scumberland unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53276 | 68.849 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MT771349 | Mycobacterium phage Tyson | Mycobacterium phage Tyson unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74429 | 58.72 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT771347 | Arthrobacter phage Paella | Arthrobacter phage Paella Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113698 | 47.624 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT771343 | Gordonia phage Clown | Gordonia phage Clown unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58203 | 65.741 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MT897910 | Mycobacterium phage VioletZ | Mycobacterium phage VioletZ unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71294 | 68.916 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT897909 | Mycobacterium phage Maru | Mycobacterium phage Maru unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68684 | 66.438 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT897908 | Streptomyces phage ShakeNBake | Streptomyces phage ShakeNBake Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69299 | 64.236 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT897906 | Arthrobacter phage Tweety19 | Arthrobacter phage Tweety19 unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40873 | 66.124 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT897905 | Streptomyces phage MulchMansion | Streptomyces phage MulchMansion unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134105 | 49.422 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MT897903 | Mycobacterium phage DirtJuice | Mycobacterium phage DirtJuice unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69166 | 66.407 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT897902 | Mycobacterium phage Boehler | Mycobacterium phage Boehler unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68341 | 66.543 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT897901 | Mycobacterium phage Adriana | Mycobacterium phage Adriana unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69122 | 66.419 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT684601 | Mycobacterium phage Hlubikazi | Mycobacterium phage Hlubikazi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55471 | 61.439 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684599 | Gordonia Phage Sephiroth | Gordonia Phage Sephiroth Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76058 | 58.642 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT684598 | Streptomyces phage Faust | Streptomyces phage Faust unclassified Annadreamyvirus Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130666 | 47.183 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MT684594 | Mycobacterium phage Hadrien | Mycobacterium phage Hadrien unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.445 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684592 | Microbacterium phage Aesir | Microbacterium phage Aesir unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56285 | 63.582 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT684591 | Microbacterium phage Chivey | Microbacterium phage Chivey unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56082 | 63.66 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT684590 | Streptomyces phage LilMartin | Streptomyces phage LilMartin unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132470 | 49.483 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MT657342 | Mycobacterium phage Mule | Mycobacterium phage Mule unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51468 | 63.583 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT657341 | Mycobacterium phage Estes | Mycobacterium phage Estes unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83601 | 60.866 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MT657340 | Streptomyces phage Thiqqums | Streptomyces phage Thiqqums unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57624 | 58.075 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MT657339 | Gordonia phage SpeedDemon | Gordonia phage SpeedDemon unclassified Bantamvirus Bantamvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94031 | 64.557 | Bantamvirus | Unclassified | Siphoviridae | Gordonia |
| MT657338 | Mycobacterium phage SuperCallie99 | Mycobacterium phage SuperCallie99 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52086 | 61.46 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT657337 | Microbacterium phage Mazun | Microbacterium phage Mazun unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54839 | 59.813 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT657336 | Microbacterium phage ClearAsMud | Microbacterium phage ClearAsMud unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52987 | 68.991 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MT657335 | Gordonia phage Mariokart | Gordonia phage Mariokart unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60762 | 70.506 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MT657333 | Microbacterium phage Casend | Microbacterium phage Casend unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62461 | 66.941 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MT657332 | Microbacterium phage NeumannU | Microbacterium phage NeumannU Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.747 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT657331 | Mycobacterium phage Manu | Mycobacterium phage Manu unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50577 | 63.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT657330 | Mycobacterium phage Webster2 | Mycobacterium phage Webster2 unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.443 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT776813 | Mycobacterium phage Magic8 | Mycobacterium phage Magic8 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68865 | 66.373 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT776812 | Mycobacterium phage HarryOW | Mycobacterium phage HarryOW unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52935 | 63.774 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT776811 | Arthrobacter phage King2 | Arthrobacter phage King2 unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 63.232 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MT776810 | Gordonia phage BigChungus | Gordonia phage BigChungus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47166 | 60.703 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT776809 | Gordonia phage Buttrmlkdreams | Gordonia phage Buttrmlkdreams unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45999 | 60.397 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MT776808 | Gordonia phage Feastonyeet | Gordonia phage Feastonyeet Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47166 | 60.709 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT776807 | Microbacterium phage Majesty | Microbacterium phage Majesty Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17444 | 68.746 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT723945 | Mycobacterium phage Quasimodo | Mycobacterium phage Quasimodo unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156354 | 64.624 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT723944 | Mycobacterium phage ChaylaJr | Mycobacterium phage ChaylaJr unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154973 | 64.642 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT723943 | Mycobacterium phage Cactus | Mycobacterium phage Cactus unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75603 | 63.012 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MT723941 | Gordonia phage YorkOnyx | Gordonia phage YorkOnyx unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57430 | 68.247 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MT723938 | Gordonia phage Bunnybear | Gordonia phage Bunnybear unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51268 | 66.933 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MT723937 | Gordonia phage JKSyngboy | Gordonia phage JKSyngboy unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58176 | 67.985 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MT723936 | Gordonia phage Hitter | Gordonia phage Hitter Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47297 | 67.362 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT723935 | Gordonia phage Azula | Gordonia phage Azula Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53622 | 67.092 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT723934 | Gordonia phage Love | Gordonia phage Love unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60042 | 67.268 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MT723933 | Streptomyces phage Keanu | Streptomyces phage Keanu unclassified Hiyaavirus Hiyaavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82100 | 66.755 | Hiyaavirus | Unclassified | Siphoviridae | Streptomyces |
| MT658805 | Gordonia phage Rabbitrun | Gordonia phage Rabbitrun Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76821 | 58.79 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT658804 | Mycobacterium phage BengiVuitton | Mycobacterium phage BengiVuitton unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52965 | 63.414 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT658803 | Mycobacterium phage Reindeer | Mycobacterium phage Reindeer unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82356 | 60.315 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MT658802 | Mycobacterium phage Aziz | Mycobacterium phage Aziz unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83412 | 60.749 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MT639653 | Arthrobacter phage Elezi | Arthrobacter phage Elezi unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43471 | 66.55 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT639652 | Mycobacterium phage MasterPo | Mycobacterium phage MasterPo unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67361 | 68.923 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT639651 | Gordonia phage Portcullis | Gordonia phage Portcullis unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58364 | 67.759 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MT639649 | Microbacterium phage Burritobowl | Microbacterium phage Burritobowl unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41417 | 66.516 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT639648 | Mycobacterium phage Heath | Mycobacterium phage Heath unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69258 | 68.727 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT639647 | Microbacterium phage JRok | Microbacterium phage JRok Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17382 | 68.749 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT639646 | Microbacterium phage Rasputia | Microbacterium phage Rasputia unclassified Rogerhendrixvirus Rogerhendrixvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97105 | 66.31 | Rogerhendrixvirus | Unclassified | Siphoviridae | Microbacterium |
| MT639645 | Mycobacterium phage PainterBoy | Mycobacterium phage PainterBoy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52698 | 61.456 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT639643 | Microbacterium phage FreddieHg | Microbacterium phage FreddieHg unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56100 | 63.651 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT639642 | Microbacterium phage YertPhresh | Microbacterium phage YertPhresh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.512 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT639641 | Microbacterium phage Phinky | Microbacterium phage Phinky unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63064 | 67.357 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MT639640 | Gordonia phage Paries | Gordonia phage Paries unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60127 | 67.354 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MT522006 | Gordonia phage BBQValindra | Gordonia phage BBQValindra Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46875 | 67.334 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT522005 | Mycobacterium phage Misfit | Mycobacterium phage Misfit unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76169 | 63.055 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MT522003 | Microbacterium phage Rie18 | Microbacterium phage Rie18 unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56435 | 63.886 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT522001 | Microbacterium phage Karate | Microbacterium phage Karate unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56428 | 63.874 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT522000 | Mycobacterium phage Soul22 | Mycobacterium phage Soul22 unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54814 | 61.501 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| MT521998 | Gordonia phage Jambalaya | Gordonia phage Jambalaya unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57764 | 67.816 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MT521995 | Gordonia phage Moosehead | Gordonia phage Moosehead Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42671 | 65.065 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT521993 | Gordonia phage RedWattleHog | Gordonia phage RedWattleHog Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135372 | 64.937 | Unclassified | Unclassified | Myoviridae | Gordonia |
| MT521992 | Rhodococcus phage NiceHouse | Rhodococcus phage NiceHouse Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142586 | 44.936 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| MT521991 | Streptomyces phage Eklok | Streptomyces phage Eklok Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37409 | 71.224 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT521990 | Microbacterium phage Bri160 | Microbacterium phage Bri160 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17442 | 68.559 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498066 | Mycobacterium phage Georgie2 | Mycobacterium phage Georgie2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53399 | 63.526 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT498057 | Microbacterium phage Cicada | Microbacterium phage Cicada unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42350 | 66.519 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MT498067 | Microbacterium phage Bullzi2019 | Microbacterium phage Bullzi2019 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17143 | 68.815 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498065 | Microbacterium phage PoRanda | Microbacterium phage PoRanda Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17510 | 69.109 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498064 | Microbacterium phage Pabst | Microbacterium phage Pabst unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53620 | 58.995 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT498061 | Mycobacterium phage Jung | Mycobacterium phage Jung unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46561 | 67.078 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MT498060 | Mycobacterium Phage FiringLine | Mycobacterium Phage FiringLine unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53348 | 63.524 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT498059 | Gordonia phage Doggs | Gordonia phage Doggs Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45384 | 66.352 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT498058 | Gordonia phage Clawz | Gordonia phage Clawz Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57664 | 64.949 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT498055 | Arthrobacter phage Bumble | Arthrobacter phage Bumble Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49500 | 70.416 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| MT498054 | Gordonia Phage Engineer | Gordonia Phage Engineer unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93340 | 61.916 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MT498053 | Streptomyces phage PHTowN | Streptomyces phage PHTowN Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69271 | 64.264 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT498051 | Microbacterium phage Jahseh | Microbacterium phage Jahseh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17368 | 68.528 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498050 | Arthrobacter phage Gisselle | Arthrobacter phage Gisselle unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43089 | 60.735 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MT498047 | Microbacterium phage TinyTruffula | Microbacterium phage TinyTruffula Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17050 | 68.974 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498046 | Microbacterium phage Hernandez44 | Microbacterium phage Hernandez44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17477 | 68.822 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498045 | Arthrobacter phage MamaPearl | Arthrobacter phage MamaPearl unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43746 | 61.263 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MT498043 | Arthrobacter phage Jinkies | Arthrobacter phage Jinkies Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49118 | 64.687 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| MT498041 | Mycobacterium phage KingCyrus | Mycobacterium phage KingCyrus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53946 | 62.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT498040 | Microbacterium phage Livingwater | Microbacterium phage Livingwater Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17450 | 68.642 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498038 | Microbacterium phage Luxx | Microbacterium phage Luxx Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17463 | 68.791 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT498037 | Streptomyces phage Dryad | Streptomyces phage Dryad Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68994 | 63.846 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT498036 | Gordonia Phage Zitch | Gordonia Phage Zitch Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58928 | 67.71 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT498035 | Gordonia Phage Barsten | Gordonia Phage Barsten unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59501 | 67.666 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MT553345 | Mycobacterium phage LastJedi | Mycobacterium phage LastJedi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55149 | 61.755 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT553344 | Gordonia phage TZGordon | Gordonia phage TZGordon unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44904 | 66.094 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MT553342 | Microbacterium phage Kelcole | Microbacterium phage Kelcole unclassified Gillianvirus Gillianvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62812 | 67.161 | Gillianvirus | Unclassified | Siphoviridae | Microbacterium |
| MT553336 | Gordonia phage BlingBling | Gordonia phage BlingBling Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48746 | 67.148 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT553335 | Mycobacterium phage Ronan | Mycobacterium phage Ronan unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154852 | 64.606 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MT451982 | Streptomyces phage Cumberbatch | Streptomyces phage Cumberbatch Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37566 | 71.275 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT451981 | Streptomyces phage Vondra | Streptomyces phage Vondra Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37129 | 71.182 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT451980 | Streptomyces phage Ignacio | Streptomyces phage Ignacio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38137 | 71.12 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT310898 | Streptomyces phage Kela | Streptomyces phage Kela Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 185323 | 66.516 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT310896 | Microbacterium phage Lynlen | Microbacterium phage Lynlen unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41448 | 66.925 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310895 | Arthrobacter phage StevieBAY | Arthrobacter phage StevieBAY unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51319 | 60.866 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MT310894 | Microbacterium phage Tempo | Microbacterium phage Tempo unclassified Gillianvirus Gillianvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62869 | 67.03 | Gillianvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310893 | Mycobacterium phage DRBy19 | Mycobacterium phage DRBy19 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58368 | 61.292 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310892 | Mycobacterium phage Compostia | Mycobacterium phage Compostia unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69129 | 67.497 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT310891 | Microbacterium phage Pharky | Microbacterium phage Pharky unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54442 | 59.711 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310890 | Microbacterium phage Phedro | Microbacterium phage Phedro Microbacterium virus Phedro Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54442 | 59.713 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310889 | Microbacterium phage Phractured | Microbacterium phage Phractured unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54442 | 59.7 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310888 | Microbacterium phage Phogo | Microbacterium phage Phogo unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53959 | 60.036 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310887 | Microbacterium phage Unphazed | Microbacterium phage Unphazed unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53919 | 60.227 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310886 | Mycobacterium phage AN3 | Mycobacterium phage AN3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51037 | 63.405 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310884 | Mycobacterium phage ANI8 | Mycobacterium phage ANI8 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51141 | 63.098 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310877 | Mycobacterium phage VA6 | Mycobacterium phage VA6 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52608 | 63.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310875 | Microbacterium phage Ashton | Microbacterium phage Ashton unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54519 | 60.113 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310874 | Arthrobacter phage Boersma | Arthrobacter phage Boersma unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59188 | 52.948 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MT310872 | Gordonia phage Evamon | Gordonia phage Evamon unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58460 | 67.805 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MT310871 | Mycobacterium phage Jackstina | Mycobacterium phage Jackstina unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68546 | 67.506 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT310870 | Mycobacterium phage RawrgerThat | Mycobacterium phage RawrgerThat unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70841 | 68.874 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT310869 | Microbacterium phage Truong | Microbacterium phage Truong unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54309 | 60.075 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT310868 | Mycobacterium phage Telesworld | Mycobacterium phage Telesworld unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68458 | 66.509 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT310867 | Mycobacterium phage Chaelin | Mycobacterium phage Chaelin unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68618 | 66.478 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT310865 | Streptomyces phage Bmoc | Streptomyces phage Bmoc unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132885 | 49.639 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MT310864 | Microbacterium phage Nobel | Microbacterium phage Nobel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17049 | 68.989 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT310863 | Streptomyces phage Issmi | Streptomyces phage Issmi unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50643 | 67.565 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT310862 | Streptomyces phage Itza | Streptomyces phage Itza unclassified Arquatrovirinae Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50426 | 68.532 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT310858 | Mycobacterium phage Iwokeuplikedis | Mycobacterium phage Iwokeuplikedis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53261 | 62.746 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310857 | Mycobacterium phage Danforth | Mycobacterium phage Danforth unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52651 | 61.349 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310854 | Streptomyces phage CricKo | Streptomyces phage CricKo unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57623 | 58.072 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MT310853 | Microbacterium phage AvGardian | Microbacterium phage AvGardian unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41522 | 68.658 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310852 | Streptomyces phage ClubPenguin | Streptomyces phage ClubPenguin unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56205 | 59.046 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MT310850 | Gordonia phage Secretariat | Gordonia phage Secretariat Gordonia virus Secretariat Secretariatvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57731 | 52.816 | Secretariatvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MT316460 | Mycobacterium phage Kimbrough | Mycobacterium phage Kimbrough unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68910 | 66.529 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT316463 | Mycobacterium phage Slatt | Mycobacterium phage Slatt unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68983 | 66.403 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT316461 | Streptomyces phage Galactica | Streptomyces phage Galactica unclassified Hiyaavirus Hiyaavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81216 | 66.935 | Hiyaavirus | Unclassified | Siphoviridae | Streptomyces |
| MT316462 | Microbacterium phage Danno | Microbacterium phage Danno Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17452 | 68.737 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT316459 | Microbacterium phage Otwor | Microbacterium phage Otwor Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17454 | 68.735 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MT316458 | Microbacterium phage McShie | Microbacterium phage McShie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17434 | 68.94 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271317 | Corynebacterium phage TouchMeNot | Corynebacterium phage TouchMeNot unclassified Poushouvirus Poushouvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40526 | 60.031 | Poushouvirus | Unclassified | Siphoviridae | Corynebacterium |
| MF197383 | Corynebacterium phage Poushou | Corynebacterium phage Poushou Corynebacterium virus Poushou Poushouvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40353 | 60.008 | Poushouvirus | Unclassified | Siphoviridae | Corynebacterium |
| MT114167 | Mycobacterium phage Phanphagia | Mycobacterium phage Phanphagia unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59699 | 61.55 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT114166 | Microbacterium phage Nike | Microbacterium phage Nike unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62393 | 66.663 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MT114165 | Mycobacterium phage BadStone | Mycobacterium phage BadStone unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75679 | 63.053 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MT114164 | Mycobacterium phage Veteran | Mycobacterium phage Veteran unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56147 | 61.499 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT114163 | Mycobacterium phage Settecandela | Mycobacterium phage Settecandela Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145208 | 67.447 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| MT114162 | Streptomyces phage Moab | Streptomyces phage Moab unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129370 | 47.202 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MT114161 | Arthrobacter phage JKerns | Arthrobacter phage JKerns unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50789 | 60.999 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MT114160 | Gordonia phage Breezic | Gordonia phage Breezic unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76057 | 58.899 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MT114159 | Mycobacterium phage Ein37 | Mycobacterium phage Ein37 unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53748 | 67.446 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT024872 | Streptomyces phage Muntaha | Streptomyces phage Muntaha unclassified Wilnyevirus Wilnyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127551 | 52.577 | Wilnyevirus | Unclassified | Siphoviridae | Streptomyces |
| MT024871 | Arthrobacter phage YesChef | Arthrobacter phage YesChef unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43510 | 67.672 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT024870 | Arthrobacter phage Whytu | Arthrobacter phage Whytu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15369 | 64.76 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT024869 | Arthrobacter phage Lego | Arthrobacter phage Lego unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43446 | 67.539 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MT024868 | Arthrobacter phage Abba | Arthrobacter phage Abba Arthrobacter virus Abba Ayohtrevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46623 | 64.957 | Ayohtrevirus | Unclassified | Myoviridae | Arthrobacter |
| MT024866 | Mycobacterium phage Willsammy | Mycobacterium phage Willsammy unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48399 | 67.026 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MT024865 | Streptomyces phage Wakanda | Streptomyces phage Wakanda unclassified Wilnyevirus Wilnyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126614 | 52.571 | Wilnyevirus | Unclassified | Siphoviridae | Streptomyces |
| MT024863 | Microbacterium phage Arete | Microbacterium phage Arete Microbacterium virus Arete Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53692 | 64.609 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| MT024861 | Mycobacterium phage Pmask | Mycobacterium phage Pmask unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52576 | 61.562 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT024860 | Microbacterium phage Stromboli | Microbacterium phage Stromboli unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41594 | 68.801 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MK880124 | Microbacterium phage IAmGroot | Microbacterium phage IAmGroot Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45625 | 70.005 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN945903 | Mycobacterium phage Jiminy | Mycobacterium phage Jiminy unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68777 | 66.423 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN945900 | Mycobacterium phage BodEinwohner17 | Mycobacterium phage BodEinwohner17 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60222 | 61.343 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN945899 | Mycobacterium phage Skippy | Mycobacterium phage Skippy unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68726 | 66.492 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN908694 | Microbacterium phage RubyRalph | Microbacterium phage RubyRalph Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61880 | 68.979 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN908693 | Gordonia phage AnClar | Gordonia phage AnClar unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61856 | 69.823 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MN908691 | Gordonia phage DatBoi | Gordonia phage DatBoi unclassified Bantamvirus Bantamvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92498 | 65.254 | Bantamvirus | Unclassified | Siphoviridae | Gordonia |
| MN908689 | Arthrobacter phage DrSierra | Arthrobacter phage DrSierra unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42622 | 66.466 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN908687 | Gordonia phage Skog | Gordonia phage Skog Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152435 | 65.764 | Unclassified | Unclassified | Myoviridae | Gordonia |
| MN908685 | Microbacterium phage PauloDiaboli | Microbacterium phage PauloDiaboli Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191968 | 60.088 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN908684 | Arthrobacter phage Shoya | Arthrobacter phage Shoya Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37783 | 63.722 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN908683 | Mycobacterium phage StressBall | Mycobacterium phage StressBall unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47915 | 67.282 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN892487 | Mycobacterium phage Mabel | Mycobacterium phage Mabel unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52577 | 63.737 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN892486 | Mycobacterium phage KilKor | Mycobacterium phage KilKor unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48916 | 67.209 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN892484 | Mycobacterium phage CactusJack | Mycobacterium phage CactusJack unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48222 | 67.285 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN735434 | Mycobacterium phage Mangeria | Mycobacterium phage Mangeria unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156170 | 64.833 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN735433 | Mycobacterium phage Scottish | Mycobacterium phage Scottish unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56825 | 61.454 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN735429 | Arthrobacter phage AppleCider | Arthrobacter phage AppleCider unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43699 | 61.12 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MN807250 | Mycobacterium phage Glaske | Mycobacterium phage Glaske unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48222 | 67.287 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN807249 | Mycobacterium phage Megiddo | Mycobacterium phage Megiddo unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48783 | 67.12 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN807248 | Mycobacterium phage Phalm | Mycobacterium phage Phalm unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48213 | 67.285 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN813700 | Mycobacterium phage Atkinbua | Mycobacterium phage Atkinbua unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52955 | 63.312 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN813693 | Mycobacterium phage Imvubu | Mycobacterium phage Imvubu unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69897 | 70.497 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN813686 | Mycobacterium phage BirdsNest | Mycobacterium phage BirdsNest unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70668 | 70.081 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN813684 | Microbacterium phage Terij | Microbacterium phage Terij unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55691 | 70.159 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MN813682 | Microbacterium phage Matzah | Microbacterium phage Matzah unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55686 | 69.863 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MN813680 | Arthrobacter phage LilStuart | Arthrobacter phage LilStuart unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42950 | 60.738 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MN813678 | Mycobacterium phage Lephleur | Mycobacterium phage Lephleur unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67349 | 68.942 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN813676 | Arthrobacter phage Adolin | Arthrobacter phage Adolin unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43022 | 65.862 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN703415 | Mycobacterium phage Mcshane | Mycobacterium phage Mcshane unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68929 | 66.402 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN703414 | Microbacterium phage Dewdrop | Microbacterium phage Dewdrop unclassified Rogerhendrixvirus Rogerhendrixvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95409 | 67.108 | Rogerhendrixvirus | Unclassified | Siphoviridae | Microbacterium |
| MN703413 | Arthrobacter phage Powerpuff | Arthrobacter phage Powerpuff unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44651 | 67.647 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN703412 | Arthrobacter phage Giantsbane | Arthrobacter phage Giantsbane unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56734 | 50.078 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN703409 | Arthrobacter phage LittleTokyo | Arthrobacter phage LittleTokyo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38969 | 66.009 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN703408 | Mycobacterium phage Phaded | Mycobacterium phage Phaded unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53598 | 63.118 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN703407 | Microbacterium phage Leaf | Microbacterium phage Leaf unclassified Rogerhendrixvirus Rogerhendrixvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95340 | 67.108 | Rogerhendrixvirus | Unclassified | Siphoviridae | Microbacterium |
| MN703405 | Mycobacterium phage Aneem | Mycobacterium phage Aneem unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52595 | 63.687 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN728683 | Mycobacterium phage Yokurt | Mycobacterium phage Yokurt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52123 | 61.512 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK814760 | Gordonia phage Forza | Gordonia phage Forza Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114174 | 53.209 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN585990 | Microbacterium phage Azizam | Microbacterium phage Azizam Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17385 | 68.749 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN586061 | Arthrobacter phage Maria1952 | Arthrobacter phage Maria1952 unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43949 | 61.863 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MN586059 | Microbacterium phage Teamocil | Microbacterium phage Teamocil unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53496 | 68.633 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MN586058 | Mycobacterium phage Vaishali24 | Mycobacterium phage Vaishali24 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68398 | 66.636 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN586057 | Gordonia phage Cleo | Gordonia phage Cleo unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44795 | 60.601 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN586055 | Gordonia phage Cynthia | Gordonia phage Cynthia unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46219 | 66.633 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MN586053 | Arthrobacter phage BeatusComedenti | Arthrobacter phage BeatusComedenti unclassified Kelleziovirus Kelleziovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58687 | 63.36 | Kelleziovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN586049 | Gordonia phage Yarn | Gordonia phage Yarn unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44619 | 59.694 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN586047 | Gordonia phage EMoore | Gordonia phage EMoore Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58844 | 68.031 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN586046 | Mycobacterium phage Tinciduntsolum | Mycobacterium phage Tinciduntsolum unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67496 | 68.958 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN586045 | Mycobacterium phage Brownie5 | Mycobacterium phage Brownie5 unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67497 | 68.957 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN586042 | Microbacterium phage Jayden | Microbacterium phage Jayden unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53352 | 68.288 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MN586040 | Gordonia phage Stormageddon | Gordonia phage Stormageddon Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136319 | 65.036 | Unclassified | Unclassified | Myoviridae | Gordonia |
| MN586039 | Mycobacterium phage Blinn1 | Mycobacterium phage Blinn1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51800 | 61.222 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586038 | Mycobacterium phage WalterMcMickey | Mycobacterium phage WalterMcMickey unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51094 | 64.959 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586036 | Gordonia phage AndPeggy | Gordonia phage AndPeggy unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 59.73 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN586033 | Corynebacterium phage EmiRose | Corynebacterium phage EmiRose Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37429 | 56.403 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| MN586028 | Gordonia phage MelBins | Gordonia phage MelBins Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59761 | 68.337 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN586027 | Arthrobacter phage Mufasa8 | Arthrobacter phage Mufasa8 Arthrobacter virus Mufasa8 Mufasoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60205 | 66.136 | Mufasoctovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN586026 | Gordonia phage Leonard | Gordonia phage Leonard Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59268 | 68.435 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN586023 | Mycobacterium phage Zaria | Mycobacterium phage Zaria unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74365 | 58.737 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586022 | Gordonia phage Chidiebere | Gordonia phage Chidiebere unclassified Bendigovirus Bendigovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91072 | 60.196 | Bendigovirus | Unclassified | Myoviridae | Gordonia |
| MN586021 | Arthrobacter phage Beagle | Arthrobacter phage Beagle Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70671 | 65.72 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN586020 | Microbacterium phage Megan | Microbacterium phage Megan Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55210 | 69.641 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN586018 | Mycobacterium phage Wyatt2 | Mycobacterium phage Wyatt2 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74010 | 58.743 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586015 | Gordonia phage Lauer | Gordonia phage Lauer Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48123 | 60.133 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN586012 | Mycobacterium phage Blexus | Mycobacterium phage Blexus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58824 | 61.432 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586011 | Mycobacterium phage LilMoolah | Mycobacterium phage LilMoolah unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58136 | 61.454 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586010 | Microbacterium phage HarperAnne | Microbacterium phage HarperAnne Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17116 | 68.848 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN586009 | Streptomyces phage Phettuccine | Streptomyces phage Phettuccine unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49530 | 66.21 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN586008 | Microbacterium phage Scrunchy | Microbacterium phage Scrunchy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.759 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN586007 | Microbacterium phage Smarties | Microbacterium phage Smarties unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54344 | 68.444 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MN586006 | Mycobacterium phage Bachome | Mycobacterium phage Bachome unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51653 | 63.708 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586004 | Mycobacterium phage Strokeseat | Mycobacterium phage Strokeseat unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55470 | 61.439 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586003 | Gordonia phage Yikes | Gordonia phage Yikes Gordonia virus Yikes Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68155 | 58.222 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN586001 | Mycobacterium phage Donkeykong | Mycobacterium phage Donkeykong unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59478 | 61.863 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586000 | Mycobacterium phage Sagefire | Mycobacterium phage Sagefire unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51482 | 63.877 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585999 | Mycobacterium phage Briton15 | Mycobacterium phage Briton15 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53032 | 63.697 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585998 | Mycobacterium phage Bugsy | Mycobacterium phage Bugsy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49937 | 63.186 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585997 | Mycobacterium phage Traft412 | Mycobacterium phage Traft412 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51223 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585996 | Gordonia phage Crocheter | Gordonia phage Crocheter Gordonia virus Crocheter Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60531 | 51.623 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN585995 | Mycobacterium phage Chuckly | Mycobacterium phage Chuckly unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55189 | 61.559 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585994 | Mycobacterium phage SheaKeira | Mycobacterium phage SheaKeira unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52431 | 61.986 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585993 | Mycobacterium phage Indlulamithi | Mycobacterium phage Indlulamithi Mycobacterium virus Indlulamithi Indlulamithivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71538 | 52.689 | Indlulamithivirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585992 | Mycobacterium phage MaryV | Mycobacterium phage MaryV unclassified Wildcatvirus Wildcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76413 | 56.875 | Wildcatvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585991 | Mycobacterium phage Snazzy | Mycobacterium phage Snazzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52220 | 63.61 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585989 | Mycobacterium phage Superchunk | Mycobacterium phage Superchunk unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52772 | 62.433 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585988 | Gordonia Phage Odesza | Gordonia Phage Odesza unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59746 | 51.113 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN585986 | Microbacterium phage Ariadne | Microbacterium phage Ariadne unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54347 | 68.444 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MN585985 | Mycobacterium phage Watermelon | Mycobacterium phage Watermelon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50965 | 63.518 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585983 | Mycobacterium phage Retro23 | Mycobacterium phage Retro23 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53044 | 63.423 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585982 | Gordonia phage Untouchable | Gordonia phage Untouchable Gordonia virus Untouchable Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61623 | 51.552 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN585981 | Mycobacterium phage QueenB2 | Mycobacterium phage QueenB2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52732 | 63.4 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585980 | Microbacterium phage Gina | Microbacterium phage Gina unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53403 | 68.629 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MN585979 | Mycobacterium phage Yecey3 | Mycobacterium phage Yecey3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53277 | 62.862 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585978 | Mycobacterium phage Beemo | Mycobacterium phage Beemo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53220 | 62.58 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585977 | Mycobacterium phage Atcoo | Mycobacterium phage Atcoo unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49075 | 67.016 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN585976 | Mycobacterium phage Kanely | Mycobacterium phage Kanely unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52539 | 63.545 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585975 | Mycobacterium phage Eish | Mycobacterium phage Eish unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58150 | 61.214 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585974 | Mycobacterium phage DreamTeam1 | Mycobacterium phage DreamTeam1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52556 | 61.921 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585973 | Mycobacterium phage StAnnes | Mycobacterium phage StAnnes unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59357 | 61.802 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585972 | Arthrobacter phage Edmundo | Arthrobacter phage Edmundo unclassified Laroyevirus Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60191 | 64.555 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| MN585971 | Gordonia phage Avazak | Gordonia phage Avazak Gordonia virus Avazak Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60278 | 51.45 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN585970 | Mycobacterium phage Killigrew | Mycobacterium phage Killigrew unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51163 | 63.775 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585968 | Mycobacterium phage Leogania | Mycobacterium phage Leogania unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49006 | 63.368 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585967 | Mycobacterium phage Kloppinator | Mycobacterium phage Kloppinator unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68480 | 66.503 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN585966 | Gordonia phage Faith5x5 | Gordonia phage Faith5x5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41982 | 65.116 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN585965 | Mycobacterium phage Duggie | Mycobacterium phage Duggie unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68885 | 66.377 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN585964 | Mycobacterium phage Blessica | Mycobacterium phage Blessica unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71470 | 65.444 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369765 | Microbacterium phage Zanella | Microbacterium phage Zanella unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42108 | 67.013 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MN369764 | Mycobacterium phage Rahalelujah | Mycobacterium phage Rahalelujah unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52816 | 62.4 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369763 | Mycobacterium phage Aliter | Mycobacterium phage Aliter unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50480 | 62.716 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369760 | Mycobacterium phage KyMonks1A | Mycobacterium phage KyMonks1A unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52145 | 63.619 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369759 | Mycobacterium phage Mithril | Mycobacterium phage Mithril unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70937 | 68.954 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN369758 | Gordonia phage NHagos | Gordonia phage NHagos unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59580 | 68.216 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MN369757 | Streptomyces phage Bordeaux | Streptomyces phage Bordeaux unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131317 | 49.489 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MN369755 | Mycobacterium phage Minnie | Mycobacterium phage Minnie unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59268 | 61.485 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369754 | Streptomyces phage Limpid | Streptomyces phage Limpid unclassified Annadreamyvirus Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131038 | 47.556 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MN369750 | Streptomyces phage TomSawyer | Streptomyces phage TomSawyer unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133961 | 49.274 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MN369749 | Mycobacterium phage MinionDave | Mycobacterium phage MinionDave unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58027 | 61.496 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369748 | Mycobacterium phage Miko | Mycobacterium phage Miko unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51881 | 63.534 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369745 | Microbacterium phage Kaijohn | Microbacterium phage Kaijohn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17426 | 68.759 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN369744 | Mycobacterium phage Beaglebox | Mycobacterium phage Beaglebox unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68418 | 66.528 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN369743 | Streptomyces phage Tribute | Streptomyces phage Tribute unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133471 | 50.027 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MN369741 | Mycobacterium phage LoneWolf | Mycobacterium phage LoneWolf unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53186 | 63.002 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369738 | Mycobacterium phage Hocus | Mycobacterium phage Hocus unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68083 | 66.372 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN444878 | Gordonia phage Quasar | Gordonia phage Quasar unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46960 | 60.019 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN444877 | Microbacterium phage Antares | Microbacterium phage Antares unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53362 | 68.888 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MN444876 | Streptomyces phage Daubenski | Streptomyces phage Daubenski unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133090 | 49.397 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MN444875 | Mycobacterium phage Jonghyun | Mycobacterium phage Jonghyun unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41904 | 66.636 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MN444873 | Mycobacterium phage Abinghost | Mycobacterium phage Abinghost unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68634 | 67.558 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN444872 | Mycobacterium phage SynergyX | Mycobacterium phage SynergyX unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68980 | 67.531 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN444871 | Gordonia phage Axym | Gordonia phage Axym unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46473 | 60.175 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN444870 | Gordonia phage Syleon | Gordonia phage Syleon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75606 | 58.735 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN444869 | Mycobacterium phage DreamCatcher | Mycobacterium phage DreamCatcher unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53408 | 63.358 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN444868 | Mycobacterium phage Prickles | Mycobacterium phage Prickles unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68454 | 66.445 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN497954 | Microbacterium phage Hiddenleaf | Microbacterium phage Hiddenleaf unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56082 | 63.664 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MN617845 | Mycobacterium phage BaconJack | Mycobacterium phage BaconJack unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53793 | 63.759 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN617844 | Mycobacterium phage Bluefalcon | Mycobacterium phage Bluefalcon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51052 | 60.854 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN617843 | Mycobacterium phage Quesadilla | Mycobacterium phage Quesadilla unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70992 | 69.86 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN617841 | Mycobacterium phage DirkDirk | Mycobacterium phage DirkDirk unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68021 | 58.673 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN484600 | Arthrobacter phage Kuleana | Arthrobacter phage Kuleana Arthrobacter virus Kuleana Kuleanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38850 | 65.833 | Kuleanavirus | Unclassified | Siphoviridae | Arthrobacter |
| MN484599 | Streptomyces phage Wipeout | Streptomyces phage Wipeout unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132935 | 49.344 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MN484601 | Gordonia phage Sixama | Gordonia phage Sixama Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115454 | 52.691 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN428066 | Mycobacterium phage JPickles | Mycobacterium phage JPickles unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155116 | 64.674 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN428065 | Mycobacterium phage Cactojaque | Mycobacterium phage Cactojaque unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154918 | 64.625 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN428064 | Microbacterium phage Owens | Microbacterium phage Owens Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.759 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN428062 | Microbacterium phage TeddyBoy | Microbacterium phage TeddyBoy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17449 | 68.76 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN428061 | Mycobacterium phage Corazon | Mycobacterium phage Corazon unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64931 | 63.409 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428060 | Streptomyces phage IchabodCrane | Streptomyces phage IchabodCrane unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130730 | 49.417 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MN428058 | Mycobacterium phage Krili | Mycobacterium phage Krili unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71670 | 65.366 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428056 | Mycobacterium phage Pringar | Mycobacterium phage Pringar unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64817 | 63.372 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428054 | Mycobacterium phage JackSparrow | Mycobacterium phage JackSparrow unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51576 | 63.609 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428053 | Gordonia phage Denise | Gordonia phage Denise Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45043 | 65.415 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN428052 | Mycobacterium phage Smooch | Mycobacterium phage Smooch unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71415 | 65.409 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428051 | Mycobacterium phage Modragons | Mycobacterium phage Modragons unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58461 | 61.104 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428050 | Mycobacterium phage Apex | Mycobacterium phage Apex unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71244 | 68.852 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN428049 | Mycobacterium phage West99 | Mycobacterium phage West99 unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67516 | 68.924 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN428047 | Mycobacterium phage Doomphist | Mycobacterium phage Doomphist unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58666 | 61.489 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN284908 | Mycobacterium phage Helpful | Mycobacterium phage Helpful Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64794 | 59.584 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MN284907 | Gordonia phage Fireball | Gordonia phage Fireball unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58924 | 67.753 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MN284906 | Gordonia phage CherryonLim | Gordonia phage CherryonLim Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48948 | 60.229 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN284904 | Gordonia phage Tanis | Gordonia phage Tanis Gordonia virus Tanis Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59727 | 51.009 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN284903 | Mycobacterium phage GaugeLDP | Mycobacterium phage GaugeLDP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53035 | 63.375 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN284899 | Streptomyces phage Whatever | Streptomyces phage Whatever unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51591 | 65.688 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN284898 | Gordonia phage MinecraftSteve | Gordonia phage MinecraftSteve unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52549 | 61.99 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MN284896 | Gordonia phage OhMyWard | Gordonia phage OhMyWard Gordonia virus OhMyWard Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60978 | 52.153 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN284893 | Mycobacterium phage LilMcDreamy | Mycobacterium phage LilMcDreamy unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71552 | 69.347 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN284892 | Microbacterium phage Bradley2 | Microbacterium phage Bradley2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17367 | 68.509 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN329679 | Microbacterium phage Vanisius | Microbacterium phage Vanisius Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17453 | 68.727 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN329677 | Gordonia phage Keitabear | Gordonia phage Keitabear unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59759 | 67.299 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MN329676 | Mycobacterium phage Poise | Mycobacterium phage Poise unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64824 | 63.385 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN329675 | Microbacterium phage SansAfet | Microbacterium phage SansAfet unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41531 | 66.716 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MN329674 | Mycobacterium phage JoieB | Mycobacterium phage JoieB unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65404 | 63.377 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN329673 | Gordonia phage Jabberwocky | Gordonia phage Jabberwocky unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59449 | 67.592 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MN234233 | Corynebacterium phage Colleen | Corynebacterium phage Colleen unclassified Poushouvirus Poushouvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40524 | 60.031 | Poushouvirus | Unclassified | Siphoviridae | Corynebacterium |
| MN234232 | Gordonia phage Sadboi | Gordonia phage Sadboi Gordonia virus Sadboi Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67485 | 58.078 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN234228 | Mycobacterium phage Tourach | Mycobacterium phage Tourach unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77876 | 58.755 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234227 | Mycobacterium phage Giuseppe | Mycobacterium phage Giuseppe Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64604 | 59.639 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MN234225 | Mycobacterium phage Rhinoforte | Mycobacterium phage Rhinoforte unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67411 | 68.926 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN234224 | Arthrobacter phage Jaek | Arthrobacter phage Jaek unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59173 | 52.948 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234223 | Mycobacterium phage Philly | Mycobacterium phage Philly unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68523 | 67.542 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN234221 | Arthrobacter phage Shepard | Arthrobacter phage Shepard unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57272 | 50.155 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234220 | Gordonia phage Lambo | Gordonia phage Lambo Gordonia virus Lambo Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67813 | 58.296 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN234217 | Arthrobacter phage Ichor | Arthrobacter phage Ichor unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59173 | 52.948 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234216 | Streptomyces phage Gilgamesh | Streptomyces phage Gilgamesh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129136 | 71.331 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MN234215 | Gordonia phage SteamedHams | Gordonia phage SteamedHams unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44571 | 59.873 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN234214 | Arthrobacter phage Amelia | Arthrobacter phage Amelia unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38128 | 66.636 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234212 | Mycobacterium phage Paphu | Mycobacterium phage Paphu unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51159 | 64.122 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234211 | Arthrobacter phage Mimi | Arthrobacter phage Mimi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172660 | 53.107 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234208 | Streptomyces phage Alvy | Streptomyces phage Alvy unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53011 | 67.548 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN234207 | Gordonia phage Ranch | Gordonia phage Ranch Gordonia virus Ranch Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66892 | 58.346 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN234206 | Arthrobacter phage Racecar | Arthrobacter phage Racecar Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173709 | 53.075 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234203 | Mycobacterium phage SemperFi | Mycobacterium phage SemperFi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53235 | 63.443 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234196 | Gordonia phage CloverMinnie | Gordonia phage CloverMinnie unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61098 | 68.67 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MN234193 | Mycobacterium phage Kboogie | Mycobacterium phage Kboogie unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155680 | 64.695 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN234192 | Streptomyces phage Animus | Streptomyces phage Animus unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50793 | 67.021 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN234191 | Microbacterium phage Naby | Microbacterium phage Naby Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17448 | 68.185 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN234190 | Gordonia phage Gibbin | Gordonia phage Gibbin Gordonia virus Gibbin Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67785 | 58.267 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN234189 | Streptomyces phage Urza | Streptomyces phage Urza unclassified Arquatrovirinae Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50448 | 68.528 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN234187 | Mycobacterium phage DyoEdafos | Mycobacterium phage DyoEdafos unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77566 | 58.539 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234184 | Mycobacterium phage IdentityCrisis | Mycobacterium phage IdentityCrisis Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38341 | 65.032 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN234180 | Arthrobacter phage Shiba | Arthrobacter phage Shiba Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54204 | 51.67 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234178 | Arthrobacter phage TripleJ | Arthrobacter phage TripleJ Arthrobacter virus TripleJ Triplejayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45507 | 64.504 | Triplejayvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234172 | Mycobacterium phage ThulaThula | Mycobacterium phage ThulaThula unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50415 | 66.53 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN234171 | Mycobacterium phage Magpie | Mycobacterium phage Magpie unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71014 | 68.996 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN234169 | Mycobacterium phage Naji | Mycobacterium phage Naji unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49127 | 63.541 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234164 | Arthrobacter phage Yeezus | Arthrobacter phage Yeezus unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59171 | 52.95 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234162 | Gordonia phage GretelLyn | Gordonia phage GretelLyn unclassified Lambovirus Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67605 | 58.077 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| MN224566 | Mycobacterium phage Herbertwm | Mycobacterium phage Herbertwm unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51546 | 62.598 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN276192 | Mycobacterium phage Trinitium | Mycobacterium phage Trinitium unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154512 | 64.665 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN310549 | Gordonia phage Gibbous | Gordonia phage Gibbous unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45810 | 60.537 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MN310547 | Mycobacterium phage Hutc2 | Mycobacterium phage Hutc2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51393 | 63.777 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN310546 | Microbacterium phage PhriedRice | Microbacterium phage PhriedRice unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54546 | 59.682 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MN310545 | Mycobacterium phage Nitzel | Mycobacterium phage Nitzel unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54857 | 61.316 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN310541 | Mycobacterium phage Jeeves | Mycobacterium phage Jeeves unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52466 | 62.116 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN204504 | Streptomyces phage Soshi | Streptomyces phage Soshi unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56311 | 59.466 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN204502 | Mycobacterium phage KingMidas | Mycobacterium phage KingMidas unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57386 | 61.407 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN204501 | Gordonia phage Chikenjars | Gordonia phage Chikenjars Gordonia virus Chikenjars Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61544 | 51.334 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN204500 | Streptomyces phage Hoshi | Streptomyces phage Hoshi unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55728 | 59.475 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN204498 | Streptomyces phage Saftant | Streptomyces phage Saftant unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48883 | 65.642 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN204497 | Streptomyces phage Meibysrarus | Streptomyces phage Meibysrarus unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55097 | 59.504 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN204496 | Streptomyces phage Kardashian | Streptomyces phage Kardashian unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55961 | 59.777 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MN204495 | Streptomyces phage CherryBlossom | Streptomyces phage CherryBlossom unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55563 | 59.558 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN204494 | Streptomyces phage Jaylociraptor | Streptomyces phage Jaylociraptor unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55699 | 59.488 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN204493 | Streptomyces phage Zuko | Streptomyces phage Zuko Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82302 | 63.73 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MN204492 | Streptomyces phage GirlPower | Streptomyces phage GirlPower unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55521 | 59.097 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MK894436 | Gordonia phage Boopy | Gordonia phage Boopy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114220 | 53.212 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN183286 | Mycobacterium phage Yeet | Mycobacterium phage Yeet unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111437 | 60.818 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN183285 | Mycobacterium phage YoungMoneyMata | Mycobacterium phage YoungMoneyMata unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156223 | 64.658 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN183284 | Microbacterium phage Mashley | Microbacterium phage Mashley unclassified Squashvirus Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61917 | 66.999 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MN183283 | Arthrobacter phage BossLady | Arthrobacter phage BossLady unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51178 | 60.985 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MN183282 | Arthrobacter phage Qui | Arthrobacter phage Qui Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113685 | 47.614 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN183281 | Microbacterium phage Vitas | Microbacterium phage Vitas unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39994 | 67.1 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MN183280 | Mycobacterium phage LolaVinca | Mycobacterium phage LolaVinca unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156412 | 64.649 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN183279 | Arthrobacter phage Zartrosa | Arthrobacter phage Zartrosa Arthrobacter virus Zartrosa Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50782 | 60.953 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MN119379 | Mycobacterium phage Anglerfish | Mycobacterium phage Anglerfish unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52380 | 63.477 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN119378 | Mycobacterium phage Penelope2018 | Mycobacterium phage Penelope2018 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64554 | 59.691 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MN119377 | Mycobacterium phage Buttons | Mycobacterium phage Buttons unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49420 | 63.667 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062720 | Microbacterium phage FuzzBuster | Microbacterium phage FuzzBuster Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54844 | 68.037 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN062719 | Microbacterium phage PiperSansNom | Microbacterium phage PiperSansNom unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53277 | 68.75 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MN062717 | Microbacterium phage Sharkboy | Microbacterium phage Sharkboy unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41947 | 69.662 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MN062716 | Gordonia phage Mayweather | Gordonia phage Mayweather Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48382 | 60.626 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN062714 | Microbacterium Phage DirtyBubble | Microbacterium Phage DirtyBubble unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41692 | 68.673 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MN062712 | Mycobacterium phage Pocahontas | Mycobacterium phage Pocahontas unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50077 | 64.049 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062711 | Microbacterium phage Araxxi | Microbacterium phage Araxxi Microbacterium virus Araxxi Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53981 | 64.836 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| MN062710 | Mycobacterium phage Ajay | Mycobacterium phage Ajay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49516 | 63.638 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062708 | Mycobacterium phage NothingSpecial | Mycobacterium phage NothingSpecial unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52441 | 63.195 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062707 | Mycobacterium phage Grungle | Mycobacterium phage Grungle unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153739 | 64.611 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MN062706 | Mycobacterium phage Heathen | Mycobacterium phage Heathen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50143 | 63.979 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062705 | Streptomyces phage Celia | Streptomyces phage Celia unclassified Arquatrovirinae Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50493 | 68.552 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN062704 | Gordonia phage JuJu | Gordonia phage JuJu Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54000 | 66.904 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN062701 | Mycobacterium phage Dallas | Mycobacterium phage Dallas unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111843 | 60.755 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN175604 | Gordonia phage PhorbesPhlower | Gordonia phage PhorbesPhlower Luckytenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41486 | 65.08 | Luckytenvirus | Unclassified | Siphoviridae | Gordonia |
| MN175603 | Microbacterium phage Tyrumbra | Microbacterium phage Tyrumbra unclassified Metamorphoovirus Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53975 | 68.663 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MK937610 | Microbacterium phage Arroyo | Microbacterium phage Arroyo unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42129 | 66.607 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MK967402 | Mycobacterium phage NihilNomen | Mycobacterium phage NihilNomen unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110439 | 60.82 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967401 | Mycobacterium phage Carlyle | Mycobacterium phage Carlyle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51220 | 63.647 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967396 | Gordonia phage Tangerine | Gordonia phage Tangerine unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57306 | 68.244 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MK967395 | Mycobacterium phage WideWale | Mycobacterium phage WideWale unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53040 | 63.503 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967393 | Rhodococcus phage Whack | Rhodococcus phage Whack Rhodococcus virus Whack Whackvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49660 | 61.911 | Whackvirus | Unclassified | Siphoviridae | Rhodococcus |
| MK967392 | Gordonia phage GrandSlam | Gordonia phage GrandSlam unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50919 | 67.032 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MK967389 | Gordonia phage JasperJr | Gordonia phage JasperJr Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 67.188 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK967387 | Mycobacterium phage Big3 | Mycobacterium phage Big3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53442 | 63.439 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967385 | Gordonia phage SheckWes | Gordonia phage SheckWes Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47618 | 60.761 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK967384 | Gordonia phage Sidious | Gordonia phage Sidious Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51789 | 66.599 | Unclassified | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK967383 | Gordonia phage SweatNTears | Gordonia phage SweatNTears unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45917 | 59.93 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MK967382 | Gordonia phage ReMo | Gordonia phage ReMo unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52601 | 61.94 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MK967381 | Gordonia phage RogerDodger | Gordonia phage RogerDodger unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59122 | 67.858 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK967378 | Gordonia phage Trax | Gordonia phage Trax Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75943 | 58.706 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK967379 | Mycobacterium phage HokkenD | Mycobacterium phage HokkenD unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113067 | 60.791 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096382 | Mycobacterium phage AvadaKedavra | Mycobacterium phage AvadaKedavra unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73721 | 58.713 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096381 | Streptomyces phage Rusticus | Streptomyces phage Rusticus unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50066 | 65.927 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN096380 | Mycobacterium phage Lewan | Mycobacterium phage Lewan unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76734 | 59.038 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096379 | Streptomyces phage Yasdnil | Streptomyces phage Yasdnil unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51631 | 65.699 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN096375 | Streptomyces phage Leviticus | Streptomyces phage Leviticus unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50095 | 66.17 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN096373 | Streptomyces phage TuanPN | Streptomyces phage TuanPN unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51517 | 65.685 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MN096372 | Mycobacterium phage Zolita | Mycobacterium phage Zolita unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51182 | 60.853 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096371 | Streptomyces phage Braelyn | Streptomyces phage Braelyn unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131234 | 50.052 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MN096369 | Gordonia phage Phendrix | Gordonia phage Phendrix unclassified Godonkavirus Godonkavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121350 | 54.613 | Godonkavirus | Unclassified | Siphoviridae | Gordonia |
| MN096366 | Mycobacterium phage AbsoluteMadLad | Mycobacterium phage AbsoluteMadLad unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68803 | 66.504 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN096365 | Gordonia phage Samba | Gordonia phage Samba Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51500 | 67.243 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN096362 | Arthrobacter phage Vibaki | Arthrobacter phage Vibaki Arthrobacter virus Vibaki Vibakivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48201 | 66.7 | Vibakivirus | Unclassified | Myoviridae | Arthrobacter |
| MN096360 | Mycobacterium phage Rita1961 | Mycobacterium phage Rita1961 unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69026 | 67.466 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN096359 | Mycobacterium phage Obutu | Mycobacterium phage Obutu unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69021 | 67.429 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MN096358 | Microbacterium phage LeeroyJenkins | Microbacterium phage LeeroyJenkins Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62439 | 58.127 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN096357 | Microbacterium phage WaterT | Microbacterium phage WaterT Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61090 | 58.227 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN096356 | Mycobacterium phage Bunnies | Mycobacterium phage Bunnies unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48822 | 67.103 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK977711 | Streptomyces phage Evy | Streptomyces phage Evy unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132977 | 49.568 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MK937612 | Mycobacterium phage LilhomieP | Mycobacterium phage LilhomieP unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81071 | 61.583 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MK937611 | Gordonia phage Verity | Gordonia phage Verity Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61248 | 67.418 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK937608 | Microbacterium phage Cressida | Microbacterium phage Cressida unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55957 | 70.299 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MK937607 | Gordonia phage Zipp | Gordonia phage Zipp Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61219 | 67.378 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK937606 | Microbacterium phage Margaery | Microbacterium phage Margaery unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56033 | 69.532 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MK937604 | Mycobacterium phage Necropolis | Mycobacterium phage Necropolis unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46263 | 67.315 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK937603 | Gordonia phage Bakery | Gordonia phage Bakery unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60468 | 67.642 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK937601 | Gordonia phage Charming | Gordonia phage Charming unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58000 | 67.141 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MK937600 | Mycobacterium phage Wigglewiggle | Mycobacterium phage Wigglewiggle unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76414 | 58.971 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK937598 | Microbacterium phage Celaena | Microbacterium phage Celaena unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42169 | 67.751 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MK937595 | Streptomyces phage Dubu | Streptomyces phage Dubu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39535 | 68.683 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK937594 | Gordonia phage Galadriel | Gordonia phage Galadriel unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59400 | 67.446 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MK937591 | Microbacterium phage Cinna | Microbacterium phage Cinna unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55366 | 70.001 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MN010762 | Gordonia phage MintFen | Gordonia phage MintFen Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48512 | 67.105 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN010761 | Gordonia phage Kenosha | Gordonia phage Kenosha Gordonia virus Kenosha Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60899 | 51.753 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MN010760 | Gordonia phage Nubi | Gordonia phage Nubi unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58718 | 67.87 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MN010759 | Microbacterium phage Ciel | Microbacterium phage Ciel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.765 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN010758 | Gordonia phage Dardanus | Gordonia phage Dardanus unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43143 | 66.595 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MN010757 | Mycobacterium phage HUHilltop | Mycobacterium phage HUHilltop unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46896 | 67.172 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MN010755 | Microbacterium phage Slentz | Microbacterium phage Slentz Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.736 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK919484 | Arthrobacter phage DreamTeam | Arthrobacter phage DreamTeam unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43089 | 60.73 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK919482 | Mycobacterium phage DeepSoil15 | Mycobacterium phage DeepSoil15 unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.449 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK919480 | Mycobacterium phage Techage | Mycobacterium phage Techage unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47094 | 67.427 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK919479 | Arthrobacter phage Grekaycon | Arthrobacter phage Grekaycon unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51091 | 60.997 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MK919476 | Arthrobacter phage EstebanJulior | Arthrobacter phage EstebanJulior unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43746 | 61.265 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK919471 | Microbacterium phage Shee | Microbacterium phage Shee unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41800 | 63.524 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK919469 | Gordonia phage Madeline | Gordonia phage Madeline unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51668 | 66.262 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK977713 | Mycobacterium phage Tubs | Mycobacterium phage Tubs unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53300 | 62.627 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK977709 | Mycobacterium phage ThetaBob | Mycobacterium phage ThetaBob Mycobacterium virus ThetaBob Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56714 | 61.71 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK977704 | Gordonia Terrae phage RoadKill | Gordonia Terrae phage RoadKill Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55939 | 67.602 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK977702 | Gordonia terrae phage McKinley | Gordonia terrae phage McKinley unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59022 | 67.605 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MK977698 | Microbacterium phage Busephilis | Microbacterium phage Busephilis unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52986 | 68.756 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MK977696 | Gordonia phage SCentae | Gordonia phage SCentae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151316 | 66 | Unclassified | Unclassified | Myoviridae | Gordonia |
| MK977695 | Gordonia phage Pupper | Gordonia phage Pupper Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150830 | 66.106 | Unclassified | Unclassified | Myoviridae | Gordonia |
| MK801735 | Mycobacterium Phage Rachaly | Mycobacterium Phage Rachaly unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52009 | 63.433 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK801734 | Gordonia phage LordFarquaad | Gordonia phage LordFarquaad unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47863 | 66.826 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MK801732 | Microbacterium phage NarutoRun | Microbacterium phage NarutoRun unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56665 | 63.77 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MK801731 | Microbacterium phage Anakin | Microbacterium phage Anakin unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56665 | 63.773 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MK801730 | Microbacterium phage Belthelas | Microbacterium phage Belthelas Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17502 | 68.598 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK801727 | Gordonia phage ThankyouJordi | Gordonia phage ThankyouJordi unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53453 | 66.355 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK801726 | Mycobacterium phage ChotaBhai | Mycobacterium phage ChotaBhai unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76145 | 62.906 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK801725 | Gordonia phage Yago84 | Gordonia phage Yago84 unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61890 | 70.016 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MK801724 | Gordonia phage PhrostedPhlake | Gordonia phage PhrostedPhlake Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52042 | 67.376 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK801722 | Streptomyces phage Birchlyn | Streptomyces phage Birchlyn unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126524 | 49.245 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MK801721 | Gordonia phage William | Gordonia phage William Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50678 | 66.536 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK878904 | Microbacterium phage TinyTimothy | Microbacterium phage TinyTimothy Microbacterium virus TinyTimothy Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53932 | 56.751 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK878902 | Gordonia phage Twinkle | Gordonia phage Twinkle Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54241 | 65.565 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK878899 | Microbacterium phage Hulk | Microbacterium phage Hulk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17395 | 68.652 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK878897 | Mycobacterium phage Blackbrain | Mycobacterium phage Blackbrain unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155713 | 64.638 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK878896 | Gordonia phage Begonia | Gordonia phage Begonia Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50867 | 67.417 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK894435 | Microbacterium phage Alex44 | Microbacterium phage Alex44 Microbacterium virus Alex44 Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54396 | 59.949 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK894434 | Microbacterium phage LaviMo | Microbacterium phage LaviMo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17453 | 68.733 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK894433 | Mycobacterium phage Kipper29 | Mycobacterium phage Kipper29 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52009 | 61.57 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK977705 | Gordonia phage Mollymur | Gordonia phage Mollymur unclassified Bantamvirus Bantamvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90669 | 65.534 | Bantamvirus | Unclassified | Siphoviridae | Gordonia |
| MK864267 | Gordonia phage Valary | Gordonia phage Valary unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59448 | 67.787 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK864266 | Gordonia phage Arri | Gordonia phage Arri unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58650 | 67.812 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK864265 | Gordonia phage Barb | Gordonia phage Barb unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59514 | 67.831 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK864264 | Gordonia phage VanDeWege | Gordonia phage VanDeWege unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58972 | 67.822 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK864263 | Gordonia phage Melba | Gordonia phage Melba Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48350 | 67.115 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK820639 | Mycobacterium phage Phineas | Mycobacterium phage Phineas unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47229 | 67.188 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK820638 | Mycobacterium phage HermioneGrange | Mycobacterium phage HermioneGrange unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53147 | 63.58 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK875800 | Mycobacterium phage MoneyMay | Mycobacterium phage MoneyMay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 64.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK875799 | Mycobacterium phage Xena | Mycobacterium phage Xena unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51306 | 63.681 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK875798 | Microbacterium phage Piperis | Microbacterium phage Piperis unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53241 | 68.862 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MK875795 | Gordonia phage Agatha | Gordonia phage Agatha unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46543 | 60.153 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MK875793 | Gordonia phage WelcomeAyanna | Gordonia phage WelcomeAyanna unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53600 | 66.405 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK875792 | Mycobacterium phage Polka14 | Mycobacterium phage Polka14 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58758 | 61.438 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK875791 | Gordonia phage Yakult | Gordonia phage Yakult Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47197 | 62.873 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK737941 | Microbacterium phage Rachella | Microbacterium phage Rachella unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56699 | 63.812 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MK814761 | Gordonia phage SmokingBunny | Gordonia phage SmokingBunny unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59454 | 67.906 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK814758 | Mycobacterium phage FirstPlacePfu | Mycobacterium phage FirstPlacePfu unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45680 | 67.261 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK814757 | Gordonia phage Fairfaxidum | Gordonia phage Fairfaxidum Gordonia virus Fairfaxidum Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50187 | 66.585 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK814754 | Mycobacterium phage Sumter | Mycobacterium phage Sumter unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52656 | 63.67 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK814753 | Microbacterium phage BubbaBear | Microbacterium phage BubbaBear unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41814 | 66.578 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MK814752 | Gordonia phage Lutum | Gordonia phage Lutum unclassified Getalongvirus Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54534 | 62.988 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MK814751 | Mycobacterium phage Bananafish | Mycobacterium phage Bananafish unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67366 | 68.932 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK814749 | Arthrobacter phage Scuttle | Arthrobacter phage Scuttle unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43729 | 61.053 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK814748 | Mycobacterium phage XianYue | Mycobacterium phage XianYue unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52907 | 62.648 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK757448 | Arthrobacter phage Idaho | Arthrobacter phage Idaho Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15675 | 63.579 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MK757445 | Mycobacterium phage Lilizi | Mycobacterium phage Lilizi unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75360 | 63.01 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524531 | Mycobacterium phage Rutherferd | Mycobacterium phage Rutherferd unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52364 | 63.746 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524521 | Mycobacterium phage Schatzie | Mycobacterium phage Schatzie unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111345 | 60.834 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK686072 | Streptomyces phage RosaAsantewaa | Streptomyces phage RosaAsantewaa unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42730 | 58.863 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MK686071 | Streptomyces phage Mischief19 | Streptomyces phage Mischief19 unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68430 | 68.451 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| MK686069 | Streptomyces phage Heather | Streptomyces phage Heather Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41238 | 62.845 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK686068 | Streptomyces phage RemusLoopin | Streptomyces phage RemusLoopin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41376 | 62.005 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK660714 | Arthrobacter phage MrGloopy | Arthrobacter phage MrGloopy unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43701 | 60.882 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK660713 | Microbacterium phage Theresita | Microbacterium phage Theresita Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40234 | 65.852 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK620900 | Streptomyces phage Forthebois | Streptomyces phage Forthebois Streptomyces virus Forthebois Deltatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 18251 | 53.586 | Deltatectivirus | Unclassified | Tectiviridae | Streptomyces |
| MK620899 | Gordonia phage GodonK | Gordonia phage GodonK Gordonia virus GodonK Godonkavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122563 | 54.382 | Godonkavirus | Unclassified | Siphoviridae | Gordonia |
| MK620898 | Mycobacterium phage Coffee | Mycobacterium phage Coffee unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67481 | 68.951 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK620897 | Microbacterium phage Rhysand | Microbacterium phage Rhysand Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17453 | 68.727 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK620896 | Streptomyces phage Circinus | Streptomyces phage Circinus unclassified Wilnyevirus Wilnyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126383 | 52.884 | Wilnyevirus | Unclassified | Siphoviridae | Streptomyces |
| MK620894 | Streptomyces phage Kradal | Streptomyces phage Kradal Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186383 | 66.718 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK620893 | Mycobacterium phage HanKaySha | Mycobacterium phage HanKaySha unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75528 | 63.024 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494123 | Mycobacterium phage Blackbeetle | Mycobacterium phage Blackbeetle unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64824 | 63.385 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494095 | Mycobacterium phage Daegal | Mycobacterium phage Daegal unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54858 | 67.476 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524501 | Gordonia phage BrutonGaster | Gordonia phage BrutonGaster Gordonia virus BrutonGaster Oneupvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91510 | 61.48 | Oneupvirus | Unclassified | Siphoviridae | Gordonia |
| MK494132 | Mycobacterium phage Cepens | Mycobacterium phage Cepens Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60656 | 67.558 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK494131 | Mycobacterium phage GodPhather | Mycobacterium phage GodPhather Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61415 | 67.57 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK494130 | Mycobacterium phage Fibonacci | Mycobacterium phage Fibonacci unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52462 | 63.753 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494122 | Mycobacterium phage GreaseLightnin | Mycobacterium phage GreaseLightnin unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48424 | 67.091 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK494116 | Mycobacterium phage Dulcie | Mycobacterium phage Dulcie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53970 | 63.728 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494114 | Mycobacterium phage Argie | Mycobacterium phage Argie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61547 | 67.516 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK494112 | Streptomyces phage EGole | Streptomyces phage EGole unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135653 | 49.905 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MK494108 | Mycobacterium phage Charm | Mycobacterium phage Charm unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52596 | 61.86 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494105 | Mycobacterium phage Pinnie | Mycobacterium phage Pinnie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44842 | 68.761 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK494104 | Mycobacterium phage HenryJackson | Mycobacterium phage HenryJackson unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69033 | 66.509 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK494102 | Mycobacterium phage WaldoWhy | Mycobacterium phage WaldoWhy Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64529 | 59.658 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MK494100 | Mycobacterium phage Halena | Mycobacterium phage Halena unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73871 | 58.82 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494099 | Mycobacterium phage Typha | Mycobacterium phage Typha Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75687 | 65.953 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK494092 | Mycobacterium phage Silverleaf | Mycobacterium phage Silverleaf unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73210 | 58.78 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494091 | Mycobacterium phage Scorpia | Mycobacterium phage Scorpia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50693 | 59.724 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494090 | Mycobacterium phage Jordennis | Mycobacterium phage Jordennis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52234 | 61.588 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494087 | Mycobacterium phage DBQu4n | Mycobacterium phage DBQu4n unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52724 | 63.497 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK559429 | Mycobacterium phage Moldemort | Mycobacterium phage Moldemort unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75240 | 63.032 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK559428 | Mycobacterium phage Elephantoon | Mycobacterium phage Elephantoon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52975 | 62.748 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK559427 | Mycobacterium phage Delton | Mycobacterium phage Delton Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64963 | 59.632 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MK524530 | Mycobacterium phage Pharaoh | Mycobacterium phage Pharaoh unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 63.729 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524529 | Mycobacterium phage Phoebus | Mycobacterium phage Phoebus unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114320 | 60.817 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524527 | Mycobacterium phage ThreeRngTarjay | Mycobacterium phage ThreeRngTarjay unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113254 | 60.959 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524526 | Mycobacterium phage Robyn | Mycobacterium phage Robyn unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68467 | 66.566 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK524525 | Mycobacterium phage Ringer | Mycobacterium phage Ringer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52018 | 63.572 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524524 | Streptomyces phage Caelum | Streptomyces phage Caelum unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50374 | 66.969 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK524523 | Mycobacterium phage Naira | Mycobacterium phage Naira unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53581 | 63.704 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524519 | Mycobacterium phage Salz | Mycobacterium phage Salz unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51333 | 63.846 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524517 | Mycobacterium phage James | Mycobacterium phage James unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59617 | 61.394 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524516 | Mycobacterium phage Bobby | Mycobacterium phage Bobby unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110636 | 60.742 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524514 | Mycobacterium phage Bud | Mycobacterium phage Bud unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51304 | 63.785 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524513 | Mycobacterium phage Munch | Mycobacterium phage Munch unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52595 | 63.692 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524512 | Mycobacterium phage Carthage | Mycobacterium phage Carthage unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67952 | 66.587 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK524511 | Mycobacterium phage Bowtie | Mycobacterium phage Bowtie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51202 | 63.79 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524510 | Microbacterium phage Fireman | Microbacterium phage Fireman Microbacterium virus Fireman Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54579 | 68.574 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MK524508 | Mycobacterium phage Zilizebeth | Mycobacterium phage Zilizebeth unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48056 | 67.259 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK524507 | Mycobacterium phage Orange | Mycobacterium phage Orange unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52147 | 63.628 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524504 | Mycobacterium phage Hughesyang | Mycobacterium phage Hughesyang unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112884 | 60.948 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524503 | Mycobacterium phage Fancypants | Mycobacterium phage Fancypants unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60228 | 61.085 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524502 | Microbacterium phage TimoTea | Microbacterium phage TimoTea Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17427 | 68.709 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK524498 | Mycobacterium phage Chill | Mycobacterium phage Chill Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64529 | 59.657 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MK524497 | Mycobacterium phage Francis47 | Mycobacterium phage Francis47 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53936 | 63.564 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524494 | Mycobacterium phage Rabbs | Mycobacterium phage Rabbs unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42430 | 66.432 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524492 | Mycobacterium phage Expelliarmus | Mycobacterium phage Expelliarmus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52274 | 61.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524491 | Mycobacterium phage Whabigail7 | Mycobacterium phage Whabigail7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53167 | 63.338 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524490 | Mycobacterium phage Donny | Mycobacterium phage Donny unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69691 | 68.454 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK524489 | Mycobacterium phage Ugenie5 | Mycobacterium phage Ugenie5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50799 | 62.716 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524486 | Mycobacterium phage Waleliano | Mycobacterium phage Waleliano unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70963 | 68.908 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK501730 | Gordonia phage Barco | Gordonia phage Barco Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48905 | 67.279 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK501729 | Gordonia phage Walrus | Gordonia phage Walrus Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52024 | 67.3 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK433278 | Streptomyces phage Nerdos | Streptomyces phage Nerdos unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49557 | 66.223 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK433276 | Streptomyces phage Asten | Streptomyces phage Asten unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51571 | 65.671 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK433271 | Streptomyces phage Indigo | Streptomyces phage Indigo unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49558 | 66.203 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK433270 | Streptomyces phage Bovely | Streptomyces phage Bovely unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49576 | 66.173 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK433269 | Streptomyces phage Madamato | Streptomyces phage Madamato unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55349 | 59.584 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MK433268 | Streptomyces phage Popy | Streptomyces phage Popy unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56470 | 59.412 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MK433264 | Gordonia phage Tiamoceli | Gordonia phage Tiamoceli Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55829 | 67.698 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK433262 | Mycobacterium phage Nimrod | Mycobacterium phage Nimrod unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76435 | 62.914 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK433260 | Streptomyces phage Namo | Streptomyces phage Namo unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56168 | 59.553 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MK433259 | Streptomyces phage IceWarrior | Streptomyces phage IceWarrior unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55532 | 59.546 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MK433266 | Streptomyces phage Shawty | Streptomyces phage Shawty unclassified Lomovskayavirus Lomovskayavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40733 | 63.15 | Lomovskayavirus | Unclassified | Siphoviridae | Streptomyces |
| MK450422 | Mycobacterium phage RoMag | Mycobacterium phage RoMag unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156319 | 64.713 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359318 | Mycobacterium phage Specks | Mycobacterium phage Specks unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156374 | 64.667 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK411746 | Arthrobacter phage Sonali | Arthrobacter phage Sonali Arthrobacter virus Sonali Sonalivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61073 | 67.74 | Sonalivirus | Unclassified | Siphoviridae | Arthrobacter |
| MK392368 | Arthrobacter phage Elesar | Arthrobacter phage Elesar Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42954 | 64.96 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MK392366 | Streptomyces phage Janus | Streptomyces phage Janus unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50632 | 67.047 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK392365 | Streptomyces phage TinaBelcher | Streptomyces phage TinaBelcher unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52490 | 67.718 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK392364 | Streptomyces phage Nishikigoi | Streptomyces phage Nishikigoi unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50660 | 66.824 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK392363 | Streptomyces phage Bowden | Streptomyces phage Bowden unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52603 | 67.703 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK359366 | Mycobacterium phage Basquiat | Mycobacterium phage Basquiat unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154283 | 64.726 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359365 | Mycobacterium phage Loanshark | Mycobacterium phage Loanshark unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154593 | 64.78 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359364 | Mycobacterium phage ChickenPhender | Mycobacterium phage ChickenPhender unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154397 | 64.709 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359363 | Mycobacterium phage Fludd | Mycobacterium phage Fludd unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154456 | 64.717 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359362 | Mycobacterium phage Atlantean | Mycobacterium phage Atlantean unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154553 | 64.747 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359361 | Mycobacterium phage Darko | Mycobacterium phage Darko unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155951 | 64.713 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359360 | Mycobacterium phage Hexamo | Mycobacterium phage Hexamo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52359 | 61.414 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359359 | Mycobacterium phage Colt | Mycobacterium phage Colt unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158508 | 64.698 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359358 | Mycobacterium phage Essence | Mycobacterium phage Essence unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155700 | 64.64 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359357 | Mycobacterium phage RomaT | Mycobacterium phage RomaT unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69239 | 67.489 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK359355 | Mycobacterium phage Sprinklers | Mycobacterium phage Sprinklers unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156060 | 64.66 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359354 | Mycobacterium phage PinkPlastic | Mycobacterium phage PinkPlastic unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49593 | 63.648 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359353 | Mycobacterium phage BadAgartude | Mycobacterium phage BadAgartude unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155658 | 64.672 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359352 | Mycobacterium phage JewelBug | Mycobacterium phage JewelBug unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50341 | 61.602 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359351 | Streptomyces phage BoomerJR | Streptomyces phage BoomerJR unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131220 | 49.249 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MK359350 | Mycobacterium phage Nidhogg | Mycobacterium phage Nidhogg unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156342 | 64.695 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359349 | Mycobacterium phage FoxtrotP1 | Mycobacterium phage FoxtrotP1 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156926 | 64.718 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359348 | Mycobacterium phage Rahel | Mycobacterium phage Rahel unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155955 | 64.705 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359347 | Mycobacterium phage Ading | Mycobacterium phage Ading unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157399 | 64.648 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359346 | Mycobacterium phage NoodleTree | Mycobacterium phage NoodleTree unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151100 | 64.746 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359345 | Mycobacterium phage QBert | Mycobacterium phage QBert unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153772 | 64.812 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359344 | Mycobacterium phage Napoleon13 | Mycobacterium phage Napoleon13 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154072 | 64.674 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359342 | Mycobacterium phage ShiaLabeouf | Mycobacterium phage ShiaLabeouf unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154471 | 64.7 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359339 | Mycobacterium phage Naija | Mycobacterium phage Naija unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155256 | 64.714 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359338 | Mycobacterium phage Amataga | Mycobacterium phage Amataga unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154553 | 64.747 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359337 | Mycobacterium phage Flabslab | Mycobacterium phage Flabslab unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152593 | 64.649 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359336 | Mycobacterium phage Chargie21 | Mycobacterium phage Chargie21 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154283 | 64.725 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359335 | Mycobacterium phage Patter | Mycobacterium phage Patter unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155470 | 64.708 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359334 | Mycobacterium phage Delilah | Mycobacterium phage Delilah unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155127 | 64.755 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359333 | Mycobacterium phage JustHall | Mycobacterium phage JustHall unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154933 | 64.702 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359332 | Streptomyces phage Genie2 | Streptomyces phage Genie2 unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131333 | 49.288 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MK359331 | Mycobacterium phage Rajelicia | Mycobacterium phage Rajelicia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52843 | 63.592 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359330 | Mycobacterium phage Iota | Mycobacterium phage Iota unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156068 | 64.71 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359329 | Mycobacterium phage Kamryn | Mycobacterium phage Kamryn unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156360 | 64.724 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359328 | Mycobacterium phage EmmaElysia | Mycobacterium phage EmmaElysia unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157365 | 64.646 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359327 | Mycobacterium phage EmToTheThree | Mycobacterium phage EmToTheThree unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155601 | 64.779 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359326 | Mycobacterium phage Pivoine | Mycobacterium phage Pivoine unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155256 | 64.716 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359324 | Mycobacterium phage Tortoise16 | Mycobacterium phage Tortoise16 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154584 | 64.758 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359321 | Mycobacterium phage Phusco | Mycobacterium phage Phusco unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155060 | 64.726 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359320 | Mycobacterium phage Cookies | Mycobacterium phage Cookies unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76340 | 62.979 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359319 | Gordonia phage Parada | Gordonia phage Parada unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49909 | 67.3 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MK359317 | Mycobacterium phage FrayBell | Mycobacterium phage FrayBell unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154543 | 64.753 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359314 | Mycobacterium phage Ewok | Mycobacterium phage Ewok unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155005 | 64.722 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359313 | Gordonia phage Vasanti | Gordonia phage Vasanti unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46463 | 66.984 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MK359312 | Mycobacterium phage Fushigi | Mycobacterium phage Fushigi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50350 | 63.668 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359311 | Mycobacterium phage SmallFry | Mycobacterium phage SmallFry unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155433 | 64.691 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359309 | Mycobacterium phage Czyszczon1 | Mycobacterium phage Czyszczon1 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75075 | 62.942 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359308 | Mycobacterium phage Shaqnato | Mycobacterium phage Shaqnato unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155596 | 64.631 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359306 | Mycobacterium phage Norm | Mycobacterium phage Norm unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155550 | 64.742 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359305 | Mycobacterium phage Teardrop | Mycobacterium phage Teardrop unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155389 | 64.674 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359304 | Mycobacterium phage SpikeBT | Mycobacterium phage SpikeBT unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50181 | 63.741 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359303 | Mycobacterium phage Shnickers | Mycobacterium phage Shnickers unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154529 | 64.71 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK359299 | Mycobacterium phage Petp2012 | Mycobacterium phage Petp2012 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51703 | 63.681 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310146 | Mycobacterium phage Crespo | Mycobacterium phage Crespo unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41902 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310144 | Mycobacterium phage CicholasNage | Mycobacterium phage CicholasNage unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71059 | 58.612 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310143 | Mycobacterium phage Dietrick | Mycobacterium phage Dietrick unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153582 | 64.728 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK310142 | Mycobacterium phage MetalQZJ | Mycobacterium phage MetalQZJ unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51854 | 63.54 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310141 | Mycobacterium phage Fenn | Mycobacterium phage Fenn unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53449 | 63.67 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310139 | Mycobacterium phage Emiris | Mycobacterium phage Emiris unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69027 | 66.377 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK460248 | Streptomyces phage Teutsch | Streptomyces phage Teutsch unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132885 | 50.01 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MK460245 | Streptomyces phage BartholomewSD | Streptomyces phage BartholomewSD unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52131 | 67.576 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MK450434 | Mycobacterium phage DrPhinkDaddy | Mycobacterium phage DrPhinkDaddy unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156574 | 64.678 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450433 | Streptomyces phage Sebastisaurus | Streptomyces phage Sebastisaurus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41609 | 62.078 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK450432 | Mycobacterium phage ValleyTerrace | Mycobacterium phage ValleyTerrace unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154553 | 64.746 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450431 | Streptomyces phage Lilbooboo | Streptomyces phage Lilbooboo Streptomyces virus Lilbooboo Vashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41267 | 62.66 | Vashvirus | Unclassified | Siphoviridae | Streptomyces |
| MK450429 | Mycobacterium phage Bipolarisk | Mycobacterium phage Bipolarisk unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154734 | 64.739 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450427 | Mycobacterium phage Adlitam | Mycobacterium phage Adlitam unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155007 | 64.682 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450426 | Streptomyces phage Euratis | Streptomyces phage Euratis unclassified Vashvirus Vashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41403 | 62.94 | Vashvirus | Unclassified | Siphoviridae | Streptomyces |
| MK450425 | Mycobacterium phage Shelob | Mycobacterium phage Shelob unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156606 | 64.764 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450424 | Mycobacterium phage Melpomini | Mycobacterium phage Melpomini unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154855 | 64.708 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450423 | Mycobacterium phage Stubby | Mycobacterium phage Stubby unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153369 | 64.715 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK450421 | Streptomyces phage Vash | Streptomyces phage Vash Streptomyces virus Vash Vashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41578 | 62.485 | Vashvirus | Unclassified | Siphoviridae | Streptomyces |
| MK305893 | Mycobacterium phage Beatrix | Mycobacterium phage Beatrix unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52295 | 63.941 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK305892 | Mycobacterium phage RedRaider77 | Mycobacterium phage RedRaider77 unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64827 | 63.404 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK305891 | Streptomyces phage Gibson | Streptomyces phage Gibson Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69439 | 64.351 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK305890 | Streptomyces phage WheeHeim | Streptomyces phage WheeHeim Streptomyces virus WheeHeim Deltatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 18266 | 54.621 | Deltatectivirus | Unclassified | Tectiviridae | Streptomyces |
| MK305889 | Gordonia phage Mutzi | Gordonia phage Mutzi unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59655 | 67.562 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MK305888 | Streptomyces phage Austintatious | Streptomyces phage Austintatious Streptomyces virus Austintatious Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36213 | 72.598 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| MK305886 | Mycobacterium phage Poenanya | Mycobacterium phage Poenanya unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58666 | 61.492 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK305885 | Gordonia phage Mulch | Gordonia phage Mulch unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49909 | 67.3 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MK376958 | Gordonia phage Brylie | Gordonia phage Brylie unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49870 | 67.303 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MK376957 | Gordonia phage Duffington | Gordonia phage Duffington Gordonia virus Duffington Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61343 | 51.373 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MK376956 | Gordonia phage WilliamBoone | Gordonia phage WilliamBoone unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92688 | 61.939 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MK376953 | Gordonia phage Rickmore | Gordonia phage Rickmore Gordonia virus Rickmore Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60218 | 50.538 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MH825713 | Mycobacterium phage Zalkecks | Mycobacterium phage Zalkecks unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154764 | 64.739 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK284522 | Mycobacterium phage Malec | Mycobacterium phage Malec unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53377 | 63.647 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061412 | Streptomyces phage Gilson | Streptomyces phage Gilson Streptomyces virus Gilson Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128338 | 47.439 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MK279881 | Mycobacterium phage Solosis | Mycobacterium phage Solosis unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68068 | 66.425 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279912 | Gordonia phage Savage | Gordonia phage Savage unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.628 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MK279911 | Gordonia phage Sproutie | Gordonia phage Sproutie unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.63 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MK279910 | Mycobacterium phage Antonia | Mycobacterium phage Antonia unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68573 | 66.466 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279908 | Mycobacterium phage Roliet | Mycobacterium phage Roliet unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67687 | 66.482 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279907 | Arthrobacter phage HeadNerd | Arthrobacter phage HeadNerd unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42968 | 61.357 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279906 | Gordonia phage Kenna | Gordonia phage Kenna Gordonia virus Kenna Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53215 | 62.975 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MK279905 | Mycobacterium phage Acquire49 | Mycobacterium phage Acquire49 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73639 | 58.7 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279904 | Mycobacterium phage RedMaple | Mycobacterium phage RedMaple unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68985 | 66.427 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279901 | Arthrobacter phage Potatoes | Arthrobacter phage Potatoes unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43661 | 60.972 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279900 | Gordonia phage Nina | Gordonia phage Nina unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46702 | 59.533 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MK279897 | Mycobacterium phage Bishoperium | Mycobacterium phage Bishoperium unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68865 | 66.495 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279896 | Arthrobacter phage Zorro | Arthrobacter phage Zorro unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43562 | 61.072 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279895 | Mycobacterium phage Zaider | Mycobacterium phage Zaider unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68983 | 66.569 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279894 | Mycobacterium phage YouGoGlencoco | Mycobacterium phage YouGoGlencoco unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68608 | 66.432 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279893 | Gordonia phage WheatThin | Gordonia phage WheatThin unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49897 | 67.291 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MK279892 | Arthrobacter phage Wawa | Arthrobacter phage Wawa unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43850 | 60.899 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279891 | Mycobacterium phage Wallhey | Mycobacterium phage Wallhey unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68821 | 66.385 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279890 | Mycobacterium phage Veritas | Mycobacterium phage Veritas unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68718 | 66.339 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279889 | Mycobacterium phage Valjean | Mycobacterium phage Valjean unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68560 | 66.495 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279888 | Mycobacterium phage TomBombadil | Mycobacterium phage TomBombadil unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68880 | 66.418 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279887 | Mycobacterium phage Timmi | Mycobacterium phage Timmi unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68355 | 66.481 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279886 | Arthrobacter phage TattModd | Arthrobacter phage TattModd unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43546 | 61.002 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279885 | Mycobacterium phage Surely | Mycobacterium phage Surely unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68888 | 66.431 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279884 | Arthrobacter phage Supakev | Arthrobacter phage Supakev unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43764 | 61.701 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279883 | Mycobacterium phage Struggle | Mycobacterium phage Struggle unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68669 | 66.403 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279882 | Mycobacterium phage Sophia | Mycobacterium phage Sophia unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68306 | 66.498 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279880 | Arthrobacter phage Savage2526 | Arthrobacter phage Savage2526 unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43418 | 60.977 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279879 | Mycobacterium phage SassyCat97 | Mycobacterium phage SassyCat97 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68318 | 66.463 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279878 | Mycobacterium phage Samaymay | Mycobacterium phage Samaymay unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68650 | 66.471 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279877 | Mycobacterium phage Roscoe | Mycobacterium phage Roscoe unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68603 | 66.545 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279876 | Arthrobacter phage Rozby | Arthrobacter phage Rozby unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43195 | 60.982 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279875 | Arthrobacter phage Riverdale | Arthrobacter phage Riverdale unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43391 | 60.99 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279874 | Arthrobacter phage Riovina | Arthrobacter phage Riovina unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43739 | 61.551 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279873 | Mycobacterium phage QueenBeane | Mycobacterium phage QueenBeane unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68650 | 66.472 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279872 | Arthrobacter phage Pterodactyl | Arthrobacter phage Pterodactyl unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43274 | 60.769 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279871 | Mycobacterium phage Plmatters | Mycobacterium phage Plmatters unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68318 | 66.473 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279870 | Mycobacterium phage Omniscient | Mycobacterium phage Omniscient unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69197 | 66.51 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279869 | Arthrobacter phage OurGirlNessie | Arthrobacter phage OurGirlNessie unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42561 | 60.678 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279868 | Arthrobacter phage OMalley | Arthrobacter phage OMalley unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43739 | 61.554 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279867 | Arthrobacter phage Nancia | Arthrobacter phage Nancia unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43238 | 60.701 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279866 | Mycobacterium phage MRabcd | Mycobacterium phage MRabcd unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68791 | 66.491 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279865 | Mycobacterium phage Mecca | Mycobacterium phage Mecca unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68890 | 66.532 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279864 | Mycobacterium phage Mag7 | Mycobacterium phage Mag7 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68497 | 66.356 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279863 | Arthrobacter phage Litotes | Arthrobacter phage Litotes unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43496 | 61.081 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279862 | Arthrobacter phage Lennox | Arthrobacter phage Lennox unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43087 | 60.726 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279861 | Mycobacterium phage Legolas | Mycobacterium phage Legolas unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68664 | 66.438 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279860 | Arthrobacter phage Lasagna | Arthrobacter phage Lasagna unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42602 | 60.795 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279859 | Mycobacterium phage Kwadwo | Mycobacterium phage Kwadwo unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68334 | 66.471 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279858 | Mycobacterium phage JakeO | Mycobacterium phage JakeO unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68341 | 66.456 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279857 | Mycobacterium phage IronMan | Mycobacterium phage IronMan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53467 | 63.316 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279856 | Arthrobacter phage Huckleberry | Arthrobacter phage Huckleberry unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42972 | 61.382 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279855 | Mycobacterium phage Haleema | Mycobacterium phage Haleema unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68415 | 66.5 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279854 | Arthrobacter phage GreenHearts | Arthrobacter phage GreenHearts unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44251 | 60.753 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279853 | Gordonia phage Gray | Gordonia phage Gray unclassified Bendigovirus Bendigovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88970 | 60.298 | Bendigovirus | Unclassified | Myoviridae | Gordonia |
| MK279852 | Mycobacterium phage Fringe | Mycobacterium phage Fringe unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69191 | 66.442 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279851 | Arthrobacter phage Eunoia | Arthrobacter phage Eunoia unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44416 | 61.525 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279850 | Mycobacterium phage Durga | Mycobacterium phage Durga unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68866 | 66.416 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279849 | Mycobacterium phage Duke13 | Mycobacterium phage Duke13 unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111970 | 60.734 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279847 | Arthrobacter phage CristinaYang | Arthrobacter phage CristinaYang unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43366 | 60.681 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279846 | Mycobacterium phage Cosmolli16 | Mycobacterium phage Cosmolli16 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68978 | 66.473 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279845 | Arthrobacter phage Cholula | Arthrobacter phage Cholula unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43661 | 60.983 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279844 | Arthrobacter phage ChewChew | Arthrobacter phage ChewChew unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43439 | 60.701 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK279843 | Mycobacterium phage CamL | Mycobacterium phage CamL unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68650 | 66.472 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK279842 | Mycobacterium phage Raela | Mycobacterium phage Raela unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65380 | 63.396 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279841 | Streptomyces phage Hiyaa | Streptomyces phage Hiyaa Streptomyces virus Hiyaa Hiyaavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83219 | 66.661 | Hiyaavirus | Unclassified | Siphoviridae | Streptomyces |
| MK279840 | Mycobacterium phage JacoRen57 | Mycobacterium phage JacoRen57 unclassified Mapvirus Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52411 | 56.738 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112547 | Gordonia phage Maridalia | Gordonia phage Maridalia unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51628 | 66.731 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK112526 | Arthrobacter phage Aledel | Arthrobacter phage Aledel unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43739 | 61.551 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112527 | Mycobacterium phage Altwerkus | Mycobacterium phage Altwerkus unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68179 | 66.519 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112528 | Arthrobacter phage Beethoven | Arthrobacter phage Beethoven unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43934 | 60.957 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112529 | Arthrobacter phage BigMack | Arthrobacter phage BigMack unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43357 | 61.266 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112530 | Mycobacterium phage BigPaolini | Mycobacterium phage BigPaolini unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49601 | 63.454 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112531 | Arthrobacter phage Bodacious | Arthrobacter phage Bodacious unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43239 | 60.723 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112532 | Mycobacterium phage Bones | Mycobacterium phage Bones unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52528 | 63.796 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112533 | Mycobacterium phage Bonray | Mycobacterium phage Bonray unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157364 | 64.671 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112534 | Mycobacterium phage Burrough | Mycobacterium phage Burrough unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155885 | 64.763 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112535 | Arthrobacter phage CallieOMalley | Arthrobacter phage CallieOMalley unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43862 | 61.183 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112536 | Mycobacterium phage Cannibal | Mycobacterium phage Cannibal unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68753 | 66.423 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112538 | Arthrobacter phage Carpal | Arthrobacter phage Carpal unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43706 | 61.015 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MK112539 | Mycobacterium phage Dione | Mycobacterium phage Dione unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68057 | 66.534 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112540 | Mycobacterium phage Froghopper | Mycobacterium phage Froghopper unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48149 | 64.005 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112541 | Mycobacterium phage Jessibeth14 | Mycobacterium phage Jessibeth14 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155784 | 64.709 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112542 | Mycobacterium phage Jillium | Mycobacterium phage Jillium unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68750 | 66.543 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112544 | Mycobacterium phage Keitherie | Mycobacterium phage Keitherie unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69085 | 66.414 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112545 | Mycobacterium phage Khaleesi | Mycobacterium phage Khaleesi unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155251 | 64.724 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112546 | Mycobacterium phage LuckyMarjie | Mycobacterium phage LuckyMarjie unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68075 | 66.546 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112548 | Mycobacterium phage McWolfish | Mycobacterium phage McWolfish unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154462 | 64.741 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112549 | Mycobacterium phage Morizzled23 | Mycobacterium phage Morizzled23 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155131 | 64.617 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112551 | Mycobacterium phage Riggan | Mycobacterium phage Riggan unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68831 | 66.461 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112552 | Mycobacterium phage Spartan300 | Mycobacterium phage Spartan300 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68568 | 66.496 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112553 | Mycobacterium Phage Squiggle | Mycobacterium Phage Squiggle unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68325 | 66.471 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112554 | Mycobacterium phage TinyTim | Mycobacterium phage TinyTim unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153817 | 64.779 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MK112555 | Mycobacterium phage Zelda | Mycobacterium phage Zelda unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68939 | 66.496 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK112556 | Mycobacterium phage Zerg | Mycobacterium phage Zerg unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58079 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061415 | Mycobacterium phage Rhynn | Mycobacterium phage Rhynn unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52522 | 63.611 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061414 | Mycobacterium phage Marley1013 | Mycobacterium phage Marley1013 unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69164 | 67.479 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK061413 | Arthrobacter phage Liebe | Arthrobacter phage Liebe Arthrobacter virus Liebe Liebevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45803 | 68.819 | Liebevirus | Unclassified | Siphoviridae | Arthrobacter |
| MK061411 | Mycobacterium phage Faze9 | Mycobacterium phage Faze9 unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67503 | 68.899 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MK061409 | Mycobacterium phage ExplosioNervosa | Mycobacterium phage ExplosioNervosa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53014 | 62.54 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061408 | Mycobacterium phage Centaur | Mycobacterium phage Centaur unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53001 | 63.499 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061407 | Mycobacterium phage Beelzebub | Mycobacterium phage Beelzebub unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65529 | 63.357 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK061406 | Gordonia phage Adora | Gordonia phage Adora Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51827 | 65.736 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH976519 | Gordonia phage Tangent | Gordonia phage Tangent unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58281 | 67.18 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976518 | Gordonia phage Stultus | Gordonia phage Stultus unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57916 | 67.738 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976515 | Gordonia phage Octobien14 | Gordonia phage Octobien14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76358 | 58.733 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH976514 | Gordonia phage Nordenberg | Gordonia phage Nordenberg unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57572 | 67.253 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976513 | Gordonia phage Kroos | Gordonia phage Kroos unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57973 | 68.13 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976512 | Gordonia phage Geodirt | Gordonia phage Geodirt unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59495 | 67.656 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976511 | Gordonia phage Gaea | Gordonia phage Gaea unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57472 | 68.127 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976510 | Gordonia phage Fosterous | Gordonia phage Fosterous unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59136 | 67.225 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976509 | Gordonia phage Bizzy | Gordonia phage Bizzy unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57312 | 68.321 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976508 | Gordonia phage Bibwit | Gordonia phage Bibwit unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57880 | 67.457 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976507 | Gordonia phage Ailee | Gordonia phage Ailee unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58474 | 67.505 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH976506 | Gordonia phage Affeca | Gordonia phage Affeca unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59156 | 67.454 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH910040 | Mycobacterium phage Sauce | Mycobacterium phage Sauce unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156554 | 64.721 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH910037 | Arthrobacter phage Isolde | Arthrobacter phage Isolde Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53234 | 62.65 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH910036 | Arthrobacter phage Hestia | Arthrobacter phage Hestia Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51476 | 62.283 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MK016504 | Mycobacterium phage Whouxphf | Mycobacterium phage Whouxphf unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59093 | 61.058 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016502 | Mycobacterium phage Pat3 | Mycobacterium phage Pat3 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75675 | 62.871 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016499 | Mycobacterium phage Mangethe | Mycobacterium phage Mangethe unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47612 | 67.37 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK016498 | Mycobacterium phage Manda | Mycobacterium phage Manda unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76023 | 62.813 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016497 | Microbacterium phage Johann | Microbacterium phage Johann unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42363 | 66.306 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MK016495 | Microbacterium phage Goodman | Microbacterium phage Goodman Microbacterium virus Goodman Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42363 | 66.31 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MK016494 | Mycobacterium phage Filuzino | Mycobacterium phage Filuzino unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59306 | 62.095 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016493 | Brevibacterium phage Cantare | Brevibacterium phage Cantare Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90743 | 52.897 | Unclassified | Unclassified | Myoviridae | Brevibacterium |
| MK016491 | Mycobacterium phage BaboJay | Mycobacterium phage BaboJay unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76377 | 62.887 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH926062 | Mycobacterium phage Vorrps | Mycobacterium phage Vorrps unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71722 | 65.311 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH926060 | Mycobacterium phage Schadenfreude | Mycobacterium phage Schadenfreude unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68591 | 66.48 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH727543 | Mycobacterium phage CharlieB | Mycobacterium phage CharlieB unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155886 | 64.688 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH834629 | Arthrobacter phage Yang | Arthrobacter phage Yang Arthobacter virus Yang Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43206 | 68.389 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834627 | Arthrobacter phage Ryan | Arthrobacter phage Ryan Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43182 | 65.069 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834626 | Microbacterium phage Rollins | Microbacterium phage Rollins unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39926 | 67.134 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MH834624 | Arthrobacter phage Polka | Arthrobacter phage Polka unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38077 | 66.699 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834621 | Arthrobacter phage Nandita | Arthrobacter phage Nandita Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42378 | 64.897 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834620 | Arthrobacter phage Melons | Arthrobacter phage Melons unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38457 | 66.784 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834619 | Arthrobacter phage Maureen | Arthrobacter phage Maureen unclassified Liebevirus Liebevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45802 | 68.816 | Liebevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834618 | Arthrobacter phage Lunar | Arthrobacter phage Lunar unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38450 | 66.687 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834617 | Mycobacterium phage Lizziana | Mycobacterium phage Lizziana unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58423 | 61.335 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH834616 | Arthrobacter phage Kepler | Arthrobacter phage Kepler Arthrobacter virus Kepler Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38449 | 66.735 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834615 | Mycobacterium phage KandZ | Mycobacterium phage KandZ Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64596 | 59.683 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MH834613 | Mycobacterium phage Fowlmouth | Mycobacterium phage Fowlmouth Mycobacterium virus Fowlmouth Fowlmouthvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69660 | 48.666 | Fowlmouthvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH834612 | Arthrobacter phage Faja | Arthrobacter phage Faja Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52490 | 63.124 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834610 | Arthrobacter phage DrManhattan | Arthrobacter phage DrManhattan Arthrobacter virus DrManhattan Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42577 | 65.953 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834609 | Arthrobacter phage Daob | Arthrobacter phage Daob unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37967 | 66.595 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834608 | Arthrobacter phage Cote | Arthrobacter phage Cote unclassified Coralvirus Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39044 | 66.722 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834607 | Arthrobacter phage Corgi | Arthrobacter phage Corgi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15771 | 67.58 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834606 | Arthrobacter phage Coral | Arthrobacter phage Coral Arthrobacter virus Coral Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38317 | 66.66 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834604 | Microbacterium phage Coltrane | Microbacterium phage Coltrane unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39828 | 67.071 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MH834599 | Microbacterium phage Bernstein | Microbacterium phage Bernstein unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39926 | 67.134 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MH834596 | Microbacterium phage Armstrong | Microbacterium phage Armstrong Microbacterium virus Armstrong Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39928 | 67.138 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MH834595 | Arthrobacter phage Andrew | Arthrobacter phage Andrew Arthrobacter virus Andrew Andrewvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38802 | 65.533 | Andrewvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834622 | Arthrobacter phage Noely | Arthrobacter phage Noely Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15013 | 68.321 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834602 | Microbacterium phage Brahms | Microbacterium phage Brahms unclassified Armstrongvirus Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39828 | 67.068 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| MH744416 | Microbacterium phage Dongwon | Microbacterium phage Dongwon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.517 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH744415 | Mycobacterium phage ArcusAngelus | Mycobacterium phage ArcusAngelus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58569 | 61.396 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727558 | Mycobacterium phage Salacia | Mycobacterium phage Salacia unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156610 | 64.679 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH825711 | Arthrobacter phage Tenno | Arthrobacter phage Tenno unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58193 | 49.626 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH825709 | Mycobacterium phage NicoleTera | Mycobacterium phage NicoleTera unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52944 | 63.973 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825707 | Mycobacterium phage MilleniumForce | Mycobacterium phage MilleniumForce unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58106 | 61.782 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825705 | Mycobacterium phage Mesh1 | Mycobacterium phage Mesh1 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68774 | 66.438 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH825704 | Mycobacterium phage LilTurb | Mycobacterium phage LilTurb unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53153 | 63.342 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825703 | Arthrobacter phage LeeroyJ | Arthrobacter phage LeeroyJ unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51025 | 60.941 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MH825702 | Mycobacterium phage Hamish | Mycobacterium phage Hamish unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68584 | 66.51 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH825701 | Mycobacterium phage Flare16 | Mycobacterium phage Flare16 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52858 | 63.472 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825700 | Microbacterium phage Efeko | Microbacterium phage Efeko Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17491 | 68.59 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH825699 | Streptomyces phage Darolandstone | Streptomyces phage Darolandstone Streptomyces virus Darolandstone Raleighvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40725 | 72.309 | Raleighvirus | Unclassified | Siphoviridae | Streptomyces |
| MH825698 | Microbacterium phage Burro | Microbacterium phage Burro Microbacterium virus Burro Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54473 | 64.338 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| MH825697 | Mycobacterium phage Bangla1971 | Mycobacterium phage Bangla1971 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154722 | 64.737 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH779517 | Mycobacterium phage Zulu | Mycobacterium phage Zulu unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52499 | 61.418 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779516 | Mycobacterium phage Waterdiva | Mycobacterium phage Waterdiva unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68866 | 66.505 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH779512 | Microbacterium phage Miaurora | Microbacterium phage Miaurora Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17032 | 68.982 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH779511 | Mycobacterium phage LilDestine | Mycobacterium phage LilDestine unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75440 | 58.946 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779507 | Mycobacterium phage Hotshotbaby7 | Mycobacterium phage Hotshotbaby7 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41903 | 66.616 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779504 | Gordonia phage Getalong | Gordonia phage Getalong Gordonia virus Getalong Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56214 | 62.821 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MH779503 | Mycobacterium phage Gemini | Mycobacterium phage Gemini unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75649 | 62.92 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779502 | Mycobacterium phage FudgeTart | Mycobacterium phage FudgeTart unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154658 | 64.795 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH779500 | Mycobacterium phage Crownjwl | Mycobacterium phage Crownjwl unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68521 | 66.561 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH779498 | Mycobacterium phage Bread | Mycobacterium phage Bread unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153796 | 64.787 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH744423 | Mycobacterium phage Saguaro | Mycobacterium phage Saguaro unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69448 | 69.538 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH744422 | Mycobacterium phage Roots515 | Mycobacterium phage Roots515 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156288 | 64.699 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH744421 | Arthrobacter phage Moki | Arthrobacter phage Moki unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43161 | 61.335 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MH744420 | Streptomyces phage Kromp | Streptomyces phage Kromp Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58268 | 71.435 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH744418 | Streptomyces phage JustBecause | Streptomyces phage JustBecause Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 184281 | 66.565 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH744417 | Mycobacterium phage Grand2040 | Mycobacterium phage Grand2040 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68591 | 66.443 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH744414 | Mycobacterium phage Arcanine | Mycobacterium phage Arcanine unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52273 | 63.704 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727564 | Streptomyces phage Yaboi | Streptomyces phage Yaboi Streptomyces virus Yaboi Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131250 | 49.301 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH727562 | Mycobacterium phage VasuNzinga | Mycobacterium phage VasuNzinga unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64911 | 63.398 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727561 | Mycobacterium phage Serendipitous | Mycobacterium phage Serendipitous unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69867 | 68.008 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH727557 | Mycobacterium phage Paperbeatsrock | Mycobacterium phage Paperbeatsrock unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75640 | 63.02 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727556 | Microbacterium phage OneinaGillian | Microbacterium phage OneinaGillian Microbacterium virus OneinaGillian Gillianvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61703 | 67.071 | Gillianvirus | Unclassified | Siphoviridae | Microbacterium |
| MH727555 | Mycobacterium phage Mulan | Mycobacterium phage Mulan unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68521 | 66.549 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH727553 | Mycobacterium phage LittleLaf | Mycobacterium phage LittleLaf unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64834 | 63.368 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727551 | Mycobacterium phage Kalah2 | Mycobacterium phage Kalah2 unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110713 | 60.987 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727550 | Corynebacterium phage Juicebox | Corynebacterium phage Juicebox Corynebacterium virus Juicebox Juiceboxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41394 | 67.858 | Juiceboxvirus | Unclassified | Siphoviridae | Corynebacterium |
| MH727549 | Mycobacterium phage Gancho | Mycobacterium phage Gancho unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54125 | 67.52 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727546 | Mycobacterium phage Eaglehorse | Mycobacterium phage Eaglehorse unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67391 | 68.956 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH697590 | Mycobacterium phage Phlegm | Mycobacterium phage Phlegm unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155959 | 64.673 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH697586 | Mycobacterium phage InigoMontoya | Mycobacterium phage InigoMontoya unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155217 | 64.709 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH651190 | Streptomyces phage Thestral | Streptomyces phage Thestral unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52628 | 67.677 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH651189 | Gordonia phage Schmidt | Gordonia phage Schmidt unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43099 | 65.67 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MH651188 | Mycobacterium phage Ruby | Mycobacterium phage Ruby unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57726 | 61.367 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651187 | Mycobacterium phage Renaud18 | Mycobacterium phage Renaud18 Mycobacterium virus Renaud18 Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58078 | 61.665 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651186 | Mycobacterium phage Podrick | Mycobacterium phage Podrick unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68406 | 66.425 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH651184 | Mycobacterium phage Phareon | Mycobacterium phage Phareon unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68040 | 66.484 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH651183 | Mycobacterium phage Nivrat | Mycobacterium phage Nivrat unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58009 | 61.454 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651182 | Gordonia phage Neville | Gordonia phage Neville Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75813 | 58.752 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH651181 | Microbacterium phage Minima | Microbacterium phage Minima Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.512 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH651180 | Mycobacterium phage Maroc7 | Mycobacterium phage Maroc7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53350 | 63.5 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651178 | Mycobacterium phage KristaRAM | Mycobacterium phage KristaRAM unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58701 | 61.643 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651177 | Mycobacterium phage KlimbOn | Mycobacterium phage KlimbOn unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68596 | 66.453 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH651175 | Mycobacterium phage Gophee | Mycobacterium phage Gophee unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68856 | 66.508 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH651174 | Mycobacterium phage Easy2Say | Mycobacterium phage Easy2Say unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75598 | 62.924 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651173 | Mycobacterium phage Dixon | Mycobacterium phage Dixon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52055 | 61.395 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651172 | Streptomyces phage Comrade | Streptomyces phage Comrade Streptomyces virus Comrade Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129015 | 47.06 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MH651169 | Mycobacterium phage Burwell21 | Mycobacterium phage Burwell21 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58098 | 61.458 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651168 | Gordonia phage Ashertheman | Gordonia phage Ashertheman unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57555 | 68.031 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH651167 | Gordonia phage Angelique | Gordonia phage Angelique Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50970 | 66.643 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH651166 | Gordonia phage Angelicage | Gordonia phage Angelicage unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59215 | 67.447 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH669017 | Mycobacterium phage Zelink | Mycobacterium phage Zelink unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111740 | 60.828 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669016 | Streptomyces phage TrvxScott | Streptomyces phage TrvxScott unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52600 | 67.793 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH669015 | Gordonia phage TillyBobJoe | Gordonia phage TillyBobJoe unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58677 | 67.761 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MH669011 | Mycobacterium phage PherrisBueller | Mycobacterium phage PherrisBueller unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49209 | 63.958 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669010 | Mycobacterium phage PHappiness | Mycobacterium phage PHappiness unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57989 | 61.603 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669009 | Mycobacterium phage Nimbo | Mycobacterium phage Nimbo unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55767 | 61.513 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669008 | Mycobacterium phage Megamind | Mycobacterium phage Megamind unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154780 | 64.758 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH669006 | Streptomyces phage LibertyBell | Streptomyces phage LibertyBell Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52733 | 59.119 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH669005 | Mycobacterium phage JoeyJr | Mycobacterium phage JoeyJr unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58962 | 61.445 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669004 | Streptomyces phage Hank144 | Streptomyces phage Hank144 unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50547 | 66.307 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH669003 | Mycobacterium phage Girr | Mycobacterium phage Girr unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57754 | 61.355 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669002 | Mycobacterium phage Emmina | Mycobacterium phage Emmina unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75295 | 62.886 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669000 | Gordonia phage Ali17 | Gordonia phage Ali17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58301 | 68.404 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH697591 | Mycobacterium phage QueenBeesly | Mycobacterium phage QueenBeesly unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53348 | 63.524 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697584 | Mycobacterium phage FrenchFry | Mycobacterium phage FrenchFry unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67494 | 68.948 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH697583 | Mycobacterium phage EricMillard | Mycobacterium phage EricMillard unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113536 | 60.933 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697582 | Mycobacterium phage Ejimix | Mycobacterium phage Ejimix unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111924 | 60.895 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697581 | Mycobacterium phage Crispicous1 | Mycobacterium phage Crispicous1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50827 | 63.694 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697580 | Gordonia phage Catfish | Gordonia phage Catfish unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46888 | 64.976 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MH697579 | Mycobacterium phage Cane17 | Mycobacterium phage Cane17 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160330 | 64.613 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH697577 | Mycobacterium phage Amochick | Mycobacterium phage Amochick unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54145 | 67.434 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697575 | Mycobacterium phage Adahisdi | Mycobacterium phage Adahisdi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51703 | 63.801 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590593 | Mycobacterium phage Rabinovish | Mycobacterium phage Rabinovish unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154861 | 64.69 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH632120 | Mycobacterium phage Thonko | Mycobacterium phage Thonko unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69471 | 68.633 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH632119 | Mycobacterium phage Harley | Mycobacterium phage Harley unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58731 | 61.422 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH632118 | Mycobacterium phage Zeeculate | Mycobacterium phage Zeeculate unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53828 | 63.837 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH632117 | Mycobacterium phage Homines | Mycobacterium phage Homines unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48502 | 63.83 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590605 | Mycobacterium phage Cherrybomb426 | Mycobacterium phage Cherrybomb426 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41456 | 66.68 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590603 | Gordonia phage Daredevil | Gordonia phage Daredevil Gordonia virus DareDevil Daredevilvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90490 | 64.843 | Daredevilvirus | Unclassified | Siphoviridae | Gordonia |
| MH590602 | Mycobacterium phage Gorge | Mycobacterium phage Gorge unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57955 | 61.38 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590601 | Streptomyces phage Haizum | Streptomyces phage Haizum unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50660 | 66.842 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH590600 | Microbacterium phage KaiHaiDragon | Microbacterium phage KaiHaiDragon Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52992 | 68.901 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MH590599 | Streptomyces phage Karimac | Streptomyces phage Karimac Streptomyces virus Karimac Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131909 | 49.411 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH590598 | Mycobacterium phage Krakatau | Mycobacterium phage Krakatau unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53058 | 61.501 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590597 | Streptomyces phage LukeCage | Streptomyces phage LukeCage Streptomyces virus LukeCage Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133195 | 49.05 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH590594 | Mycobacterium phage PinheadLarry | Mycobacterium phage PinheadLarry unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68390 | 66.435 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH590592 | Mycobacterium phage Ryadel | Mycobacterium phage Ryadel unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72658 | 65.207 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590591 | Mycobacterium phage Sabella | Mycobacterium phage Sabella unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67304 | 68.935 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH590589 | Streptomyces phage SparkleGoddess | Streptomyces phage SparkleGoddess unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129742 | 47.063 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MH590588 | Mycobacterium phage Vaticameos | Mycobacterium phage Vaticameos unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66887 | 66.535 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH590587 | Mycobacterium phage xkcd | Mycobacterium phage xkcd unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76915 | 62.838 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576977 | Mycobacterium phage Tydolla | Mycobacterium phage Tydolla unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68657 | 67.48 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH576974 | Mycobacterium phage Sibs6 | Mycobacterium phage Sibs6 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50210 | 63.766 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576972 | Gordonia phage Beyoncage | Gordonia phage Beyoncage unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66610 | 59.551 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MH576971 | Mycobacterium phage Arlo | Mycobacterium phage Arlo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52960 | 63.75 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576970 | Mycobacterium phage DonSanchon | Mycobacterium phage DonSanchon unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68015 | 66.515 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH576968 | Streptomyces phage Wollford | Streptomyces phage Wollford Streptomyces virus Wollford Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133007 | 47.731 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH576967 | Mycobacterium phage UAch1 | Mycobacterium phage UAch1 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68405 | 66.445 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH576965 | Streptomyces phage StarPlatinum | Streptomyces phage StarPlatinum Streptomyces virus StarPlatinum Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133886 | 49.466 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH576964 | Streptomyces phage Starbow | Streptomyces phage Starbow unclassified Karimacvirus Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131427 | 49.504 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| MH576963 | Microbacterium phage Scamander | Microbacterium phage Scamander Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17452 | 68.731 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH576962 | Streptomyces phage Satis | Streptomyces phage Satis Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186702 | 66.685 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH576958 | Mycobacterium phage MichaelPhcott | Mycobacterium phage MichaelPhcott unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68494 | 66.353 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH576956 | Mycobacterium phage Inca | Mycobacterium phage Inca unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74070 | 62.801 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576954 | Mycobacterium phage HSavage | Mycobacterium phage HSavage unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67993 | 66.52 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH576953 | Mycobacterium phage Hopey | Mycobacterium phage Hopey unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75586 | 62.988 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513985 | Mycobacterium phage Tomathan | Mycobacterium phage Tomathan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52876 | 63.397 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513983 | Mycobacterium phage ShereKhan | Mycobacterium phage ShereKhan unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76133 | 63.062 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513980 | Mycobacterium phage Roy17 | Mycobacterium phage Roy17 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68056 | 66.582 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH513978 | Mycobacterium phage Phaja | Mycobacterium phage Phaja unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75685 | 62.866 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513976 | Mycobacterium phage Morty007 | Mycobacterium phage Morty007 unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69431 | 67.48 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH513974 | Gordonia phage LastResort | Gordonia phage LastResort unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52619 | 61.981 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MH513973 | Mycobacterium phage Kwksand96 | Mycobacterium phage Kwksand96 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68822 | 66.351 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH513972 | Mycobacterium phage IHOP | Mycobacterium phage IHOP unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75668 | 62.898 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513971 | Mycobacterium phage Hope4ever | Mycobacterium phage Hope4ever unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50455 | 63.583 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513970 | Mycobacterium phage Hangman | Mycobacterium phage Hangman unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71376 | 68.854 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH536829 | Mycobacterium phage TBrady12 | Mycobacterium phage TBrady12 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75359 | 62.837 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536828 | Mycobacterium phage Snape | Mycobacterium phage Snape unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52138 | 63.737 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536827 | Mycobacterium phage Simpliphy | Mycobacterium phage Simpliphy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75366 | 63.028 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536826 | Gordonia phage Ruthy | Gordonia phage Ruthy Gordonia virus Ruthy Ruthyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51265 | 66.681 | Ruthyvirus | Unclassified | Siphoviridae | Gordonia |
| MH536820 | Mycobacterium phage Glexan | Mycobacterium phage Glexan unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76498 | 62.94 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536816 | Mycobacterium phage DrFeelGood | Mycobacterium phage DrFeelGood unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50155 | 63.91 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536814 | Streptomyces phage Blueeyedbeauty | Streptomyces phage Blueeyedbeauty Streptomyces virus Blueeyedbeauty Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130473 | 47.88 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MH509447 | Microbacterium phage Eden | Microbacterium phage Eden Microbacterium virus Eden Edenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40833 | 66.282 | Edenvirus | Unclassified | Siphoviridae | Microbacterium |
| MH509446 | Gordonia phage DelRio | Gordonia phage DelRio unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50961 | 66.985 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MH479926 | Arthrobacter phage Synepsis | Arthrobacter phage Synepsis unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57542 | 49.948 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH479924 | Gordonia phage Rofo | Gordonia phage Rofo unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58767 | 67.291 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH479921 | Mycobacterium phage Placalicious | Mycobacterium phage Placalicious unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69085 | 66.46 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH479920 | Mycobacterium phage Mowgli | Mycobacterium phage Mowgli unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41902 | 66.589 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH479919 | Mycobacterium phage Moose | Mycobacterium phage Moose unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52695 | 63.708 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH479918 | Mycobacterium phage Labeouficaum | Mycobacterium phage Labeouficaum unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68637 | 66.374 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH479917 | Gordonia phage KimmyK | Gordonia phage KimmyK unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58755 | 67.768 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MH479916 | Mycobacterium phage GeneCoco | Mycobacterium phage GeneCoco unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68547 | 66.36 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH479915 | Gordonia phage GEazy | Gordonia phage GEazy unclassified Bowservirus Bowservirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46728 | 66.949 | Bowservirus | Unclassified | Siphoviridae | Gordonia |
| MH479914 | Mycobacterium phage FugateOSU | Mycobacterium phage FugateOSU unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69476 | 66.459 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH479910 | Gordonia phage Danyall | Gordonia phage Danyall unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58695 | 67.757 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| MH450133 | Mycobacterium phage SwissCheese | Mycobacterium phage SwissCheese unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51822 | 63.544 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450132 | Gordonia phage SketchMex | Gordonia phage SketchMex unclassified Emalynvirus Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45847 | 60.497 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MH450131 | Arthrobacter phage Sergei | Arthrobacter phage Sergei unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43950 | 61.863 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450130 | Mycobacterium phage Rohr | Mycobacterium phage Rohr unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53483 | 63.559 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450129 | Gordonia phage Ribeye | Gordonia phage Ribeye unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57448 | 68.14 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH450127 | Mycobacterium phage Plagueis | Mycobacterium phage Plagueis unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41722 | 66.634 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450125 | Arthrobacter phage MediumFry | Arthrobacter phage MediumFry unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56978 | 50.874 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450122 | Mycobacterium phage KingTut | Mycobacterium phage KingTut unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64827 | 66.462 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH450121 | Arthrobacter phage KingBob | Arthrobacter phage KingBob unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43950 | 61.857 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450118 | Arthrobacter phage Herb | Arthrobacter phage Herb unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43949 | 61.86 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450117 | Arthrobacter phage Daiboju | Arthrobacter phage Daiboju unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43950 | 61.863 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450116 | Mycobacterium phage Buckeye | Mycobacterium phage Buckeye unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69174 | 66.453 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH450115 | Arthrobacter phage Breylor17 | Arthrobacter phage Breylor17 unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57822 | 49.934 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH450113 | Mycobacterium phage BigMau | Mycobacterium phage BigMau unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52632 | 63.725 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536811 | Streptomyces phage Annadreamy | Streptomyces phage Annadreamy Streptomyces virus Annadreamy Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125726 | 47.604 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| MH399772 | Mycobacterium phage Constella | Mycobacterium phage Constella unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110169 | 60.82 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399789 | Arthrobacter phage Tatanka | Arthrobacter phage Tatanka unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58120 | 49.981 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH399788 | Mycobacterium phage Visconti | Mycobacterium phage Visconti Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64570 | 59.684 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MH399786 | Mycobacterium phage PhrodoBaggins | Mycobacterium phage PhrodoBaggins unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68873 | 66.415 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH399784 | Mycobacterium phage NormanBulbieJr | Mycobacterium phage NormanBulbieJr unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58102 | 61.277 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399783 | Microbacterium phage Noelani | Microbacterium phage Noelani Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17349 | 68.154 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH399782 | Mycobacterium phage Niza | Mycobacterium phage Niza unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51672 | 63.448 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399781 | Gordonia phage Nadeem | Gordonia phage Nadeem unclassified Betterkatzvirus Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49897 | 67.289 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| MH399780 | Mycobacterium phage Mutante | Mycobacterium phage Mutante unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68494 | 66.509 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH399778 | Mycobacterium phage Icee | Mycobacterium phage Icee unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74169 | 63.002 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399775 | Mycobacterium phage Gareth | Mycobacterium phage Gareth unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68866 | 66.468 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH399774 | Mycobacterium phage DoctorDiddles | Mycobacterium phage DoctorDiddles unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75070 | 62.815 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399773 | Mycobacterium phage Craff | Mycobacterium phage Craff unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69263 | 66.432 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH371107 | Mycobacterium phage Doddsville | Mycobacterium phage Doddsville unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68243 | 66.417 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH371111 | Microbacterium phage Quaker | Microbacterium phage Quaker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17450 | 68.636 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH371112 | Mycobacterium phage Adnama | Mycobacterium phage Adnama unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75208 | 62.886 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371114 | Mycobacterium phage Childish | Mycobacterium phage Childish unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69044 | 66.478 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH371115 | Mycobacterium phage Clifton | Mycobacterium phage Clifton unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55727 | 61.658 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371116 | Mycobacterium phage DMoney | Mycobacterium phage DMoney unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41880 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371117 | Mycobacterium phage Kahve | Mycobacterium phage Kahve unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69031 | 66.486 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH371118 | Microbacterium phage KayPaulus | Microbacterium phage KayPaulus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17455 | 68.536 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH371119 | Mycobacterium phage OctaviousRex | Mycobacterium phage OctaviousRex unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41880 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371120 | Mycobacterium phage OlympiaSaint | Mycobacterium phage OlympiaSaint unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58297 | 61.399 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371122 | Mycobacterium phage Priya | Mycobacterium phage Priya unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52949 | 62.532 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371123 | Mycobacterium phage Sandalphon | Mycobacterium phage Sandalphon unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59540 | 61.407 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371124 | Microbacterium phage VitulaEligans | Microbacterium phage VitulaEligans Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17534 | 68.769 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH371125 | Mycobacterium phage Morty | Mycobacterium phage Morty unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68878 | 66.506 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH338241 | Mycobacterium phage Tarynearal | Mycobacterium phage Tarynearal unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51143 | 60.9 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338239 | Mycobacterium phage Mryolo | Mycobacterium phage Mryolo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50505 | 63.974 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338238 | Mycobacterium phage Michley | Mycobacterium phage Michley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51527 | 63.706 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338237 | Mycobacterium phage Joselito | Mycobacterium phage Joselito unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52347 | 63.694 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338236 | Mycobacterium phage Ebony | Mycobacterium phage Ebony unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52152 | 63.771 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338235 | Mycobacterium phage Dublin | Mycobacterium phage Dublin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50280 | 60.903 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338234 | Mycobacterium phage Crucio | Mycobacterium phage Crucio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53256 | 63.491 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH316567 | Mycobacterium phage ParkTD | Mycobacterium phage ParkTD unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155006 | 64.756 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH316561 | Mycobacterium phage Derek | Mycobacterium phage Derek unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156199 | 64.685 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH248947 | Streptomyces phage Yosif | Streptomyces phage Yosif unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50129 | 66.117 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH316568 | Mycobacterium phage Phleuron | Mycobacterium phage Phleuron unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68527 | 66.368 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH316566 | Gordonia phage Nedarya | Gordonia phage Nedarya unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53140 | 62.036 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MH316565 | Mycobacterium phage Mortcellus | Mycobacterium phage Mortcellus unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69800 | 67.463 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH316564 | Mycobacterium phage Koella | Mycobacterium phage Koella unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54118 | 61.499 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH316563 | Mycobacterium phage Hookmount | Mycobacterium phage Hookmount unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50061 | 64.062 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH316562 | Mycobacterium phage Erk16 | Mycobacterium phage Erk16 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64877 | 59.625 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MH271321 | Microbacterium phage Quhwah | Microbacterium phage Quhwah Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53549 | 68.847 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MH271320 | Microbacterium phage Zeta1847 | Microbacterium phage Zeta1847 Microbacterium virus Zeta1847 Zetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47921 | 71.415 | Zetavirus | Unclassified | Siphoviridae | Microbacterium |
| MH271313 | Gordonia phage Suzy | Gordonia phage Suzy Gordonia virus Suzy Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67118 | 59.094 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MH271312 | Microbacterium phage RobsFeet | Microbacterium phage RobsFeet Microbacterium virus RobsFeet Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54189 | 68.602 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MH271308 | Microbacterium phage Percival | Microbacterium phage Percival Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47364 | 69.622 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271304 | Microbacterium phage Metamorphoo | Microbacterium phage Metamorphoo Microbacterium virus Metamorphoo Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54148 | 68.527 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| MH271303 | Microbacterium phage MementoMori | Microbacterium phage MementoMori Microbacterium virus MementoMori Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55572 | 69.978 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| MH271302 | Gordonia phage Margaret | Gordonia phage Margaret Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46950 | 62.775 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH271301 | Microbacterium phage Krampus | Microbacterium phage Krampus Microbacterium virus Krampus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56708 | 63.799 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MH271298 | Microbacterium phage Floof | Microbacterium phage Floof Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48500 | 69.023 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271295 | Gordonia phage Ebert | Gordonia phage Ebert unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46653 | 66.611 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MH271292 | Microbacterium phage AnnaSerena | Microbacterium phage AnnaSerena unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56707 | 63.809 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153809 | Gordonia phage Sitar | Gordonia phage Sitar unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59641 | 67.331 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MH178382 | Streptomyces phage Salete | Streptomyces phage Salete unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57243 | 69.228 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| MH178381 | Streptomyces phage BayC | Streptomyces phage BayC unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57243 | 69.226 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| MH183162 | Microbacterium phage Hendrix | Microbacterium phage Hendrix Microbacterium virus Hendrix Rogerhendrixvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97757 | 66.075 | Rogerhendrixvirus | Unclassified | Siphoviridae | Microbacterium |
| MH171098 | Streptomyces phage Toma | Streptomyces phage Toma unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51396 | 65.813 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH171097 | Streptomyces phage Goby | Streptomyces phage Goby unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51393 | 65.803 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH171096 | Streptomyces phage Eddasa | Streptomyces phage Eddasa unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50605 | 65.912 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH171095 | Streptomyces phage Maneekul | Streptomyces phage Maneekul unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51612 | 65.688 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH171094 | Streptomyces phage Nabi | Streptomyces phage Nabi unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51127 | 65.77 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH171093 | Streptomyces phage Rana | Streptomyces phage Rana unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50980 | 65.785 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH155880 | Streptomyces phage SendItCS | Streptomyces phage SendItCS unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55993 | 58.159 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MH155877 | Streptomyces phage Rainydai | Streptomyces phage Rainydai unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57623 | 58.076 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MH155876 | Mycobacterium phage Priamo | Mycobacterium phage Priamo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51633 | 61.391 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH155878 | Streptomyces phage RavenPuff | Streptomyces phage RavenPuff unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43709 | 60.903 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MH155874 | Mycobacterium phage PhrankReynolds | Mycobacterium phage PhrankReynolds unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68721 | 66.32 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH155873 | Microbacterium phage Paschalis | Microbacterium phage Paschalis Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52935 | 68.837 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MH155872 | Streptomyces phage Moozy | Streptomyces phage Moozy unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43545 | 61.178 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MH155869 | Streptomyces phage HotFries | Streptomyces phage HotFries unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43699 | 60.908 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MH155866 | Mycobacterium phage Byougenkin | Mycobacterium phage Byougenkin unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54685 | 61.425 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH153813 | Microbacterium phage Squash | Microbacterium phage Squash Microbacterium virus Squash Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62312 | 66.788 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153812 | Rhodococcus phage Grayson | Rhodococcus phage Grayson unclassified Weaselvirus Weaselvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131801 | 41.246 | Weaselvirus | Unclassified | Siphoviridae | Rhodococcus |
| MH153810 | Gordonia phage Sour | Gordonia phage Sour Gordonia virus Sour Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61670 | 68.009 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MH153808 | Gordonia phage Petra | Gordonia phage Petra Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52304 | 67.083 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MH153807 | Rhodococcus phage Peregrin | Rhodococcus phage Peregrin unclassified Weaselvirus Weaselvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133006 | 41.442 | Weaselvirus | Unclassified | Siphoviridae | Rhodococcus |
| MH153804 | Gordonia phage Jace | Gordonia phage Jace Gordonia virus Jace Jacevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53912 | 66.981 | Jacevirus | Unclassified | Siphoviridae | Gordonia |
| MH153803 | Microbacterium phage Hyperion | Microbacterium phage Hyperion Microbacterium virus Hyperion Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61769 | 67.009 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| MH230879 | Mycobacterium phage Xavia | Mycobacterium phage Xavia unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49808 | 65.859 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| MH230878 | Mycobacterium phage Oogway | Mycobacterium phage Oogway unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51745 | 63.573 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH230876 | Mycobacterium phage ConceptII | Mycobacterium phage ConceptII unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53287 | 63.522 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH230875 | Mycobacterium phage CheetO | Mycobacterium phage CheetO unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68503 | 66.523 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH230874 | Mycobacterium phage Banjo | Mycobacterium phage Banjo unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68296 | 66.468 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH077584 | Mycobacterium phage Phish | Mycobacterium phage Phish unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.559 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077582 | Mycobacterium phage Olive | Mycobacterium phage Olive unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68740 | 66.525 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH077580 | Mycobacterium phage Melissauren88 | Mycobacterium phage Melissauren88 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54356 | 61.783 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077579 | Mycobacterium phage Halley | Mycobacterium phage Halley unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112351 | 60.797 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077578 | Mycobacterium phage DillTech15 | Mycobacterium phage DillTech15 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57797 | 61.861 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077576 | Mycobacterium phage AbbyPaige | Mycobacterium phage AbbyPaige unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53225 | 63.371 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051261 | Mycobacterium phage Tyke | Mycobacterium phage Tyke unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156679 | 64.651 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH051260 | Mycobacterium phage InterFolia | Mycobacterium phage InterFolia unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156221 | 64.697 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH051257 | Mycobacterium phage OwlsT2W | Mycobacterium phage OwlsT2W unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56515 | 61.219 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051253 | Mycobacterium phage GreedyLawyer | Mycobacterium phage GreedyLawyer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52091 | 61.531 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051252 | Mycobacterium phage Fameo | Mycobacterium phage Fameo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52618 | 63.408 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051251 | Mycobacterium phage DuchessDung | Mycobacterium phage DuchessDung unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68235 | 66.353 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH051249 | Mycobacterium phage Boyle | Mycobacterium phage Boyle unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67491 | 68.963 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH051248 | Mycobacterium phage BigPhil | Mycobacterium phage BigPhil unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53618 | 61.496 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051266 | Mycobacterium phage MikeLiesIn | Mycobacterium phage MikeLiesIn unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155916 | 64.708 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MH045568 | Microbacterium phage Kieran | Microbacterium phage Kieran unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41417 | 69.715 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045569 | Mycobacterium phage Schiebel | Mycobacterium phage Schiebel unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41441 | 66.688 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH045567 | Microbacterium phage Elva | Microbacterium phage Elva unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42139 | 68.165 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045566 | Microbacterium phage Didgeridoo | Microbacterium phage Didgeridoo unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42655 | 66.067 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045561 | Microbacterium phage PaoPu | Microbacterium phage PaoPu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.529 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH045557 | Microbacterium phage BurtonThePup | Microbacterium phage BurtonThePup Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17445 | 68.753 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH045556 | Microbacterium phage BonaeVitae | Microbacterium phage BonaeVitae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17451 | 68.22 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH051265 | Mycobacterium phage Yahalom | Mycobacterium phage Yahalom unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68482 | 67.438 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH051264 | Mycobacterium phage Cobra | Mycobacterium phage Cobra unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68875 | 66.412 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH025890 | Gordonia phage Djokovic | Gordonia phage Djokovic unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66609 | 59.552 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MH025888 | Mycobacterium phage BigMama | Mycobacterium phage BigMama Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64592 | 59.698 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MH020248 | Mycobacterium phage Panamaxus | Mycobacterium phage Panamaxus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50080 | 64.073 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020247 | Mycobacterium phage MPhalcon | Mycobacterium phage MPhalcon unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75605 | 63.078 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020246 | Mycobacterium phage SchoolBus | Mycobacterium phage SchoolBus unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71651 | 65.417 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020244 | Mycobacterium phage Lopton | Mycobacterium phage Lopton unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52394 | 63.544 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020240 | Mycobacterium phage SuperAwesome | Mycobacterium phage SuperAwesome unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52686 | 61.595 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020237 | Mycobacterium phage Jsquared | Mycobacterium phage Jsquared unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52967 | 62.59 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020236 | Mycobacterium phage GageAP | Mycobacterium phage GageAP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51904 | 63.465 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH019216 | Streptomyces phage Wentworth | Streptomyces phage Wentworth Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68260 | 64.122 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH019215 | Streptomyces phage Yara | Streptomyces phage Yara Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68671 | 63.89 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MG962375 | Mycobacterium phage ProfessorX | Mycobacterium phage ProfessorX unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68086 | 66.545 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG962372 | Mycobacterium phage McGuire | Mycobacterium phage McGuire unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51521 | 63.731 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG962370 | Mycobacterium phage Kykar | Mycobacterium phage Kykar unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50294 | 63.656 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG962369 | Gordonia phage Kerry | Gordonia phage Kerry unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59608 | 51.062 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MG962368 | Gordonia phage Gravy | Gordonia phage Gravy unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59545 | 51.071 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MG962366 | Rhodococcus phage Finch | Rhodococcus phage Finch Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138896 | 63.079 | Unclassified | Unclassified | Myoviridae | Rhodococcus |
| MG962365 | Mycobacterium phage DoesntMatter | Mycobacterium phage DoesntMatter unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68331 | 66.516 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG962363 | Mycobacterium phage BatteryCK | Mycobacterium phage BatteryCK unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68509 | 66.35 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG962362 | Mycobacterium phage AltPhacts | Mycobacterium phage AltPhacts unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68056 | 66.475 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG962361 | Mycobacterium phage Alexphander | Mycobacterium phage Alexphander unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57734 | 61.163 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH000607 | Mycobacterium phage RiverMonster | Mycobacterium phage RiverMonster unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75565 | 63.052 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH000606 | Mycobacterium phage Fascinus | Mycobacterium phage Fascinus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52157 | 63.704 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001460 | Streptomyces phage Paedore | Streptomyces phage Paedore unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52743 | 66.998 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MH001458 | Mycobacterium phage Smeagol | Mycobacterium phage Smeagol unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53134 | 63.675 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001457 | Streptomyces phage Nesbitt | Streptomyces phage Nesbitt unclassified Abbeymikolonvirus Abbeymikolonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42446 | 66.831 | Abbeymikolonvirus | Unclassified | Siphoviridae | Streptomyces |
| MH001456 | Mycobacterium phage CLED96 | Mycobacterium phage CLED96 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41456 | 66.678 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001455 | Mycobacterium phage Remy19 | Mycobacterium phage Remy19 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.566 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001454 | Gordonia phage Waits | Gordonia phage Waits Gordonia virus Waits Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52856 | 61.972 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MH001452 | Mycobacterium phage Kheth | Mycobacterium phage Kheth unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67406 | 68.936 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MH001450 | Mycobacterium phage Jeon | Mycobacterium phage Jeon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60908 | 67.566 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MH001449 | Mycobacterium phage MooMoo | Mycobacterium phage MooMoo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55178 | 61.969 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MH001446 | Mycobacterium phage Megabear | Mycobacterium phage Megabear Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60579 | 67.52 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MG944223 | Mycobacterium phage Trypo | Mycobacterium phage Trypo unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68798 | 66.407 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG944225 | Mycobacterium phage Xavier | Mycobacterium phage Xavier unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68493 | 66.433 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG944221 | Mycobacterium phage Scowl | Mycobacterium phage Scowl unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51690 | 63.784 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG944220 | Mycobacterium phage Ruotula | Mycobacterium phage Ruotula unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52601 | 63.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925356 | Mycobacterium phage OliverWalter | Mycobacterium phage OliverWalter unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68792 | 66.514 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925353 | Mycobacterium phage Nozo | Mycobacterium phage Nozo unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69439 | 67.484 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925351 | Mycobacterium phage MPlant7149 | Mycobacterium phage MPlant7149 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.655 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925350 | Mycobacterium phage Mosaic | Mycobacterium phage Mosaic unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68488 | 66.511 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925348 | Mycobacterium phage Megatron | Mycobacterium phage Megatron unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69009 | 66.506 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925346 | Mycobacterium phage LeeLot | Mycobacterium phage LeeLot unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68921 | 66.467 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925344 | Mycobacterium phage Ichabod | Mycobacterium phage Ichabod unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53317 | 63.751 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925342 | Mycobacterium phage Forsytheast | Mycobacterium phage Forsytheast unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52695 | 63.708 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925354 | Mycobacterium phage Ogopogo | Mycobacterium phage Ogopogo unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56867 | 61.139 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925340 | Mycobacterium phage Corvo | Mycobacterium phage Corvo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53325 | 63.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925339 | Mycobacterium phage Chunky | Mycobacterium phage Chunky unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68249 | 66.476 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG925338 | Mycobacterium phage ChaChing | Mycobacterium phage ChaChing unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68668 | 67.505 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG920060 | Mycobacterium phage Bob3 | Mycobacterium phage Bob3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52233 | 63.833 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG920059 | Mycobacterium phage Baloo | Mycobacterium phage Baloo unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68514 | 67.484 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG812494 | Mycobacterium phage Pier | Mycobacterium phage Pier unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156471 | 64.712 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MG812493 | Mycobacterium phage NuevoMundo | Mycobacterium phage NuevoMundo unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155943 | 64.745 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MG872839 | Mycobacterium phage LifeSavor | Mycobacterium phage LifeSavor unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156804 | 64.632 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MG872843 | Mycobacterium phage Sotrice96 | Mycobacterium phage Sotrice96 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76299 | 63.058 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872842 | Mycobacterium phage Scherzo | Mycobacterium phage Scherzo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53355 | 62.667 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872841 | Mycobacterium phage Priscilla | Mycobacterium phage Priscilla unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58230 | 61.278 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872840 | Mycobacterium phage Phonnegut | Mycobacterium phage Phonnegut unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53219 | 62.594 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872838 | Mycobacterium phage JangDynasty | Mycobacterium phage JangDynasty unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70883 | 65.426 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872837 | Mycobacterium phage Gage | Mycobacterium phage Gage unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71650 | 63.197 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872836 | Mycobacterium phage Fajezeel | Mycobacterium phage Fajezeel unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 63.411 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872835 | Mycobacterium phage Conquerage | Mycobacterium phage Conquerage unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52953 | 62.533 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872833 | Mycobacterium phage BeesKnees | Mycobacterium phage BeesKnees unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51702 | 63.715 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872832 | Mycobacterium phage Barbarian | Mycobacterium phage Barbarian unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74594 | 62.787 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872831 | Mycobacterium phage Asriel | Mycobacterium phage Asriel unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74594 | 62.788 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG757167 | Mycobacterium phage Tesla | Mycobacterium phage Tesla unclassified Marvinvirus Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64625 | 63.451 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG757165 | Mycobacterium phage Phergie | Mycobacterium phage Phergie unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68716 | 66.321 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG757164 | Mycobacterium phage PhenghisKhan | Mycobacterium phage PhenghisKhan unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68727 | 66.322 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG757163 | Streptomyces phage OzzyJ | Streptomyces phage OzzyJ unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51464 | 65.693 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MG757162 | Mycobacterium phage Opia | Mycobacterium phage Opia unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67401 | 68.923 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG757160 | Mycobacterium phage Kimchi | Mycobacterium phage Kimchi unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75829 | 62.87 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG757159 | Mycobacterium phage JangoPhett | Mycobacterium phage JangoPhett unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68673 | 66.476 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG757158 | Mycobacterium phage HighStump | Mycobacterium phage HighStump unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68821 | 66.456 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG757154 | Streptomyces phage Bing | Streptomyces phage Bing Streptomyces virus Bing Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56341 | 59.424 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MG757153 | Streptomyces phage BillNye | Streptomyces phage BillNye Streptomyces virus BillNye Wilnyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127084 | 52.833 | Wilnyevirus | Unclassified | Siphoviridae | Streptomyces |
| KP027207 | Mycobacterium phage Chandler | Mycobacterium phage Chandler unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69451 | 67.511 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX619650 | Mycobacterium phage Jerm | Mycobacterium phage Jerm unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53163 | 63.347 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX670790 | Streptomyces phage Rima | Streptomyces phage Rima Strepomyces virus Rima Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56168 | 59.557 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| KT347314 | Mycobacterium phage Phreak | Mycobacterium phage Phreak unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.566 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KM591906 | Mycobacterium phage SweetiePie | Mycobacterium phage SweetiePie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53184 | 63.472 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC691255 | Mycobacterium phage Dumbo | Mycobacterium phage Dumbo unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75736 | 62.996 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JX649097 | Mycobacterium phage Piglet | Mycobacterium phage Piglet Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68992 | 66.544 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN020141 | Mycobacterium phage Shauna1 | Mycobacterium phage Shauna1 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59315 | 61.708 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| EU826469 | Mycobacterium phage ScottMcG | Mycobacterium phage ScottMcG Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154017 | 64.829 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MG839014 | Mycobacterium phage RagingRooster | Mycobacterium phage RagingRooster unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68670 | 67.578 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG812492 | Mycobacterium phage Lambert1 | Mycobacterium phage Lambert1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50042 | 64.058 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG812491 | Mycobacterium phage ItsyBitsy1 | Mycobacterium phage ItsyBitsy1 unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67570 | 68.927 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG812490 | Mycobacterium phage Holeinone | Mycobacterium phage Holeinone unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67044 | 68.9 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG812488 | Mycobacterium phage Frankie | Mycobacterium phage Frankie unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57036 | 61.896 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG812487 | Gordonia phage Brandonk123 | Gordonia phage Brandonk123 Gordonia virus Brandonk123 Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58842 | 67.287 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MG812486 | Mycobacterium phage Acme | Mycobacterium phage Acme unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51793 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG770216 | Mycobacterium phage Rem711 | Mycobacterium phage Rem711 unclassified Trigintaduovirus Trigintaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50832 | 66.214 | Trigintaduovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG770215 | Gordonia phage Troje | Gordonia phage Troje Gordonia virus Troje Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45909 | 60.378 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| MG770213 | Mycobacterium phage OldBen | Mycobacterium phage OldBen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57159 | 61.485 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG770212 | Mycobacterium phage Haimas | Mycobacterium phage Haimas unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68296 | 66.513 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG593803 | Streptomyces phage Rowa | Streptomyces phage Rowa Streptomyces virus Rowa Rowavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42890 | 61.203 | Rowavirus | Unclassified | Siphoviridae | Streptomyces |
| MG593802 | Streptomyces phage Omar | Streptomyces phage Omar unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49299 | 67.293 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MG593801 | Streptomyces phage Attoomi | Streptomyces phage Attoomi Streptomyces virus Attoomi Attoomivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41872 | 69.689 | Attoomivirus | Unclassified | Siphoviridae | Streptomyces |
| MG593800 | Streptomyces phage AbbeyMikolon | Streptomyces phage AbbeyMikolon Streptomyces virus AbbeyMikolon Abbeymikolonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42551 | 66.826 | Abbeymikolonvirus | Unclassified | Siphoviridae | Streptomyces |
| MG670586 | Microbacterium phage Dismas | Microbacterium phage Dismas Microbacterium virus Dismas Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41593 | 69.632 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MF919537 | Gordonia phage Toniann | Gordonia phage Toniann unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92546 | 61.896 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MG099934 | Mycobacterium phage AgentM | Mycobacterium phage AgentM unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50503 | 60.884 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099937 | Mycobacterium phage Aragog | Mycobacterium phage Aragog unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50812 | 60.842 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099938 | Mycobacterium phage Archetta | Mycobacterium phage Archetta unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47409 | 60.609 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099939 | Gordonia phage BENtherdunthat | Gordonia phage BENtherdunthat Gordonia virus BENtherdunthat Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54867 | 63.364 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MG099941 | Gordonia phage Bjanes7 | Gordonia phage Bjanes7 unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46042 | 66.578 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MG099943 | Mycobacterium phage Familton | Mycobacterium phage Familton unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71807 | 65.438 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099944 | Mycobacterium phage ForGetIt | Mycobacterium phage ForGetIt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51050 | 60.578 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099945 | Mycobacterium phage Koko | Mycobacterium phage Koko unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52879 | 61.342 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099946 | Mycobacterium phage LouisV14 | Mycobacterium phage LouisV14 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44145 | 66.578 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099947 | Mycobacterium phage Ph8s | Mycobacterium phage Ph8s unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52874 | 62.566 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099949 | Mycobacterium phage Phlorence | Mycobacterium phage Phlorence unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50403 | 60.852 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099950 | Arthrobacter phage Waltz | Arthrobacter phage Waltz unclassified Laroyevirus Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58499 | 64.343 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| MG099951 | Mycobacterium phage Wilkins | Mycobacterium phage Wilkins unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50907 | 63.932 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099952 | Mycobacterium phage WunderPhul | Mycobacterium phage WunderPhul unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48724 | 61.536 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG099953 | Mycobacterium phage Youngblood | Mycobacterium phage Youngblood unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75896 | 62.949 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG298964 | Streptomyces phage Alsaber | Streptomyces phage Alsaber unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48803 | 65.912 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MG210949 | Arthrobacter phage Huntingdon | Arthrobacter phage Huntingdon unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43891 | 60.65 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MG198785 | Mycobacterium phage Thespis | Mycobacterium phage Thespis unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47618 | 66.996 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MG198782 | Arthrobacter phage MeganNoll | Arthrobacter phage MeganNoll unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44258 | 60.943 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MG198781 | Arthrobacter phage Fluke | Arthrobacter phage Fluke unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43812 | 60.917 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MG198779 | Arthrobacter phage Urla | Arthrobacter phage Urla unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43940 | 61.181 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MG198776 | Corynebacterium phage C3PO | Corynebacterium phage C3PO Corynebacterium virus C3PO Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67383 | 52.305 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MF919534 | Mycobacterium phage Superphikiman | Mycobacterium phage Superphikiman unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109799 | 60.981 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919524 | Mycobacterium phage MISSy | Mycobacterium phage MISSy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75808 | 63.074 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668286 | Rhodococcus phage Trina | Rhodococcus phage Trina Rhodococcus virus Trina Trinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139262 | 44.724 | Trinavirus | Unclassified | Siphoviridae | Rhodococcus |
| MF919541 | Mycobacterium phage YassJohnny | Mycobacterium phage YassJohnny unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73697 | 62.886 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919540 | Mycobacterium phage Willez | Mycobacterium phage Willez unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74576 | 62.94 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919539 | Mycobacterium phage Virapocalypse | Mycobacterium phage Virapocalypse unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68682 | 66.479 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919538 | Mycobacterium phage Updawg | Mycobacterium phage Updawg unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53043 | 63.501 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919536 | Mycobacterium phage TipsytheTRex | Mycobacterium phage TipsytheTRex unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52350 | 62.52 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919535 | Mycobacterium phage Terminus | Mycobacterium phage Terminus unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76169 | 63.058 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919531 | Mycobacterium phage SnapTap | Mycobacterium phage SnapTap unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53051 | 63.45 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919530 | Mycobacterium phage Sheila | Mycobacterium phage Sheila unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67880 | 66.537 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919529 | Mycobacterium phage Sassay | Mycobacterium phage Sassay unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73495 | 63.034 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919528 | Mycobacterium phage Phunky | Mycobacterium phage Phunky unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68375 | 66.501 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919527 | Mycobacterium phage Phox | Mycobacterium phage Phox unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154874 | 64.726 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919526 | Mycobacterium phage Phasih | Mycobacterium phage Phasih unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56125 | 61.055 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919525 | Mycobacterium phage Murica | Mycobacterium phage Murica unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77053 | 63.002 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919523 | Mycobacterium phage Mikota | Mycobacterium phage Mikota unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67768 | 66.586 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919520 | Gordonia phage Lozinak | Gordonia phage Lozinak unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93201 | 61.917 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MF919519 | Mycobacterium phage Longacauda | Mycobacterium phage Longacauda unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68804 | 66.405 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919517 | Mycobacterium phage LilHazelnut | Mycobacterium phage LilHazelnut unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.452 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919514 | Gordonia phage Lennon | Gordonia phage Lennon Gordonia virus Lennon Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60237 | 67.364 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MF919513 | Mycobacterium phage Koguma | Mycobacterium phage Koguma unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155759 | 64.706 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919511 | Mycobacterium phage Kailash | Mycobacterium phage Kailash unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68953 | 66.303 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919508 | Mycobacterium phage ILeeKay | Mycobacterium phage ILeeKay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51017 | 64 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919507 | Mycobacterium phage Horchata | Mycobacterium phage Horchata unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68171 | 66.411 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919506 | Mycobacterium phage FireRed | Mycobacterium phage FireRed unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76217 | 62.982 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919504 | Mycobacterium phage DmpstrDiver | Mycobacterium phage DmpstrDiver unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112285 | 60.633 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919503 | Mycobacterium phage Dingo | Mycobacterium phage Dingo unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67776 | 66.569 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF919501 | Mycobacterium phage DaWorst | Mycobacterium phage DaWorst unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56827 | 60.982 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919500 | Mycobacterium phage Dalmatian | Mycobacterium phage Dalmatian unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52850 | 63.502 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919499 | Mycobacterium phage Daffodil | Mycobacterium phage Daffodil unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155034 | 64.742 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919497 | Mycobacterium phage Chance64 | Mycobacterium phage Chance64 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41903 | 66.635 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919495 | Mycobacterium phage Bigswole | Mycobacterium phage Bigswole unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156514 | 64.775 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919494 | Mycobacterium phage BeanWater | Mycobacterium phage BeanWater unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154061 | 64.723 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919493 | Mycobacterium phage Audrick | Mycobacterium phage Audrick unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155205 | 64.661 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MF919492 | Mycobacterium phage Arib1 | Mycobacterium phage Arib1 unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46732 | 67.474 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF919490 | Mycobacterium phage Anselm | Mycobacterium phage Anselm unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53545 | 63.461 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG009575 | Mycobacterium phage Kumao | Mycobacterium phage Kumao Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70373 | 62.072 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MF975638 | Streptomyces phage Abt2graduatex2 | Streptomyces phage Abt2graduatex2 unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57385 | 69.185 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| MF975637 | Streptomyces phage Scap1 | Streptomyces phage Scap1 Streptomyces virus Scap1 Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43060 | 60.876 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MF766048 | Streptomyces phage Tefunt | Streptomyces phage Tefunt unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50574 | 66.839 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF766047 | Streptomyces phage SqueakyClean | Streptomyces phage SqueakyClean unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50837 | 67.02 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF766046 | Streptomyces phage Diane | Streptomyces phage Diane unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50483 | 66.694 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF766045 | Streptomyces phage Daudau | Streptomyces phage Daudau unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50602 | 67.12 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF766044 | Streptomyces phage Amethyst | Streptomyces phage Amethyst unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49372 | 67.224 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541410 | Streptomyces phage Ozzie | Streptomyces phage Ozzie unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49961 | 66.216 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541409 | Streptomyces phage Oliynyk | Streptomyces phage Oliynyk unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49976 | 65.938 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541408 | Streptomyces phage Jash | Streptomyces phage Jash unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50066 | 65.923 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541407 | Streptomyces phage Esperer | Streptomyces phage Esperer unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49908 | 66.21 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541406 | Streptomyces phage Dattran | Streptomyces phage Dattran unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50976 | 65.792 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541405 | Streptomyces phage Celeste | Streptomyces phage Celeste unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50536 | 65.775 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541404 | Streptomyces phage BryanRecycles | Streptomyces phage BryanRecycles unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50066 | 65.929 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF541403 | Streptomyces phage BeardedLady | Streptomyces phage BeardedLady unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49941 | 66.24 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MF668280 | Mycobacterium phage Phabba | Mycobacterium phage Phabba Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164254 | 65.209 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| MF472895 | Mycobacterium phage Kimona | Mycobacterium phage Kimona unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50283 | 64.378 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF472896 | Mycobacterium phage JoshKayV | Mycobacterium phage JoshKayV unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53366 | 63.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF472894 | Mycobacterium phage Majeke | Mycobacterium phage Majeke unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47612 | 67.367 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF472893 | Mycobacterium phage MissWhite | Mycobacterium phage MissWhite unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50263 | 63.299 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668288 | Mycobacterium phage Wiks | Mycobacterium phage Wiks unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52122 | 61.521 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668287 | Mycobacterium phage Wachhund | Mycobacterium phage Wachhund unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54513 | 61.681 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668285 | Arthrobacter phage Temper16 | Arthrobacter phage Temper16 unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43950 | 61.859 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF668283 | Mycobacterium phage Smairt | Mycobacterium phage Smairt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54655 | 63.555 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668282 | Gordonia phage ShayRa | Gordonia phage ShayRa unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51238 | 62.052 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MF668281 | Mycobacterium phage RitaG | Mycobacterium phage RitaG unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57789 | 61.603 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668277 | Mycobacterium phage MadamMonkfish | Mycobacterium phage MadamMonkfish unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75554 | 63.022 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668276 | Mycobacterium phage Lulumae | Mycobacterium phage Lulumae unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68056 | 66.583 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF668272 | Mycobacterium phage Gideon | Mycobacterium phage Gideon unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41903 | 66.592 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668271 | Mycobacterium phage Geralt | Mycobacterium phage Geralt unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55765 | 61.528 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668270 | Mycobacterium phage Emma | Mycobacterium phage Emma unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56418 | 61.321 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668269 | Mycobacterium phage Drake55 | Mycobacterium phage Drake55 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52719 | 63.433 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668268 | Mycobacterium phage Aroostook | Mycobacterium phage Aroostook unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41903 | 66.63 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185731 | Arthrobacter phage Molivia | Arthrobacter phage Molivia Arthrobacteria virus Molivia Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58247 | 53.462 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF324910 | Mycobacterium phage GuuelaD | Mycobacterium phage GuuelaD unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76315 | 58.889 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324899 | Mycobacterium phage Lokk | Mycobacterium phage Lokk unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51008 | 63.431 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140422 | Mycobacterium phage Nicholasp3 | Mycobacterium phage Nicholasp3 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75822 | 58.876 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140403 | Mycobacterium phage Changeling | Mycobacterium phage Changeling unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52991 | 64.588 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140399 | Mycobacterium phage Bartholomew | Mycobacterium phage Bartholomew unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46484 | 67.189 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF185727 | Mycobacterium phage BobSwaget | Mycobacterium phage BobSwaget unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50400 | 63.286 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185726 | Mycobacterium phage Appletree2 | Mycobacterium phage Appletree2 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73808 | 58.923 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF281061 | Mycobacterium phage Ksquared | Mycobacterium phage Ksquared unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48699 | 67.094 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF373843 | Mycobacterium phage StevieRay | Mycobacterium phage StevieRay unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48815 | 66.875 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF467949 | Streptomyces phage Spectropatronm | Streptomyces phage Spectropatronm unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55707 | 59.537 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MF467948 | Streptomyces phage DrGrey | Streptomyces phage DrGrey Strepomyces virus Drgrey Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56076 | 59.48 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MF347639 | Streptomyces phage Samisti12 | Streptomyces phage Samisti12 Streptomyces virus Samisti12 Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133710 | 49.924 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF324907 | Arthrobacter phage Beans | Arthrobacter phage Beans Arthrobacter virus Beans Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49797 | 63.566 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF140434 | Arthrobacter phage Wheelbite | Arthrobacter phage Wheelbite Arthrobacter virus Wheelbite Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59879 | 64.766 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140432 | Arthrobacter phage Teacup | Arthrobacter phage Teacup unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58238 | 49.821 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140427 | Arthrobacter phage Shrooms | Arthrobacter phage Shrooms unclassified Laroyevirus Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59707 | 64.498 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140425 | Arthrobacter phage PitaDog | Arthrobacter phage PitaDog unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42963 | 60.699 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140424 | Arthrobacter phage Nubia | Arthrobacter phage Nubia unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44045 | 60.695 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140418 | Arthrobacter phage LiSara | Arthrobacter phage LiSara Arthrobacter virus LiSara Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60137 | 64.745 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140407 | Arthrobacter phage Dino | Arthrobacter phage Dino unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43562 | 61.062 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140404 | Arthrobacter phage Christian | Arthrobacter phage Christian unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43081 | 60.707 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140400 | Arthrobacter phage Canowicakte | Arthrobacter phage Canowicakte unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43914 | 61.22 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140397 | Arthrobacter phage Abidatro | Arthrobacter phage Abidatro Arthrobacter virus Abidatro Galaxyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39122 | 68.545 | Galaxyvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189178 | Arthrobacter phage Shade | Arthrobacter phage Shade Arthrobacter virus Shade Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50930 | 61.082 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF189177 | Arthrobacter phage Piccoletto | Arthrobacter phage Piccoletto Arthrobacter virus Piccoletto Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49808 | 63.556 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF189176 | Arthrobacter phage Jordan | Arthrobacter phage Jordan unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51347 | 61.057 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF377442 | Arthrobacter phage Franzy | Arthrobacter phage Franzy unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49976 | 63.346 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF377441 | Arthrobacter phage Timinator | Arthrobacter phage Timinator unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51138 | 60.916 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| MF347638 | Streptomyces phage Peebs | Streptomyces phage Peebs Streptomyces virus Peebs Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133047 | 50.057 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF347637 | Streptomyces phage Paradiddles | Streptomyces phage Paradiddles Streptomyces virus Paradiddles Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133486 | 50.123 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF347636 | Streptomyces phage NootNoot | Streptomyces phage NootNoot Streptomyces virus NootNoot Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131086 | 50.246 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF141540 | Mycobacterium phage Avocado | Mycobacterium phage Avocado Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45389 | 68.739 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MF358542 | Streptomyces phage Sushi23 | Streptomyces phage Sushi23 unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133917 | 49.979 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF319184 | Mycobacterium phage GenevaB15 | Mycobacterium phage GenevaB15 unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80123 | 60.785 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MF190168 | Mycobacterium phage Spoonbill | Mycobacterium phage Spoonbill unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55735 | 61.303 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF155947 | Mycobacterium phage LemonSlice | Mycobacterium phage LemonSlice unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68324 | 66.466 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MF155946 | Streptomyces phage Mildred21 | Streptomyces phage Mildred21 Streptomyces virus Mildred21 Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131976 | 49.469 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF358541 | Streptomyces phage Warpy | Streptomyces phage Warpy unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132996 | 49.563 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MF185730 | Mycobacterium phage Miley16 | Mycobacterium phage Miley16 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76653 | 58.932 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140423 | Arthrobacter phage Nightmare | Arthrobacter phage Nightmare unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58839 | 49.855 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| KY965066 | Mycobacterium phage BlackStallion | Mycobacterium phage BlackStallion unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68714 | 66.499 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY965065 | Mycobacterium phage Heffalump | Mycobacterium phage Heffalump unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53085 | 63.577 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY965064 | Mycobacterium phage Pippy | Mycobacterium phage Pippy unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56663 | 61.504 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF133446 | Mycobacterium phage Bogie | Mycobacterium phage Bogie unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48639 | 66.887 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| MF038791 | Arthrobacter phage ElephantMan | Arthrobacter phage ElephantMan unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58405 | 49.912 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF038790 | Arthrobacter phage Niktson | Arthrobacter phage Niktson unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58405 | 49.912 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| KY798216 | Mycobacterium phage Journey13 | Mycobacterium phage Journey13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48502 | 62.67 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY702574 | Mycobacterium phage Kingsley | Mycobacterium phage Kingsley unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60300 | 61.38 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KU160653 | Arthrobacter phage Korra | Arthrobacter phage Korra Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43707 | 61.075 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160672 | Arthrobacter phage Wayne | Arthrobacter phage Wayne Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44371 | 61.139 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160640 | Arthrobacter phage Bennie | Arthrobacter phage Bennie Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43075 | 61.416 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KY676784 | Streptomyces phage ToastyFinz | Streptomyces phage ToastyFinz unclassified Raleighvirus Raleighvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39693 | 72.512 | Raleighvirus | Unclassified | Siphoviridae | Streptomyces |
| KY676783 | Mycobacterium phage Chorkpop | Mycobacterium phage Chorkpop Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68506 | 66.447 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY471269 | Gordonia phage DinoDaryn | Gordonia phage DinoDaryn unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44936 | 66.183 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| KY471268 | Gordonia phage Huffy | Gordonia phage Huffy unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44936 | 66.181 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| KY471267 | Mycobacterium phage SassyB | Mycobacterium phage SassyB unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55094 | 61.646 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY471460 | Mycobacterium phage Jabith | Mycobacterium phage Jabith unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52715 | 63.731 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY385383 | Mycobacterium phage Maskar | Mycobacterium phage Maskar Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69082 | 66.343 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY385382 | Mycobacterium phage ImtiyazSitla | Mycobacterium phage ImtiyazSitla Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69082 | 66.347 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY385380 | Mycobacterium phage Ashraf | Mycobacterium phage Ashraf Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69082 | 66.341 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY385384 | Mycobacterium phage SimranZ1 | Mycobacterium phage SimranZ1 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56335 | 61.26 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY385381 | Mycobacterium phage EniyanLRS | Mycobacterium phage EniyanLRS unclassified Wildcatvirus Wildcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78536 | 56.889 | Wildcatvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY549152 | Mycobacterium phage Maxxinista | Mycobacterium phage Maxxinista unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75215 | 62.989 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KY365739 | Streptomyces phage BabyGotBac | Streptomyces phage BabyGotBac unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57165 | 69.214 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KY319168 | Mycobacterium phage CrystalP | Mycobacterium phage CrystalP unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76483 | 62.981 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KY224001 | Mycobacterium phage Blue | Mycobacterium phage Blue unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50864 | 63.935 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY224000 | Mycobacterium phage Rich | Mycobacterium phage Rich unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69819 | 67.643 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY223999 | Mycobacterium phage MrMagoo | Mycobacterium phage MrMagoo unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84303 | 60.796 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| KX712238 | Mycobacterium phage Zephyr | Mycobacterium phage Zephyr unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52320 | 63.939 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY213952 | Mycobacterium phage Petruchio | Mycobacterium phage Petruchio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49531 | 63.821 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY130461 | Mycobacterium phage Taptic | Mycobacterium phage Taptic Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60973 | 67.436 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KY092484 | Streptomyces phage Raleigh | Streptomyces phage Raleigh Streptomyces virus Raleigh Raleighvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40785 | 71.801 | Raleighvirus | Unclassified | Siphoviridae | Streptomyces |
| KY092483 | Streptomyces phage Bioscum | Streptomyces phage Bioscum unclassified Austintatiousvirus Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37830 | 72.342 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| KY092482 | Streptomyces phage Mojorita | Streptomyces phage Mojorita unclassified Picardvirus Picardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38496 | 72.537 | Picardvirus | Unclassified | Siphoviridae | Streptomyces |
| KY092481 | Streptomyces phage PapayaSalad | Streptomyces phage PapayaSalad Streptomyces virus PapayaSalad Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38411 | 72.635 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| KY092480 | Streptomyces phage Picard | Streptomyces phage Picard Streptomyces virus Picard Picardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39522 | 72.605 | Picardvirus | Unclassified | Siphoviridae | Streptomyces |
| KY092479 | Streptomyces phage Ididsumtinwong | Streptomyces phage Ididsumtinwong Streptomyces virus Ididsumtinwong Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37817 | 72.367 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| KX831080 | Mycobacterium phage Lukilu | Mycobacterium phage Lukilu Mycobacterium virus Lukilu Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157034 | 64.656 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KY012363 | Mycobacterium phage Empress | Mycobacterium phage Empress unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57155 | 61.519 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX897981 | Mycobacterium phage StarStuff | Mycobacterium phage StarStuff unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52785 | 63.503 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX817173 | Mycobacterium phage Tuco | Mycobacterium phage Tuco unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76944 | 63.019 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KX808131 | Mycobacterium phage SuperGrey | Mycobacterium phage SuperGrey unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59346 | 61.723 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX758539 | Mycobacterium phage Jaan | Mycobacterium phage Jaan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52462 | 63.152 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX758538 | Mycobacterium phage Kerberos | Mycobacterium phage Kerberos unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52753 | 63.471 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX688103 | Arthrobacter phage Greenhouse | Arthrobacter phage Greenhouse unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43977 | 60.827 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX688102 | Arthrobacter phage Oxynfrius | Arthrobacter phage Oxynfrius unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44163 | 60.829 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX670789 | Streptomyces phage OlympicHelado | Streptomyces phage OlympicHelado unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56189 | 59.535 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| KX683293 | Mycobacterium phage Daffy | Mycobacterium phage Daffy Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68492 | 66.468 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX683292 | Mycobacterium phage Held | Mycobacterium phage Held Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68314 | 66.455 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX683876 | Mycobacterium phage CactusRose | Mycobacterium phage CactusRose unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52659 | 63.812 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX683875 | Mycobacterium phage Baehexic | Mycobacterium phage Baehexic unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53111 | 63.467 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX774321 | Rhodococcus phage Weasels2 | Rhodococcus phage Weasels2 Rhodococcus virus Weasel Weaselvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134973 | 41.293 | Weaselvirus | Unclassified | Siphoviridae | Rhodococcus |
| KX781992 | Mycobacterium phage Gabriel | Mycobacterium phage Gabriel unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154474 | 64.786 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KX721256 | Mycobacterium phage Erdmann | Mycobacterium phage Erdmann unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155565 | 64.747 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KX721255 | Mycobacterium phage Yucca | Mycobacterium phage Yucca unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155582 | 64.706 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KX657793 | Mycobacterium phage DarthPhader | Mycobacterium phage DarthPhader unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53432 | 63.159 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX664447 | Mycobacterium phage Phaeder | Mycobacterium phage Phaeder unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53300 | 62.619 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX588251 | Mycobacterium phage Jane | Mycobacterium phage Jane unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KX550443 | Mycobacterium phage Magnito | Mycobacterium phage Magnito unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51743 | 63.819 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX620786 | Mycobacterium phage Lego3393 | Mycobacterium phage Lego3393 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69037 | 66.508 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX589269 | Mycobacterium phage Fortunato | Mycobacterium phage Fortunato unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70679 | 68.958 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX685355 | Mycobacterium phage Qobbit | Mycobacterium phage Qobbit unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52911 | 62.573 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX610764 | Mycobacterium phage Kersh | Mycobacterium phage Kersh unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60190 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX648391 | Mycobacterium phage Tortellini | Mycobacterium phage Tortellini Mycobacterium virus Tortellini Tortellinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49658 | 65.818 | Tortellinivirus | Unclassified | Siphoviridae | Mycobacterium |
| KX664455 | Mycobacterium phage Zombie | Mycobacterium phage Zombie unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.566 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KX683423 | Mycobacterium phage BabyRay | Mycobacterium phage BabyRay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50657 | 63.555 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641266 | Mycobacterium phage Terror | Mycobacterium phage Terror unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41077 | 66.431 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641265 | Mycobacterium phage Taheera | Mycobacterium phage Taheera unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41077 | 66.434 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641263 | Mycobacterium phage Kalpine | Mycobacterium phage Kalpine unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53330 | 64.367 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641262 | Mycobacterium phage Nazo | Mycobacterium phage Nazo unclassified Bignuzvirus Bignuzvirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48870 | 66.775 | Bignuzvirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KX641261 | Mycobacterium phage Isiphiwo | Mycobacterium phage Isiphiwo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51910 | 61.562 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX620754 | Propionibacterium phage G4 | Propionibacterium phage G4 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38555 | 65.706 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX620753 | Propionibacterium phage E6 | Propionibacterium phage E6 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38067 | 65.201 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX620751 | Propionibacterium phage Doucette | Propionibacterium phage Doucette Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37429 | 65.583 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX620750 | Propionibacterium phage B22 | Propionibacterium phage B22 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37219 | 65.45 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX621005 | Arthrobacter phage Vallejo | Arthrobacter phage Vallejo unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43607 | 60.963 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX621004 | Arthrobacter phage Suppi | Arthrobacter phage Suppi unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43914 | 61.22 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX611831 | Mycobacterium phage Pharsalus | Mycobacterium phage Pharsalus unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75779 | 63.012 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KX670813 | Mycobacterium phage MitKao | Mycobacterium phage MitKao Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68493 | 66.49 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX592589 | Mycobacterium phage Iridoclysis | Mycobacterium phage Iridoclysis Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68597 | 66.41 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX580962 | Mycobacterium phage Wilder | Mycobacterium phage Wilder unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75806 | 58.896 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX580961 | Mycobacterium phage Zakai | Mycobacterium phage Zakai unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76363 | 58.909 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX702320 | Mycobacterium phage Bigfoot | Mycobacterium phage Bigfoot unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51646 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX702319 | Mycobacterium phage Pinkman | Mycobacterium phage Pinkman Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68938 | 66.374 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX585252 | Mycobacterium phage HortumSL17 | Mycobacterium phage HortumSL17 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53426 | 62.649 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX576647 | Mycobacterium phage CharlieGBrown | Mycobacterium phage CharlieGBrown Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68604 | 66.394 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX576646 | Mycobacterium phage TyrionL | Mycobacterium phage TyrionL Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68637 | 66.375 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX576645 | Mycobacterium phage Derpp | Mycobacterium phage Derpp Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68637 | 66.372 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX576644 | Mycobacterium phage WillSterrel | Mycobacterium phage WillSterrel unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58608 | 61.637 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX576643 | Mycobacterium phage FriarPreacher | Mycobacterium phage FriarPreacher Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68415 | 66.427 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX576641 | Arthrobacter phage Lucy | Arthrobacter phage Lucy unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42944 | 60.705 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX576640 | Arthrobacter phage RcigaStruga | Arthrobacter phage RcigaStruga unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43891 | 60.65 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KX576638 | Mycobacterium phage Gruunaga | Mycobacterium phage Gruunaga unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52195 | 61.383 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX574455 | Mycobacterium phage Pomar16 | Mycobacterium phage Pomar16 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52833 | 63.428 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX574454 | Mycobacterium phage PacerPaul | Mycobacterium phage PacerPaul unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52732 | 63.576 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX578071 | Mycobacterium phage Mana | Mycobacterium phage Mana Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68479 | 66.436 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX507345 | Streptomyces phage Brataylor | Streptomyces phage Brataylor unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51067 | 65.694 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KX507344 | Streptomyces phage Godpower | Streptomyces phage Godpower unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50701 | 65.715 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KX507343 | Streptomyces phage Lorelei | Streptomyces phage Lorelei unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50558 | 65.772 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KX557288 | Rhodococcus phage ChewyVIII | Rhodococcus phage ChewyVIII Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69165 | 61.804 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| KX752698 | Mycobacterium phage Tonenili | Mycobacterium phage Tonenili unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160985 | 64.09 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KX557286 | Gordonia phage Twister6 | Gordonia phage Twister6 Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57804 | 67.722 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| KX557285 | Gordonia phage Terapin | Gordonia phage Terapin Gordonia virus Terapin Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66611 | 59.552 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| KX557284 | Gordonia phage Strosahl | Gordonia phage Strosahl Gordonia virus Strosahl Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52738 | 61.963 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KX557283 | Gordonia phage Remus | Gordonia phage Remus unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52738 | 61.967 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KX557281 | Gordonia phage Jumbo | Gordonia phage Jumbo Gordonia virus Jumbo Gorjumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78302 | 54.452 | Gorjumvirus | Unclassified | Siphoviridae | Gordonia |
| KX557280 | Gordonia phage JSwag | Gordonia phage JSwag unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52726 | 61.918 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KX557278 | Gordonia phage Ghobes | Gordonia phage Ghobes Gordonia virus Ghobes Ghobesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45285 | 65.247 | Ghobesvirus | Unclassified | Siphoviridae | Gordonia |
| KX557277 | Gordonia phage Eyre | Gordonia phage Eyre Gordonia virus Eyre Eyrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44929 | 67.455 | Eyrevirus | Unclassified | Siphoviridae | Gordonia |
| KX557276 | Gordonia phage Cucurbita | Gordonia phage Cucurbita unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93686 | 61.985 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| KX557272 | Gordonia phage Bantam | Gordonia phage Bantam Gordonia virus Bantam Bantamvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92580 | 64.705 | Bantamvirus | Unclassified | Siphoviridae | Gordonia |
| KX443696 | Mycobacterium phage Laurie | Mycobacterium phage Laurie Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66507 | 68.955 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX534004 | Mycobacterium phage Sneeze | Mycobacterium phage Sneeze unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42429 | 66.455 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KX522649 | Mycobacterium phage Bircsak | Mycobacterium phage Bircsak unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53622 | 63.56 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX522943 | Mycobacterium phage Gompeii16 | Mycobacterium phage Gompeii16 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53623 | 63.559 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX507361 | Mycobacterium phage Gardann | Mycobacterium phage Gardann unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76012 | 58.903 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX375815 | Mycobacterium phage ToneTone | Mycobacterium phage ToneTone unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52044 | 61.481 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX369586 | Mycobacterium phage Papez | Mycobacterium phage Papez unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52501 | 63.787 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX369585 | Mycobacterium phage PhatCats2014 | Mycobacterium phage PhatCats2014 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69000 | 66.513 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX369584 | Mycobacterium phage Makemake | Mycobacterium phage Makemake unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52496 | 63.761 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX369583 | Mycobacterium phage Littleton | Mycobacterium phage Littleton Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155800 | 64.724 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KX344445 | Streptomyces phage Nanodon | Streptomyces phage Nanodon unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50082 | 65.698 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KX443326 | Mycobacterium phage BruceB | Mycobacterium phage BruceB unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.564 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KU963253 | Gordonia phage Phinally | Gordonia phage Phinally Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59265 | 68.437 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KX159477 | Mycobacterium phage Gattaca | Mycobacterium phage Gattaca Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65237 | 63.349 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU997639 | Mycobacterium phage Loadrie | Mycobacterium phage Loadrie Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76492 | 58.954 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU998256 | Gordonia phage Jeanie | Gordonia phage Jeanie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17118 | 68.618 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KU998255 | Gordonia phage McGonagall | Gordonia phage McGonagall Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17119 | 68.614 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KU998251 | Gordonia phage KatherineG | Gordonia phage KatherineG unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52689 | 61.926 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KU998250 | Gordonia phage Rosalind | Gordonia phage Rosalind unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52684 | 61.93 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KU998249 | Gordonia phage Soups | Gordonia phage Soups Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52924 | 61.864 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| KU998247 | Gordonia phage Bachita | Gordonia phage Bachita Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93843 | 61.948 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| KU998246 | Gordonia phage ClubL | Gordonia phage ClubL Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92618 | 61.92 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| KU998245 | Gordonia phage OneUp | Gordonia phage OneUp Oneupvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93577 | 61.483 | Oneupvirus | Unclassified | Siphoviridae | Gordonia |
| KU998244 | Gordonia phage Smoothie | Gordonia phage Smoothie Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93139 | 61.932 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| KU998239 | Gordonia phage Cozz | Gordonia phage Cozz Gordonia virus Cozz Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46600 | 60.043 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| KU998238 | Gordonia phage Splinter | Gordonia phage Splinter Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45858 | 66.13 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| KU998237 | Gordonia phage Vendetta | Gordonia phage Vendetta Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45858 | 66.135 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| KU998235 | Gordonia phage Bowser | Gordonia phage Bowser Gordonia virus Bowser Bowservirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46570 | 67.112 | Bowservirus | Unclassified | Siphoviridae | Gordonia |
| KU998234 | Gordonia phage Wizard | Gordonia phage Wizard Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58308 | 67.943 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| KU985096 | Mycobacterium phage Kazan | Mycobacterium phage Kazan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52160 | 61.555 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU985095 | Mycobacterium phage ChipMunk | Mycobacterium phage ChipMunk unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53932 | 63.059 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU985093 | Mycobacterium phage EvilGenius | Mycobacterium phage EvilGenius unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53935 | 63.007 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU985090 | Mycobacterium phage Shipwreck | Mycobacterium phage Shipwreck unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48670 | 66.926 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KU935729 | Mycobacterium phage SkinnyPete | Mycobacterium phage SkinnyPete Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43478 | 66.353 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KU963251 | Gordonia phage UmaThurman | Gordonia phage UmaThurman Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50127 | 67.026 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KU963263 | Gordonia phage BaxterFox | Gordonia phage BaxterFox Baxtervirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53717 | 66.543 | Baxtervirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KU963261 | Gordonia phage BetterKatz | Gordonia phage BetterKatz Gordonia virus BetterKatz Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50636 | 67.114 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| KU963260 | Gordonia phage Emalyn | Gordonia phage Emalyn Gordonia virus Emalyn Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43982 | 61.214 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| KU963256 | Gordonia phage Lucky10 | Gordonia phage Lucky10 Gordonia virus Lucky10 Luckytenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42979 | 65.378 | Luckytenvirus | Unclassified | Siphoviridae | Gordonia |
| KU963250 | Gordonia phage Vivi2 | Gordonia phage Vivi2 Gordonia virus Vivi2 Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59337 | 67.133 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| KU963249 | Gordonia phage Yeezy | Gordonia phage Yeezy Baxtervirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51884 | 66.695 | Baxtervirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KU963248 | Gordonia phage Yvonnetastic | Gordonia phage Yvonnetastic Gordonia virus Yvonnetastic Yvonnevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98136 | 59.749 | Yvonnevirus | Unclassified | Siphoviridae | Gordonia |
| KU963247 | Gordonia phage Attis | Gordonia phage Attis Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47881 | 66.82 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| KU963246 | Gordonia phage SoilAssassin | Gordonia phage SoilAssassin unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47880 | 66.819 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| KU865303 | Mycobacterium phage TeardropMSU | Mycobacterium phage TeardropMSU unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74896 | 63.041 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KU865302 | Mycobacterium phage Brocalys | Mycobacterium phage Brocalys unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54751 | 61.776 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KU958700 | Streptomyces phage Chymera | Streptomyces phage Chymera Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34742 | 71.369 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| KU867907 | Mycobacterium phage Potter | Mycobacterium phage Potter Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68327 | 66.513 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KU761559 | Mycobacterium phage ArcherNM | Mycobacterium phage ArcherNM unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52561 | 64.177 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU761558 | Mycobacterium phage Loser | Mycobacterium phage Loser unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53486 | 64.464 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU716094 | Mycobacterium phage Eidsmoe | Mycobacterium phage Eidsmoe Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52946 | 62.532 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU728633 | Mycobacterium phage Bipper | Mycobacterium phage Bipper Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77832 | 67.341 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KU695582 | Mycobacterium phage McFly | Mycobacterium phage McFly unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52502 | 61.541 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU695581 | Mycobacterium phage Mulciber | Mycobacterium phage Mulciber unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52428 | 63.724 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU568494 | Mycobacterium phage Bactobuster | Mycobacterium phage Bactobuster unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52129 | 63.074 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU716095 | Mycobacterium phage Myxus | Mycobacterium phage Myxus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53425 | 62.647 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU613353 | Mycobacterium phage Catalina | Mycobacterium phage Catalina unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53411 | 62.641 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU647629 | Arthrobacter phage BarretLemon | Arthrobacter phage BarretLemon Arthrobacter virus BarretLemon Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 60.903 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KU647627 | Arthrobacter phage Kitkat | Arthrobacter phage Kitkat Kelleziovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58560 | 63.391 | Kelleziovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU647626 | Arthrobacter phage KellEzio | Arthrobacter phage KellEzio Kelleziovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58871 | 63.344 | Kelleziovirus | Unclassified | Siphoviridae | Arthrobacter |
| KT626047 | Mycobacterium phage Dynamix | Mycobacterium phage Dynamix unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50628 | 63.724 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU530220 | Streptomyces phage Xkcd426 | Streptomyces phage Xkcd426 unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64477 | 68.828 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KU297783 | Mycobacterium phage NaSiaTalie | Mycobacterium phage NaSiaTalie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52920 | 63.398 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT880194 | Mycobacterium phage Glass | Mycobacterium phage Glass unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67509 | 68.966 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KU160673 | Arthrobacter phage Wilde | Arthrobacter phage Wilde unclassified Tankvirus Tankvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68203 | 62.868 | Tankvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160669 | Arthrobacter phage Tank | Arthrobacter phage Tank Tankvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67592 | 62.87 | Tankvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160668 | Arthrobacter phage TaeYoung | Arthrobacter phage TaeYoung unclassified Marthavirus Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50999 | 60.984 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160666 | Arthrobacter phage SorJuana | Arthrobacter phage SorJuana unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58979 | 53.009 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160665 | Arthrobacter phage Sonny | Arthrobacter phage Sonny Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50909 | 61.091 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160664 | Arthrobacter phage Salgado | Arthrobacter phage Salgado Arthrobacter virus Salgado Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59807 | 64.635 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160663 | Arthrobacter phage Rings | Arthrobacter phage Rings unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59167 | 53.026 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160662 | Arthrobacter phage RAP15 | Arthrobacter phage RAP15 unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44259 | 60.944 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160661 | Arthrobacter phage Pumancara | Arthrobacter phage Pumancara Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42830 | 61.749 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160659 | Arthrobacter phage Preamble | Arthrobacter phage Preamble Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43374 | 60.691 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160656 | Arthrobacter phage Martha | Arthrobacter phage Martha Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51027 | 60.954 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160654 | Arthrobacter phage Laroye | Arthrobacter phage Laroye Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60005 | 64.788 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160652 | Arthrobacter phage Joann | Arthrobacter phage Joann Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44183 | 60.725 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160651 | Arthrobacter phage Jawnski | Arthrobacter phage Jawnski Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49419 | 63.439 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160649 | Arthrobacter phage Immaculata | Arthrobacter phage Immaculata unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43661 | 60.995 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160647 | Arthrobacter phage Gorgeous | Arthrobacter phage Gorgeous unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58979 | 53.01 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160646 | Arthrobacter phage Gordon | Arthrobacter phage Gordon Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58279 | 49.841 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160645 | Arthrobacter phage Glenn | Arthrobacter phage Glenn Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44389 | 60.819 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160644 | Arthrobacter phage Galaxy | Arthrobacter phage Galaxy Arthrobacter virus Galaxy Galaxyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37809 | 68.433 | Galaxyvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160643 | Arthrobacter phage DrRobert | Arthrobacter phage DrRobert Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42601 | 60.611 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160641 | Arthrobacter phage CaptnMurica | Arthrobacter phage CaptnMurica Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58159 | 49.588 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160639 | Arthrobacter phage Anansi | Arthrobacter phage Anansi unclassified Amigovirus Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58848 | 53.009 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160638 | Arthrobacter phage Amigo | Arthrobacter phage Amigo Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59173 | 52.948 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU252585 | Gordonia phage Howe | Gordonia phage Howe Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53182 | 65.601 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KU234099 | Mycobacterium phage MkaliMitinis3 | Mycobacterium phage MkaliMitinis3 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75844 | 58.881 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU189325 | Streptomyces phage Maih | Streptomyces phage Maih unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57256 | 69.256 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KT895281 | Mycobacterium phage Cabrinians | Mycobacterium phage Cabrinians unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56669 | 61.208 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT895280 | Mycobacterium phage Sparkdehlily | Mycobacterium phage Sparkdehlily unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 61.235 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT818595 | Mycobacterium phage Pepe | Mycobacterium phage Pepe unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50515 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT438500 | Mycobacterium phage Pari | Mycobacterium phage Pari unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50614 | 63.532 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT599441 | Mycobacterium phage Squid | Mycobacterium phage Squid Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68596 | 66.456 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT365400 | Mycobacterium phage ErnieJ | Mycobacterium phage ErnieJ Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153243 | 64.733 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KT365398 | Mycobacterium phage DTDevon | Mycobacterium phage DTDevon Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156754 | 64.636 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KT388014 | Mycobacterium phage Turj99 | Mycobacterium phage Turj99 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51161 | 63.742 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT588442 | Mycobacterium phage LadyBird | Mycobacterium phage LadyBird unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53141 | 63.459 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281794 | Mycobacterium phage Snenia | Mycobacterium phage Snenia unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75626 | 59.276 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT365402 | Mycobacterium phage Tres | Mycobacterium phage Tres Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67349 | 68.939 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT365401 | Arthrobacter phage Brent | Arthrobacter phage Brent Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49879 | 63.425 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| KT365399 | Mycobacterium phage Annihilator | Mycobacterium phage Annihilator unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41893 | 66.598 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KT381276 | Mycobacterium phage Serenity | Mycobacterium phage Serenity unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52088 | 62.642 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT355474 | Mycobacterium phage Frosty24 | Mycobacterium phage Frosty24 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.571 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KT355472 | Mycobacterium phage Cedasite | Mycobacterium phage Cedasite unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KT591489 | Mycobacterium phage Archie | Mycobacterium phage Archie unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76271 | 58.722 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT454972 | Mycobacterium phage Evanesce | Mycobacterium phage Evanesce Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.445 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281796 | Mycobacterium phage Zakhe101 | Mycobacterium phage Zakhe101 unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69653 | 65.457 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281795 | Mycobacterium phage XFactor | Mycobacterium phage XFactor unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55617 | 61.715 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281793 | Mycobacterium phage Seagreen | Mycobacterium phage Seagreen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57766 | 61.832 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281792 | Mycobacterium phage PopTart | Mycobacterium phage PopTart unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55094 | 61.644 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281791 | Mycobacterium phage Lolly9 | Mycobacterium phage Lolly9 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75816 | 59.316 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281790 | Mycobacterium phage HyRo | Mycobacterium phage HyRo Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153714 | 64.686 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KT259047 | Mycobacterium phage Rufus | Mycobacterium phage Rufus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52357 | 63.854 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT375356 | Rhodococcus phage Rhodalysa | Rhodococcus phage Rhodalysa unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.548 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KT364588 | Mycobacterium phage Hetaeria | Mycobacterium phage Hetaeria Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68405 | 66.451 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT326768 | Mycobacterium phage Tasp14 | Mycobacterium phage Tasp14 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51409 | 63.923 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT285706 | Mycobacterium phage Pioneer | Mycobacterium phage Pioneer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53219 | 62.587 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT246486 | Mycobacterium phage Chadwick | Mycobacterium phage Chadwick unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49421 | 59.786 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT246485 | Mycobacterium phage OBUpride | Mycobacterium phage OBUpride Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53790 | 67.505 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT372003 | Mycobacterium phage Lumos | Mycobacterium phage Lumos unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75586 | 59.27 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT373978 | Mycobacterium phage Ukulele | Mycobacterium phage Ukulele unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75114 | 63.019 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KT363731 | Mycobacterium phage TheloniousMonk | Mycobacterium phage TheloniousMonk unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52055 | 63.648 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT186229 | Streptomyces phage Verse | Streptomyces phage Verse Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49483 | 65.602 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT186228 | Streptomyces phage Amela | Streptomyces phage Amela Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49452 | 65.613 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT184391 | Streptomyces phage Lannister | Streptomyces phage Lannister Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50165 | 65.743 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT184390 | Streptomyces phage Izzy | Streptomyces phage Izzy Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50113 | 65.911 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT124229 | Streptomyces phage Hydra | Streptomyces phage Hydra Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50727 | 66.225 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT124228 | Streptomyces phage Danzina | Streptomyces phage Danzina Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50773 | 65.667 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT124227 | Streptomyces phage Aaronocolus | Streptomyces phage Aaronocolus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49562 | 66.176 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT152029 | Streptomyces phage Caliburn | Streptomyces phage Caliburn Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49949 | 66.194 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT309034 | Mycobacterium phage Dante | Mycobacterium phage Dante unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59652 | 61.793 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT279576 | Mycobacterium phage Phatniss | Mycobacterium phage Phatniss unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57293 | 61.336 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT270441 | Mycobacterium phage BrownCNA | Mycobacterium phage BrownCNA Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71214 | 68.878 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT347313 | Mycobacterium phage Brusacoram | Mycobacterium phage Brusacoram Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47618 | 66.994 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KT321476 | Mycobacterium phage Zeenon | Mycobacterium phage Zeenon Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155292 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KR997967 | Mycobacterium phage Pops | Mycobacterium phage Pops Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68367 | 66.601 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR997934 | Mycobacterium phage Nhonho | Mycobacterium phage Nhonho unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51355 | 63.75 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KR997932 | Mycobacterium phage Godines | Mycobacterium phage Godines Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67277 | 69.025 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR997930 | Mycobacterium phage Florinda | Mycobacterium phage Florinda unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59416 | 61.748 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KR997929 | Mycobacterium phage Barriga | Mycobacterium phage Barriga unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52643 | 63.395 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT020852 | Mycobacterium phage NoSleep | Mycobacterium phage NoSleep unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74655 | 62.983 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KT222942 | Mycobacterium phage Dusk | Mycobacterium phage Dusk unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75339 | 63.018 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KT222941 | Mycobacterium phage NelitzaMV | Mycobacterium phage NelitzaMV unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72790 | 63.054 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KT222940 | Mycobacterium phage Kinbote | Mycobacterium phage Kinbote Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.445 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| KR824843 | Mycobacterium phage Ovechkin | Mycobacterium phage Ovechkin unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58338 | 62.004 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KR816508 | Mycobacterium phage Phamished | Mycobacterium phage Phamished Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68515 | 66.476 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR781349 | Mycobacterium phage Apizium | Mycobacterium phage Apizium Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68227 | 66.44 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR080205 | Mycobacterium phage Corofin | Mycobacterium phage Corofin unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68685 | 67.513 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR080196 | Mycobacterium phage Momo | Mycobacterium phage Momo Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154553 | 64.744 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KR080204 | Mycobacterium phage Mindy | Mycobacterium phage Mindy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75796 | 62.965 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KR080203 | Mycobacterium phage OrangeOswald | Mycobacterium phage OrangeOswald unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68674 | 67.536 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR080202 | Mycobacterium phage MOOREtheMARYer | Mycobacterium phage MOOREtheMARYer Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 68.581 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KR080201 | Mycobacterium phage Nerujay | Mycobacterium phage Nerujay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53455 | 63.693 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KR080200 | Mycobacterium phage AlanGrant | Mycobacterium phage AlanGrant Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72109 | 68.926 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR080199 | Mycobacterium phage Baee | Mycobacterium phage Baee Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70270 | 67.645 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR080198 | Mycobacterium phage Cambiare | Mycobacterium phage Cambiare Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45161 | 68.798 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KR080197 | Mycobacterium phage FlagStaff | Mycobacterium phage FlagStaff Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44576 | 68.521 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KR080195 | Mycobacterium phage Phayonce | Mycobacterium phage Phayonce Phayoncevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49203 | 66.673 | Phayoncevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KR080194 | Mycobacterium phage Vincenzo | Mycobacterium phage Vincenzo Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72139 | 68.909 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KP876466 | Streptomyces phage TP1604 | Streptomyces phage TP1604 Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57168 | 69.208 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KP876465 | Streptomyces phage YDN12 | Streptomyces phage YDN12 Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56528 | 69.21 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KM923970 | Mycobacterium phage Gomashi | Mycobacterium phage Gomashi unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41903 | 66.592 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027200 | Mycobacterium phage Malithi | Mycobacterium phage Malithi Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46869 | 67.093 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KP027205 | Mycobacterium phage Alvin | Mycobacterium phage Alvin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49577 | 63.628 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP273225 | Mycobacterium phage Sheen | Mycobacterium phage Sheen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52927 | 63.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027208 | Mycobacterium phage Pipsqueak | Mycobacterium phage Pipsqueak Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68328 | 66.517 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KP027195 | Mycobacterium phage Cosmo | Mycobacterium phage Cosmo unclassified Wildcatvirus Wildcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78229 | 56.843 | Wildcatvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027209 | Mycobacterium phage Sigman | Mycobacterium phage Sigman Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68311 | 66.509 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KP027204 | Mycobacterium phage Thor | Mycobacterium phage Thor unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53058 | 63.745 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027203 | Mycobacterium phage Treddle | Mycobacterium phage Treddle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53008 | 63.86 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027201 | Mycobacterium phage Sbash | Mycobacterium phage Sbash Chenonavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55832 | 65.647 | Chenonavirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027197 | Mycobacterium phage FluffyNinja | Mycobacterium phage FluffyNinja Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68716 | 66.5 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KP027196 | Mycobacterium phage Edtherson | Mycobacterium phage Edtherson unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51492 | 63.977 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM881426 | Mycobacterium phage ZygoTaiga | Mycobacterium phage ZygoTaiga Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157204 | 64.692 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KM652554 | Streptomyces phage Jay2Jay | Streptomyces phage Jay2Jay Streptomyces virus Jay2Jay Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133531 | 49.497 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| KM652553 | Mycobacterium phage CaptainTrips | Mycobacterium phage CaptainTrips unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57328 | 61.523 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM591905 | Mycobacterium phage RonRayGun | Mycobacterium phage RonRayGun unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42596 | 66.133 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ959632 | Mycobacterium phage Equemioh13 | Mycobacterium phage Equemioh13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53042 | 63.506 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM597531 | Mycobacterium phage Abrogate | Mycobacterium phage Abrogate unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52530 | 63.756 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM597530 | Mycobacterium phage Bipolar | Mycobacterium phage Bipolar unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58985 | 61.372 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM463009 | Mycobacterium phage Power | Mycobacterium phage Power unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53395 | 63.493 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM401838 | Mycobacterium phage VohminGhazi | Mycobacterium phage VohminGhazi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52155 | 61.524 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM233454 | Mycobacterium phage MarQuardt | Mycobacterium phage MarQuardt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50882 | 64.036 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM402757 | Mycobacterium phage Llama | Mycobacterium phage Llama unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58472 | 61.067 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM363597 | Mycobacterium phage Zonia | Mycobacterium phage Zonia Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69271 | 66.348 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KM347890 | Mycobacterium phage Vivaldi | Mycobacterium phage Vivaldi Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68873 | 66.396 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KM408320 | Mycobacterium phage Lasso | Mycobacterium phage Lasso Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68576 | 66.414 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KM236502 | Mycobacterium phage Eremos | Mycobacterium phage Eremos Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68472 | 66.531 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KM279937 | Mycobacterium phage Estave1 | Mycobacterium phage Estave1 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60727 | 61.324 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101123 | Mycobacterium phage Minerva | Mycobacterium phage Minerva unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109871 | 60.745 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101122 | Mycobacterium phage Hades | Mycobacterium phage Hades unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54986 | 61.396 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101118 | Mycobacterium phage Swirley | Mycobacterium phage Swirley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49717 | 61.044 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101117 | Mycobacterium phage LizLemon | Mycobacterium phage LizLemon Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67496 | 68.949 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KM066034 | Mycobacterium phage Inventum | Mycobacterium phage Inventum unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57052 | 61.376 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM215148 | Mycobacterium phage Cerasum | Mycobacterium phage Cerasum unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53636 | 61.494 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ829260 | Mycobacterium phage YungJamal | Mycobacterium phage YungJamal unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70214 | 65.34 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM083128 | Mycobacterium phage Sparky | Mycobacterium phage Sparky Mycobacterium virus Sparky Sparkyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63334 | 65.164 | Sparkyvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ567044 | Mycobacterium phage EmpTee | Mycobacterium phage EmpTee Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68428 | 66.476 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ567043 | Mycobacterium phage Gaia | Mycobacterium phage Gaia Gaiavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90460 | 56.831 | Gaiavirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ595575 | Mycobacterium phage Willis | Mycobacterium phage Willis Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155476 | 64.724 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KJ567041 | Mycobacterium phage Zapner | Mycobacterium phage Zapner unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55307 | 61.106 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ595576 | Mycobacterium phage Manad | Mycobacterium phage Manad Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68807 | 66.431 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ690250 | Mycobacterium phage Pinto | Mycobacterium phage Pinto unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50610 | 63.533 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ538723 | Mycobacterium phage KingVeVeVe | Mycobacterium phage KingVeVeVe Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68043 | 66.502 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ538722 | Mycobacterium phage HH92 | Mycobacterium phage HH92 Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.447 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ538721 | Mycobacterium phage MosMoris | Mycobacterium phage MosMoris Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65243 | 63.352 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ194579 | Mycobacterium phage Swish | Mycobacterium phage Swish Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68735 | 66.48 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ409696 | Mycobacterium phage Lamina13 | Mycobacterium phage Lamina13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53255 | 63.664 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ194585 | Mycobacterium phage Seabiscuit | Mycobacterium phage Seabiscuit unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51781 | 63.657 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ194584 | Mycobacterium phage Heathcliff | Mycobacterium phage Heathcliff unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68628 | 67.522 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ194583 | Mycobacterium phage Numberten | Mycobacterium phage Numberten Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68607 | 66.517 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ194582 | Mycobacterium phage Hawkeye | Mycobacterium phage Hawkeye Hawkeyevirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67383 | 57.05 | Hawkeyevirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| KJ194581 | Mycobacterium phage Audrey | Mycobacterium phage Audrey unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68662 | 67.502 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ194580 | Mycobacterium phage Badfish | Mycobacterium phage Badfish Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69030 | 66.522 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ174157 | Mycobacterium phage Soto | Mycobacterium phage Soto Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67744 | 66.558 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ174156 | Mycobacterium phage Alsfro | Mycobacterium phage Alsfro unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52136 | 63.57 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ025956 | Mycobacterium phage Saal | Mycobacterium phage Saal unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57775 | 61.289 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF959563 | Mycobacterium phage Obama12 | Mycobacterium phage Obama12 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51797 | 63.963 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF861510 | Mycobacterium phage EagleEye | Mycobacterium phage EagleEye unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52974 | 61.423 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF841477 | Mycobacterium phage Donovan | Mycobacterium phage Donovan Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47162 | 67.181 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KF713485 | Mycobacterium phage Suffolk | Mycobacterium phage Suffolk Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68262 | 66.561 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF493881 | Mycobacterium phage JAMaL | Mycobacterium phage JAMaL Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70841 | 68.751 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF493883 | Mycobacterium phage Mosby | Mycobacterium phage Mosby unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74533 | 63.054 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF493882 | Mycobacterium phage Jovo | Mycobacterium phage Jovo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51319 | 60.771 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF493880 | Mycobacterium phage HanShotFirst | Mycobacterium phage HanShotFirst unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52390 | 63.814 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF493879 | Mycobacterium phage Bernardo | Mycobacterium phage Bernardo Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68196 | 67.428 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF562102 | Mycobacterium phage PhatBacter | Mycobacterium phage PhatBacter unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76217 | 62.979 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF562101 | Mycobacterium phage Nala | Mycobacterium phage Nala unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75894 | 63.052 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF562100 | Mycobacterium phage HufflyPuff | Mycobacterium phage HufflyPuff unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76323 | 63.064 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF562099 | Mycobacterium phage Bruin | Mycobacterium phage Bruin unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74210 | 63.023 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF560334 | Mycobacterium phage Zaka | Mycobacterium phage Zaka unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52122 | 61.504 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF560333 | Mycobacterium phage Artemis2UCLA | Mycobacterium phage Artemis2UCLA unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52344 | 61.405 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF560332 | Mycobacterium phage CloudWang3 | Mycobacterium phage CloudWang3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52873 | 61.353 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF560331 | Mycobacterium phage Graduation | Mycobacterium phage Graduation unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52823 | 63.452 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024734 | Mycobacterium phage Shrimp | Mycobacterium phage Shrimp Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155714 | 64.717 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KF006817 | Mycobacterium phage Breeniome | Mycobacterium phage Breeniome Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154434 | 64.78 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KF279415 | Mycobacterium phage PhrostyMug | Mycobacterium phage PhrostyMug unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53636 | 63.677 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024730 | Mycobacterium phage Dylan | Mycobacterium phage Dylan unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69815 | 65.416 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024726 | Mycobacterium phage SarFire | Mycobacterium phage SarFire unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53701 | 63.813 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF416344 | Mycobacterium phage Adzzy | Mycobacterium phage Adzzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52519 | 62.556 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF416343 | Mycobacterium phage Goku | Mycobacterium phage Goku unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76483 | 62.83 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF416341 | Mycobacterium phage Phelemich | Mycobacterium phage Phelemich unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70115 | 68.252 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF416340 | Mycobacterium phage Wheeler | Mycobacterium phage Wheeler unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53588 | 63.316 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF306380 | Mycobacterium phage DrDrey | Mycobacterium phage DrDrey unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77367 | 63 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024732 | Mycobacterium phage Contagion | Mycobacterium phage Contagion unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74533 | 63.057 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024731 | Mycobacterium phage Crossroads | Mycobacterium phage Crossroads unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76129 | 58.945 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024729 | Mycobacterium phage KayaCho | Mycobacterium phage KayaCho unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70838 | 70.006 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF024728 | Mycobacterium phage Muddy | Mycobacterium phage Muddy Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48228 | 58.827 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024727 | Mycobacterium phage Reprobate | Mycobacterium phage Reprobate Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70120 | 68.269 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF024725 | Mycobacterium phage Whirlwind | Mycobacterium phage Whirlwind unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76050 | 59.344 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024724 | Mycobacterium phage Trouble | Mycobacterium phage Trouble unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52102 | 63.602 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024723 | Mycobacterium phage Hamulus | Mycobacterium phage Hamulus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57155 | 61.758 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017005 | Mycobacterium phage Daenerys | Mycobacterium phage Daenerys unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58043 | 61.553 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017004 | Mycobacterium phage Velveteen | Mycobacterium phage Velveteen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54314 | 61.465 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017003 | Mycobacterium phage Jabbawokkie | Mycobacterium phage Jabbawokkie unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55213 | 61.073 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017002 | Mycobacterium phage Catdawg | Mycobacterium phage Catdawg unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72108 | 65.394 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF006818 | Mycobacterium phage Wanda | Mycobacterium phage Wanda unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109960 | 60.75 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024721 | Mycobacterium phage AnnaL29 | Mycobacterium phage AnnaL29 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53253 | 64.389 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF114875 | Mycobacterium phage Redno2 | Mycobacterium phage Redno2 unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108297 | 60.879 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF114874 | Mycobacterium phage Bobi | Mycobacterium phage Bobi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59179 | 61.664 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024722 | Mycobacterium virus TA17a | Mycobacterium virus TA17a Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67324 | 68.984 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF188414 | Mycobacterium phage ABCat | Mycobacterium phage ABCat unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76131 | 63.045 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017927 | Mycobacterium phage Odin | Mycobacterium phage Odin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52807 | 62.3 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661280 | Mycobacterium phage Job42 | Mycobacterium phage Job42 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59626 | 61.158 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KC748971 | Mycobacterium phage Murphy | Mycobacterium phage Murphy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76179 | 62.924 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KC748970 | Mycobacterium phage ArcherS7 | Mycobacterium phage ArcherS7 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156558 | 64.692 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KC748969 | Mycobacterium phage Phaux | Mycobacterium phage Phaux unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76479 | 62.919 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KC748968 | Mycobacterium phage Gizmo | Mycobacterium phage Gizmo Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157482 | 64.642 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KC691258 | Mycobacterium phage Newman | Mycobacterium phage Newman Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68598 | 66.518 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KC691257 | Mycobacterium phage Astraea | Mycobacterium phage Astraea Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154872 | 64.683 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| KC691256 | Mycobacterium phage Fishburne | Mycobacterium phage Fishburne Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47109 | 67.267 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| KC691254 | Mycobacterium phage Breezona | Mycobacterium phage Breezona Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76652 | 58.931 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC700558 | Streptomyces phage Zemlya | Streptomyces phage Zemlya Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51077 | 65.673 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KC700557 | Streptomyces phage Sujidade | Streptomyces phage Sujidade Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51552 | 65.706 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KC700556 | Streptomyces phage Lika | Streptomyces phage Lika Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51252 | 65.796 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KC661278 | Mycobacterium phage SiSi | Mycobacterium phage SiSi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56279 | 61.492 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661277 | Mycobacterium phage Phrux | Mycobacterium phage Phrux unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74711 | 63.054 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661276 | Mycobacterium phage Winky | Mycobacterium phage Winky Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76653 | 58.932 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661274 | Mycobacterium phage SDcharge11 | Mycobacterium phage SDcharge11 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67702 | 66.521 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KC661273 | Mycobacterium phage PattyP | Mycobacterium phage PattyP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52057 | 63.59 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC576784 | Mycobacterium phage ShiVal | Mycobacterium phage ShiVal Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68355 | 66.498 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JX649099 | Mycobacterium phage Gyarad | Mycobacterium phage Gyarad Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68004 | 66.486 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JX649098 | Mycobacterium phage Nacho | Mycobacterium phage Nacho Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69321 | 66.561 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JX649096 | Mycobacterium phage Serpentine | Mycobacterium phage Serpentine Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68884 | 66.515 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JX182372 | Streptomyces virus TG1 | Streptomyces virus TG1 Tigunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40474 | 64.649 | Tigunavirus | Unclassified | Siphoviridae | Streptomyces |
| JX182371 | Streptomyces phage SV1 | Streptomyces phage SV1 unclassified Picardvirus Picardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37612 | 72.692 | Picardvirus | Unclassified | Siphoviridae | Streptomyces |
| JX182370 | Streptomyces phage R4 | Streptomyces phage R4 Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51071 | 66.962 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| JX307705 | Mycobacterium virus Marcell | Mycobacterium virus Marcell Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49186 | 63.971 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX262225 | Propionibacterium phage ATCC29399B_C | Propionibacterium phage ATCC29399B_C Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29516 | 53.998 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262224 | Propionibacterium phage ATCC29399B_T | Propionibacterium phage ATCC29399B_T Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29516 | 54.005 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262223 | Propionibacterium phage P1.1 | Propionibacterium phage P1.1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29348 | 54.365 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262222 | Propionibacterium phage P100_1 | Propionibacterium phage P100_1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29612 | 54.069 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262221 | Propionibacterium phage P100_A | Propionibacterium phage P100_A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29505 | 53.832 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262220 | Propionibacterium phage P100D | Propionibacterium phage P100D Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29506 | 53.762 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262219 | Propionibacterium phage P105 | Propionibacterium phage P105 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29202 | 54.174 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262218 | Propionibacterium phage P104A | Propionibacterium phage P104A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29371 | 53.982 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262217 | Propionibacterium phage P101A | Propionibacterium phage P101A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29574 | 54.135 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX411620 | Mycobacterium virus Dorothy | Mycobacterium virus Dorothy Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58866 | 61.407 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JX262376 | Streptomyces phage phiELB20 | Streptomyces phage phiELB20 Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51160 | 66.986 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| JX121091 | Mycobacterium phage Taj | Mycobacterium phage Taj Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58550 | 61.863 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ809703 | Mycobacterium phage Aeneas | Mycobacterium phage Aeneas unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53684 | 63.587 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ809702 | Mycobacterium phage Avani | Mycobacterium phage Avani Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54470 | 60.973 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ911768 | Mycobacterium phage Ava3 | Mycobacterium phage Ava3 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154466 | 64.762 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JQ300538 | Mycobacterium phage Spartacus | Mycobacterium phage Spartacus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61164 | 61.652 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ398052 | Mycobacterium phage Wildcat | Mycobacterium phage Wildcat Wildcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78296 | 56.852 | Wildcatvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU203571 | Mycobacterium phage Giles | Mycobacterium phage Giles Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.447 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699019 | Mycobacterium virus Jeffabunny | Mycobacterium virus Jeffabunny Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48963 | 61.573 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699018 | Mycobacterium phage Kamiyu | Mycobacterium phage Kamiyu unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68633 | 67.478 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699017 | Mycobacterium phage Kikipoo | Mycobacterium phage Kikipoo Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68839 | 66.52 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699016 | Mycobacterium virus Kugel | Mycobacterium virus Kugel Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52379 | 63.837 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699014 | Mycobacterium phage Nova | Mycobacterium phage Nova Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65108 | 59.678 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| JN699013 | Mycobacterium phage Pio | Mycobacterium phage Pio Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156758 | 64.76 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN699012 | Mycobacterium virus SG4 | Mycobacterium virus SG4 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59016 | 61.871 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699011 | Mycobacterium phage Stinger | Mycobacterium phage Stinger Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69641 | 68.557 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699010 | Mycobacterium phage TallGrassMM | Mycobacterium phage TallGrassMM unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68133 | 66.455 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699009 | Mycobacterium phage ThreeOh3D2 | Mycobacterium phage ThreeOh3D2 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68992 | 66.515 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699008 | Mycobacterium phage Vista | Mycobacterium phage Vista Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68494 | 66.48 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699007 | Mycobacterium phage Acadian | Mycobacterium phage Acadian Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69864 | 68.423 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699006 | Mycobacterium phage Akoma | Mycobacterium phage Akoma unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68711 | 67.541 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699004 | Mycobacterium phage Ares | Mycobacterium phage Ares Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67436 | 68.993 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699003 | Mycobacterium phage Athena | Mycobacterium phage Athena Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69409 | 67.526 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699002 | Mycobacterium phage Avrafan | Mycobacterium phage Avrafan unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.569 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699001 | Mycobacterium phage Babsiella | Mycobacterium phage Babsiella Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48420 | 67.078 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| JN698999 | Mycobacterium phage Blue7 | Mycobacterium phage Blue7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52288 | 61.358 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698998 | Mycobacterium phage Bruns | Mycobacterium phage Bruns Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53003 | 63.617 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698997 | Mycobacterium virus Courthouse | Mycobacterium virus Courthouse Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110569 | 60.922 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698996 | Mycobacterium virus DeadP | Mycobacterium virus DeadP Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56461 | 61.552 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698995 | Mycobacterium phage Dori | Mycobacterium phage Dori Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64613 | 65.988 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| JN698993 | Mycobacterium virus Firecracker | Mycobacterium virus Firecracker Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71341 | 65.498 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698992 | Mycobacterium phage Gadjet | Mycobacterium phage Gadjet Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67949 | 67.486 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN698991 | Mycobacterium phage Hedgerow | Mycobacterium phage Hedgerow Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67451 | 68.955 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN698990 | Mycobacterium phage IsaacEli | Mycobacterium phage IsaacEli Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68839 | 66.519 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN698989 | Mycobacterium phage JacAttac | Mycobacterium phage JacAttac Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68311 | 66.505 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN699627 | Mycobacterium phage Nappy | Mycobacterium phage Nappy Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156646 | 64.652 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN699626 | Mycobacterium phage MoMoMixon | Mycobacterium phage MoMoMixon Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154573 | 64.767 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN699625 | Mycobacterium phage Wally | Mycobacterium phage Wally Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155299 | 64.666 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN859129 | Mycobacterium virus DotProduct | Mycobacterium virus DotProduct Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55363 | 61.752 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN687951 | Mycobacterium phage Violet | Mycobacterium phage Violet Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52481 | 63.846 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN680858 | Mycobacterium virus Rumpelstiltskin | Mycobacterium virus Rumpelstiltskin Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69279 | 58.911 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN660814 | Mycobacterium phage Dreamboat | Mycobacterium phage Dreamboat unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 63.908 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN638753 | Mycobacterium phage Morgushi | Mycobacterium phage Morgushi Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68307 | 66.444 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN638752 | Mycobacterium phage Murdoc | Mycobacterium phage Murdoc Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68600 | 66.417 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN624850 | Mycobacterium phage Pleione | Mycobacterium phage Pleione Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155586 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN618996 | Mycobacterium phage Arbiter | Mycobacterium phage Arbiter unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67169 | 68.91 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN572061 | Mycobacterium phage Jebeks | Mycobacterium phage Jebeks Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45580 | 67.26 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| JN572689 | Mycobacterium virus Perseus | Mycobacterium virus Perseus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53142 | 63.712 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN412593 | Mycobacterium virus Liefie | Mycobacterium virus Liefie Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41650 | 66.783 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| JN412592 | Mycobacterium phage LinStu | Mycobacterium phage LinStu Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153882 | 64.837 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN412591 | Mycobacterium phage BigNuz | Mycobacterium phage BigNuz Bignuzvirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48984 | 66.736 | Bignuzvirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| JN412590 | Mycobacterium virus Eureka | Mycobacterium virus Eureka Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76174 | 62.852 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN412588 | Mycobacterium phage Dandelion | Mycobacterium phage Dandelion Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157568 | 64.726 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN391441 | Mycobacterium phage Elph10 | Mycobacterium phage Elph10 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74675 | 63.008 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN382248 | Mycobacterium phage Lilac | Mycobacterium phage Lilac unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76260 | 62.969 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN398369 | Mycobacterium phage RidgeCB | Mycobacterium phage RidgeCB Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50844 | 63.952 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN398368 | Mycobacterium virus GUmbie | Mycobacterium virus GUmbie Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57387 | 61.369 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN408461 | Mycobacterium phage Trixie | Mycobacterium phage Trixie Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53526 | 64.464 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN408460 | Mycobacterium phage Turbido | Mycobacterium phage Turbido Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53169 | 63.283 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937110 | Mycobacterium phage KSSJEB | Mycobacterium phage KSSJEB Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51381 | 63.631 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937109 | Mycobacterium phage Yoshand | Mycobacterium phage Yoshand Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68719 | 66.488 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF937108 | Mycobacterium phage Switzer | Mycobacterium phage Switzer Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52298 | 63.756 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937107 | Mycobacterium phage SirHarley | Mycobacterium phage SirHarley Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64791 | 59.567 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| JF937106 | Mycobacterium phage SirDuracell | Mycobacterium phage SirDuracell Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75973 | 62.922 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937105 | Mycobacterium phage Rey | Mycobacterium phage Rey Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83724 | 60.873 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| JF937103 | Mycobacterium phage Museum | Mycobacterium phage Museum Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51426 | 63.637 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937102 | Mycobacterium phage Mozy | Mycobacterium phage Mozy Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57278 | 61.055 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937101 | Mycobacterium phage LittleE | Mycobacterium phage LittleE Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109086 | 61.345 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937100 | Mycobacterium phage Lesedi | Mycobacterium phage Lesedi Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50486 | 63.76 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937099 | Mycobacterium phage JC27 | Mycobacterium phage JC27 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52169 | 63.639 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937098 | Mycobacterium phage Ibhubesi | Mycobacterium phage Ibhubesi Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55600 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937097 | Mycobacterium phage Hertubise | Mycobacterium phage Hertubise Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68675 | 66.449 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF937096 | Mycobacterium phage Henry | Mycobacterium phage Henry Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76049 | 62.962 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937095 | Mycobacterium phage Harvey | Mycobacterium phage Harvey Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68193 | 66.475 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF937094 | Mycobacterium phage Hammer | Mycobacterium phage Hammer Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51889 | 61.331 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937093 | Mycobacterium phage DLane | Mycobacterium phage DLane Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58899 | 61.884 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937092 | Mycobacterium phage DaVinci | Mycobacterium phage DaVinci unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51547 | 61.546 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704113 | Mycobacterium phage UPIE | Mycobacterium phage UPIE unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73784 | 58.793 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704109 | Mycobacterium phage Oosterbaan | Mycobacterium phage Oosterbaan Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68735 | 66.452 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704116 | Mycobacterium phage Drazdys | Mycobacterium phage Drazdys Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156281 | 64.685 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JN204348 | Mycobacterium phage Sebata | Mycobacterium phage Sebata Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155286 | 64.782 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JF704104 | Mycobacterium phage Zemanar | Mycobacterium phage Zemanar Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71092 | 68.876 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704103 | Mycobacterium phage Vortex | Mycobacterium phage Vortex unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68346 | 66.45 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704102 | Mycobacterium phage Phipps | Mycobacterium phage Phipps Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68293 | 66.461 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704100 | Mycobacterium phage Marvin | Mycobacterium phage Marvin Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65100 | 63.424 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704099 | Mycobacterium phage KLucky39 | Mycobacterium phage KLucky39 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68138 | 66.509 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704096 | Mycobacterium phage Ghost | Mycobacterium phage Ghost Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155167 | 64.648 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JF704095 | Mycobacterium phage Daisy | Mycobacterium phage Daisy unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68245 | 67.564 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704094 | Mycobacterium phage ChrisnMich | Mycobacterium phage ChrisnMich Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70428 | 69.075 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JF704092 | Mycobacterium phage Alice | Mycobacterium phage Alice Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153401 | 64.681 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| JF704091 | Mycobacterium phage ABU | Mycobacterium phage ABU Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68850 | 66.53 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN201525 | Mycobacterium phage Thibault | Mycobacterium phage Thibault Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106327 | 60.794 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN192463 | Mycobacterium phage Oline | Mycobacterium phage Oline Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68720 | 66.388 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN153086 | Mycobacterium phage Euphoria | Mycobacterium phage Euphoria Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53597 | 63.675 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN153085 | Mycobacterium phage Doom | Mycobacterium phage Doom Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51421 | 63.807 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN049605 | Mycobacterium phage EricB | Mycobacterium phage EricB Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51702 | 61.533 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN020143 | Mycobacterium phage ShiLan | Mycobacterium phage ShiLan Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59794 | 61.446 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN020142 | Mycobacterium phage Mutaforma13 | Mycobacterium phage Mutaforma13 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57701 | 61.283 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN020140 | Mycobacterium phage MrGordon | Mycobacterium phage MrGordon Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50988 | 63.752 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN006064 | Mycobacterium phage OSmaximus | Mycobacterium phage OSmaximus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69118 | 66.327 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN006063 | Mycobacterium phage Serendipity | Mycobacterium phage Serendipity Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68804 | 66.457 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| JN006062 | Mycobacterium phage Rakim | Mycobacterium phage Rakim unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75706 | 62.854 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JN006061 | Mycobacterium phage Toto | Mycobacterium phage Toto Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75933 | 63.017 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF957059 | Mycobacterium phage Optimus | Mycobacterium phage Optimus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109270 | 60.791 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| JF957058 | Mycobacterium phage HelDan | Mycobacterium phage HelDan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50364 | 63.984 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF957056 | Mycobacterium phage Thora | Mycobacterium phage Thora Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68839 | 66.47 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| HQ728524 | Mycobacterium phage Wee | Mycobacterium phage Wee Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59230 | 61.786 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| GU247134 | Mycobacterium phage Scoot17C | Mycobacterium phage Scoot17C Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68432 | 66.503 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| GU247133 | Mycobacterium phage Fang | Mycobacterium phage Fang Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68569 | 66.456 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| GU247132 | Mycobacterium phage SkiPole | Mycobacterium phage SkiPole Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53137 | 63.649 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| GQ303266 | Mycobacterium phage UncleHowie | Mycobacterium phage UncleHowie Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68016 | 66.508 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| GQ303264 | Mycobacterium phage Puhltonio | Mycobacterium phage Puhltonio Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68323 | 66.382 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| GQ303262 | Mycobacterium phage LRRHood | Mycobacterium phage LRRHood Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154349 | 64.732 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| GQ303261 | Mycobacterium phage Hope | Mycobacterium phage Hope unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.564 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| GQ303260 | Mycobacterium phage ET08 | Mycobacterium phage ET08 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155445 | 64.603 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| GQ303259 | Mycobacterium phage Colbert | Mycobacterium phage Colbert Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67774 | 66.495 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| FJ973624 | Mycobacterium phage Angel | Mycobacterium phage Angel unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41441 | 66.69 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ641182 | Mycobacterium phage Phlyer | Mycobacterium phage Phlyer unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69378 | 67.484 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| FJ174694 | Mycobacterium phage Chah | Mycobacterium phage Chah Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68450 | 66.498 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| FJ174691 | Mycobacterium phage Konstantine | Mycobacterium phage Konstantine Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68952 | 57.337 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| FJ168662 | Mycobacterium phage Troll4 | Mycobacterium phage Troll4 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64618 | 59.612 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| FJ168661 | Mycobacterium phage Gumball | Mycobacterium phage Gumball Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64807 | 59.554 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| FJ168660 | Mycobacterium phage Butterscotch | Mycobacterium phage Butterscotch Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64562 | 59.674 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| EU826471 | Mycobacterium phage Cali | Mycobacterium phage Cali Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155372 | 64.72 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| EU826470 | Mycobacterium phage Solon | Mycobacterium phage Solon Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49487 | 63.843 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU826468 | Mycobacterium phage Spud | Mycobacterium phage Spud Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154906 | 64.777 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| EU826467 | Mycobacterium phage Rizal | Mycobacterium phage Rizal Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153894 | 64.743 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| EU826466 | Mycobacterium phage Myrna | Mycobacterium phage Myrna Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164602 | 65.436 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| EU816589 | Mycobacterium phage Phaedrus | Mycobacterium phage Phaedrus unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68090 | 67.566 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| EU770221 | Mycobacterium phage Nigel | Mycobacterium phage Nigel Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69904 | 68.342 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| EU744252 | Mycobacterium phage DD5 | Mycobacterium phage DD5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51621 | 63.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| EU676000 | Mycobacterium phage Adjutor | Mycobacterium phage Adjutor Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64511 | 59.729 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| EU568876 | Mycobacterium phage BPs | Mycobacterium phage BPs unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.559 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| EF536069 | Mycobacterium phage Tweety | Mycobacterium phage Tweety Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58692 | 61.739 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ398053 | Mycobacterium phage Catera | Mycobacterium phage Catera Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153766 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| DQ398048 | Mycobacterium phage Qyrzula | Mycobacterium phage Qyrzula Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67188 | 68.963 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| DQ398051 | Mycobacterium phage PLot | Mycobacterium phage PLot Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64787 | 59.663 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| DQ398049 | Mycobacterium phage Pipefish | Mycobacterium phage Pipefish Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69059 | 67.335 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| DQ398047 | Mycobacterium phage PBI1 | Mycobacterium phage PBI1 Mycobacterium virus PBI1 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64494 | 59.711 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| DQ398046 | Mycobacterium phage Orion | Mycobacterium phage Orion Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68427 | 66.51 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| DQ398045 | Mycobacterium phage Llij | Mycobacterium phage Llij Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56852 | 61.53 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ398044 | Mycobacterium phage Cooper | Mycobacterium phage Cooper Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70654 | 69.106 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| AY500152 | Mycobacterium phage U2 | Mycobacterium phage U2 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 63.699 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AF547430 | Mycobacterium phage PG1 | Mycobacterium phage PG1 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68999 | 66.502 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| AY129337 | Mycobacterium phage Bxz1 | Mycobacterium phage Bxz1 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156102 | 64.77 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| AY129334 | Mycobacterium phage Rosebush | Mycobacterium phage Rosebush Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67480 | 68.957 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ514874 | Acinetobacter phage Ab105-2phideltaCI404ad | Acinetobacter phage Ab105-2phideltaCI404ad unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53211 | 39.479 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| MZ501055 | Escherichia phage CarlSpitteler | Escherichia phage CarlSpitteler unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39466 | 48.667 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| AF009630 | Lactococcus phage bIL170 | Lactococcus phage bIL170 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31754 | 34.345 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| OK570375 | Escherichia phage UoN_LDJ77_1 | Escherichia phage UoN_LDJ77_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111068 | 35.425 | Unclassified | Unclassified | Myoviridae | Escherichia |
| OK570374 | Escherichia phage UoN_LG358_1 | Escherichia phage UoN_LG358_1 unclassified Gelderlandvirus Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136254 | 36.764 | Gelderlandvirus | Tevenvirinae | Myoviridae | Escherichia |
| OL512807 | Vibrio phage V5 | Vibrio phage V5 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6658 | 43.677 | Unclassified | Unclassified | Inoviridae | Vibrio |
| OL441337 | Pseudomonas phage PaP_Se | Pseudomonas phage PaP_Se unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45938 | 52.534 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| OK625527 | Klebsiella phage vB_KpnP_ZK1 | Klebsiella phage vB_KpnP_ZK1 unclassified Molineuxvirinae Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45839 | 51.502 | Unclassified | Molineuxvirinae | Autographiviridae | Klebsiella |
| OK624832 | Xanthomonas phage DES1 | Xanthomonas phage DES1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64732 | 43.161 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| OK562670 | Stenotrophomonas phage vB_SM_ytsc_ply2008005c | Stenotrophomonas phage vB_SM_ytsc_ply2008005c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42318 | 63.02 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| OK562429 | Klebsiella phage P929 | Klebsiella phage P929 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44764 | 53.661 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OK546191 | Acinetobacter phage vB_AbaP_ABWU2101 | Acinetobacter phage vB_AbaP_ABWU2101 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41496 | 39.454 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ675741 | Acinetobacter phage Ab1656-2 | Acinetobacter phage Ab1656-2 unclassified bacterial viruses Viruses | 50929 | 38.593 | Unclassified | Unclassified | Unclassified | Acinetobacter |
| NC_054459 | Xanthomonas phage XPV1 | Xanthomonas phage XPV1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46503 | 61.121 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| NC_054458 | Xanthomonas phage XPP1 | Xanthomonas phage XPP1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46195 | 60.989 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| NC_048647 | Klebsiella phage KNP2 | Klebsiella phage KNP2 Klebsiella virus KNP2 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146203 | 38.548 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| NC_047881 | Shigella phage SSP1 | Shigella phage SSP1 Shigella virus SSP1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113299 | 39.076 | Tequintavirus | Markadamsvirinae | Demerecviridae | Shigella |
| OK490361 | Salmonella phage ZC-S1_prp | Salmonella phage ZC-S1_prp unclassified bacterial viruses Viruses | 19611 | 47.825 | Unclassified | Unclassified | Unclassified | Salmonella |
| OK483342 | Hafnia phage vB_HalM_SPARTY | Hafnia phage vB_HalM_SPARTY unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85494 | 43.735 | Kolesnikvirus | Ounavirinae | Myoviridae | Hafnia |
| OK490494 | Stenotrophomonas phage vB_SmaS_P15 | Stenotrophomonas phage vB_SmaS_P15 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43707 | 60.178 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| OK483199 | Vibrio phage vB_ValP_VA-RY-4 | Vibrio phage vB_ValP_VA-RY-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79532 | 45.692 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| OK428602 | Vibrio phage BX-1 | Vibrio phage BX-1 unclassified Vapseptimavirus Vapseptimavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144654 | 41.893 | Vapseptimavirus | Unclassified | Ackermannviridae | Vibrio |
| OK412919 | Stretomyces phage Pablito | Stretomyces phage Pablito unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49492 | 66.411 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| OV032902 | Vibrio phage vB_VpaM_sm033 | Vibrio phage vB_VpaM_sm033 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 320253 | 43.23 | Unclassified | Unclassified | Myoviridae | Vibrio |
| OV032860 | Vibrio phage vB_VpaS_sm030 | Vibrio phage vB_VpaS_sm030 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79660 | 45.674 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| OK638203 | Pseudomonas phage BHU-17 | Pseudomonas phage BHU-17 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85479 | 55.29 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| OK638202 | Pseudomonas phage BHU-15 | Pseudomonas phage BHU-15 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85815 | 55.265 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| OK638201 | Pseudomonas phage BHU-1 | Pseudomonas phage BHU-1 unclassified Samunavirus Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85609 | 55.261 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| OK663600 | Enterobacter phage vB_SEqdws-315 | Enterobacter phage vB_SEqdws-315 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40081 | 48.514 | Berlinvirus | Studiervirinae | Autographiviridae | Enterobacter |
| OK665842 | Burkholderia phage BgManors32 | Burkholderia phage BgManors32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59762 | 65.998 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| OK665841 | Burkholderia phage BgVeeders33 | Burkholderia phage BgVeeders33 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32945 | 62.076 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| OK655936 | Klebsiella phage phiW14 | Klebsiella phage phiW14 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50247 | 50.787 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OK913680 | Xanthomonas phage MET23-P3 | Xanthomonas phage MET23-P3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43010 | 66.743 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| OK913679 | Xanthomonas phage JUN5 | Xanthomonas phage JUN5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42912 | 66.813 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MZ571833 | Klebsiella phage vB_Kpn-VAC112 | Klebsiella phage vB_Kpn-VAC112 unclassified bacterial viruses Viruses | 49068 | 50.22 | Unclassified | Unclassified | Unclassified | Klebsiella |
| MZ571831 | Klebsiella phage vB_Kpn-VAC70 | Klebsiella phage vB_Kpn-VAC70 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49631 | 50.7 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ571827 | Klebsiella phage vB_Kpn-VAC25 | Klebsiella phage vB_Kpn-VAC25 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43777 | 53.747 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| OK210077 | Serratia phage KKP_3263 | Serratia phage KKP_3263 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148182 | 37.509 | Unclassified | Vequintavirinae | Myoviridae | Serratia |
| OK210076 | Enterobacter phage KKP_3262 | Enterobacter phage KKP_3262 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84075 | 39.508 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| OK210075 | Citrobacter phage KKP_3664 | Citrobacter phage KKP_3664 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61608 | 43.509 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| OK210074 | Enterobacter phage KKP_3263 | Enterobacter phage KKP_3263 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39418 | 51.142 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| LC644976 | Komagataeibacter phage phiKX2 | Komagataeibacter phage phiKX2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53798 | 60.004 | Unclassified | Unclassified | Myoviridae | Komagataeibacter |
| LC644975 | Komagataeibacter phage phiKX1 | Komagataeibacter phage phiKX1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53192 | 60.161 | Unclassified | Unclassified | Myoviridae | Komagataeibacter |
| LC644974 | Komagataeibacter phage phiKM1 | Komagataeibacter phage phiKM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51977 | 62.185 | Unclassified | Unclassified | Myoviridae | Komagataeibacter |
| LC644973 | Acetobacter phage phiAX1 | Acetobacter phage phiAX1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50652 | 56.047 | Unclassified | Unclassified | Podoviridae | Acetobacter |
| LC644972 | Acetobacter phage phiAP1 | Acetobacter phage phiAP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42805 | 52.391 | Unclassified | Unclassified | Myoviridae | Acetobacter |
| LC644971 | Acetobacter phage phiAO1 | Acetobacter phage phiAO1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34100 | 61.364 | Unclassified | Unclassified | Myoviridae | Acetobacter |
| NC_028749 | Brevibacillus phage Sundance | Brevibacillus phage Sundance Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134270 | 35.504 | Unclassified | Unclassified | Siphoviridae | Brevibacillus |
| MZ612110 | Klebsiella phage vB_Kpn-VAC111 | Klebsiella phage vB_Kpn-VAC111 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15134 | 51.625 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ571834 | Klebsiella phage vB_Kpn-VAC113 | Klebsiella phage vB_Kpn-VAC113 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49568 | 50.456 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ571832 | Klebsiella phage vB_Kpn-VAC71 | Klebsiella phage vB_Kpn-VAC71 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40388 | 52.979 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MZ571830 | Klebsiella phage vB_Kpn-VAC51 | Klebsiella phage vB_Kpn-VAC51 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113149 | 45.402 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MZ571829 | Klebsiella phage vB_Kpn-VAC36 | Klebsiella phage vB_Kpn-VAC36 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169970 | 40.904 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MZ571828 | Klebsiella phage vB_Kpn-VAC35 | Klebsiella phage vB_Kpn-VAC35 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112862 | 45.438 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| OK138556 | Klebsiella phage vB_Kp_IME531 | Klebsiella phage vB_Kp_IME531 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40314 | 53.031 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| OK138555 | Klebsiella phage vB_Kp_IME328 | Klebsiella phage vB_Kp_IME328 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48841 | 50.347 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| OK094665 | Pseudomonas phage vB_Pae_W3 | Pseudomonas phage vB_Pae_W3 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43014 | 53.81 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| OK094664 | Shewanella phage vB_Sb_QDWS | Shewanella phage vB_Sb_QDWS Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47902 | 63.169 | Unclassified | Unclassified | Unclassified | Shewanella |
| MZ955867 | Pseudomonas phage vB_PaeP_TUMS_P121 | Pseudomonas phage vB_PaeP_TUMS_P121 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73001 | 54.928 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MZ958750 | Streptomyces phage Targaryen | Streptomyces phage Targaryen unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134920 | 49.529 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| MZ958749 | Microbacterium phage SJay | Microbacterium phage SJay unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41847 | 63.484 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ958748 | Mycobacterium phage JuicyJay | Mycobacterium phage JuicyJay unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113075 | 60.85 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ958747 | Mycobacterium phage InvictusManeo | Mycobacterium phage InvictusManeo unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61147 | 65.632 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ958746 | Mycobacterium phage Enceladus | Mycobacterium phage Enceladus unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66420 | 58.71 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ958745 | Mycobacterium phage Pinkcreek | Mycobacterium phage Pinkcreek unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153184 | 64.654 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MZ958744 | Mycobacterium phage JalFarm20 | Mycobacterium phage JalFarm20 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58768 | 61.396 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ501113 | Escherichia phage WilhelmHis | Escherichia phage WilhelmHis unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166861 | 35.437 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501112 | Escherichia phage WildeMaa | Escherichia phage WildeMaa unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60090 | 44.299 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501111 | Escherichia phage WalterGehring | Escherichia phage WalterGehring unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140659 | 43.561 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501110 | Escherichia phage VogelGryff | Escherichia phage VogelGryff unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58342 | 44.207 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501109 | Escherichia phage TrudiRoth | Escherichia phage TrudiRoth unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114488 | 39.719 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501108 | Escherichia phage TrudiGerster | Escherichia phage TrudiGerster unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114179 | 40.683 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501107 | Escherichia phage TheodorHerzl | Escherichia phage TheodorHerzl unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43627 | 54.436 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ501106 | Escherichia phage TadeuszReichstein | Escherichia phage TadeuszReichstein unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166115 | 35.428 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501105 | Escherichia phage SuperGirl | Escherichia phage SuperGirl unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110821 | 40.054 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501104 | Escherichia phage SeppeHuegi | Escherichia phage SeppeHuegi unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60068 | 44.654 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501103 | Escherichia phage SelmaRatti | Escherichia phage SelmaRatti unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107080 | 38.821 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501102 | Escherichia phage RudolfGeigy | Escherichia phage RudolfGeigy unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137380 | 43.664 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501101 | Escherichia phage PeterMerian | Escherichia phage PeterMerian unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47775 | 44.716 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501100 | Escherichia phage PaulScherrer | Escherichia phage PaulScherrer unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150425 | 37.501 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MZ501099 | Escherichia phage PaulSarasin | Escherichia phage PaulSarasin unclassified Changchunvirus Changchunvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51285 | 46.386 | Changchunvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501098 | Escherichia phage PaulHMueller | Escherichia phage PaulHMueller unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170053 | 35.645 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501097 | Escherichia phage PaulFeyerabend | Escherichia phage PaulFeyerabend unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44550 | 54.382 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ501096 | Escherichia phage Paracelsus | Escherichia phage Paracelsus unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167732 | 35.412 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501095 | Escherichia phage Oekolampad | Escherichia phage Oekolampad unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44882 | 54.519 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ501094 | Escherichia phage MaxTheCat | Escherichia phage MaxTheCat unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135061 | 43.636 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501093 | Escherichia phage MaxBurger | Escherichia phage MaxBurger unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136346 | 43.522 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501092 | Escherichia phage LeonhardEuler | Escherichia phage LeonhardEuler unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50192 | 45.848 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MZ501091 | Escherichia phage KurtStettler | Escherichia phage KurtStettler unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58567 | 44.655 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501090 | Escherichia phage KarlJaspers | Escherichia phage KarlJaspers unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51474 | 43.956 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501089 | Escherichia phage KarlGJung | Escherichia phage KarlGJung unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167832 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501088 | Escherichia phage KarlBarth | Escherichia phage KarlBarth unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43104 | 54.61 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ501087 | Escherichia phage JulesPiccard | Escherichia phage JulesPiccard unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47731 | 44.665 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MZ501086 | Escherichia phage JohannRWettstein | Escherichia phage JohannRWettstein unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87100 | 39.029 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MZ501085 | Escherichia phage JohannLBurckhardt | Escherichia phage JohannLBurckhardt unclassified Henuseptimavirus Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49715 | 48.579 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501084 | Escherichia phage JohannJBalmer | Escherichia phage JohannJBalmer unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144831 | 37.436 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MZ501083 | Escherichia phage JohannBauhin | Escherichia phage JohannBauhin unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135602 | 43.634 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501082 | Escherichia phage JeffSchatz | Escherichia phage JeffSchatz unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135630 | 43.687 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501081 | Escherichia phage JeanTinguely | Escherichia phage JeanTinguely Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39842 | 48.492 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ501080 | Escherichia phage JeanPiccard | Escherichia phage JeanPiccard unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47149 | 44.531 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MZ501079 | Escherichia phage JakobBernoulli | Escherichia phage JakobBernoulli unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51129 | 43.934 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501078 | Escherichia phage JacobBurckhardt | Escherichia phage JacobBurckhardt Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39451 | 48.711 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ501077 | Escherichia phage IsaakIselin | Escherichia phage IsaakIselin unclassified Henuseptimavirus Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49911 | 48.212 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501076 | Escherichia phage IrmaTschudi | Escherichia phage IrmaTschudi unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115446 | 40.145 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501075 | Escherichia phage IrisVonRoten | Escherichia phage IrisVonRoten unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112239 | 39.027 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501074 | Escherichia phage HildyBeyeler | Escherichia phage HildyBeyeler unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111607 | 39.12 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501073 | Escherichia phage HeinrichReichert | Escherichia phage HeinrichReichert unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136906 | 43.613 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501072 | Escherichia phage GreteKellenberger | Escherichia phage GreteKellenberger unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114196 | 40.189 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501071 | Escherichia phage GottfriedDienst | Escherichia phage GottfriedDienst unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56589 | 44.692 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501070 | Escherichia phage GeorgBuechner | Escherichia phage GeorgBuechner unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44295 | 54.647 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ501069 | Escherichia phage FritzSarasin | Escherichia phage FritzSarasin unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51777 | 44.775 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501068 | Escherichia phage FritzHoffmann | Escherichia phage FritzHoffmann unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59834 | 43.5 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501067 | Escherichia phage FriedrichZschokke | Escherichia phage FriedrichZschokke unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165574 | 35.419 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501066 | Escherichia phage FriedrichMiescher | Escherichia phage FriedrichMiescher unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169509 | 35.447 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501065 | Escherichia phage FelixPlatter | Escherichia phage FelixPlatter unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164714 | 35.492 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501064 | Escherichia phage ErnstBeyeler | Escherichia phage ErnstBeyeler unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39315 | 48.956 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ501063 | Escherichia phage EmilieFrey | Escherichia phage EmilieFrey unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146666 | 37.433 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MZ501062 | Escherichia phage EmilHeitz | Escherichia phage EmilHeitz unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139980 | 43.684 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501061 | Escherichia phage EduardKellenberger | Escherichia phage EduardKellenberger unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131526 | 43.695 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501060 | Escherichia phage DrSchubert | Escherichia phage DrSchubert unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138240 | 43.448 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501059 | Escherichia phage DanielBernoulli | Escherichia phage DanielBernoulli unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50731 | 43.159 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ501058 | Escherichia phage DaisyDussoix | Escherichia phage DaisyDussoix unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113532 | 38.969 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ501057 | Escherichia phage ChristophMerian | Escherichia phage ChristophMerian unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58073 | 43.478 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MZ501056 | Escherichia phage ChristianSchoenbein | Escherichia phage ChristianSchoenbein unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169031 | 37.693 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501054 | Escherichia phage CarlMeissner | Escherichia phage CarlMeissner unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137623 | 43.703 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501053 | Escherichia phage BrunoManser | Escherichia phage BrunoManser unclassified Sertoctavirus Sertoctavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49298 | 48.111 | Sertoctavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MZ501052 | Escherichia phage AugustSocin | Escherichia phage AugustSocin unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167411 | 35.395 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501051 | Escherichia phage AugustePiccard | Escherichia phage AugustePiccard unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50126 | 44.755 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MZ501050 | Escherichia phage AndreasVesalius | Escherichia phage AndreasVesalius unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168255 | 35.457 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501049 | Escherichia phage AlfredRasser | Escherichia phage AlfredRasser unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70849 | 41.327 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MZ501048 | Escherichia phage AlexBoehm | Escherichia phage AlexBoehm unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135345 | 43.762 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ501046 | Escherichia phage AdolfPortmann | Escherichia phage AdolfPortmann unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166773 | 35.454 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ501047 | Escherichia phage AlbertHofmann | Escherichia phage AlbertHofmann unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168968 | 37.672 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234053 | Escherichia phage vB_EcoM-PM136 | Escherichia phage vB_EcoM-PM136 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169519 | 35.334 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234051 | Escherichia phage vB_EcoM-PM108 | Escherichia phage vB_EcoM-PM108 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169415 | 35.368 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234050 | Escherichia phage vB_EcoP-UTI89UKE3 | Escherichia phage vB_EcoP-UTI89UKE3 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44294 | 45.076 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234049 | Escherichia phage vB_EcoP-UTI89UKE2 | Escherichia phage vB_EcoP-UTI89UKE2 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44293 | 45.079 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234047 | Escherichia phage vB_EcoP-PM139 | Escherichia phage vB_EcoP-PM139 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40193 | 49.874 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234046 | Escherichia phage vB_EcoP-PM137 | Escherichia phage vB_EcoP-PM137 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40193 | 49.877 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234045 | Escherichia phage vB_EcoP-PM131 | Escherichia phage vB_EcoP-PM131 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40193 | 49.879 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234044 | Escherichia phage vB_EcoP-PM129 | Escherichia phage vB_EcoP-PM129 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40193 | 49.879 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234043 | Escherichia phage vB_EcoM-PM123 | Escherichia phage vB_EcoM-PM123 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169448 | 35.341 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234042 | Escherichia phage vB_EcoM-PM107 | Escherichia phage vB_EcoM-PM107 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169416 | 35.364 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234041 | Escherichia phage vB_EcoM-PM105 | Escherichia phage vB_EcoM-PM105 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169499 | 35.367 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234040 | Escherichia phage vB_EcoM-G3G7 | Escherichia phage vB_EcoM-G3G7 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168649 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234039 | Escherichia phage vB_EcoM-G3G6 | Escherichia phage vB_EcoM-G3G6 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168522 | 35.37 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234038 | Escherichia phage vB_EcoM-G3F10 | Escherichia phage vB_EcoM-G3F10 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167867 | 35.448 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234037 | Escherichia phage vB_EcoM-G3F9 | Escherichia phage vB_EcoM-G3F9 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168524 | 35.369 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234036 | Escherichia phage vB_EcoM-G3F8 | Escherichia phage vB_EcoM-G3F8 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168524 | 35.37 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234035 | Escherichia phage vB_EcoM-G3F7 | Escherichia phage vB_EcoM-G3F7 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167839 | 35.444 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234034 | Escherichia phage vB_EcoM-G3F6 | Escherichia phage vB_EcoM-G3F6 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168524 | 35.37 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234033 | Escherichia phage vB_EcoS-G3B1 | Escherichia phage vB_EcoS-G3B1 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51299 | 44.553 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MZ234032 | Escherichia phage vB_EcoP-G3A1 | Escherichia phage vB_EcoP-G3A1 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44471 | 44.989 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234031 | Escherichia phage vB_EcoM-CHD94UKE2 | Escherichia phage vB_EcoM-CHD94UKE2 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167922 | 35.452 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234030 | Escherichia phage vB_EcoM-CHD16UKE1 | Escherichia phage vB_EcoM-CHD16UKE1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168543 | 35.362 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234028 | Escherichia phage vB_EcoP-CHD5UKE1 | Escherichia phage vB_EcoP-CHD5UKE1 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77359 | 42.155 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MZ234027 | Escherichia phage vB_EcoM-CHD2BS1 | Escherichia phage vB_EcoM-CHD2BS1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168577 | 35.369 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234025 | Escherichia phage vB_EcoM-172859UKE1 | Escherichia phage vB_EcoM-172859UKE1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168667 | 37.661 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234024 | Escherichia phage vB_EcoP-101136BS1 | Escherichia phage vB_EcoP-101136BS1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39375 | 50.154 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234023 | Escherichia phage vB_EcoP-101120B2 | Escherichia phage vB_EcoP-101120B2 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39899 | 49.926 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234022 | Escherichia phage vB_EcoP-101120B1-2 | Escherichia phage vB_EcoP-101120B1-2 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39845 | 49.933 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234021 | Escherichia phage vB_EcoP-101118UKE1 | Escherichia phage vB_EcoP-101118UKE1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40233 | 50.068 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234020 | Escherichia phage vB_EcoP-101118B1 | Escherichia phage vB_EcoP-101118B1 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44526 | 44.996 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234019 | Escherichia phage vB_EcoP-101117UKE2 | Escherichia phage vB_EcoP-101117UKE2 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44526 | 44.985 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234018 | Escherichia phage vB_EcoM-101117BS1 | Escherichia phage vB_EcoM-101117BS1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167080 | 35.413 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234017 | Escherichia phage vB_EcoP-101114UKE3 | Escherichia phage vB_EcoP-101114UKE3 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75747 | 42.018 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MZ234016 | Escherichia phage vB_EcoS-101114BS4 | Escherichia phage vB_EcoS-101114BS4 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45251 | 54.363 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ234014 | Escherichia phage vB_EcoS-101114B2 | Escherichia phage vB_EcoS-101114B2 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44971 | 54.491 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MZ234013 | Escherichia phage vB_EcoM-101112UKE3-1 | Escherichia phage vB_EcoM-101112UKE3-1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169555 | 35.222 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ234012 | Escherichia phage vB_EcoP-101101UKE1 | Escherichia phage vB_EcoP-101101UKE1 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44450 | 45.066 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ234011 | Escherichia phage vB_EcoP-22664UKE3-2 | Escherichia phage vB_EcoP-22664UKE3-2 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40482 | 49.827 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234010 | Escherichia phage vB_EcoS-22664BS2 | Escherichia phage vB_EcoS-22664BS2 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45176 | 50.39 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MZ234009 | Escherichia phage vB_EcoP-22664BS1 | Escherichia phage vB_EcoP-22664BS1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39133 | 50.211 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ234008 | Escherichia phage vB_EcoS-22664B1 | Escherichia phage vB_EcoS-22664B1 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45019 | 54.506 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| OK275722 | Escherichia phage HP3.1 | Escherichia phage HP3.1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168195 | 35.374 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ398021 | Klebsiella phage BUCT_47333 | Klebsiella phage BUCT_47333 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47333 | 49.16 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MZ374361 | Klebsiella phage BUCT_49532 | Klebsiella phage BUCT_49532 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49532 | 48.815 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MW258712 | Kosakonia phage Kc259 | Kosakonia phage Kc259 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41192 | 58.133 | Unclassified | Unclassified | Podoviridae | Kosakonia |
| MW317014 | Kosakonia phage Kc166BP | Kosakonia phage Kc166BP unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 6014 | 49.618 | Unclassified | Peduovirinae | Myoviridae | Kosakonia |
| MG913376 | Lactobacillus phage PMBT4 | Lactobacillus phage PMBT4 unclassified Cequinquevirus Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31399 | 41.629 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| LC625742 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 37025 | 38.612 | Unclassified | Unclassified | Unclassified | Unspecified |
| OK905446 | Vibrio phage VS1 | Vibrio phage VS1 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 248270 | 42.5 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MK568540 | Vibrio phage Va3 | Vibrio phage Va3 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 247567 | 41.337 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MK387337 | Vibrio phage Va1 | Vibrio phage Va1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194900 | 34.855 | Unclassified | Unclassified | Myoviridae | Vibrio |
| OK018184 | Shigella phage CT01 | Shigella phage CT01 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168362 | 35.594 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MZ364294 | robinz microvirus RP_188 | robinz microvirus RP_188 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4136 | 41.779 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364293 | robinz microvirus RP_178 | robinz microvirus RP_178 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4260 | 59.272 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364292 | robinz microvirus RP_175 | robinz microvirus RP_175 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4267 | 53.105 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364291 | robinz microvirus RP_172 | robinz microvirus RP_172 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4315 | 55.226 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364290 | robinz microvirus RP_171 | robinz microvirus RP_171 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4322 | 46.923 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364289 | robinz microvirus RP_165 | robinz microvirus RP_165 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4329 | 51.582 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364288 | robinz microvirus RP_163 | robinz microvirus RP_163 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4329 | 43.405 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364287 | robinz microvirus RP_160 | robinz microvirus RP_160 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4144 | 56.781 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364286 | robinz microvirus RP_153 | robinz microvirus RP_153 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4343 | 60.465 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364285 | robinz microvirus RP_152 | robinz microvirus RP_152 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4415 | 50.419 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364284 | robinz microvirus RP_147 | robinz microvirus RP_147 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4447 | 50.978 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364283 | robinz microvirus RP_145 | robinz microvirus RP_145 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4333 | 54.927 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364282 | robinz microvirus RP_144 | robinz microvirus RP_144 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4140 | 39.686 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364281 | robinz microvirus RP_143 | robinz microvirus RP_143 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4506 | 50.644 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364280 | robinz microvirus RP_142 | robinz microvirus RP_142 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4480 | 51.674 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364279 | robinz microvirus RP_139 | robinz microvirus RP_139 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4559 | 53.389 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364278 | robinz microvirus RP_136 | robinz microvirus RP_136 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4534 | 55.845 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364277 | robinz microvirus RP_133 | robinz microvirus RP_133 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4539 | 49.725 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364276 | robinz microvirus RP_132 | robinz microvirus RP_132 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4548 | 51.209 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364275 | robinz microvirus RP_131 | robinz microvirus RP_131 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4554 | 48.573 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364274 | robinz microvirus RP_121 | robinz microvirus RP_121 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4566 | 56.045 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364273 | robinz microvirus RP_120 | robinz microvirus RP_120 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4572 | 49.213 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364272 | robinz microvirus RP_117 | robinz microvirus RP_117 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4687 | 47.408 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364271 | robinz microvirus RP_115 | robinz microvirus RP_115 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4662 | 48.498 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364270 | robinz microvirus RP_112 | robinz microvirus RP_112 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4694 | 53.749 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364269 | robinz microvirus RP_111 | robinz microvirus RP_111 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4693 | 48.562 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364268 | robinz microvirus RP_110 | robinz microvirus RP_110 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4720 | 48.962 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364267 | robinz microvirus RP_109 | robinz microvirus RP_109 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4717 | 48.442 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364266 | robinz microvirus RP_107 | robinz microvirus RP_107 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4580 | 52.205 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364265 | robinz microvirus RP_106 | robinz microvirus RP_106 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4726 | 47.778 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364264 | robinz microvirus RP_105 | robinz microvirus RP_105 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4673 | 49.925 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364263 | robinz microvirus RP_104 | robinz microvirus RP_104 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4750 | 55.979 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364262 | robinz microvirus RP_103 | robinz microvirus RP_103 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4745 | 46.807 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364261 | robinz microvirus RP_102 | robinz microvirus RP_102 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4670 | 48.844 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364260 | robinz microvirus RP_101 | robinz microvirus RP_101 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4767 | 56.094 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364259 | robinz microvirus RP_97 | robinz microvirus RP_97 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4771 | 49.235 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364258 | robinz microvirus RP_96 | robinz microvirus RP_96 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4724 | 52.477 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364257 | robinz microvirus RP_95 | robinz microvirus RP_95 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4491 | 52.973 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364256 | robinz microvirus RP_94 | robinz microvirus RP_94 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4642 | 54.718 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364255 | robinz microvirus RP_93 | robinz microvirus RP_93 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4811 | 46.518 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364254 | robinz microvirus RP_92 | robinz microvirus RP_92 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4763 | 50.892 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364253 | robinz microvirus RP_87 | robinz microvirus RP_87 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4776 | 57.684 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364252 | robinz microvirus RP_86 | robinz microvirus RP_86 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4291 | 58.425 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364251 | robinz microvirus RP_84 | robinz microvirus RP_84 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4162 | 41.567 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364250 | robinz microvirus RP_82 | robinz microvirus RP_82 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4963 | 54.604 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364249 | robinz microvirus RP_81 | robinz microvirus RP_81 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4964 | 54.392 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364248 | robinz microvirus RP_78 | robinz microvirus RP_78 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4642 | 49.203 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364247 | robinz microvirus RP_77 | robinz microvirus RP_77 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4854 | 54.141 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364246 | robinz microvirus RP_76 | robinz microvirus RP_76 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4623 | 52.65 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364245 | robinz microvirus RP_75 | robinz microvirus RP_75 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4932 | 54.4 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364244 | robinz microvirus RP_74 | robinz microvirus RP_74 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4681 | 53.514 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364243 | robinz microvirus RP_71 | robinz microvirus RP_71 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 55.686 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364242 | robinz microvirus RP_65 | robinz microvirus RP_65 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4561 | 47.183 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364241 | robinz microvirus RP_62 | robinz microvirus RP_62 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4812 | 54.385 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364240 | robinz microvirus RP_61 | robinz microvirus RP_61 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4603 | 51.532 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364239 | robinz microvirus RP_58 | robinz microvirus RP_58 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4289 | 51.993 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364238 | robinz microvirus RP_56 | robinz microvirus RP_56 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4403 | 51.737 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364237 | robinz microvirus RP_48 | robinz microvirus RP_48 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4642 | 46.704 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364236 | robinz microvirus RP_45 | robinz microvirus RP_45 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4288 | 52.215 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364235 | robinz microvirus RP_44 | robinz microvirus RP_44 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 52.472 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364234 | robinz microvirus RP_42 | robinz microvirus RP_42 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4499 | 41.743 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364233 | robinz microvirus RP_41 | robinz microvirus RP_41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4567 | 50.055 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364232 | robinz microvirus RP_40 | robinz microvirus RP_40 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4686 | 49.168 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364231 | robinz microvirus RP_39 | robinz microvirus RP_39 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4466 | 50.94 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364230 | robinz microvirus RP_38 | robinz microvirus RP_38 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4474 | 40.165 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364229 | robinz microvirus RP_37 | robinz microvirus RP_37 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4320 | 43.356 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364228 | robinz microvirus RP_36 | robinz microvirus RP_36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4718 | 48.432 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364227 | robinz microvirus RP_35 | robinz microvirus RP_35 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4364 | 57.195 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364226 | robinz microvirus RP_34 | robinz microvirus RP_34 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4780 | 54.226 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364225 | robinz microvirus RP_33 | robinz microvirus RP_33 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4384 | 50.091 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364224 | robinz microvirus RP_32 | robinz microvirus RP_32 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4384 | 50.091 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364223 | robinz microvirus RP_30 | robinz microvirus RP_30 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4722 | 50.339 | Unclassified | Unclassified | Microviridae | Unspecified |
| MZ364222 | robinz microvirus RP_29 | robinz microvirus RP_29 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4731 | 50.328 | Unclassified | Unclassified | Microviridae | Unspecified |
| OK041469 | Achromobacter phage AXY1 | Achromobacter phage AXY1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61950 | 60.316 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| OK041468 | Burkholderia phage BCE1 | Burkholderia phage BCE1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14800 | 48.209 | Unclassified | Unclassified | Podoviridae | Burkholderia |
| OK041467 | Pseudomonas phage PAE2 | Pseudomonas phage PAE2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35826 | 57.123 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| OK136181 | Escherichia phage 18-1-2 | Escherichia phage 18-1-2 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54435 | 39.535 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK136180 | Escherichia phage 18-1-2 | Escherichia phage 18-1-2 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79447 | 40.244 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| OK173051 | Streptococcus phage SF39 | Streptococcus phage SF39 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41257 | 39.877 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MZ622163 | Mycobacterium phage Filch | Mycobacterium phage Filch unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75692 | 62.923 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ477002 | Acinetobacter phage Phab24 | Acinetobacter phage Phab24 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93604 | 33.653 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MT178275 | Klebsiella phage KpS8 | Klebsiella phage KpS8 Klebsiella virus KpS8 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143800 | 44.583 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| MN478483 | Klebsiella phage UPM 2146 | Klebsiella phage UPM 2146 Klebsiella virus UPM2146 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160795 | 46.306 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MN316588 | Escherichia phage VEc33 | Escherichia phage VEc33 Escherichia virus VEc33 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108640 | 39.109 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KX581096 | Vibrio phage vB_VspP_pVa5_12Jun | Vibrio phage vB_VspP_pVa5_12Jun Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53227 | 43.021 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| NC_031129 | Salmonella phage SJ46 | Salmonella phage SJ46 Salmonella virus SJ46 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103445 | 48.578 | Punavirus | Unclassified | Myoviridae | Salmonella |
| NC_028903 | Escherichia phage 172-1 | Escherichia phage 172-1 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77266 | 41.984 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| NC_027350 | Citrobacter phage Stevie | Citrobacter phage Stevie Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49816 | 42.838 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| NC_025824 | Escherichia phage fd | Escherichia phage fd Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6408 | 40.886 | Inovirus | Unclassified | Inoviridae | Escherichia |
| NC_023593 | Escherichia phage KBNP1711 | Escherichia phage KBNP1711 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76184 | 42.37 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| NC_022989 | Lactobacillus phage LL-Ku | Lactobacillus phage LL-Ku Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31080 | 41.602 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_019917 | Burkholderia phage BcepMigl | Burkholderia phage BcepMigl Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62952 | 65.505 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| NC_019916 | Lactobacillus phage ATCC8014 | Lactobacillus phage ATCC8014 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38002 | 47.597 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_019449 | Lactobacillus phage c5 | Lactobacillus phage c5 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31841 | 42.018 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| JX486087 | Lactobacillus phage ATCC8014 | Lactobacillus phage ATCC8014 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38002 | 47.597 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_017688 | Lactococcus phage ASCC191 | Lactococcus phage ASCC191 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32414 | 34.112 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740813 | Lactococcus phage ASCC532 | Lactococcus phage ASCC532 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32384 | 34.687 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740789 | Lactococcus phage ASCC281 | Lactococcus phage ASCC281 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32023 | 34.766 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740788 | Lactococcus phage ASCC273 | Lactococcus phage ASCC273 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32582 | 34.528 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740787 | Lactococcus phage ASCC191 | Lactococcus phage ASCC191 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32414 | 34.112 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740804 | Lactococcus phage ASCC465 | Lactococcus phage ASCC465 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31772 | 34.644 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_016570 | Escherichia phage Cba120 | Escherichia phage Cba120 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157304 | 44.499 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| NC_016562 | Campylobacter phage CPX | Campylobacter phage CPX Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132662 | 26.043 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_012743 | Burkholderia phage BcepIL02 | Burkholderia phage BcepIL02 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62715 | 66.203 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| EU340421 | Lactobacillus phage c5 | Lactobacillus phage c5 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31841 | 42.018 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| AY739900 | Lactobacillus phage LL-Ku | Lactobacillus phage LL-Ku Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31080 | 41.602 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_015464 | Campylobacter phage NCTC12673 | Campylobacter phage NCTC12673 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135041 | 26.154 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| FJ822135 | Lactobacillus phage Lb338-1 | Lactobacillus phage Lb338-1 Mooreparkvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142111 | 37.398 | Mooreparkvirus | Unclassified | Herelleviridae | Lactobacillus |
| CP000917 | Escherichia phage Eps7 | Escherichia phage Eps7 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111382 | 39.901 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_010324 | Escherichia phage phiEco32 | Escherichia phage phiEco32 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77554 | 42.274 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| DQ227763 | Lactococcus phage 712 | Lactococcus phage 712 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30510 | 33.884 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| AY682195 | Lactobacillus phage LP65 | Lactobacillus phage LP65 Salchichonvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131522 | 37.321 | Salchichonvirus | Unclassified | Herelleviridae | Lactobacillus |
| AY543070 | Escherichia phage T5 | Escherichia phage T5 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121750 | 39.27 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| AY308796 | Escherichia phage Tls | Escherichia phage Tls Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49902 | 42.68 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| AY236756 | Lactobacillus phage phiJL-1 | Lactobacillus phage phiJL-1 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36674 | 39.355 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_001422 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| PFDCG | Escherichia phage fd | Escherichia phage fd Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6408 | 40.886 | Inovirus | Unclassified | Inoviridae | Escherichia |
| MZ398245 | Klebsiella phage IME183 | Klebsiella phage IME183 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41384 | 52.921 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| OK040807 | Escherichia phage C600M2 | Escherichia phage C600M2 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88159 | 38.983 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MW882907 | Escherichia phage vB_EcoM_SQ17 | Escherichia phage vB_EcoM_SQ17 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166457 | 37.524 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| OK018136 | Stenotrophomonas phage TS-10 | Stenotrophomonas phage TS-10 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42931 | 59.929 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| OK040794 | Arthrobacter phage Atuin | Arthrobacter phage Atuin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176888 | 54.004 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| OK040793 | Microbacterium phage Zooman | Microbacterium phage Zooman Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192730 | 58.592 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK040792 | Microbacterium phage Kozie | Microbacterium phage Kozie unclassified Mementomorivirus Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53094 | 71.117 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| OK040791 | Mycobacterium phage Parliament | Mycobacterium phage Parliament unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53727 | 63.927 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OK040790 | Microbacterium phage Pumpernickel | Microbacterium phage Pumpernickel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186095 | 56.485 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK040789 | Mycobacterium phage Izajani | Mycobacterium phage Izajani unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156009 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| OK040788 | Microbacterium phage Cybele | Microbacterium phage Cybele unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56694 | 63.832 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| OK040787 | Microbacterium phage Pepe25 | Microbacterium phage Pepe25 unclassified Gillianvirus Gillianvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60998 | 67.26 | Gillianvirus | Unclassified | Siphoviridae | Microbacterium |
| OK040786 | Gordonia phage Kudefre | Gordonia phage Kudefre Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76731 | 58.619 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| OK040785 | Gordonia phage Pons | Gordonia phage Pons Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47982 | 60.646 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| OK040784 | Mycobacterium phage CaseJules | Mycobacterium phage CaseJules unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59905 | 66.557 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| OK040783 | Microbacterium phage Eightball | Microbacterium phage Eightball Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17439 | 68.737 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK040782 | Mycobacterium phage Illumine | Mycobacterium phage Illumine unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60620 | 66.349 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| OK040781 | Microbacterium phage Gardevoir | Microbacterium phage Gardevoir Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17396 | 68.521 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK040780 | Arthrobacter phage Sarge | Arthrobacter phage Sarge Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36407 | 63.485 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| OK040779 | Microbacterium phage Gretchen | Microbacterium phage Gretchen Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47463 | 69.574 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| OK040778 | Mycobacterium phage Devera | Mycobacterium phage Devera unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60618 | 66.348 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| OK040777 | Arthrobacter phage KeAlii | Arthrobacter phage KeAlii unclassified Manhattanvirus Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41850 | 65.46 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| OK108609 | Salmonella phage vB_SenS_S532 | Salmonella phage vB_SenS_S532 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43620 | 49.723 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OK108608 | Salmonella phage vB_SenS_S528 | Salmonella phage vB_SenS_S528 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42662 | 49.674 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| OK108607 | Salmonella phage vB_SenS_S124 | Salmonella phage vB_SenS_S124 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110832 | 39.179 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| OK169295 | Vibrio phage Rostov 13 | Vibrio phage Rostov 13 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14072 | 42.418 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| OK169294 | Vibrio phage Rostov 13 | Vibrio phage Rostov 13 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22254 | 43.075 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| NC_042038 | Escherichia phage phiKP26 | Escherichia phage phiKP26 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47285 | 44.257 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KX279892 | Escherichia phage AAPEc6 | Escherichia phage AAPEc6 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44559 | 45.176 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| KM979355 | Escherichia phage DT571/2 | Escherichia phage DT571/2 Escherichia virus DT5712 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108418 | 39.678 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KC579452 | Escherichia phage phiKP26 | Escherichia phage phiKP26 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47285 | 44.257 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| JX104231 | Burkholderia phage BcepMigl | Burkholderia phage BcepMigl Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62952 | 65.505 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| JN672684 | Enterobacter phage F20 | Enterobacter phage F20 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51543 | 47.902 | Webervirus | Unclassified | Drexlerviridae | Enterobacter |
| FJ937737 | Burkholderia phage BcepIL02 | Burkholderia phage BcepIL02 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62715 | 66.203 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| JN593240 | Escherichia phage Cba120 | Escherichia phage Cba120 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157304 | 44.499 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| EU330206 | Escherichia phage phiEco32 | Escherichia phage phiEco32 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77554 | 42.274 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| EF153632 | Burkholderia phage BcepF1 | Burkholderia phage BcepF1 Bcepfunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72415 | 55.893 | Bcepfunavirus | Unclassified | Myoviridae | Burkholderia |
| EF056009 | Escherichia phage N4 | Escherichia phage N4 Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70153 | 41.304 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| DQ121662 | Escherichia phage Jk06 | Escherichia phage Jk06 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46072 | 44.001 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| AY967407 | Escherichia phage RB43 | Escherichia phage RB43 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180500 | 43.199 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| AY216660 | Escherichia phage T1 | Escherichia phage T1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48836 | 45.552 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| AF234172 | Escherichia phage P1 | Escherichia phage P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94800 | 47.311 | Punavirus | Unclassified | Myoviridae | Escherichia |
| AF069529 | Escherichia phage HK97 | Escherichia phage HK97 Byrnievirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39732 | 49.786 | Byrnievirus | Hendrixvirinae | Siphoviridae | Escherichia |
| AF069308 | Escherichia phage HK022 | Escherichia phage HK022 Shamshuipovirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40751 | 49.483 | Shamshuipovirus | Hendrixvirinae | Siphoviridae | Escherichia |
| AF064539 | Escherichia phage N15 | Escherichia phage N15 Ravinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46375 | 51.172 | Ravinvirus | Unclassified | Siphoviridae | Escherichia |
| PX1CG | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| CP067347 | Clostridioides phage ES-S-0107-01 | Clostridioides phage ES-S-0107-01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132924 | 26.159 | Unclassified | Unclassified | Siphoviridae | Clostridioides |
| CP067352 | Clostridioides phage ES-S-0173-01 | Clostridioides phage ES-S-0173-01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113019 | 26.487 | Unclassified | Unclassified | Siphoviridae | Clostridioides |
| OK318961 | Microbacterium phage Saratos | Microbacterium phage Saratos unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40350 | 61.988 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| OK318960 | Mycobacterium phage LestyG | Mycobacterium phage LestyG unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68198 | 67.569 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| OK318959 | Mycobacterium phage Colbster | Mycobacterium phage Colbster unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50884 | 63.995 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| OK318958 | Microbacterium phage IndyLu | Microbacterium phage IndyLu unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41958 | 66.235 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| OK318957 | Microbacterium phage Neptune | Microbacterium phage Neptune unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56694 | 63.83 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| OK539620 | Escherichia phage NTEC3 | Escherichia phage NTEC3 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44240 | 50.958 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN545971 | Citrobacter phage HCF1 | Citrobacter phage HCF1 Citrobacter virus HCF1 Hicfunavirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45806 | 44.509 | Hicfunavirus | Unclassified | Drexlerviridae | Citrobacter |
| MH375600 | Streptococcus phage pST | Streptococcus phage pST unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35581 | 38.546 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375599 | Streptococcus phage 142 | Streptococcus phage 142 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34639 | 39.014 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375598 | Streptococcus phage 140 | Streptococcus phage 140 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36064 | 38.285 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375597 | Streptococcus phage 123 | Streptococcus phage 123 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36129 | 38.67 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375595 | Streptococcus phage 107 | Streptococcus phage 107 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36938 | 38.678 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375594 | Streptococcus phage 94 | Streptococcus phage 94 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37611 | 38.135 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH375593 | Streptococcus phage 93 | Streptococcus phage 93 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40563 | 39.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH375596 | Streptococcus phage 115 | Streptococcus phage 115 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42017 | 38.796 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_031091 | Pseudomonas phage MD8 | Pseudomonas phage MD8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43277 | 61.134 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KU760857 | Salmonella phage SJ46 | Salmonella phage SJ46 Salmonella virus SJ46 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103445 | 48.578 | Punavirus | Unclassified | Myoviridae | Salmonella |
| KM236241 | Citrobacter phage Stevie | Citrobacter phage Stevie Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49816 | 42.838 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| JN132397 | Campylobacter phage CPX | Campylobacter phage CPX Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132662 | 26.043 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| GU296433 | Campylobacter phage NCTC12673 | Campylobacter phage NCTC12673 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135041 | 26.154 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| DQ289555 | Bacillus phage WBeta | Bacillus phage WBeta Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40867 | 35.258 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| AY029185 | Bordetella phage BPP-1 | Bordetella phage BPP-1 Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42493 | 65.408 | Rauchvirus | Unclassified | Podoviridae | Bordetella |
| MZ727334 | Pseudomonas phage PPAT | Pseudomonas phage PPAT unclassified bacterial viruses Viruses | 15211 | 44.573 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| MZ727333 | Pseudomonas phage PPAT | Pseudomonas phage PPAT unclassified bacterial viruses Viruses | 22812 | 43.718 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| MZ727332 | Pseudomonas phage PPAT | Pseudomonas phage PPAT unclassified bacterial viruses Viruses | 34072 | 51.027 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| OK483201 | Vibrio phage F23s1 | Vibrio phage F23s1 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76648 | 48.818 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MN364664 | Xanthomonas phage Xoo-sp15 | Xanthomonas phage Xoo-sp15 unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157091 | 39.86 | Caeruleovirus | Bastillevirinae | Herelleviridae | Xanthomonas |
| LC628082 | Streptococcus phage SG005 | Streptococcus phage SG005 Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16127 | 34.52 | Unclassified | Unclassified | Rountreeviridae | Streptococcus |
| MZ643249 | Proteus phage vB_PmiS_PM-CJR | Proteus phage vB_PmiS_PM-CJR unclassified Gorganvirus Gorganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54169 | 35.982 | Gorganvirus | Unclassified | Siphoviridae | Proteus |
| MT757392 | Klebsiella phage vB_KleM_KB2 | Klebsiella phage vB_KleM_KB2 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48245 | 48.888 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| NC_011285 | Mycobacterium phage Troll4 | Mycobacterium phage Troll4 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64618 | 59.612 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| CP045566 | Liberibacter phage CLasMV1 | Liberibacter phage CLasMV1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 8869 | 36.791 | Unclassified | Unclassified | Microviridae | Liberibacter |
| OK042081 | Yersinia phage vB_YenS_P400 | Yersinia phage vB_YenS_P400 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46585 | 43.744 | Unclassified | Unclassified | Siphoviridae | Yersinia |
| OK042080 | Yersinia phage vB_YenP_Rambo | Yersinia phage vB_YenP_Rambo Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40327 | 46.827 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| MZ209302 | Gordonia phage Jojo24 | Gordonia phage Jojo24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40579 | 67.542 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| OK283619 | Escherichia phage LPEK22 | Escherichia phage LPEK22 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158981 | 44.592 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| OK283618 | Listeria phage LPML1 | Listeria phage LPML1 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87038 | 38.707 | Unclassified | Ounavirinae | Myoviridae | Listeria |
| OK149216 | Acinetobacter phage vB_IME_Ab2712 | Acinetobacter phage vB_IME_Ab2712 unclassified Presleyvirus Presleyvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74579 | 43.302 | Presleyvirus | Unclassified | Schitoviridae | Acinetobacter |
| OK149215 | Klebsiella phage vB_KpnP_IME305 | Klebsiella phage vB_KpnP_IME305 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38641 | 50.943 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MZ648215 | Escherichia phage vB_EcoP_IMEP24 | Escherichia phage vB_EcoP_IMEP24 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39577 | 49.938 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ648214 | Escherichia phage vB_EcoP_IMEP8 | Escherichia phage vB_EcoP_IMEP8 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75809 | 42.103 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MZ997841 | Escherichia phage ULINTec7 | Escherichia phage ULINTec7 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46142 | 45.076 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ997840 | Escherichia phage ULINTec6 | Escherichia phage ULINTec6 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44728 | 45.07 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ997839 | Escherichia phage ULINTec4 | Escherichia phage ULINTec4 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45159 | 44.609 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| OK181169 | Vibrio phage 536 | Vibrio phage 536 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12676 | 41.464 | Unclassified | Unclassified | Podoviridae | Vibrio |
| OK274245 | Ralstonia phage RpT1 | Ralstonia phage RpT1 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40788 | 62.651 | Unclassified | Unclassified | Autographiviridae | Ralstonia |
| OK322699 | Escherichia phage Sa157lw | Escherichia phage Sa157lw unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155887 | 44.944 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| OK318991 | Ralstonia phage RpY2 | Ralstonia phage RpY2 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40147 | 62.65 | Unclassified | Unclassified | Autographiviridae | Ralstonia |
| OK319016 | Brevundimonas phage AA | Brevundimonas phage AA unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41678 | 61.795 | Unclassified | Okabevirinae | Autographiviridae | Brevundimonas |
| OK335783 | Brevundimonas phage BC | Brevundimonas phage BC unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41678 | 61.793 | Unclassified | Okabevirinae | Autographiviridae | Brevundimonas |
| OK335775 | Escherichia phage PSD2002 | Escherichia phage PSD2002 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172439 | 39.459 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ997838 | Escherichia phage ULINTec2 | Escherichia phage ULINTec2 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40815 | 51.114 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| OK143207 | Escherichia phage Lindwurm | Escherichia phage Lindwurm unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110040 | 39.052 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| OK095358 | Burkholderia phage phiBt-TXDOH | Burkholderia phage phiBt-TXDOH unclassified Stanholtvirus Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56453 | 60.3 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| OK172329 | Alcaligenes phage vB_Af_QDWS535 | Alcaligenes phage vB_Af_QDWS535 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50121 | 47.962 | Unclassified | Unclassified | Unclassified | Alcaligenes |
| OK076929 | Escherichia phage vB_EcoM_C2-3 | Escherichia phage vB_EcoM_C2-3 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167069 | 35.336 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| OK040170 | Serratia phage BUCT660 | Serratia phage BUCT660 unclassified Moabitevirus Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 272720 | 46.763 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| OK019720 | Klebsiella phage KL3 | Klebsiella phage KL3 unclassified Drexlerviridae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52742 | 49.226 | Unclassified | Unclassified | Drexlerviridae | Klebsiella |
| MZ969012 | Mycobacterium phage Saroj | Mycobacterium phage Saroj Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54958 | 61.381 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MT956952 | Lactobacillus phage MLC-A | Lactobacillus phage MLC-A unclassified Ashduovirus Ashduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41511 | 44.706 | Ashduovirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_048734 | Mycobacterium phage Fowlmouth | Mycobacterium phage Fowlmouth Mycobacterium virus Fowlmouth Fowlmouthvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69660 | 48.666 | Fowlmouthvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048696 | Mycobacterium phage Cuke | Mycobacterium phage Cuke Mycobacterium virus Cuke Cukevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68869 | 49.08 | Cukevirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029071 | Salmonella phage 36 | Salmonella phage 36 Salmonella virus 36 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41085 | 43.522 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_011318 | Clostridium phage phiCP39-O | Clostridium phage phiCP39-O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38753 | 30.421 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_049456 | Pseudomonas phage Zuri | Pseudomonas phage Zuri Pseudomonas virus Zuri Zurivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75873 | 53.528 | Zurivirus | Unclassified | Schitoviridae | Pseudomonas |
| NC_042351 | Salinibacter virus M8CRM1 | Salinibacter virus M8CRM1 Kryptosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53197 | 53.381 | Kryptosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_019423 | Escherichia phage IME11 | Escherichia phage IME11 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72570 | 43.139 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MW450817 | Halorubrum virus VOLN27B | Halorubrum virus VOLN27B unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76891 | 56.157 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| NC_052967 | Xanthomonas phage Bosa | Xanthomonas phage Bosa unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63828 | 64.837 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_052966 | Xanthomonas phage Xoo-sp2 | Xanthomonas phage Xoo-sp2 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60497 | 66.532 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_048742 | Streptomyces phage Gilson | Streptomyces phage Gilson Streptomyces virus Gilson Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128338 | 47.439 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047879 | Bordetella phage MW2 | Bordetella phage MW2 Bordetella virus MW2 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60160 | 64.058 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| NC_047878 | Bordetella phage FP1 | Bordetella phage FP1 Bordetella virus FP1 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60332 | 64.182 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| NC_047877 | Bordetella phage CN2 | Bordetella phage CN2 Bordetella virus CN2 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62030 | 64.034 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| NC_047876 | Bordetella phage CN1 | Bordetella phage CN1 Bordetella virus CN1 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59779 | 63.974 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| MN857617 | Bacillus phage SRT01hs | Bacillus phage SRT01hs Bacillus virus SRT01hs Gaunavirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20784 | 34.945 | Gaunavirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| NC_041958 | Roseobacter phage RDJL Phi 2 | Roseobacter phage RDJL Phi 2 Xiamenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63513 | 57.295 | Xiamenvirus | Unclassified | Siphoviridae | Roseobacter |
| MF360957 | Bacillus phage PBS1 | Bacillus phage PBS1 Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252197 | 27.71 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| NC_029024 | Bacillus phage 250 | Bacillus phage 250 Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56505 | 36.441 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| NC_028980 | Pseudomonas phage PAE1 | Pseudomonas phage PAE1 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62181 | 64.233 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_027397 | Vibrio phage QH | Vibrio phage QH unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39725 | 50.505 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| NC_027393 | Vibrio phage J2 | Vibrio phage J2 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39530 | 50.582 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| NC_027118 | Vibrio phage phiVC8 | Vibrio phage phiVC8 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39422 | 50.807 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| NC_026594 | Pseudomonas phage vB_PaeS_PAO1_Ab18 | Pseudomonas phage vB_PaeS_PAO1_Ab18 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56537 | 63.498 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_022773 | Bacillus phage Blastoid | Bacillus phage Blastoid Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50354 | 42.233 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KF669659 | Bacillus phage Riggi | Bacillus phage Riggi Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49836 | 41.46 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KF669651 | Bacillus phage Glittering | Bacillus phage Glittering Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49246 | 42.054 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_020479 | Bacillus phage Curly | Bacillus phage Curly Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49425 | 41.821 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_020478 | Bacillus phage Andromeda | Bacillus phage Andromeda Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49259 | 41.907 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_020477 | Bacillus phage Eoghan | Bacillus phage Eoghan Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49458 | 42.208 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KC330683 | Bacillus phage Finn | Bacillus phage Finn Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50161 | 41.69 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KC330682 | Bacillus phage Taylor | Bacillus phage Taylor Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49492 | 42.294 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| JX887877 | Bacillus phage vB_BtS_BMBtp2 | Bacillus phage vB_BtS_BMBtp2 Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36932 | 37.788 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| NC_018452 | Burkholderia phage DC1 | Burkholderia phage DC1 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61847 | 66.212 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| NC_018282 | Pseudomonas phage MP1412 | Pseudomonas phage MP1412 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61167 | 64.283 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| JN638751 | Bacillus phage G | Bacillus phage G Donellivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 497513 | 29.926 | Donellivirus | Unclassified | Myoviridae | Bacillus |
| NC_015466 | Roseobacter phage RDJL Phi 1 | Roseobacter phage RDJL Phi 1 Xiamenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62668 | 57.881 | Xiamenvirus | Unclassified | Siphoviridae | Roseobacter |
| FJ230960 | Bacillus phage SPO1 | Bacillus phage SPO1 Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132562 | 39.965 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| EU874396 | Bacillus phage IEBH | Bacillus phage IEBH Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53104 | 36.423 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| EU771092 | Bacillus phage phi29 | Bacillus phage phi29 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19282 | 39.991 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_009604 | Burkholderia phage BcepNY3 | Burkholderia phage BcepNY3 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47382 | 63.64 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| NC_009235 | Burkholderia phage phi644-2 | Burkholderia phage phi644-2 Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48674 | 60.449 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| NC_009236 | Burkholderia phage phiE12-2 | Burkholderia phage phiE12-2 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36690 | 64.617 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_007024 | Xanthomonas phage Xp15 | Xanthomonas phage Xp15 Xanthomonas virus Xp15 Alachuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55770 | 51.888 | Alachuavirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_005891 | Vibrio phage VP5 | Vibrio phage VP5 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39786 | 50.485 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| NC_005879 | Vibrio phage VP2 | Vibrio phage VP2 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39853 | 50.558 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| NC_055956 | Klebsiella phage RAD2 | Klebsiella phage RAD2 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49276 | 50.388 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_052978 | Proteus phage Saba | Proteus phage Saba Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60056 | 48.818 | Unclassified | Unclassified | Siphoviridae | Proteus |
| NC_052974 | Achromobacter phage vB_AchrS_AchV4 | Achromobacter phage vB_AchrS_AchV4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59489 | 62.83 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| OK040806 | Escherichia phage CL1 | Escherichia phage CL1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87817 | 39.024 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_055920 | Gordonia phage Phlop | Gordonia phage Phlop unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58358 | 67.944 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055918 | Mycobacterium phage OhShagHennessy | Mycobacterium phage OhShagHennessy unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72623 | 58.875 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055916 | Staphylococcus phage LSA2366 | Staphylococcus phage LSA2366 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17056 | 29.397 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_055914 | Bacillus phage Thornton | Bacillus phage Thornton unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26319 | 30.487 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_055912 | Gordonia phage Blino | Gordonia phage Blino Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54168 | 66.903 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055911 | Burkholderia phage phiE094 | Burkholderia phage phiE094 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37727 | 64.484 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_055909 | Mycobacterium phage Plumbus | Mycobacterium phage Plumbus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54468 | 61.133 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055908 | Bacillus phage DLc1 | Bacillus phage DLc1 Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28950 | 31.095 | Unclassified | Northropvirinae | Salasmaviridae | Bacillus |
| NC_055906 | Staphylococcus phage vB_SauP_EBHT | Staphylococcus phage vB_SauP_EBHT unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17471 | 29.569 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_055905 | Bacillus phage Baseball_field | Bacillus phage Baseball_field unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26863 | 30.436 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_055900 | crAssphage cr125_1 | crAssphage cr125_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98095 | 32.751 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055898 | crAssphage cr115_1 | crAssphage cr115_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99848 | 32.553 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055897 | crAssphage cr114_1 | crAssphage cr114_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98308 | 32.79 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055894 | crAssphage cr6_1 | crAssphage cr6_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97422 | 30.93 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055893 | crAssphage cr4_1 | crAssphage cr4_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96055 | 33.495 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055890 | crAssphage cr56_1 | crAssphage cr56_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97290 | 33.008 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055888 | crAssphage cr52_1 | crAssphage cr52_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103665 | 34.658 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055867 | Gordonia phage Clown | Gordonia phage Clown unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58203 | 65.741 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055865 | Gordonia phage Azula | Gordonia phage Azula Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53622 | 67.092 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055864 | Gordonia phage Love | Gordonia phage Love unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60042 | 67.268 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055863 | Burkholderia phage Mana | Burkholderia phage Mana Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38038 | 64.315 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_055862 | Vibrio phage ValB1MD-2 | Vibrio phage ValB1MD-2 unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36616 | 43.224 | Unclassified | Peduovirinae | Myoviridae | Vibrio |
| NC_055861 | Gordonia phage Portcullis | Gordonia phage Portcullis unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58364 | 67.759 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055859 | Gordonia phage Paries | Gordonia phage Paries unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60127 | 67.354 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055858 | Mycobacterium phage Gail | Mycobacterium phage Gail unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53702 | 62.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055857 | Gordonia phage Jambalaya | Gordonia phage Jambalaya unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57764 | 67.816 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055846 | Yersinia phage vB_YpM_50 | Yersinia phage vB_YpM_50 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31096 | 52.126 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| NC_055845 | Yersinia phage vB_YpM_46 | Yersinia phage vB_YpM_46 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32336 | 51.661 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| NC_055844 | Yersinia phage vB_YpM_22 | Yersinia phage vB_YpM_22 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31809 | 51.467 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| NC_055843 | Gordonia phage Evamon | Gordonia phage Evamon unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58460 | 67.805 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055842 | Streptomyces phage Bmoc | Streptomyces phage Bmoc unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132885 | 49.639 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_055838 | Xanthomonas phage FoX5 | Xanthomonas phage FoX5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43872 | 59.466 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| NC_055835 | Xanthomonas phage FoX1 | Xanthomonas phage FoX1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44763 | 59.429 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| NC_055831 | Phage NBEco003 | Phage NBEco003 Escherichia virus NBEco003 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166579 | 35.515 | Tequatrovirus | Tevenvirinae | Myoviridae | Unspecified |
| NC_055722 | Burkholderia phage FLC5 | Burkholderia phage FLC5 unclassified Kisquattuordecimvirus Kisquattuordecimvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32090 | 61.792 | Kisquattuordecimvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_055040 | Staphylococcus phage vB_SauS_JS02 | Staphylococcus phage vB_SauS_JS02 Staphylococcus virus JS02 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46435 | 33.143 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055026 | Ralstonia phage Dina | Ralstonia phage Dina Ralstonia virus Dina Dinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39161 | 65.274 | Dinavirus | Unclassified | Siphoviridae | Ralstonia |
| NC_054981 | Staphylococcus phage vB_SauS-SAP27 | Staphylococcus phage vB_SauS-SAP27 Staphylococcus virus SAP27 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43444 | 34.946 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054965 | Klebsiella phage LASTA | Klebsiella phage LASTA Klebsiella virus LASTA Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62420 | 56.237 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| NC_054961 | Ralstonia phage Firinga | Ralstonia phage Firinga Ralstonia virus Firinga Firingavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39744 | 59.692 | Firingavirus | Unclassified | Podoviridae | Ralstonia |
| NC_054955 | Ralstonia phage Raharianne | Ralstonia phage Raharianne Ralstonia virus Raharianne Rahariannevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46321 | 60.547 | Rahariannevirus | Unclassified | Myoviridae | Ralstonia |
| NC_054938 | Serratia phage PhiZZ30 | Serratia phage PhiZZ30 Serratia virus PhiZZ30 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167484 | 35.345 | Tequatrovirus | Tevenvirinae | Myoviridae | Serratia |
| NC_054934 | Escherichia phage teqskov | Escherichia phage teqskov Escherichia virus teqskov Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165017 | 35.359 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054933 | Escherichia phage teqhad | Escherichia phage teqhad Escherichia virus teqhad Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167892 | 35.327 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054932 | Escherichia phage teqdroes | Escherichia phage teqdroes Escherichia virus teqdroes Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166833 | 35.438 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054927 | Escherichia phage vB_EcoM_Ozark | Escherichia phage vB_EcoM_Ozark Escherichia virus Ozark Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167600 | 35.404 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054924 | Escherichia phage vB_EcoM_Lutter | Escherichia phage vB_EcoM_Lutter Escherichia virus Lutter Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170054 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054921 | Escherichia phage vB_EcoM_IME537 | Escherichia phage vB_EcoM_IME537 Escherichia virus IME537 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168642 | 35.425 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054913 | Escherichia phage F2 | Escherichia phage F2 Escherichia virus F2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168241 | 37.324 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054912 | Escherichia phage vB_EcoM_F1 | Escherichia phage vB_EcoM_F1 Escherichia virus EcoMF1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168410 | 35.355 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054901 | Citrobacter phage PhiZZ23 | Citrobacter phage PhiZZ23 Citrobacter virus PhiZZ23 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167516 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_054900 | Citrobacter phage PhiZZ6 | Citrobacter phage PhiZZ6 Citrobacter virus PhiZZ6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165795 | 35.432 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_054896 | Escherichia phage tiwna | Escherichia phage tiwna Escherichia virus tiwna Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51014 | 44.631 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_054891 | Ralstonia phage Anchaing | Ralstonia phage Anchaing Ralstonia virus Anchaing Anchaingvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43597 | 63.087 | Anchaingvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_054890 | Pseudomonas phage vB_PsyP_3MF5 | Pseudomonas phage vB_PsyP_3MF5 Pseudomonas virus 3MF5 Warsawvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41608 | 56.528 | Warsawvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_054810 | Mycobacterium phage Georgie2 | Mycobacterium phage Georgie2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53399 | 63.526 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054789 | Mycobacterium phage DroogsArmy | Mycobacterium phage DroogsArmy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53254 | 63.015 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054771 | Mycobacterium phage MA5 | Mycobacterium phage MA5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47895 | 64.092 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054679 | Streptomyces phage Werner | Streptomyces phage Werner unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51566 | 65.698 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054676 | Streptomyces phage Sitrop | Streptomyces phage Sitrop unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48481 | 65.597 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054673 | Streptomyces phage Endor2 | Streptomyces phage Endor2 unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48829 | 65.08 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054672 | Streptomyces phage Endor1 | Streptomyces phage Endor1 unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49395 | 65.79 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054667 | Streptomyces phage Issmi | Streptomyces phage Issmi unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50643 | 67.565 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054659 | Streptomyces phage Spernnie | Streptomyces phage Spernnie unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50834 | 65.694 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054658 | Streptomyces phage Sentinel | Streptomyces phage Sentinel unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50272 | 66.862 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054657 | Streptomyces phage Salutena | Streptomyces phage Salutena unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51956 | 67.501 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054654 | Klebsiella phage vB_KpnS_ZX4 | Klebsiella phage vB_KpnS_ZX4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45424 | 48.371 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| NC_054653 | Klebsiella virus KpV2811 | Klebsiella virus KpV2811 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46391 | 48.208 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| NC_054649 | Escherichia phage vB_EcoS_swi2 | Escherichia phage vB_EcoS_swi2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47611 | 46.254 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| NC_054648 | Salmonella phage vB_Se_STGO-35-1 | Salmonella phage vB_Se_STGO-35-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47483 | 46.488 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054456 | Mycobacterium phage Jamie19 | Mycobacterium phage Jamie19 unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41278 | 66.244 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054454 | Mycobacterium phage Rebel | Mycobacterium phage Rebel unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40580 | 66.518 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054448 | Mycobacterium phage Raymond7 | Mycobacterium phage Raymond7 unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42382 | 65.964 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054432 | Mycobacterium phage phiT46-1 | Mycobacterium phage phiT46-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52849 | 63.708 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054149 | Pseudomonas phage Persinger | Pseudomonas phage Persinger Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44033 | 62.86 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| NC_054148 | Burkholderia phage Mica | Burkholderia phage Mica Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43707 | 62.153 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| NC_054147 | Achromobacter phage Mano | Achromobacter phage Mano Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42452 | 64.303 | Unclassified | Unclassified | Myoviridae | Achromobacter |
| NC_053516 | Mycobacterium phage Reindeer | Mycobacterium phage Reindeer unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82356 | 60.315 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_053514 | Mycobacterium phage Estes | Mycobacterium phage Estes unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83601 | 60.866 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_053513 | Mycobacterium phage Aziz | Mycobacterium phage Aziz unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83412 | 60.749 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_053506 | Mycobacterium phage CELFI | Mycobacterium phage CELFI unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77086 | 59.033 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053209 | Mycobacterium phage KilKor | Mycobacterium phage KilKor unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48916 | 67.209 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_053172 | Gordonia phage DumpsterDude | Gordonia phage DumpsterDude Gordonia virus DumpsterDude Ruthyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52018 | 66.61 | Ruthyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_053010 | Providencia phage vB_PreS-Stilesk | Providencia phage vB_PreS-Stilesk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60924 | 49.47 | Unclassified | Unclassified | Siphoviridae | Providencia |
| NC_053009 | Providencia phage vB_PreS-PibeRecoleta | Providencia phage vB_PreS-PibeRecoleta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60727 | 49.27 | Unclassified | Unclassified | Siphoviridae | Providencia |
| NC_053008 | Salmonella phage SAP012 | Salmonella phage SAP012 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59618 | 54.056 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_053007 | Salmonella phage vB_SenS_ER6 | Salmonella phage vB_SenS_ER6 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58872 | 56.405 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_053006 | Salmonella phage vB_SenS_ER3 | Salmonella phage vB_SenS_ER3 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58971 | 56.385 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_053003 | Salmonella phage vB_SalS_ABTNLsp1 | Salmonella phage vB_SalS_ABTNLsp1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58922 | 56.324 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052999 | Salmonella phage vB_SenS_ER24 | Salmonella phage vB_SenS_ER24 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60438 | 56.58 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052996 | Salmonella phage vB_SenS_BPS4 | Salmonella phage vB_SenS_BPS4 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58529 | 56.623 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052995 | Salmonella phage vB_SenS_BPS2 | Salmonella phage vB_SenS_BPS2 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58526 | 56.623 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052994 | Salmonella phage vB_SenS_BPS1 | Salmonella phage vB_SenS_BPS1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58852 | 56.591 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052993 | Salmonella phage vB_SenS_ER19 | Salmonella phage vB_SenS_ER19 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58344 | 56.585 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052991 | Serratia phage KpYy_1_41 | Serratia phage KpYy_1_41 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54417 | 56.899 | Chivirus | Unclassified | Siphoviridae | Serratia |
| NC_052986 | Aeromonas phage BUCT551 | Aeromonas phage BUCT551 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61382 | 61.71 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| NC_052467 | Mycobacterium phage STLscum | Mycobacterium phage STLscum unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52265 | 63.674 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052448 | Mycobacterium phage Mule | Mycobacterium phage Mule unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51468 | 63.583 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052444 | Mycobacterium phage MaryBeth | Mycobacterium phage MaryBeth unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51854 | 63.534 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052442 | Mycobacterium phage Manatee | Mycobacterium phage Manatee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51040 | 63.791 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052436 | Mycobacterium phage Jorgensen | Mycobacterium phage Jorgensen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53771 | 63.638 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052432 | Mycobacterium phage HarryOW | Mycobacterium phage HarryOW unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52935 | 63.774 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051730 | Mycobacterium phage Jung | Mycobacterium phage Jung unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46561 | 67.078 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051727 | Mycobacterium phage Willsammy | Mycobacterium phage Willsammy unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48399 | 67.026 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051723 | Mycobacterium phage Soul22 | Mycobacterium phage Soul22 unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54814 | 61.501 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051716 | Mycobacterium phage Veteran | Mycobacterium phage Veteran unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56147 | 61.499 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051708 | Mycobacterium phage Seabastian | Mycobacterium phage Seabastian unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58618 | 61.096 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051705 | Mycobacterium phage Royals2015 | Mycobacterium phage Royals2015 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57356 | 61.708 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051688 | Mycobacterium phage Mova | Mycobacterium phage Mova unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57599 | 61.253 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051687 | Mycobacterium phage Moonbeam | Mycobacterium phage Moonbeam unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57984 | 61.1 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051677 | Mycobacterium phage LastJedi | Mycobacterium phage LastJedi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55149 | 61.755 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051668 | Mycobacterium phage Hlubikazi | Mycobacterium phage Hlubikazi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55471 | 61.439 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051662 | Mycobacterium phage Gandalph | Mycobacterium phage Gandalph unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54974 | 61.369 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051658 | Mycobacterium phage Firehouse51 | Mycobacterium phage Firehouse51 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58166 | 61.476 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051650 | Mycobacterium phage DRBy19 | Mycobacterium phage DRBy19 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58368 | 61.292 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051644 | Mycobacterium phage Coco12 | Mycobacterium phage Coco12 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57693 | 61.136 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051637 | Mycobacterium phage BodEinwohner17 | Mycobacterium phage BodEinwohner17 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60222 | 61.343 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_050151 | Pseudomonas phage crassa | Pseudomonas phage crassa unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66295 | 55.218 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050150 | Pseudomonas phage antinowhere | Pseudomonas phage antinowhere unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65852 | 55.189 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050149 | Pseudomonas phage vB_PaeM_USP_1 | Pseudomonas phage vB_PaeM_USP_1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65918 | 55.578 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050147 | Pseudomonas phage Epa13 | Pseudomonas phage Epa13 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65680 | 55.504 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050146 | Pseudomonas phage Epa7 | Pseudomonas phage Epa7 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65629 | 55.457 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050144 | Pseudomonas phage Epa14 | Pseudomonas phage Epa14 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65797 | 55.282 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050143 | Pseudomonas phage datas | Pseudomonas phage datas unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60746 | 54.807 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_049974 | Bacillus phage Gxv1 | Bacillus phage Gxv1 Bacillus virus Gxv1 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21781 | 39.686 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_049955 | Escherichia phage Lambda_ev243 | Escherichia phage Lambda_ev243 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45560 | 50.316 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049953 | Escherichia phage Lambda_ev099 | Escherichia phage Lambda_ev099 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47224 | 50.43 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049949 | Escherichia phage Lambda_ev207 | Escherichia phage Lambda_ev207 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46685 | 50.288 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049948 | Escherichia phage Lambda_ev017 | Escherichia phage Lambda_ev017 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50126 | 49.689 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049900 | Listeria phage LP-HM00113468 | Listeria phage LP-HM00113468 Listeria virus LPHM00113468 Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40298 | 35.488 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| NC_049852 | Escherichia phage egaa | Escherichia phage egaa Escherichia virus egaa Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51643 | 44.128 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049832 | Escherichia phage vB_EcoS-DELF2 | Escherichia phage vB_EcoS-DELF2 Escherichia virus DELF2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50168 | 45.505 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_049830 | Escherichia phage PGN590 | Escherichia phage PGN590 Escherichia virus PGN590 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49043 | 43.806 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049829 | Escherichia phage aalborv | Escherichia phage aalborv Escherichia virus aalborv Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46660 | 43.89 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049828 | Escherichia phage haarsle | Escherichia phage haarsle Escherichia virus haarsle Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48613 | 44.038 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049827 | Escherichia phage damhaus | Escherichia phage damhaus Escherichia virus damhaus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51154 | 44.145 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049826 | Escherichia phage aaroes | Escherichia phage aaroes Escherichia virus aaroes Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51662 | 44.125 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049825 | Escherichia phage grams | Escherichia phage grams Escherichia virus grams Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49530 | 44.099 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049824 | Escherichia phage vojen | Escherichia phage vojen Escherichia virus vojen Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50709 | 44.13 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049823 | Escherichia phage herni | Escherichia phage herni Escherichia virus herni Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50971 | 44.088 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049821 | Salmonella phage slyngel | Salmonella phage slyngel Salmonella virus slyngel Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51048 | 44.012 | Hanrivervirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_049820 | Shigella virus 2019SD1 | Shigella virus 2019SD1 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53145 | 44.512 | Hanrivervirus | Tempevirinae | Drexlerviridae | Shigella |
| NC_049819 | Escherichia phage atuna | Escherichia phage atuna Escherichia virus atuna Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50732 | 44.599 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049818 | Escherichia phage tonijn | Escherichia phage tonijn Escherichia virus tonijn Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51627 | 44.637 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049817 | Escherichia phage tonnikala | Escherichia phage tonnikala Escherichia virus tonnikala Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 44.788 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049816 | Escherichia phage tunus | Escherichia phage tunus Escherichia virus tunus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51111 | 44.799 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049815 | Escherichia phage tonn | Escherichia phage tonn Escherichia virus tonn Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51012 | 44.472 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049395 | Escherichia phage ESSI2_ev015 | Escherichia phage ESSI2_ev015 Escherichia virus ev015 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30584 | 50.876 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049344 | Escherichia phage 520873 | Escherichia phage 520873 Escherichia virus 520873 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37583 | 52.005 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049343 | Escherichia phage 500465-2 | Escherichia phage 500465-2 Escherichia virus 500465-2 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33041 | 52.441 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049342 | Escherichia phage 500465-1 | Escherichia phage 500465-1 Escherichia virus 500465-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39179 | 52.651 | Felsduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049341 | Escherichia phage 503458 | Escherichia phage 503458 Escherichia virus 503458 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31194 | 53.446 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_048876 | Gordonia phage Secretariat | Gordonia phage Secretariat Gordonia virus Secretariat Secretariatvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57731 | 52.816 | Secretariatvirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048873 | Klebsiella virus KpS8 | Klebsiella virus KpS8 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143800 | 44.583 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| NC_048864 | Salmonella phage birk | Salmonella phage birk Salmonella virus birk Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52767 | 45.845 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| NC_048863 | Salmonella phage yarpen | Salmonella phage yarpen Salmonella virus yarpen Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52464 | 45.961 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| NC_048850 | Mycobacterium phage Cornie | Mycobacterium phage Cornie Mycobacterium virus Cornie Cornievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56391 | 61.386 | Cornievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048849 | Enterobacter phage vB_EclM_CIP9 | Enterobacter phage vB_EclM_CIP9 Enterobacter virus CIP9 Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174924 | 39.934 | Kanagawavirus | Unclassified | Myoviridae | Enterobacter |
| NC_048848 | Butyrivibrio virus Arawn | Butyrivibrio virus Arawn Arawnvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31118 | 46.944 | Arawnvirus | Unclassified | Siphoviridae | Butyrivibrio |
| NC_048666 | Salmonella phage YSD1 | Salmonella phage YSD1 Salmonella virus YSD1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58916 | 56.531 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_041878 | Pectinobacterium phage vB_PcaM_CBB | Pectinobacterium phage vB_PcaM_CBB Pectinobacterium virus CBB Mimasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 378379 | 35.919 | Mimasvirus | Unclassified | Myoviridae | Pectobacterium |
| KY608967 | Escherichia phage HP3 | Escherichia phage HP3 Escherichia virus HP3 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168188 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KY608966 | Escherichia phage CF2 | Escherichia phage CF2 Escherichia virus CF2 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53242 | 45.945 | Rosemountvirus | Unclassified | Myoviridae | Escherichia |
| AY349011 | Burkholderia phage Bcep22 | Burkholderia phage Bcep22 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63882 | 65.309 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| NC_015209 | Vibrio phage CTXphi | Vibrio phage CTXphi Affertcholeramvirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10638 | 44.623 | Affertcholeramvirus | Unclassified | Inoviridae | Vibrio |
| EU719189 | Clostridioides phage phiCD27 | Clostridioides phage phiCD27 Clostridioides virus phiCD27 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50930 | 29.327 | Colneyvirus | Unclassified | Myoviridae | Clostridioides |
| DQ466086 | Clostridioides phage phiC2 | Clostridioides phage phiC2 Clostridioides virus phiC2 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56538 | 28.726 | Yongloolinvirus | Unclassified | Myoviridae | Clostridioides |
| AY855346 | Clostridium phage phi CD119 | Clostridium phage phi CD119 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53325 | 28.773 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| AY605181 | Burkholderia phage BcepC6B | Burkholderia phage BcepC6B Ryyoungvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42415 | 65.194 | Ryyoungvirus | Unclassified | Podoviridae | Burkholderia |
| AY369265 | Burkholderia phage Bcep1 | Burkholderia phage Bcep1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48177 | 63.64 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| AY368235 | Burkholderia phage Bcep43 | Burkholderia phage Bcep43 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48024 | 63.435 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| AF543311 | Burkholderia phage Bcep781 | Burkholderia phage Bcep781 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48247 | 63.33 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| AF447491 | Burkholderia phage phiE125 | Burkholderia phage phiE125 Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53373 | 61.192 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| OU784750 | Escherichia virus P2 | Escherichia virus P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32541 | 51.572 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| OU784746 | Escherichia virus P2 | Escherichia virus P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31490 | 52.455 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| OU784549 | Escherichia virus P2 | Escherichia virus P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33562 | 51.082 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_056724 | Lactococcus phage BK5-T | Lactococcus phage BK5-T Lactococcus virus BK5T Sandinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40003 | 34.975 | Sandinevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_020414 | Escherichia phage UAB_Phi78 | Escherichia phage UAB_Phi78 Escherichia virus UAB78 Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43984 | 47.449 | Zindervirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_001900 | Mycobacterium phage D29 | Mycobacterium phage D29 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49127 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054935 | Escherichia phage YUEEL01 | Escherichia phage YUEEL01 Escherichia virus YUEEL01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168266 | 35.36 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055827 | Mycobacterium phage DirkDirk | Mycobacterium phage DirkDirk unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68021 | 58.673 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055825 | Mycobacterium phage LilMoolah | Mycobacterium phage LilMoolah unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58136 | 61.454 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055822 | Streptomyces phage Daubenski | Streptomyces phage Daubenski unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133090 | 49.397 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_055817 | Mycobacterium phage Jeeves | Mycobacterium phage Jeeves unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52466 | 62.116 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055816 | Gordonia phage Fireball | Gordonia phage Fireball unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58924 | 67.753 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055815 | Mycobacterium phage Xula | Mycobacterium phage Xula unclassified Brujitavirus Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48540 | 66.745 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| NC_055814 | Staphylococcus phage Portland | Staphylococcus phage Portland unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17711 | 29.293 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_055813 | Streptomyces phage Braelyn | Streptomyces phage Braelyn unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131234 | 50.052 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_055812 | Mycobacterium phage Obutu | Mycobacterium phage Obutu unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69021 | 67.429 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_055811 | Gordonia phage JuJu | Gordonia phage JuJu Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54000 | 66.904 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055810 | Gordonia phage Nubi | Gordonia phage Nubi unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58718 | 67.87 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055809 | Streptomyces phage Evy | Streptomyces phage Evy unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132977 | 49.568 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_055805 | Gordonia phage RogerDodger | Gordonia phage RogerDodger unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59122 | 67.858 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055804 | Gordonia phage Bakery | Gordonia phage Bakery unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60468 | 67.642 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055803 | Gordonia phage Galadriel | Gordonia phage Galadriel unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59400 | 67.446 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055802 | Staphylococcus phage SA46-CL1 | Staphylococcus phage SA46-CL1 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17508 | 28.81 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_055799 | Gordonia phage Valary | Gordonia phage Valary unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59448 | 67.787 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055798 | Gordonia phage Arri | Gordonia phage Arri unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58650 | 67.812 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055797 | Gordonia phage Barb | Gordonia phage Barb unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59514 | 67.831 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055796 | Gordonia phage VanDeWege | Gordonia phage VanDeWege unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58972 | 67.822 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055795 | Gordonia phage SmokingBunny | Gordonia phage SmokingBunny unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59454 | 67.906 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055793 | Pseudomonas phage vB_Pae_BR319a | Pseudomonas phage vB_Pae_BR319a unclassified Citexvirus Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35343 | 62.42 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| NC_055792 | Gordonia phage Walrus | Gordonia phage Walrus Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52024 | 67.3 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055791 | Streptomyces phage EGole | Streptomyces phage EGole unclassified Samistivirus Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135653 | 49.905 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_055790 | Mycobacterium phage Silverleaf | Mycobacterium phage Silverleaf unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73210 | 58.78 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055787 | Mycobacterium phage Ochi17 | Mycobacterium phage Ochi17 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58069 | 61.014 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055786 | Mycobacterium phage CicholasNage | Mycobacterium phage CicholasNage unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71059 | 58.612 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055785 | Gordonia phage Mutzi | Gordonia phage Mutzi unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59655 | 67.562 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055783 | Escherichia phage vB_EcoM_005 | Escherichia phage vB_EcoM_005 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155101 | 39.327 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055782 | Escherichia phage AnYang | Escherichia phage AnYang unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169215 | 39.494 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055781 | Escherichia phage p000v | Escherichia phage p000v unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167803 | 37.62 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055780 | Escherichia phage MLF4 | Escherichia phage MLF4 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167379 | 35.387 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055777 | Gordonia phage Nordenberg | Gordonia phage Nordenberg unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57572 | 67.253 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055768 | Klebsiella phage KP179 | Klebsiella phage KP179 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162630 | 39.346 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_055765 | Gordonia phage TillyBobJoe | Gordonia phage TillyBobJoe unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58677 | 67.761 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055762 | Gordonia phage Angelicage | Gordonia phage Angelicage unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59215 | 67.447 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055761 | Synechococcus virus S-PRM1 | Synechococcus virus S-PRM1 unclassified Kanaloavirus Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144311 | 40.743 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| NC_055757 | Escherichia virus KFS-EC | Escherichia virus KFS-EC unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164715 | 40.47 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055754 | Gordonia phage Frokostdame | Gordonia phage Frokostdame Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52531 | 66.919 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055752 | Gordonia phage KimmyK | Gordonia phage KimmyK unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58755 | 67.768 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055751 | Gordonia phage Danyall | Gordonia phage Danyall unclassified Wizardvirus Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58695 | 67.757 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055749 | Escherichia phage SF | Escherichia phage SF unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168695 | 37.614 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055748 | Klebsiella phage Mineola | Klebsiella phage Mineola unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166130 | 39.513 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_055745 | Gordonia phage Petra | Gordonia phage Petra Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52304 | 67.083 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_055744 | Synechococcus phage S-B64 | Synechococcus phage S-B64 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151867 | 41.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_055743 | Erwinia phage Cronus | Erwinia phage Cronus unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175774 | 38.378 | Unclassified | Tevenvirinae | Myoviridae | Erwinia |
| NC_055742 | Enterobacteria phage vB_EcoM_IME341 | Enterobacteria phage vB_EcoM_IME341 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172379 | 39.515 | Dhakavirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_055741 | Enterobacteria phage vB_EcoM_IME339 | Enterobacteria phage vB_EcoM_IME339 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164366 | 35.64 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_055740 | Enterobacteria phage vB_EcoM_IME281 | Enterobacteria phage vB_EcoM_IME281 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170531 | 39.442 | Dhakavirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_055739 | Enterobacter phage myPSH1140 | Enterobacter phage myPSH1140 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172614 | 39.883 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| NC_055737 | Proteus phage phiP4-3 | Proteus phage phiP4-3 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167849 | 31.507 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| NC_055734 | Shigella phage phi25-307 | Shigella phage phi25-307 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167544 | 37.55 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| NC_055728 | Serratia phage X20 | Serratia phage X20 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172450 | 38.655 | Winklervirus | Unclassified | Myoviridae | Serratia |
| NC_055727 | Proteus phage PM2 | Proteus phage PM2 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163469 | 31.157 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| NC_055726 | Cronobacter phage Pet-CM3-4 | Cronobacter phage Pet-CM3-4 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171975 | 39.806 | Karamvirus | Tevenvirinae | Myoviridae | Cronobacter |
| NC_055724 | Yersinia phage fPS-65 | Yersinia phage fPS-65 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167058 | 35.327 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_055723 | Yersinia phage fPS-90 | Yersinia phage fPS-90 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167132 | 35.384 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_055721 | Escherichia phage EcS1 | Escherichia phage EcS1 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175437 | 37.829 | Winklervirus | Unclassified | Myoviridae | Escherichia |
| NC_055720 | Pseudomonas phage PspYZU05 | Pseudomonas phage PspYZU05 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166442 | 35.479 | Unclassified | Tevenvirinae | Myoviridae | Pseudomonas |
| NC_055719 | Synechococcus phage S-H35 | Synechococcus phage S-H35 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174231 | 41.188 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_055718 | Escherichia phage phiC120 | Escherichia phage phiC120 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186570 | 37.626 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055715 | Klebsiella phage KPV15 | Klebsiella phage KPV15 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167034 | 39.522 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_055710 | Synechococcus phage S-CAM22 | Synechococcus phage S-CAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172230 | 39.919 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_055709 | Escherichia phage phiE142 | Escherichia phage phiE142 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121442 | 37.355 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055708 | Enterobacteria phage ATK47 | Enterobacteria phage ATK47 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170020 | 37.649 | Mosigvirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_055707 | Mycobacterium phage HC | Mycobacterium phage HC unclassified Brujitavirus Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46305 | 67.172 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| NC_055049 | Staphylococcus phage phiSa2wa_st121mssa | Staphylococcus phage phiSa2wa_st121mssa Staphylococcus virus st121mssa Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45621 | 33.057 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055048 | Staphylococcus phage phiSa2wa_st78 | Staphylococcus phage phiSa2wa_st78 Staphylococcus virus st78 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45878 | 33.216 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055047 | Staphylococcus phage phiSa2wa_st30 | Staphylococcus phage phiSa2wa_st30 Staphylococcus virus st30 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45780 | 33.438 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055046 | Staphylococcus phage phiSa2wa_st5 | Staphylococcus phage phiSa2wa_st5 Staphylococcus virus st5 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44823 | 33.407 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055045 | Staphylococcus phage phiSa2wa_st1 | Staphylococcus phage phiSa2wa_st1 Staphylococcus virus st1 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45585 | 33.281 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055044 | Staphylococcus phage SAP11 | Staphylococcus phage SAP11 Staphylococcus virus SAP11 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45346 | 33.419 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055043 | Staphylococcus phage SA137ruMSSAST121PVL | Staphylococcus phage SA137ruMSSAST121PVL Staphylococcus virus SA137ruMSSAST121PVL Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45999 | 33.305 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055042 | Staphylococcus phage P240 | Staphylococcus phage P240 Staphylococcus virus P240 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45985 | 33.111 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055041 | Staphylococcus phage LH1 | Staphylococcus phage LH1 Staphylococcus virus LH1 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46048 | 33.191 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055039 | Staphylococcus phage vB_SauS_fPfSau02 | Staphylococcus phage vB_SauS_fPfSau02 Staphylococcus virus fPfSau02 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45108 | 33.708 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055038 | Staphylococcus phage SH-St 15644 | Staphylococcus phage SH-St 15644 Staphylococcus virus 15644 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45111 | 33.349 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055037 | Leuconostoc phage phiMH1 | Leuconostoc phage phiMH1 Leuconostoc virus MH1 Seongbukvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38709 | 38.678 | Seongbukvirus | Unclassified | Siphoviridae | Leuconostoc |
| NC_055036 | Staphylococcus phage IME1348_01 | Staphylococcus phage IME1348_01 Staphylococcus virus IME1348-01 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42371 | 34.686 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055031 | Paenibacillus phage Unity | Paenibacillus phage Unity Paenibacillus virus Unity Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50316 | 49.11 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_055028 | Paenibacillus phage C7Cdelta | Paenibacillus phage C7Cdelta Paenibacillus virus C7Cdelta Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55774 | 48.008 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_055025 | Staphylococcus phage vB_SpsM_WIS42 | Staphylococcus phage vB_SpsM_WIS42 Staphylococcus virus WIS42 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40093 | 35.695 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055024 | Staphylococcus phage SpT152 | Staphylococcus phage SpT152 Staphylococcus virus SpT152 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41087 | 35.537 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055023 | Staphylococcus phage SP276 | Staphylococcus phage SP276 Staphylococcus virus SP276 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40147 | 35.353 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055022 | Staphylococcus phage SP197 | Staphylococcus phage SP197 Staphylococcus virus SP197 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 35.673 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055021 | Staphylococcus phage SP120 | Staphylococcus phage SP120 Staphylococcus virus SP120 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40530 | 34.991 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055020 | Staphylococcus phage SN8 | Staphylococcus phage SN8 Staphylococcus virus SN8 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39803 | 35.791 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055019 | Staphylococcus phage phiSP44-1 | Staphylococcus phage phiSP44-1 Staphylococcus virus phiSP44-1 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42565 | 35.9 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_055018 | Lactococcus phage PC_S1 | Lactococcus phage PC_S1 Lactococcus virus PCS1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21629 | 36.104 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055017 | Lactococcus phage PC_B1 | Lactococcus phage PC_B1 Lactococcus virus PCB1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22099 | 36.092 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055016 | Lactococcus phage vB_LacS_15 | Lactococcus phage vB_LacS_15 Lactococcus virus LacS15 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21006 | 36.142 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055015 | Lactococcus phage CHPC1242 | Lactococcus phage CHPC1242 Lactococcus virus CHPC1242 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21144 | 35.873 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055014 | Lactococcus phage CHPC1183 | Lactococcus phage CHPC1183 Lactococcus virus CHPC1183 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20030 | 36.216 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055013 | Lactococcus phage CHPC1182 | Lactococcus phage CHPC1182 Lactococcus virus CHPC1182 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20751 | 35.96 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055012 | Lactococcus phage CHPC1170 | Lactococcus phage CHPC1170 Lactococcus virus CHPC1170 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21746 | 35.616 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055011 | Lactococcus phage CHPC1020 | Lactococcus phage CHPC1020 Lactococcus virus CHPC1020 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22410 | 35.573 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055010 | Lactococcus phage CHPC972 | Lactococcus phage CHPC972 Lactococcus virus CHPC972 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23281 | 35.51 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055009 | Lactococcus phage CHPC967 | Lactococcus phage CHPC967 Lactococcus virus CHPC967 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22405 | 35.729 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055008 | Lactococcus phage CHPC966 | Lactococcus phage CHPC966 Lactococcus virus CHPC966 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21749 | 36.264 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055007 | Lactococcus phage CHPC122 | Lactococcus phage CHPC122 Lactococcus virus CHPC122 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22109 | 35.474 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055006 | Lactococcus phage CHPC116 | Lactococcus phage CHPC116 Lactococcus virus CHPC116 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21869 | 35.731 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055005 | Lactococcus phage vB_Llc_bIBB94p4 | Lactococcus phage vB_Llc_bIBB94p4 Lactococcus virus blBB94p4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21834 | 36.077 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055004 | Lactococcus phage vB_Llc_bIBBp6/4 | Lactococcus phage vB_Llc_bIBBp6/4 Lactococcus virus bIBBp6-4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22220 | 35.482 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055003 | Lactococcus phage vB_Llc_bIBB12 | Lactococcus phage vB_Llc_bIBB12 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21458 | 36.206 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055002 | Lactococcus phage vB_Llc_bIBBE1 | Lactococcus phage vB_Llc_bIBBE1 Lactococcus virus bIBBE1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21749 | 36.043 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055001 | Lactococcus phage vB_Llc_bIBBAm4 | Lactococcus phage vB_Llc_bIBBAm4 Lactococcus virus bIBBAm4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22703 | 35.872 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055000 | Lactococcus phage vB_Llc_bIBBA3 | Lactococcus phage vB_Llc_bIBBA3 Lactococcus virus bIBBA3 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21906 | 36.159 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054999 | Lactococcus phage vB_LLc_bIBB14 | Lactococcus phage vB_LLc_bIBB14 Lactococcus virus bIBB14 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21710 | 36.039 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054998 | Lactococcus phage 62606 | Lactococcus phage 62606 Lactococcus virus 62606 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22185 | 36.407 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054997 | Lactococcus phage 62403 | Lactococcus phage 62403 Lactococcus virus 62403 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21704 | 35.851 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054996 | Lactococcus phage 62402 | Lactococcus phage 62402 Lactococcus virus 62402 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21980 | 35.983 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054995 | Lactococcus phage 5171F | Lactococcus phage 5171F Lactococcus virus 5171F Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21078 | 36.218 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054994 | Lactococcus phage 50504 | Lactococcus phage 50504 Lactococcus virus 50504 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21562 | 35.92 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054993 | Lactococcus phage 50102 | Lactococcus phage 50102 Lactococcus virus 50102 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22242 | 36.022 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054992 | Lactococcus phage 37203 | Lactococcus phage 37203 Lactococcus virus 37203 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22029 | 35.921 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054991 | Lactococcus phage 20R03M | Lactococcus phage 20R03M Lactococcus virus 20R03M Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21750 | 35.959 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054990 | Lactococcus phage 05802 | Lactococcus phage 05802 Lactococcus virus 05802 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22434 | 35.727 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054989 | Brevibacterium phage AGM1 | Brevibacterium phage AGM1 Brevibacterium virus AGM1 Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36032 | 60.774 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| NC_054988 | Leuconostoc phage Ln-7 | Leuconostoc phage Ln-7 Leuconostoc virus Ln7 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28093 | 36.301 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_054987 | Leuconostoc phage P965 | Leuconostoc phage P965 Leuconostoc virus P965 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26526 | 36.504 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_054986 | Leuconostoc phage LDG | Leuconostoc phage LDG Leuconostoc virus LDG Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26559 | 36.274 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_054985 | Paenibacillus phage Wanderer | Paenibacillus phage Wanderer Paenibacillus virus Wanderer Wanderervirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40448 | 42.353 | Wanderervirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| NC_054984 | Paenibacillus phage Dragolir | Paenibacillus phage Dragolir Paenibacillus virus Dragolir Dragolirvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41131 | 43.981 | Dragolirvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| NC_054983 | Staphylococcus phage UPMK_2 | Staphylococcus phage UPMK_2 Staphylococcus virus UPMK2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40955 | 35.393 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054982 | Staphylococcus phage Sebago | Staphylococcus phage Sebago Staphylococcus virus Sebago Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43878 | 35.314 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054980 | Staphylococcus phage Henu2 | Staphylococcus phage Henu2 Staphylococcus virus Henu2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43513 | 35.04 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054979 | Staphylococcus virus B122 | Staphylococcus virus B122 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43108 | 34.987 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054978 | Staphylococcus phage TEM126 | Staphylococcus phage TEM126 Staphylococcus virus TEM126 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41882 | 33.676 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054977 | Staphylococcus phage SP5 | Staphylococcus phage SP5 Staphylococcus virus SP5 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43305 | 34.446 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054976 | Staphylococcus phage SAP33 | Staphylococcus phage SAP33 Staphylococcus virus SAP33 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42414 | 33.963 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054975 | Staphylococcus phage SA75 | Staphylococcus phage SA75 Staphylococcus virus SA75 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43134 | 34.437 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054974 | Staphylococcus phage HSA84 | Staphylococcus phage HSA84 Staphylococcus virus HSA84 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42650 | 34.567 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054973 | Staphylococcus virus Baq_Sau1 | Staphylococcus virus Baq_Sau1 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44384 | 34.046 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_054954 | Natrialba phage PhiCh1 | Natrialba phage PhiCh1 Natrialba virus PhiCh1 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58487 | 61.898 | Myohalovirus | Unclassified | Myoviridae | Natrialba |
| NC_054953 | Halobacterium virus ChaoS9 | Halobacterium virus ChaoS9 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55145 | 65.273 | Myohalovirus | Unclassified | Myoviridae | Halobacterium |
| NC_054945 | Yersinia phage vB_YepM_ZN18 | Yersinia phage vB_YepM_ZN18 Yersinia virus ZN18 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167948 | 35.353 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_054944 | Yersinia phage PYPS2T | Yersinia phage PYPS2T Yersinia virus PYPS2T Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169604 | 35.404 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_054943 | Yersinia phage fPS-2 | Yersinia phage fPS-2 Yersinia virus fPS-2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169060 | 35.347 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_054942 | Shigella phage SH7 | Shigella phage SH7 Shigella virus SH7 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164870 | 35.446 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_054941 | Shigella phage Sf23 | Shigella phage Sf23 Shigella virus Sf23 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167678 | 35.382 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_054940 | Shigella phage JK45 | Shigella phage JK45 Shigella virus JK45 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170740 | 37.639 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_054939 | Shigella phage CM8 | Shigella phage CM8 Shigella virus CM8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167247 | 35.28 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_054937 | Salmonella phage pSe_SNUABM_01 | Salmonella phage pSe_SNUABM_01 Salmonella virus SNUABM-01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172360 | 35.36 | Tequatrovirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_054936 | Salmonella phage SG1 | Salmonella phage SG1 Salmonella virus SG1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169805 | 35.276 | Tequatrovirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_054931 | Escherichia phage T2 | Escherichia phage T2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163825 | 35.318 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054930 | Escherichia phage PP01 | Escherichia phage PP01 Escherichia virus PP01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167812 | 35.459 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054929 | Escherichia phage PE37 | Escherichia phage PE37 Escherichia virus PE37 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166423 | 35.353 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054928 | Escherichia phage vB_vPM_PD112 | Escherichia phage vB_vPM_PD112 Escherichia virus PD112 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168084 | 35.364 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054926 | Escherichia phage vB_EcoM_OE5505 | Escherichia phage vB_EcoM_OE5505 Escherichia virus OE5505 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168756 | 35.221 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054925 | Escherichia phage vB_EcoM_NBG2 | Escherichia phage vB_EcoM_NBG2 Escherichia virus NBG2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166083 | 35.448 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054923 | Escherichia phage KIT03 | Escherichia phage KIT03 Escherichia virus KIT03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166848 | 35.342 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054922 | Escherichia phage vB_EcoM_KAW1E185 | Escherichia phage vB_EcoM_KAW1E185 Escherichia virus KAW1E185 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164987 | 35.368 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054920 | Escherichia phage vB_EcoM_G9062 | Escherichia phage vB_EcoM_G9062 Escherichia virus G9062 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168670 | 35.296 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054919 | Escherichia phage vB_EcoM_G4507 | Escherichia phage vB_EcoM_G4507 Escherichia virus G4507 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168828 | 35.367 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054918 | Escherichia phage vB_EcoM_G4498 | Escherichia phage vB_EcoM_G4498 Escherichia virus G4498 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167826 | 35.46 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054917 | Escherichia phage vB_EcoM_G50 | Escherichia phage vB_EcoM_G50 Escherichia virus G50 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167728 | 35.515 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054916 | Escherichia phage vB_EcoM-G28 | Escherichia phage vB_EcoM-G28 Escherichia virus G28 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170121 | 35.346 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054915 | Escherichia phage vB_EcoM_G8 | Escherichia phage vB_EcoM_G8 Escherichia virus G8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169650 | 35.326 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054914 | Escherichia phage vB_EcoM-fFiEco06 | Escherichia phage vB_EcoM-fFiEco06 Escherichia virus fFiEco06 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167076 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054911 | Escherichia phage ECO4 | Escherichia phage ECO4 Escherichia virus ECO4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168638 | 35.466 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054910 | Escherichia phage EcNP1 | Escherichia phage EcNP1 Escherichia virus EcNP1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166176 | 35.497 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054909 | Escherichia phage EC121 | Escherichia phage EC121 Escherichia virus EC121 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168805 | 35.529 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054908 | Escherichia phage vB_EcoM_DalCa | Escherichia phage vB_EcoM_DalCa Escherichia virus DalCa Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166040 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_054907 | Enterobacteria phage T6 | Enterobacteria phage T6 Enterobacteria virus T6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168697 | 35.211 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054906 | Enterobacteria phage RB18 | Enterobacteria phage RB18 Enterobacteria virus RB18 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166677 | 35.371 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054905 | Enterobacteria phage Kha5h | Enterobacteria phage Kha5h Enterobacteria virus Kha5h Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167318 | 35.548 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054904 | Enterobacteria phage vB_EcoM_IME340 | Enterobacteria phage vB_EcoM_IME340 Enterobacteria virus IME340 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165549 | 35.549 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054903 | Enterobacteria phage GiZh | Enterobacteria phage GiZh Enterobacteria virus GiZh Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166514 | 35.451 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054902 | Enterobacteria phage Aplg8 | Enterobacteria phage Aplg8 Enterobacteria virus Aplg8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168496 | 35.533 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_054899 | Citrobacter phage vB_CroM_CrRp10 | Citrobacter phage vB_CroM_CrRp10 Citrobacter virus CrRp10 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168403 | 35.508 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_054895 | Escherichia phage vB_Eco_SLUR29 | Escherichia phage vB_Eco_SLUR29 Escherichia virus SLUR29 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48466 | 44.739 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_054894 | Escherichia phage Henu7 | Escherichia phage Henu7 Escherichia virus Henu7 Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49526 | 48.205 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_054893 | Escherichia phage vB_EcoS_W011D | Escherichia phage vB_EcoS_W011D Escherichia virus W011D Changchunvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49847 | 46.241 | Changchunvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_054892 | Escherichia phage vB_EcoS_PHB17 | Escherichia phage vB_EcoS_PHB17 Escherichia virus PHB17 Changchunvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48939 | 46.897 | Changchunvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MW722821 | Escherichia phage 590B | Escherichia phage 590B unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44390 | 50.892 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_054815 | Mycobacterium phage Phaded | Mycobacterium phage Phaded unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53598 | 63.118 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054814 | Mycobacterium phage IronMan | Mycobacterium phage IronMan unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53467 | 63.316 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054813 | Mycobacterium phage Malec | Mycobacterium phage Malec unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53377 | 63.647 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054812 | Mycobacterium phage JoshKayV | Mycobacterium phage JoshKayV unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53366 | 63.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054811 | Mycobacterium phage Heffalump | Mycobacterium phage Heffalump unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53085 | 63.577 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054809 | Mycobacterium phage Fameo | Mycobacterium phage Fameo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52618 | 63.408 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054808 | Mycobacterium phage Centaur | Mycobacterium phage Centaur unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53001 | 63.499 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054807 | Mycobacterium phage Bugsy | Mycobacterium phage Bugsy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49937 | 63.186 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054806 | Mycobacterium phage Benvolio | Mycobacterium phage Benvolio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53136 | 63.735 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054805 | Mycobacterium phage Anselm | Mycobacterium phage Anselm unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53545 | 63.461 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054804 | Mycobacterium phage Cindaradix | Mycobacterium phage Cindaradix unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51575 | 64.012 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054803 | Mycobacterium phage Mundrea | Mycobacterium phage Mundrea unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51257 | 63.952 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054802 | Mycobacterium phage Miramae | Mycobacterium phage Miramae unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51300 | 63.871 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054801 | Mycobacterium phage Noelle | Mycobacterium phage Noelle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51483 | 63.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054800 | Mycobacterium phage ICleared | Mycobacterium phage ICleared unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51440 | 63.861 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054799 | Mycobacterium phage Camperdownii | Mycobacterium phage Camperdownii unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51131 | 63.944 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054798 | Mycobacterium phage Wizard007 | Mycobacterium phage Wizard007 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51029 | 63.85 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054797 | Mycobacterium phage Cintron | Mycobacterium phage Cintron unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51679 | 64.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054796 | Mycobacterium phage LeoAvram | Mycobacterium phage LeoAvram unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47079 | 63.827 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054795 | Mycobacterium phage Cerulean | Mycobacterium phage Cerulean unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51809 | 63.927 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054794 | Mycobacterium phage Stasia | Mycobacterium phage Stasia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51591 | 63.591 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054793 | Mycobacterium phage Kimona | Mycobacterium phage Kimona unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50283 | 64.378 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054792 | Mycobacterium phage PP | Mycobacterium phage PP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51509 | 64.486 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054791 | Mycobacterium phage MyraDee | Mycobacterium phage MyraDee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50514 | 62.694 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054788 | Mycobacterium phage SWU2 | Mycobacterium phage SWU2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50013 | 62.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054787 | Mycobacterium phage Steamy | Mycobacterium phage Steamy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53420 | 63.401 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054786 | Mycobacterium phage Refuge | Mycobacterium phage Refuge unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53594 | 63.548 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054785 | Mycobacterium phage DarthPhader | Mycobacterium phage DarthPhader unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53432 | 63.159 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054784 | Mycobacterium phage Pharaoh | Mycobacterium phage Pharaoh unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 63.729 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054783 | Mycobacterium phage Yecey3 | Mycobacterium phage Yecey3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53277 | 62.862 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054782 | Mycobacterium phage Koko | Mycobacterium phage Koko unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52879 | 61.342 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054781 | Mycobacterium phage Priamo | Mycobacterium phage Priamo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51633 | 61.391 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054780 | Mycobacterium phage JewelBug | Mycobacterium phage JewelBug unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50341 | 61.602 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054779 | Mycobacterium phage Blinn1 | Mycobacterium phage Blinn1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51800 | 61.222 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054778 | Mycobacterium phage Archetta | Mycobacterium phage Archetta unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47409 | 60.609 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054777 | Mycobacterium phage Bluefalcon | Mycobacterium phage Bluefalcon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51052 | 60.854 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054776 | Mycobacterium phage Zolita | Mycobacterium phage Zolita unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51182 | 60.853 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054775 | Mycobacterium phage Scorpia | Mycobacterium phage Scorpia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50693 | 59.724 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054774 | Mycobacterium phage Naca | Mycobacterium phage Naca unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51594 | 59.831 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054773 | Mycobacterium phage Pistachio | Mycobacterium phage Pistachio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50006 | 63.782 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054772 | Mycobacterium phage BabyRay | Mycobacterium phage BabyRay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50657 | 63.555 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054770 | Mycobacterium phage Veracruz | Mycobacterium phage Veracruz unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50062 | 64.049 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054761 | Mycobacterium phage Zetzy | Mycobacterium phage Zetzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48229 | 64.028 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054760 | Mycobacterium phage SoilDragon | Mycobacterium phage SoilDragon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50293 | 64.019 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054759 | Mycobacterium phage PurpleHaze | Mycobacterium phage PurpleHaze unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48596 | 63.999 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054727 | Mycobacterium phage Phrappuccino | Mycobacterium phage Phrappuccino Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136361 | 67.378 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| NC_054726 | Gordonia phage Stormageddon | Gordonia phage Stormageddon Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136319 | 65.036 | Unclassified | Unclassified | Myoviridae | Gordonia |
| NC_054714 | Mycobacterium phage Phabba | Mycobacterium phage Phabba Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164254 | 65.209 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| NC_054681 | Streptomyces phage Lorelei | Streptomyces phage Lorelei unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50558 | 65.772 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054680 | Streptomyces phage Goby | Streptomyces phage Goby unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51393 | 65.803 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054678 | Streptomyces phage Yasdnil | Streptomyces phage Yasdnil unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51631 | 65.699 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054677 | Streptomyces phage Alsaber | Streptomyces phage Alsaber unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48803 | 65.912 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054675 | Streptomyces phage Saftant | Streptomyces phage Saftant unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48883 | 65.642 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054671 | Streptomyces phage Paedore | Streptomyces phage Paedore unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52743 | 66.998 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054670 | Streptomyces phage Thestral | Streptomyces phage Thestral unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52628 | 67.677 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054669 | Streptomyces phage Alvy | Streptomyces phage Alvy unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53011 | 67.548 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054668 | Streptomyces phage Daudau | Streptomyces phage Daudau unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50602 | 67.12 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054666 | Streptomyces phage Caelum | Streptomyces phage Caelum unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50374 | 66.969 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054665 | Streptomyces phage Tefunt | Streptomyces phage Tefunt unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50574 | 66.839 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054664 | Streptomyces phage Amethyst | Streptomyces phage Amethyst unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49372 | 67.224 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054663 | Streptomyces phage Diane | Streptomyces phage Diane unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50483 | 66.694 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054662 | Streptomyces phage Omar | Streptomyces phage Omar unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49299 | 67.293 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054661 | Streptomyces phage Hank144 | Streptomyces phage Hank144 unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50547 | 66.307 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054660 | Streptomyces phage Janus | Streptomyces phage Janus unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50632 | 67.047 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054656 | Streptomyces phage Yosif | Streptomyces phage Yosif unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50129 | 66.117 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054655 | Streptomyces phage Celia | Streptomyces phage Celia unclassified Arquatrovirinae Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50493 | 68.552 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054652 | Klebsiella phage YX3973 | Klebsiella phage YX3973 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46907 | 46.899 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| NC_054651 | Escherichia phage C1 | Escherichia phage C1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46667 | 46.596 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| NC_054650 | Shigella phage DS8 | Shigella phage DS8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44605 | 46.517 | Unclassified | Unclassified | Siphoviridae | Shigella |
| NC_054647 | Salmonella phage Akira | Salmonella phage Akira Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45367 | 46.022 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054646 | Salmonella phage VB_StyS_BS5 | Salmonella phage VB_StyS_BS5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47604 | 45.526 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054645 | Salmonella phage LPST10 | Salmonella phage LPST10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47657 | 45.532 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054644 | Salmonella virus VSt472 | Salmonella virus VSt472 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46905 | 45.643 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054643 | Salmonella phage SeSz-2 | Salmonella phage SeSz-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45049 | 45.972 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054642 | Salmonella phage Seszw_1 | Salmonella phage Seszw_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45881 | 45.938 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054640 | Salmonella phage Segz_1 | Salmonella phage Segz_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48285 | 46.416 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054639 | Salmonella phage Skate | Salmonella phage Skate Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47393 | 45.804 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054638 | Salmonella phage vB_SenS_SB28 | Salmonella phage vB_SenS_SB28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45126 | 46.164 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054637 | Escherichia phage vB_EcoS Sa179lw | Escherichia phage vB_EcoS Sa179lw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46833 | 45.974 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| NC_054636 | Shigella phage Sf11 SMD-2017 | Shigella phage Sf11 SMD-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46454 | 45.985 | Unclassified | Unclassified | Siphoviridae | Shigella |
| NC_054464 | Edwardsiella phage Edno5 | Edwardsiella phage Edno5 unclassified Gofduovirus Gofduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40844 | 50.803 | Gofduovirus | Unclassified | Myoviridae | Edwardsiella |
| NC_054455 | Mycobacterium phage Tapioca | Mycobacterium phage Tapioca unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44205 | 66.137 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054453 | Mycobacterium phage Fulbright | Mycobacterium phage Fulbright unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42396 | 66.275 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054452 | Mycobacterium phage Andies | Mycobacterium phage Andies unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43779 | 66.171 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054451 | Mycobacterium phage Kevin1 | Mycobacterium phage Kevin1 unclassified Buttersvirus Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41988 | 66.231 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054450 | Mycobacterium phage Aggie | Mycobacterium phage Aggie unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44333 | 66.537 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054449 | Mycobacterium phage Philonius | Mycobacterium phage Philonius unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43886 | 66.541 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_054439 | Mycobacterium phage Paito | Mycobacterium phage Paito unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42311 | 66.023 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054433 | Mycobacterium phage Weirdo19 | Mycobacterium phage Weirdo19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52581 | 70.195 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_053515 | Mycobacterium phage MrMagoo | Mycobacterium phage MrMagoo unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84303 | 60.796 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_053511 | Mycobacterium phage DyoEdafos | Mycobacterium phage DyoEdafos unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77566 | 58.539 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053510 | Mycobacterium phage Bromden | Mycobacterium phage Bromden unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70183 | 58.228 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053509 | Mycobacterium phage Tourach | Mycobacterium phage Tourach unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77876 | 58.755 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053508 | Mycobacterium phage GuuelaD | Mycobacterium phage GuuelaD unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76315 | 58.889 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053507 | Mycobacterium phage Finemlucis | Mycobacterium phage Finemlucis unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77031 | 58.859 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053505 | Mycobacterium phage Krypton555 | Mycobacterium phage Krypton555 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76075 | 59.268 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053504 | Mycobacterium phage Lumos | Mycobacterium phage Lumos unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75586 | 59.27 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_053240 | Gordonia phage Tangerine | Gordonia phage Tangerine unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57306 | 68.244 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053239 | Gordonia phage Bizzy | Gordonia phage Bizzy unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57312 | 68.321 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053238 | Gordonia phage Gaea | Gordonia phage Gaea unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57472 | 68.127 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053237 | Gordonia phage Ashertheman | Gordonia phage Ashertheman unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57555 | 68.031 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053234 | Gordonia phage Kroos | Gordonia phage Kroos unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57973 | 68.13 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053174 | Gordonia phage William | Gordonia phage William Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50678 | 66.536 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_053173 | Gordonia phage Toast | Gordonia phage Toast Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49604 | 66.785 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_053013 | Pseudomonas phage vB_Pae-SS2019XI | Pseudomonas phage vB_Pae-SS2019XI Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57567 | 60.132 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_053004 | Salmonella phage TS13 | Salmonella phage TS13 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59168 | 56.356 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_053002 | Escherichia phage Utah | Escherichia phage Utah unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59024 | 56.418 | Chivirus | Unclassified | Siphoviridae | Escherichia |
| NC_053001 | Salmonella phage SeZq-1 | Salmonella phage SeZq-1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60188 | 56.588 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_053000 | Salmonella phage SeWh-1 | Salmonella phage SeWh-1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59878 | 56.583 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052998 | Salmonella phage ST-374 | Salmonella phage ST-374 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59934 | 56.602 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052997 | Salmonella phage ST-118 | Salmonella phage ST-118 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59106 | 56.698 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052990 | Salmonella phage Season12 | Salmonella phage Season12 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59047 | 56.467 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_052988 | Klebsiella phage KPN U2874 | Klebsiella phage KPN U2874 unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59087 | 56.302 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| NC_052987 | Klebsiella phage KPN N98 | Klebsiella phage KPN N98 unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59214 | 56.289 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| NC_052985 | Aeromonas phage LAh_7 | Aeromonas phage LAh_7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61426 | 61.972 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| NC_052984 | Aeromonas phage vB_AhyS-A18P4 | Aeromonas phage vB_AhyS-A18P4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60975 | 62.021 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| NC_052650 | Halobacterium phage phiH | Halobacterium phage phiH Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58072 | 63.678 | Myohalovirus | Unclassified | Myoviridae | Halobacterium |
| NC_052470 | Mycobacterium phage Magnito | Mycobacterium phage Magnito unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51743 | 63.819 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052469 | Mycobacterium phage Traft412 | Mycobacterium phage Traft412 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51223 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052468 | Mycobacterium phage Sumter | Mycobacterium phage Sumter unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52656 | 63.67 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052466 | Mycobacterium phage SpikeBT | Mycobacterium phage SpikeBT unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50181 | 63.741 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052465 | Mycobacterium phage Snazzy | Mycobacterium phage Snazzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52220 | 63.61 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052464 | Mycobacterium phage Smairt | Mycobacterium phage Smairt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54655 | 63.555 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052463 | Mycobacterium phage Sibs6 | Mycobacterium phage Sibs6 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50210 | 63.766 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052462 | Mycobacterium phage Sagefire | Mycobacterium phage Sagefire unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51482 | 63.877 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052461 | Mycobacterium phage Rutherferd | Mycobacterium phage Rutherferd unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52364 | 63.746 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052460 | Mycobacterium phage Ruotula | Mycobacterium phage Ruotula unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52601 | 63.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052458 | Mycobacterium phage Ringer | Mycobacterium phage Ringer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52018 | 63.572 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052457 | Mycobacterium phage Rhynn | Mycobacterium phage Rhynn unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52522 | 63.611 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052456 | Mycobacterium phage Rajelicia | Mycobacterium phage Rajelicia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52843 | 63.592 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052455 | Mycobacterium phage Pita2 | Mycobacterium phage Pita2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51486 | 63.4 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052454 | Mycobacterium phage Pippin | Mycobacterium phage Pippin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52034 | 63.753 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052453 | Mycobacterium phage Petruchio | Mycobacterium phage Petruchio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49531 | 63.821 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052452 | Mycobacterium phage PacerPaul | Mycobacterium phage PacerPaul unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52732 | 63.576 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052451 | Mycobacterium phage Oogway | Mycobacterium phage Oogway unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51745 | 63.573 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052450 | Mycobacterium phage Niza | Mycobacterium phage Niza unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51672 | 63.448 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052449 | Mycobacterium phage NEHalo | Mycobacterium phage NEHalo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50535 | 63.819 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052447 | Mycobacterium phage Mryolo | Mycobacterium phage Mryolo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50505 | 63.974 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052446 | Mycobacterium phage MPlant7149 | Mycobacterium phage MPlant7149 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.655 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052445 | Mycobacterium phage Michley | Mycobacterium phage Michley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51527 | 63.706 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052443 | Mycobacterium phage Maroc7 | Mycobacterium phage Maroc7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53350 | 63.5 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052441 | Mycobacterium phage Magnar | Mycobacterium phage Magnar unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51428 | 63.726 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052440 | Mycobacterium phage Lopton | Mycobacterium phage Lopton unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52394 | 63.544 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052439 | Mycobacterium phage KyMonks1A | Mycobacterium phage KyMonks1A unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52145 | 63.619 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052438 | Mycobacterium phage Killigrew | Mycobacterium phage Killigrew unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51163 | 63.775 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052437 | Mycobacterium phage Kanely | Mycobacterium phage Kanely unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52539 | 63.545 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052435 | Mycobacterium phage JackSparrow | Mycobacterium phage JackSparrow unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51576 | 63.609 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052434 | Mycobacterium phage ILeeKay | Mycobacterium phage ILeeKay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51017 | 64 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052433 | Mycobacterium phage Ichabod | Mycobacterium phage Ichabod unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53317 | 63.751 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052431 | Mycobacterium phage GageAP | Mycobacterium phage GageAP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51904 | 63.465 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052430 | Mycobacterium phage Fajezeel | Mycobacterium phage Fajezeel unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 63.411 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052429 | Mycobacterium phage Dulcie | Mycobacterium phage Dulcie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53970 | 63.728 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052428 | Mycobacterium phage Crispicous1 | Mycobacterium phage Crispicous1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50827 | 63.694 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052427 | Mycobacterium phage Corvo | Mycobacterium phage Corvo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53325 | 63.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052426 | Mycobacterium phage ConceptII | Mycobacterium phage ConceptII unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53287 | 63.522 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052425 | Mycobacterium phage Bones | Mycobacterium phage Bones unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52528 | 63.796 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052424 | Mycobacterium phage Bob3 | Mycobacterium phage Bob3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52233 | 63.833 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052423 | Mycobacterium phage Blue | Mycobacterium phage Blue unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50864 | 63.935 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052422 | Mycobacterium phage BK1 | Mycobacterium phage BK1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49728 | 63.761 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052421 | Mycobacterium phage Bircsak | Mycobacterium phage Bircsak unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53622 | 63.56 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052420 | Mycobacterium phage BigPaolini | Mycobacterium phage BigPaolini unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49601 | 63.454 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052419 | Mycobacterium phage Big3 | Mycobacterium phage Big3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53442 | 63.439 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052418 | Mycobacterium phage BaconJack | Mycobacterium phage BaconJack unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53793 | 63.759 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052417 | Mycobacterium phage Atkinbua | Mycobacterium phage Atkinbua unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52955 | 63.312 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052416 | Mycobacterium phage Arlo | Mycobacterium phage Arlo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52960 | 63.75 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052415 | Mycobacterium phage Anglerfish | Mycobacterium phage Anglerfish unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52380 | 63.477 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052414 | Mycobacterium phage Ajay | Mycobacterium phage Ajay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49516 | 63.638 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052413 | Mycobacterium phage AFIS | Mycobacterium phage AFIS unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51737 | 63.687 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052412 | Mycobacterium phage Adahisdi | Mycobacterium phage Adahisdi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51703 | 63.801 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052411 | Mycobacterium phage Acme | Mycobacterium phage Acme unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51793 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_052410 | Mycobacterium phage Scowl | Mycobacterium phage Scowl unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51690 | 63.784 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051740 | Mycobacterium phage Xavia | Mycobacterium phage Xavia unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49808 | 65.859 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051739 | Mycobacterium phage Purky | Mycobacterium phage Purky unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50513 | 66.385 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051738 | Mycobacterium phage ThulaThula | Mycobacterium phage ThulaThula unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50415 | 66.53 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051737 | Mycobacterium phage Majeke | Mycobacterium phage Majeke unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47612 | 67.367 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051736 | Mycobacterium phage Arib1 | Mycobacterium phage Arib1 unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46732 | 67.474 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051735 | Mycobacterium phage FirstPlacePfu | Mycobacterium phage FirstPlacePfu unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45680 | 67.261 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051734 | Mycobacterium phage Bartholomew | Mycobacterium phage Bartholomew unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46484 | 67.189 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051733 | Mycobacterium phage Phineas | Mycobacterium phage Phineas unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47229 | 67.188 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051732 | Mycobacterium phage Ksquared | Mycobacterium phage Ksquared unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48699 | 67.094 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051731 | Mycobacterium phage GreaseLightnin | Mycobacterium phage GreaseLightnin unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48424 | 67.091 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051729 | Mycobacterium phage Atcoo | Mycobacterium phage Atcoo unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49075 | 67.016 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051728 | Mycobacterium phage Megiddo | Mycobacterium phage Megiddo unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48783 | 67.12 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_051725 | Mycobacterium phage Rem711 | Mycobacterium phage Rem711 unclassified Trigintaduovirus Trigintaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50832 | 66.214 | Trigintaduovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051724 | Mycobacterium phage Zapner | Mycobacterium phage Zapner unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55307 | 61.106 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051722 | Mycobacterium phage Demsculpinboyz | Mycobacterium phage Demsculpinboyz unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57437 | 60.865 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051720 | Mycobacterium phage Zerg | Mycobacterium phage Zerg unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58079 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051719 | Mycobacterium phage WillSterrel | Mycobacterium phage WillSterrel unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58608 | 61.637 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051718 | Mycobacterium phage Whouxphf | Mycobacterium phage Whouxphf unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59093 | 61.058 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051717 | Mycobacterium phage Wachhund | Mycobacterium phage Wachhund unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54513 | 61.681 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051715 | Mycobacterium phage TDanisky | Mycobacterium phage TDanisky unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 61.233 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051714 | Mycobacterium phage SwagPigglett | Mycobacterium phage SwagPigglett unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58930 | 61.259 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051713 | Mycobacterium phage SuperGrey | Mycobacterium phage SuperGrey unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59346 | 61.723 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051712 | Mycobacterium phage StAnnes | Mycobacterium phage StAnnes unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59357 | 61.802 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051711 | Mycobacterium phage Spoonbill | Mycobacterium phage Spoonbill unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55735 | 61.303 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051710 | Mycobacterium phage Spikelee | Mycobacterium phage Spikelee unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58604 | 61.038 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051709 | Mycobacterium phage SimranZ1 | Mycobacterium phage SimranZ1 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56335 | 61.26 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051707 | Mycobacterium phage SassyB | Mycobacterium phage SassyB unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55094 | 61.646 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051706 | Mycobacterium phage Sandalphon | Mycobacterium phage Sandalphon unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59540 | 61.407 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051704 | Mycobacterium phage RitaG | Mycobacterium phage RitaG unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57789 | 61.603 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051703 | Mycobacterium phage QuickMath | Mycobacterium phage QuickMath unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57526 | 61.706 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051702 | Mycobacterium phage Priscilla | Mycobacterium phage Priscilla unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58230 | 61.278 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051701 | Mycobacterium phage Pollywog | Mycobacterium phage Pollywog unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58397 | 61.251 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051700 | Mycobacterium phage Polka14 | Mycobacterium phage Polka14 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58758 | 61.438 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051699 | Mycobacterium phage Poenanya | Mycobacterium phage Poenanya unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58666 | 61.492 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051698 | Mycobacterium phage Pippy | Mycobacterium phage Pippy unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56663 | 61.504 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051697 | Mycobacterium phage Piper2020 | Mycobacterium phage Piper2020 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57703 | 62.676 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051696 | Mycobacterium phage Phasih | Mycobacterium phage Phasih unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56125 | 61.055 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051695 | Mycobacterium phage OwlsT2W | Mycobacterium phage OwlsT2W unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56515 | 61.219 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051694 | Mycobacterium phage OlympiaSaint | Mycobacterium phage OlympiaSaint unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58297 | 61.399 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051693 | Mycobacterium phage OldBen | Mycobacterium phage OldBen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57159 | 61.485 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051692 | Mycobacterium phage Ogopogo | Mycobacterium phage Ogopogo unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56867 | 61.139 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051691 | Mycobacterium phage Nivrat | Mycobacterium phage Nivrat unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58009 | 61.454 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051690 | Mycobacterium phage Nitzel | Mycobacterium phage Nitzel unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54857 | 61.316 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051689 | Mycobacterium phage Nimbo | Mycobacterium phage Nimbo unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55767 | 61.513 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051686 | Mycobacterium phage Misha28 | Mycobacterium phage Misha28 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57455 | 62.945 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051685 | Mycobacterium phage Minnie | Mycobacterium phage Minnie unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59268 | 61.485 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051684 | Mycobacterium phage MinionDave | Mycobacterium phage MinionDave unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58027 | 61.496 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051683 | Mycobacterium phage MilleniumForce | Mycobacterium phage MilleniumForce unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58106 | 61.782 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051682 | Mycobacterium phage Melissauren88 | Mycobacterium phage Melissauren88 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54356 | 61.783 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051681 | Mycobacterium phage Mattes | Mycobacterium phage Mattes unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58074 | 61.31 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051680 | Mycobacterium phage Mantra | Mycobacterium phage Mantra unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57001 | 61.545 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051679 | Mycobacterium phage Mahavrat | Mycobacterium phage Mahavrat unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55945 | 61.285 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051678 | Mycobacterium phage Lizziana | Mycobacterium phage Lizziana unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58423 | 61.335 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051676 | Mycobacterium phage KristaRAM | Mycobacterium phage KristaRAM unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58701 | 61.643 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051675 | Mycobacterium phage Krakatau | Mycobacterium phage Krakatau unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53058 | 61.501 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051674 | Mycobacterium phage Koella | Mycobacterium phage Koella unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54118 | 61.499 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051673 | Mycobacterium phage Kingsley | Mycobacterium phage Kingsley unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60300 | 61.38 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051672 | Mycobacterium phage Kersh | Mycobacterium phage Kersh unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60190 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051671 | Mycobacterium phage Kenuha5 | Mycobacterium phage Kenuha5 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56502 | 61.693 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051670 | Mycobacterium phage Juice456 | Mycobacterium phage Juice456 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56726 | 61.612 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051669 | Mycobacterium phage JoeyJr | Mycobacterium phage JoeyJr unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58962 | 61.445 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051667 | Mycobacterium phage Henu3 PeY-2017 | Mycobacterium phage Henu3 PeY-2017 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61276 | 61.172 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051666 | Mycobacterium phage Hegedechwinu | Mycobacterium phage Hegedechwinu unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57346 | 61.5 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051665 | Mycobacterium phage Harley | Mycobacterium phage Harley unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58731 | 61.422 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051664 | Mycobacterium phage Gorge | Mycobacterium phage Gorge unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57955 | 61.38 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051663 | Mycobacterium phage Girr | Mycobacterium phage Girr unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57754 | 61.355 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051661 | Mycobacterium phage Galactic | Mycobacterium phage Galactic unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57861 | 61.447 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051660 | Mycobacterium phage Frankie | Mycobacterium phage Frankie unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57036 | 61.896 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051659 | Mycobacterium phage Flathead | Mycobacterium phage Flathead unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57663 | 61.181 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051657 | Mycobacterium phage Filuzino | Mycobacterium phage Filuzino unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59306 | 62.095 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051656 | Mycobacterium phage Fancypants | Mycobacterium phage Fancypants unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60228 | 61.085 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051655 | Mycobacterium phage Enby | Mycobacterium phage Enby unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54687 | 61.276 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051654 | Mycobacterium phage Empress | Mycobacterium phage Empress unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57155 | 61.519 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051653 | Mycobacterium phage Emma | Mycobacterium phage Emma unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56418 | 61.321 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051652 | Mycobacterium phage EleanorGeorge | Mycobacterium phage EleanorGeorge unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59482 | 61.731 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051651 | Mycobacterium phage Eish | Mycobacterium phage Eish unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58150 | 61.214 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051649 | Mycobacterium phage Doug | Mycobacterium phage Doug unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58397 | 61.145 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051647 | Mycobacterium phage DillTech15 | Mycobacterium phage DillTech15 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57797 | 61.861 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051646 | Mycobacterium phage DaWorst | Mycobacterium phage DaWorst unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56827 | 60.982 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051645 | Mycobacterium phage Cornucopia | Mycobacterium phage Cornucopia unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55422 | 61.394 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051643 | Mycobacterium phage Clifton | Mycobacterium phage Clifton unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55727 | 61.658 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051642 | Mycobacterium phage Chuckly | Mycobacterium phage Chuckly unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55189 | 61.559 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051639 | Mycobacterium phage Burwell21 | Mycobacterium phage Burwell21 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58098 | 61.458 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051638 | Mycobacterium phage Bubbles123 | Mycobacterium phage Bubbles123 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57647 | 61.375 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051636 | Mycobacterium phage BobaPhett | Mycobacterium phage BobaPhett unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59815 | 61.326 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051635 | Mycobacterium phage Blexus | Mycobacterium phage Blexus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58824 | 61.432 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051634 | Mycobacterium phage Batiatus | Mycobacterium phage Batiatus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57737 | 61.836 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051633 | Mycobacterium phage ArcusAngelus | Mycobacterium phage ArcusAngelus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58569 | 61.396 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051632 | Mycobacterium phage Alexphander | Mycobacterium phage Alexphander unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57734 | 61.163 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051630 | Mycobacterium phage Yunkel11 | Mycobacterium phage Yunkel11 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60757 | 66.559 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051625 | Mycobacterium Phage Niklas | Mycobacterium Phage Niklas unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60989 | 68.509 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051602 | Mycobacterium phage Shandong1 | Mycobacterium phage Shandong1 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60618 | 67.46 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051598 | Mycobacterium phage Findley | Mycobacterium phage Findley unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58150 | 68.339 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051593 | Mycobacterium phage Collard | Mycobacterium phage Collard unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61395 | 65.556 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_050154 | Escherichia phage D6 | Escherichia phage D6 unclassified Punavirus Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91159 | 47.01 | Punavirus | Unclassified | Myoviridae | Escherichia |
| NC_050153 | Escherichia phage vB_EcoM-Ro157c2YLVW | Escherichia phage vB_EcoM-Ro157c2YLVW unclassified Punavirus Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80732 | 47.764 | Punavirus | Unclassified | Myoviridae | Escherichia |
| NC_050152 | Enterobacteria phage P7 | Enterobacteria phage P7 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101660 | 47.411 | Punavirus | Unclassified | Myoviridae | Enterobacteria |
| NC_050148 | Pseudomonas virus Pa193 | Pseudomonas virus Pa193 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66657 | 55.672 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_050145 | Pseudomonas phage PaGU11 | Pseudomonas phage PaGU11 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65554 | 55.551 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_049976 | Bacillus phage Nf | Bacillus phage Nf Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18753 | 37.317 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_049975 | Bacillus phage vB_BsuP-Goe1 | Bacillus phage vB_BsuP-Goe1 Bacillus virus Goe1 Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18379 | 37.494 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_049969 | Bacillus phage DK3 | Bacillus phage DK3 Bacillus virus DK3 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26865 | 31.055 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049968 | Bacillus phage DK2 | Bacillus phage DK2 Bacillus virus DK2 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26357 | 30.914 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049967 | Bacillus phage DK1 | Bacillus phage DK1 Bacillus virus DK1 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27180 | 30.868 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049966 | Bacillus phage vB_BthP-Goe4 | Bacillus phage vB_BthP-Goe4 Bacillus virus Goe4 Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25722 | 30.433 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049965 | Bacillus phage vB_BveP-Goe6 | Bacillus phage vB_BveP-Goe6 Bacillus virus Goe6 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19105 | 39.78 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_049964 | Bacillus phage KonjoTrouble | Bacillus phage KonjoTrouble Bacillus virus KonjoTrouble Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26061 | 30.125 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049963 | Bacillus phage Juan | Bacillus phage Juan Bacillus virus Juan Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25032 | 30.565 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049962 | Bacillus phage SerPounce | Bacillus phage SerPounce Bacillus virus SerPounce Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27206 | 30.431 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049959 | Bacillus phage QCM11 | Bacillus phage QCM11 Bacillus virus QCM11 Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26054 | 30.441 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_049954 | Escherichia virus Lambda_2G7b | Escherichia virus Lambda_2G7b unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47142 | 49.879 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049952 | Escherichia virus Lambda_2B8 | Escherichia virus Lambda_2B8 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48763 | 50.188 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049951 | Enterobacteria phage O276 | Enterobacteria phage O276 Enterobacteria virus O276 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41969 | 50.683 | Lambdavirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_049946 | Escherichia virus Lambda_4A7 | Escherichia virus Lambda_4A7 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49168 | 49.77 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_049945 | Stx2-converting phage Stx2a_WGPS6 | Stx2-converting phage Stx2a_WGPS6 Escherichia virus WGPS6 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57849 | 50.678 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| NC_049944 | Stx2-converting phage Stx2a_WGPS8 | Stx2-converting phage Stx2a_WGPS8 Escherichia virus WGPS8 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55023 | 51.03 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| NC_049943 | Enterobacteria phage 2851 | Enterobacteria phage 2851 Enterobacteria virus 2851 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57248 | 51.053 | Pankowvirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_049942 | Escherichia phage JLK-2012 | Escherichia phage JLK-2012 Escherichia virus JLK2012 Marienburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57198 | 50.719 | Marienburgvirus | Unclassified | Siphoviridae | Escherichia |
| NC_049941 | Stx2-converting phage Stx2a_WGPS2 | Stx2-converting phage Stx2a_WGPS2 Escherichia virus WGPS2 Sawaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58326 | 51.38 | Sawaravirus | Unclassified | Siphoviridae | Unspecified |
| NC_049926 | Escherichia phage ECP1 | Escherichia phage ECP1 Escherichia virus ECP1 Wongtaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40469 | 51.009 | Wongtaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_049925 | Stx converting phage vB_EcoS_P27 | Stx converting phage vB_EcoS_P27 Escherichia virus P27 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61580 | 49.169 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049924 | Stx2-converting phage Stx2a_F451 | Stx2-converting phage Stx2a_F451 Escherichia virus F451 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64900 | 49.767 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049923 | Stx2-converting phage Stx2a_WGPS9 | Stx2-converting phage Stx2a_WGPS9 Escherichia virus WGPS9 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56224 | 49.945 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049922 | Stx converting phage vB_EcoS_ST2-8624 | Stx converting phage vB_EcoS_ST2-8624 Enterobacteria virus ST2-8624 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62822 | 49.607 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049921 | Stx1 converting phage AU6Stx1 | Stx1 converting phage AU6Stx1 Escherichia virus AU6Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57123 | 48.732 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049920 | Stx1 converting phage AU5Stx1 | Stx1 converting phage AU5Stx1 Escherichia virus AU5Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58371 | 49.775 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_049919 | Escherichia phage SH2026Stx1 | Escherichia phage SH2026Stx1 Escherichia virus SH2026Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61564 | 49.391 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_049918 | Escherichia phage ArgO145 | Escherichia phage ArgO145 Escherichia virus ArgO145 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62020 | 50.002 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_049917 | Escherichia phage Lyz12581Vzw | Escherichia phage Lyz12581Vzw Eschericia virus Lyz12581Vzw Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62668 | 50.137 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_049855 | Lactococcus phage LW81 | Lactococcus phage LW81 Lactococcus virus LW81 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128179 | 32.57 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049854 | Lactococcus phage AM1 | Lactococcus phage AM1 Lactococcus virus AM1 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125658 | 32.551 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049847 | Klebsiella phage Sin4 | Klebsiella phage Sin4 Klebsiella virus Sin4 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49916 | 50.288 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049839 | Klebsiella phage Sweeny | Klebsiella phage Sweeny Klebsiella virus Sweeny Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50241 | 50.672 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049853 | Escherichia phage Henu8 | Escherichia phage Henu8 Escherichia virus Henu8 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49890 | 44.171 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049848 | Klebsiella phage KP1801 | Klebsiella phage KP1801 Klebsiella virus KP1801 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49835 | 50.137 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049846 | Klebsiella phage Shelby | Klebsiella phage Shelby Klebsiella virus Shelby Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49045 | 50.407 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049845 | Klebsiella phage PhiKpNIH-2 | Klebsiella phage PhiKpNIH-2 Klebsiella virus PhiKpNIH2 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49477 | 50.419 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049844 | Klebsiella phage 13 | Klebsiella phage 13 Klebsiella virus 13 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43094 | 50.629 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049843 | Klebsiella phage JY917 | Klebsiella phage JY917 Klebsiella virus JY917 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37655 | 50.432 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049842 | Klebsiella phage vB_KpnS_15-38_KLPPOU149 | Klebsiella phage vB_KpnS_15-38_KLPPOU149 Klebsiella virus KLPPOU149 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49316 | 51.032 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049841 | Klebsiella phage Skenny | Klebsiella phage Skenny Klebsiella virus Skenny Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49935 | 50.794 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049840 | Klebsiella phage vB_KpnS_FZ10 | Klebsiella phage vB_KpnS_FZ10 Klebsiella virus FZ10 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50381 | 50.66 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049838 | Klebsiella phage KL | Klebsiella phage KL Klebsiella virus KL Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47844 | 50.907 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049837 | Klebsiella phage vB_KpnS_Alina | Klebsiella phage vB_KpnS_Alina Klebsiella virus Alina Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51780 | 51.613 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049836 | Klebsiella phage vB_KpnS_Call | Klebsiella phage vB_KpnS_Call Klebsiella virus Call Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51487 | 51.549 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049835 | Klebsiella phage vB_KpnS_Domnhall | Klebsiella phage vB_KpnS_Domnhall Klebsiella virus Domnhall Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54438 | 51.587 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049834 | Klebsiella phage vB_KpnS_IMGroot | Klebsiella phage vB_KpnS_IMGroot Klebsiella virus IMGroot Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52866 | 51.347 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049833 | Klebsiella phage vB_KpnS_SegesCirculi | Klebsiella phage vB_KpnS_SegesCirculi Klebsiella virus SegesCirculi Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50713 | 51.121 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_049831 | Shigella virus Sfin-3 | Shigella virus Sfin-3 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50309 | 45.352 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_049822 | Escherichia phage vB_EcoS_G29-2 | Escherichia phage vB_EcoS_G29-2 Escherichia virus G29-2 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51739 | 44.017 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_049814 | Shigella phage JK16 | Shigella phage JK16 Shigella virus JK16 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51854 | 44.546 | Warwickvirus | Tempevirinae | Drexlerviridae | Shigella |
| NC_049813 | Escherichia phage vB_EcoS-12210I | Escherichia phage vB_EcoS-12210I Escherichia virus 12210I Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.413 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_049812 | Lactococcus phage P1045 | Lactococcus phage P1045 Lactococcus virus P1045 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31050 | 36.135 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049811 | Lactococcus phage 62503 | Lactococcus phage 62503 Lactococcus virus 62503 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30556 | 36.32 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049810 | Lactococcus phage 50902 | Lactococcus phage 50902 Lactococcus virus 50902 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30745 | 36.045 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049809 | Lactococcus phage Q33 | Lactococcus phage Q33 Lactococcus virus Q33 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31106 | 36.215 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_023601 | Pseudomonas phage phiPsa374 | Pseudomonas phage phiPsa374 Pseudomonas virus Psa374 Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98287 | 47.716 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| NC_049511 | Acinetobacter phage AM101 | Acinetobacter phage AM101 Acinetobacter virus AM101 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166487 | 36.708 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049510 | Erwinia phage phiEaP8 | Erwinia phage phiEaP8 Erwinia virus phiEaP8 Yonginvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75929 | 46.841 | Yonginvirus | Erskinevirinae | Schitoviridae | Erwinia |
| NC_049494 | Acinetobacter phage vB_AbaM_PhT2 | Acinetobacter phage vB_AbaM_PhT2 Acinetobacter virus PhT2 Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166330 | 36.372 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049493 | Acinetobacter phage vB_AbaM_Lazarus | Acinetobacter phage vB_AbaM_Lazarus Acinetobacter virus Lazarus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165151 | 36.763 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049492 | Acinetobacter phage vB_AbaM_Kimel | Acinetobacter phage vB_AbaM_Kimel Acinetobacter virus Kimel Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167922 | 36.729 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049491 | Acinetobacter phage vB_AbaM_Apostate | Acinetobacter phage vB_AbaM_Apostate Acinetobacter virus Apostate Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164746 | 36.752 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049490 | Acinetobacter phage vB_AbaM_Berthold | Acinetobacter phage vB_AbaM_Berthold Acinetobacter virus Berthold Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165927 | 36.714 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049488 | Lactococcus phage CHPC965 | Lactococcus phage CHPC965 Lactococcus virus CHPC965 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25327 | 34.876 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049487 | Lactococcus phage CHPC964 | Lactococcus phage CHPC964 Lactococcus virus CHPC964 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29925 | 34.399 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049486 | Lactococcus phage CHPC959 | Lactococcus phage CHPC959 Lactococcus virus CHPC959 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29340 | 34.315 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049485 | Lactococcus phage CHPC958 | Lactococcus phage CHPC958 Lactococcus virus CHPC958 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32653 | 34.478 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049484 | Lactococcus phage CHPC781 | Lactococcus phage CHPC781 Lactococcus virus CHPC781 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29218 | 34.475 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049483 | Lactococcus phage CHPC52 | Lactococcus phage CHPC52 Lactococcus virus CHPC52 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29652 | 34.149 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049482 | Lactococcus phage CHPC362 | Lactococcus phage CHPC362 Lactococcus virus CHPC362 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27642 | 35.428 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049481 | Lactococcus phage CHPC361 | Lactococcus phage CHPC361 Lactococcus virus CHPC361 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30162 | 34.451 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049480 | Lactococcus phage CHPC129 | Lactococcus phage CHPC129 Lactococcus virus CHPC129 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30832 | 34.561 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049479 | Acinetobacter phage vB_AbaM_Konradin | Acinetobacter phage vB_AbaM_Konradin Acinetobacter virus Konradin Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166210 | 36.746 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049478 | Arthrobacter phage Mufasa8 | Arthrobacter phage Mufasa8 Arthrobacter virus Mufasa8 Mufasoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60205 | 66.136 | Mufasoctovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_049477 | Vibrio phage Valm-yong1 | Vibrio phage Valm-yong1 Vibrio virus Yong1 Yongunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33851 | 45.564 | Yongunavirus | Peduovirinae | Myoviridae | Vibrio |
| NC_049476 | Lactococcus phage P4565 | Lactococcus phage P4565 Lactococcus virus P4565 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32697 | 34.236 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049475 | Lactococcus phage P656 | Lactococcus phage P656 Lactococcus virus P656 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28396 | 35.075 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049474 | Lactococcus phage P1532 | Lactococcus phage P1532 Lactococcus virus P1532 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29075 | 35.384 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049470 | Arthrobacter phage TripleJ | Arthrobacter phage TripleJ Arthrobacter virus TripleJ Triplejayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45507 | 64.504 | Triplejayvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_049469 | Sinorhizobium phage ort11 | Sinorhizobium phage ort11 Sinorhizobium virus ort11 Huelvavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75239 | 44.246 | Huelvavirus | Unclassified | Schitoviridae | Sinorhizobium |
| NC_049464 | Serratia phage Muldoon | Serratia phage Muldoon Serratia virus Muldoon Muldoonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167547 | 41.802 | Muldoonvirus | Unclassified | Myoviridae | Serratia |
| NC_049461 | Bacteriophage R18C | Bacteriophage R18C Citrobacter virus R18C Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31834 | 51.602 | Peduovirus | Peduovirinae | Myoviridae | Unspecified |
| NC_049460 | Salmonella phage SI7 | Salmonella phage SI7 Salmonella virus SI7 Irrigatiovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30164 | 54.363 | Irrigatiovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_049459 | Salmonella phage SW9 | Salmonella phage SW9 Salmonella virus SW9 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31123 | 53.006 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| NC_049457 | Escherichia phage vB_EcoM-12474III | Escherichia phage vB_EcoM-12474III Escherichia virus 12474III Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33807 | 50.918 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049454 | Klebsiella phage ST15-OXA48phi14.1 | Klebsiella phage ST15-OXA48phi14.1 Klebsiella virus ST15OXA48phi14-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33839 | 52.747 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049453 | Klebsiella phage ST13-OXA48phi12.1 | Klebsiella phage ST13-OXA48phi12.1 Klebsiella virus ST13OXA48phi12-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39086 | 53.526 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049452 | Klebsiella phage ST512-KPC3phi13.2 | Klebsiella phage ST512-KPC3phi13.2 Klebsiella virus ST512KPC3phi13-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32302 | 51.873 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049451 | Klebsiella phage ST147-VIM1phi7.1 | Klebsiella phage ST147-VIM1phi7.1 Klebsiella virus ST147VIM1phi7-1 Vimunumvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34141 | 52.969 | Vimunumvirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049450 | Klebsiella phage ST16-OXA48phi5.4 | Klebsiella phage ST16-OXA48phi5.4 Klebsiella virus ST16OXA48phi5-4 Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38322 | 50.06 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049449 | Klebsiella phage ST437-OXA245phi4.2 | Klebsiella phage ST437-OXA245phi4.2 Klebsiella virus ST437OXA245phi4-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18281 | 50.501 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049448 | Klebsiella phage ST437-OXA245phi4.1 | Klebsiella phage ST437-OXA245phi4.1 Klebsiella virus ST437OXA245phi4-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39642 | 52.76 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_049447 | Lactococcus phage MV10L | Lactococcus phage MV10L Lactococcus virus MV10L Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31374 | 34.876 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049446 | Lactococcus phage 8V08 | Lactococcus phage 8V08 Lactococcus virus 8V08 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32312 | 35.071 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049441 | Acinetobacter phage KARL-1 | Acinetobacter phage KARL-1 Acinetobacter virus KARL1 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166560 | 36.786 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049438 | Acinetobacter phage vB_ApiM_fHyAci03 | Acinetobacter phage vB_ApiM_fHyAci03 Acinetobacter virus fHyAci03 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165975 | 36.758 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_049434 | Lactococcus phage MP1 | Lactococcus phage MP1 Lactococcus virus MP1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32575 | 35.008 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049432 | Ralstonia phage RsoM1USA | Ralstonia phage RsoM1USA Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39309 | 65.153 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| NC_049429 | Enterobacter phage phiT5282H | Enterobacter phage phiT5282H Enterobacter virus phiT5282H Novemvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31978 | 52.458 | Novemvirus | Peduovirinae | Myoviridae | Enterobacter |
| NC_049426 | Lactococcus phage vB_Llc_bIBBF12 | Lactococcus phage vB_Llc_bIBBF12 Lactococcus virus bIBBF12 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29454 | 34.824 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049425 | Lactococcus phage vB_Llc_bIBB5g1 | Lactococcus phage vB_Llc_bIBB5g1 Lactococcus virus bIBB5g1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30132 | 34.667 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049424 | Lactococcus phage LP1502a | Lactococcus phage LP1502a Lactococcus virus LP1502a Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30247 | 35.121 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049423 | Lactococcus phage LP0903 | Lactococcus phage LP0903 Lactococcus virus LP0903 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31049 | 34.9 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049422 | Lactococcus phage LP0209 | Lactococcus phage LP0209 Lactococcus virus LP0209 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30339 | 34.761 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049421 | Lactococcus phage LP9908 | Lactococcus phage LP9908 Lactococcus virus LP9908 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29668 | 34.829 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049420 | Lactococcus phage LP9903 | Lactococcus phage LP9903 Lactococcus virus LP9903 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29557 | 34.96 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049419 | Lactococcus phage LP9609 | Lactococcus phage LP9609 Lactococcus virus LP9609 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30945 | 34.765 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049418 | Lactococcus phage LP9404 | Lactococcus phage LP9404 Lactococcus virus LP9404 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30072 | 34.946 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049417 | Lactococcus phage LP9207 | Lactococcus phage LP9207 Lactococcus virus LP9207 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29203 | 35.01 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049416 | Lactococcus phage LP8511 | Lactococcus phage LP8511 Lactococcus virus LP8511 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29953 | 35.112 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049415 | Lactococcus phage 96603 | Lactococcus phage 96603 Lactococcus virus 96603 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30690 | 34.904 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049414 | Lactococcus phage 96403 | Lactococcus phage 96403 Lactococcus virus 96403 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26874 | 34.476 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049413 | Lactococcus phage 96401 | Lactococcus phage 96401 Lactococcus virus 96401 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29306 | 35.16 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049412 | Lactococcus phage 88605 | Lactococcus phage 88605 Lactococcus virus 88605 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31508 | 34.467 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049411 | Lactococcus phage 79201 | Lactococcus phage 79201 Lactococcus virus 79201 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31123 | 34.932 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049410 | Lactococcus phage 66901 | Lactococcus phage 66901 Lactococcus virus 66901 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28917 | 34.71 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049409 | Lactococcus phage 63302 | Lactococcus phage 63302 Lactococcus virus 63302 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31883 | 34.774 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049408 | Lactococcus phage 62605 | Lactococcus phage 62605 Lactococcus virus 62605 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31014 | 34.562 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049407 | Lactococcus phage 62601 | Lactococcus phage 62601 Lactococcus virus 62601 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30852 | 34.62 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049406 | Lactococcus phage 57001 | Lactococcus phage 57001 Lactococcus virus 57001 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31819 | 34.3 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049405 | Lactococcus phage 56301 | Lactococcus phage 56301 Lactococcus virus 56301 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32555 | 33.958 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049404 | Lactococcus phage 56003 | Lactococcus phage 56003 Lactococcus virus 56003 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30926 | 34.524 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049403 | Lactococcus phage 51701 | Lactococcus phage 51701 Lactococcus virus 51701 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32529 | 34.357 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049402 | Lactococcus phage 38507 | Lactococcus phage 38507 Lactococcus virus 38507 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32458 | 33.56 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049401 | Lactococcus phage 38503 | Lactococcus phage 38503 Lactococcus virus 38503 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27948 | 34.414 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049400 | Lactococcus phage 37201 | Lactococcus phage 37201 Lactococcus virus 37201 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30989 | 34.79 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049399 | Lactococcus phage 30804 | Lactococcus phage 30804 Lactococcus virus 30804 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30401 | 34.193 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049398 | Lactococcus phage 16802 | Lactococcus phage 16802 Lactococcus virus 16802 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28721 | 35.239 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049397 | Lactococcus phage 05601 | Lactococcus phage 05601 Lactococcus virus 05601 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32895 | 34.084 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049394 | Escherichia phage ESSI2_ev129 | Escherichia phage ESSI2_ev129 Escherichia virus ev129 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30927 | 51.483 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049393 | Escherichia phage ESSI2_ev040 | Escherichia phage ESSI2_ev040 Escherichia virus ev040 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30597 | 50.822 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049392 | Escherichia phage ESSI2_ev239 | Escherichia phage ESSI2_ev239 Escherichia virus ev239 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29203 | 51.378 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049383 | Tenacibaculum phage pT24 | Tenacibaculum phage pT24 Tenacibaculum virus pT24 Kungbxnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 234670 | 28.717 | Kungbxnavirus | Unclassified | Myoviridae | Tenacibaculum |
| NC_049381 | Lactococcus phage R31 | Lactococcus phage R31 Lactococcus virus R31 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27203 | 35.378 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049378 | Salmonella phage ST-W77 | Salmonella phage ST-W77 Salmonella virus STW77 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157458 | 44.482 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049377 | Lactococcus phage 13W11L | Lactococcus phage 13W11L Lactococcus virus 13W11L Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29952 | 34.709 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049376 | Lactococcus phage 3R16S | Lactococcus phage 3R16S Lactococcus virus 3R16S Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30446 | 34.98 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049375 | Lactococcus phage i0139 | Lactococcus phage i0139 Lactococcus virus i0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29513 | 35.1 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049374 | Lactococcus phage 10W18 | Lactococcus phage 10W18 Lactococcus virus 10W18 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32393 | 34.949 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049373 | Lactococcus phage 16W23 | Lactococcus phage 16W23 Lactococcus virus 16W23 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32477 | 34.76 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049370 | Lactococcus phage 936 group phage PhiS0139 | Lactococcus phage 936 group phage PhiS0139 Lactococcus virus PhiS0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30116 | 35.197 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049369 | Lactococcus phage 936 group phage PhiLj | Lactococcus phage 936 group phage PhiLj Lactococcus virus PhiLj Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28983 | 35.138 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049368 | Lactococcus phage 936 group phage PhiM1127 | Lactococcus phage 936 group phage PhiM1127 Lactococcus virus PhiM1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32682 | 34.79 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049367 | Lactococcus phage 936 group phage PhiE1127 | Lactococcus phage 936 group phage PhiE1127 Lactococcus virus PhiE1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32270 | 34.62 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049366 | Lactococcus phage 936 group phage Phi155 | Lactococcus phage 936 group phage Phi155 Lactococcus virus Phi155 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32150 | 34.936 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049365 | Lactococcus phage 936 group phage PhiJF1 | Lactococcus phage 936 group phage PhiJF1 Lactococcus virus PhiJF1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31901 | 34.579 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049364 | Lactococcus phage 936 group phage Phi91127 | Lactococcus phage 936 group phage Phi91127 Lactococcus virus Phi91127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31495 | 34.821 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049363 | Lactococcus phage 936 group phage Phi44 | Lactococcus phage 936 group phage Phi44 Lactococcus virus Phi44 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31379 | 34.622 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049362 | Lactococcus phage 936 group phage Phi4.2 | Lactococcus phage 936 group phage Phi4.2 Lactococcus virus Phi42 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31375 | 34.489 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049361 | Lactococcus phage 936 group phage PhiL.6 | Lactococcus phage 936 group phage PhiL.6 Lactococcus virus PhiL6 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31181 | 34.55 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049360 | Lactococcus phage 936 group phage Phi109 | Lactococcus phage 936 group phage Phi109 Lactococcus virus Phi109 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31033 | 34.921 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049359 | Lactococcus phage 936 group phage PhiL.18 | Lactococcus phage 936 group phage PhiL.18 Lactococcus virus PhiL18 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30882 | 35.007 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049358 | Lactococcus phage 936 group phage PhiF0139 | Lactococcus phage 936 group phage PhiF0139 Lactococcus virus PhiF0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30811 | 34.669 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049357 | Lactococcus phage 936 group phage Phi5.12 | Lactococcus phage 936 group phage Phi5.12 Lactococcus virus Phi512 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29026 | 34.745 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049356 | Lactococcus phage 936 group phage Phi19.3 | Lactococcus phage 936 group phage Phi19.3 Lactococcus virus Phi193 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28811 | 34.792 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049355 | Lactococcus phage 936 group phage PhiB1127 | Lactococcus phage 936 group phage PhiB1127 Lactococcus virus PhiB1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28785 | 35.459 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049354 | Lactococcus phage 936 group phage PhiA.16 | Lactococcus phage 936 group phage PhiA.16 Lactococcus virus PhiA16 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28349 | 35.246 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049353 | Lactococcus phage phi145 | Lactococcus phage phi145 Lactococcus virus phi145 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30862 | 34.901 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049352 | Lactococcus phage phi15 | Lactococcus phage phi15 Lactococcus virus phi15 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31945 | 34.575 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049351 | Lactococcus phage CaseusJM1 | Lactococcus phage CaseusJM1 Lactococcus virus CaseusJM1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30692 | 34.836 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049349 | Lactococcus lactis phage P475 | Lactococcus lactis phage P475 Lactococcus virus P475 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30961 | 34.324 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049348 | Lactococcus lactis phage p272 | Lactococcus lactis phage p272 Lactococcus virus p272 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30778 | 34.145 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049347 | Lactococcus phage fd13 | Lactococcus phage fd13 Lactococcus virus fd13 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30674 | 34.717 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049346 | Lactococcus phage 936 | Lactococcus phage 936 Lactococcus virus 936 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27302 | 34.518 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049340 | Tenacibaculum phage PTm1 | Tenacibaculum phage PTm1 Tenacibaculum virus PTm1 Shirahamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224680 | 29.744 | Shirahamavirus | Unclassified | Myoviridae | Tenacibaculum |
| NC_048892 | Leptospira phage LE1 | Leptospira phage LE1 Leptospira virus LE1 Saintgironsvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73623 | 38.527 | Saintgironsvirus | Unclassified | Myoviridae | Leptospira |
| NC_048625 | Lactobacillus phage T25 | Lactobacillus phage T25 Lactobacillus virus T25 Sukhumvitvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38421 | 44.541 | Sukhumvitvirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_048624 | Staphylococcus phage phi7247PVL | Staphylococcus phage phi7247PVL Staphylococcus virus phi7247PVL Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42481 | 33.309 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048628 | Bacillus phage BceA1 | Bacillus phage BceA1 Bacillus virus BceA1 Beceayunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42932 | 35.663 | Beceayunavirus | Unclassified | Siphoviridae | Bacillus |
| NC_048626 | Pseudomonas phage PA01 | Pseudomonas phage PA01 Pseudomonas virus PA01 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66220 | 55.402 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048656 | Klebsiella phage vB_KpnM_BIS47 | Klebsiella phage vB_KpnM_BIS47 Klebsiella virus BIS47 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147443 | 44.622 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| NC_048658 | Staphylococcus phage SA7 | Staphylococcus phage SA7 Staphylococcus virus SA7 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34730 | 34.129 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048841 | Flavobacterium phage vB_FspS_snusmum6-1 | Flavobacterium phage vB_FspS_snusmum6-1 Flavobacterium virus Snusmum Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39365 | 29.051 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048840 | Flavobacterium phage vB_FspS_snork6-1 | Flavobacterium phage vB_FspS_snork6-1 Flavobacterium virus Snork Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38214 | 29.45 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048839 | Flavobacterium phage vB_FspS_sniff9-1 | Flavobacterium phage vB_FspS_sniff9-1 Flavobacterium virus Sniff Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38134 | 29.404 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048836 | Flavobacterium phage vB_FspS_morran9-1 | Flavobacterium phage vB_FspS_morran9-1 Flavobacterium virus Morran Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38385 | 29.462 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048835 | Flavobacterium phage vB_FspS_lillamy9-1 | Flavobacterium phage vB_FspS_lillamy9-1 Flavobacterium virus Lillamy Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.364 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048834 | Flavobacterium phage vB_FspS_laban6-1 | Flavobacterium phage vB_FspS_laban6-1 Flavobacterium virus Laban Labanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43236 | 31.141 | Labanvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048832 | Flavobacterium phage vB_FspS_hattifnatt9-1 | Flavobacterium phage vB_FspS_hattifnatt9-1 Flavobacterium virus Hattifnatt Hattifnattvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39262 | 29.196 | Hattifnattvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048828 | Erwinia phage Hena1 | Erwinia phage Hena1 Erwinia virus Hena1 Henunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148842 | 48.404 | Henunavirus | Vequintavirinae | Myoviridae | Erwinia |
| NC_048827 | Vibrio phage vB_VchM_Kuja | Vibrio phage vB_VchM_Kuja Vibrio virus Kuja Kujavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148180 | 36.378 | Kujavirus | Unclassified | Ackermannviridae | Vibrio |
| NC_048825 | Gordonia phage Crocheter | Gordonia phage Crocheter Gordonia virus Crocheter Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60531 | 51.623 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048823 | Gordonia phage Untouchable | Gordonia phage Untouchable Gordonia virus Untouchable Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61623 | 51.552 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048822 | Gordonia phage Avazak | Gordonia phage Avazak Gordonia virus Avazak Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60278 | 51.45 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048817 | Gordonia phage Tanis | Gordonia phage Tanis Gordonia virus Tanis Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59727 | 51.009 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048816 | Gordonia phage OhMyWard | Gordonia phage OhMyWard Gordonia virus OhMyWard Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60978 | 52.153 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048810 | Gordonia phage Chikenjars | Gordonia phage Chikenjars Gordonia virus Chikenjars Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61544 | 51.334 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048807 | Erwinia phage pEp_SNUABM_01 | Erwinia phage pEp_SNUABM_01 Erwinia virus SNUABM01 Henunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147321 | 48.706 | Henunavirus | Vequintavirinae | Myoviridae | Erwinia |
| NC_048806 | Pseudomonas virus PA8P1 | Pseudomonas virus PA8P1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65690 | 55.695 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048805 | Klebsiella phage Soft | Klebsiella phage Soft Klebsiella virus Soft Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57805 | 55.871 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| NC_048799 | Escherichia phage Mangalitsa | Escherichia phage Mangalitsa Escherichia virus Mangalitsa Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52329 | 44.012 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| NC_048798 | Klebsiella phage Marfa | Klebsiella phage Marfa Klebsiella virus Marfa Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168532 | 40.993 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| NC_048797 | Bacillus phage vB_BmeM-Goe8 | Bacillus phage vB_BmeM-Goe8 Bacillus virus vBBmeMGoe8 Goettingenvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161583 | 38.872 | Goettingenvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_048793 | Gordonia phage Kenosha | Gordonia phage Kenosha Gordonia virus Kenosha Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60899 | 51.753 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048791 | Corynebacterium phage Adelaide | Corynebacterium phage Adelaide Corynebacterium virus Adelaide Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44104 | 67.982 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048790 | Corynebacterium phage Lederberg | Corynebacterium phage Lederberg unclassified Samwavirus Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45644 | 68.072 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048789 | Corynebacterium phage Stiles | Corynebacterium phage Stiles Corynebacterium virus Stiles Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42846 | 67.663 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048788 | Mycobacterium phage ThetaBob | Mycobacterium phage ThetaBob Mycobacterium virus ThetaBob Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56714 | 61.71 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048787 | Corynebacterium phage Dina | Corynebacterium phage Dina Corynebacterium virus Dina Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44381 | 67.95 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048784 | Rhodococcus phage Whack | Rhodococcus phage Whack Rhodococcus virus Whack Whackvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49660 | 61.911 | Whackvirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_048782 | Rhodococcus phage Sleepyhead | Rhodococcus phage Sleepyhead Rhodococcus virus Sleepyhead Sleepyheadvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43943 | 61.022 | Sleepyheadvirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_048780 | Corynebacterium phage StAB | Corynebacterium phage StAB Corynebacterium virus StAB Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45751 | 67.861 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048765 | Vibrio phage VAP7 | Vibrio phage VAP7 Vibrio virus VAP7 Vapseptimavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144685 | 41.864 | Vapseptimavirus | Unclassified | Ackermannviridae | Vibrio |
| NC_048764 | Salmonella phage SE4 | Salmonella phage SE4 unclassified Loughboroughvirus Loughboroughvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53494 | 45.478 | Loughboroughvirus | Unclassified | Myoviridae | Salmonella |
| NC_048763 | Salmonella phage SE13 | Salmonella phage SE13 Salmonella virus SE13 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52438 | 45.78 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| NC_048762 | Bacillus phage vB_BtS_B83 | Bacillus phage vB_BtS_B83 Bacillus virus B83 Pushchinovirus Skryabinvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49952 | 35.79 | Pushchinovirus | Skryabinvirinae | Siphoviridae | Bacillus |
| NC_048758 | Serratia phage Parlo | Serratia phage Parlo Serratia virus Parlo Parlovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62853 | 58.042 | Parlovirus | Unclassified | Podoviridae | Serratia |
| NC_048756 | Nodularia phage vB_NspS-kac65v151 | Nodularia phage vB_NspS-kac65v151 Nodularia virus kac65v151 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146618 | 40.31 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| NC_048750 | Gordonia phage Duffington | Gordonia phage Duffington Gordonia virus Duffington Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61343 | 51.373 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048745 | Pseudomonas phage vB_PaeM_SCUT-S1 | Pseudomonas phage vB_PaeM_SCUT-S1 Pseudomonas virus S1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66086 | 55.432 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048744 | Pseudomonas phage EPa61 | Pseudomonas phage EPa61 Pseudomonas virus EPa61 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65905 | 55.134 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048741 | Proteus phage Mydo | Proteus phage Mydo Proteus virus Mydo Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145127 | 44.809 | Mydovirus | Vequintavirinae | Myoviridae | Proteus |
| NC_048737 | Lactobacillus phage BH1 | Lactobacillus phage BH1 Lactobacillus virus BH1 Behunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40721 | 44.834 | Behunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_048732 | Klebsiella phage Seifer | Klebsiella phage Seifer Klebsiella virus Seifer Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58197 | 56.106 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| NC_048730 | Streptomyces phage Yaboi | Streptomyces phage Yaboi Streptomyces virus Yaboi Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131250 | 49.301 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048727 | Salmonella phage Siskin | Salmonella phage Siskin Salmonella virus Siskin Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58476 | 56.604 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_048726 | Bacillus phage BSP38 | Bacillus phage BSP38 Bacillus virus BSP38 Jeonjuvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153268 | 41.747 | Jeonjuvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_048724 | Streptomyces phage Karimac | Streptomyces phage Karimac Streptomyces virus Karimac Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131909 | 49.411 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048723 | Streptomyces phage LukeCage | Streptomyces phage LukeCage Streptomyces virus LukeCage Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133195 | 49.05 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048722 | Streptomyces phage Wollford | Streptomyces phage Wollford Streptomyces virus Wollford Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133007 | 47.731 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048721 | Streptomyces phage StarPlatinum | Streptomyces phage StarPlatinum Streptomyces virus StarPlatinum Karimacvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133886 | 49.466 | Karimacvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048718 | Paenibacillus phage Jacopo | Paenibacillus phage Jacopo Paenibacillus virus Jacopo Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38526 | 41.559 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048717 | Paenibacillus phage Eltigre | Paenibacillus phage Eltigre Paenibacillus virus Eltigre Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38675 | 41.417 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048716 | Paenibacillus phage Lucielle | Paenibacillus phage Lucielle Paenibacillus virus Lucielle Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37947 | 41.821 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048715 | Paenibacillus phage Kawika | Paenibacillus phage Kawika Paenibacillus virus Kawika Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40769 | 41.782 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048714 | Paenibacillus phage Yyerffej | Paenibacillus phage Yyerffej Paenibacillus virus Yyerffej Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43126 | 40.551 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048713 | Staphylococcus phage SA345ruMSSAST8 | Staphylococcus phage SA345ruMSSAST8 Staphylococcus virus SA345ruMSSAST8 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42735 | 33.062 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048711 | Staphylococcus phage SA780ruMSSAST101 | Staphylococcus phage SA780ruMSSAST101 Staphylococcus virus SA780ruMSSAST101 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43184 | 33.554 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048710 | Staphylococcus phage SA1014ruMSSAST7 | Staphylococcus phage SA1014ruMSSAST7 Staphylococcus virus SA1014ruMSSAST7 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43504 | 33.08 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048705 | Mycobacterium phage TChen | Mycobacterium phage TChen Mycobacterium virus TChen Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57742 | 62.338 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048701 | Anoxybacillus phage A403 | Anoxybacillus phage A403 Anoxybacillus virus A403 Tandoganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40847 | 36.769 | Tandoganvirus | Unclassified | Siphoviridae | Anoxybacillus |
| NC_048700 | Klebsiella phage KP8 | Klebsiella phage KP8 Klebsiella virus KP8 Kaypoctavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73679 | 44.201 | Kaypoctavirus | Enquatrovirinae | Schitoviridae | Klebsiella |
| NC_048699 | Pseudomonas phage vB_PaeM_LS1 | Pseudomonas phage vB_PaeM_LS1 Pseudomonas virus LS1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66095 | 55.693 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048694 | Klebsiella phage vB_Kpn_F48 | Klebsiella phage vB_Kpn_F48 Klebsiella virus F48 Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170764 | 40.783 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| NC_048693 | Paenibacillus phage Likha | Paenibacillus phage Likha Paenibacillus virus Likha Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39778 | 41.302 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048692 | Paenibacillus phage Leyra | Paenibacillus phage Leyra Paenibacillus virus Leyra Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42276 | 41.395 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048691 | Paenibacillus phage Tadhana | Paenibacillus phage Tadhana Paenibacillus virus Tadhana Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37880 | 42.112 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048690 | Paenibacillus phage Pagassa | Paenibacillus phage Pagassa Paenibacillus virus Pagassa Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40035 | 42.001 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048689 | Paenibacillus phage PBL1c | Paenibacillus phage PBL1c Paenibacillus virus PBL1c Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40611 | 41.247 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_048683 | Salmonella phage KFS-SE1 | Salmonella phage KFS-SE1 Salmonella virus KFSSE1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59715 | 56.711 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_048682 | Raoultella phage Ro1 | Raoultella phage Ro1 Raoultella virus Ro1 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145759 | 44.548 | Mydovirus | Vequintavirinae | Myoviridae | Raoultella |
| NC_048681 | Staphylococcus phage phiSa2wa_st22 | Staphylococcus phage phiSa2wa_st22 Staphylococcus virus st22 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38576 | 33.241 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048680 | Lactobacillus phage LJ | Lactobacillus phage LJ Lactobacillus virus LJ Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44260 | 44.781 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_048676 | Pseudomonas phage SL1 | Pseudomonas phage SL1 Pseudomonas virus SL1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65847 | 55.585 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048670 | Klebsiella phage KPN N137 | Klebsiella phage KPN N137 Klebsiella virus N137 Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59100 | 56.301 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| NC_048667 | Bacillus phage Deep-Purple | Bacillus phage Deep-Purple Bacillus virus DeepPurple Deurplevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36278 | 38.357 | Deurplevirus | Unclassified | Siphoviridae | Bacillus |
| NC_048665 | Clostridium phage phiCDHM14 | Clostridium phage phiCDHM14 Clostridioides virus phiCDHM14 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32651 | 29.54 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_048663 | Pseudomonas phage R26 | Pseudomonas phage R26 Pseudomonas virus R26 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65737 | 55.071 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048662 | Pseudomonas phage R12 | Pseudomonas phage R12 Pseudomonas virus R12 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65415 | 55.397 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048657 | Staphylococcus phage IME1361_01 | Staphylococcus phage IME1361_01 Staphylococcus virus IME1361-01 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43516 | 32.85 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048655 | Salmonella phage BSPM4 | Salmonella phage BSPM4 Salmonella virus BSPM4 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59097 | 56.541 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_048652 | Bacillus phage vB_BsuM-Goe3 | Bacillus phage vB_BsuM-Goe3 Bacillus virus vBBsuMGoe3 Grisebachstrassevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156430 | 41.927 | Grisebachstrassevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_048651 | Aeribacillus phage AP45 | Aeribacillus phage AP45 Aeribacillus virus AP45 Kamchatkavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51606 | 38.273 | Kamchatkavirus | Unclassified | Siphoviridae | Aeribacillus |
| NC_048649 | Salmonella phage UPF_BP2 | Salmonella phage UPF_BP2 Salmonella virus UPFBP2 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54894 | 46.009 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| NC_048648 | Salmonella phage ST-101 | Salmonella phage ST-101 Salmonella virus ST101 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59132 | 56.702 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_048644 | Staphylococcus phage 3 AJ-2017 | Staphylococcus phage 3 AJ-2017 Staphylococcus virus 3AJ2017 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43922 | 33.277 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048643 | Clostridium phage CDKM15 | Clostridium phage CDKM15 Clostridioides virus CDKM15 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50605 | 28.683 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| NC_048642 | Clostridium phage CDKM9 | Clostridium phage CDKM9 Clostridioides virus CDKM9 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49822 | 28.975 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| NC_048641 | Bacillus phage PfEFR-4 | Bacillus phage PfEFR-4 Bacillus virus PfEFR4 Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43223 | 35.437 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| NC_048640 | Bacillus phage vB_BtS_BMBtp14 | Bacillus phage vB_BtS_BMBtp14 Bacillus virus BMBtp14 Bembunaquatrovirus Skryabinvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50740 | 36.762 | Bembunaquatrovirus | Skryabinvirinae | Siphoviridae | Bacillus |
| NC_048636 | Staphylococcus phage P1105 | Staphylococcus phage P1105 Staphylococcus virus P1105 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42282 | 33.463 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048635 | Staphylococcus phage P630 | Staphylococcus phage P630 Staphylococcus virus P630 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40448 | 33.747 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048634 | Staphylococcus phage P282 | Staphylococcus phage P282 Staphylococcus virus P282 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41960 | 32.96 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_048633 | Bacillus phage vB_BceS-MY192 | Bacillus phage vB_BceS-MY192 Bacillus virus MY192 Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44696 | 34.979 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| NC_048632 | Salmonella phage 35 | Salmonella phage 35 Salmonella virus 35 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55391 | 56.845 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_048631 | Bacillus phage phi105 | Bacillus phage phi105 Bacillus virus phi105 Spizizenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39318 | 42.673 | Spizizenvirus | Unclassified | Siphoviridae | Bacillus |
| MT366944 | Yersinia phage vB_YenM_281 | Yersinia phage vB_YenM_281 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50481 | 46.814 | Unclassified | Unclassified | Myoviridae | Yersinia |
| NC_042139 | Corynebacterium phage Poushou | Corynebacterium phage Poushou Corynebacterium virus Poushou Poushouvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40353 | 60.008 | Poushouvirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048024 | Klebsiella phage NJS1 | Klebsiella phage NJS1 Klebsiella virus NJS1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49292 | 50.665 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048208 | Escherichia phage vB_EcoS_IME542 | Escherichia phage vB_EcoS_IME542 Escherichia virus IME542 Christensenvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46553 | 43.568 | Christensenvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_048206 | Escherichia phage vB_Eco_mar001J1 | Escherichia phage vB_Eco_mar001J1 Escherichia virus mar001J1 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50343 | 44.395 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_048204 | Escherichia phage vB_Eco_mar001J1 | Escherichia phage vB_Eco_mar001J1 Escherichia virus mar001J1 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50343 | 44.382 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_048202 | Escherichia phage vB_EcoS_swan01 | Escherichia phage vB_EcoS_swan01 Escherichia virus swan01 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50865 | 44.74 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_048201 | Pseudomonas phage 17A | Pseudomonas phage 17A Pseudomonas virus 17A Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40242 | 57.711 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_048200 | Pseudomonas phage shl2 | Pseudomonas phage shl2 Pseudomonas virus shl2 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40466 | 56.606 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_048197 | Erwinia phage vB_EhrS_49 | Erwinia phage vB_EhrS_49 Erwinia virus Eho49 Feofaniavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46835 | 50.844 | Feofaniavirus | Unclassified | Siphoviridae | Erwinia |
| NC_048192 | Staphylococcus phage vB_SpsS_QT1 | Staphylococcus phage vB_SpsS_QT1 Staphylococcus virus QT1 Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43029 | 36.882 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_048185 | Gordonia phage Fairfaxidum | Gordonia phage Fairfaxidum Gordonia virus Fairfaxidum Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50187 | 66.585 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048183 | Synechococcus phage S-CBP4 | Synechococcus phage S-CBP4 Synechococcus virus SCBP4 Poseidonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41824 | 44.293 | Poseidonvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_048181 | Raoultella phage RP180 | Raoultella phage RP180 Raoultella virus RP180 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44851 | 50.342 | Kagunavirus | Guernseyvirinae | Siphoviridae | Raoultella |
| NC_048180 | Shigella phage VB_Ship_A7 | Shigella phage VB_Ship_A7 Shigella virus A7 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40058 | 48.405 | Berlinvirus | Studiervirinae | Autographiviridae | Shigella |
| NC_048179 | Salmonella phage ZCSE2 | Salmonella phage ZCSE2 Salmonella virus ZCSE2 Loughboroughvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53965 | 45.843 | Loughboroughvirus | Unclassified | Myoviridae | Salmonella |
| NC_048178 | Escherichia phage Sortsne | Escherichia phage Sortsne Escherichia virus Sortsne Sortsnevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41912 | 59.973 | Sortsnevirus | Unclassified | Podoviridae | Escherichia |
| NC_048177 | Escherichia phage Lidtsur | Escherichia phage Lidtsur Escherichia virus Lidtsur Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42291 | 54.633 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| NC_048176 | Gordonia phage GodonK | Gordonia phage GodonK Gordonia virus GodonK Godonkavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122563 | 54.382 | Godonkavirus | Unclassified | Siphoviridae | Gordonia |
| NC_048175 | Klebsiella phage Pharr | Klebsiella phage Pharr Klebsiella virus Pharr Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40599 | 53.267 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048174 | Shigella phage Buco | Shigella phage Buco Shigella virus Buco Bucovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43630 | 54.082 | Bucovirus | Slopekvirinae | Autographiviridae | Shigella |
| NC_048172 | Escherichia phage Minorna | Escherichia phage Minorna Escherichia virus Minorna Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43264 | 51.456 | Drulisvirus | Slopekvirinae | Autographiviridae | Escherichia |
| NC_048171 | Synechococcus phage S-B28 | Synechococcus phage S-B28 Synechococcus virus SB28 Qadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45424 | 42.059 | Qadamvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_048170 | Escherichia phage vB_EcoM_Goslar | Escherichia phage vB_EcoM_Goslar Escherichia virus Goslar Goslarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237307 | 46.53 | Goslarvirus | Unclassified | Myoviridae | Escherichia |
| NC_048169 | Gordonia phage BrutonGaster | Gordonia phage BrutonGaster Gordonia virus BrutonGaster Oneupvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91510 | 61.48 | Oneupvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048168 | Pseudomonas phage RLP | Pseudomonas phage RLP Pseudomonas virus RLP Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43446 | 62.093 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_048165 | Microbacterium phage Fireman | Microbacterium phage Fireman Microbacterium virus Fireman Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54579 | 68.574 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048164 | Escherichia phage HZP2 | Escherichia phage HZP2 Escherichia virus HZP2 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32448 | 48.305 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048163 | Escherichia phage NC-A | Escherichia phage NC-A Escherichia virus NCA Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39536 | 48.816 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048162 | Klebsiella phage GH-K3 | Klebsiella phage GH-K3 Klebsiella virus GHK3 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49427 | 50.169 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048161 | Cronobacter phage GW1 | Cronobacter phage GW1 Cronobacter virus GW1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39695 | 53.183 | Kayfunavirus | Studiervirinae | Autographiviridae | Cronobacter |
| NC_048160 | Halorubrum pleomorphic virus 9 | Halorubrum pleomorphic virus 9 Betapleolipovirus HRPV9 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16159 | 60.326 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_048159 | Staphylococcus phage CSA13 | Staphylococcus phage CSA13 Staphylococcus virus CSA13 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17034 | 28.966 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_048155 | Streptomyces phage Lilbooboo | Streptomyces phage Lilbooboo Streptomyces virus Lilbooboo Vashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41267 | 62.66 | Vashvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048154 | Streptomyces phage Vash | Streptomyces phage Vash Streptomyces virus Vash Vashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41578 | 62.485 | Vashvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048153 | Streptomyces phage Austintatious | Streptomyces phage Austintatious Streptomyces virus Austintatious Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36213 | 72.598 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048150 | Shigella phage vB_SsoS_008 | Shigella phage vB_SsoS_008 Shigella virus 008 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50414 | 45.656 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_048146 | Gordonia phage Asapag | Gordonia phage Asapag Gordonia virus Asapag Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55119 | 63.071 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_048143 | Klebsiella phage KpKT21phi1 | Klebsiella phage KpKT21phi1 Klebsiella virus KpKT21phi1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49106 | 50.89 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048141 | Gordonia phage Kenna | Gordonia phage Kenna Gordonia virus Kenna Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53215 | 62.975 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_048140 | Arthrobacter phage Maja | Arthrobacter phage Maja Arthrobacter virus Maja Majavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36776 | 65.053 | Majavirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048138 | Klebsiella phage Henu1 | Klebsiella phage Henu1 Klebsiella virus Henu1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40352 | 53.142 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048136 | Escherichia phage Ro45lw | Escherichia phage Ro45lw Escherichia virus Ro45lw Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39793 | 52.19 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048135 | Enterobacter phage EcpYZU01 | Enterobacter phage EcpYZU01 Enterobacter virus EcpYZU01 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39767 | 51.751 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| NC_048132 | Escherichia phage vB_EcoS-95 | Escherichia phage vB_EcoS-95 Escherichia virus 95 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50910 | 44.83 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_048131 | Klebsiella phage KN3-1 | Klebsiella phage KN3-1 Klebsiella virus KN3-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41059 | 53.491 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048130 | Klebsiella phage KN4-1 | Klebsiella phage KN4-1 Klebsiella virus KN4-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41219 | 52.944 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048129 | Klebsiella phage KN1-1 | Klebsiella phage KN1-1 Klebsiella virus KN1-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40236 | 52.843 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048126 | Klebsiella phage TSK1 | Klebsiella phage TSK1 Klebsiella virus TSK1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49861 | 50.452 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048114 | Klebsiella phage kpssk3 | Klebsiella phage kpssk3 Klebsiella virus kpssk3 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40539 | 52.803 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048113 | Salmonella phage Lumpael | Salmonella phage Lumpael Salmonella virus Lumpael Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41126 | 59.466 | Murrayvirus | Unclassified | Siphoviridae | Salmonella |
| NC_048112 | Escherichia phage Skarpretter | Escherichia phage Skarpretter Escherichia virus Skarpretter Skarprettervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42042 | 55.82 | Skarprettervirus | Unclassified | Podoviridae | Escherichia |
| NC_048109 | Pseudomonas phage Dobby | Pseudomonas phage Dobby Pseudomonas virus Dobby Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37152 | 62.215 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| NC_048108 | Yersinia phage PYPS50 | Yersinia phage PYPS50 Yersinia virus PYPS50 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39648 | 48.736 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_048107 | Staphylococcus phage Pabna | Staphylococcus phage Pabna Staphylococcus virus Pabna Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17700 | 29.362 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_048106 | Synechococcus phage S-CBWM1 | Synechococcus phage S-CBWM1 Synechococcus virus SCBWM1 Aokuangvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139069 | 51.623 | Aokuangvirus | Unclassified | Myoviridae | Synechococcus |
| NC_048105 | Salmonella phage BSP161 | Salmonella phage BSP161 Salmonella virus BSP161 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39688 | 48.72 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| NC_048103 | Pectobacterium phage PPWS2 | Pectobacterium phage PPWS2 Pectobacterium virus PPWS2 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44716 | 50.968 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048102 | Synechococcus phage S-P4 | Synechococcus phage S-P4 Synechococcus virus SP4 Leucotheavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158485 | 39.946 | Leucotheavirus | Unclassified | Myoviridae | Synechococcus |
| NC_048097 | Arthrobacter phage Yang | Arthrobacter phage Yang Arthobacter virus Yang Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43206 | 68.389 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048096 | Arthrobacter phage Peas | Arthrobacter phage Peas Arthrobacter virus Peas Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44103 | 64.712 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048095 | Arthrobacter phage Judy | Arthrobacter phage Judy Arthrobacter virus Judy Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43440 | 65.168 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048094 | Arthrobacter phage Eileen | Arthrobacter phage Eileen Arthrobacter virus Eileen Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41165 | 65.126 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048093 | Arthrobacter phage DrManhattan | Arthrobacter phage DrManhattan Arthrobacter virus DrManhattan Manhattanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42577 | 65.953 | Manhattanvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048092 | Arthrobacter phage Constance | Arthrobacter phage Constance Arthrobacter virus Constance Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43706 | 65.128 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048091 | Arthrobacter phage Bridgette | Arthrobacter phage Bridgette Arthrobacter virus Bridgette Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43113 | 65.089 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048088 | Cronobacter phage CS01 | Cronobacter phage CS01 Cronobacter virus CS01 Gyeonggidovirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48195 | 50.109 | Gyeonggidovirus | Unclassified | Drexlerviridae | Cronobacter |
| NC_048087 | Enterococcus phage EfV12-phi1 | Enterococcus phage EfV12-phi1 Enterococcus virus EfV12 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152770 | 37.02 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_048085 | Lactobacillus phage Bromius | Lactobacillus phage Bromius Lactobacillus virus Bromius Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140527 | 36.525 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048084 | Lactobacillus phage Iacchus | Lactobacillus phage Iacchus Lactobacillus virus Iacchus Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137973 | 36.505 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048083 | Gordonia phage Getalong | Gordonia phage Getalong Gordonia virus Getalong Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56214 | 62.821 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_048082 | Enterobacteria phage vB_EcoP_IME390 | Enterobacteria phage vB_EcoP_IME390 Enterobacteria virus IME390 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38396 | 49.065 | Teseptimavirus | Studiervirinae | Autographiviridae | Enterobacteria |
| NC_048079 | Escherichia phage C5 | Escherichia phage C5 Escherichia virus C5 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39551 | 48.815 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048078 | Escherichia phage N30 | Escherichia phage N30 Escherichia virus N30 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39266 | 48.615 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048072 | Streptomyces phage Darolandstone | Streptomyces phage Darolandstone Streptomyces virus Darolandstone Raleighvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40725 | 72.309 | Raleighvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048071 | Escherichia phage IMM-002 | Escherichia phage IMM-002 Escherichia virus IMM002 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40060 | 53.19 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048070 | Corynebacterium phage Juicebox | Corynebacterium phage Juicebox Corynebacterium virus Juicebox Juiceboxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41394 | 67.858 | Juiceboxvirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048069 | Corynebacterium phage SamW | Corynebacterium phage SamW Corynebacterium virus SamW Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 68.134 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_048066 | Vibrio phage 1.202.O._10N.222.45.E8 | Vibrio phage 1.202.O._10N.222.45.E8 Vibrio virus Canoe Canoevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32014 | 44.559 | Canoevirus | Peduovirinae | Myoviridae | Vibrio |
| NC_048065 | Citrobacter phage Sazh | Citrobacter phage Sazh Citrobacter virus Sazh Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49665 | 42.811 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| NC_048064 | Escherichia phage LL11 | Escherichia phage LL11 Escherichia virus LL11 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45185 | 45.072 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_048063 | Escherichia phage LL2 | Escherichia phage LL2 Escherichia virus LL2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39382 | 50.34 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048055 | Curvibacter phage P26059B | Curvibacter phage P26059B Curvibacter virus P26059B Kalppathivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41344 | 54.252 | Kalppathivirus | Unclassified | Autographiviridae | Curvibacter |
| NC_048049 | Synechococcus phage S-T4 | Synechococcus phage S-T4 Synechococcus virus ST4 Tamkungvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181082 | 38.927 | Tamkungvirus | Unclassified | Myoviridae | Synechococcus |
| NC_048044 | Klebsiella phage NJR15 | Klebsiella phage NJR15 Klebsiella virus NJR15 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49468 | 51.043 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048043 | Klebsiella phage NJS2 | Klebsiella phage NJS2 Klebsiella virus NJS2 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50132 | 50.786 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048042 | Klebsiella phage TAH8 | Klebsiella phage TAH8 Klebsiella virus TAH8 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49344 | 51.092 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_048040 | Gordonia phage Phistory | Gordonia phage Phistory Gordonia virus Phistory Phistoryvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53085 | 62.978 | Phistoryvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_048039 | Gordonia phage Horus | Gordonia phage Horus Gordonia virus Horus Horusvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55668 | 62.957 | Horusvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_048026 | Synechococcus T7-like phage S-TIP37 | Synechococcus T7-like phage S-TIP37 Synechococcus virus STIP37 Igirivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45477 | 50.483 | Igirivirus | Unclassified | Autographiviridae | Synechococcus |
| NC_048025 | Shigella phage SFPH2 | Shigella phage SFPH2 Shigella virus SFPH2 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40387 | 52.41 | Kayfunavirus | Studiervirinae | Autographiviridae | Shigella |
| NC_048023 | Escherichia virus Vec13 | Escherichia virus Vec13 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39747 | 50.17 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_048020 | Microbacterium phage KaiHaiDragon | Microbacterium phage KaiHaiDragon Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52992 | 68.901 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048019 | Gordonia phage Ruthy | Gordonia phage Ruthy Gordonia virus Ruthy Ruthyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51265 | 66.681 | Ruthyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048018 | Gordonia phage Apricot | Gordonia phage Apricot Gordonia virus Apricot Apricotvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52195 | 63.06 | Apricotvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048015 | Cyanophage S-TIM4 | Cyanophage S-TIM4 Synechococcus virus STIM4 Thaumasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176495 | 37.657 | Thaumasvirus | Unclassified | Myoviridae | Synechococcus |
| NC_048014 | Klebsiella phage vB_KpnP_IME321 | Klebsiella phage vB_KpnP_IME321 Klebsiella virus IME321 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39906 | 52.767 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_048004 | Salmonella phage 3A_8767 | Salmonella phage 3A_8767 Salmonella virus 3A8767 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38821 | 49.254 | Teseptimavirus | Studiervirinae | Autographiviridae | Salmonella |
| NC_047998 | Shigella phage Sfin-1 | Shigella phage Sfin-1 Shigella virus Sfin1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50403 | 45.202 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_047996 | Escherichia phage Eco_BIFF | Escherichia phage Eco_BIFF Escherichia virus BIFF Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49372 | 45.258 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_047994 | Klebsiella phage ZCKP1 | Klebsiella phage ZCKP1 Klebsiella virus ZCKP1 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150925 | 39.102 | Phapecoctavirus | Unclassified | Myoviridae | Klebsiella |
| NC_047993 | Microbacterium phage Quhwah | Microbacterium phage Quhwah Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53549 | 68.847 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047989 | Microbacterium phage RobsFeet | Microbacterium phage RobsFeet Microbacterium virus RobsFeet Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54189 | 68.602 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047988 | Microbacterium phage Metamorphoo | Microbacterium phage Metamorphoo Microbacterium virus Metamorphoo Metamorphoovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54148 | 68.527 | Metamorphoovirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047987 | Microbacterium phage MementoMori | Microbacterium phage MementoMori Microbacterium virus MementoMori Mementomorivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55572 | 69.978 | Mementomorivirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047985 | Escherichia phage LL5 | Escherichia phage LL5 Escherichia virus LL5 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49788 | 42.464 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_047984 | Escherichia phage vB_EcoP_S523 | Escherichia phage vB_EcoP_S523 Escherichia virus S523 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39825 | 48.806 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047983 | Lactobacillus phage Lb | Lactobacillus phage Lb Lactobacillus virus Lb Heilongjiangvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43416 | 41.738 | Heilongjiangvirus | Unclassified | Myoviridae | Lactobacillus |
| NC_047980 | Klebsiella phage SH-Kp 152234 | Klebsiella phage SH-Kp 152234 Klebsiella virus SHKp152234 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40578 | 52.851 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047976 | Microbacterium phage Paschalis | Microbacterium phage Paschalis Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52935 | 68.837 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047974 | Gordonia phage Jace | Gordonia phage Jace Gordonia virus Jace Jacevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53912 | 66.981 | Jacevirus | Unclassified | Siphoviridae | Gordonia |
| NC_047972 | Microbacterium phage Appa | Microbacterium phage Appa Microbacterium virus Appa Appavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38684 | 68.041 | Appavirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047971 | Klebsiella virus KP32 | Klebsiella virus KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40337 | 53.023 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047970 | Klebsiella virus KP32 | Klebsiella virus KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40540 | 52.753 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047969 | Klebsiella virus KP32 | Klebsiella virus KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41161 | 52.836 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047968 | Klebsiella virus KP32 | Klebsiella virus KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40635 | 53.203 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047967 | Pseudomonas phage vB_PaeP_PAO1_1-15pyo | Pseudomonas phage vB_PaeP_PAO1_1-15pyo Pseudomonas virus 15pyo Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42750 | 62.18 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_047960 | Escherichia phage vB_EcoS_IME347 | Escherichia phage vB_EcoS_IME347 Escherichia virus IME347 Badaguanvirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50048 | 49.706 | Badaguanvirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_047959 | Enterobacteria phage vB_EcoS_IME18 | Enterobacteria phage vB_EcoS_IME18 Escherichia virus IME18 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50354 | 45.615 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_047958 | Burkholderia phage vB_BmuP_KL4 | Burkholderia phage vB_BmuP_KL4 Burkholderia virus KL4 Kelquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42250 | 63.183 | Kelquatrovirus | Unclassified | Podoviridae | Burkholderia |
| NC_047953 | Pseudomonas phage vB_PaeP_130_113 | Pseudomonas phage vB_PaeP_130_113 Pseudomonas virus 130-113 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44205 | 62.357 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_047952 | Pseudomonas phage PAXYB1 | Pseudomonas phage PAXYB1 Pseudomonas virus PAXYB1 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43337 | 62.289 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_047948 | Staphylococcus phage phiSA_BS2 | Staphylococcus phage phiSA_BS2 Staphylococcus virus BS2 Baoshanvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149229 | 29.608 | Baoshanvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047947 | Salmonella phage vB_SenS_PHB07 | Salmonella phage vB_SenS_PHB07 Salmonella virus PHB07 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51818 | 43.359 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_047945 | Staphylococcus phage phiSA_BS1 | Staphylococcus phage phiSA_BS1 Staphylococcus virus BS1 Baoshanvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142978 | 29.799 | Baoshanvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047944 | Klebsiella phage myPSH1235 | Klebsiella phage myPSH1235 Klebsiella virus myPSH1235 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45135 | 53.663 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047943 | Haloarcula hispanica pleomorphic virus 4 | Haloarcula hispanica pleomorphic virus 4 Betapleolipovirus HHPV4 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 15010 | 54.63 | Betapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| NC_047942 | Escherichia phage Ebrios | Escherichia phage Ebrios Escherichia virus Ebrios Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39752 | 52.853 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047940 | Yersinia phage YpsP-G | Yersinia phage YpsP-G Yersinia virus YpsPG Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38288 | 48.221 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047939 | Yersinia phage YpP-Y | Yersinia phage YpP-Y Yersinia virus YpPY Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37714 | 48.348 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047938 | Escherichia phage SECphi27 | Escherichia phage SECphi27 Escherichia virus SECphi27 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51811 | 44.691 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_047937 | Yersinia phage fPS-54-ocr | Yersinia phage fPS-54-ocr Yersinia virus fPS54ocr Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40075 | 45.535 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047936 | Yersinia phage fPS-53 | Yersinia phage fPS-53 Yersinia virus fPS53 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40694 | 45.461 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047935 | Yersinia phage fPS-59 | Yersinia phage fPS-59 Yersinia virus fPS59 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38391 | 45.714 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047934 | Yersinia phage fPS-9 | Yersinia phage fPS-9 Yersinia virus fPS9 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39034 | 45.604 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047933 | Pseudomonas phage phiNV3 | Pseudomonas phage phiNV3 Pseudomonas virus NV3 Kirikabuvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43184 | 58.29 | Kirikabuvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_047929 | Shigella phage SFN6B | Shigella phage SFN6B Shigella virus SFN6B Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43036 | 50.943 | Drulisvirus | Slopekvirinae | Autographiviridae | Shigella |
| NC_047928 | Haloarcula hispanica pleomorphic virus 3 | Haloarcula hispanica pleomorphic virus 3 Betapleolipovirus HHPV3 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 11648 | 56.104 | Betapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| NC_047927 | Escherichia phage YZ1 | Escherichia phage YZ1 Escherichia virus YZ1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38823 | 50.375 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047925 | Lactobacillus phage Nyseid | Lactobacillus phage Nyseid Lactobacillus virus Nyseid Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79221 | 37.067 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047924 | Lactobacillus phage Bacchae | Lactobacillus phage Bacchae Lactobacillus virus Bacchae Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141124 | 36.346 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_047923 | Escherichia phage HZ2R8 | Escherichia phage HZ2R8 Escherichia virus HZ2R8 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39482 | 48.615 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047922 | Pseudomonas phage Henninger | Pseudomonas phage Henninger Pseudomonas virus Henninger Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40923 | 57.586 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047920 | Citrobacter phage vB_CroP_CrRp3 | Citrobacter phage vB_CroP_CrRp3 Citrobacter CrRp3 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44349 | 45.155 | Vectrevirus | Molineuxvirinae | Autographiviridae | Citrobacter |
| NC_047919 | Staphylococcus phage vB_SauP_phiAGO1.3 | Staphylococcus phage vB_SauP_phiAGO1.3 Staphylococcus virus phiAGO13 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17603 | 28.955 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_047916 | Faecalibacterium phage FP_oengus | Faecalibacterium phage FP_oengus Faecalibacterium virus Oengus Oengusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58391 | 49.794 | Oengusvirus | Unclassified | Siphoviridae | Faecalibacterium |
| NC_047915 | Faecalibacterium phage FP_Toutatis | Faecalibacterium phage FP_Toutatis Faecalibacterium virus Toutatis Toutatisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54725 | 53.906 | Toutatisvirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047914 | Faecalibacterium phage FP_Taranis | Faecalibacterium phage FP_Taranis Faecalibacterium virus Taranis Taranisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54286 | 55.799 | Taranisvirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047913 | Faecalibacterium phage FP_Mushu | Faecalibacterium phage FP_Mushu Faecalibacterium virus Mushu Mushuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36636 | 60.31 | Mushuvirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047912 | Faecalibacterium phage FP_Lugh | Faecalibacterium phage FP_Lugh Faecalibacterium virus Lugh Lughvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34075 | 59.894 | Lughvirus | Unclassified | Siphoviridae | Faecalibacterium |
| NC_047911 | Faecalibacterium phage FP_Lagaffe | Faecalibacterium phage FP_Lagaffe Faecalibacterium virus Lagaffe Lagaffevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48211 | 55.906 | Lagaffevirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047910 | Faecalibacterium phage FP_Epona | Faecalibacterium phage FP_Epona Faecalibacterium virus Epona Eponavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49573 | 57.092 | Eponavirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047909 | Faecalibacterium phage FP_Brigit | Faecalibacterium phage FP_Brigit Faecalibacterium virus Brigit Brigitvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61716 | 55.796 | Brigitvirus | Unclassified | Myoviridae | Faecalibacterium |
| NC_047908 | Klebsiella phage SH-Kp 152410 | Klebsiella phage SH-Kp 152410 Klebsiella virus SHKp152410 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40945 | 52.346 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047907 | Klebsiella phage GML-KpCol1 | Klebsiella phage GML-KpCol1 Klebsiella virus KpCol1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50249 | 50.97 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_047906 | Streptomyces phage Rowa | Streptomyces phage Rowa Streptomyces virus Rowa Rowavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42890 | 61.203 | Rowavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047905 | Streptomyces phage Attoomi | Streptomyces phage Attoomi Streptomyces virus Attoomi Attoomivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41872 | 69.689 | Attoomivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047904 | Streptomyces phage AbbeyMikolon | Streptomyces phage AbbeyMikolon Streptomyces virus AbbeyMikolon Abbeymikolonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42551 | 66.826 | Abbeymikolonvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047899 | Escherichia virus VEc3 | Escherichia virus VEc3 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44895 | 44.972 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047898 | Salmonella phage YSP2 | Salmonella phage YSP2 Salmonella virus YSP2 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50316 | 42.873 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_047897 | Lactobacillus phage Lenus | Lactobacillus phage Lenus Lactobacillus virus Lenus Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78891 | 37.204 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047895 | Escherichia phage EG1 | Escherichia phage EG1 Escherichia virus EG1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39919 | 48.461 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047893 | Escherichia phage DTL | Escherichia phage DTL Escherichia virus DTL Loudonvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45814 | 44.251 | Loudonvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_047891 | Gordonia phage BENtherdunthat | Gordonia phage BENtherdunthat Gordonia virus BENtherdunthat Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54867 | 63.364 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| NC_047889 | Escherichia virus mutPK1A2 | Escherichia virus mutPK1A2 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44784 | 45.067 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047886 | Pectobacterium phage DU_PP_II | Pectobacterium phage DU_PP_II Pectobacterium virus DUPPII Unyawovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40142 | 47.917 | Unyawovirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_047875 | Salmonella phage UPF_BP1 | Salmonella phage UPF_BP1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39902 | 47.559 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_047874 | Pseudomonas virus Pf1 ERZ-2017 | Pseudomonas virus Pf1 ERZ-2017 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39195 | 57.199 | Teseptimavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047871 | Listeria phage LMTA-57 | Listeria phage LMTA-57 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136589 | 35.848 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_047867 | Enterobacteria phage T7M | Enterobacteria phage T7M Enterobacteria virus T7M Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38202 | 49.906 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacteria |
| NC_047864 | Enterobacteria phage T3 | Enterobacteria phage T3 Escherichia virus T3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38209 | 49.918 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047862 | Klebsiella phage vB_KpnS_IME279 | Klebsiella phage vB_KpnS_IME279 Klebsiella virus IME279 Sortsnevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42518 | 59.307 | Sortsnevirus | Unclassified | Podoviridae | Klebsiella |
| NC_047861 | Bordetella phage vB_BbrM_PHB04 | Bordetella phage vB_BbrM_PHB04 Bordetella virus PHB04 Phabquatrovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94005 | 65.464 | Phabquatrovirus | Unclassified | Myoviridae | Bordetella |
| NC_047858 | Proteus phage PM 116 | Proteus phage PM 116 Proteus virus PM116 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44601 | 39.217 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| NC_047857 | Klebsiella phage phiKpS2 | Klebsiella phage phiKpS2 Klebsiella virus KpS2 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44024 | 53.998 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047855 | Staphylococcus phage PSa3 | Staphylococcus phage PSa3 Staphylococcus virus PSa3 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17602 | 29.588 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_047854 | Cronobacter phage ESSI-2 | Cronobacter phage ESSI-2 Cronobacter virus ESSI2 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28765 | 55.168 | Seongnamvirus | Peduovirinae | Myoviridae | Cronobacter |
| NC_047851 | Brevibacterium phage LuckyBarnes | Brevibacterium phage LuckyBarnes Brevibacterium virus LuckyBarnes Luckybarnesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50774 | 61.933 | Luckybarnesvirus | Unclassified | Siphoviridae | Brevibacterium |
| NC_047850 | Klebsiella phage MezzoGao | Klebsiella phage MezzoGao Klebsiella virus MezzoGao Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49807 | 50.977 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_047849 | Klebsiella phage AltoGao | Klebsiella phage AltoGao Klebsiella virus AltoGao Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43012 | 53.292 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047848 | Shigella phage Sf12 | Shigella phage Sf12 Shigella virus Sf12 Eastlansingvirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47647 | 44.328 | Eastlansingvirus | Rogunavirinae | Drexlerviridae | Shigella |
| NC_047847 | Shigella phage Sd1 | Shigella phage Sd1 Shigella virus Sd1 Wilsonroadvirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48262 | 44.499 | Wilsonroadvirus | Rogunavirinae | Drexlerviridae | Shigella |
| NC_047844 | Serratia phage 2050H2 | Serratia phage 2050H2 Serratia virus 2050H2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39216 | 50.418 | Teetrevirus | Studiervirinae | Autographiviridae | Serratia |
| NC_047843 | Leclercia phage 10164-302 | Leclercia phage 10164-302 Leclercia virus 10164-302 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39064 | 50.819 | Teetrevirus | Studiervirinae | Autographiviridae | Leclercia |
| NC_047842 | Klebsiella phage 2044-307w | Klebsiella phage 2044-307w Klebsiella virus 2044-307w Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40048 | 52.892 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047841 | Klebsiella phage KPN N141 | Klebsiella phage KPN N141 Klebsiella virus KPN N141 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49090 | 50.959 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_047838 | Synechococcus phage Bellamy | Synechococcus phage Bellamy Synechococcus virus Bellamy Bellamyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 204930 | 41.105 | Bellamyvirus | Unclassified | Myoviridae | Synechococcus |
| NC_047833 | Erwinia phage EtG | Erwinia phage EtG Erwinia virus EtG Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30413 | 54.138 | Eganvirus | Peduovirinae | Myoviridae | Erwinia |
| NC_047830 | Escherichia phage ST32 | Escherichia phage ST32 Escherichia virus ST32 Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53092 | 44.137 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| NC_047829 | Escherichia phage ST31 | Escherichia phage ST31 Escherichia virus ST31 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39693 | 49.989 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047828 | Escherichia phage vB_EcoS_SH2 | Escherichia phage vB_EcoS_SH2 Escherichia virus SH2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49088 | 45.475 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_047827 | Pseudomonas virus KNP | Pseudomonas virus KNP Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40491 | 57.294 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047826 | Pseudomonas virus WRT | Pseudomonas virus WRT Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40214 | 57.42 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047825 | Klebsiella phage KOX1 | Klebsiella phage KOX1 Klebsiella virus KOX1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50526 | 51.237 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_047823 | Citrobacter phage CF1 DK-2017 | Citrobacter phage CF1 DK-2017 Citrobacter virus DK2017 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50339 | 42.647 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| NC_047820 | Escherichia phage vB_EcoS_ESCO41 | Escherichia phage vB_EcoS_ESCO41 Escherichia virus ESCO41 Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50800 | 46.085 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| NC_047818 | Klebsiella phage 4LV2017 | Klebsiella phage 4LV2017 Klebsiella virus 4LV2017 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33540 | 50.403 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_047817 | Klebsiella phage 3LV2017 | Klebsiella phage 3LV2017 Klebsiella virus 3LV2017 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35100 | 54.43 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| NC_047816 | Ralstonia phage RS-PI-1 | Ralstonia phage RS-PI-1 Ralstonia virus RSPI1 Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43211 | 61.482 | Ampunavirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_047814 | Staphylococcus phage St 134 | Staphylococcus phage St 134 Staphylococcus virus St134 Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18275 | 30.096 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_047813 | Staphylococcus phage Andhra | Staphylococcus phage Andhra Staphylococcus virus Andhra Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18546 | 29.823 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_047812 | Escherichia phage vB_EcoS_CEB_EC3a | Escherichia phage vB_EcoS_CEB_EC3a Escherichia virus EC3a Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44234 | 44.217 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_047811 | Klebsiella phage vB_KpnP_KpV74 | Klebsiella phage vB_KpnP_KpV74 Klebsiella virus KpV74 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44094 | 54.141 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047810 | Escherichia phage vB_EcoS-IME253 | Escherichia phage vB_EcoS-IME253 Escherichia virus IME253 Rtpvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46717 | 44.215 | Rtpvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_047809 | Escherichia phage vB_EcoP_K | Escherichia phage vB_EcoP_K Escherichia virus K Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37775 | 45.117 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047808 | Escherichia phage vB_EcoP_F | Escherichia phage vB_EcoP_F Escherichia virus F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39300 | 49.954 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047807 | Escherichia phage vB_EcoP_C | Escherichia phage vB_EcoP_C Escherichia virus C Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44970 | 44.966 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047806 | Escherichia phage vB_EcoP_B | Escherichia phage vB_EcoP_B Escherichia virus B Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44018 | 45.036 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047804 | Ralstonia phage RS-PII-1 | Ralstonia phage RS-PII-1 Ralstonia virus RSPII1 Sukuvirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42042 | 63.177 | Sukuvirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_047796 | Enterococcus phage EFP01 | Enterococcus phage EFP01 Enterococcus virus EFP01 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155053 | 37.018 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_047795 | Streptomyces phage Raleigh | Streptomyces phage Raleigh Streptomyces virus Raleigh Raleighvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40785 | 71.801 | Raleighvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047794 | Streptomyces phage Picard | Streptomyces phage Picard Streptomyces virus Picard Picardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39522 | 72.605 | Picardvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047793 | Streptomyces phage PapayaSalad | Streptomyces phage PapayaSalad Streptomyces virus PapayaSalad Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38411 | 72.635 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047792 | Streptomyces phage Ididsumtinwong | Streptomyces phage Ididsumtinwong Streptomyces virus Ididsumtinwong Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37817 | 72.367 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_047790 | Pseudoalteromonas phage C5a | Pseudoalteromonas phage C5a Pseudoalteromonas virus C5a Catalunyavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35209 | 42.231 | Catalunyavirus | Peduovirinae | Myoviridae | Pseudoalteromonas |
| NC_047789 | Pectobacterium phage PP74 | Pectobacterium phage PP74 Pectobacterium virus PP74 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39790 | 48.635 | Berlinvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_047788 | Staphylococcus phage SCH1 | Staphylococcus phage SCH1 Staphylococcus virus SCH1 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18023 | 29.34 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_047787 | Klebsiella phage KPV811 | Klebsiella phage KPV811 Klebsiella virus KPV811 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42641 | 51.347 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047786 | Salmonella phage vB_SenS_Sasha | Salmonella phage vB_SenS_Sasha Salmonella virus Sasha Sashavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53263 | 43.732 | Sashavirus | Unclassified | Siphoviridae | Salmonella |
| NC_047785 | Shigella phage SH6 | Shigella phage SH6 Shigella virus SH6 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.828 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_047784 | Klebsiella phage vB_KpnS_KpV522 | Klebsiella phage vB_KpnS_KpV522 Klebsiella virus KpV522 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51099 | 50.848 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_047783 | Klebsiella phage vB_KpnP_KpV48 | Klebsiella phage vB_KpnP_KpV48 Klebsiella virus KpV48 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44623 | 54.129 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047782 | Klebsiella phage vB_KpnP_IL33 | Klebsiella phage vB_KpnP_IL33 Klebsiella virus IL33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41335 | 52.495 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047781 | Klebsiella phage vB_KpnP_BIS33 | Klebsiella phage vB_KpnP_BIS33 Klebsiella virus BIS33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41697 | 52.704 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047780 | Klebsiella phage vB_KpnP_PRA33 | Klebsiella phage vB_KpnP_PRA33 Klebsiella virus PRA33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40605 | 52.516 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047779 | Klebsiella phage KP-Rio/2015 | Klebsiella phage KP-Rio/2015 Klebsiella virus KPRio2015 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43557 | 54.134 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047777 | Escherichia phage ZG49 | Escherichia phage ZG49 Escherichia virus ZG49 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40291 | 49.815 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047775 | Salinivibrio phage SMHB1 | Salinivibrio phage SMHB1 Salinivibrio virus SMHB1 Playavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32826 | 50.795 | Playavirus | Peduovirinae | Myoviridae | Salinivibrio |
| NC_047774 | Serratia phage SM9-3Y | Serratia phage SM9-3Y Serratia virus SM9-3Y Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39631 | 50.741 | Teetrevirus | Studiervirinae | Autographiviridae | Serratia |
| NC_047773 | Klebsiella phage vB_KpnP_KpV766 | Klebsiella phage vB_KpnP_KpV766 Klebsiella virus KpV766 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41283 | 52.583 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047772 | Klebsiella phage vB_KpnP_KpV767 | Klebsiella phage vB_KpnP_KpV767 Klebsiella virus KpV767 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40395 | 52.281 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047771 | Klebsiella phage vB_KpnP_KpV763 | Klebsiella phage vB_KpnP_KpV763 Klebsiella virus KpV763 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40765 | 53.222 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047769 | Escherichia virus AAPEc6 | Escherichia virus AAPEc6 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44559 | 45.176 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_047767 | Staphylococcus phage vB_SscM-1 | Staphylococcus phage vB_SscM-1 Staphylococcus virus SscM1 Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139681 | 31.827 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047766 | Escherichia phage ECA2 | Escherichia phage ECA2 Escherichia virus ECA2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38890 | 50.463 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047763 | Streptococcus virus 9873 | Streptococcus virus 9873 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32813 | 36.9 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_047761 | Klebsiella phage vB_KpnP_IME205 | Klebsiella phage vB_KpnP_IME205 Klebsiella virus IME205 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41310 | 52.213 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047760 | Lactobacillus phage SA-C12 | Lactobacillus phage SA-C12 Lactobacillus virus SAC12 Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79099 | 37.491 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047755 | Yersinia phage phiYe-F10 | Yersinia phage phiYe-F10 Yersinia virus YeF10 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39210 | 50.699 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_047753 | Salmonella phage SEN8 | Salmonella phage SEN8 Salmonella virus SEN8 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35203 | 51.936 | Felsduovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_047752 | Burkholderia phage AP3 | Burkholderia phage AP3 Burkholderia virus AP3 Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36499 | 64.407 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_047750 | Mannheimia phage vB_MhM_1127AP1 | Mannheimia phage vB_MhM_1127AP1 Mannheimia virus 1127AP1 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36745 | 41.984 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| NC_047748 | Klebsiella phage phiBO1E | Klebsiella phage phiBO1E Klebsiella virus BO1E Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43865 | 53.852 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_047747 | Pseudomonas phage phiPsa17 | Pseudomonas phage phiPsa17 Pseudomonas virus PhiPsa17 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40525 | 57.291 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047744 | Bacillus phage Bp8p-T | Bacillus phage Bp8p-T Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151419 | 41.154 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_047743 | Burkholderia phage Bp-AMP1 | Burkholderia phage Bp-AMP1 Burkholderia virus BpAMP1 Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42409 | 61.749 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| NC_047742 | Enterobacteria phage IME_EC2 | Enterobacteria phage IME_EC2 Escherichia virus EC2 Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41510 | 59.157 | Murrayvirus | Unclassified | Siphoviridae | Escherichia |
| NC_047740 | Enterobacteria phage phiJLA23 | Enterobacteria phage phiJLA23 Escherichia virus phiJLA23 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43017 | 44.19 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_047739 | Lactobacillus phage ATCC 8014-B2 | Lactobacillus phage ATCC 8014-B2 Lactobacillus virus B2 Douglaswolinvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80617 | 36.962 | Douglaswolinvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047738 | Rhizobium phage RHEph01 | Rhizobium phage RHEph01 Rhizobium virus RHEph01 Paadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43444 | 59.177 | Paadamvirus | Unclassified | Autographiviridae | Rhizobium |
| NC_047736 | Bacillus phage B5S | Bacillus phage B5S Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162598 | 37.71 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_047735 | Bacillus phage BCU4 | Bacillus phage BCU4 Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154371 | 39.857 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_047734 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197450 | 40.367 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| NC_047733 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171216 | 40.63 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| NC_047732 | Staphylococcus phage IME-SA119 | Staphylococcus phage IME-SA119 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141028 | 30.326 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047731 | Staphylococcus phage IME-SA118 | Staphylococcus phage IME-SA118 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139750 | 30.323 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047730 | Staphylococcus phage IME-SA2 | Staphylococcus phage IME-SA2 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140906 | 30.328 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047729 | Staphylococcus phage IME-SA1 | Staphylococcus phage IME-SA1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140218 | 30.328 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047728 | Staphylococcus phage A5W | Staphylococcus phage A5W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145542 | 30.416 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047727 | Staphylococcus phage P4W | Staphylococcus phage P4W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147590 | 30.392 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047726 | Staphylococcus phage MSA6 | Staphylococcus phage MSA6 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148243 | 30.243 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047725 | Staphylococcus phage Fi200W | Staphylococcus phage Fi200W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148481 | 30.393 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047724 | Staphylococcus phage 676Z | Staphylococcus phage 676Z Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148564 | 30.405 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047723 | Staphylococcus phage A3R | Staphylococcus phage A3R Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141018 | 30.498 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047722 | Staphylococcus phage Staph1N | Staphylococcus phage Staph1N Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145647 | 30.412 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047721 | Staphylococcus phage SA5 | Staphylococcus phage SA5 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137031 | 30.421 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047720 | Staphylococcus phage ISP | Staphylococcus phage ISP Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138339 | 30.422 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_047719 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172874 | 39.112 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| NC_047718 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154990 | 39.226 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| NC_047717 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175559 | 39.519 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| NC_047716 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175557 | 39.526 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| NC_047715 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174670 | 39.513 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| NC_047714 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 196842 | 41.536 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| NC_047713 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223270 | 41.607 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| NC_047712 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216628 | 41.575 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| NC_007636 | Escherichia phage K1F | Escherichia phage K1F Escherichia virus K1F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39699 | 49.777 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_004927 | Haloferax virus HF1 | Haloferax virus HF1 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75898 | 55.793 | Haloferacalesvirus | Unclassified | Myoviridae | Haloferax |
| NC_003345 | Halorubrum phage HF2 | Halorubrum phage HF2 Halorubrum virus HF2 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77672 | 55.849 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| NC_044940 | Pectinobacterium phage PEAT2 | Pectinobacterium phage PEAT2 Pectinobacterium virus PEAT2 Peatvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48659 | 49.126 | Peatvirus | Unclassified | Myoviridae | Pectobacterium |
| NC_031005 | Propionibacterium phage QueenBey | Propionibacterium phage QueenBey unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29338 | 54.145 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027623 | Propionibacterium phage Pirate | Propionibacterium phage Pirate Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29324 | 54.085 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_019524 | Enterobacter phage Enc34 | Enterobacter phage Enc34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60496 | 51.084 | Unclassified | Unclassified | Siphoviridae | Enterobacter |
| NC_042346 | Salmonella virus BTP1 | Salmonella virus BTP1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40525 | 47.45 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_042343 | Pseudomonas phage PaP4 | Pseudomonas phage PaP4 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43895 | 52.464 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_042342 | Pseudomonas virus H66 | Pseudomonas virus H66 Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65270 | 62.628 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_042341 | Mycobacterium virus Rebeuca | Mycobacterium virus Rebeuca Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51235 | 65.057 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042340 | Mycobacterium virus Goose | Mycobacterium virus Goose Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50645 | 65.138 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042339 | Mycobacterium phage Arturo | Mycobacterium phage Arturo Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51500 | 64.052 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042337 | Mycobacterium phage Astro | Mycobacterium phage Astro Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52494 | 61.38 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042336 | Mycobacterium virus DotProduct | Mycobacterium virus DotProduct Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55363 | 61.752 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042335 | Mycobacterium virus Saintus | Mycobacterium virus Saintus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49228 | 61.235 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042334 | Mycobacterium virus Jeffabunny | Mycobacterium virus Jeffabunny Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48963 | 61.573 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042333 | Mycobacterium virus Cuco | Mycobacterium virus Cuco Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50965 | 60.857 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042331 | Mycobacterium phage Benedict | Mycobacterium phage Benedict Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 59.797 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042330 | Mycobacterium virus Ericb | Mycobacterium virus Ericb Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51702 | 61.533 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042329 | Mycobacterium virus Mrgordo | Mycobacterium virus Mrgordo Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50988 | 63.752 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042328 | Mycobacterium virus Heldan | Mycobacterium virus Heldan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50364 | 63.984 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042327 | Mycobacterium phage BPBiebs31 | Mycobacterium phage BPBiebs31 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53171 | 63.448 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042326 | Mycobacterium virus Kssjeb | Mycobacterium virus Kssjeb Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51381 | 63.631 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042324 | Mycobacterium virus Museum | Mycobacterium virus Museum Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51426 | 63.637 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042323 | Mycobacterium virus Mozy | Mycobacterium virus Mozy Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57278 | 61.055 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042322 | Mycobacterium virus Littlee | Mycobacterium virus Littlee Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109086 | 61.345 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042321 | Mycobacterium virus Lesedi | Mycobacterium virus Lesedi Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50486 | 63.76 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042320 | Mycobacterium virus JC27 | Mycobacterium virus JC27 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52169 | 63.639 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042319 | Mycobacterium virus Ibhubesi | Mycobacterium virus Ibhubesi Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55600 | 61.239 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042318 | Mycobacterium virus Hammer | Mycobacterium virus Hammer Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51889 | 61.331 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042317 | Mycobacterium virus Dlane | Mycobacterium virus Dlane Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58899 | 61.884 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042316 | Mycobacterium phage Baka | Mycobacterium phage Baka Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111688 | 60.732 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042315 | Mycobacterium phage RockyHorror | Mycobacterium phage RockyHorror Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56719 | 61.128 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042314 | Mycobacterium phage PackMan | Mycobacterium phage PackMan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51339 | 62.576 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042313 | Mycobacterium phage JoeDirt | Mycobacterium phage JoeDirt Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74914 | 58.781 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042312 | Mycobacterium phage George | Mycobacterium phage George Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51578 | 60.954 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042311 | Mycobacterium phage Microwolf | Mycobacterium phage Microwolf Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50864 | 64.041 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042310 | Mycobacterium phage JHC117 | Mycobacterium phage JHC117 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 63.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042309 | Mycobacterium phage Gladiator | Mycobacterium phage Gladiator Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52213 | 61.427 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042308 | Mycobacterium phage Backyardigan | Mycobacterium phage Backyardigan Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51308 | 63.688 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042140 | Bacillus virus BM15 | Bacillus virus BM15 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165213 | 39.605 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_042135 | Pseudomonas phage JBD67 | Pseudomonas phage JBD67 Pseudomonas virus JBD67 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38232 | 63.018 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_042133 | Bacillus phage SIOphi | Bacillus phage SIOphi Bacillus virus SIOphi Siophivirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146698 | 39.019 | Siophivirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_042130 | Escherichia phage slur04 | Escherichia phage slur04 Escherichia virus slur04 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167298 | 35.505 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_042129 | Escherichia phage slur03 | Escherichia phage slur03 Escherichia virus slur03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167467 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_042128 | Escherichia phage RCS47 | Escherichia phage RCS47 Escherichia virus RCS47 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115154 | 49.35 | Punavirus | Unclassified | Myoviridae | Escherichia |
| NC_042123 | Gordonia phage Waits | Gordonia phage Waits Gordonia virus Waits Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52856 | 61.972 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_042121 | Pseudoalteromonas phage Cr39582 | Pseudoalteromonas phage Cr39582 Pseudoalteromonas virus Cr39582 Corticovirus Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10584 | 41.903 | Corticovirus | Unclassified | Corticoviridae | Pseudoalteromonas |
| NC_042116 | Yersinia phage fHe-Yen9-04 | Yersinia phage fHe-Yen9-04 Yersinia virus Yen9-04 Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 354378 | 31.646 | Eneladusvirus | Unclassified | Myoviridae | Yersinia |
| NC_042114 | Streptococcus phage CP-7 | Streptococcus phage CP-7 Cepunavirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19741 | 38.544 | Cepunavirus | Unclassified | Salasmaviridae | Streptococcus |
| NC_042107 | Pseudomonas phage NV1 | Pseudomonas phage NV1 Pseudomonas virus NV1 Vicosavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45058 | 52.115 | Vicosavirus | Unclassified | Podoviridae | Pseudomonas |
| NC_042106 | Streptomyces phage Bing | Streptomyces phage Bing Streptomyces virus Bing Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56341 | 59.424 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042103 | Pseudomonas phage Bjorn | Pseudomonas phage Bjorn Pseudomonas virus Bjorn Bjornvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45936 | 53.076 | Bjornvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_042097 | Salmonella phage BPS17W1 | Salmonella phage BPS17W1 Salmonella virus BPS17W1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87609 | 38.802 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_042096 | Salmonella phage BPS17L1 | Salmonella phage BPS17L1 Salmonella virus BPS17L1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84916 | 38.861 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_042091 | Pseudomonas phage nickie | Pseudomonas phage nickie Pseudomonas virus nickie Nickievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112225 | 57.36 | Nickievirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_042084 | Escherichia phage VB_EcoS-Golestan | Escherichia phage VB_EcoS-Golestan Escherichia virus Golestan Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44829 | 50.601 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_042081 | Pseudomonas phage SL2 | Pseudomonas phage SL2 Pseudomonas virus SL2 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279696 | 36.9 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042080 | Pseudomonas phage vB_PaeM_E215 | Pseudomonas phage vB_PaeM_E215 Pseudomonas virus E215 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66789 | 55.629 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042079 | Pseudomonas phage vB_PaeM_E217 | Pseudomonas phage vB_PaeM_E217 Pseudomonas virus E217 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66291 | 55.647 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042078 | Shigella phage Sf24 | Shigella phage Sf24 Shigella virus Sf24 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168112 | 35.321 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_042077 | Shigella phage Sf21 | Shigella phage Sf21 Shigella virus Sf21 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166002 | 35.451 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_042076 | Shigella phage Sf17 | Shigella phage Sf17 Shigella virus Sf17 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90092 | 39.032 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| NC_042075 | Shigella phage Sf14 | Shigella phage Sf14 Shigella virus Sf14 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87575 | 39.065 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| NC_042067 | Citrobacter phage CF1 ERZ-2017 | Citrobacter phage CF1 ERZ-2017 Citrobacter virus CF1 Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171911 | 38.741 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_042066 | Salmonella virus SP126 | Salmonella virus SP126 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51092 | 42.868 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_042065 | Salmonella phage FSL SP-101 | Salmonella phage FSL SP-101 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41873 | 50.269 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_042063 | Salmonella phage SP069 | Salmonella phage SP069 Nonanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13598 | 43.506 | Nonanavirus | Unclassified | Siphoviridae | Salmonella |
| NC_042062 | Salmonella phage SP069 | Salmonella phage SP069 Nonanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41391 | 42.475 | Nonanavirus | Unclassified | Siphoviridae | Salmonella |
| NC_042060 | Pseudomonas phage PA7 | Pseudomonas phage PA7 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266743 | 36.933 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042058 | Mycobacterium virus Wonder | Mycobacterium virus Wonder Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48491 | 63.95 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042057 | Escherichia phage DE3 | Escherichia phage DE3 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42925 | 51.884 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_042056 | Erwinia phage Machina | Erwinia phage Machina Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241654 | 51.013 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| NC_042055 | Mycobacterium phage Pipsqueaks | Mycobacterium phage Pipsqueaks Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43679 | 66.277 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_042054 | Pseudomonas phage vB_PsyM_KIL4 | Pseudomonas phage vB_PsyM_KIL4 Pseudomonas virus KIL2 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92816 | 44.897 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042052 | Streptomyces phage Hydra | Streptomyces phage Hydra Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50727 | 66.225 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_042051 | Streptomyces phage Aaronocolus | Streptomyces phage Aaronocolus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49562 | 66.176 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_042048 | Listeria phage LMTA-34 | Listeria phage LMTA-34 Listeria virus LMTA34 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138036 | 35.87 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_042046 | Escherichia phage C119 | Escherichia phage C119 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47319 | 44.238 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_042044 | Salmonella phage Melville | Salmonella phage Melville Salmonella virus Melville Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159323 | 36.947 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_042042 | Streptomyces phage Scap1 | Streptomyces phage Scap1 Streptomyces virus Scap1 Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43060 | 60.876 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042041 | Klebsiella phage vB_KpnM_KpV79 | Klebsiella phage vB_KpnM_KpV79 Klebsiella virus KpV52 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47760 | 48.955 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| NC_042039 | Shigella phage Sf22 | Shigella phage Sf22 Shigella virus Sf22 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166283 | 35.509 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_042036 | Mycobacterium phage Rockstar | Mycobacterium phage Rockstar Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47780 | 64.28 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042030 | Mycobacterium phage Yoshi | Mycobacterium phage Yoshi Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58714 | 61.023 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042028 | Acinetobacter phage AB1 | Acinetobacter phage AB1 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45159 | 37.689 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_042027 | Mycobacterium phage Pumpkin | Mycobacterium phage Pumpkin Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74491 | 62.958 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042025 | Salmonella phage SE1 (in:Nonagvirus) | Salmonella phage SE1 (in:Nonagvirus) Salmonella virus SE1Kor Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60030 | 43.385 | Nonagvirus | Queuovirinae | Siphoviridae | Salmonella |
| NC_042024 | Lactococcus virus P2 | Lactococcus virus P2 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27595 | 34.727 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_042017 | Shigella phage Sf13 | Shigella phage Sf13 Shigella virus Sf13 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87570 | 38.911 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| NC_042012 | Streptomyces phage Samisti12 | Streptomyces phage Samisti12 Streptomyces virus Samisti12 Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133710 | 49.924 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042011 | Streptomyces phage Peebs | Streptomyces phage Peebs Streptomyces virus Peebs Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133047 | 50.057 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042010 | Streptomyces phage Paradiddles | Streptomyces phage Paradiddles Streptomyces virus Paradiddles Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133486 | 50.123 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042009 | Streptomyces phage NootNoot | Streptomyces phage NootNoot Streptomyces virus NootNoot Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131086 | 50.246 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042008 | Streptomyces phage Mildred21 | Streptomyces phage Mildred21 Streptomyces virus Mildred21 Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131976 | 49.469 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_041998 | Acinetobacter phage AbP2 | Acinetobacter phage AbP2 Acinetobacter virus AbP2 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45373 | 37.837 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_041996 | Serratia phage CHI14 | Serratia phage CHI14 Serratia virus CHI14 Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171175 | 38.735 | Winklervirus | Unclassified | Myoviridae | Serratia |
| NC_041995 | Shigella phage vB_SsoS-ISF002 | Shigella phage vB_SsoS-ISF002 Shigella virus ISF002 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50564 | 45.532 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_041992 | Salmonella phage wksl3 | Salmonella phage wksl3 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42633 | 49.797 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_041991 | Salmonella phage vB_SenS_AG11 | Salmonella phage vB_SenS_AG11 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41546 | 49.909 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_041990 | Escherichia phage ST0 | Escherichia phage ST0 Escherichia virus ST0 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170496 | 37.665 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_041989 | Mycobacterium phage Shauna1 | Mycobacterium phage Shauna1 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59315 | 61.708 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041988 | Mycobacterium phage ShiLan | Mycobacterium phage ShiLan Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59794 | 61.446 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041987 | Mycobacterium phage SirDuracell | Mycobacterium phage SirDuracell Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75973 | 62.922 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041986 | Mycobacterium phage Spartacus | Mycobacterium phage Spartacus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61164 | 61.652 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041985 | Mycobacterium phage Switzer | Mycobacterium phage Switzer Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52298 | 63.756 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041984 | Mycobacterium phage Tiger | Mycobacterium phage Tiger Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50332 | 60.665 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041983 | Mycobacterium phage Timshel | Mycobacterium phage Timshel Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53278 | 63.144 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041982 | Mycobacterium phage Twister | Mycobacterium phage Twister Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51094 | 64.957 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041979 | Escherichia typing phage 1 | Escherichia typing phage 1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88531 | 38.849 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_041978 | Erwinia phage vB_Eam-MM7 | Erwinia phage vB_Eam-MM7 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84694 | 43.387 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| NC_041977 | Citrobacter phage Mordin | Citrobacter phage Mordin Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89596 | 38.808 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| NC_041976 | Brevibacillus phage Jimmer2 | Brevibacillus phage Jimmer2 Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54312 | 38.097 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| NC_041971 | Mycobacterium phage Rey | Mycobacterium phage Rey Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83724 | 60.873 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_041970 | Mycobacterium phage Bongo | Mycobacterium phage Bongo Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80228 | 61.621 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_041969 | Mycobacterium phage Jebeks | Mycobacterium phage Jebeks Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45580 | 67.26 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_041966 | Acinetobacter phage WCHABP1 | Acinetobacter phage WCHABP1 Acinetobacter virus WCHABP1 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45888 | 37.626 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_041965 | Mycobacterium phage Athena | Mycobacterium phage Athena Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69409 | 67.526 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_041957 | Propionibacterium phage PHL151N00 | Propionibacterium phage PHL151N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.99 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041956 | Propionibacterium phage PHL117M01 | Propionibacterium phage PHL117M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29422 | 53.912 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041955 | Propionibacterium phage PHL082M03 | Propionibacterium phage PHL082M03 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.383 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041954 | Propionibacterium phage Keiki | Propionibacterium phage Keiki Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29339 | 54.027 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041937 | Escherichia phage V18 | Escherichia phage V18 Escherichia virus V18 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129454 | 43.626 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_041936 | Escherichia phage ECD7 | Escherichia phage ECD7 Escherichia virus ECD7 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164706 | 40.542 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_041935 | Escherichia phage PA28 | Escherichia phage PA28 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61292 | 49.741 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_041929 | Sinorhizobium phage phiM7 | Sinorhizobium phage phiM7 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 188427 | 49.037 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| NC_041928 | Staphylococcus phage phiIBB-SEP1 | Staphylococcus phage phiIBB-SEP1 Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139928 | 27.93 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_041926 | Escherichia phage vB_EcoM_ECOO78 | Escherichia phage vB_EcoM_ECOO78 Escherichia virus ECOO78 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41289 | 53.075 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| NC_041924 | Acinetobacter phage WCHABP12 | Acinetobacter phage WCHABP12 Acinetobacter virus WCHABP12 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45415 | 37.578 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_041923 | Salmonella phage Mushroom | Salmonella phage Mushroom Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87709 | 39.035 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_041922 | Salmonella phage Si3 | Salmonella phage Si3 Salmonella virus Si3 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84419 | 39.013 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_041910 | Vibrio phage SSP002 | Vibrio phage SSP002 Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76350 | 48.777 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| NC_041909 | Paenibacillus phage Shelly | Paenibacillus phage Shelly Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41152 | 41.534 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_041906 | Escherichia phage EC1-UPM | Escherichia phage EC1-UPM Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70912 | 42.887 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_041902 | Pseudomonas phage PA5 | Pseudomonas phage PA5 Pseudomonas virus PA5 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66182 | 55.489 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041900 | Klebsiella phage vB_KpnM_KpV52 | Klebsiella phage vB_KpnM_KpV52 Klebsiella virus KpV80 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47405 | 48.955 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| NC_041898 | Escherichia phage K1ind2 | Escherichia phage K1ind2 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42765 | 51.348 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_041897 | Escherichia phage K1ind1 | Escherichia phage K1ind1 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42292 | 51.272 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_041895 | Propionibacterium phage G4 | Propionibacterium phage G4 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38555 | 65.706 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041894 | Propionibacterium phage E6 | Propionibacterium phage E6 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38067 | 65.201 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041893 | Propionibacterium phage Doucette | Propionibacterium phage Doucette Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37429 | 65.583 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041892 | Propionibacterium phage B3 | Propionibacterium phage B3 Anatolevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35948 | 64.424 | Anatolevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041891 | Propionibacterium phage B22 | Propionibacterium phage B22 Doucettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37219 | 65.45 | Doucettevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041890 | Propionibacterium phage Anatole | Propionibacterium phage Anatole Anatolevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35284 | 64.431 | Anatolevirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_041888 | Mycobacterium phage Tortellini | Mycobacterium phage Tortellini Mycobacterium virus Tortellini Tortellinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49658 | 65.818 | Tortellinivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041886 | Gordonia phage Strosahl | Gordonia phage Strosahl Gordonia virus Strosahl Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52738 | 61.963 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_041883 | Gordonia phage Attis | Gordonia phage Attis Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47881 | 66.82 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| NC_041882 | Mycobacterium phage SkinnyPete | Mycobacterium phage SkinnyPete Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43478 | 66.353 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_041873 | Escherichia phage JMPW2 | Escherichia phage JMPW2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50298 | 45.376 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_041871 | Escherichia phage Murica | Escherichia phage Murica Escherichia virus Murica Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135391 | 43.614 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_041870 | Pseudomonas phage vB_PaeM_CEB_DP1 | Pseudomonas phage vB_PaeM_CEB_DP1 Pseudomonas virus DP1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66158 | 55.55 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041869 | Escherichia phage APECc02 | Escherichia phage APECc02 Escherichia virus APECc02 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135400 | 43.592 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_041868 | Paracoccus phage Shpa | Paracoccus phage Shpa Paracoccus virus Shpa Vhulanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38261 | 64.674 | Vhulanivirus | Unclassified | Siphoviridae | Paracoccus |
| NC_041867 | Paenibacillus phage Willow | Paenibacillus phage Willow unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37994 | 41.857 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_041866 | Acinetobacter phage YMC11/11/R3177 | Acinetobacter phage YMC11/11/R3177 Acinetobacter virus R3177 Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47575 | 39.832 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| NC_041865 | Pseudomonas phage phiKTN6 | Pseudomonas phage phiKTN6 Pseudomonas virus KTN6 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65994 | 55.513 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041864 | Escherichia coli O157 typing phage 6 | Escherichia coli O157 typing phage 6 Escherichia virus O157tp6 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160570 | 37.628 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_041863 | Escherichia coli O157 typing phage 3 | Escherichia coli O157 typing phage 3 Escherichia virus O157tp3 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168733 | 37.627 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_041860 | Streptomyces phage Danzina | Streptomyces phage Danzina Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50773 | 65.667 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_041859 | Flavobacterium phage 23T | Flavobacterium phage 23T Flavobacterium virus 23T Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43226 | 31.389 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_041857 | Acinetobacter phage vB_AbaM-IME-AB2 | Acinetobacter phage vB_AbaM-IME-AB2 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43665 | 37.497 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_041855 | Mycobacterium virus Dorothy | Mycobacterium virus Dorothy Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58866 | 61.407 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041853 | Mycobacterium virus Marcell | Mycobacterium virus Marcell Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49186 | 63.971 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041852 | Mycobacterium virus Nepal | Mycobacterium virus Nepal Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53972 | 63.548 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041851 | Pseudomonas virus FHA0480 | Pseudomonas virus FHA0480 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37374 | 64.141 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_041850 | Mycobacterium virus Eureka | Mycobacterium virus Eureka Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76174 | 62.852 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041849 | Streptomyces phage phiELB20 | Streptomyces phage phiELB20 Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51160 | 66.986 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_041848 | Bacteriophage P2 | Bacteriophage P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31200 | 52.603 | Peduovirus | Peduovirinae | Myoviridae | Unspecified |
| NC_041847 | Mycobacterium virus Mutaforma13 | Mycobacterium virus Mutaforma13 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57701 | 61.283 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041846 | Mycobacterium phage Bask21 | Mycobacterium phage Bask21 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74997 | 62.857 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041844 | Mycobacterium virus Optimus | Mycobacterium virus Optimus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109270 | 60.791 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_041834 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24608 | 27.329 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_041833 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51534 | 27.632 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_041832 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14701 | 26.427 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_041831 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27985 | 28.426 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_031107 | Erwinia phage vB_EamM_Asesino | Erwinia phage vB_EamM_Asesino Erskinevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246290 | 51.211 | Erskinevirus | Unclassified | Myoviridae | Erwinia |
| NC_009552 | Geobacillus virus E2 | Geobacillus virus E2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40863 | 44.786 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| NC_031068 | Pectobacterium phage PP16 | Pectobacterium phage PP16 Pectobacterium virus PP16 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44268 | 51.936 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_028808 | Escherichia phage phiSUSP1 | Escherichia phage phiSUSP1 Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90743 | 39.761 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| NC_028671 | Enterococcus phage vB_EfaS_IME197 | Enterococcus phage vB_EfaS_IME197 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41098 | 34.058 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_004775 | Salmonella phage epsilon15 | Salmonella phage epsilon15 Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39672 | 50.834 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| NC_028935 | Escherichia phage SUSP2 | Escherichia phage SUSP2 Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88698 | 40.158 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| NC_004686 | Mycobacterium phage Che9d | Mycobacterium phage Che9d Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 60.901 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029043 | Mycobacterium phage SweetiePie | Mycobacterium phage SweetiePie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53184 | 63.472 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029018 | Mycobacterium phage Anubis | Mycobacterium phage Anubis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50854 | 64.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028960 | Mycobacterium phage Theia | Mycobacterium phage Theia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51543 | 60.802 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028876 | Mycobacterium phage Ariel | Mycobacterium phage Ariel unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109801 | 61.018 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021306 | Mycobacterium phage Dumbo | Mycobacterium phage Dumbo unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75736 | 62.996 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031255 | Gordonia phage Schnabeltier | Gordonia phage Schnabeltier Gordonia virus Schnabeltier Schnabeltiervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46895 | 67.037 | Schnabeltiervirus | Unclassified | Siphoviridae | Gordonia |
| NC_029048 | Clostridium phage phiCD211 | Clostridium phage phiCD211 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131704 | 26.415 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_031062 | Erwinia phage vB_EamP_Frozen | Erwinia phage vB_EamP_Frozen Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75147 | 46.92 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| NC_034248 | Rhizobium phage RHEph10 | Rhizobium phage RHEph10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115042 | 60.323 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| NC_029003 | Salmonella phage SEN1 | Salmonella phage SEN1 Salmonella virus SEN1 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29733 | 53.012 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| NC_028696 | Salmonella phage SEN22 | Salmonella phage SEN22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41338 | 47.828 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_031021 | Salmonella phage MA12 | Salmonella phage MA12 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41224 | 49.876 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_029015 | Salmonella phage SEN4 | Salmonella phage SEN4 Salmonella virus SEN4 Senquatrovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33509 | 53.365 | Senquatrovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_028701 | Salmonella phage SEN5 | Salmonella phage SEN5 unclassified Senquatrovirus Senquatrovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33509 | 53.365 | Senquatrovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_031937 | Escherichia phage LM33_P1 | Escherichia phage LM33_P1 Escherichia virus LM33P1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38979 | 50.186 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_031925 | Salmonella phage BPS11Q3 | Salmonella phage BPS11Q3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43788 | 49.705 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_031918 | Salmonella phage 64795_sal3 | Salmonella phage 64795_sal3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45342 | 45.596 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_031946 | Salmonella phage 103203_sal5 | Salmonella phage 103203_sal5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40443 | 46.525 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_031944 | Synechococcus phage S-WAM1 | Synechococcus phage S-WAM1 unclassified Libanvirus Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 185102 | 44.659 | Libanvirus | Unclassified | Myoviridae | Synechococcus |
| NC_031943 | Escherichia phage GA2A | Escherichia phage GA2A Escherichia virus GA2A Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40470 | 51.097 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_031942 | Streptococcus phage phiARI0462 | Streptococcus phage phiARI0462 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40498 | 40.543 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031941 | Streptococcus phage phiARI0131-2 | Streptococcus phage phiARI0131-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34563 | 40.187 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031940 | Salmonella phage 118970_sal3 | Salmonella phage 118970_sal3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77375 | 50.701 | Unclassified | Unclassified | Myoviridae | Salmonella |
| NC_031939 | Salmonella phage BPS15Q2 | Salmonella phage BPS15Q2 Salmonella virus BPS15Q2 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89817 | 38.857 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_031938 | Flavobacterium phage Fpv6 | Flavobacterium phage Fpv6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42820 | 31.985 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031936 | Gordonia phage GAL1 | Gordonia phage GAL1 Gordonia virus GAL1 Galunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49979 | 63.547 | Galunavirus | Unclassified | Siphoviridae | Gordonia |
| NC_031935 | Synechococcus phage S-WAM2 | Synechococcus phage S-WAM2 Synechococcus virus SWAM2 Cymopoleiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186386 | 41.321 | Cymopoleiavirus | Unclassified | Myoviridae | Synechococcus |
| NC_031934 | Escherichia phage MX01 | Escherichia phage MX01 Escherichia virus MX01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168929 | 39.564 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_031930 | Salmonella phage 118970_sal1 | Salmonella phage 118970_sal1 Salmonella virus 118970sal1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59518 | 56.492 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_031929 | Streptococcus phage phiARI0468-1 | Streptococcus phage phiARI0468-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41063 | 40.806 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031928 | Escherichia phage WG01 | Escherichia phage WG01 Escherichia virus WG01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169936 | 39.623 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_031926 | Flavobacterium phage 2A | Flavobacterium phage 2A Flavobacterium virus 2A Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46575 | 31.744 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_031924 | Salmonella phage IME207 | Salmonella phage IME207 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47564 | 46.415 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_031923 | Streptococcus phage phiARI0468-2 | Streptococcus phage phiARI0468-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42186 | 40.362 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031922 | Synechococcus phage S-CAM9 | Synechococcus phage S-CAM9 Synechococcus virus SCAM9 Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174830 | 39.046 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| NC_031920 | Streptococcus phage phiARI0004 | Streptococcus phage phiARI0004 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41105 | 40.659 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031919 | Flavobacterium phage Fpv8 | Flavobacterium phage Fpv8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031915 | Streptococcus phage phiARI0468-4 | Streptococcus phage phiARI0468-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37395 | 38.131 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031913 | Streptococcus phage phiARI0460-1 | Streptococcus phage phiARI0460-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41807 | 39.69 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031911 | Flavobacterium phage 1H | Flavobacterium phage 1H Flavobacterium virus 1H Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39290 | 31.441 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_031910 | Streptococcus phage phiARI0031 | Streptococcus phage phiARI0031 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41887 | 41.208 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031907 | Streptococcus phage phiARI0746 | Streptococcus phage phiARI0746 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33181 | 39.586 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031906 | Synechococcus phage S-CAM3 | Synechococcus phage S-CAM3 Synechococcus virus SCAM3 Charybdisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 198190 | 41.558 | Charybdisvirus | Unclassified | Myoviridae | Synechococcus |
| NC_031905 | Flavobacterium phage Fpv7 | Flavobacterium phage Fpv7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42894 | 32.037 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031903 | Synechococcus phage S-CAM22 | Synechococcus phage S-CAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172345 | 39.913 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_031901 | Streptococcus phage phiARI0131-1 | Streptococcus phage phiARI0131-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42927 | 39.597 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031921 | Flavobacterium phage Fpv5 | Flavobacterium phage Fpv5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031900 | Synechococcus phage S-CAM4 | Synechococcus phage S-CAM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191983 | 38.583 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_031279 | Mycobacterium phage Bactobuster | Mycobacterium phage Bactobuster unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52129 | 63.074 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031277 | Mycobacterium phage ArcherNM | Mycobacterium phage ArcherNM unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52561 | 64.177 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031269 | Gordonia phage Yeezy | Gordonia phage Yeezy Baxtervirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51884 | 66.695 | Baxtervirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| NC_031266 | Mycobacterium phage Phrann | Mycobacterium phage Phrann Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44872 | 66.28 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_031265 | Gordonia phage Guacamole | Gordonia phage Guacamole Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 67.186 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031264 | Brucella phage BiPBO1 | Brucella phage BiPBO1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46877 | 53.32 | Unclassified | Unclassified | Siphoviridae | Brucella |
| NC_031263 | Mycobacterium phage EvilGenius | Mycobacterium phage EvilGenius unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53935 | 63.007 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031262 | Mycobacterium phage Brocalys | Mycobacterium phage Brocalys unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54751 | 61.776 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031261 | Mycobacterium phage Shipwreck | Mycobacterium phage Shipwreck unclassified Fishburnevirus Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48670 | 66.926 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_031256 | Mycobacterium phage Loser | Mycobacterium phage Loser unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53486 | 64.464 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031243 | Mycobacterium phage Xeno | Mycobacterium phage Xeno Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42395 | 66.803 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_031233 | Gordonia phage Kita | Gordonia phage Kita Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 66.665 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| NC_031228 | Salmonella phage BP12C | Salmonella phage BP12C Salmonella virus BP12C Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60606 | 56.395 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_031251 | Gordonia phage SoilAssassin | Gordonia phage SoilAssassin unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47880 | 66.819 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031250 | Salmonella phage BP63 | Salmonella phage BP63 Salmonella virus BP63 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52437 | 46.021 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| NC_031249 | Gordonia phage UmaThurman | Gordonia phage UmaThurman Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50127 | 67.026 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031246 | Klebsiella phage KpV71 | Klebsiella phage KpV71 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43267 | 53.979 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_031242 | Cyanophage S-RIM50 | Cyanophage S-RIM50 Synechococcus virus SRIM50 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174307 | 40.281 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| NC_031237 | Gordonia phage Obliviate | Gordonia phage Obliviate Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49286 | 67.53 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031226 | Gordonia phage BaxterFox | Gordonia phage BaxterFox Baxtervirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53717 | 66.543 | Baxtervirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| NC_031281 | Mycobacterium phage Panchino | Mycobacterium phage Panchino Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43516 | 65.948 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_031271 | Salmonella phage BP12B | Salmonella phage BP12B Salmonella virus BP12B Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43602 | 47.557 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| NC_031267 | Gordonia phage Lucky10 | Gordonia phage Lucky10 Gordonia virus Lucky10 Luckytenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42979 | 65.378 | Luckytenvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031260 | Enterococcus phage Ec-ZZ2 | Enterococcus phage Ec-ZZ2 Enterococcus virus EcZZ2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41170 | 34.596 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_031258 | Salmonella phage BP12A | Salmonella phage BP12A Salmonella virus BP12A Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39696 | 48.952 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| NC_031257 | Mycobacterium phage Mulciber | Mycobacterium phage Mulciber unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52428 | 63.724 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031241 | Staphylococcus phage CNPx | Staphylococcus phage CNPx Staphylococcus virus CNPx Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43293 | 34.661 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_031238 | Mycobacterium phage Catalina | Mycobacterium phage Catalina unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53411 | 62.641 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031235 | Cyanophage S-RIM32 | Cyanophage S-RIM32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194437 | 39.868 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_031056 | Bacillus phage Deep Blue | Bacillus phage Deep Blue Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157501 | 39.947 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031127 | Erwinia phage vB_EamM_Huxley | Erwinia phage vB_EamM_Huxley unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240761 | 51.069 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| NC_031124 | Mycobacterium phage Gardann | Mycobacterium phage Gardann unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76012 | 58.903 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031122 | Gordonia phage Eyre | Gordonia phage Eyre Gordonia virus Eyre Eyrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44929 | 67.455 | Eyrevirus | Unclassified | Siphoviridae | Gordonia |
| NC_031121 | Bacillus phage Aurora | Bacillus phage Aurora Bacillus virus Aurora Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25908 | 30.662 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_031120 | Erwinia phage vB_EamM_Caitlin | Erwinia phage vB_EamM_Caitlin unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241147 | 52.17 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| NC_031119 | Propionibacterium phage Enoki | Propionibacterium phage Enoki unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29347 | 54.588 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_031118 | Gordonia phage CarolAnn | Gordonia phage CarolAnn Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54167 | 66.902 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031117 | Acinetobacter phage LZ35 | Acinetobacter phage LZ35 Acinetobacter virus LZ35 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44885 | 37.948 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_031115 | Morganella phage vB_MmoP_MP2 | Morganella phage vB_MmoP_MP2 Morganella virus MP2 Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39616 | 46.913 | Minipunavirus | Studiervirinae | Autographiviridae | Morganella |
| NC_031114 | Escherichia phage 64795_ec1 | Escherichia phage 64795_ec1 Escherichia virus 64795ec1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39235 | 48.658 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_031108 | Propionibacterium phage PFR2 | Propionibacterium phage PFR2 unclassified Pulverervirus Pulverervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39640 | 64.955 | Pulverervirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_031103 | Escherichia phage vB_EcoM-UFV13 | Escherichia phage vB_EcoM-UFV13 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165772 | 35.537 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_031101 | Weissella phage WCP30 | Weissella phage WCP30 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33697 | 41.229 | Unclassified | Unclassified | Siphoviridae | Weissella |
| NC_031100 | Bacillus phage Phrodo | Bacillus phage Phrodo unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164443 | 37.802 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031098 | Acinetobacter phage vB_AbaS_TRS1 | Acinetobacter phage vB_AbaS_TRS1 unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40749 | 40.256 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| NC_031097 | Gordonia phage Zirinka | Gordonia phage Zirinka Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52077 | 66.667 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| NC_031095 | Klebsiella phage PKO111 | Klebsiella phage PKO111 Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168758 | 39.394 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_031092 | Citrobacter phage SH2 | Citrobacter phage SH2 Citrobacter virus SH2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39158 | 50.723 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_031090 | Shigella phage SHSML-52-1 | Shigella phage SHSML-52-1 Shigella virus SHSML521 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169621 | 37.576 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| NC_031087 | Klebsiella phage vB_KpnM_KpV477 | Klebsiella phage vB_KpnM_KpV477 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168272 | 39.321 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_031084 | Propionibacterium phage BruceLethal | Propionibacterium phage BruceLethal unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29249 | 54.039 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_031082 | Escherichia phage vB_EcoM_Alf5 | Escherichia phage vB_EcoM_Alf5 Escherichia virus Alf5 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87662 | 39.018 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_031081 | Escherichia phage Envy | Escherichia phage Envy unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45206 | 54.457 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_031078 | Streptomyces phage Nanodon | Streptomyces phage Nanodon unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50082 | 65.698 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_031077 | Enterobacter phage Tyrion | Enterobacter phage Tyrion unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41760 | 50.587 | Uetakevirus | Unclassified | Podoviridae | Enterobacter |
| NC_031076 | Propionibacterium phage PFR1 | Propionibacterium phage PFR1 Pulverervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38071 | 64.842 | Pulverervirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_031075 | Lactococcus phage M5938 | Lactococcus phage M5938 Lactococcus virus M5938 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22240 | 35.863 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_031072 | Gordonia phage CaptainKirk2 | Gordonia phage CaptainKirk2 Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47898 | 67.368 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031071 | Lactococcus phage D4410 | Lactococcus phage D4410 Lactococcus virus D4410 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22825 | 35.706 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_031070 | Bacillus phage Nemo | Bacillus phage Nemo unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162375 | 38.787 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031067 | Mycobacterium phage Makemake | Mycobacterium phage Makemake unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52496 | 63.761 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031066 | Citrobacter phage SH1 | Citrobacter phage SH1 Citrobacter virus SH1 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39434 | 50.953 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_031064 | Lactococcus phage 98201 | Lactococcus phage 98201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36202 | 35.951 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_031061 | Gordonia phage Nymphadora | Gordonia phage Nymphadora Gordonia virus Nymphadora Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53431 | 66.555 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| NC_031060 | Gordonia phage JSwag | Gordonia phage JSwag unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52726 | 61.918 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031059 | Rhodovulum phage vB_RhkS_P1 | Rhodovulum phage vB_RhkS_P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38773 | 67.454 | Unclassified | Unclassified | Siphoviridae | Rhodovulum |
| NC_031055 | Bacillus phage PfEFR-5 | Bacillus phage PfEFR-5 unclassified Hubeivirus Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43773 | 35.392 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| NC_031054 | Bacillus phage DirtyBetty | Bacillus phage DirtyBetty unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162415 | 38.724 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031053 | Klebsiella phage PKP126 | Klebsiella phage PKP126 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50934 | 50.73 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_031052 | Gordonia phage Twister6 | Gordonia phage Twister6 Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57804 | 67.722 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031050 | Escherichia phage vB_EcoS_NBD2 | Escherichia phage vB_EcoS_NBD2 Escherichia virus NBD2 Vilniusvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51802 | 49.817 | Vilniusvirus | Unclassified | Drexlerviridae | Escherichia |
| NC_031049 | Bacillus phage NotTheCreek | Bacillus phage NotTheCreek unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161929 | 38.711 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031048 | Enterobacter phage Arya | Enterobacter phage Arya unclassified Jilinvirus Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41918 | 54.108 | Jilinvirus | Unclassified | Myoviridae | Enterobacter |
| NC_031047 | Escherichia phage HY03 | Escherichia phage HY03 Escherichia virus HY03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170770 | 35.302 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_031046 | Staphylococcus phage BP39 | Staphylococcus phage BP39 Staphylococcus virus BP39 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17641 | 29.018 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_031044 | Lactococcus phage M6165 | Lactococcus phage M6165 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22179 | 35.732 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_031040 | Lactococcus phage 50101 | Lactococcus phage 50101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31987 | 35.618 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_031037 | Bacillus phage Nigalana | Bacillus phage Nigalana unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163041 | 38.75 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031036 | Lactobacillus phage PLE2 | Lactobacillus phage PLE2 Lactobacillus virus PLE2 Pleeduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35068 | 45.326 | Pleeduovirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_031032 | Bacillus phage Stitch | Bacillus phage Stitch Bacillus virus Stitch Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24320 | 30.362 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_031031 | Gordonia phage Remus | Gordonia phage Remus unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52738 | 61.967 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031030 | Escherichia phage UFV-AREG1 | Escherichia phage UFV-AREG1 Escherichia virus AREG1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170788 | 35.279 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_031029 | Gordonia phage Cucurbita | Gordonia phage Cucurbita unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93686 | 61.985 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| NC_031027 | Bacillus phage SageFayge | Bacillus phage SageFayge unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162359 | 38.698 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031026 | Salmonella phage phSE-2 | Salmonella phage phSE-2 Salmonella virus phSE2 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49167 | 42.889 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| NC_031025 | Klebsiella phage KpV475 | Klebsiella phage KpV475 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42201 | 54.198 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_031018 | Citrobacter phage SH4 | Citrobacter phage SH4 Citrobacter virus SH4 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39274 | 52.587 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_031017 | Lactococcus phage 63301 | Lactococcus phage 63301 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32409 | 35.132 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_031015 | Bacillus phage Claudi | Bacillus phage Claudi Bacillus virus Claudi Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26504 | 30.26 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_031013 | Lactococcus phage 28201 | Lactococcus phage 28201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34396 | 35.821 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_031012 | Bacillus phage Kida | Bacillus phage Kida unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162151 | 38.666 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031011 | Shigella phage SHFML-26 | Shigella phage SHFML-26 Shigella virus SHFML26 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168993 | 35.382 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_031009 | Lactococcus phage D4412 | Lactococcus phage D4412 Lactococcus virus D4412 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22884 | 35.431 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_031008 | Staphylococcus phage SLPW | Staphylococcus phage SLPW Staphylococcus virus SLPW Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17861 | 29.354 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_031004 | Gordonia phage Nyceirae | Gordonia phage Nyceirae Gordonia virus Nyceirae Nyceiraevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41857 | 67.513 | Nyceiraevirus | Unclassified | Siphoviridae | Gordonia |
| NC_031113 | Escherichia phage Gluttony | Escherichia phage Gluttony unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44513 | 54.456 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_031019 | Enterobacteria phage UAB_Phi20 | Enterobacteria phage UAB_Phi20 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41809 | 47.241 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| NC_031111 | Mycobacterium phage Gompeii16 | Mycobacterium phage Gompeii16 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53623 | 63.559 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031085 | Shigella phage SHBML-50-1 | Shigella phage SHBML-50-1 Shigella virus SHBML501 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166634 | 35.372 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_031065 | Salmonella phage vB_SnwM_CGG4-1 | Salmonella phage vB_SnwM_CGG4-1 unclassified Gelderlandvirus Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159878 | 37.148 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_031041 | Mycobacterium phage Papez | Mycobacterium phage Papez unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52501 | 63.787 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031034 | Bacillus phage SalinJah | Bacillus phage SalinJah unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161140 | 38.72 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031024 | Bacillus phage Belinda | Bacillus phage Belinda unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162308 | 38.784 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031020 | Morganella phage vB_MmoM_MP1 | Morganella phage vB_MmoM_MP1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163201 | 34.747 | Unclassified | Tevenvirinae | Myoviridae | Morganella |
| NC_031016 | Lactococcus phage PLgT-1 | Lactococcus phage PLgT-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40273 | 35.401 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_031002 | Lactococcus phage M6162 | Lactococcus phage M6162 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21697 | 35.793 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_031123 | Citrobacter phage SH3 | Citrobacter phage SH3 Citrobacter virus SH3 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39444 | 50.641 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_031006 | Bacillus phage DIGNKC | Bacillus phage DIGNKC unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161552 | 38.67 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031003 | Propionibacterium phage Moyashi | Propionibacterium phage Moyashi unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29254 | 54.587 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_031126 | Erwinia phage vB_EamM_ChrisDB | Erwinia phage vB_EamM_ChrisDB unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244840 | 49.414 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| NC_031125 | Lactobacillus phage PLE3 | Lactobacillus phage PLE3 Lactobacillus virus PLE3 Pleetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41005 | 44.931 | Pleetrevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_031116 | Bacillus phage Zuko | Bacillus phage Zuko unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163345 | 38.63 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_031099 | Gordonia phage Hedwig | Gordonia phage Hedwig Gordonia virus Hedwig Hedwigvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44536 | 67.193 | Hedwigvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031094 | Streptococcus virus 9872 | Streptococcus virus 9872 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33105 | 36.84 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031073 | Pseudomonas phage vB_PaeM_MAG1 | Pseudomonas phage vB_PaeM_MAG1 Pseudomonas virus MAG1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94555 | 49.248 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_031069 | Streptococcus virus 9871 | Streptococcus virus 9871 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32729 | 37.059 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_031023 | Streptococcus virus 9874 | Streptococcus virus 9874 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32649 | 36.617 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_030953 | Shigella phage SHFML-11 | Shigella phage SHFML-11 Shigella virus SHFML11 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170650 | 35.238 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_030951 | Clostridium phage phiCTC2B | Clostridium phage phiCTC2B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38243 | 29.056 | Unclassified | Unclassified | Myoviridae | Clostridium |
| NC_030949 | Clostridium phage phiCTC2A | Clostridium phage phiCTC2A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47020 | 28.405 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_030946 | Streptococcus phage phiARI0923 | Streptococcus phage phiARI0923 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33477 | 38.952 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_030945 | Bacillus phage BalMu-1 | Bacillus phage BalMu-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39873 | 42.761 | Unclassified | Unclassified | Myoviridae | Bacillus |
| NC_030943 | Gordonia phage Blueberry | Gordonia phage Blueberry Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54990 | 67.03 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030942 | Gordonia phage BritBrat | Gordonia phage BritBrat Gordonia virus Britbrat Britbratvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55524 | 64.999 | Britbratvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030934 | Pseudomonas phage vB_PsyM_KIL1 | Pseudomonas phage vB_PsyM_KIL1 Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90552 | 44.811 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| NC_030933 | Staphylococcus phage Stau2 | Staphylococcus phage Stau2 Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133798 | 29.974 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_030929 | Pseudomonas phage JBD44 | Pseudomonas phage JBD44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49033 | 58.493 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_030921 | Gordonia phage Utz | Gordonia phage Utz Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49768 | 67.652 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030920 | Bacillus phage Eldridge | Bacillus phage Eldridge Bacillus virus Eldridge Eldridgevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165868 | 40.407 | Eldridgevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_030916 | Tsukamurella phage TPA4 | Tsukamurella phage TPA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56212 | 70.542 | Unclassified | Unclassified | Siphoviridae | Tsukamurella |
| NC_030912 | Gordonia phage McGonagall | Gordonia phage McGonagall Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17119 | 68.614 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_030904 | Bacillus phage vB_BhaS-171 | Bacillus phage vB_BhaS-171 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38975 | 40.816 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_030940 | Pseudomonas phage phi3 | Pseudomonas phage phi3 Pseudomonas virus phi3 Phitrevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32637 | 63.854 | Phitrevirus | Peduovirinae | Myoviridae | Pseudomonas |
| NC_030939 | Gordonia phage GMA4 | Gordonia phage GMA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45537 | 66.368 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_030931 | Pseudomonas phage phi2 | Pseudomonas phage phi2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41871 | 62.115 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_030930 | Gordonia phage Bowser | Gordonia phage Bowser Gordonia virus Bowser Bowservirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46570 | 67.112 | Bowservirus | Unclassified | Siphoviridae | Gordonia |
| NC_030925 | Bacillus phage Shbh1 | Bacillus phage Shbh1 Bacillus virus Shbh1 Shalavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138081 | 36.428 | Shalavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_030919 | Salmonella phage 118970_sal4 | Salmonella phage 118970_sal4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42418 | 46.813 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_030918 | Pseudomonas phage JBD93 | Pseudomonas phage JBD93 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36629 | 64.293 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_030917 | Gordonia phage OneUp | Gordonia phage OneUp Oneupvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93577 | 61.483 | Oneupvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030913 | Gordonia phage Wizard | Gordonia phage Wizard Wizardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58308 | 67.943 | Wizardvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030909 | Pseudomonas phage YMC11/07/P54_PAE_BP | Pseudomonas phage YMC11/07/P54_PAE_BP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50183 | 62.18 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_030908 | Pseudomonas phage JBD69 | Pseudomonas phage JBD69 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36938 | 63.931 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_030902 | Gordonia phage GMA1 | Gordonia phage GMA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41207 | 65.712 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_030950 | Clostridium phage phiCT19406A | Clostridium phage phiCT19406A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47020 | 28.396 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_030947 | Clostridium phage phiCT19406B | Clostridium phage phiCT19406B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38243 | 29.048 | Unclassified | Unclassified | Myoviridae | Clostridium |
| NC_030884 | Sulfolobus islandicus rudivirus 3 | Sulfolobus islandicus rudivirus 3 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32995 | 25.519 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_030698 | Gordonia phage Soups | Gordonia phage Soups Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52924 | 61.864 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030695 | Gordonia phage KatherineG | Gordonia phage KatherineG unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52689 | 61.926 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030694 | Gordonia phage Rosalind | Gordonia phage Rosalind unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52684 | 61.93 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030652 | Staphylococcus phage 80 | Staphylococcus phage 80 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42140 | 35.565 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028748 | Bacillus phage vB_BtS_BMBtp3 | Bacillus phage vB_BtS_BMBtp3 Bacillus virus BMBtp3 Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 35.446 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| NC_028695 | Enterobacter phage phiEap-2 | Enterobacter phage phiEap-2 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40491 | 51.972 | Cornellvirus | Guernseyvirinae | Siphoviridae | Enterobacter |
| NC_019919 | Yersinia phage phiR2-01 | Yersinia phage phiR2-01 Yersinia virus phiR201 Haartmanvirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122696 | 40.512 | Haartmanvirus | Markadamsvirinae | Demerecviridae | Yersinia |
| NC_029316 | Acidianus tailed spindle virus | Acidianus tailed spindle virus unclassified Bicaudaviridae Bicaudaviridae Viruses | 70812 | 36.41 | Unclassified | Unclassified | Bicaudaviridae | Acidianus |
| NC_029083 | Pseudomonas phage PaoP5 | Pseudomonas phage PaoP5 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93464 | 49.508 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_029121 | Bacillus phage Bp8p-C | Bacillus phage Bp8p-C Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151417 | 41.153 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_029120 | Shigella phage 75/02 Stx | Shigella phage 75/02 Stx Diegovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60875 | 49.117 | Diegovirus | Sepvirinae | Podoviridae | Shigella |
| NC_029119 | Staphylococcus phage SPbeta-like | Staphylococcus phage SPbeta-like Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127726 | 30.534 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_029116 | Clostridium phage phiCDHM13 | Clostridium phage phiCDHM13 Clostridioides virus phiCDHM13 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33596 | 29.989 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_029107 | Ralstonia phage RS138 | Ralstonia phage RS138 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 65.103 | Unclassified | Unclassified | Siphoviridae | Ralstonia |
| NC_029104 | Brevibacillus phage Jimmer1 | Brevibacillus phage Jimmer1 unclassified Jimmervirus Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54312 | 38.109 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| NC_029103 | Sulfolobus monocaudavirus SMV3 | Sulfolobus monocaudavirus SMV3 unclassified Bicaudaviridae Bicaudaviridae Viruses | 64323 | 38.363 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_029102 | Enterobacter phage E-2 | Enterobacter phage E-2 Enterobacter virus E2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36051 | 50.639 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| NC_029099 | Klebsiella phage KP36 | Klebsiella phage KP36 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49797 | 50.702 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_029098 | Streptomyces phage Jay2Jay | Streptomyces phage Jay2Jay Streptomyces virus Jay2Jay Samistivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133531 | 49.497 | Samistivirus | Unclassified | Siphoviridae | Streptomyces |
| NC_029093 | Mycobacterium phage Mindy | Mycobacterium phage Mindy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75796 | 62.965 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029091 | Escherichia phage APCEc01 | Escherichia phage APCEc01 Escherichia virus APCEc01 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168771 | 37.707 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_029084 | Paenibacillus phage Rani | Paenibacillus phage Rani Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37990 | 41.779 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_029080 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142096 | 30.396 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_029079 | Mycobacterium phage Dusk | Mycobacterium phage Dusk unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75339 | 63.018 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029077 | Mycobacterium phage VohminGhazi | Mycobacterium phage VohminGhazi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52155 | 61.524 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029073 | Geobacillus virus E3 | Geobacillus virus E3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141298 | 29.635 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| NC_029068 | Lactobacillus phage LfeSau | Lactobacillus phage LfeSau Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31703 | 48.024 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_029066 | Pseudomonas phage PS-1 | Pseudomonas phage PS-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48666 | 59.834 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_029059 | Mycobacterium phage Edtherson | Mycobacterium phage Edtherson unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51492 | 63.977 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_029058 | Lactobacillus phage LfeInf | Lactobacillus phage LfeInf Lactobacillus virus LfeInf Hopescreekvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106071 | 38.162 | Hopescreekvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_029046 | Sinorhizobium phage phiLM21 | Sinorhizobium phage phiLM21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50827 | 60.661 | Unclassified | Unclassified | Siphoviridae | Sinorhizobium |
| NC_029045 | Salmonella phage 37 | Salmonella phage 37 Salmonella virus 37 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60216 | 56.527 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_024206 | Lactobacillus phage phi jlb1 | Lactobacillus phage phi jlb1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38269 | 37.359 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| NC_024329 | Podovirus Lau218 | Podovirus Lau218 Lauvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39425 | 34.699 | Lauvirus | Unclassified | Autographiviridae | Unspecified |
| NC_029022 | Clostridium phage phiCT9441A | Clostridium phage phiCT9441A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46895 | 28.692 | Unclassified | Unclassified | Myoviridae | Clostridium |
| NC_029008 | Bacillus phage phi4J1 | Bacillus phage phi4J1 Bacillus virus phi4J1 Rockvillevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41486 | 35.889 | Rockvillevirus | Unclassified | Siphoviridae | Bacillus |
| NC_029006 | Clostridium phage phiCT19406C | Clostridium phage phiCT19406C Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35703 | 29.188 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_029004 | Clostridium phage phiCT453B | Clostridium phage phiCT453B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36712 | 29.129 | Unclassified | Unclassified | Myoviridae | Clostridium |
| NC_028991 | Clostridium phage phiCT453A | Clostridium phage phiCT453A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42158 | 29.456 | Unclassified | Unclassified | Myoviridae | Clostridium |
| NC_028886 | Bacillus phage phi4B1 | Bacillus phage phi4B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38663 | 35.892 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_028880 | Citrobacter phage phiCFP-1 | Citrobacter phage phiCFP-1 Citrobacter virus CFP1 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38625 | 50.278 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_028857 | Citrobacter phage Merlin | Citrobacter phage Merlin Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172733 | 38.763 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_028772 | Enterobacter phage phiEap-1 | Enterobacter phage phiEap-1 Enterobacter virus Eap1 Eapunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39133 | 51.688 | Eapunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| NC_028959 | Clostridium phage phiMMP03 | Clostridium phage phiMMP03 Clostridioides virus phiMMP03 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52261 | 28.87 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| NC_028958 | Clostridium phage phiCD146 | Clostridium phage phiCD146 Clostridioides virus phiCD146 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41507 | 30.689 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| NC_028951 | Clostridium phage phiCD481-1 | Clostridium phage phiCD481-1 Clostridioides virus phiCD481-1 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32846 | 30.25 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_028905 | Clostridium phage phiCD111 | Clostridium phage phiCD111 Clostridioides virus phiCD111 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41560 | 30.893 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| NC_028883 | Clostridium phage phiMMP01 | Clostridium phage phiMMP01 Clostridioides virus phiMMP01 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44461 | 28.92 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| NC_028838 | Clostridium phage phiCD506 | Clostridium phage phiCD506 Clostridioides virus phiCD506 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33274 | 29.612 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_028764 | Clostridium phage phiCD505 | Clostridium phage phiCD505 Clostridioides virus phiCD505 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49316 | 29.384 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| NC_029030 | Streptococcus phage APCM01 | Streptococcus phage APCM01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31075 | 39.028 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_029029 | Brevibacillus phage Abouo | Brevibacillus phage Abouo Abouovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45552 | 39.16 | Abouovirus | Unclassified | Myoviridae | Brevibacillus |
| NC_029028 | Enterobacteria phage JenP1 | Enterobacteria phage JenP1 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60754 | 43.232 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| NC_029027 | Sulfolobales virus YNP1 | Sulfolobales virus YNP1 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 40860 | 38.338 | Unclassified | Unclassified | Unclassified | Unspecified |
| NC_029025 | Staphylococcus phage IME-SA4 | Staphylococcus phage IME-SA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41843 | 33.982 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_029023 | Escherichia phage JH2 | Escherichia phage JH2 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87712 | 38.816 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_029021 | Enterobacteria phage JenK1 | Enterobacteria phage JenK1 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60747 | 43.862 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| NC_029020 | Sulfolobus monocaudavirus SMV2 | Sulfolobus monocaudavirus SMV2 unclassified Bicaudaviridae Bicaudaviridae Viruses | 50918 | 43.552 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_029010 | Staphylococcus phage SA97 | Staphylococcus phage SA97 Staphylococcus virus SA97 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40592 | 34.246 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_029009 | Enterococcus phage EFDG1 | Enterococcus phage EFDG1 Enterococcus virus EFDG1 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147589 | 37.203 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_029001 | Clostridium phage phiCDHM11 | Clostridium phage phiCDHM11 Clostridioides virus phiCDHM11 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32000 | 29.256 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_028997 | Enterobacteria phage JenP2 | Enterobacteria phage JenP2 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59802 | 43.214 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| NC_028996 | Clostridium phage phiCDHM19 | Clostridium phage phiCDHM19 Clostridioides virus phiCDHM19 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54295 | 28.249 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| NC_028995 | Acinetobacter phage phiAC-1 | Acinetobacter phage phiAC-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43216 | 38.476 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| NC_028984 | Streptococcus phage M102AD | Streptococcus phage M102AD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30664 | 39.62 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_028977 | Klebsiella phage vB_KpnP_KpV289 | Klebsiella phage vB_KpnP_KpV289 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41054 | 52.565 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_028976 | Streptomyces phage Izzy | Streptomyces phage Izzy Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50113 | 65.911 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_028971 | Pseudomonas phage DL68 | Pseudomonas phage DL68 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66111 | 55.724 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028969 | Brevibacillus phage Osiris | Brevibacillus phage Osiris Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52955 | 38.102 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| NC_028967 | Propionibacterium phage PAC1 | Propionibacterium phage PAC1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29605 | 54.042 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_028965 | Mycobacterium phage Pioneer | Mycobacterium phage Pioneer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53219 | 62.587 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028962 | Staphylococcus phage phiIPLA-C1C | Staphylococcus phage phiIPLA-C1C Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140961 | 27.96 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_028961 | Mycobacterium phage PopTart | Mycobacterium phage PopTart unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55094 | 61.644 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028957 | Escherichia phage vB_EcoM_VR26 | Escherichia phage vB_EcoM_VR26 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171541 | 40.222 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028956 | Mannheimia phage vB_MhS_1152AP2 | Mannheimia phage vB_MhS_1152AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52139 | 41.094 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| NC_028955 | Prochlorococcus phage P-TIM68 | Prochlorococcus phage P-TIM68 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197361 | 34.321 | Unclassified | Unclassified | Myoviridae | Prochlorococcus |
| NC_028953 | Mycobacterium phage MiaZeal | Mycobacterium phage MiaZeal unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110764 | 61.153 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028947 | Mycobacterium phage Kratio | Mycobacterium phage Kratio unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62738 | 65.67 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028946 | Mycobacterium phage Dante | Mycobacterium phage Dante unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59652 | 61.793 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028945 | Sinorhizobium phage phiN3 | Sinorhizobium phage phiN3 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 206713 | 49.075 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| NC_028944 | Bacillus phage Eyuki | Bacillus phage Eyuki Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162252 | 38.773 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_028943 | Escherichia phage pro483 | Escherichia phage pro483 Escherichia virus pro483 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29237 | 52.981 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_028941 | Mycobacterium phage Turj99 | Mycobacterium phage Turj99 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51161 | 63.742 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028940 | Pectobacterium bacteriophage PM2 | Pectobacterium bacteriophage PM2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170286 | 34.785 | Unclassified | Unclassified | Myoviridae | Pectobacterium |
| NC_028939 | Pseudomonas phage vB_Pae_PS44 | Pseudomonas phage vB_Pae_PS44 Pseudomonas virus PS44 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68871 | 55.242 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028937 | Mycobacterium phage Ovechkin | Mycobacterium phage Ovechkin unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58338 | 62.004 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028936 | Mycobacterium phage Enkosi | Mycobacterium phage Enkosi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59052 | 67.183 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028933 | Pseudomonas phage PhiCHU | Pseudomonas phage PhiCHU Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45626 | 52.021 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_028932 | Mycobacterium phage Serenity | Mycobacterium phage Serenity unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52088 | 62.642 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028930 | Paenibacillus phage Tripp | Paenibacillus phage Tripp Paenibacillus virus Tripp Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54439 | 48.315 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028929 | Listeria phage vB_LmoS_293 | Listeria phage vB_LmoS_293 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40759 | 36.861 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_028928 | Mycobacterium phage Nerujay | Mycobacterium phage Nerujay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53455 | 63.693 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028927 | Escherichia phage slur02 | Escherichia phage slur02 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167298 | 35.516 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028925 | Escherichia phage vB_EcoM_VR25 | Escherichia phage vB_EcoM_VR25 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170822 | 40.418 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028923 | Mycobacterium phage Llama | Mycobacterium phage Llama unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58472 | 61.067 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028920 | Mycobacterium phage Abrogate | Mycobacterium phage Abrogate unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52530 | 63.756 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028919 | Pseudomonas phage DL54 | Pseudomonas phage DL54 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45673 | 52.394 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_028917 | Staphylococcus phage 3MRA | Staphylococcus phage 3MRA Staphylococcus virus 3MRA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41931 | 35.411 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028916 | Proteus phage vB_PmiP_Pm5460 | Proteus phage vB_PmiP_Pm5460 Proteus virus Pm5460 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44573 | 39.576 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| NC_028915 | Staphylococcus phage B236 | Staphylococcus phage B236 Staphylococcus virus B236 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43228 | 35.553 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028914 | Mycobacterium phage Sheen | Mycobacterium phage Sheen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52927 | 63.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028912 | Mycobacterium phage Swirley | Mycobacterium phage Swirley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49717 | 61.044 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028911 | Lactobacillus phage iLp1308 | Lactobacillus phage iLp1308 Lactobacillus virus iLp1308 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34176 | 45.049 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_028910 | Mycobacterium phage XFactor | Mycobacterium phage XFactor unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55617 | 61.715 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028906 | Mycobacterium phage Toto | Mycobacterium phage Toto Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75933 | 63.017 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028904 | Streptomyces phage Amela | Streptomyces phage Amela Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49452 | 65.613 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_028901 | Escherichia phage slur05 | Escherichia phage slur05 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43900 | 54.544 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_028900 | Lactococcus phage SL4 | Lactococcus phage SL4 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28144 | 34.967 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_028898 | Mannheimia phage vB_MhM_587AP1 | Mannheimia phage vB_MhM_587AP1 unclassified Baylorvirus Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35764 | 42.062 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| NC_028897 | Mycobacterium phage Chadwick | Mycobacterium phage Chadwick unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49421 | 59.786 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028896 | Escherichia phage pro147 | Escherichia phage pro147 Escherichia virus pro147 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32675 | 50.742 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_028894 | Escherichia phage vB_EcoM_VR20 | Escherichia phage vB_EcoM_VR20 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170336 | 40.445 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028892 | Streptomyces phage Caliburn | Streptomyces phage Caliburn Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49949 | 66.194 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_028890 | Bacillus phage TsarBomba | Bacillus phage TsarBomba Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162486 | 40.074 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_028889 | Mycobacterium phage Florinda | Mycobacterium phage Florinda unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59416 | 61.748 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028888 | Lactobacillus phage CL1 | Lactobacillus phage CL1 Lactobacillus virus CL1 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39474 | 44.774 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_028887 | Bacillus phage AvesoBmore | Bacillus phage AvesoBmore Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167431 | 37.69 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_028881 | Escherichia phage vb_EcoM-VR5 | Escherichia phage vb_EcoM-VR5 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170473 | 39.303 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028878 | Mycobacterium phage Archie | Mycobacterium phage Archie unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76271 | 58.722 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028877 | Mycobacterium phage Larenn | Mycobacterium phage Larenn unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52967 | 63.54 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028874 | Mycobacterium phage Pari | Mycobacterium phage Pari unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50614 | 63.532 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028872 | Escherichia phage HY02 | Escherichia phage HY02 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86252 | 38.921 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_028871 | Listeria phage vB_LmoS_188 | Listeria phage vB_LmoS_188 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38392 | 35.872 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_028870 | Klebsiella phage vB_KpnP_SU552A | Klebsiella phage vB_KpnP_SU552A Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43595 | 54.153 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_028863 | Escherichia phage P694 | Escherichia phage P694 Escherichia virus P694 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40477 | 48.42 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_028862 | Staphylococcus phage vB_SauS_phi2 | Staphylococcus phage vB_SauS_phi2 Staphylococcus virus phi2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44222 | 33.716 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_028860 | Mycobacterium phage Smeadley | Mycobacterium phage Smeadley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52392 | 61.414 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028859 | Staphylococcus phage B166 | Staphylococcus phage B166 Staphylococcus virus B166 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42881 | 34.829 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028855 | Acinetobacter phage YMC11/12/R2315 | Acinetobacter phage YMC11/12/R2315 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44846 | 37.6 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_028854 | Paenibacillus phage Sitara | Paenibacillus phage Sitara Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43724 | 41.593 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028853 | Mannheimia phage vB_MhS_535AP2 | Mannheimia phage vB_MhS_535AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50078 | 41.31 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| NC_028852 | Mycobacterium phage Equemioh13 | Mycobacterium phage Equemioh13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53042 | 63.506 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028851 | Paenibacillus phage Fern | Paenibacillus phage Fern Paenibacillus virus Fern Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37995 | 41.858 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028849 | Mycobacterium phage Luchador | Mycobacterium phage Luchador unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53387 | 62.111 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028847 | Escherichia phage QL01 | Escherichia phage QL01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170527 | 39.647 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028844 | Mycobacterium phage Phatniss | Mycobacterium phage Phatniss unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57293 | 61.336 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028843 | Mycobacterium phage Lolly9 | Mycobacterium phage Lolly9 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75816 | 59.316 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028842 | Mycobacterium phage CaptainTrips | Mycobacterium phage CaptainTrips unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57328 | 61.523 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028841 | Bacteriophage Lily | Bacteriophage Lily Paenibacillus virus Lily Lilyvirus Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44952 | 42.73 | Lilyvirus | Unclassified | Unclassified | Unspecified |
| NC_028837 | Paenibacillus phage Xenia | Paenibacillus phage Xenia unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 41.525 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028836 | Pseudomonas phage DL62 | Pseudomonas phage DL62 Pseudomonas virus DL62 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42508 | 62.195 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_028835 | Lactobacillus phage CL2 | Lactobacillus phage CL2 Lactobacillus virus CL2 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38751 | 44.879 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_028834 | Achromobacter phage 83-24 | Achromobacter phage 83-24 Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48216 | 54.853 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| NC_028832 | Mycobacterium phage Omnicron | Mycobacterium phage Omnicron unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61511 | 64.034 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028830 | Lactobacillus phage iA2 | Lactobacillus phage iA2 Lactobacillus virus iA2 Iaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34155 | 45.639 | Iaduovirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_028828 | Mycobacterium phage TheloniousMonk | Mycobacterium phage TheloniousMonk unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52055 | 63.648 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028827 | Streptomyces phage Lannister | Streptomyces phage Lannister Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50165 | 65.743 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_028825 | Escherichia phage vB_EcoM_AYO145A | Escherichia phage vB_EcoM_AYO145A Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87372 | 39.003 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_028823 | Mycobacterium phage Bipolar | Mycobacterium phage Bipolar unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58985 | 61.372 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028822 | Escherichia phage P483 | Escherichia phage P483 Escherichia virus P483 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40829 | 48.444 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_028821 | Staphylococcus phage StB20-like | Staphylococcus phage StB20-like Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40670 | 33.29 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_028820 | Yersinia phage vB_YenM_TG1 | Yersinia phage vB_YenM_TG1 Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162101 | 34.56 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| NC_028816 | Klebsiella phage vB_KpnP_SU503 | Klebsiella phage vB_KpnP_SU503 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43809 | 53.749 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_028815 | Mycobacterium phage Nhonho | Mycobacterium phage Nhonho unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51355 | 63.75 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028813 | Mycobacterium phage Seagreen | Mycobacterium phage Seagreen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57766 | 61.832 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028810 | Mycobacterium phage Rufus | Mycobacterium phage Rufus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52357 | 63.854 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028805 | Brevibacillus phage Jenst | Brevibacillus phage Jenst Jenstvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126341 | 42.886 | Jenstvirus | Unclassified | Siphoviridae | Brevibacillus |
| NC_028804 | Mycobacterium phage Barriga | Mycobacterium phage Barriga unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52643 | 63.395 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028802 | Mycobacterium phage UnionJack | Mycobacterium phage UnionJack unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49158 | 59.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028800 | Klebsiella phage K5 | Klebsiella phage K5 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41698 | 52.492 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_028798 | Mycobacterium phage MarQuardt | Mycobacterium phage MarQuardt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50882 | 64.036 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028796 | Mycobacterium phage Phayonce | Mycobacterium phage Phayonce Phayoncevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49203 | 66.673 | Phayoncevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_028795 | Enterobacter phage E-3 | Enterobacter phage E-3 Enterobacter virus E3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31522 | 50.771 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| NC_028794 | Mycobacterium phage Piro94 | Mycobacterium phage Piro94 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52647 | 63.419 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028792 | Mycobacterium phage Kimberlium | Mycobacterium phage Kimberlium unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56826 | 61.44 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028788 | Paenibacillus phage Diva | Paenibacillus phage Diva Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37246 | 42.072 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028786 | Klebsiella phage 1513 | Klebsiella phage 1513 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49462 | 50.611 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_028785 | Mycobacterium phage NelitzaMV | Mycobacterium phage NelitzaMV unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72790 | 63.054 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028784 | Mycobacterium phage Tasp14 | Mycobacterium phage Tasp14 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51409 | 63.923 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028783 | Lactobacillus phage iLp84 | Lactobacillus phage iLp84 Lactobacillus virus iLp84 Pleetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39399 | 45.085 | Pleetrevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_028780 | Escherichia phage slur07 | Escherichia phage slur07 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167124 | 35.429 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028779 | Mycobacterium phage Carcharodon | Mycobacterium phage Carcharodon unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43680 | 66.241 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_028778 | Mycobacterium phage Snenia | Mycobacterium phage Snenia unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75626 | 59.276 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028775 | Staphylococcus phage 23MRA | Staphylococcus phage 23MRA Staphylococcus virus 23MRA Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43098 | 32.902 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_028774 | Klebsiella phage Sushi | Klebsiella phage Sushi Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48754 | 50.769 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_028768 | Achromobacter phage JWX | Achromobacter phage JWX Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49714 | 55.413 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| NC_028767 | Paenibacillus phage Vegas | Paenibacillus phage Vegas Vegasvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45653 | 43.642 | Vegasvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| NC_028766 | Mannheimia phage vB_MhM_3927AP2 | Mannheimia phage vB_MhM_3927AP2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33755 | 43.137 | Unclassified | Unclassified | Myoviridae | Mannheimia |
| NC_028765 | Staphylococcus phage phiIPLA-RODI | Staphylococcus phage phiIPLA-RODI Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142348 | 30.425 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_028762 | Proteus phage vB_PmiM_Pm5461 | Proteus phage vB_PmiM_Pm5461 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161989 | 31.082 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| NC_028760 | Klebsiella phage KLPN1 | Klebsiella phage KLPN1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49037 | 50.525 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| NC_028759 | Mycobacterium phage Mufasa | Mycobacterium phage Mufasa unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58065 | 68.232 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028758 | Paenibacillus phage HB10c2 | Paenibacillus phage HB10c2 Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35644 | 41.844 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_028757 | Mycobacterium phage Tiffany | Mycobacterium phage Tiffany unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50768 | 63.991 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028751 | Mycobacterium phage Quico | Mycobacterium phage Quico unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58671 | 61.688 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028747 | Mycobacterium phage Brusacoram | Mycobacterium phage Brusacoram Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47618 | 66.994 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_028746 | Paenibacillus phage Harrison | Paenibacillus phage Harrison Harrisonvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44249 | 40.157 | Harrisonvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| NC_028745 | Pseudomonas phage DL60 | Pseudomonas phage DL60 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66103 | 54.917 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028247 | Citrobacter phage Michonne | Citrobacter phage Michonne unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90000 | 38.849 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| NC_028744 | Mycobacterium phage LadyBird | Mycobacterium phage LadyBird unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53141 | 63.459 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028743 | Mannheimia phage vB_MhS_587AP2 | Mannheimia phage vB_MhS_587AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48594 | 41.285 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| NC_028698 | Salmonella phage f18SE | Salmonella phage f18SE Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41868 | 49.799 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_028659 | Klebsiella phage vB_KpnM_KB57 | Klebsiella phage vB_KpnM_KB57 Klebsiella virus KB57 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142987 | 44.616 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| NC_028700 | Streptococcus phage T12 | Streptococcus phage T12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37976 | 38.106 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_028699 | Salmonella phage SEN34 | Salmonella phage SEN34 Salmonella virus SEN34 Brunovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40740 | 49.912 | Brunovirus | Unclassified | Myoviridae | Salmonella |
| NC_028697 | Streptococcus phage A25 | Streptococcus phage A25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33900 | 38.448 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_028694 | Propionibacterium phage PA1-14 | Propionibacterium phage PA1-14 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29407 | 53.851 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_028688 | Klebsiella phage vB_Kp1 | Klebsiella phage vB_Kp1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40114 | 53.283 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_028687 | Mycobacterium phage Iracema64 | Mycobacterium phage Iracema64 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51637 | 64.014 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028686 | Klebsiella phage JD18 | Klebsiella phage JD18 Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166313 | 39.599 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_028685 | Shigella phage Ss-VASD | Shigella phage Ss-VASD Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62851 | 50.071 | Oslovirus | Sepvirinae | Podoviridae | Shigella |
| NC_028683 | Edwardsiella phage PEi20 | Edwardsiella phage PEi20 Edwardsiella virus PEi20 Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177643 | 40.637 | Kanagawavirus | Unclassified | Myoviridae | Edwardsiella |
| NC_028680 | Mycobacterium phage Pepe | Mycobacterium phage Pepe unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50515 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028678 | Mycobacterium phage Cabrinians | Mycobacterium phage Cabrinians unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56669 | 61.208 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028670 | Klebsiella phage KpV41 | Klebsiella phage KpV41 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44203 | 53.967 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_028669 | Staphylococcus phage phiJB | Staphylococcus phage phiJB Staphylococcus virus phiJB Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43012 | 34.73 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_028667 | Pseudomonas phage vB_PaeS_PM105 | Pseudomonas phage vB_PaeS_PM105 Pseudomonas virus PM105 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39593 | 63.1 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_028666 | Streptococcus phage Str-PAP-1 | Streptococcus phage Str-PAP-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36595 | 35.289 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_028664 | Klebsiella phage Kp2 | Klebsiella phage Kp2 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43963 | 53.784 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_028663 | Cyanophage P-TIM40 | Cyanophage P-TIM40 Prochlorococcus virus PTIM40 Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 188632 | 40.717 | Libanvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_028661 | Pseudomonas phage PPPL-1 | Pseudomonas phage PPPL-1 Pseudomonas virus PPPL1 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 57.003 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_028660 | Escherichia phage phi191 | Escherichia phage phi191 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61035 | 50.189 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_028657 | Pseudomonas phage YMC11/02/R656 | Pseudomonas phage YMC11/02/R656 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60919 | 58.739 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_028656 | Enterobacteria phage VT2phi_272 | Enterobacteria phage VT2phi_272 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65955 | 50.104 | Oslovirus | Sepvirinae | Podoviridae | Enterobacteria |
| NC_028655 | Yersinia phage vB_YenP_AP10 | Yersinia phage vB_YenP_AP10 Yersinia virus AP10 Apdecimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39235 | 51.757 | Apdecimavirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_028654 | Mycobacterium phage Sparkdehlily | Mycobacterium phage Sparkdehlily unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 61.235 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028449 | Escherichia phage PA2 | Escherichia phage PA2 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63569 | 50.169 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_028448 | Escherichia phage slur14 | Escherichia phage slur14 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167467 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_028248 | Escherichia phage slur16 | Escherichia phage slur16 Escherichia virus slur16 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136896 | 43.613 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_024142 | Escherichia phage Bp4 | Escherichia phage Bp4 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72583 | 42.883 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_020490 | Staphylococcus phage StB12 | Staphylococcus phage StB12 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44714 | 34.171 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_010275 | Vibrio virus Kappa | Vibrio virus Kappa unclassified Longwoodvirus Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33507 | 48.835 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| NC_009763 | Staphylococcus phage tp310-3 | Staphylococcus phage tp310-3 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42973 | 33.454 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_009761 | Staphylococcus phage tp310-1 | Staphylococcus phage tp310-1 Staphylococcus virus tp310-1 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42232 | 33.503 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_009762 | Staphylococcus phage tp310-2 | Staphylococcus phage tp310-2 Staphylococcus virus tp310-2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47785 | 33.751 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_005964 | Mycoplasma phage phiMFV1 | Mycoplasma phage phiMFV1 unclassified bacterial viruses Viruses | 18855 | 24.826 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| NC_004914 | Escherichia phage Stx2 II | Escherichia phage Stx2 II Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62706 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_004913 | Escherichia Stx1 converting phage | Escherichia Stx1 converting phage unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59866 | 49.694 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_002628 | Methanothermobacter phage psiM100 | Methanothermobacter phage psiM100 Psimunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31007 | 45.745 | Psimunavirus | Unclassified | Siphoviridae | Methanothermobacter |
| NC_002484 | Pseudomonas virus D3 | Pseudomonas virus D3 Detrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56426 | 57.805 | Detrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_009799 | Corynebacterium phage BFK20 | Corynebacterium phage BFK20 Corynebacterium virus BFK20 Sasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42972 | 56.162 | Sasvirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_005262 | Burkholderia phage Bcep22 | Burkholderia phage Bcep22 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63882 | 65.309 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| NC_001978 | Streptomyces virus phiC31 | Streptomyces virus phiC31 Lomovskayavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41491 | 63.619 | Lomovskayavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_027995 | Escherichia phage vB_EcoM_ECO1230-10 | Escherichia phage vB_EcoM_ECO1230-10 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41666 | 53.374 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| NC_027986 | Pseudomonas phage JD18 | Pseudomonas phage JD18 Pseudomonas virus JD18 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39014 | 63.388 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_027992 | Pseudomonas phage JBD25 | Pseudomonas phage JBD25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39552 | 62.535 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_027983 | Escherichia phage AR1 | Escherichia phage AR1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167435 | 35.331 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_027979 | Enterobacteria phage RB68 | Enterobacteria phage RB68 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168401 | 35.252 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_027997 | Campylobacter virus CP220 | Campylobacter virus CP220 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177534 | 27.359 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_027996 | Campylobacter virus CPt10 | Campylobacter virus CPt10 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175720 | 27.337 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_027994 | Escherichia phage K1H | Escherichia phage K1H Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41632 | 51.17 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_027993 | Escherichia phage K1G | Escherichia phage K1G Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43587 | 51.068 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| NC_027991 | Staphylococcus phage SA1 | Staphylococcus phage SA1 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147303 | 45.827 | Unclassified | Ounavirinae | Myoviridae | Staphylococcus |
| NC_027990 | Lactobacillus phage LBR48 | Lactobacillus phage LBR48 Lactobacillus virus LBR48 Anamdongvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48211 | 45.877 | Anamdongvirus | Unclassified | Myoviridae | Lactobacillus |
| NC_027989 | Campylobacter phage vB_CcoM-IBB_35 | Campylobacter phage vB_CcoM-IBB_35 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53237 | 27.265 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_027982 | Lactobacillus phage phiPYB5 | Lactobacillus phage phiPYB5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32847 | 45.468 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_027981 | Vibrio phage VP585 | Vibrio phage VP585 Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42612 | 50.878 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| NC_027980 | Enterobacter phage phiKDA1 | Enterobacter phage phiKDA1 Enterobacter virus KDA1 Koutsourovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43804 | 51.678 | Koutsourovirus | Slopekvirinae | Autographiviridae | Enterobacter |
| NC_027630 | Propionibacterium phage Ouroboros | Propionibacterium phage Ouroboros Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29506 | 53.928 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027629 | Propionibacterium phage Attacne | Propionibacterium phage Attacne Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28876 | 54.706 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027628 | Propionibacterium phage Lauchelly | Propionibacterium phage Lauchelly Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29517 | 53.86 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027627 | Propionibacterium phage Solid | Propionibacterium phage Solid Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29440 | 53.916 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027626 | Propionibacterium phage Procrass1 | Propionibacterium phage Procrass1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29347 | 54.04 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027625 | Propionibacterium phage Kubed | Propionibacterium phage Kubed Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29461 | 54.526 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027624 | Propionibacterium phage SKKY | Propionibacterium phage SKKY Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29594 | 54.599 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027622 | Propionibacterium phage Stormborn | Propionibacterium phage Stormborn Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29330 | 53.802 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027621 | Propionibacterium phage Wizzo | Propionibacterium phage Wizzo Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29463 | 54.441 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027620 | Propionibacterium phage MrAK | Propionibacterium phage MrAK Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29726 | 54.417 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_024124 | Escherichia phage vB_EcoM_JS09 | Escherichia phage vB_EcoM_JS09 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169148 | 37.649 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_026607 | Salmonella phage STP4-a | Salmonella phage STP4-a Salmonella virus STP4a Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159914 | 36.862 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_027396 | Streptococcus phage SpSL1 | Streptococcus phage SpSL1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33756 | 38.595 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_027392 | Propionibacterium phage PHL194M00 | Propionibacterium phage PHL194M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.159 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027391 | Propionibacterium phage PHL041M10 | Propionibacterium phage PHL041M10 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29412 | 54.27 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027390 | Proteus phage PM 93 | Proteus phage PM 93 Proteus virus PM93 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45169 | 39.363 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| NC_027389 | Propionibacterium phage PHL141N00 | Propionibacterium phage PHL141N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29494 | 53.943 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027386 | Propionibacterium phage PHL152M00 | Propionibacterium phage PHL152M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29247 | 54.153 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027385 | Propionibacterium phage PHL092M00 | Propionibacterium phage PHL092M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29261 | 53.918 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027384 | Pseudomonas phage H70 | Pseudomonas phage H70 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37359 | 64.25 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_027383 | Escherichia phage YD-2008.s | Escherichia phage YD-2008.s Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44613 | 54.614 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_027382 | Shigella phage SfMu | Shigella phage SfMu Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37146 | 51.946 | Muvirus | Unclassified | Myoviridae | Shigella |
| NC_027380 | Propionibacterium phage PHL067M01 | Propionibacterium phage PHL067M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29377 | 54.267 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027379 | Proteus phage PM 85 | Proteus phage PM 85 Proteus virus PM85 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43642 | 39.322 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| NC_027377 | Leuconostoc phage Ln-8 | Leuconostoc phage Ln-8 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28921 | 36.14 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_027376 | Propionibacterium phage PHL308M00 | Propionibacterium phage PHL308M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29442 | 54.249 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027375 | Pseudomonas phage vB_PaeP_PPA-ABTNL | Pseudomonas phage vB_PaeP_PPA-ABTNL Pseudomonas virus ABTNL Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43227 | 62.442 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_027373 | Propionibacterium phage PHL030N00 | Propionibacterium phage PHL030N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29423 | 53.91 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027370 | Propionibacterium phage PHL179M00 | Propionibacterium phage PHL179M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29428 | 53.928 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027367 | Propionibacterium phage PHL132N00 | Propionibacterium phage PHL132N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29003 | 54.329 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027362 | Propionibacterium phage PHL116M00 | Propionibacterium phage PHL116M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29394 | 53.96 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027361 | Propionibacterium phage PHL085N00 | Propionibacterium phage PHL085N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29454 | 53.833 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027360 | Enterobacteriaphage UAB_Phi87 | Enterobacteriaphage UAB_Phi87 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87603 | 38.887 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| NC_027359 | Propionibacterium phage PHL082M00 | Propionibacterium phage PHL082M00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.379 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027357 | Propionibacterium phage PHL025M00 | Propionibacterium phage PHL025M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29496 | 53.984 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027355 | Bacillus phage BCP8-2 | Bacillus phage BCP8-2 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159071 | 39.452 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_027354 | Propionibacterium phage PHL301M00 | Propionibacterium phage PHL301M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29323 | 54.408 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027352 | Bacillus phage JBP901 | Bacillus phage JBP901 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159492 | 39.658 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_027349 | Escherichia phage HY01 | Escherichia phage HY01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166977 | 35.453 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_027347 | Propionibacterium phage PHL151M00 | Propionibacterium phage PHL151M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.993 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027346 | Propionibacterium phage PHL171M01 | Propionibacterium phage PHL171M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29327 | 53.712 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027339 | Enterobacteria phage SfI | Enterobacteria phage SfI Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38389 | 50.116 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| NC_027337 | Escherichia phage vB_EcoM-VpaE1 | Escherichia phage vB_EcoM-VpaE1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88403 | 38.943 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_027336 | Propionibacterium phage PHL009M11 | Propionibacterium phage PHL009M11 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29503 | 53.937 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027333 | Propionibacterium phage PHL070N00 | Propionibacterium phage PHL070N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29421 | 54.04 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027331 | Citrobacter phage Moon | Citrobacter phage Moon Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170341 | 38.876 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_027329 | Salmonella phage vB_SPuM_SP116 | Salmonella phage vB_SPuM_SP116 Salmonella virus SP116 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87510 | 38.839 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_027405 | Propionibacterium phage PHL163M00 | Propionibacterium phage PHL163M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.155 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027404 | Yersinia phage PST | Yersinia phage PST Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167785 | 35.32 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_027403 | Propionibacterium phage PHL117M00 | Propionibacterium phage PHL117M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29255 | 54.162 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027402 | Salmonella phage SPN3US | Salmonella phage SPN3US Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240413 | 48.54 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| NC_027401 | Propionibacterium phage PHL095N00 | Propionibacterium phage PHL095N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29751 | 53.995 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027400 | Propionibacterium phage PHL055N00 | Propionibacterium phage PHL055N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.159 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027399 | Klebsiella phage K64-1 | Klebsiella phage K64-1 Klebsiella virus K64-1 Alcyoneusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 346602 | 31.717 | Alcyoneusvirus | Unclassified | Myoviridae | Klebsiella |
| NC_027398 | Enterobacteria phage Sf101 | Enterobacteria phage Sf101 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38742 | 47.445 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| NC_027387 | Escherichia phage CICC 80001 | Escherichia phage CICC 80001 Escherichia virus CICC80001 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38810 | 48.82 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_027371 | Propionibacterium phage Pacnes 2012-15 | Propionibacterium phage Pacnes 2012-15 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29741 | 54.423 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027369 | Escherichia phage EC6 | Escherichia phage EC6 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86231 | 38.898 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_027368 | Vibrio phage PV94 | Vibrio phage PV94 Vibrio virus PV94 Vulnificusvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33828 | 48.17 | Vulnificusvirus | Peduovirinae | Myoviridae | Vibrio |
| NC_027364 | Escherichia phage PBECO4 | Escherichia phage PBECO4 Escherichia virus PBECO4 Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348113 | 34.087 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| NC_027353 | Yersinia phage phiD1 | Yersinia phage phiD1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167063 | 35.442 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| NC_027351 | Salmonella phage SSE121 | Salmonella phage SSE121 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147745 | 45.292 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| NC_027344 | Salmonella phage STML-198 | Salmonella phage STML-198 Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158099 | 36.888 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_027334 | Aurantimonas phage AmM-1 | Aurantimonas phage AmM-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47800 | 66.684 | Unclassified | Unclassified | Myoviridae | Aurantimonas |
| NC_027295 | Propionibacterium phage PHL199M00 | Propionibacterium phage PHL199M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29806 | 54.043 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027299 | Sulfitobacter phage NYA-2014a | Sulfitobacter phage NYA-2014a Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42092 | 58.541 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| NC_027298 | Pseudomonas virus LPB1 | Pseudomonas virus LPB1 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36814 | 64.364 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_027294 | Propionibacterium phage PHL150M00 | Propionibacterium phage PHL150M00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29438 | 54.25 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_027293 | Citrobacter phage Moogle | Citrobacter phage Moogle Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87999 | 38.982 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| NC_027204 | Sinorhizobium phage phiM12 | Sinorhizobium phage phiM12 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194701 | 49.013 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| NC_027130 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.135 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| NC_027132 | Synechococcus phage ACG-2014i | Synechococcus phage ACG-2014i Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 190768 | 39.008 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_021190 | Enterobacteria phage phi80 | Enterobacteria phage phi80 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46150 | 52.132 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_026928 | Synechococcus phage ACG-2014e | Synechococcus phage ACG-2014e Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189418 | 38.938 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_026927 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 228143 | 41.619 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| NC_026924 | Synechococcus phage ACG-2014g | Synechococcus phage ACG-2014g unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174885 | 39.258 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| NC_026923 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179110 | 40.265 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_026926 | Synechococcus phage ACG-2014j | Synechococcus phage ACG-2014j Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192108 | 38.566 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_026611 | Edwardsiella phage GF-2 | Edwardsiella phage GF-2 Edwardsiella virus GF2 Gofduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43129 | 51.269 | Gofduovirus | Unclassified | Myoviridae | Edwardsiella |
| NC_026605 | Mycobacterium phage Malithi | Mycobacterium phage Malithi Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46869 | 67.093 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_026604 | Mycobacterium phage Cerasum | Mycobacterium phage Cerasum unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53636 | 61.494 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026602 | Pseudomonas phage vB_PaeP_PAO1_Ab05 | Pseudomonas phage vB_PaeP_PAO1_Ab05 Pseudomonas virus Ab05 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43639 | 62.346 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_026601 | Pseudomonas phage vB_PaeS_PAO1_Ab30 | Pseudomonas phage vB_PaeS_PAO1_Ab30 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37238 | 64.08 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_026600 | Pseudomonas phage vB_PaeM_C1-14_Ab28 | Pseudomonas phage vB_PaeM_C1-14_Ab28 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66181 | 54.902 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_026599 | Pseudomonas phage vB_PaeP_C2-10_Ab22 | Pseudomonas phage vB_PaeP_C2-10_Ab22 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45808 | 52.353 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_026598 | Mycobacterium phage Milly | Mycobacterium phage Milly unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58211 | 68.312 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026597 | Mycobacterium phage Sparky | Mycobacterium phage Sparky Mycobacterium virus Sparky Sparkyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63334 | 65.164 | Sparkyvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026596 | Mycobacterium phage Inventum | Mycobacterium phage Inventum unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57052 | 61.376 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026593 | Mycobacterium phage BuzzLyseyear | Mycobacterium phage BuzzLyseyear unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59419 | 61.132 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026591 | Mycobacterium phage Trike | Mycobacterium phage Trike unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44716 | 64.498 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026590 | Mycobacterium phage Gaia | Mycobacterium phage Gaia Gaiavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90460 | 56.831 | Gaiavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026589 | Mycobacterium phage Sbash | Mycobacterium phage Sbash Chenonavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55832 | 65.647 | Chenonavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026588 | Mycobacterium phage Squirty | Mycobacterium phage Squirty Mycobacterium virus Squirty Squirtyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60285 | 62.405 | Squirtyvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026586 | Pseudomonas phage vB_PaeM_PAO1_Ab27 | Pseudomonas phage vB_PaeM_PAO1_Ab27 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66299 | 55.668 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_026585 | Mycobacterium phage Estave1 | Mycobacterium phage Estave1 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60727 | 61.324 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026584 | Mycobacterium phage Minerva | Mycobacterium phage Minerva unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109871 | 60.745 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026583 | Mycobacterium phage Alvin | Mycobacterium phage Alvin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49577 | 63.628 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026017 | Salmonella phage LSPA1 | Salmonella phage LSPA1 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41880 | 50.487 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_026016 | Staphylococcus phage phiBU01 | Staphylococcus phage phiBU01 Staphylococcus virus BU01 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43748 | 33.016 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_026014 | Enterobacteria phage P88 | Enterobacteria phage P88 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35814 | 52.87 | Peduovirus | Peduovirinae | Myoviridae | Enterobacteria |
| NC_025830 | Escherichia phage Av-05 | Escherichia phage Av-05 Escherichia virus Av05 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120938 | 40.044 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_025829 | Shigella phage pSs-1 | Shigella phage pSs-1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164999 | 35.544 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_025453 | Enterococcus phage EFC-1 | Enterococcus phage EFC-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40286 | 35.047 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_025445 | Escherichia phage J8-65 | Escherichia phage J8-65 Escherichia virus J8-65 Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40981 | 55.687 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| NC_025437 | Shigella phage Shf125875 | Shigella phage Shf125875 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169062 | 37.558 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| NC_025434 | Shigella phage POCJ13 | Shigella phage POCJ13 Diegovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62699 | 49.347 | Diegovirus | Sepvirinae | Podoviridae | Shigella |
| NC_025418 | Klebsiella phage NTUH-K2044-K1-1 | Klebsiella phage NTUH-K2044-K1-1 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43871 | 54.179 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_025417 | Staphylococcus phage Team1 | Staphylococcus phage Team1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140903 | 30.327 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_025416 | Staphylococcus phage MCE-2014 | Staphylococcus phage MCE-2014 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141907 | 30.385 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_025460 | Staphylococcus phage phiSa119 | Staphylococcus phage phiSa119 unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42600 | 33.404 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_025448 | Enterobacteria phage RB27 | Enterobacteria phage RB27 Enterobacteria virus RB27 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165179 | 35.485 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_025425 | Enterobacteria phage GEC-3S | Enterobacteria phage GEC-3S Escherichia virus GEC3S Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163424 | 40.549 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_025419 | Escherichia phage RB3 | Escherichia phage RB3 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168402 | 35.374 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_025464 | Synechococcus phage S-CBP4 | Synechococcus phage S-CBP4 Synechococcus virus SCBP4 Poseidonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44147 | 44.381 | Poseidonvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_025461 | Synechococcus phage S-CBP3 | Synechococcus phage S-CBP3 Synechococcus virus SCBP3 Lirvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45871 | 46.949 | Lirvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_025456 | Synechococcus phage S-CBP1 | Synechococcus phage S-CBP1 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46547 | 47.578 | Unclassified | Unclassified | Autographiviridae | Synechococcus |
| NC_025451 | Yersinia phage vB_YenP_AP5 | Yersinia phage vB_YenP_AP5 Yersinia virus AP5 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38646 | 50.696 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_025450 | Lelliottia phage phD2B | Lelliottia phage phD2B Lelliottia virus phD2B Tuodvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44366 | 51.003 | Tuodvirus | Molineuxvirinae | Autographiviridae | Lelliottia |
| NC_025449 | Escherichia phage ECML-134 | Escherichia phage ECML-134 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166783 | 35.411 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_025447 | Escherichia phage 121Q | Escherichia phage 121Q Escherichia virus 121Q Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348532 | 34.114 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| NC_025443 | Salmonella phage 9NA | Salmonella phage 9NA Nonanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52869 | 42.908 | Nonanavirus | Unclassified | Siphoviridae | Salmonella |
| NC_025442 | Salmonella virus Chi | Salmonella virus Chi Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59578 | 56.511 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_025431 | Mesorhizobium phage vB_MloP_Lo5R7ANS | Mesorhizobium phage vB_MloP_Lo5R7ANS Mesorhizobium virus Lo5R7ANS Pairvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45718 | 61.053 | Pairvirus | Unclassified | Autographiviridae | Mesorhizobium |
| NC_025430 | Escherichia phage vB_EcoM-ep3 | Escherichia phage vB_EcoM-ep3 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42351 | 53.347 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| NC_025427 | Rhizobium phage vB_RleM_PPF1 | Rhizobium phage vB_RleM_PPF1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54506 | 61.9 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| NC_025426 | Staphylococcus phage P108 | Staphylococcus phage P108 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140807 | 30.217 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_025424 | Bacillus phage Waukesha92 | Bacillus phage Waukesha92 Bacillus virus Waukesha92 Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45648 | 35.452 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| NC_024125 | Escherichia phage vB_EcoM_112 | Escherichia phage vB_EcoM_112 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168470 | 35.275 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_025115 | Ralstonia phage RSY1 | Ralstonia phage RSY1 Ralstonia virus RSY1 Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40002 | 64.822 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| NC_024794 | Escherichia phage vB_EcoM_PhAPEC2 | Escherichia phage vB_EcoM_PhAPEC2 Escherichia virus PhAPEC2 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167318 | 37.725 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_024793 | Escherichia phage vB_EcoS_AHP42 | Escherichia phage vB_EcoS_AHP42 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46847 | 43.977 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_024792 | Bacillus phage Bobb | Bacillus phage Bobb Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160281 | 41.331 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024791 | Vibrio phage ICP2_2013_A_Haiti | Vibrio phage ICP2_2013_A_Haiti Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50440 | 42.797 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| NC_024790 | Escherichia phage vB_EcoP_PhAPEC7 | Escherichia phage vB_EcoP_PhAPEC7 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71778 | 43.345 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_024789 | Escherichia phage vB_EcoS_AKS96 | Escherichia phage vB_EcoS_AKS96 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45746 | 43.943 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_024788 | Bacillus phage Riley | Bacillus phage Riley Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162816 | 37.784 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024787 | Listeria phage LMTA-148 | Listeria phage LMTA-148 Listeria virus LMTA148 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132541 | 36.027 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_024786 | Escherichia phage vB_EcoP_PhAPEC5 | Escherichia phage vB_EcoP_PhAPEC5 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71248 | 43.492 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_024785 | Acinetobacter phage YMC-13-01-C62 | Acinetobacter phage YMC-13-01-C62 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44844 | 37.599 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_024784 | Escherichia phage vB_EcoS_AHS24 | Escherichia phage vB_EcoS_AHS24 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46440 | 43.811 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_024782 | Pseudomonas phage MP48 | Pseudomonas phage MP48 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36838 | 64.091 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_020858 | Sulfitobacter phage pCB2047-A | Sulfitobacter phage pCB2047-A Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40929 | 58.802 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| NC_020856 | Sulfitobacter phage pCB2047-C | Sulfitobacter phage pCB2047-C Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40931 | 58.994 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| NC_024391 | Staphylococcus phage DW2 | Staphylococcus phage DW2 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 35.977 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_024390 | Leuconostoc phage LN03 | Leuconostoc phage LN03 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26752 | 36.779 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024387 | Listeria phage LP-101 | Listeria phage LP-101 Listeria virus LP101 Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43767 | 35.451 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| NC_024386 | Leuconostoc phage LN25 | Leuconostoc phage LN25 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28427 | 36.191 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024385 | Leuconostoc phage phiLN12 | Leuconostoc phage phiLN12 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28193 | 36.637 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024384 | Listeria phage LP-030-3 | Listeria phage LP-030-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41156 | 36.554 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_024380 | Leuconostoc phage phiLN6B | Leuconostoc phage phiLN6B Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25740 | 36.465 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024379 | Escherichia phage PE3-1 | Escherichia phage PE3-1 Escherichia virus PE3-1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39093 | 49.927 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_024368 | Mycobacterium phage Pinto | Mycobacterium phage Pinto unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50610 | 63.533 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024362 | Pseudomonas phage phiPSA2 | Pseudomonas phage phiPSA2 Pseudomonas virus PhiPSA2 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40472 | 57.371 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_024370 | Streptococcus phage IC1 | Streptococcus phage IC1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39965 | 41.121 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_024365 | Pseudomonas phage phiPSA1 | Pseudomonas phage phiPSA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51090 | 58.565 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_024364 | Listeria phage List-36 | Listeria phage List-36 Listeria virus List36 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131952 | 36.013 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_024361 | Streptococcus phage DCC1738 | Streptococcus phage DCC1738 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38392 | 40.813 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_024360 | Listeria phage LMSP-25 | Listeria phage LMSP-25 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138036 | 35.87 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_024357 | Streptococcus phage K13 | Streptococcus phage K13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39360 | 40.615 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_024330 | Pseudomonas phage JD024 | Pseudomonas phage JD024 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37380 | 64.152 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_005880 | Staphylococcus virus K | Staphylococcus virus K Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148317 | 30.386 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_023558 | Cronobacter phage Dev2 | Cronobacter phage Dev2 Cronobacter virus Dev2 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38966 | 52.559 | Kayfunavirus | Studiervirinae | Autographiviridae | Cronobacter |
| NC_024216 | Bacillus phage CAM003 | Bacillus phage CAM003 Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160541 | 38.031 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024213 | Bacillus phage Hakuna | Bacillus phage Hakuna Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158100 | 38.696 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024211 | Bacillus phage Megatron | Bacillus phage Megatron Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158750 | 38.803 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024210 | Escherichia phage e4/1c | Escherichia phage e4/1c Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47112 | 44.078 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_024207 | Bacillus phage Evoli | Bacillus phage Evoli Bacillus virus Evoli Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159656 | 38.059 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024205 | Bacillus phage Hoody T | Bacillus phage Hoody T Bacillus virus HoodyT Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159837 | 38.014 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024204 | Salmonella phage vB_SenS-Ent3 | Salmonella phage vB_SenS-Ent3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42764 | 49.792 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_024148 | Mycobacterium phage Phantastic | Mycobacterium phage Phantastic unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50101 | 63.805 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024147 | Mycobacterium phage ZoeJ | Mycobacterium phage ZoeJ unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57315 | 68.525 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024146 | Enterobacteria phage 9g | Enterobacteria phage 9g Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56702 | 43.894 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| NC_024144 | Clostridium phage CDMH1 | Clostridium phage CDMH1 Clostridioides virus CDMH1 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54279 | 28.446 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| NC_024143 | Mycobacterium phage Lamina13 | Mycobacterium phage Lamina13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53255 | 63.664 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024141 | Mycobacterium phage Kampy | Mycobacterium phage Kampy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51378 | 63.891 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024137 | Bacillus phage Bcp1 | Bacillus phage Bcp1 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152778 | 39.757 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_024136 | Mycobacterium phage Seabiscuit | Mycobacterium phage Seabiscuit unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51781 | 63.657 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024134 | Escherichia phage FFH2 | Escherichia phage FFH2 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139020 | 43.609 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_024121 | Serratia phage PS2 | Serratia phage PS2 Serratia virus PS2 Muldoonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167266 | 41.698 | Muldoonvirus | Unclassified | Myoviridae | Serratia |
| NC_024123 | Pseudomonas phage KPP25 | Pseudomonas phage KPP25 Kochitakasuvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64113 | 60.287 | Kochitakasuvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_023567 | Klebsiella phage F19 | Klebsiella phage F19 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43766 | 53.82 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_022088 | Bacillus phage Troll | Bacillus phage Troll Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163019 | 37.825 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_023748 | Mycobacterium phage SkiPole | Mycobacterium phage SkiPole Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53137 | 63.649 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023746 | Mycobacterium virus GUmbie | Mycobacterium virus GUmbie Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57387 | 61.369 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023745 | Mycobacterium virus LHTSCC | Mycobacterium virus LHTSCC Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51813 | 63.932 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023744 | Mycobacterium phage DS6A | Mycobacterium phage DS6A Mycobacterium virus DS6A Hnatkovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60588 | 68.438 | Hnatkovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023742 | Mycobacterium phage Akoma | Mycobacterium phage Akoma unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68711 | 67.541 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022980 | Brevibacillus phage Davies | Brevibacillus phage Davies Abouovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45798 | 39.1 | Abouovirus | Unclassified | Myoviridae | Brevibacillus |
| NC_022972 | Arthrobacter phage vB_ArS-ArV2 | Arthrobacter phage vB_ArS-ArV2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37372 | 62.731 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| NC_023862 | Mycobacterium phage Alsfro | Mycobacterium phage Alsfro unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52136 | 63.57 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023743 | Escherichia phage phiEB49 | Escherichia phage phiEB49 Escherichia virus phiEB49 Lindendrivevirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47180 | 44.019 | Lindendrivevirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_023738 | Mycobacterium phage Thibault | Mycobacterium phage Thibault Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106327 | 60.794 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023735 | Rhodococcus phage ReqiPepy6 | Rhodococcus phage ReqiPepy6 Pepyhexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76797 | 53.411 | Pepyhexavirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_023734 | Psychrobacter phage Psymv2 | Psychrobacter phage Psymv2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35725 | 44.467 | Unclassified | Unclassified | Siphoviridae | Psychrobacter |
| NC_023732 | Mycobacterium virus Rumpelstiltskin | Mycobacterium virus Rumpelstiltskin Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69279 | 58.911 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023731 | Mycobacterium phage Trixie | Mycobacterium phage Trixie Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53526 | 64.464 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023730 | Mycobacterium phage Redi | Mycobacterium phage Redi Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42594 | 66.145 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_023729 | Mycobacterium phage Charlie | Mycobacterium phage Charlie Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43036 | 66.331 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_023728 | Mycobacterium virus DeadP | Mycobacterium virus DeadP Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56461 | 61.552 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023726 | Mycobacterium virus Euphoria | Mycobacterium virus Euphoria Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53597 | 63.675 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023724 | Mycobacterium virus Larva | Mycobacterium virus Larva Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62991 | 65.295 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023723 | Mycobacterium phage Aeneas | Mycobacterium phage Aeneas unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53684 | 63.587 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023721 | Mycobacterium virus Drago | Mycobacterium virus Drago Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54411 | 61.17 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023720 | Mycobacterium virus Perseus | Mycobacterium virus Perseus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53142 | 63.712 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023719 | Bacillus phage G | Bacillus phage G Donellivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 497513 | 29.926 | Donellivirus | Unclassified | Myoviridae | Bacillus |
| NC_023715 | Yersinia phage Yep-phi | Yersinia phage Yep-phi Yersinia virus Yepf Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38616 | 47.1 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_023713 | Mycobacterium phage Blue7 | Mycobacterium phage Blue7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52288 | 61.358 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023710 | Mycobacterium phage RidgeCB | Mycobacterium phage RidgeCB Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50844 | 63.952 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023709 | Mycobacterium phage Wile | Mycobacterium phage Wile unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51308 | 63.69 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023708 | Mycobacterium phage Dreamboat | Mycobacterium phage Dreamboat unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 63.908 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023707 | Mycobacterium phage Turbido | Mycobacterium phage Turbido Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53169 | 63.283 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023704 | Mycobacterium virus Doom | Mycobacterium virus Doom Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51421 | 63.807 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023703 | Mycobacterium phage Dori | Mycobacterium phage Dori Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64613 | 65.988 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_023702 | Mycobacterium virus Kugel | Mycobacterium virus Kugel Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52379 | 63.837 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023700 | Pseudomonas phage PA1/KOR/2010 | Pseudomonas phage PA1/KOR/2010 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34553 | 64.767 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_023699 | Mycobacterium virus SG4 | Mycobacterium virus SG4 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59016 | 61.871 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023698 | Mycobacterium phage Avani | Mycobacterium phage Avani Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54470 | 60.973 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023697 | Mycobacterium phage Babsiella | Mycobacterium phage Babsiella Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48420 | 67.078 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| NC_023695 | Mycobacterium phage Violet | Mycobacterium phage Violet Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52481 | 63.846 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023694 | Rhodococcus phage ReqiPoco6 | Rhodococcus phage ReqiPoco6 Pepyhexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78064 | 53.265 | Pepyhexavirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_023692 | Mycobacterium phage BigNuz | Mycobacterium phage BigNuz Bignuzvirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48984 | 66.736 | Bignuzvirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_023690 | Mycobacterium virus Courthouse | Mycobacterium virus Courthouse Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110569 | 60.922 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023689 | Mycobacterium phage Lilac | Mycobacterium phage Lilac unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76260 | 62.969 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023687 | Mycobacterium phage Bruns | Mycobacterium phage Bruns Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53003 | 63.617 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023686 | Mycobacterium phage Gadjet | Mycobacterium phage Gadjet Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67949 | 67.486 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023609 | Mycobacterium phage RhynO | Mycobacterium phage RhynO unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46739 | 65.262 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023607 | Mycobacterium phage 40AC | Mycobacterium phage 40AC unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53396 | 63.338 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023602 | Mycobacterium phage 32HC | Mycobacterium phage 32HC Trigintaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50781 | 65.731 | Trigintaduovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023553 | Mycobacterium phage Echild | Mycobacterium phage Echild unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53159 | 63.69 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023552 | Mycobacterium phage Donovan | Mycobacterium phage Donovan Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47162 | 67.181 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_023551 | Enterococcus phage IME-EF4 | Enterococcus phage IME-EF4 Enterococcus virus EF4 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40692 | 34.624 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_023550 | Staphylococcus phage GRCS | Staphylococcus phage GRCS Staphylococcus virus GRCS Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17869 | 28.899 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_023606 | Mycobacterium phage CRB1 | Mycobacterium phage CRB1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52963 | 63.227 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023599 | Bacillus phage phiCM3 | Bacillus phage phiCM3 Bacillus virus phiCM3 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38772 | 35.482 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| NC_023597 | Mycobacterium phage 20ES | Mycobacterium phage 20ES unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53124 | 63.427 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023588 | Stenotrophomonas phage Smp131 | Stenotrophomonas phage Smp131 Stenotrophomonas virus Smp131 Simpcentumvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33525 | 65.023 | Simpcentumvirus | Peduovirinae | Myoviridae | Stenotrophomonas |
| NC_023587 | Synechococcus phage ACG-2014h | Synechococcus phage ACG-2014h Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189311 | 40.491 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_023584 | Synechococcus phage S-MbCM100 | Synechococcus phage S-MbCM100 Synechococcus virus SMbCM100 Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170438 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| NC_023582 | Staphylococcus phage vB_SepS_SEP9 | Staphylococcus phage vB_SepS_SEP9 Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92417 | 29.568 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_023580 | Mycobacterium phage Saal | Mycobacterium phage Saal unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57775 | 61.289 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023579 | Erwinia phage Ea9-2 | Erwinia phage Ea9-2 Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75568 | 47.041 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| NC_023577 | Mycobacterium phage Obama12 | Mycobacterium phage Obama12 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51797 | 63.963 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023572 | Mycobacterium phage MichelleMyBell | Mycobacterium phage MichelleMyBell Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42240 | 66.03 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_023565 | Mycobacterium phage Nyxis | Mycobacterium phage Nyxis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51250 | 63.924 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023564 | Mycobacterium phage EagleEye | Mycobacterium phage EagleEye unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52974 | 61.423 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023562 | Mycobacterium phage BellusTerra | Mycobacterium phage BellusTerra unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51236 | 63.857 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023561 | Enterobacter phage PG7 | Enterobacter phage PG7 Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173276 | 39.822 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| NC_023608 | Salmonella phage vB_SenS-Ent2 | Salmonella phage vB_SenS-Ent2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42093 | 49.916 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_023585 | Sulfolobus monocaudavirus SMV1 | Sulfolobus monocaudavirus SMV1 unclassified Bicaudaviridae Bicaudaviridae Viruses | 48775 | 38.421 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_023583 | Pseudomonas phage TL | Pseudomonas phage TL Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45696 | 52.366 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_023576 | Citrobacter phage CR44b | Citrobacter phage CR44b Citrobacter virus CR44b Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39207 | 50.491 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_023575 | Pseudomonas phage vB_PaeP_Tr60_Ab31 | Pseudomonas phage vB_PaeP_Tr60_Ab31 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45550 | 57.104 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| NC_023573 | Staphylococcus phage phiSA12 | Staphylococcus phage phiSA12 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142094 | 30.31 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_023571 | Oenococcus phage phiS11 | Oenococcus phage phiS11 Oenococcus virus phiS11 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46243 | 38.603 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| NC_023560 | Oenococcus phage phiS13 | Oenococcus phage phiS13 Oenococcus virus phiS13 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43454 | 38.97 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| NC_023559 | Oenococcus phage phi9805 | Oenococcus phage phi9805 Oenococcus virus phi9805 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46145 | 38.86 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| NC_023548 | Citrobacter phage CR8 | Citrobacter phage CR8 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39651 | 49.681 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| NC_023501 | Bacillus phage BPS10C | Bacillus phage BPS10C Bacillus virus BPS10C Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159590 | 38.742 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_023500 | Staphylococcus phage StauST398-5 | Staphylococcus phage StauST398-5 Staphylococcus virus StauST398-5 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43301 | 35.135 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_023499 | Staphylococcus phage StauST398-4 | Staphylococcus phage StauST398-4 Staphylococcus virus StauST398-4 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42906 | 33.084 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_023498 | Mycobacterium phage Validus | Mycobacterium phage Validus unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62466 | 68.431 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023503 | Streptococcus phage 20617 | Streptococcus phage 20617 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48800 | 39.975 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_021539 | Listeria phage LP-030-2 | Listeria phage LP-030-2 Psavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38275 | 34.751 | Psavirus | Unclassified | Siphoviridae | Listeria |
| NC_022775 | Lactobacillus phage phiJB | Lactobacillus phage phiJB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36969 | 47.702 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_022771 | Bacillus phage Grass | Bacillus phage Grass Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156648 | 42.247 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_022769 | Bacillus phage BigBertha | Bacillus phage BigBertha Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165238 | 37.773 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_022763 | Bacillus phage Spock | Bacillus phage Spock Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164297 | 37.621 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_020081 | Bacillus phage phiAGATE | Bacillus phage phiAGATE Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149844 | 40.966 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_023009 | Staphylococcus phage Sb1 | Staphylococcus phage Sb1 Staphylococcus virus Sb1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127188 | 30.475 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_023006 | Pseudomonas phage PPpW-3 | Pseudomonas phage PPpW-3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43564 | 61.11 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| NC_009232 | Salmonella phage SETP3 | Salmonella phage SETP3 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42572 | 49.852 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_022988 | Mycobacterium phage Bruin | Mycobacterium phage Bruin unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74210 | 63.023 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022985 | Mycobacterium phage Zaka | Mycobacterium phage Zaka unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52122 | 61.504 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022984 | Mycobacterium phage Jovo | Mycobacterium phage Jovo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51319 | 60.771 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022983 | Mycobacterium phage Bernardo | Mycobacterium phage Bernardo Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68196 | 67.428 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022981 | Mycobacterium phage HufflyPuff | Mycobacterium phage HufflyPuff unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76323 | 63.064 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022979 | Mycobacterium phage Graduation | Mycobacterium phage Graduation unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52823 | 63.452 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022977 | Mycobacterium phage Artemis2UCLA | Mycobacterium phage Artemis2UCLA unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52344 | 61.405 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022976 | Mycobacterium phage Nala | Mycobacterium phage Nala unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75894 | 63.052 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022975 | Mycobacterium phage HanShotFirst | Mycobacterium phage HanShotFirst unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52390 | 63.814 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022973 | Mycobacterium phage Conspiracy | Mycobacterium phage Conspiracy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50755 | 60.615 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022971 | Pseudomonas phage phiIBB-PAA2 | Pseudomonas phage phiIBB-PAA2 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45344 | 52.327 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_022969 | Mycobacterium phage PhatBacter | Mycobacterium phage PhatBacter unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76217 | 62.979 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022968 | Escherichia phage 4MG | Escherichia phage 4MG Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148567 | 46.333 | Seunavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_022965 | Mycobacterium phage CloudWang3 | Mycobacterium phage CloudWang3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52873 | 61.353 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022920 | Staphylococcus phage S25-3 | Staphylococcus phage S25-3 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139738 | 30.219 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_022918 | Staphylococcus phage S25-4 | Staphylococcus phage S25-4 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132123 | 30.309 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_022915 | Ralstonia phage RSK1 | Ralstonia phage RSK1 Ralstonia virus RSK1 Firingavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40471 | 59.591 | Firingavirus | Unclassified | Podoviridae | Ralstonia |
| NC_022914 | Staphylococcus phage phiRS7 | Staphylococcus phage phiRS7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43394 | 34.267 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_022791 | Streptococcus phage phiBHN167 | Streptococcus phage phiBHN167 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37375 | 38.341 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_022758 | Staphylococcus phage YMC/09/04/R1988 | Staphylococcus phage YMC/09/04/R1988 Staphylococcus virus R1988 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44459 | 33.37 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_022757 | Lactobacillus phage PL-1 | Lactobacillus phage PL-1 Lactobacillus virus PL1 Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38880 | 44.882 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_022756 | Lactobacillus phage J-1 | Lactobacillus phage J-1 Lactobacillus virus J1 Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40931 | 44.768 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_022754 | Salmonella phage SETP7 | Salmonella phage SETP7 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42749 | 49.901 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_022753 | Mycobacterium phage Fredward | Mycobacterium phage Fredward unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52282 | 61.241 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022752 | Salmonella phage SETP13 | Salmonella phage SETP13 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42665 | 49.788 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_022750 | Enterobacteria phage fiAA91-ss | Enterobacteria phage fiAA91-ss Escherichia virus fiAA91ss Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33628 | 51.906 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_022749 | Shigella phage SfIV | Shigella phage SfIV Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39758 | 50.297 | Unclassified | Unclassified | Myoviridae | Shigella |
| NC_022747 | Vibrio phage VPUSM 8 | Vibrio phage VPUSM 8 unclassified Longwoodvirus Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34145 | 48.783 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| NC_022746 | Pseudomonas phage MPK6 | Pseudomonas phage MPK6 Pseudomonas virus MPK6 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42957 | 62.279 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_022744 | Erwinia phage FE44 | Erwinia phage FE44 Erwinia virus FE44 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39860 | 48.62 | Berlinvirus | Studiervirinae | Autographiviridae | Erwinia |
| NC_022342 | Propionibacterium phage PHL111M01 | Propionibacterium phage PHL111M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29140 | 54.331 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022341 | Propionibacterium phage PHL113M01 | Propionibacterium phage PHL113M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29200 | 54.096 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022340 | Propionibacterium phage PHL114L00 | Propionibacterium phage PHL114L00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29464 | 54.205 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022339 | Propionibacterium phage PHL037M02 | Propionibacterium phage PHL037M02 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29443 | 53.778 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022338 | Propionibacterium phage PHL060L00 | Propionibacterium phage PHL060L00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29514 | 54.049 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022337 | Propionibacterium phage PHL071N05 | Propionibacterium phage PHL071N05 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29467 | 53.921 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022336 | Propionibacterium phage PHL010M04 | Propionibacterium phage PHL010M04 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.987 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022335 | Propionibacterium phage PHL067M10 | Propionibacterium phage PHL067M10 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29377 | 54.264 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022334 | Propionibacterium phage PHL112N00 | Propionibacterium phage PHL112N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29266 | 54.483 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_022330 | Mycobacterium phage Quink | Mycobacterium phage Quink unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76586 | 62.993 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022329 | Mycobacterium phage PhrostyMug | Mycobacterium phage PhrostyMug unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53636 | 63.677 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022324 | Mycobacterium phage SarFire | Mycobacterium phage SarFire unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53701 | 63.813 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022323 | Escherichia phage JES2013 | Escherichia phage JES2013 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136910 | 43.572 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_021773 | Staphylococcus phage JS01 | Staphylococcus phage JS01 Staphylococcus virus JS01 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43458 | 33.32 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_022094 | Bacillus phage Wip1 | Bacillus phage Wip1 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14319 | 36.846 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| NC_022091 | Pseudomonas phage MPK7 | Pseudomonas phage MPK7 Pseudomonas virus MPK7 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42874 | 62.138 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_022090 | Staphylococcus phage vB_SauM_Remus | Staphylococcus phage vB_SauM_Remus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134643 | 29.973 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_022087 | Mycobacterium phage AnnaL29 | Mycobacterium phage AnnaL29 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53253 | 64.389 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022086 | Mycobacterium phage LittleCherry | Mycobacterium phage LittleCherry unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50690 | 60.933 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022085 | Mycobacterium phage Goku | Mycobacterium phage Goku unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76483 | 62.83 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022071 | Mycobacterium phage Crossroads | Mycobacterium phage Crossroads unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76129 | 58.945 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022070 | Mycobacterium phage Wheeler | Mycobacterium phage Wheeler unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53588 | 63.316 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022069 | Mycobacterium phage Jabbawokkie | Mycobacterium phage Jabbawokkie unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55213 | 61.073 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022068 | Mycobacterium phage Daenerys | Mycobacterium phage Daenerys unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58043 | 61.553 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022067 | Mycobacterium phage Wanda | Mycobacterium phage Wanda unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109960 | 60.75 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022066 | Mycobacterium phage Redno2 | Mycobacterium phage Redno2 unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108297 | 60.879 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022065 | Mycobacterium phage Contagion | Mycobacterium phage Contagion unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74533 | 63.057 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022062 | Mycobacterium phage Trouble | Mycobacterium phage Trouble unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52102 | 63.602 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022060 | Mycobacterium phage Velveteen | Mycobacterium phage Velveteen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54314 | 61.465 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022059 | Mycobacterium phage DrDrey | Mycobacterium phage DrDrey unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77367 | 63 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022058 | Mycobacterium phage Adzzy | Mycobacterium phage Adzzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52519 | 62.556 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022056 | Mycobacterium phage Hamulus | Mycobacterium phage Hamulus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57155 | 61.758 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022055 | Mycobacterium phage Bobi | Mycobacterium phage Bobi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59179 | 61.664 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022054 | Mycobacterium phage Muddy | Mycobacterium phage Muddy Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48228 | 58.827 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022052 | Mycobacterium phage Whirlwind | Mycobacterium phage Whirlwind unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76050 | 59.344 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021865 | Paenibacillus phage phiIBB_P123 | Paenibacillus phage phiIBB_P123 Paenibacillus virus P123 Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41294 | 40.936 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| NC_021864 | Puniceispirillum phage HMO-2011 | Puniceispirillum phage HMO-2011 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55282 | 43.088 | Unclassified | Unclassified | Podoviridae | Puniceispirillum |
| NC_021863 | Staphylococcus phage SA13 | Staphylococcus phage SA13 Staphylococcus virus SA13 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42652 | 35.419 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_021861 | Lactococcus phage BM13 | Lactococcus phage BM13 Lactococcus virus BM13 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30910 | 35.988 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021860 | Lactococcus phage jm2 | Lactococcus phage jm2 Lactococcus virus jm2 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31090 | 34.31 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021857 | Shigella phage SfII | Shigella phage SfII Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41475 | 49.174 | Unclassified | Unclassified | Myoviridae | Shigella |
| NC_021855 | Lactococcus phage phi7 | Lactococcus phage phi7 Lactococcus virus phi7 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32382 | 34.217 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021854 | Lactococcus phage jm3 | Lactococcus phage jm3 Lactococcus virus jm3 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28674 | 34.428 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021853 | Lactococcus phage 340 | Lactococcus phage 340 Lactococcus virus 340 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32337 | 34.487 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021852 | Lactococcus phage P680 | Lactococcus phage P680 Lactococcus virus P680 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29631 | 35.115 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_021801 | Staphylococcus phage SA12 | Staphylococcus phage SA12 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42902 | 34.542 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_021789 | Cellulophaga phage phi19:3 | Cellulophaga phage phi19:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75994 | 32.547 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021783 | Salmonella virus iEPS5 | Salmonella virus iEPS5 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59254 | 56.308 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_021867 | Flavobacterium phage 6H | Flavobacterium phage 6H Flavobacterium virus 6H Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46978 | 31.604 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_021856 | Bacillus phage phiNIT1 | Bacillus phage phiNIT1 Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155631 | 42.118 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_021784 | Thermus phage phi OH2 | Thermus phage phi OH2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38099 | 44.689 | Unclassified | Unclassified | Myoviridae | Thermus |
| NC_021774 | Salmonella phage FSLSP004 | Salmonella phage FSLSP004 Salmonella virus FSLSP004 Stockinghallvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29742 | 52.838 | Stockinghallvirus | Peduovirinae | Myoviridae | Salmonella |
| NC_021780 | Salmonella virus FSLSP088 | Salmonella virus FSLSP088 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59454 | 56.442 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_021779 | Salmonella virus FSLSP030 | Salmonella virus FSLSP030 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59746 | 56.556 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_021777 | Salmonella phage Jersey | Salmonella phage Jersey Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43447 | 49.971 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_021775 | Salmonella phage FSL SP-031 | Salmonella phage FSL SP-031 Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42215 | 51.067 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_021560 | Rhizobium phage RR1-A | Rhizobium phage RR1-A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53102 | 57.164 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| NC_021559 | Prochlorococcus phage P-SSM3 | Prochlorococcus phage P-SSM3 Prochlorococcus virus PSSM3 Ronodorvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179063 | 36.731 | Ronodorvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_021558 | Paenibacillus phage PG1 | Paenibacillus phage PG1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37644 | 42.4 | Unclassified | Unclassified | Siphoviridae | Paenibacillus |
| NC_021557 | Rhizobium phage RR1-B | Rhizobium phage RR1-B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37378 | 58.272 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| NC_021538 | Mycobacterium phage Job42 | Mycobacterium phage Job42 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59626 | 61.158 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021535 | Mycobacterium phage Jobu08 | Mycobacterium phage Jobu08 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50679 | 64.032 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021533 | Mycobacterium phage BTCU-1 | Mycobacterium phage BTCU-1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45942 | 64.246 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021532 | Alteromonas phage vB_AmaP_AD45-P1 | Alteromonas phage vB_AmaP_AD45-P1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103910 | 43.154 | Unclassified | Unclassified | Schitoviridae | Alteromonas |
| NC_021536 | Synechococcus phage S-IOM18 | Synechococcus phage S-IOM18 Synechococcus virus SIOM18 Tefnutvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171797 | 40.572 | Tefnutvirus | Unclassified | Myoviridae | Synechococcus |
| NC_021530 | Synechococcus phage S-CAM8 | Synechococcus phage S-CAM8 unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171407 | 39.333 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| NC_021343 | Burkholderia phage ST79 | Burkholderia phage ST79 Burkholderia virus ST79 Nampongvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35430 | 62.501 | Nampongvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_021339 | Streptomyces phage Zemlya | Streptomyces phage Zemlya Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51077 | 65.673 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_021338 | Mycobacterium phage Chy4 | Mycobacterium phage Chy4 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46639 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021336 | Bacillus phage MG-B1 | Bacillus phage MG-B1 Bacillus virus MGB1 Klosterneuburgvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27190 | 30.747 | Klosterneuburgvirus | Northropvirinae | Salasmaviridae | Bacillus |
| NC_021334 | Mycobacterium phage WIVsmall | Mycobacterium phage WIVsmall unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53359 | 61.009 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021332 | Staphylococcus phage StauST398-3 | Staphylococcus phage StauST398-3 Staphylococcus virus StauST398-3 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41392 | 35.599 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_021326 | Staphylococcus phage StauST398-1 | Staphylococcus phage StauST398-1 Staphylococcus virus StauST398-1 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45242 | 34.464 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_021325 | Clostridium phage vB_CpeS-CP51 | Clostridium phage vB_CpeS-CP51 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39108 | 28.076 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_021324 | Mycobacterium phage CASbig | Mycobacterium phage CASbig unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53369 | 63.612 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021323 | Staphylococcus phage StauST398-2 | Staphylococcus phage StauST398-2 Staphylococcus virus StauST398-2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45572 | 33.341 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_021321 | Halorubrum Tailed Virus 8 | Halorubrum Tailed Virus 8 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74519 | 57.097 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| NC_021320 | Halorubrum Tailed Virus 5 | Halorubrum Tailed Virus 5 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76134 | 56.408 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| NC_021318 | Mycobacterium phage Chy5 | Mycobacterium phage Chy5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51214 | 63.6 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021317 | Salmonella phage L13 | Salmonella phage L13 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21248 | 50.08 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_021315 | Salmonella virus Chi | Salmonella virus Chi Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59407 | 56.51 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_021311 | Mycobacterium phage Phaux | Mycobacterium phage Phaux unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76479 | 62.919 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021309 | Mycobacterium phage Phrux | Mycobacterium phage Phrux unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74711 | 63.054 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021308 | Mycobacterium phage HINdeR | Mycobacterium phage HINdeR unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52617 | 62.839 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021307 | Mycobacterium phage Severus | Mycobacterium phage Severus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 64.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021305 | Mycobacterium phage Murphy | Mycobacterium phage Murphy unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76179 | 62.924 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021304 | Streptomyces phage Sujidade | Streptomyces phage Sujidade Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51552 | 65.706 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_021302 | Mycobacterium phage Fishburne | Mycobacterium phage Fishburne Fishburnevirus Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47109 | 67.267 | Fishburnevirus | Pclasvirinae | Siphoviridae | Mycobacterium |
| NC_021301 | Mycobacterium phage SiSi | Mycobacterium phage SiSi unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56279 | 61.492 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021299 | Mycobacterium phage PegLeg | Mycobacterium phage PegLeg unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80955 | 61.549 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| NC_021298 | Streptomyces phage Lika | Streptomyces phage Lika Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51252 | 65.796 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_021297 | Mycobacterium phage PattyP | Mycobacterium phage PattyP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52057 | 63.59 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_018850 | Pseudomonas phage UFV-P2 | Pseudomonas phage UFV-P2 Pseudomonas virus UFVP2 Vicosavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45517 | 51.471 | Vicosavirus | Unclassified | Podoviridae | Pseudomonas |
| NC_021072 | Synechococcus phage Syn30 | Synechococcus phage Syn30 Synechococcus virus Syn30 Leucotheavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178807 | 39.948 | Leucotheavirus | Unclassified | Myoviridae | Synechococcus |
| NC_021071 | Cyanophage P-RSM1 | Cyanophage P-RSM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177211 | 40.194 | Unclassified | Unclassified | Myoviridae | Prochlorococcus |
| NC_021070 | Vibrio phage martha 12B12 | Vibrio phage martha 12B12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33277 | 45.755 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_021061 | Mycobacterium phage Butters | Mycobacterium phage Butters Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41491 | 65.838 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| NC_020880 | Leuconostoc phage P793 | Leuconostoc phage P793 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26769 | 36.628 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_020879 | Aeromonas phage Aes012 | Aeromonas phage Aes012 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161978 | 41.3 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_020877 | Staphylococcus phage vB_SauM_Romulus | Staphylococcus phage vB_SauM_Romulus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131332 | 30.011 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_020876 | Mycobacterium phage First | Mycobacterium phage First unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53028 | 63.425 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_020873 | Bacillus phage vB_BceM_Bc431v3 | Bacillus phage vB_BceM_Bc431v3 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158621 | 39.978 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_020871 | Listeria phage vB_LmoM_AG20 | Listeria phage vB_LmoM_AG20 Listeria virus AG20 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133057 | 35.933 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_020870 | Leuconostoc phage LN04 | Leuconostoc phage LN04 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25865 | 36.598 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_020878 | Prochlorococcus phage P-GSP1 | Prochlorococcus phage P-GSP1 Prochlorococcus virus PGSP1 Lingvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44945 | 39.551 | Lingvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_020875 | Synechococcus phage S-SSM4 | Synechococcus phage S-SSM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182801 | 39.393 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_020874 | Prochlorococcus phage P-SSP3 | Prochlorococcus phage P-SSP3 Prochlorococcus virus PSSP3 Tritonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46198 | 37.941 | Tritonvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_020867 | Synechococcus phage S-RIP1 | Synechococcus phage S-RIP1 Synechococcus virus SRIP1 Kajamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44892 | 42.93 | Kajamvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_020866 | Rhodovulum phage RS1 | Rhodovulum phage RS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40231 | 62.119 | Unclassified | Unclassified | Siphoviridae | Rhodovulum |
| NC_020865 | Cyanophage KBS-P-1A | Cyanophage KBS-P-1A Sednavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45730 | 47.354 | Sednavirus | Unclassified | Autographiviridae | Synechococcus |
| NC_020859 | Synechococcus phage S-RIM2 R1_1999 | Synechococcus phage S-RIM2 R1_1999 Synechococcus virus SRIM2 Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175430 | 42.155 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| NC_020855 | Cyanophage P-RSM6 | Cyanophage P-RSM6 unclassified Libanvirus Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192497 | 39.316 | Libanvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_020851 | Synechococcus phage S-SKS1 | Synechococcus phage S-SKS1 Synechococcus virus SSKS1 Llyrvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 208007 | 36.027 | Llyrvirus | Unclassified | Myoviridae | Synechococcus |
| NC_020845 | Prochlorococcus phage MED4-213 | Prochlorococcus phage MED4-213 Prochlorococcus virus MED4-213 Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180977 | 37.764 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_020844 | Salicola phage CGphi29 | Salicola phage CGphi29 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40695 | 56.422 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_020841 | Psychrobacter phage pOW20-A | Psychrobacter phage pOW20-A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40803 | 44.056 | Unclassified | Unclassified | Myoviridae | Psychrobacter |
| NC_020839 | Rhodobacter phage RC1 | Rhodobacter phage RC1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39573 | 62.272 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| NC_020838 | Synechococcus phage S-RIP2 | Synechococcus phage S-RIP2 Synechococcus virus SRIP2 Sednavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45728 | 47.343 | Sednavirus | Unclassified | Autographiviridae | Synechococcus |
| NC_020837 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 198013 | 43.025 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| NC_020835 | Prochlorococcus phage P-SSP10 | Prochlorococcus phage P-SSP10 Prochlorococcus virus PSSP10 Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47325 | 39.231 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_020489 | Rhodobacter phage RcapNL | Rhodobacter phage RcapNL Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40489 | 65.112 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| NC_020486 | Synechococcus phage S-RIM8 A.HR1 | Synechococcus phage S-RIM8 A.HR1 unclassified Neptunevirus Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171211 | 40.631 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| NC_020483 | Pelagibacter phage HTVC019P | Pelagibacter phage HTVC019P Pelagibacter virus HTVC019P Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42084 | 34.046 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| NC_020416 | Salmonella phage vB_SenM-S16 | Salmonella phage vB_SenM-S16 Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160221 | 36.877 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| NC_020077 | Sulfolobus virus STSV2 | Sulfolobus virus STSV2 unclassified Bicaudaviridae Bicaudaviridae Viruses | 76107 | 35.098 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_020204 | Klebsiella phage JD001 | Klebsiella phage JD001 Klebsiella virus JD001 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48814 | 48.535 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| NC_020201 | Pectobacterium phage phiTE | Pectobacterium phage phiTE Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142349 | 50.137 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| NC_020199 | Staphylococcus phage phi7401PVL | Staphylococcus phage phi7401PVL unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47252 | 33.044 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_019782 | Lactobacillus phage phiAQ113 | Lactobacillus phage phiAQ113 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36566 | 36.52 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| NC_020197 | Streptococcus phage TP-J34 | Streptococcus phage TP-J34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45606 | 38.819 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_019704 | Enterobacteria phage mEp237 | Enterobacteria phage mEp237 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44375 | 51.432 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_019935 | Pseudomonas phage KPP12 | Pseudomonas phage KPP12 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64144 | 55.623 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_019934 | Cronobacter phage ENT39118 | Cronobacter phage ENT39118 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39012 | 53.063 | Unclassified | Hendrixvirinae | Siphoviridae | Cronobacter |
| NC_019932 | Erwinia phage ENT90 | Erwinia phage ENT90 Erwinia virus ENT90 Entnonagintavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29564 | 55.811 | Entnonagintavirus | Peduovirinae | Myoviridae | Erwinia |
| NC_019931 | Helicobacter phage KHP40 | Helicobacter phage KHP40 Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26449 | 35.941 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| NC_019929 | Erwinia phage phiEaH2 | Erwinia phage phiEaH2 Erskinevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 243050 | 51.29 | Erskinevirus | Unclassified | Myoviridae | Erwinia |
| NC_019928 | Helicobacter phage KHP30 | Helicobacter phage KHP30 Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26215 | 35.77 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| NC_019927 | Cronobacter phage ENT47670 | Cronobacter phage ENT47670 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47611 | 51.593 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| NC_019921 | Staphylococcus phage phi5967PVL | Staphylococcus phage phi5967PVL unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42461 | 33.311 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_019915 | Staphylococcus phage StB20 | Staphylococcus phage StB20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40917 | 34.067 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_019914 | Staphylococcus phage StB27 | Staphylococcus phage StB27 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40071 | 33.216 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_019912 | Bacillus phage vB_BtS_BMBtp2 | Bacillus phage vB_BtS_BMBtp2 Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36932 | 37.788 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| NC_019909 | Yersinia phage phiR1-RT | Yersinia phage phiR1-RT Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168809 | 34.544 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| NC_019813 | Pseudomonas phage vB_PaeP_p2-10_Or1 | Pseudomonas phage vB_PaeP_p2-10_Or1 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44030 | 51.994 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_019769 | Escherichia phage HK542 | Escherichia phage HK542 Wongtaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38964 | 50.914 | Wongtaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019768 | Escherichia phage HK106 | Escherichia phage HK106 Wanchaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41468 | 49.342 | Wanchaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019767 | Escherichia phage HK544 | Escherichia phage HK544 Kwaitsingvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40155 | 49.797 | Kwaitsingvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019726 | Staphylococcus phage JD007 | Staphylococcus phage JD007 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141836 | 30.371 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_019723 | Escherichia phage HK630 | Escherichia phage HK630 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47090 | 50.053 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_019722 | Vibrio phage vB_VpaM_MAR | Vibrio phage vB_VpaM_MAR Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41351 | 51.336 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| NC_019721 | Enterobacterial phage mEp390 | Enterobacterial phage mEp390 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40029 | 51.678 | Unclassified | Hendrixvirinae | Siphoviridae | Enterobacteria |
| NC_019720 | Enterobacteria phage mEp213 | Enterobacteria phage mEp213 Enterobacteria virus mEp213 Aguilavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44120 | 50.977 | Aguilavirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_019719 | Escherichia phage HK633 | Escherichia phage HK633 Saikungvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41528 | 49.653 | Saikungvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019718 | Enterobacteria phage vB_EcoS_Rogue1 | Enterobacteria phage vB_EcoS_Rogue1 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45805 | 44.214 | Rogunavirus | Rogunavirinae | Drexlerviridae | Enterobacteria |
| NC_019717 | Enterobacteria phage HK225 | Enterobacteria phage HK225 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45366 | 51.955 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_019716 | Enterobacteria phage mEp460 | Enterobacteria phage mEp460 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44510 | 50.874 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_019715 | Escherichia phage mEp234 | Escherichia phage mEp234 Wanchaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39578 | 49.586 | Wanchaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019714 | Escherichia phage HK446 | Escherichia phage HK446 Kwaitsingvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39026 | 50.102 | Kwaitsingvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019713 | Vibrio phage vB_VpaS_MAR10 | Vibrio phage vB_VpaS_MAR10 Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78751 | 49.703 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| NC_019711 | Escherichia phage HK629 | Escherichia phage HK629 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47288 | 49.638 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_019710 | Enterobacteria phage HK140 | Enterobacteria phage HK140 Enterobacteria virus HK140 Yautsimvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40710 | 49.912 | Yautsimvirus | Hendrixvirinae | Siphoviridae | Enterobacteria |
| NC_019709 | Escherichia phage mEpX1 | Escherichia phage mEpX1 Cuauhtlivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41567 | 49.311 | Cuauhtlivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019708 | Enterobacteria phage mEp235 | Enterobacteria phage mEp235 Enterobacteria virus mEp235 Nochtlivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37595 | 50.015 | Nochtlivirus | Hendrixvirinae | Siphoviridae | Enterobacteria |
| NC_019706 | Enterobacteria phage mEp043 c-1 | Enterobacteria phage mEp043 c-1 Enterobacteria virus mEp043 Aguilavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42780 | 50.79 | Aguilavirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_019705 | Escherichia phage mEpX2 | Escherichia phage mEpX2 Shamshuipovirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38759 | 50.076 | Shamshuipovirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_019550 | Liberibacter phage SC2 | Liberibacter phage SC2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38997 | 39.377 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| NC_019549 | Liberibacter phage SC1 | Liberibacter phage SC1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40048 | 41.196 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| NC_019545 | Salmonella phage SPN3UB | Salmonella phage SPN3UB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47355 | 49.613 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_019544 | Deep-sea thermophilic phage D6E | Deep-sea thermophilic phage D6E Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49335 | 45.998 | Unclassified | Unclassified | Myoviridae | Unspecified |
| NC_019541 | Acinetobacter phage YMC/09/02/B1251 | Acinetobacter phage YMC/09/02/B1251 Acinetobacter virus B1251 Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45364 | 39.053 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| NC_019539 | Salmonella phage Ent1 | Salmonella phage Ent1 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42391 | 49.789 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_019527 | Aeromonas phage vB_AsaM-56 | Aeromonas phage vB_AsaM-56 Aeromonas virus 56 Popoffvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43551 | 55.425 | Popoffvirus | Unclassified | Myoviridae | Aeromonas |
| NC_019526 | Klebsiella phage vB_KleM_RaK2 | Klebsiella phage vB_KleM_RaK2 Klebsiella virus RaK2 Alcyoneusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 345809 | 31.702 | Alcyoneusvirus | Unclassified | Myoviridae | Klebsiella |
| NC_019522 | Pectobacterium phage ZF40 | Pectobacterium phage ZF40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48454 | 50.204 | Unclassified | Unclassified | Myoviridae | Pectobacterium |
| NC_019517 | Escherichia phage FV3 | Escherichia phage FV3 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136947 | 43.699 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_019516 | Cyanophage S-TIM5 | Cyanophage S-TIM5 Synechococcus virus STIM5 Aurunvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161440 | 40.458 | Aurunvirus | Unclassified | Myoviridae | Synechococcus |
| NC_019515 | Bacillus phage BCD7 | Bacillus phage BCD7 Bacillus virus BCD7 Becedseptimavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93839 | 38.043 | Becedseptimavirus | Unclassified | Myoviridae | Bacillus |
| NC_019513 | Staphylococcus phage SMSAP5 | Staphylococcus phage SMSAP5 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45552 | 33.415 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_019512 | Helicobacter phage 1961P | Helicobacter phage 1961P Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26836 | 35.516 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| NC_019511 | Staphylococcus virus SA11 | Staphylococcus virus SA11 Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136326 | 30.038 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_019509 | Cronobacter phage ESP2949-1 | Cronobacter phage ESP2949-1 Kyungwonvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49116 | 50.092 | Kyungwonvirus | Unclassified | Drexlerviridae | Cronobacter |
| NC_019505 | Escherichia phage wV7 | Escherichia phage wV7 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166452 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_019503 | Escherichia phage ime09 | Escherichia phage ime09 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166499 | 35.675 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_019502 | Bacillus phage phIS3501 | Bacillus phage phIS3501 Bacillus virus phIS3501 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44401 | 34.864 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| NC_019501 | Enterobacteria phage IME10 | Enterobacteria phage IME10 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39646 | 47.495 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| NC_019500 | Escherichia phage Bp7 | Escherichia phage Bp7 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168066 | 39.493 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_019490 | Riemerella phage RAP44 | Riemerella phage RAP44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49329 | 34.74 | Unclassified | Unclassified | Siphoviridae | Riemerella |
| NC_019489 | Lactobacillus phage Sha1 | Lactobacillus phage Sha1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41726 | 40.605 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_019488 | Salmonella phage RE2010 | Salmonella phage RE2010 Salmonella virus RE2010 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34117 | 51.021 | Felsduovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_019487 | Bacillus phage SP-10 | Bacillus phage SP-10 unclassified Spounavirinae Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143986 | 40.487 | Unclassified | Spounavirinae | Herelleviridae | Bacillus |
| NC_019486 | Lactobacillus phage LF1 | Lactobacillus phage LF1 Lactobacillus virus LF1 Lafunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42606 | 45.005 | Lafunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_019456 | Lactobacillus phage JCL1032 | Lactobacillus phage JCL1032 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49433 | 44.246 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_019455 | Haemophilus phage SuMu | Haemophilus phage SuMu Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37151 | 41.872 | Unclassified | Unclassified | Myoviridae | Haemophilus |
| NC_019451 | Pseudomonas phage NH-4 | Pseudomonas phage NH-4 Pseudomonas virus NH4 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66116 | 55.549 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_019448 | Staphylococcus phage G15 | Staphylococcus phage G15 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139806 | 30.228 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_019447 | Brucella phage Pr | Brucella phage Pr Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.182 | Perisivirus | Unclassified | Podoviridae | Brucella |
| NC_019446 | Brucella phage Tb | Brucella phage Tb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41148 | 48.194 | Perisivirus | Unclassified | Podoviridae | Brucella |
| NC_019445 | Escherichia phage TL-2011b | Escherichia phage TL-2011b Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44784 | 47.053 | Unclassified | Unclassified | Podoviridae | Escherichia |
| NC_019444 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176043 | 39.119 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| NC_019443 | Synechococcus phage metaG-MbCM1 | Synechococcus phage metaG-MbCM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172879 | 39.762 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_019442 | Escherichia phage TL-2011c | Escherichia phage TL-2011c Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60523 | 50.297 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_017674 | Pseudomonas phage JG024 | Pseudomonas phage JG024 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66275 | 55.621 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_019422 | Clostridium phage phiMMP04 | Clostridium phage phiMMP04 Clostridioides virus phiMMP04 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31674 | 29.971 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| NC_019421 | Clostridium phage phiMMP02 | Clostridium phage phiMMP02 Clostridioides virus phiMMP02 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48396 | 29.552 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| NC_019418 | Streptococcus phage phiNJ2 | Streptococcus phage phiNJ2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37282 | 39.418 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_019417 | Salmonella virus SPN19 | Salmonella virus SPN19 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59203 | 56.524 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| NC_019416 | Stenotrophomonas phage IME15 | Stenotrophomonas phage IME15 Stenotrophomonas virus IME15 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38513 | 53.652 | Teseptimavirus | Studiervirinae | Autographiviridae | Stenotrophomonas |
| NC_019414 | Streptomyces phage R4 | Streptomyces phage R4 Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51071 | 66.962 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_019404 | Escherichia phage vB_EcoS_ACG-M12 | Escherichia phage vB_EcoS_ACG-M12 Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46054 | 43.512 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_019403 | Escherichia phage vB_EcoP_ACG-C91 | Escherichia phage vB_EcoP_ACG-C91 Escherichia virus ACGC91 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43731 | 45.281 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_019401 | Cronobacter phage vB_CsaM_GAP32 | Cronobacter phage vB_CsaM_GAP32 Cronobacter virus GAP32 Mimasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 358663 | 35.551 | Mimasvirus | Unclassified | Myoviridae | Cronobacter |
| NC_019400 | Cronobacter phage vB_CsaM_GAP31 | Cronobacter phage vB_CsaM_GAP31 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147940 | 46.298 | Seunavirus | Vequintavirinae | Myoviridae | Cronobacter |
| NC_019399 | Escherichia phage vB_EcoM_ACG-C40 | Escherichia phage vB_EcoM_ACG-C40 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167396 | 35.237 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_018863 | Bacillus phage B4 | Bacillus phage B4 Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162596 | 37.71 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_018857 | Bacillus phage BPS13 | Bacillus phage BPS13 Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158305 | 38.751 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_018856 | Bacillus phage Bastille | Bacillus phage Bastille Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153962 | 38.144 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_018854 | Escherichia phage ECBP1 | Escherichia phage ECBP1 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69855 | 42.73 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_018853 | Streptomyces virus TG1 | Streptomyces virus TG1 Tigunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40474 | 64.649 | Tigunavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_018852 | Propionibacterium phage P100D | Propionibacterium phage P100D Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29506 | 53.762 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018851 | Propionibacterium phage ATCC29399B_C | Propionibacterium phage ATCC29399B_C Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29516 | 53.998 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018849 | Propionibacterium phage P105 | Propionibacterium phage P105 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29202 | 54.174 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018848 | Streptomyces phage SV1 | Streptomyces phage SV1 unclassified Picardvirus Picardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37612 | 72.692 | Picardvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_018847 | Propionibacterium phage ATCC29399B_T | Propionibacterium phage ATCC29399B_T Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29516 | 54.005 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018846 | Escherichia phage P13374 | Escherichia phage P13374 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60894 | 50.23 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| NC_018845 | Propionibacterium phage P104A | Propionibacterium phage P104A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29371 | 53.982 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018843 | Salmonella phage SSU5 | Salmonella phage SSU5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103299 | 51.105 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_018842 | Propionibacterium phage P1.1 | Propionibacterium phage P1.1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29348 | 54.365 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018841 | Propionibacterium phage P101A | Propionibacterium phage P101A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29574 | 54.135 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018840 | Propionibacterium phage P100_1 | Propionibacterium phage P100_1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29612 | 54.069 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018839 | Propionibacterium phage P14 | Propionibacterium phage P14 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29729 | 54.082 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018838 | Propionibacterium phage P100_A | Propionibacterium phage P100_A Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29505 | 53.832 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018836 | Streptomyces phage phiHau3 | Streptomyces phage phiHau3 Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50255 | 67.834 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_018834 | Propionibacterium phage P9.1 | Propionibacterium phage P9.1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29214 | 54.121 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_018832 | Providencia phage Redjac | Providencia phage Redjac Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58104 | 49.499 | Unclassified | Unclassified | Siphoviridae | Providencia |
| NC_018454 | Cronobacter phage phiES15 | Cronobacter phage phiES15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39974 | 53.542 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| NC_018284 | Staphylococcus virus IPLA7 | Staphylococcus virus IPLA7 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42123 | 34.755 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_018281 | Staphylococcus virus IPLA5 | Staphylococcus virus IPLA5 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43581 | 34.726 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_018279 | Salmonella phage vB_SosS_Oslo | Salmonella phage vB_SosS_Oslo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49116 | 48.738 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_018274 | Pseudomonas phage MP42 | Pseudomonas phage MP42 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36847 | 64.198 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_018273 | Leuconostoc phage Lmd1 | Leuconostoc phage Lmd1 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26201 | 36.403 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_018277 | Staphylococcus phage SpaA1 | Staphylococcus phage SpaA1 unclassified Beceayunavirus Beceayunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42784 | 35.635 | Beceayunavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_018275 | Salmonella phage vB_SemP_Emek | Salmonella phage vB_SemP_Emek Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39783 | 47.651 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_018285 | Streptococcus phage YMC-2011 | Streptococcus phage YMC-2011 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40758 | 41.239 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_018264 | Thermoanaerobacterium phage THSA-485A | Thermoanaerobacterium phage THSA-485A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41938 | 36.771 | Unclassified | Unclassified | Siphoviridae | Thermoanaerobacterium |
| NC_008721 | Streptococcus phage SMP | Streptococcus phage SMP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36019 | 41.581 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_018085 | Bacillus phage BtCS33 | Bacillus phage BtCS33 Bacillus virus BtCS33 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41992 | 35.223 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| NC_017985 | Salmonella phage SPN9CC | Salmonella phage SPN9CC Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40128 | 47.326 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_017984 | Acinetobacter phage AP22 | Acinetobacter phage AP22 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46387 | 37.743 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_017978 | Clostridium phage PhiS63 | Clostridium phage PhiS63 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33609 | 27.478 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_017973 | Mycobacterium phage SWU1 | Mycobacterium phage SWU1 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52474 | 62.406 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_017968 | Staphylococcus phage TEM123 | Staphylococcus phage TEM123 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43786 | 34.036 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_017865 | Pseudomonas phage vB_Pae-TbilisiM32 | Pseudomonas phage vB_Pae-TbilisiM32 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42966 | 62.27 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_017732 | Enterococcus phage EF62phi | Enterococcus phage EF62phi Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30505 | 32.657 | Unclassified | Unclassified | Podoviridae | Enterococcus |
| NC_017702 | Lactococcus phage ASCC281 | Lactococcus phage ASCC281 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32023 | 34.766 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_017698 | Lactococcus phage ASCC465 | Lactococcus phage ASCC465 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31772 | 34.644 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_017696 | Lactococcus phage ASCC532 | Lactococcus phage ASCC532 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32384 | 34.687 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_017695 | Lactococcus phage ASCC273 | Lactococcus phage ASCC273 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32582 | 34.528 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_016899 | Thermococcus prieurii virus 1 | Thermococcus prieurii virus 1 unclassified archaeal viruses Viruses | 21592 | 49.537 | Unclassified | Unclassified | Unclassified | Thermococcus |
| NC_016763 | Salmonella phage SE2 | Salmonella phage SE2 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43221 | 49.645 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_016770 | Bacteroides phage B124-14 | Bacteroides phage B124-14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47159 | 38.75 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| NC_016767 | Erwinia phage PEp14 | Erwinia phage PEp14 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60714 | 62.78 | Unclassified | Unclassified | Podoviridae | Erwinia |
| NC_016765 | Pseudomonas virus PMG1 | Pseudomonas virus PMG1 Detrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54024 | 57.467 | Detrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_016762 | Pseudomonas phage phi297 | Pseudomonas phage phi297 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49135 | 62.147 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_016761 | Salmonella phage SPN1S | Salmonella phage SPN1S Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38684 | 50.163 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| NC_006940 | Salmonella phage SS3e | Salmonella phage SS3e Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40793 | 50.08 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| NC_016652 | Rhodococcus phage REQ2 | Rhodococcus phage REQ2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49330 | 65.406 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_016659 | Cyanophage NATL2A-133 | Cyanophage NATL2A-133 Prochlorococcus virus NATL2A133 Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47536 | 39.871 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_016658 | Cyanophage NATL1A-7 | Cyanophage NATL1A-7 Prochlorococcus virus NATL1A7 Cheungvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47741 | 38.732 | Cheungvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_016657 | Cyanophage 9515-10a | Cyanophage 9515-10a Prochlorococcus virus 951510a Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47055 | 39.197 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_016656 | Cyanophage P-SSP2 | Cyanophage P-SSP2 Synechococcus virus PSSP2 Tritonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45890 | 37.932 | Tritonvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_016571 | Pseudomonas phage OBP | Pseudomonas phage OBP Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 284757 | 43.455 | Petsuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_016568 | Helicobacter phage phiHP33 | Helicobacter phage phiHP33 unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24645 | 37.338 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| NC_016565 | Staphylococcus phage S24-1 | Staphylococcus phage S24-1 Staphylococcus virus S24-1 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18168 | 28.908 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_016563 | Bacillus phage W.Ph. | Bacillus phage W.Ph. Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156897 | 36.447 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_016564 | Planktothrix phage PaV-LD | Planktothrix phage PaV-LD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95299 | 41.45 | Unclassified | Unclassified | Siphoviridae | Planktothrix |
| NC_016071 | Salmonella phage PVPSE1 | Salmonella phage PVPSE1 Salmonella virus PVPSE1 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145964 | 45.607 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| NC_016165 | Rhodobacter phage RcapMu | Rhodobacter phage RcapMu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39283 | 64.896 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| NC_016161 | Pediococcus phage cIP1 | Pediococcus phage cIP1 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38013 | 47.607 | Coetzeevirus | Unclassified | Siphoviridae | Pediococcus |
| NC_016160 | Escherichia phage HK75 | Escherichia phage HK75 Saikungvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36661 | 50.19 | Saikungvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_016158 | Escherichia phage HK639 | Escherichia phage HK639 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49576 | 52.451 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| NC_015933 | Escherichia phage vB_EcoP_G7C | Escherichia phage vB_EcoP_G7C Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72917 | 43.416 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_015719 | Escherichia phage K30 | Escherichia phage K30 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40940 | 51.426 | Przondovirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_015569 | Synechococcus phage S-CRM01 | Synechococcus phage S-CRM01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178563 | 39.745 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_015568 | Clostridium phage phiCD38-2 | Clostridium phage phiCD38-2 Clostridioides virus phiCD38-2 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41090 | 30.83 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| NC_015457 | Shigella phage Shfl2 | Shigella phage Shfl2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165919 | 35.567 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_015456 | Shigella phage Shfl1 | Shigella phage Shfl1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50661 | 45.408 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_015454 | Propionibacterium phage PAD20 | Propionibacterium phage PAD20 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29074 | 54.1 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_015453 | Propionibacterium phage PAS50 | Propionibacterium phage PAS50 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29017 | 53.972 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_015254 | Brochothrix phage BL3 | Brochothrix phage BL3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41518 | 36.377 | Unclassified | Unclassified | Siphoviridae | Brochothrix |
| NC_015295 | Erwinia phage phiEt88 | Erwinia phage phiEt88 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47279 | 47.33 | Unclassified | Unclassified | Myoviridae | Erwinia |
| NC_015274 | Streptococcus phage Dp-1 | Streptococcus phage Dp-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56506 | 40.337 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_015273 | Burkholderia phage KS14 | Burkholderia phage KS14 Burkholderia virus KS14 Kisquattuordecimvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32317 | 62.28 | Kisquattuordecimvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_015266 | Burkholderia phage KL3 | Burkholderia phage KL3 Burkholderia virus KL3 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40555 | 63.225 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_015265 | Burkholderia phage KS5 | Burkholderia phage KS5 Burkholderia virus KS5 Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37236 | 63.707 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_015263 | Lactococcus phage 949 | Lactococcus phage 949 Lactococcus virus 949 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114768 | 32.689 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_015253 | Brochothrix phage A9 | Brochothrix phage A9 Brochothrix virus A9 Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127065 | 41.424 | Unclassified | Unclassified | Herelleviridae | Brochothrix |
| NC_015252 | Brochothrix phage NF5 | Brochothrix phage NF5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36953 | 37.296 | Unclassified | Unclassified | Siphoviridae | Brochothrix |
| NC_015290 | Prochlorococcus phage P-SSM7 | Prochlorococcus phage P-SSM7 Prochlorococcus virus PSSM7 Palaemonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182180 | 37.086 | Palaemonvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015289 | Synechococcus phage S-SSM5 | Synechococcus phage S-SSM5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176184 | 39.962 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_015288 | Prochlorococcus phage Syn1 | Prochlorococcus phage Syn1 Prochlorococcus virus Syn1 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191195 | 40.617 | Vellamovirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015287 | Synechococcus phage S-SSM7 | Synechococcus phage S-SSM7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 232878 | 39.116 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_015286 | Synechococcus phage Syn19 | Synechococcus phage Syn19 Synechococcus virus Syn19 Pontusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175230 | 41.251 | Pontusvirus | Unclassified | Myoviridae | Synechococcus |
| NC_015285 | Prochlorococcus phage Syn33 | Prochlorococcus phage Syn33 Prochlorococcus virus Syn33 Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174285 | 39.57 | Brizovirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015284 | Prochlorococcus phage P-HM2 | Prochlorococcus phage P-HM2 unclassified Eurybiavirus Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 183806 | 38.115 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015283 | Prochlorococcus phage P-RSM4 | Prochlorococcus phage P-RSM4 unclassified Thaumasvirus Thaumasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176428 | 37.644 | Thaumasvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015282 | Synechococcus phage S-SM1 | Synechococcus phage S-SM1 Synechococcus virus SSM1 Thetisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174079 | 41.138 | Thetisvirus | Unclassified | Myoviridae | Synechococcus |
| NC_015281 | Synechococcus phage S-ShM2 | Synechococcus phage S-ShM2 Synechococcus virus SShM2 Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179563 | 41.107 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| NC_015280 | Prochlorococcus phage P-HM1 | Prochlorococcus phage P-HM1 unclassified Eurybiavirus Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181044 | 37.849 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_015279 | Synechococcus phage S-SM2 | Synechococcus phage S-SM2 unclassified Bellamyvirus Bellamyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 190789 | 40.426 | Bellamyvirus | Unclassified | Myoviridae | Synechococcus |
| NC_015271 | Salmonella phage Vi06 | Salmonella phage Vi06 Salmonella virus Vi06 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38368 | 48.926 | Teseptimavirus | Studiervirinae | Autographiviridae | Salmonella |
| NC_015262 | Clostridium phage phiCD6356 | Clostridium phage phiCD6356 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37664 | 28.38 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_015250 | Acinetobacter phage 133 | Acinetobacter phage 133 Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159801 | 39.672 | Unclassified | Tevenvirinae | Myoviridae | Acinetobacter |
| NC_015249 | Escherichia phage 285P | Escherichia phage 285P Escherichia virus 285P Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39270 | 48.732 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_015158 | Vibrio phage ICP2 | Vibrio phage ICP2 Vibrio virus ICP2 Icepovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49675 | 42.677 | Icepovirus | Unclassified | Zobellviridae | Vibrio |
| NC_014900 | Salmonella phage ST160 | Salmonella phage ST160 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40986 | 47.06 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_014792 | Escherichia phage vB_EcoM_VR7 | Escherichia phage vB_EcoM_VR7 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169285 | 40.333 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_006884 | Prochlorococcus phage P-SSM4 | Prochlorococcus phage P-SSM4 Prochlorococcus virus PSSM4 Ronodorvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178249 | 36.744 | Ronodorvirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_006883 | Prochlorococcus phage P-SSM2 | Prochlorococcus phage P-SSM2 Prochlorococcus virus PSSM2 Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252401 | 35.506 | Salacisavirus | Unclassified | Myoviridae | Prochlorococcus |
| NC_014662 | Enterobacter phage CC31 | Enterobacter phage CC31 Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165540 | 39.918 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| NC_014661 | Acinetobacter phage Acj61 | Acinetobacter phage Acj61 Acinetobacter virus Acj61 Lasallevirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164093 | 39.006 | Lasallevirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_014660 | Acinetobacter phage Ac42 | Acinetobacter phage Ac42 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167716 | 36.366 | Unclassified | Tevenvirinae | Myoviridae | Acinetobacter |
| NC_014595 | Shigella phage SP18 | Shigella phage SP18 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170605 | 40.445 | Gaprivervirus | Tevenvirinae | Myoviridae | Shigella |
| NC_014460 | Staphylococcus phage SAP-26 | Staphylococcus phage SAP-26 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41207 | 34.016 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_014260 | Escherichia phage IME08 | Escherichia phage IME08 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172253 | 39.591 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_014229 | Streptomyces phage phiSASD1 | Streptomyces phage phiSASD1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37068 | 66.254 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| NC_013649 | Klebsiella phage KP34 | Klebsiella phage KP34 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43809 | 54.073 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| NC_013696 | Enterococcus phage phiEf11 | Enterococcus phage phiEf11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42822 | 34.389 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_013643 | Enterococcus phage phiFL2A | Enterococcus phage phiFL2A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36270 | 34.607 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| NC_013644 | Enterococcus phage phiFL4A | Enterococcus phage phiFL4A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37856 | 37.777 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_013645 | Streptococcus phage Abc2 | Streptococcus phage Abc2 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34882 | 38.986 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_013646 | Enterococcus phage phiFL1A | Enterococcus phage phiFL1A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38764 | 34.014 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| NC_013647 | Klebsiella phage KP32 | Klebsiella phage KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41119 | 52.37 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_013648 | Enterococcus phage phiFL3A | Enterococcus phage phiFL3A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39576 | 34.546 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| NC_013594 | Escherichia phage D108 | Escherichia phage D108 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37235 | 51.76 | Muvirus | Unclassified | Myoviridae | Escherichia |
| NC_013600 | Sodalis phage SO1 | Sodalis phage SO1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45169 | 54.582 | Dhillonvirus | Unclassified | Siphoviridae | Sodalis |
| NC_013599 | Xylella phage Xfas53 | Xylella phage Xfas53 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36674 | 56.757 | Unclassified | Unclassified | Podoviridae | Xylella |
| NC_013598 | Streptococcus phage ALQ13.2 | Streptococcus phage ALQ13.2 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35525 | 39.395 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_013597 | Aggregatibacter phage S1249 | Aggregatibacter phage S1249 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43970 | 42.443 | Unclassified | Unclassified | Myoviridae | Aggregatibacter |
| NC_013588 | Sulfolobus spindle-shaped virus 7 | Sulfolobus spindle-shaped virus 7 Alphafusellovirus Fuselloviridae Viruses | 17602 | 38.984 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_013587 | Sulfolobus spindle-shaped virus 6 | Sulfolobus spindle-shaped virus 6 Betafusellovirus Fuselloviridae Viruses | 15684 | 38.192 | Betafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_013197 | Thermus phage P23-77 | Thermus phage P23-77 Gammasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 17036 | 67.95 | Gammasphaerolipovirus | Unclassified | Sphaerolipoviridae | Thermus |
| NC_013195 | Staphylococcus phage P954 | Staphylococcus phage P954 Staphylococcus virus P954 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40761 | 33.993 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_013152 | Lactococcus phage CB14 | Lactococcus phage CB14 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29459 | 34.804 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_013155 | Lactococcus phage CB13 | Lactococcus phage CB13 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32182 | 34.7 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_013154 | Lactococcus virus CB20 | Lactococcus virus CB20 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28625 | 35.025 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_013153 | Lactococcus phage CB19 | Lactococcus phage CB19 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28643 | 35.146 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_013085 | Synechococcus phage S-RSM4 | Synechococcus phage S-RSM4 unclassified Cymopoleiavirus Cymopoleiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194454 | 41.12 | Cymopoleiavirus | Unclassified | Myoviridae | Synechococcus |
| NC_013059 | Salmonella phage g341c | Salmonella phage g341c Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 47.402 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_013055 | Burkholderia phage KS9 | Burkholderia phage KS9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39896 | 60.68 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| NC_012884 | Streptococcus phage M102 | Streptococcus phage M102 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31147 | 39.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_012784 | Staphylococcus phage phiPVL-CN125 | Staphylococcus phage phiPVL-CN125 Staphylococcus virus CN125 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 33.568 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_012756 | Streptococcus phage PH10 | Streptococcus phage PH10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31276 | 39.49 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_012753 | Streptococcus phage 5093 | Streptococcus phage 5093 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37184 | 38.03 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_012749 | Escherichia phage wV8 | Escherichia phage wV8 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88487 | 38.894 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| NC_012741 | Escherichia phage JS10 | Escherichia phage JS10 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171451 | 39.515 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_012740 | Escherichia phage JSE | Escherichia phage JSE Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166418 | 40.512 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_012638 | Escherichia virus RB14 | Escherichia virus RB14 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165429 | 35.325 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_012635 | Enterobacteria phage RB51 | Enterobacteria phage RB51 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168394 | 35.253 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| NC_012418 | Pseudomonas phage phikF77 | Pseudomonas phage phikF77 Pseudomonas virus kF77 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43152 | 62.868 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_012027 | Mycobacterium phage Phlyer | Mycobacterium phage Phlyer unclassified Pipefishvirus Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69378 | 67.484 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_011976 | Salmonella phage epsilon34 | Salmonella phage epsilon34 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43016 | 47.264 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_011802 | Salmonella phage SE1 (in:P22virus) | Salmonella phage SE1 (in:P22virus) Salmonella virus SE1Spa Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 46.985 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_011801 | Lactobacillus phage Lv-1 | Lactobacillus phage Lv-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38934 | 37.042 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_011810 | Pseudomonas phage PB1 | Pseudomonas phage PB1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65764 | 54.928 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_011756 | Pseudomonas phage SN | Pseudomonas phage SN Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66390 | 55.582 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_011703 | Pseudomonas phage 14-1 | Pseudomonas phage 14-1 Pseudomonas virus 14-1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66235 | 55.593 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_011645 | Bacillus phage TP21-L | Bacillus phage TP21-L Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37456 | 37.802 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| NC_011613 | Pseudomonas phage MP29 | Pseudomonas phage MP29 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36632 | 64.329 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_011611 | Pseudomonas phage MP38 | Pseudomonas phage MP38 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36885 | 64.514 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_011614 | Staphylococcus phage vB_SauS-phiIPLA88 | Staphylococcus phage vB_SauS-phiIPLA88 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42526 | 34.913 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_011612 | Staphylococcus phage phiSauS-IPLA35 | Staphylococcus phage phiSauS-IPLA35 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45344 | 33.246 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_011589 | Stenotrophomonas phage S1 | Stenotrophomonas phage S1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40287 | 63.691 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| NC_011534 | Kluyvera phage Kvp1 | Kluyvera phage Kvp1 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39472 | 48.642 | Berlinvirus | Studiervirinae | Autographiviridae | Kluyvera |
| NC_011551 | Bacteriophage APSE-2 | Bacteriophage APSE-2 Hamiltonella virus APSE2 Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39867 | 42.91 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| NC_011398 | Clostridium virus phiCD27 | Clostridium virus phiCD27 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50930 | 29.327 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| NC_011373 | Pseudomonas phage PAJU2 | Pseudomonas phage PAJU2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46872 | 56.264 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_011357 | Stx2-converting phage 1717 | Stx2-converting phage 1717 Escherichia virus 1717 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62147 | 50.915 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| NC_011356 | Enterobacteria phage YYZ-2008 | Enterobacteria phage YYZ-2008 Enterobacteria virus YYZ2008 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54896 | 51.115 | Pankowvirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_011344 | Staphylococcus phage phi2958PVL | Staphylococcus phage phi2958PVL Staphylococcus virus phi2958PVL Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47342 | 33.04 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_011289 | Mycobacterium phage Ramsey | Mycobacterium phage Ramsey Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58578 | 61.219 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011287 | Mycobacterium phage Pacc40 | Mycobacterium phage Pacc40 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58554 | 61.263 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011288 | Mycobacterium phage Fruitloop | Mycobacterium phage Fruitloop Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58471 | 61.773 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011291 | Mycobacterium phage Brujita | Mycobacterium phage Brujita Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47057 | 66.772 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| NC_011273 | Mycobacterium phage Myrna | Mycobacterium phage Myrna Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164602 | 65.436 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| NC_011216 | Burkholderia phage KS10 | Burkholderia phage KS10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37635 | 62.87 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| NC_009811 | Listeria phage A511 | Listeria phage A511 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137619 | 35.929 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_011167 | Bacillus phage IEBH | Bacillus phage IEBH Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53104 | 36.423 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| NC_011166 | Pseudomonas phage LMA2 | Pseudomonas phage LMA2 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66530 | 55.617 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_011165 | Pseudomonas phage LBL3 | Pseudomonas phage LBL3 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64427 | 55.469 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_011142 | Iodobacter phage PhiPLPE | Iodobacter phage PhiPLPE Iodobacter virus PLPE Iodovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47453 | 46.105 | Iodovirus | Unclassified | Myoviridae | Iodobacter |
| NC_011104 | Lactobacillus phage Lrm1 | Lactobacillus phage Lrm1 Lactobacillus virus Lrm1 Larmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39989 | 45.49 | Larmunavirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_011107 | Pseudomonas phage PT2 | Pseudomonas phage PT2 Pseudomonas virus PT2 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42961 | 62.182 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_011105 | Pseudomonas phage PT5 | Pseudomonas phage PT5 Pseudomonas virus PT5 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42954 | 62.332 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_011103 | Rhizobium phage 16-3 | Rhizobium phage 16-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60195 | 58.952 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| NC_011048 | Bacillus phage phi29 | Bacillus phage phi29 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19282 | 39.991 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_011041 | Escherichia phage V5 | Escherichia phage V5 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137947 | 43.561 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| NC_011039 | Mycobacterium phage Predator | Mycobacterium phage Predator Predatorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70110 | 56.286 | Predatorvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011038 | Yersinia phage Yepe2 | Yersinia phage Yepe2 Yersinia virus Yepe2 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38677 | 47.266 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_011043 | Klebsiella phage K11 | Klebsiella phage K11 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41181 | 53.248 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_011045 | Escherichia phage 13a | Escherichia phage 13a Escherichia virus 13a Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38841 | 48.39 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_011042 | Escherichia phage EcoDS1 | Escherichia phage EcoDS1 Escherichia virus EcoDS1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39252 | 49.944 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_011040 | Escherichia phage BA14 | Escherichia phage BA14 Escherichia virus BA14 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39816 | 48.777 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_011046 | Lactococcus phage bIBB29 | Lactococcus phage bIBB29 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29305 | 34.724 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_009737 | Bacillus virus 1 | Bacillus virus 1 Svunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35055 | 44.85 | Svunavirus | Unclassified | Myoviridae | Bacillus |
| NC_010945 | Streptococcus phage PH15 | Streptococcus phage PH15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39136 | 40.507 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_010807 | Salmonella phage phiSG-JL2 | Salmonella phage phiSG-JL2 Salmonella virus SG-JL2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38815 | 50.864 | Teetrevirus | Studiervirinae | Autographiviridae | Salmonella |
| NC_010808 | Staphylococcus virus phiMR25 | Staphylococcus virus phiMR25 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44342 | 34.326 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_010237 | Escherichia phage Min27 | Escherichia phage Min27 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63395 | 49.504 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_010495 | Salmonella phage Vi II-E1 | Salmonella phage Vi II-E1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45051 | 46.126 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_010179 | Lactobacillus prophage Lj771 | Lactobacillus prophage Lj771 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40881 | 36.401 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| NC_010355 | Azospirillum phage Cd | Azospirillum phage Cd Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62337 | 63.932 | Unclassified | Unclassified | Siphoviridae | Azospirillum |
| NC_010353 | Streptococcus virus 858 | Streptococcus virus 858 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35543 | 39.786 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_010342 | Halomonas phage phiHAP-1 | Halomonas phage phiHAP-1 Hapunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39245 | 58.978 | Hapunavirus | Unclassified | Myoviridae | Halomonas |
| NC_010326 | Pseudomonas phage LUZ19 | Pseudomonas phage LUZ19 Pseudomonas virus LUZ19 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43548 | 62.278 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_010325 | Pseudomonas virus LUZ24 | Pseudomonas virus LUZ24 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45625 | 52.248 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_008296 | Synechococcus phage syn9 | Synechococcus phage syn9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177300 | 40.634 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| NC_010147 | Staphylococcus virus phiMR11 | Staphylococcus virus phiMR11 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43011 | 35.63 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_010105 | Escherichia phage JS98 | Escherichia phage JS98 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170523 | 39.513 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_010106 | Escherichia phage phiEcoM-GJ1 | Escherichia phage phiEcoM-GJ1 Escherichia virus GJ1 Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52975 | 44.023 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| NC_009986 | Sulfolobus spindle-shaped virus 4 | Sulfolobus spindle-shaped virus 4 Alphafusellovirus Fuselloviridae Viruses | 15135 | 38.474 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_009965 | Acidianus rod-shaped virus 1 | Acidianus rod-shaped virus 1 Itarudivirus ARV1 Itarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 24655 | 39.128 | Itarudivirus | Unclassified | Rudiviridae | Acidianus |
| NC_009542 | Aeromonas phage phiO18P | Aeromonas phage phiO18P Bielevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33985 | 61.745 | Bielevirus | Peduovirinae | Myoviridae | Aeromonas |
| NC_009935 | Pseudomonas phage LKD16 | Pseudomonas phage LKD16 Pseudomonas virus LKD16 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43200 | 62.317 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_009818 | Pseudomonas phage MP22 | Pseudomonas phage MP22 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36409 | 64.19 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_009878 | Mycobacterium phage Bethlehem | Mycobacterium phage Bethlehem Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52250 | 63.246 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_009875 | Staphylococcus phage SAP-2 | Staphylococcus phage SAP-2 Staphylococcus virus SAP2 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17938 | 28.944 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_009815 | Listeria phage A006 | Listeria phage A006 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38124 | 35.497 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_009813 | Listeria phage B054 | Listeria phage B054 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48172 | 36.212 | Unclassified | Unclassified | Myoviridae | Listeria |
| NC_009812 | Listeria phage B025 | Listeria phage B025 unclassified Slepowronvirus Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42653 | 35.1 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| NC_009810 | Listeria phage A500 | Listeria phage A500 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38867 | 36.684 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_009821 | Escherichia phage Phi1 | Escherichia phage Phi1 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164270 | 40.499 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_009820 | Mycobacterium phage Tweety | Mycobacterium phage Tweety Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58692 | 61.739 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_009819 | Streptococcus phage P9 | Streptococcus phage P9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40539 | 39.651 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_009817 | Lactococcus phage KSY1 | Lactococcus phage KSY1 Chopinvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79232 | 35.111 | Chopinvirus | Unclassified | Podoviridae | Lactococcus |
| NC_009804 | Thermus phage P74-26 | Thermus phage P74-26 Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83319 | 57.759 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| NC_009803 | Thermus phage P23-45 | Thermus phage P23-45 Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84201 | 57.762 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| NC_007804 | Escherichia phage phiV10 | Escherichia phage phiV10 Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39104 | 48.972 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| NC_009603 | Microbacterium phage Min1 | Microbacterium phage Min1 Microbacterium virus Min1 Minunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46365 | 68.314 | Minunavirus | Unclassified | Siphoviridae | Microbacterium |
| NC_009554 | Lactobacillus phage LL-H | Lactobacillus phage LL-H Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34659 | 47.817 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_009540 | Escherichia phage Tls | Escherichia phage Tls Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49902 | 42.68 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| NC_009543 | Xanthomonas phage Xop411 | Xanthomonas phage Xop411 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44520 | 51.903 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_009541 | Propionibacterium phage PA6 | Propionibacterium phage PA6 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29739 | 54.023 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| NC_009526 | Staphylococcus phage 80alpha | Staphylococcus phage 80alpha Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43864 | 34.103 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_009514 | Enterobacteria phage cdtI | Enterobacteria phage cdtI Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47021 | 49.116 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_009382 | Ralstonia virus RSA1 | Ralstonia virus RSA1 Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38760 | 65.348 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| NC_009237 | Burkholderia phage phiE255 | Burkholderia phage phiE255 Bcepmuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37446 | 63.051 | Bcepmuvirus | Unclassified | Myoviridae | Burkholderia |
| NC_009234 | Burkholderia phage phiE202 | Burkholderia phage phiE202 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35741 | 65.429 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_009231 | Clostridium virus phiC2 | Clostridium virus phiC2 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56538 | 28.726 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| NC_009018 | Streptococcus phage phi3396 | Streptococcus phage phi3396 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38528 | 37.482 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| NC_009016 | Vibrio phage VP882 | Vibrio phage VP882 Hapunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38197 | 56.952 | Hapunavirus | Unclassified | Myoviridae | Vibrio |
| NC_008376 | Geobacillus phage GBSV1 | Geobacillus phage GBSV1 Svunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34683 | 44.379 | Svunavirus | Unclassified | Myoviridae | Geobacillus |
| NC_008799 | Staphylococcus virus phiETA3 | Staphylococcus virus phiETA3 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43282 | 34.899 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_008798 | Staphylococcus virus phiETA2 | Staphylococcus virus phiETA2 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43265 | 34.279 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_008723 | Staphylococcus phage PH15 | Staphylococcus phage PH15 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44041 | 34.908 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_008717 | Pseudomonas phage DMS3 | Pseudomonas phage DMS3 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36415 | 64.265 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_008720 | Escherichia phage N4 | Escherichia phage N4 Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70153 | 41.304 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_008694 | Yersinia phage Berlin | Yersinia phage Berlin Yersinia virus Berlin Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38564 | 47.189 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_008689 | Staphylococcus virus 108PVL | Staphylococcus virus 108PVL unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44857 | 33.469 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_008617 | Staphylococcus phage phiNM3 | Staphylococcus phage phiNM3 Staphylococcus virus phiNM3 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44061 | 32.993 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_007145 | Burkholderia virus phi52237 | Burkholderia virus phi52237 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37639 | 64.818 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| NC_008562 | Microcystis virus Ma-LMM01 | Microcystis virus Ma-LMM01 Fukuivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162109 | 45.953 | Fukuivirus | Unclassified | Myoviridae | Microcystis |
| NC_008515 | Escherichia virus RB32 | Escherichia virus RB32 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165890 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_008464 | Stx2-converting phage 86 | Stx2-converting phage 86 Escherichia virus 86 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60238 | 49.067 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| NC_008371 | Lactococcus virus jj50 | Lactococcus virus jj50 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27453 | 34.943 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_008370 | Lactococcus phage 712 | Lactococcus phage 712 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30510 | 33.884 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_008363 | Lactococcus phage P008 | Lactococcus phage P008 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28538 | 34.68 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_008265 | Clostridium phage phiSM101 | Clostridium phage phiSM101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38092 | 28.129 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_008205 | Mycobacterium phage PMC | Mycobacterium phage PMC Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56692 | 61.418 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_008199 | Mycobacterium phage Pipefish | Mycobacterium phage Pipefish Pipefishvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69059 | 67.335 | Pipefishvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_008203 | Mycobacterium phage Che12 | Mycobacterium phage Che12 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52047 | 62.882 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_008196 | Mycobacterium phage Llij | Mycobacterium phage Llij Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56852 | 61.53 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_008193 | Pasteurella phage F108 | Pasteurella phage F108 Irtavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30505 | 42.105 | Irtavirus | Peduovirinae | Myoviridae | Pasteurella |
| NC_008194 | Mycobacterium phage 244 | Mycobacterium phage 244 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74483 | 62.937 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_008201 | Mannheimia virus PHL101 | Mannheimia virus PHL101 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34525 | 41.567 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| NC_008152 | Escherichia virus K1-5 | Escherichia virus K1-5 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44385 | 45.247 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_007967 | Streptomyces phage mu1/6 | Streptomyces phage mu1/6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38194 | 71.194 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| NC_007917 | Clostridium phage phi CD119 | Clostridium phage phi CD119 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53325 | 28.773 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| NC_007924 | Lactobacillus phage KC5a | Lactobacillus phage KC5a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38239 | 36.897 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| NC_007902 | Sodalis phage phiSG1 | Sodalis phage phiSG1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52162 | 50.826 | Unclassified | Unclassified | Podoviridae | Sodalis |
| NC_007805 | Pseudomonas phage F10 | Pseudomonas phage F10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39199 | 62.083 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_007814 | Bacillus phage Fah | Bacillus phage Fah unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37974 | 34.945 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| NC_007734 | Bacillus phage WBeta | Bacillus phage WBeta Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40867 | 35.258 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| NC_007710 | Xanthomonas phage OP2 | Xanthomonas phage OP2 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46643 | 60.882 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| NC_007709 | Xanthomonas virus OP1 | Xanthomonas virus OP1 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43785 | 51.075 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_007637 | Escherichia virus K1E | Escherichia virus K1E Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45251 | 45.053 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| NC_007623 | Pseudomonas phage EL | Pseudomonas phage EL Elvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 211215 | 49.327 | Elvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_007610 | Listeria phage P100 | Listeria phage P100 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131384 | 36.039 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_007603 | Escherichia virus Rtp | Escherichia virus Rtp Rtpvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46219 | 44.283 | Rtpvirus | Braunvirinae | Drexlerviridae | Escherichia |
| NC_007581 | Clostridium phage c-st | Clostridium phage c-st Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 185683 | 26.284 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_007501 | Lactobacillus phage Lc-Nu | Lactobacillus phage Lc-Nu Lactobacillus virus LcNu Lacnuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36466 | 44.28 | Lacnuvirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_007497 | Burkholderia phage Bcep176 | Burkholderia phage Bcep176 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44856 | 61.539 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| NC_007456 | Escherichia phage K1F | Escherichia phage K1F Escherichia virus K1F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39704 | 49.776 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_007458 | Bacillus phage Gamma | Bacillus phage Gamma unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37253 | 35.221 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| NC_007409 | Acidianus two-tailed virus | Acidianus two-tailed virus Bicaudavirus Bicaudaviridae Viruses | 62730 | 41.226 | Bicaudavirus | Unclassified | Bicaudaviridae | Acidianus |
| NC_007291 | Escherichia phage Jk06 | Escherichia phage Jk06 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46072 | 44.001 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| NC_007064 | Staphylococcus phage 92 | Staphylococcus phage 92 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42431 | 35.722 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007063 | Staphylococcus phage 88 | Staphylococcus phage 88 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43231 | 35.47 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007062 | Staphylococcus phage 52A | Staphylococcus phage 52A Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41690 | 35.534 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007061 | Staphylococcus virus 29 | Staphylococcus virus 29 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42802 | 35.375 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007060 | Staphylococcus virus 55 | Staphylococcus virus 55 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41902 | 35.724 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007059 | Staphylococcus virus 71 | Staphylococcus virus 71 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43114 | 35.239 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007058 | Staphylococcus phage ROSA | Staphylococcus phage ROSA Staphylococcus virus ROSA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43155 | 35.06 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007057 | Staphylococcus virus 96 | Staphylococcus virus 96 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43576 | 35.008 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007056 | Staphylococcus virus EW | Staphylococcus virus EW Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45286 | 35.991 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007055 | Staphylococcus virus 37 | Staphylococcus virus 37 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43681 | 35.114 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007054 | Staphylococcus virus 47 | Staphylococcus virus 47 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44777 | 33.459 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_007053 | Staphylococcus virus 3a | Staphylococcus virus 3a Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43095 | 33.463 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_007052 | Staphylococcus phage 42E | Staphylococcus phage 42E Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45861 | 33.73 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_007051 | Staphylococcus phage 2638A | Staphylococcus phage 2638A Staphylococcus virus 2638A Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41318 | 36.916 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_007049 | Staphylococcus phage 53 | Staphylococcus phage 53 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43883 | 34.077 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007048 | Staphylococcus phage 69 | Staphylococcus phage 69 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42732 | 34.302 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007046 | Staphylococcus phage 66 | Staphylococcus phage 66 Staphylococcus virus 66 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18199 | 29.287 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_007045 | Staphylococcus phage PT1028 | Staphylococcus phage PT1028 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15603 | 31.417 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_007066 | Staphylococcus phage G1 | Staphylococcus phage G1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138715 | 30.388 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_007050 | Staphylococcus phage 85 | Staphylococcus phage 85 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44283 | 34.587 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007065 | Staphylococcus phage X2 | Staphylococcus phage X2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43440 | 35.958 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_007019 | Streptococcus phage 2972 | Streptococcus phage 2972 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34704 | 40.151 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_007021 | Staphylococcus phage Twort | Staphylococcus phage Twort Twortvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130706 | 30.261 | Twortvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_006949 | Enterobacteria phage ES18 | Enterobacteria phage ES18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46900 | 48.593 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| NC_006945 | Bacillus phage GIL16c | Bacillus phage GIL16c Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14844 | 40.07 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| NC_006936 | Lactobacillus phage phiJL-1 | Lactobacillus phage phiJL-1 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36674 | 39.355 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_006820 | Synechococcus phage S-PM2 | Synechococcus phage S-PM2 Synechococcus virus SPM2 Nodensvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 196280 | 37.82 | Nodensvirus | Unclassified | Myoviridae | Synechococcus |
| NC_006557 | Bacillus phage BCASJ1c | Bacillus phage BCASJ1c Bacillus virus BCAJ1 Jarrellvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41092 | 41.74 | Jarrellvirus | Unclassified | Siphoviridae | Bacillus |
| NC_006552 | Pseudomonas phage F116 | Pseudomonas phage F116 Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65195 | 63.172 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_006548 | Pseudomonas phage B3 | Pseudomonas phage B3 Pseudomonas virus B3 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38439 | 63.16 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_005345 | Streptomyces phage VWB | Streptomyces phage VWB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49220 | 71.138 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| NC_006268 | Sulfolobus virus STSV1 | Sulfolobus virus STSV1 unclassified Bicaudaviridae Bicaudaviridae Viruses | 75294 | 35.215 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_005893 | Lactobacillus phage phiAT3 | Lactobacillus phage phiAT3 Lactobacillus virus phiAT3 Fattrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39166 | 44.592 | Fattrevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_005887 | Burkholderia phage BcepC6B | Burkholderia phage BcepC6B Ryyoungvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42415 | 65.194 | Ryyoungvirus | Unclassified | Podoviridae | Burkholderia |
| NC_005263 | Burkholderia phage Bcep1 | Burkholderia phage Bcep1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48177 | 63.64 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| NC_002661 | Staphylococcus virus phiSLT | Staphylococcus virus phiSLT Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42942 | 33.312 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_005857 | Klebsiella phage phiKO2 | Klebsiella phage phiKO2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51601 | 51.485 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| NC_005856 | Escherichia phage P1 | Escherichia phage P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94800 | 47.311 | Punavirus | Unclassified | Myoviridae | Escherichia |
| NC_005841 | Enterobacteria phage ST104 | Enterobacteria phage ST104 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41391 | 47.431 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| NC_005833 | Escherichia phage T1 | Escherichia phage T1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48836 | 45.552 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_005822 | Lactococcus phage phiLC3 | Lactococcus phage phiLC3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32172 | 35.509 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_005361 | Sulfolobus virus Kamchatka 1 | Sulfolobus virus Kamchatka 1 Alphafusellovirus Fuselloviridae Viruses | 17385 | 38.746 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_005360 | Sulfolobus virus Ragged Hills | Sulfolobus virus Ragged Hills Alphafusellovirus Fuselloviridae Viruses | 16473 | 37.534 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_005356 | Staphylococcus virus 77 | Staphylococcus virus 77 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41708 | 33.533 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_005357 | Bordetella phage BPP-1 | Bordetella phage BPP-1 Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42493 | 65.408 | Rauchvirus | Unclassified | Podoviridae | Bordetella |
| NC_005354 | Lactobacillus prophage Lj928 | Lactobacillus prophage Lj928 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38384 | 34.697 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_005355 | Lactobacillus prophage Lj965 | Lactobacillus prophage Lj965 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40190 | 35.141 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_005340 | Salmonella phage PSP3 | Salmonella phage PSP3 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30636 | 52.827 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| NC_005344 | Shigella phage Sf6 | Shigella phage Sf6 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39043 | 47.468 | Lederbergvirus | Unclassified | Podoviridae | Shigella |
| NC_004664 | Streptomyces virus phiBT1 | Streptomyces virus phiBT1 Lomovskayavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41831 | 62.762 | Lomovskayavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_004466 | Pseudomonas phage PaP3 | Pseudomonas phage PaP3 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45503 | 52.227 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| NC_005294 | Streptococcus phage EJ-1 | Streptococcus phage EJ-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42935 | 39.641 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| NC_005282 | Salmonella phage FelixO1 | Salmonella phage FelixO1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86155 | 39.006 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| NC_005258 | Bacillus phage Bam35c | Bacillus phage Bam35c Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14935 | 39.719 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| NC_005265 | Sulfolobus spindle-shaped virus 2 | Sulfolobus spindle-shaped virus 2 Alphafusellovirus Fuselloviridae Viruses | 14796 | 38.497 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_005178 | Pseudomonas phage D3112 | Pseudomonas phage D3112 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37611 | 64.34 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_005069 | Yersinia phage PY54 | Yersinia phage PY54 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46339 | 44.574 | Unclassified | Unclassified | Siphoviridae | Yersinia |
| NC_005066 | Escherichia phage RB49 | Escherichia phage RB49 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164018 | 40.442 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_005056 | Escherichia phage Wphi | Escherichia phage Wphi Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32684 | 51.723 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_005045 | Pseudomonas phage phiKMV | Pseudomonas phage phiKMV Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42519 | 62.297 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_004996 | Streptococcus phage SM1 | Streptococcus phage SM1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34692 | 39.15 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004928 | Escherichia phage RB69 | Escherichia phage RB69 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167560 | 37.656 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_004902 | Xanthomonas phage Xp10 | Xanthomonas phage Xp10 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44373 | 52.038 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_004831 | Salmonella phage SP6 | Salmonella phage SP6 Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43769 | 47.227 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| NC_004827 | Haemophilus phage Aaphi23 | Haemophilus phage Aaphi23 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43033 | 42.456 | Unclassified | Unclassified | Myoviridae | Haemophilus |
| NC_004813 | Enterobacteria phage BP-4795 | Enterobacteria phage BP-4795 Enterobacteria virus BP4795 Marienburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57930 | 50.608 | Marienburgvirus | Unclassified | Siphoviridae | Enterobacteria |
| NC_004777 | Yersinia phage phiA1122 | Yersinia phage phiA1122 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37555 | 48.316 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_004746 | Lactococcus phage P335 sensu lato | Lactococcus phage P335 sensu lato Siphoviridae Caudovirales Viruses | 36596 | 35.367 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_004745 | Yersinia virus L413C | Yersinia virus L413C Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30728 | 52.106 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| NC_003291 | Listeria phage PSA | Listeria phage PSA Psavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37618 | 34.725 | Psavirus | Unclassified | Siphoviridae | Listeria |
| NC_004688 | Mycobacterium phage Omega | Mycobacterium phage Omega Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110865 | 61.375 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004683 | Mycobacterium phage Che9c | Mycobacterium phage Che9c Chenonavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57050 | 65.378 | Chenonavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004681 | Mycobacterium phage Cjw1 | Mycobacterium phage Cjw1 Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75931 | 63.115 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004680 | Mycobacterium phage Che8 | Mycobacterium phage Che8 Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59471 | 61.304 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004679 | Staphylococcus phage P68 | Staphylococcus phage P68 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18227 | 29.325 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_004678 | Staphylococcus virus 44AHJD | Staphylococcus virus 44AHJD Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16784 | 29.623 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| NC_004665 | Pseudomonas phage gh-1 | Pseudomonas phage gh-1 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37359 | 57.416 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_000866 | Escherichia virus T4 | Escherichia virus T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168903 | 35.298 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_002072 | Streptococcus virus DT1 | Streptococcus virus DT1 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34815 | 39.072 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_004629 | Pseudomonas phage phiKZ | Pseudomonas phage phiKZ Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 280334 | 36.833 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_004617 | Staphylococcus virus 13 | Staphylococcus virus 13 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42722 | 33.472 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_004616 | Staphylococcus phage phi 12 | Staphylococcus phage phi 12 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44970 | 33.347 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_004615 | Staphylococcus virus 11 | Staphylococcus virus 11 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43604 | 34.497 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_004456 | Vibrio phage VHML | Vibrio phage VHML Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43198 | 50.618 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| NC_003050 | Streptococcus phage MM1 | Streptococcus phage MM1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40248 | 38.412 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004348 | Salmonella phage ST64T | Salmonella phage ST64T Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40679 | 47.523 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_004313 | Salmonella phage ST64B | Salmonella phage ST64B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40149 | 51.015 | Unclassified | Unclassified | Myoviridae | Salmonella |
| NC_004303 | Streptococcus virus O1205 | Streptococcus virus O1205 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43075 | 38.261 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_004305 | Lactobacillus phage phig1e | Lactobacillus phage phig1e Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42259 | 43.061 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_004165 | Bacillus virus B103 | Bacillus virus B103 Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18630 | 37.66 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_004167 | Bacillus phage phi105 | Bacillus phage phi105 Bacillus virus phi105 Spizizenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39325 | 42.69 | Spizizenvirus | Unclassified | Siphoviridae | Bacillus |
| NC_004112 | Lactobacillus phage A2 | Lactobacillus phage A2 Lactobacillus virus A2 Ashduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43411 | 44.89 | Ashduovirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_004087 | Sulfolobus islandicus rod-shaped virus 1 | Sulfolobus islandicus rod-shaped virus 1 Icerudivirus SIRV1 Icerudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32308 | 25.269 | Icerudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_004084 | Natrialba phage PhiCh1 | Natrialba phage PhiCh1 Natrialba virus PhiCh1 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58498 | 61.888 | Myohalovirus | Unclassified | Myoviridae | Natrialba |
| NC_004066 | Lactococcus phage ul36 | Lactococcus phage ul36 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36798 | 35.836 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_003524 | Clostridium phage phi3626 | Clostridium phage phi3626 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33507 | 28.382 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_003444 | Enterobacteria phage SfV | Enterobacteria phage SfV Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37074 | 50.769 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| NC_003356 | Enterobacteria phage phiP27 | Enterobacteria phage phiP27 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42575 | 49.351 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| NC_003324 | Sinorhizobium phage PBC5 | Sinorhizobium phage PBC5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57416 | 61.486 | Unclassified | Unclassified | Podoviridae | Sinorhizobium |
| NC_003315 | Haemophilus virus HP2 | Haemophilus virus HP2 Hpunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31508 | 39.936 | Hpunavirus | Peduovirinae | Myoviridae | Haemophilus |
| NC_003309 | Burkholderia phage phiE125 | Burkholderia phage phiE125 Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53373 | 61.192 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| NC_003313 | Bacteriophage K139 | Bacteriophage K139 Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33106 | 48.897 | Longwoodvirus | Peduovirinae | Myoviridae | Unspecified |
| NC_003298 | Enterobacteria phage T3 | Enterobacteria phage T3 Escherichia virus T3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38208 | 49.901 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_003288 | Staphylococcus virus phiETA | Staphylococcus virus phiETA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43081 | 35.431 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| NC_003278 | Pseudomonas virus phiCTX | Pseudomonas virus phiCTX Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35580 | 62.622 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| NC_003216 | Listeria phage A118 | Listeria phage A118 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40834 | 36.107 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_003158 | Natrinema virus SNJ1 | Natrinema virus SNJ1 Betasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 16341 | 61.104 | Betasphaerolipovirus | Unclassified | Sphaerolipoviridae | Natrinema |
| NC_003085 | Myxococcus phage Mx8 | Myxococcus phage Mx8 Myxococcus virus Mx8 Myxoctovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49534 | 67.745 | Myxoctovirus | Unclassified | Podoviridae | Myxococcus |
| NC_002796 | Lactococcus phage BK5-T | Lactococcus phage BK5-T Lactococcus virus BK5T Sandinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40003 | 34.975 | Sandinevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_002747 | Lactococcus phage TP901-1 | Lactococcus phage TP901-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37667 | 35.378 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002730 | Enterobacteria phage HK620 | Enterobacteria phage HK620 Escherichia virus HK620 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38297 | 46.688 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| NC_002703 | Lactococcus phage Tuc2009 | Lactococcus phage Tuc2009 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38347 | 36.222 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002671 | Lactococcus phage bIL312 | Lactococcus phage bIL312 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15179 | 33.039 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002669 | Lactococcus phage bIL310 | Lactococcus phage bIL310 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14957 | 35.876 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002668 | Lactococcus phage bIL309 | Lactococcus phage bIL309 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36949 | 35.679 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002667 | Lactococcus phage bIL286 | Lactococcus phage bIL286 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41834 | 35.323 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002666 | Lactococcus phage bIL285 | Lactococcus phage bIL285 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35538 | 35.188 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002670 | Lactococcus phage bIL311 | Lactococcus phage bIL311 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14510 | 34.238 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| NC_002656 | Mycobacterium phage Bxb1 | Mycobacterium phage Bxb1 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50550 | 63.65 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_002486 | Staphylococcus prophage phiPV83 | Staphylococcus prophage phiPV83 Staphylococcus virus PV83 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45636 | 33.476 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| NC_002214 | Streptococcus virus Sfi11 | Streptococcus virus Sfi11 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39807 | 38.556 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_002185 | Streptococcus phage 7201 | Streptococcus phage 7201 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35466 | 38.727 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_002167 | Escherichia phage HK97 | Escherichia phage HK97 Byrnievirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39732 | 49.786 | Byrnievirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_002166 | Escherichia phage HK022 | Escherichia phage HK022 Shamshuipovirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40751 | 49.483 | Shamshuipovirus | Hendrixvirinae | Siphoviridae | Escherichia |
| NC_001317 | Escherichia phage 186 | Escherichia phage 186 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30624 | 53.086 | Eganvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_001271 | Yersinia phage phiYeO3-12 | Yersinia phage phiYeO3-12 Yersinia virus YeO3-12 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39600 | 50.631 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| NC_000935 | Hamiltonella phage APSE-1 | Hamiltonella phage APSE-1 Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36524 | 43.886 | Sendosyvirus | Unclassified | Podoviridae | Hamiltonella |
| NC_000929 | Escherichia phage Mu | Escherichia phage Mu Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36717 | 52.055 | Muvirus | Unclassified | Myoviridae | Escherichia |
| NC_000902 | Enterobacteria phage VT2-Sakai | Enterobacteria phage VT2-Sakai unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60942 | 49.913 | Traversvirus | Sepvirinae | Podoviridae | Enterobacteria |
| NC_000896 | Lactobacillus phage phiadh | Lactobacillus phage phiadh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43785 | 35.572 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| NC_000872 | Streptococcus phage Sfi21 | Streptococcus phage Sfi21 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40739 | 37.566 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_000871 | Streptococcus phage Sfi19 | Streptococcus phage Sfi19 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37370 | 38.274 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_000867 | Pseudoalteromonas phage PM2 | Pseudoalteromonas phage PM2 Corticovirus Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10079 | 42.167 | Corticovirus | Unclassified | Corticoviridae | Pseudoalteromonas |
| NC_000924 | Escherichia phage 933W | Escherichia phage 933W Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61670 | 49.368 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| NC_001942 | Mycoplasma phage MAV1 | Mycoplasma phage MAV1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 15644 | 29.033 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| NC_001902 | Methanobacterium phage psiM2 | Methanobacterium phage psiM2 Psimunavirus psiM2 Psimunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26111 | 46.306 | Psimunavirus | Unclassified | Siphoviridae | Methanobacterium |
| NC_001901 | Escherichia phage N15 | Escherichia phage N15 Ravinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46375 | 51.172 | Ravinvirus | Unclassified | Siphoviridae | Escherichia |
| NC_001895 | Bacteriophage P2 | Bacteriophage P2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33593 | 50.168 | Peduovirus | Peduovirinae | Myoviridae | Unspecified |
| NC_001884 | Bacillus virus SPbeta | Bacillus virus SPbeta Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134416 | 34.635 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| NC_001835 | Lactococcus phage SK1 | Lactococcus phage SK1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28451 | 34.512 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_001706 | Lactococcus virus c2 | Lactococcus virus c2 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22172 | 36.307 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_001697 | Haemophilus virus HP1 | Haemophilus virus HP1 Hpunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32355 | 40.006 | Hpunavirus | Peduovirinae | Myoviridae | Haemophilus |
| NC_001629 | Lactococcus phage bIL67 | Lactococcus phage bIL67 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22195 | 35.99 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| NC_001609 | Enterobacteria phage P4 | Enterobacteria phage P4 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11624 | 49.527 | Unclassified | Unclassified | Unclassified | Enterobacteria |
| NC_001604 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39937 | 48.399 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_001423 | Bacillus phage PZA | Bacillus phage PZA Bacillus virus PZA Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19366 | 39.662 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| NC_001416 | Escherichia virus Lambda | Escherichia virus Lambda Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48502 | 49.858 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| NC_001338 | Sulfolobus spindle-shaped virus 1 | Sulfolobus spindle-shaped virus 1 Alphafusellovirus Fuselloviridae Viruses | 15465 | 39.696 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_001335 | Mycobacterium virus L5 | Mycobacterium virus L5 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52297 | 62.258 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055877 | crAssphage cr8_1 | crAssphage cr8_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92668 | 30.185 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055730 | Mycobacterium phage Geralt | Mycobacterium phage Geralt unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55765 | 61.528 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055716 | Yersinia phage fHe-Yen9-01 | Yersinia phage fHe-Yen9-01 unclassified Tegunavirus Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167773 | 34.791 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| NC_054972 | Achromobacter phage vB_AxyP_19-32_Axy11 | Achromobacter phage vB_AxyP_19-32_Axy11 Achromobacter virus Axy11 Pourcelvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73413 | 54.324 | Pourcelvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_054971 | Achromobacter phage vB_AxyP_19-32_Axy10 | Achromobacter phage vB_AxyP_19-32_Axy10 Achromobacter virus Axy10 Pourcelvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73898 | 54.269 | Pourcelvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_054970 | Pseudomonas phage inbricus | Pseudomonas phage inbricus Pseudomonas virus inbricus Inbricusvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70211 | 55.856 | Inbricusvirus | Rothmandenesvirinae | Schitoviridae | Pseudomonas |
| NC_054969 | Achromobacter phage vB_AxyP_19-32_Axy24 | Achromobacter phage vB_AxyP_19-32_Axy24 Achromobacter virus Axy24 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74744 | 54.918 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_054968 | Achromobacter phage vB_AxyP_19-32_Axy12 | Achromobacter phage vB_AxyP_19-32_Axy12 Achromobacter virus Axy12 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74096 | 55.038 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_054967 | Achromobacter phage vB_AxyP_19-32_Axy04 | Achromobacter phage vB_AxyP_19-32_Axy04 Achromobacter virus Axy04 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73834 | 54.934 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_054962 | Ralstonia phage Claudette | Ralstonia phage Claudette Ralstonia virus Claudettte Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57085 | 64.367 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| NC_054960 | Ralstonia phage Heva | Ralstonia phage Heva Ralstonia virus Heva Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58352 | 64.62 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| NC_054959 | Ralstonia phage Gerry | Ralstonia phage Gerry Ralstonia virus Gerry Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60898 | 64.73 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| NC_054958 | Ralstonia phage Gamede | Ralstonia phage Gamede Ralstonia virus Gamede Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58603 | 64.602 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| NC_054957 | Ralstonia phage Eline | Ralstonia phage Eline Ralstonia virus Eline Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57352 | 64.622 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| NC_054946 | Ralstonia phage Simangalove | Ralstonia phage Simangalove Ralstonia virus Simangalove Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44834 | 62.252 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| NC_054790 | Mycobacterium phage Anthony | Mycobacterium phage Anthony unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52202 | 61.367 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054674 | Streptomyces phage Joe | Streptomyces phage Joe unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48941 | 65.477 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_054641 | Salmonella virus KFS-SE2 | Salmonella virus KFS-SE2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48608 | 45.801 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| NC_054462 | Ralstonia phage Adzire | Ralstonia phage Adzire unclassified Bakolyvirus Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44732 | 62.228 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| NC_053241 | Gordonia phage Ribeye | Gordonia phage Ribeye unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57448 | 68.14 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_052459 | Mycobacterium phage Rohr | Mycobacterium phage Rohr unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53483 | 63.559 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051648 | Mycobacterium phage Donkeykong | Mycobacterium phage Donkeykong unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59478 | 61.863 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051640 | Mycobacterium phage ByChance | Mycobacterium phage ByChance unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53123 | 61.314 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051618 | Mycobacterium phage Curiosium | Mycobacterium phage Curiosium unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61222 | 65.645 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051597 | Mycobacterium phage Thyatira | Mycobacterium phage Thyatira unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63874 | 64.574 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051588 | Mycobacterium phage Aminay | Mycobacterium phage Aminay unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60430 | 67.834 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_049856 | Lactococcus phage AM4 | Lactococcus phage AM4 Lactococcus virus AM4 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132949 | 32.508 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049445 | Acinetobacter phage AbTZA1 | Acinetobacter phage AbTZA1 Acinetobacter virus AbTZA1 Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168223 | 36.345 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_048800 | Serratia phage MyoSmar | Serratia phage MyoSmar Serratia virus MyoSmar Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68745 | 49.151 | Myosmarvirus | Unclassified | Myoviridae | Serratia |
| NC_048729 | Mycobacterium phage Renaud18 | Mycobacterium phage Renaud18 Mycobacterium virus Renaud18 Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58078 | 61.665 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048675 | Pseudomonas phage BrSP1 | Pseudomonas phage BrSP1 Pseudomonas virus BrSP1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66189 | 55.68 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_048637 | Nodularia phage vB_NpeS-2AV2 | Nodularia phage vB_NpeS-2AV2 Nodularia virus 2AV2 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139100 | 40.278 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| NC_048122 | Vibrio phage 1.188.A._10N.286.51.A6 | Vibrio phage 1.188.A._10N.286.51.A6 Vibrio virus 51A6 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72305 | 42.983 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| NC_048121 | Vibrio phage 1.169.O._10N.261.52.B1 | Vibrio phage 1.169.O._10N.261.52.B1 Vibrio virus 52B1 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72290 | 42.941 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| NC_048117 | Pectobacterium phage Khlen | Pectobacterium phage Khlen Pectobacterium virus Khlen Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44013 | 48.915 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048002 | Escherichia phage SRT7 | Escherichia phage SRT7 Escherichia virus SRT7 Foetvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39883 | 50.535 | Foetvirus | Studiervirinae | Autographiviridae | Escherichia |
| NC_047951 | Yersinia phage phiR8-01 | Yersinia phage phiR8-01 Yersinia virus R8-01 Pienvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42085 | 51.845 | Pienvirus | Melnykvirinae | Autographiviridae | Yersinia |
| NC_047946 | Ralstonia phage RsoP1EGY | Ralstonia phage RsoP1EGY Ralstonia virus RsoP1EGY Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41297 | 58.97 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_047926 | Lactobacillus phage Semele | Lactobacillus phage Semele Lactobacillus virus Semele Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139450 | 36.331 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_047918 | Lactobacillus phage Satyr | Lactobacillus phage Satyr Lactobacillus virus Satyr Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81076 | 39.417 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047890 | Escherichia phage PGT2 | Escherichia phage PGT2 Escherichia virus PGT2 Ermolevavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42686 | 51.649 | Ermolevavirus | Unclassified | Autographiviridae | Escherichia |
| NC_047872 | Listeria phage LMTA-94 | Listeria phage LMTA-94 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136468 | 35.841 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_047863 | Shigella phage vB_SflS-ISF001 | Shigella phage vB_SflS-ISF001 Shigella virus ISF001 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.585 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_047856 | Enterobacter phage KNP3 | Enterobacter phage KNP3 Enterobacter virus KPN3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44911 | 39.193 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| NC_047852 | Pseudomonas phage phiNFS | Pseudomonas phage phiNFS Pseudomonas virus NFS Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42351 | 62.263 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_047805 | Yersinia phage fHe-Yen3-01 | Yersinia phage fHe-Yen3-01 Yersinia virus fHeYen301 Pokrovskaiavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42771 | 47.672 | Pokrovskaiavirus | Melnykvirinae | Autographiviridae | Yersinia |
| NC_047799 | Klebsiella phage K5-4 | Klebsiella phage K5-4 Klebsiella virus K5-4 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40163 | 53.136 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047798 | Klebsiella phage K5-2 | Klebsiella phage K5-2 Klebsiella virus K5-2 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41116 | 53.342 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_047764 | Lactobacillus phage P1 | Lactobacillus phage P1 Lactobacillus virus P1 Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73787 | 38.962 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047759 | Dickeya phage BF25/12 | Dickeya phage BF25/12 Dickeya virus BF25-12 Stompvirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43872 | 52.186 | Stompvirus | Corkvirinae | Autographiviridae | Dickeya |
| NC_043469 | Enterobacter phage F20 | Enterobacter phage F20 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51543 | 47.902 | Webervirus | Unclassified | Drexlerviridae | Enterobacter |
| NC_042126 | Enterococcus phage vB_EfaS_AL3 | Enterococcus phage vB_EfaS_AL3 Enterococcus virus AL3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40789 | 34.838 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042125 | Enterococcus phage LY0322 | Enterococcus phage LY0322 Enterococcus virus LY0322 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40934 | 34.841 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042122 | Enterobacter phage Ec_L1 | Enterobacter phage Ec_L1 Enterobacter virus EcL1 Eclunavirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51894 | 48.241 | Eclunavirus | Unclassified | Drexlerviridae | Enterobacter |
| NC_042113 | Pseudomonas phage vB_PaeM_C2-10_Ab02 | Pseudomonas phage vB_PaeM_C2-10_Ab02 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93848 | 49.397 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042043 | Escherichia phage SRT8 | Escherichia phage SRT8 Escherichia virus SRT8 Sertoctavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49579 | 48.034 | Sertoctavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_042037 | Aeromonas phage pIS4-A | Aeromonas phage pIS4-A Roufvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47624 | 47.279 | Roufvirus | Unclassified | Siphoviridae | Aeromonas |
| NC_042029 | Edwardsiella phage eiAU | Edwardsiella phage eiAU Eiauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43015 | 55.432 | Eiauvirus | Unclassified | Siphoviridae | Edwardsiella |
| NC_041968 | Pseudomonas phage vB_PaeM_G1 | Pseudomonas phage vB_PaeM_G1 Pseudomonas virus G1 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87646 | 54.682 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041925 | Proteus phage VB_PmiS-Isfahan | Proteus phage VB_PmiS-Isfahan Proteus virus Isfahan Gorganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54836 | 36.093 | Gorganvirus | Unclassified | Siphoviridae | Proteus |
| NC_041904 | Pseudomonas phage Zigelbrucke | Pseudomonas phage Zigelbrucke Pseudomonas virus Zigelbrucke Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92338 | 49.318 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041903 | Pseudomonas phage PA10 | Pseudomonas phage PA10 Pseudomonas virus PA10 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91212 | 49.305 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041874 | Escherichia phage JMPW1 | Escherichia phage JMPW1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49628 | 45.559 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_041856 | Streptomyces phage phiCAM | Streptomyces phage phiCAM Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50348 | 65.586 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| NC_041854 | Pectobacterium phage PhiM1 | Pectobacterium phage PhiM1 Pectobacterium virus fM1 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43827 | 49.182 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_041845 | Flavobacterium phage FCV-1 | Flavobacterium phage FCV-1 Flavobacterium virus FCV1 Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46450 | 29.823 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| NC_019911 | Yersinia phage phi80-18 | Yersinia phage phi80-18 Yersinia virus Phi80-18 Pokrovskaiavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42406 | 47.637 | Pokrovskaiavirus | Melnykvirinae | Autographiviridae | Yersinia |
| NC_028908 | Achromobacter phage phiAxp-3 | Achromobacter phage phiAxp-3 Jwalphavirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72825 | 55.202 | Jwalphavirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_034628 | Sulfolobus islandicus rod-shaped virus 4 | Sulfolobus islandicus rod-shaped virus 4 Usarudivirus SIRV4 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35035 | 25.623 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034627 | Sulfolobus islandicus rod-shaped virus 6 | Sulfolobus islandicus rod-shaped virus 6 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35439 | 25.54 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034625 | Sulfolobus islandicus rod-shaped virus 10 | Sulfolobus islandicus rod-shaped virus 10 Usarudivirus SIRV10 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32735 | 25.383 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034624 | Sulfolobus islandicus rod-shaped virus 11 | Sulfolobus islandicus rod-shaped virus 11 Usarudivirus SIRV11 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 33356 | 25.8 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034623 | Sulfolobus islandicus rod-shaped virus 8 | Sulfolobus islandicus rod-shaped virus 8 Usarudivirus SIRV8 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36493 | 25.079 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034620 | Sulfolobus islandicus rod-shaped virus 9 | Sulfolobus islandicus rod-shaped virus 9 Usarudivirus SIRV9 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36391 | 24.932 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034619 | Sulfolobus islandicus rod-shaped virus 7 | Sulfolobus islandicus rod-shaped virus 7 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 34190 | 25.566 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_034621 | Sulfolobus islandicus rod-shaped virus 5 | Sulfolobus islandicus rod-shaped virus 5 Usarudivirus SIRV5 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36306 | 25.434 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_031932 | Flavobacterium phage Fpv10 | Flavobacterium phage Fpv10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42702 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031945 | Flavobacterium phage Fpv11 | Flavobacterium phage Fpv11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.01 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_031110 | Pseudomonas phage phiMK | Pseudomonas phage phiMK Pseudomonas virus phiMK Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93129 | 49.288 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_031051 | Enterococcus phage SANTOR1 | Enterococcus phage SANTOR1 Enterococcus virus SANTOR1 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37933 | 34.54 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_030910 | Pseudomonas phage K5 | Pseudomonas phage K5 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93754 | 49.353 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_030903 | Bacillus phage SPG24 | Bacillus phage SPG24 Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152069 | 42.18 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_029065 | Pseudomonas phage VCM | Pseudomonas phage VCM Pseudomonas virus VCM Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98765 | 48.508 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| NC_029070 | Cronobacter phage Dev-CD-23823 | Cronobacter phage Dev-CD-23823 Cronobacter virus DevCD23823 Cronosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41620 | 53.84 | Cronosvirus | Melnykvirinae | Autographiviridae | Cronobacter |
| NC_029002 | Microcystis phage MaMV-DC | Microcystis phage MaMV-DC Microcystis virus MVDC Fukuivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169223 | 46.032 | Fukuivirus | Unclassified | Myoviridae | Microcystis |
| NC_028952 | Streptomyces phage SF3 | Streptomyces phage SF3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60934 | 69.112 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| NC_028882 | Pseudomonas phage vB_PaeM_PS24 | Pseudomonas phage vB_PaeM_PS24 Pseudomonas virus PS24 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84583 | 54.7 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028850 | Yersinia phage vB_YenP_ISAO8 | Yersinia phage vB_YenP_ISAO8 Yersinia virus ISAO8 Aghbyvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41449 | 53.84 | Aghbyvirus | Melnykvirinae | Autographiviridae | Yersinia |
| NC_028829 | Vibrio phage ValKK3 | Vibrio phage ValKK3 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 248088 | 41.235 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| NC_028817 | Pseudomonas phage K8 | Pseudomonas phage K8 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93879 | 49.355 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028807 | Streptomyces phage SF1 | Streptomyces phage SF1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43150 | 69.089 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| NC_028763 | Polaribacter phage P12002S | Polaribacter phage P12002S Incheonvrus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49847 | 28.933 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| NC_028676 | Sinorhizobium phage phiM9 | Sinorhizobium phage phiM9 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149218 | 49.763 | Unclassified | Unclassified | Ackermannviridae | Sinorhizobium |
| NC_028662 | Mycobacterium phage Phlei | Mycobacterium phage Phlei unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50418 | 60.062 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028652 | Pseudomonas phage C11 | Pseudomonas phage C11 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94109 | 49.391 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028268 | Aeropyrum pernix spindle-shaped virus 1 | Aeropyrum pernix spindle-shaped virus 1 unclassified archaeal viruses Viruses | 38049 | 56.511 | Unclassified | Unclassified | Unclassified | Aeropyrum |
| NC_028256 | Aeropyrum pernix ovoid virus 1 | Aeropyrum pernix ovoid virus 1 Betaguttavirus Guttaviridae Viruses | 13769 | 55.501 | Betaguttavirus | Unclassified | Guttaviridae | Aeropyrum |
| NC_027125 | Flavobacterium phage FCL-2 | Flavobacterium phage FCL-2 Flavobacterium virus FCL2 Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47142 | 30.198 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| NC_026595 | Mycobacterium phage Hades | Mycobacterium phage Hades unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54986 | 61.396 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_026587 | Pseudomonas phage vB_PaeM_PAO1_Ab03 | Pseudomonas phage vB_PaeM_PAO1_Ab03 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86246 | 54.677 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_026010 | Shigella phage pSf-2 | Shigella phage pSf-2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50109 | 45.443 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| NC_024356 | Enterococcus phage IME-EFm1 | Enterococcus phage IME-EFm1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42597 | 35.181 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_024355 | Staphylococcus phage 6ec | Staphylococcus phage 6ec Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93794 | 29.296 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| NC_015272 | Pseudomonas phage KPP10 | Pseudomonas phage KPP10 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88322 | 54.794 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_023595 | Enterococcus phage IME_EF3 | Enterococcus phage IME_EF3 Enterococcus virus EF3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41687 | 34.598 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_023589 | Shigella phage pSb-1 | Shigella phage pSb-1 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71629 | 42.742 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Shigella |
| NC_023556 | Achromobacter phage JWAlpha | Achromobacter phage JWAlpha Jwalphavirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72329 | 54.412 | Jwalphavirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| NC_023555 | Edwardsiella phage eiAU-183 | Edwardsiella phage eiAU-183 unclassified Eiauvirus Eiauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43017 | 55.436 | Eiauvirus | Unclassified | Siphoviridae | Edwardsiella |
| NC_023568 | Vibrio phage VH7D | Vibrio phage VH7D unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246964 | 41.306 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| NC_023502 | Rhizobium phage vB_RleS_L338C | Rhizobium phage vB_RleS_L338C Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109558 | 59.003 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| NC_015294 | Pseudomonas phage PAK_P1 | Pseudomonas phage PAK_P1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93198 | 49.498 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022974 | Pseudomonas phage CHA_P1 | Pseudomonas phage CHA_P1 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88255 | 54.628 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022970 | Pseudomonas phage PAK_P3 | Pseudomonas phage PAK_P3 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88097 | 54.827 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022967 | Pseudomonas phage PAK_P2 | Pseudomonas phage PAK_P2 Pseudomonas virus PAKP2 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92495 | 49.288 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022966 | Pseudomonas phage PAK_P5 | Pseudomonas phage PAK_P5 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88135 | 54.73 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022776 | Streptococcus phage TP-778L | Streptococcus phage TP-778L unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41757 | 38.978 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| NC_022096 | Pseudomonas phage PaBG | Pseudomonas phage PaBG Pseudomonas virus PaBG Baikalvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 258139 | 55.815 | Baikalvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_021537 | Halorubrum phage CGphi46 | Halorubrum phage CGphi46 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39784 | 65.107 | Unclassified | Unclassified | Siphoviridae | Halorubrum |
| NC_021534 | Vibrio phage pYD38-A | Vibrio phage pYD38-A unclassified Roufvirus Roufvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47552 | 47.289 | Roufvirus | Unclassified | Siphoviridae | Vibrio |
| NC_021331 | Shigella phage pSf-1 | Shigella phage pSf-1 Shigella virus pSf1 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51821 | 44.023 | Hanrivervirus | Tempevirinae | Drexlerviridae | Shigella |
| NC_020883 | Bacillus phage PM1 | Bacillus phage PM1 Bacillus virus PM1 Pemunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50861 | 41.285 | Pemunavirus | Unclassified | Myoviridae | Bacillus |
| NC_020078 | Cronobacter phage vB_CskP_GAP227 | Cronobacter phage vB_CskP_GAP227 Cronobacter virus GAP227 Cronosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41796 | 55.74 | Cronosvirus | Melnykvirinae | Autographiviridae | Cronobacter |
| NC_020203 | Pseudomonas phage JBD24 | Pseudomonas phage JBD24 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37095 | 64.24 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_020202 | Pseudomonas phage JBD5 | Pseudomonas phage JBD5 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37740 | 64.269 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_020200 | Pseudomonas phage JBD88a | Pseudomonas phage JBD88a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36429 | 63.985 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_020198 | Pseudomonas phage JBD30 | Pseudomonas phage JBD30 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36947 | 64.276 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_019918 | Pseudomonas phage vB_PaeM_C2-10_Ab1 | Pseudomonas phage vB_PaeM_C2-10_Ab1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92777 | 49.277 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_019520 | Escherichia phage phiKT | Escherichia phage phiKT Escherichia virus PhiKT Ermolevavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42608 | 51.589 | Ermolevavirus | Unclassified | Autographiviridae | Escherichia |
| NC_019450 | Pseudomonas phage JG004 | Pseudomonas phage JG004 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93017 | 49.261 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_018860 | Bacillus phage BCP78 | Bacillus phage BCP78 Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156176 | 39.859 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_018855 | Escherichia phage HX01 | Escherichia phage HX01 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169158 | 37.588 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_016654 | Rhodococcus phage REQ3 | Rhodococcus phage REQ3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39474 | 65.95 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_015585 | Pantoea phage LIMEzero | Pantoea phage LIMEzero Waewaevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43032 | 55.352 | Waewaevirus | Unclassified | Autographiviridae | Pantoea |
| NC_015292 | Erwinia phage phiEa104 | Erwinia phage phiEa104 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84565 | 43.83 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| NC_014099 | Sulfolobus turreted icosahedral virus 2 | Sulfolobus turreted icosahedral virus 2 Alphaturrivirus Turriviridae Belfryvirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 16622 | 36.734 | Alphaturrivirus | Unclassified | Turriviridae | Sulfolobus |
| NC_013585 | Acidianus spindle-shaped virus 1 | Acidianus spindle-shaped virus 1 Betafusellovirus Fuselloviridae Viruses | 24186 | 37.695 | Betafusellovirus | Unclassified | Fuselloviridae | Acidianus |
| NC_004462 | Thermus virus IN93 | Thermus virus IN93 Gammasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 19604 | 65.936 | Gammasphaerolipovirus | Unclassified | Sphaerolipoviridae | Thermus |
| NC_011811 | Erwinia phage phiEa21-4 | Erwinia phage phiEa21-4 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84576 | 43.813 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| NC_004682 | Mycobacterium phage Bxz2 | Mycobacterium phage Bxz2 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50913 | 64.184 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021529 | Vibrio phage nt-1 | Vibrio phage nt-1 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 247511 | 41.327 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| NC_025375 | Stygiolobus rod-shaped virus | Stygiolobus rod-shaped virus Azorudivirus SRV Azorudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 28096 | 29.275 | Azorudivirus | Unclassified | Rudiviridae | Stygiolobus |
| NC_011217 | Sulfolobus spindle-shaped virus 5 | Sulfolobus spindle-shaped virus 5 Alphafusellovirus Fuselloviridae Viruses | 15330 | 38.48 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| NC_009990 | Thalassomonas phage BA3 | Thalassomonas phage BA3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37313 | 40.889 | Unclassified | Unclassified | Podoviridae | Thalassomonas |
| NC_008695 | Archaeal BJ1 virus | Archaeal BJ1 virus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42271 | 64.886 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_004166 | Bacillus phage SPP1 | Bacillus phage SPP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44010 | 43.715 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_005083 | Vibrio phage KVP40 | Vibrio phage KVP40 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244834 | 42.599 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| NC_007810 | Pseudomonas phage F8 | Pseudomonas phage F8 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66015 | 54.944 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_006565 | Lactobacillus phage LP65 | Lactobacillus phage LP65 Salchichonvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131522 | 37.321 | Salchichonvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_005892 | Sulfolobus turreted icosahedral virus 1 | Sulfolobus turreted icosahedral virus 1 Alphaturrivirus Turriviridae Belfryvirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 17663 | 36.007 | Alphaturrivirus | Unclassified | Turriviridae | Sulfolobus |
| NC_004086 | Sulfolobus islandicus rod-shaped virus 2 | Sulfolobus islandicus rod-shaped virus 2 Icerudivirus SIRV2 Icerudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35450 | 25.199 | Icerudivirus | Unclassified | Rudiviridae | Sulfolobus |
| NC_002321 | Staphylococcus phage PVL | Staphylococcus phage PVL Staphylococcus virus PVL Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41401 | 33.555 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| BK059162 | Altiarchaeum virus | Altiarchaeum virus unclassified archaeal viruses Viruses | 22586 | 35.633 | Unclassified | Unclassified | Unclassified | Altiarchaeum |
| BK059157 | Altiarchaeum virus | Altiarchaeum virus unclassified archaeal viruses Viruses | 12584 | 31.921 | Unclassified | Unclassified | Unclassified | Altiarchaeum |
| OK094519 | Pseudomonas phage PCS4 | Pseudomonas phage PCS4 unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39191 | 57.988 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| OK085710 | Myxococcus phage Mx4 | Myxococcus phage Mx4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56975 | 70.124 | Unclassified | Unclassified | Myoviridae | Myxococcus |
| MZ333462 | Enterococcus phage GVEsP-1 | Enterococcus phage GVEsP-1 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149913 | 37.002 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MZ333461 | Enterococcus phage VEsP-2 | Enterococcus phage VEsP-2 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43177 | 34.845 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MW464860 | Enterobacter phage EC151 | Enterobacter phage EC151 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60753 | 47.693 | Unclassified | Queuovirinae | Siphoviridae | Enterobacter |
| CP000625 | Burkholderia phage phi644-2 | Burkholderia phage phi644-2 Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48674 | 60.449 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| CP000624 | Burkholderia phage phiE12-2 | Burkholderia phage phiE12-2 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36690 | 64.617 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| MZ065353 | Escherichia virus KFS-EC3 | Escherichia virus KFS-EC3 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167440 | 35.549 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JN662425 | Burkholderia phage DC1 | Burkholderia phage DC1 Lessievirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61847 | 66.212 | Lessievirus | Unclassified | Podoviridae | Burkholderia |
| EF602154 | Burkholderia phage BcepNY3 | Burkholderia phage BcepNY3 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47382 | 63.64 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| AY539836 | Burkholderia phage BcepMu | Burkholderia phage BcepMu Bcepmuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36748 | 62.861 | Bcepmuvirus | Unclassified | Myoviridae | Burkholderia |
| AY453853 | Burkholderia phage phi1026b | Burkholderia phage phi1026b Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54865 | 60.683 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| MW334946 | Cyanophage S-2L | Cyanophage S-2L Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45087 | 69.432 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| KT995480 | Bacillus phage BM15 | Bacillus phage BM15 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165213 | 39.605 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KF669648 | Bacillus phage Blastoid | Bacillus phage Blastoid Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50354 | 42.233 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KC330684 | Bacillus phage Andromeda | Bacillus phage Andromeda Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49259 | 41.907 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KC330680 | Bacillus phage Eoghan | Bacillus phage Eoghan Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49458 | 42.208 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KC330679 | Bacillus phage Curly | Bacillus phage Curly Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49425 | 41.821 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| GU229986 | Bacillus phage 250 | Bacillus phage 250 Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56505 | 36.441 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| EU408779 | Bacillus phage AP50 | Bacillus phage AP50 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14398 | 38.651 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| DQ123818 | Actinomyces phage Av-1 | Actinomyces phage Av-1 Dybvigvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17171 | 49.502 | Dybvigvirus | Unclassified | Podoviridae | Actinomyces |
| AY266303 | Aeromonas phage Aeh1 | Aeromonas phage Aeh1 Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 233234 | 42.779 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| CP084202 | Akkermansia phage DTMo-2021a | Akkermansia phage DTMo-2021a Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31946 | 54.298 | Unclassified | Unclassified | Unclassified | Akkermansia |
| MZ322895 | Klebsiella phage vB_KpnM_VAC13 | Klebsiella phage vB_KpnM_VAC13 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174826 | 41.89 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MT543041 | Serratia phage vB_SmaP-UFV01 | Serratia phage vB_SmaP-UFV01 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39164 | 48.825 | Teseptimavirus | Studiervirinae | Autographiviridae | Serratia |
| MT980841 | Ruminococcus phage phiRM10 | Ruminococcus phage phiRM10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36920 | 41.533 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT980840 | Ruminococcus phage phiRgIBDN1 | Ruminococcus phage phiRgIBDN1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37233 | 41.485 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT980839 | Ruminococcus phage phiRgPS_6 | Ruminococcus phage phiRgPS_6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37780 | 41.316 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT980838 | Ruminococcus phage phiRg519T2 | Ruminococcus phage phiRg519T2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36865 | 41.809 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT980837 | Ruminococcus phage phiRg507T2_3 | Ruminococcus phage phiRg507T2_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36510 | 42.049 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT980836 | Ruminococcus phage phiRg507T2_2 | Ruminococcus phage phiRg507T2_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37304 | 41.878 | Unclassified | Unclassified | Siphoviridae | Ruminococcus |
| MT632020 | Salmonella phage UPWr_S5 | Salmonella phage UPWr_S5 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43600 | 49.989 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT632019 | Salmonella phage UPWr_S4 | Salmonella phage UPWr_S4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42506 | 50.019 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT632018 | Salmonella phage UPWr_S3 | Salmonella phage UPWr_S3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42618 | 50.012 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT632017 | Salmonella phage UPWr_S2 | Salmonella phage UPWr_S2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42487 | 50.025 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT588083 | Salmonella virus Jersey | Salmonella virus Jersey Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43193 | 49.848 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MZ666966 | Xanthomonas phage N1 | Xanthomonas phage N1 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41152 | 61.951 | Unclassified | Okabevirinae | Autographiviridae | Xanthomonas |
| MZ666965 | Xanthomonas phage N3 | Xanthomonas phage N3 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38454 | 61.84 | Unclassified | Okabevirinae | Autographiviridae | Xanthomonas |
| MZ612130 | Klebsiella phage vB_Kpn-VAC66 | Klebsiella phage vB_Kpn-VAC66 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178532 | 41.724 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| KX198612 | Pseudomonas phage MD8 | Pseudomonas phage MD8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43277 | 61.134 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ334527 | Halorubrum virus HATV-3 | Halorubrum virus HATV-3 Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42293 | 51.063 | Unclassified | Unclassified | Unclassified | Halorubrum |
| MZ334526 | Halorubrum virus HRTV-29 | Halorubrum virus HRTV-29 Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36603 | 60.793 | Unclassified | Unclassified | Unclassified | Halorubrum |
| MZ334525 | Halorubrum virus HATV-2 | Halorubrum virus HATV-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63301 | 49.713 | Unclassified | Unclassified | Myoviridae | Halorubrum |
| MZ334524 | Haloarcula virus HCTV-16 | Haloarcula virus HCTV-16 Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104681 | 56.902 | Unclassified | Unclassified | Unclassified | Haloarcula |
| MZ334523 | Haloarcula virus HVTV-2 | Haloarcula virus HVTV-2 Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102319 | 58.271 | Unclassified | Unclassified | Unclassified | Haloarcula |
| MZ334522 | Halorubrum virus HRTV-27 | Halorubrum virus HRTV-27 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56593 | 62.022 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334521 | Halorubrum virus HRTV-25 | Halorubrum virus HRTV-25 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61934 | 55.85 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334520 | Haloarcula virus HCTV-15 | Haloarcula virus HCTV-15 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71672 | 59.376 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334519 | Haloarcula virus HCTV-6 | Haloarcula virus HCTV-6 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71672 | 59.377 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334518 | Halorubrum virus HRTV-11 | Halorubrum virus HRTV-11 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71449 | 59.872 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334517 | Halorubrum virus HRTV-2 | Halorubrum virus HRTV-2 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68923 | 60.335 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334516 | Halorubrum virus HRTV-15 | Halorubrum virus HRTV-15 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76242 | 57.123 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334515 | Haloarcula virus HJTV-3 | Haloarcula virus HJTV-3 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77353 | 57.046 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334514 | Halorubrum virus HSTV-3 | Halorubrum virus HSTV-3 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76908 | 57.123 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334513 | Haloarcula virus HJTV-2 | Haloarcula virus HJTV-2 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76821 | 56.978 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334512 | Halorubrum virus HRTV-24 | Halorubrum virus HRTV-24 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77537 | 56.839 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334511 | Halorubrum virus HRTV-21 | Halorubrum virus HRTV-21 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76556 | 56.303 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334510 | Halorubrum virus HRTV-13 | Halorubrum virus HRTV-13 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76666 | 56.197 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334509 | Haloarcula virus HJTV-1 | Haloarcula virus HJTV-1 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78012 | 56.47 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334508 | Haloarcula virus HCTV-10 | Haloarcula virus HCTV-10 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75019 | 56.384 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334507 | Haloarcula virus HCTV-8 | Haloarcula virus HCTV-8 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75019 | 56.383 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334506 | Halorubrum virus HRTV-16 | Halorubrum virus HRTV-16 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77109 | 56.425 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334505 | Halorubrum virus HRTV-9 | Halorubrum virus HRTV-9 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75429 | 56.168 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334504 | Haloarcula virus HCTV-11 | Haloarcula virus HCTV-11 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76008 | 56.432 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334503 | Haloarcula virus HCTV-9 | Haloarcula virus HCTV-9 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76008 | 56.424 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334502 | Haloarcula virus HCTV-7 | Haloarcula virus HCTV-7 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76008 | 56.43 | Haloferacalesvirus | Unclassified | Myoviridae | Haloarcula |
| MZ334501 | Halorubrum virus HSTV-4 | Halorubrum virus HSTV-4 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75181 | 56.711 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334500 | Halorubrum virus HRTV-26 | Halorubrum virus HRTV-26 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77578 | 56.216 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334499 | Halorubrum virus HRTV-22 | Halorubrum virus HRTV-22 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76812 | 56.39 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334498 | Halorubrum virus HRTV-20 | Halorubrum virus HRTV-20 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76259 | 56.325 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334497 | Halorubrum virus HRTV-18 | Halorubrum virus HRTV-18 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76361 | 56.353 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334496 | Halorubrum virus HRTV-10 | Halorubrum virus HRTV-10 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76759 | 56.653 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334495 | Halorubrum virus HRTV-23 | Halorubrum virus HRTV-23 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77739 | 56.542 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334494 | Halorubrum virus HRTV-19 | Halorubrum virus HRTV-19 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77739 | 56.542 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334493 | Halorubrum virus HRTV-17 | Halorubrum virus HRTV-17 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74754 | 57.047 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MZ334492 | Halorubrum virus HRTV-14 | Halorubrum virus HRTV-14 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74355 | 57.037 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MT239398 | Yersinia phage P37 | Yersinia phage P37 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33007 | 51.68 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| OK216879 | Gordonia phage AnarQue | Gordonia phage AnarQue unclassified Sourvirus Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61822 | 68.806 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| MZ417354 | Staphylococcus phage PG-2021_9 | Staphylococcus phage PG-2021_9 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141528 | 30.771 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417353 | Staphylococcus phage PG-2021_93 | Staphylococcus phage PG-2021_93 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43459 | 34.368 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417352 | Staphylococcus phage PG-2021_91 | Staphylococcus phage PG-2021_91 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42188 | 34.967 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417351 | Staphylococcus phage PG-2021_90 | Staphylococcus phage PG-2021_90 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44493 | 34.718 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417350 | Staphylococcus phage PG-2021_8 | Staphylococcus phage PG-2021_8 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139709 | 30.936 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417349 | Staphylococcus phage PG-2021_89 | Staphylococcus phage PG-2021_89 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43039 | 35.108 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417348 | Staphylococcus phage PG-2021_88 | Staphylococcus phage PG-2021_88 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92222 | 29.413 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417347 | Staphylococcus phage PG-2021_87 | Staphylococcus phage PG-2021_87 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141212 | 30.754 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417346 | Staphylococcus phage PG-2021_86 | Staphylococcus phage PG-2021_86 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141965 | 30.829 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417345 | Staphylococcus phage PG-2021_84 | Staphylococcus phage PG-2021_84 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141291 | 30.756 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417344 | Staphylococcus phage PG-2021_76 | Staphylococcus phage PG-2021_76 unclassified Sciuriunavirus Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139439 | 31.591 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417343 | Staphylococcus phage PG-2021_74 | Staphylococcus phage PG-2021_74 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85762 | 29.688 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417342 | Staphylococcus phage PG-2021_68 | Staphylococcus phage PG-2021_68 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91947 | 30.611 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417341 | Staphylococcus phage PG-2021_67 | Staphylococcus phage PG-2021_67 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92064 | 30.573 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417340 | Staphylococcus phage PG-2021_64 | Staphylococcus phage PG-2021_64 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142287 | 30.885 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417339 | Staphylococcus phage PG-2021_5 | Staphylococcus phage PG-2021_5 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92222 | 29.415 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417338 | Staphylococcus phage PG-2021_4 | Staphylococcus phage PG-2021_4 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91860 | 30.613 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417337 | Staphylococcus phage PG-2021_47 | Staphylococcus phage PG-2021_47 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142885 | 30.702 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417336 | Staphylococcus phage PG-2021_46 | Staphylococcus phage PG-2021_46 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86018 | 29.662 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MZ417335 | Staphylococcus phage PG-2021_43 | Staphylococcus phage PG-2021_43 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145090 | 31.338 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417334 | Staphylococcus phage PG-2021_41 | Staphylococcus phage PG-2021_41 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145090 | 31.335 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417333 | Staphylococcus phage PG-2021_40 | Staphylococcus phage PG-2021_40 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142875 | 31.351 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417332 | Staphylococcus phage PG-2021_38 | Staphylococcus phage PG-2021_38 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140647 | 30.801 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417331 | Staphylococcus phage PG-2021_35 | Staphylococcus phage PG-2021_35 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138653 | 30.803 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417330 | Staphylococcus phage PG-2021_33 | Staphylococcus phage PG-2021_33 unclassified Sciuriunavirus Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135943 | 31.673 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417329 | Staphylococcus phage PG-2021_31 | Staphylococcus phage PG-2021_31 unclassified Sciuriunavirus Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139439 | 31.591 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417328 | Staphylococcus phage PG-2021_2 | Staphylococcus phage PG-2021_2 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142223 | 30.853 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417327 | Staphylococcus phage PG-2021_29 | Staphylococcus phage PG-2021_29 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131570 | 30.889 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417326 | Staphylococcus phage PG-2021_27 | Staphylococcus phage PG-2021_27 unclassified Herelleviridae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128279 | 29.667 | Unclassified | Unclassified | Herelleviridae | Staphylococcus |
| MZ417325 | Staphylococcus phage PG-2021_23 | Staphylococcus phage PG-2021_23 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139827 | 28.001 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417324 | Staphylococcus phage PG-2021_22 | Staphylococcus phage PG-2021_22 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144280 | 31.332 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417323 | Staphylococcus phage PG-2021_1 | Staphylococcus phage PG-2021_1 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143764 | 27.961 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417322 | Staphylococcus phage PG-2021_19 | Staphylococcus phage PG-2021_19 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141132 | 31.247 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417321 | Staphylococcus phage PG-2021_18 | Staphylococcus phage PG-2021_18 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138844 | 30.799 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417320 | Staphylococcus phage PG-2021_17 | Staphylococcus phage PG-2021_17 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144971 | 30.852 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417319 | Staphylococcus phage PG-2021_16 | Staphylococcus phage PG-2021_16 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141321 | 30.797 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417318 | Staphylococcus phage PG-2021_15 | Staphylococcus phage PG-2021_15 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145090 | 31.327 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417317 | Staphylococcus phage PG-2021_14 | Staphylococcus phage PG-2021_14 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145090 | 31.329 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417316 | Staphylococcus phage PG-2021_12 | Staphylococcus phage PG-2021_12 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145091 | 31.329 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ417315 | Staphylococcus phage PG-2021_10 | Staphylococcus phage PG-2021_10 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141528 | 30.77 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ422438 | Listeria phage LPJP1 | Listeria phage LPJP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223580 | 25.923 | Unclassified | Unclassified | Myoviridae | Listeria |
| MW650841 | Staphylococcus phage B_UFSM1 | Staphylococcus phage B_UFSM1 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41971 | 33.988 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MW627293 | Staphylococcus phage B_UFSM3 | Staphylococcus phage B_UFSM3 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42835 | 34.126 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MZ334528 | Halorubrum virus HRTV-28 | Halorubrum virus HRTV-28 Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35270 | 64.278 | Unclassified | Unclassified | Unclassified | Halorubrum |
| MZ443785 | Erwinia phage pEa_SNUABM_22 | Erwinia phage pEa_SNUABM_22 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265146 | 49.569 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443782 | Erwinia phage pEa_SNUABM_16 | Erwinia phage pEa_SNUABM_16 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266838 | 49.58 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ868718 | Pseudomonas phage psageK9 | Pseudomonas phage psageK9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51465 | 58.527 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ868717 | Pseudomonas phage pphageBV72 | Pseudomonas phage pphageBV72 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41435 | 56.633 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ868716 | Pseudomonas phage pphageT12 | Pseudomonas phage pphageT12 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41889 | 56.526 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ868715 | Pseudomonas phage pphageT21 | Pseudomonas phage pphageT21 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41512 | 56.617 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ868714 | Pseudomonas phage pphageB21 | Pseudomonas phage pphageB21 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41708 | 56.651 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ868713 | Pseudomonas phage psageK4e | Pseudomonas phage psageK4e unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94020 | 47.884 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MZ443778 | Erwinia phage pEa_SNUABM_30 | Erwinia phage pEa_SNUABM_30 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 259771 | 49.298 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443772 | Erwinia phage pEa_SNUABM_23 | Erwinia phage pEa_SNUABM_23 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265846 | 49.516 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ936316 | Acinetobacter phage APK20 | Acinetobacter phage APK20 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42351 | 39.279 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ936315 | Acinetobacter phage APK15 | Acinetobacter phage APK15 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41092 | 39.3 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ936314 | Acinetobacter phage APK86 | Acinetobacter phage APK86 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41297 | 39.25 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ868725 | Acinetobacter phage APK16 | Acinetobacter phage APK16 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41135 | 39.441 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ868726 | Acinetobacter phage APK77 | Acinetobacter phage APK77 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41350 | 39.156 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ868724 | Acinetobacter phage APK09 | Acinetobacter phage APK09 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41477 | 39.154 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ747522 | Arthrobacter phage Klevey | Arthrobacter phage Klevey Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50075 | 69.426 | Unclassified | Unclassified | Myoviridae | Arthrobacter |
| MZ747521 | Mycobacterium phage Timothy | Mycobacterium phage Timothy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51970 | 63.7 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747520 | Arthrobacter phage Dynamite | Arthrobacter phage Dynamite unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57846 | 45.27 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ747519 | Mycobacterium phage Tootsieroll | Mycobacterium phage Tootsieroll unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57595 | 61.441 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747518 | Mycobacterium phage Calamitous | Mycobacterium phage Calamitous unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67371 | 68.898 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ747517 | Mycobacterium phage Lucivia | Mycobacterium phage Lucivia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52626 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747516 | Arthrobacter phage Eraser | Arthrobacter phage Eraser Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43608 | 66.598 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MZ747515 | Microbacterium phage SBlackberry | Microbacterium phage SBlackberry unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42048 | 66.857 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ747514 | Mycobacterium phage HaveUMetTed | Mycobacterium phage HaveUMetTed unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49799 | 64.076 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747513 | Microbacterium phage Swervy | Microbacterium phage Swervy unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41510 | 66.748 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ747512 | Mycobacterium phage MadMen | Mycobacterium phage MadMen unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57759 | 61.279 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747511 | Mycobacterium phage Mori | Mycobacterium phage Mori unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71672 | 65.31 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747510 | Gordonia phage Whitney | Gordonia phage Whitney unclassified Getalongvirus Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55784 | 63.014 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MZ747509 | Mycobacterium phage Phelipe | Mycobacterium phage Phelipe unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51333 | 63.834 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ747508 | Mycobacterium phage Coleslaw | Mycobacterium phage Coleslaw unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41893 | 66.591 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ969646 | Bacillus phage BUCT082 | Bacillus phage BUCT082 unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125016 | 34.789 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| MZ983385 | Psychrobacillus phage PVJ1 | Psychrobacillus phage PVJ1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53187 | 37.015 | Unclassified | Unclassified | Myoviridae | Psychrobacillus |
| MZ967493 | Acinetobacter phage APK37 | Acinetobacter phage APK37 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40966 | 39.184 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ923531 | Salmonella phage PRF-SP1 | Salmonella phage PRF-SP1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39966 | 50.255 | Kayfunavirus | Studiervirinae | Autographiviridae | Salmonella |
| MZ926748 | Pseudomonas phage AIIMS-Plu-RaNi | Pseudomonas phage AIIMS-Plu-RaNi unclassified Abidjanvirus Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46647 | 64.452 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ868638 | Escherichia phage vB_EcoM-pEK20 | Escherichia phage vB_EcoM-pEK20 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169226 | 37.678 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ833439 | Enterobacteria phage AH67C600_Q5 | Enterobacteria phage AH67C600_Q5 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38824 | 48.382 | Teseptimavirus | Studiervirinae | Autographiviridae | Enterobacteria |
| MZ832314 | Escherichia phage 26 | Escherichia phage 26 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42462 | 50.422 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MZ447858 | Vibrio phage BUCT194 | Vibrio phage BUCT194 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73099 | 38.477 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MZ803112 | Synechococcus T7-like virus S-TIP28 | Synechococcus T7-like virus S-TIP28 Voetvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45749 | 52.449 | Voetvirus | Unclassified | Autographiviridae | Synechococcus |
| MZ272341 | Enterococcus phage vB_EfaS_785CC | Enterococcus phage vB_EfaS_785CC unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40956 | 34.781 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MZ826354 | Pseudomonas phage SNK | Pseudomonas phage SNK unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40818 | 57.232 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826353 | Pseudomonas phage REC | Pseudomonas phage REC unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98432 | 48.985 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MZ826352 | Pseudomonas phage QAC | Pseudomonas phage QAC unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40336 | 57.7 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826351 | Pseudomonas phage NOI | Pseudomonas phage NOI unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40496 | 57.258 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826350 | Pseudomonas phage M5.1 | Pseudomonas phage M5.1 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98536 | 49.036 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MZ826349 | Pseudomonas phage M3.1 | Pseudomonas phage M3.1 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40502 | 57.251 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826348 | Pseudomonas phage M1.2 | Pseudomonas phage M1.2 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40502 | 57.264 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826347 | Pseudomonas phage M1.1 | Pseudomonas phage M1.1 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40502 | 57.249 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826346 | Pseudomonas phage JOR | Pseudomonas phage JOR unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40531 | 57.215 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826345 | Pseudomonas phage FMS | Pseudomonas phage FMS unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40215 | 56.616 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ826344 | Pseudomonas phage ALEA | Pseudomonas phage ALEA unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40502 | 57.242 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ753803 | Escherichia phage U115 | Escherichia phage U115 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166986 | 35.424 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ779062 | Klebsiella phage pJN2-26 | Klebsiella phage pJN2-26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109952 | 48.999 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ955870 | Glutamicibacter phage BIM BV-113 | Glutamicibacter phage BIM BV-113 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43694 | 57.102 | Unclassified | Unclassified | Siphoviridae | Glutamicibacter |
| MZ955869 | Lactococcus phage BIM BV-114 | Lactococcus phage BIM BV-114 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21499 | 35.904 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MZ826699 | Escherichia phage P817 | Escherichia phage P817 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50751 | 47.087 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| OK040171 | Pseudomonas phage PHB09 | Pseudomonas phage PHB09 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94844 | 45.327 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MZ751040 | Bacillus phage phi18 | Bacillus phage phi18 unclassified Okubovirus Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147298 | 40.557 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| MZ368140 | Lactococcus phage PLG-II | Lactococcus phage PLG-II unclassified Uwajimavirus Uwajimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32271 | 37.737 | Uwajimavirus | Unclassified | Siphoviridae | Lactococcus |
| MZ773648 | Dinoroseobacter phage vB_DshP-R7L | Dinoroseobacter phage vB_DshP-R7L unclassified Plymouthvirus Plymouthvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75871 | 48.958 | Plymouthvirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| MZ851152 | Aeromonas phage PZL-Ah8 | Aeromonas phage PZL-Ah8 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40855 | 51.891 | Unclassified | Studiervirinae | Autographiviridae | Aeromonas |
| MZ520833 | Salmonella phage BIS20 | Salmonella phage BIS20 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29512 | 53.182 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| MZ520832 | Salmonella phage SSBI34 | Salmonella phage SSBI34 unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141095 | 43.926 | Certrevirus | Vequintavirinae | Myoviridae | Salmonella |
| MT577844 | Salmonella phage vB_StyS-LmqsSP1 | Salmonella phage vB_StyS-LmqsSP1 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109938 | 38.846 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MZ488273 | Staphylococcus phage vB_SarS_BM31 | Staphylococcus phage vB_SarS_BM31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42271 | 34.591 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MN905546 | Pseudomonas phage PN05 | Pseudomonas phage PN05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 214121 | 49.814 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MN432485 | Shigella phage SFP17 | Shigella phage SFP17 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51296 | 44.101 | Hanrivervirus | Tempevirinae | Drexlerviridae | Shigella |
| MZ779063 | Staphylococcus phage vB_SauM-UFV_DC4 | Staphylococcus phage vB_SauM-UFV_DC4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263185 | 25.029 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MZ681520 | Mycobacterium phage Weher20 | Mycobacterium phage Weher20 unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68684 | 66.371 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ681519 | Mycobacterium phage MilanaBonita | Mycobacterium phage MilanaBonita unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56251 | 62.058 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ681518 | Mycobacterium phage LarryKay | Mycobacterium phage LarryKay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50916 | 64.184 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ681517 | Gordonia phage Dolores | Gordonia phage Dolores Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47336 | 66.562 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ681516 | Mycobacterium phage Newrala | Mycobacterium phage Newrala unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52566 | 61.584 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ681515 | Gordonia phage Hortense | Gordonia phage Hortense Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53182 | 65.597 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ681514 | Gordonia phage Malibo | Gordonia phage Malibo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49417 | 66.601 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ681513 | Microbacterium phage TinyMiny | Microbacterium phage TinyMiny Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57042 | 62.712 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ681512 | Mycobacterium phage DirtMcgirt | Mycobacterium phage DirtMcgirt unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54396 | 61.067 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ681511 | Mycobacterium phage SilverDipper | Mycobacterium phage SilverDipper unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155812 | 64.698 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MZ681510 | Gordonia phage Baddon | Gordonia phage Baddon unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57691 | 68.186 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| MZ681509 | Gordonia phage Gizermo | Gordonia phage Gizermo unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.619 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MZ681508 | Gordonia Phage Boohoo | Gordonia Phage Boohoo unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52178 | 61.949 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MZ681507 | Gordonia phage Puppers | Gordonia phage Puppers unclassified Gustavvirus Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43098 | 67.702 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| MZ681506 | Arthrobacter phage Asa16 | Arthrobacter phage Asa16 unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43601 | 66.643 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ681505 | Mycobacterium phage Tarkin | Mycobacterium phage Tarkin unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75998 | 62.99 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MW929183 | Escherichia phage FS2B | Escherichia phage FS2B unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18619 | 41.78 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MW929182 | Escherichia phage FS2B | Escherichia phage FS2B unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58790 | 42.364 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MN384979 | Salmonella phage vB_SentM_Phi_10 | Salmonella phage vB_SentM_Phi_10 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159043 | 44.707 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN384978 | Escherichia phage vB_EcolP_P433.1 | Escherichia phage vB_EcolP_P433.1 Tuodvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43865 | 45.506 | Tuodvirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MW960038 | Rhizobium phage RHph_X3_15 | Rhizobium phage RHph_X3_15 unclassified Huelvavirus Huelvavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76185 | 45.09 | Huelvavirus | Unclassified | Schitoviridae | Rhizobium |
| MW960037 | Rhizobium phage RHph_X2_26 | Rhizobium phage RHph_X2_26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68522 | 59.131 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW960036 | Rhizobium phage RHph_X2_28B | Rhizobium phage RHph_X2_28B unclassified Huelvavirus Huelvavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76359 | 45.073 | Huelvavirus | Unclassified | Schitoviridae | Rhizobium |
| MW960035 | Rhizobium phage RHph_X92 | Rhizobium phage RHph_X92 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6054 | 59.861 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MW960034 | Rhizobium phage RHph_X3_9 | Rhizobium phage RHph_X3_9 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44896 | 56.119 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MW960033 | Rhizobium phage RHph_X3_2 | Rhizobium phage RHph_X3_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55597 | 58.922 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW960032 | Rhizobium phage RHph_X2_25 | Rhizobium phage RHph_X2_25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68757 | 58.909 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW960031 | Rhizobium phage RHph_X2_24 | Rhizobium phage RHph_X2_24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55689 | 51.358 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW960030 | Rhizobium phage RHph_X66 | Rhizobium phage RHph_X66 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120445 | 59.43 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW960029 | Rhizobium phage RHph_X94 | Rhizobium phage RHph_X94 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.402 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MW057918 | Escherichia phage Msk | Escherichia phage Msk unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45053 | 44.197 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MZ573923 | Staphylococcus phage vB_ScoM-PSC1 | Staphylococcus phage vB_ScoM-PSC1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140342 | 30.18 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ856343 | Mycobacterium phage EasyJones | Mycobacterium phage EasyJones unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154315 | 64.72 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MW749010 | Hafnia phage vB_HpaM_SarahDanielle | Hafnia phage vB_HpaM_SarahDanielle unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86185 | 37.997 | Unclassified | Ounavirinae | Myoviridae | Hafnia |
| MW749008 | Hafnia phage vB_HpaM_Meifeng | Hafnia phage vB_HpaM_Meifeng Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48503 | 45.721 | Unclassified | Unclassified | Myoviridae | Hafnia |
| MW749004 | Hafnia phage vB_HpaM_Zyzzx | Hafnia phage vB_HpaM_Zyzzx unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85869 | 38.291 | Unclassified | Ounavirinae | Myoviridae | Hafnia |
| MW633168 | Enterococcus phage MDA2 | Enterococcus phage MDA2 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140226 | 35.844 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MW623430 | Enterococcus phage MDA1 | Enterococcus phage MDA1 unclassified Copernicusvirus Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18058 | 32.96 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MW354678 | Bacillus phage BSTP12 | Bacillus phage BSTP12 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55418 | 39.908 | Unclassified | Unclassified | Podoviridae | Bacillus |
| MW354677 | Bacillus phage BSTP10 | Bacillus phage BSTP10 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55450 | 39.917 | Unclassified | Unclassified | Podoviridae | Bacillus |
| MW354676 | Bacillus phage BSTP8 | Bacillus phage BSTP8 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55448 | 39.908 | Unclassified | Unclassified | Podoviridae | Bacillus |
| KX961385 | Bordetella virus LK3 | Bordetella virus LK3 unclassified Vojvodinavirus Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59831 | 63.922 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| KY000221 | Bordetella phage CN1 | Bordetella phage CN1 Bordetella virus CN1 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59779 | 63.974 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| KY000220 | Bordetella phage FP1 | Bordetella phage FP1 Bordetella virus FP1 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60332 | 64.182 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| KY000219 | Bordetella phage CN2 | Bordetella phage CN2 Bordetella virus CN2 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62030 | 64.034 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| KY000218 | Bordetella phage MW2 | Bordetella phage MW2 Bordetella virus MW2 Vojvodinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60160 | 64.058 | Vojvodinavirus | Unclassified | Siphoviridae | Bordetella |
| MZ577098 | Methylophilales phage Melnitz-3 EXVC039M | Methylophilales phage Melnitz-3 EXVC039M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141372 | 37.623 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MZ577097 | Methylophilales phage Melnitz EXVC044M | Methylophilales phage Melnitz EXVC044M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141548 | 37.565 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MZ577096 | Methylophilales phage Melnitz-2 EXVC040M | Methylophilales phage Melnitz-2 EXVC040M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141548 | 37.565 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MZ577095 | Methylophilales phage Melnitz-1 EXVC043M | Methylophilales phage Melnitz-1 EXVC043M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141552 | 37.563 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MZ553931 | Pseudomonas phage vB_PaeA_55_1W | Pseudomonas phage vB_PaeA_55_1W unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42546 | 62.286 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ553930 | Pseudomonas phage vB_PaeM_198 | Pseudomonas phage vB_PaeM_198 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66288 | 55.668 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW824434 | Vibrio phage PS65A.1 | Vibrio phage PS65A.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62006 | 48.066 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824431 | Vibrio phage PS14A.1 | Vibrio phage PS14A.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84620 | 46.316 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824429 | Vibrio phage 82E33.2 | Vibrio phage 82E33.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47279 | 45.729 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824428 | Vibrio phage 82E32.3 | Vibrio phage 82E32.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47214 | 45.749 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824427 | Vibrio phage 82E32.2 | Vibrio phage 82E32.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47278 | 45.759 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824421 | Vibrio phage 51E28.1 | Vibrio phage 51E28.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83109 | 45.752 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824420 | Vibrio phage 38E33.6a | Vibrio phage 38E33.6a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47278 | 45.746 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824418 | Vibrio phage 268E42.1 | Vibrio phage 268E42.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60078 | 48.331 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824417 | Vibrio phage 264E42.1 | Vibrio phage 264E42.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82450 | 45.734 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824414 | Vibrio phage 19E33.1 | Vibrio phage 19E33.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47257 | 45.773 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824413 | Vibrio phage 14E30.2 | Vibrio phage 14E30.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44322 | 41.79 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824412 | Vibrio phage 14E30.1 | Vibrio phage 14E30.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46023 | 41.909 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824409 | Vibrio phage PS32B.6 | Vibrio phage PS32B.6 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48144 | 43.231 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW824408 | Vibrio phage PS32B.11 | Vibrio phage PS32B.11 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48140 | 43.222 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW824407 | Vibrio phage PS32B.1 | Vibrio phage PS32B.1 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48078 | 43.228 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW824406 | Vibrio phage PS25B.1 | Vibrio phage PS25B.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44450 | 46.448 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824405 | Vibrio phage 75E35.1 | Vibrio phage 75E35.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824404 | Vibrio phage 69E27.1 | Vibrio phage 69E27.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40715 | 49.942 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824403 | Vibrio phage 66E30.1 | Vibrio phage 66E30.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37073 | 43.471 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824402 | Vibrio phage 64E30.1 | Vibrio phage 64E30.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37073 | 43.471 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824401 | Vibrio phage 5P1a | Vibrio phage 5P1a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37984 | 43.911 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824400 | Vibrio phage 51E28.4 | Vibrio phage 51E28.4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83109 | 45.741 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824399 | Vibrio phage 44E38.2 | Vibrio phage 44E38.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824398 | Vibrio phage 44E38.1 | Vibrio phage 44E38.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824397 | Vibrio phage 431E48.2 | Vibrio phage 431E48.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49909 | 42.98 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824396 | Vibrio phage 431E46.1 | Vibrio phage 431E46.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49909 | 42.982 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824395 | Vibrio phage 431E45.1 | Vibrio phage 431E45.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49909 | 42.958 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824393 | Vibrio phage 36E38.1 | Vibrio phage 36E38.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824392 | Vibrio phage 34E29.1 | Vibrio phage 34E29.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824391 | Vibrio phage 340E47.2 | Vibrio phage 340E47.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37921 | 43.899 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824390 | Vibrio phage 252E42.2 | Vibrio phage 252E42.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58955 | 44.688 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824389 | Vibrio phage 24E35.2 | Vibrio phage 24E35.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37073 | 43.471 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824388 | Vibrio phage 24E30.2 | Vibrio phage 24E30.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37073 | 43.471 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824387 | Vibrio phage 23E28.1 | Vibrio phage 23E28.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40729 | 49.942 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824386 | Vibrio phage 234P8 | Vibrio phage 234P8 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52297 | 42.962 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824384 | Vibrio phage 219E41.1 | Vibrio phage 219E41.1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52283 | 42.735 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824383 | Vibrio phage PS34B.2 | Vibrio phage PS34B.2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48100 | 42.601 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824382 | Vibrio phage PS15B.4 | Vibrio phage PS15B.4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49407 | 44.093 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824381 | Vibrio phage PS15B.3 | Vibrio phage PS15B.3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48480 | 44.026 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824380 | Vibrio phage PS15B.2 | Vibrio phage PS15B.2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49478 | 44.068 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824379 | Vibrio phage 98E28.6a | Vibrio phage 98E28.6a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48607 | 42.51 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824378 | Vibrio phage 91E28.1a | Vibrio phage 91E28.1a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48607 | 42.549 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824375 | Vibrio phage 207E48.1 | Vibrio phage 207E48.1 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162787 | 45.592 | Unclassified | Unclassified | Ackermannviridae | Vibrio |
| MW824374 | Vibrio phage 207E29.1 | Vibrio phage 207E29.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48367 | 42.711 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824373 | Vibrio phage 18E29.1 | Vibrio phage 18E29.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48552 | 42.532 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824371 | Vibrio phage 184E37.1 | Vibrio phage 184E37.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136415 | 37.821 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824370 | Vibrio phage 144E46.1 | Vibrio phage 144E46.1 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157512 | 44.868 | Unclassified | Unclassified | Ackermannviridae | Vibrio |
| MW824369 | Vibrio phage 12E28.1 | Vibrio phage 12E28.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48552 | 42.532 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824433 | Vibrio phage PS34B.1 | Vibrio phage PS34B.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59928 | 48.688 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824432 | Vibrio phage PS17B.1 | Vibrio phage PS17B.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61131 | 48.342 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824430 | Vibrio phage PS10B.1 | Vibrio phage PS10B.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60587 | 48.573 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824426 | Vibrio phage 70E37.6 | Vibrio phage 70E37.6 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113569 | 42.564 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824425 | Vibrio phage 70E37.1 | Vibrio phage 70E37.1 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113189 | 42.553 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824424 | Vibrio phage 70E35.6 | Vibrio phage 70E35.6 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113569 | 42.547 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824423 | Vibrio phage 70E35.5a | Vibrio phage 70E35.5a unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114081 | 42.563 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824422 | Vibrio phage 70E35.2 | Vibrio phage 70E35.2 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114053 | 42.539 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824419 | Vibrio phage 294E48.1 | Vibrio phage 294E48.1 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97580 | 42.592 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824416 | Vibrio phage 234P7B | Vibrio phage 234P7B unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114079 | 42.54 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824415 | Vibrio phage 234P1 | Vibrio phage 234P1 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113569 | 42.567 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW824411 | Vibrio phage 104E43.1 | Vibrio phage 104E43.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44308 | 42.994 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824410 | Vibrio phage 103E44.1 | Vibrio phage 103E44.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44461 | 43.051 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW824394 | Vibrio phage 41E34.2 | Vibrio phage 41E34.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 43.502 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824385 | Vibrio phage 219E41.2 | Vibrio phage 219E41.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52283 | 42.74 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW824377 | Vibrio phage 6E35.1a | Vibrio phage 6E35.1a unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253432 | 43.775 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MW824376 | Vibrio phage 355E48.1 | Vibrio phage 355E48.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133102 | 37.849 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW824372 | Vibrio phage 184E37.3a | Vibrio phage 184E37.3a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136362 | 37.863 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW865341 | Vibrio phage 70E38.1 | Vibrio phage 70E38.1 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17772 | 43.197 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW865327 | Vibrio phage 70E38.1 | Vibrio phage 70E38.1 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22047 | 42.323 | Unclassified | Unclassified | Demerecviridae | Vibrio |
| MW865319 | Vibrio phage 11E33.1 | Vibrio phage 11E33.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26480 | 46.246 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW865307 | Vibrio phage PS35B.3 | Vibrio phage PS35B.3 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37694 | 43.689 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW865302 | Vibrio phage PS35B.1 | Vibrio phage PS35B.1 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30280 | 43.851 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW865299 | Vibrio phage PS32B.3 | Vibrio phage PS32B.3 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37715 | 43.696 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW865297 | Vibrio phage PS32B.2 | Vibrio phage PS32B.2 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37147 | 43.767 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW865293 | Vibrio phage 187P1 | Vibrio phage 187P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15416 | 45.913 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW865292 | Vibrio phage 172P1 | Vibrio phage 172P1 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46895 | 44.395 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MW865291 | Vibrio phage 15E36.1 | Vibrio phage 15E36.1 Gutovirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45211 | 44.558 | Gutovirus | Colwellvirinae | Autographiviridae | Vibrio |
| MZ701794 | Stenotrophomonas phage vB_SmaS_P11 | Stenotrophomonas phage vB_SmaS_P11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44600 | 63.684 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ712044 | Acinetobacter phage BUCT629 | Acinetobacter phage BUCT629 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46325 | 37.587 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MZ576217 | Escherichia phage vB_EcoS-phiEc3 | Escherichia phage vB_EcoS-phiEc3 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45288 | 50.627 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MZ443789 | Erwinia phage pEa_SNUABM_36 | Erwinia phage pEa_SNUABM_36 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261689 | 49.168 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443788 | Erwinia phage pEa_SNUABM_35 | Erwinia phage pEa_SNUABM_35 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261719 | 49.167 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443787 | Erwinia phage pEa_SNUABM_39 | Erwinia phage pEa_SNUABM_39 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265365 | 49.369 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443786 | Erwinia phage pEa_SNUABM_2 | Erwinia phage pEa_SNUABM_2 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265442 | 49.379 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443784 | Erwinia phage pEa_SNUABM_18 | Erwinia phage pEa_SNUABM_18 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265970 | 49.565 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443783 | Erwinia phage pEa_SNUABM_28 | Erwinia phage pEa_SNUABM_28 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 267143 | 49.541 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443780 | Erwinia phage pEa_SNUABM_40 | Erwinia phage pEa_SNUABM_40 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 267121 | 49.472 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443779 | Erwinia phage pEa_SNUABM_33 | Erwinia phage pEa_SNUABM_33 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263237 | 49.326 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443777 | Erwinia phage pEa_SNUABM_17 | Erwinia phage pEa_SNUABM_17 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263916 | 49 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443776 | Erwinia phage pEa_SNUABM_1 | Erwinia phage pEa_SNUABM_1 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266939 | 49.447 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443774 | Erwinia phage pEa_SNUABM_32 | Erwinia phage pEa_SNUABM_32 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265891 | 49.186 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443771 | Erwinia phage pEa_SNUABM_20 | Erwinia phage pEa_SNUABM_20 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265871 | 49.508 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443770 | Erwinia phage pEa_SNUABM_3 | Erwinia phage pEa_SNUABM_3 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266175 | 49.503 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ443769 | Erwinia phage pEp_SNUABM_52 | Erwinia phage pEp_SNUABM_52 Sasquatchvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261095 | 47.181 | Sasquatchvirus | Unclassified | Myoviridae | Erwinia |
| MZ428229 | Klebsiella phage vB_KpnP-VAC1 | Klebsiella phage vB_KpnP-VAC1 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39371 | 50.745 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MZ428228 | Klebsiella phage vB_KpnS-VAC11 | Klebsiella phage vB_KpnS-VAC11 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48826 | 50.856 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428227 | Klebsiella phage vB_kpnS-VAC10 | Klebsiella phage vB_kpnS-VAC10 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48935 | 50.655 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428226 | Klebsiella phage vB_KpnS-VAC8 | Klebsiella phage vB_KpnS-VAC8 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48933 | 50.534 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428225 | Klebsiella phage vB_KpnS-VAC7 | Klebsiella phage vB_KpnS-VAC7 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49684 | 51.254 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428224 | Klebsiella phage vB_KpnS-VAC6 | Klebsiella phage vB_KpnS-VAC6 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51554 | 51.647 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428223 | Klebsiella phage vB_KpnS-VAC5 | Klebsiella phage vB_KpnS-VAC5 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49636 | 50.457 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428222 | Klebsiella phage vB_KpnS-VAC4 | Klebsiella phage vB_KpnS-VAC4 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45558 | 51.117 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428221 | Klebsiella phage vB_kpnS-VAC2 | Klebsiella phage vB_kpnS-VAC2 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51784 | 47.87 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ428230 | Klebsiella phage vB_KpnS-VAC9 | Klebsiella phage vB_KpnS-VAC9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11444 | 50.524 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ820095 | Streptomyces phage Karp | Streptomyces phage Karp unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130557 | 46.996 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MZ820094 | Streptomyces phage Jada | Streptomyces phage Jada unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127753 | 47.086 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MZ820093 | Streptomyces phage Forrest | Streptomyces phage Forrest unclassified Gilsonvirus Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128513 | 47.052 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| MZ820092 | Streptomyces phage SarahRose | Streptomyces phage SarahRose unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51567 | 65.616 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ820091 | Streptomyces phage South40 | Streptomyces phage South40 unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51271 | 65.585 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ820090 | Mycobacterium phage Agape74 | Mycobacterium phage Agape74 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53198 | 63.39 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ820089 | Gordonia phage ChisanaKitsune | Gordonia phage ChisanaKitsune unclassified Bendigovirus Bendigovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88657 | 60.032 | Bendigovirus | Unclassified | Myoviridae | Gordonia |
| MZ820088 | Streptomyces phage Katalie | Streptomyces phage Katalie unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51271 | 65.587 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ820087 | Arthrobacter phage Niobe | Arthrobacter phage Niobe unclassified Yangvirus Yangvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43602 | 66.657 | Yangvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ820086 | Streptomyces phage Bilo | Streptomyces phage Bilo unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43835 | 60.605 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MZ820085 | Mycobacterium phage ScoobyDoobyDoo | Mycobacterium phage ScoobyDoobyDoo Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158235 | 64.196 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| MZ820084 | Microbacterium phage KatChan | Microbacterium phage KatChan unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40349 | 63.005 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ820083 | Microbacterium phage Doobus | Microbacterium phage Doobus unclassified Dismasvirus Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41195 | 66.256 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| LC648443 | Pseudomonas phage vB_PaS-HSN4 | Pseudomonas phage vB_PaS-HSN4 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44534 | 53.458 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ622183 | Mycobacterium phage Moostard | Mycobacterium phage Moostard unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69480 | 59.128 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622182 | Microbacterium phage Hasitha | Microbacterium phage Hasitha unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39952 | 63.611 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ622181 | Mycobacterium phage Forge | Mycobacterium phage Forge unclassified Gilesvirus Gilesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53746 | 67.447 | Gilesvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622180 | Microbacterium phage Lola20 | Microbacterium phage Lola20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17388 | 68.634 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ622179 | Microbacterium phage Honeyfin | Microbacterium phage Honeyfin unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53154 | 68.994 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ622178 | Gordonia phage Tracker | Gordonia phage Tracker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66607 | 65.903 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ622177 | Gordonia phage PinkCoffee | Gordonia phage PinkCoffee Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58559 | 67.771 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ622176 | Gordonia phage Looper | Gordonia phage Looper unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51468 | 62.025 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MZ622175 | Microbacterium phage Juicer | Microbacterium phage Juicer unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41099 | 62.766 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ622174 | Mycobacterium phage Sarma624 | Mycobacterium phage Sarma624 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54312 | 61.463 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622173 | Mycobacterium phage Katniss | Mycobacterium phage Katniss unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68447 | 66.508 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ622172 | Mycobacterium phage Giraffe | Mycobacterium phage Giraffe unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68869 | 66.449 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ622171 | Mycobacterium phage Dignity | Mycobacterium phage Dignity unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53305 | 63.274 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622170 | Microbacterium phage Blett | Microbacterium phage Blett unclassified Appavirus Appavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38630 | 68.002 | Appavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ622169 | Mycobacterium phage Beem | Mycobacterium phage Beem unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113128 | 60.793 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622168 | Gordonia phage Nadmeg | Gordonia phage Nadmeg unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61137 | 51.576 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MZ622167 | Gordonia phage Vine | Gordonia phage Vine Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48092 | 60.445 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ622166 | Microbacterium phage Platte | Microbacterium phage Platte Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62728 | 64.718 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ622165 | Gordonia phage LonelyBoi | Gordonia phage LonelyBoi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53604 | 66.4 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ622164 | Gordonia phage MissRona | Gordonia phage MissRona Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53623 | 67.092 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MZ622162 | Mycobacterium phage Magsby | Mycobacterium phage Magsby unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43642 | 66.271 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MZ622161 | Gordonia phage Floral | Gordonia phage Floral Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52488 | 66.791 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ622160 | Streptomyces phage RetrieverFever | Streptomyces phage RetrieverFever unclassified Flowerpowervirus Flowerpowervirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46175 | 60.671 | Flowerpowervirus | Beephvirinae | Podoviridae | Streptomyces |
| MZ622159 | Mycobacterium phage Drake94 | Mycobacterium phage Drake94 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49890 | 64.434 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622158 | Mycobacterium phage Thresher | Mycobacterium phage Thresher unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76192 | 63.047 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ622157 | Mycobacterium phage Wiggin | Mycobacterium phage Wiggin unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74903 | 62.931 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ424865 | Klebsiella phage vB_KoxP_ZX8 | Klebsiella phage vB_KoxP_ZX8 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39398 | 53.061 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW119258 | Klebsiella phage vB_KpnS_MK54 | Klebsiella phage vB_KpnS_MK54 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46218 | 48.297 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ327262 | Salmonella phage vB_Si_DR094 | Salmonella phage vB_Si_DR094 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85730 | 39.178 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MZ327261 | Salmonella phage vB_Si_35FD | Salmonella phage vB_Si_35FD unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85818 | 38.925 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MZ311865 | Escherichia phage vB_EcoM_LNA2 | Escherichia phage vB_EcoM_LNA2 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168410 | 37.666 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ311863 | Escherichia phage vB_EcoM_LNA1 | Escherichia phage vB_EcoM_LNA1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88369 | 39.101 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MZ311864 | Escherichia phage vB_EcoM_LNA4 | Escherichia phage vB_EcoM_LNA4 unclassified Seunavirus Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147613 | 46.339 | Seunavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ311862 | Escherichia phage vB_EcoP_LNA3 | Escherichia phage vB_EcoP_LNA3 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39586 | 49.853 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ311861 | Escherichia phage vB_EcoP_LNA5 | Escherichia phage vB_EcoP_LNA5 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39091 | 50.019 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ436830 | Escherichia phage JK1 | Escherichia phage JK1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166057 | 35.483 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ648031 | Microbacterium phage Jannah | Microbacterium phage Jannah Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17428 | 68.648 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW722083 | Escherichia phage vB_EcoP_ZX5 | Escherichia phage vB_EcoP_ZX5 unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39565 | 48.988 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| MW722080 | Klebsiella phage vB_KpnP_ZX1 | Klebsiella phage vB_KpnP_ZX1 unclassified Lastavirus Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60982 | 56.987 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| MZ501272 | Pseudomonas phage AH05 | Pseudomonas phage AH05 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40502 | 57.254 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ501271 | Pseudomonas phage AH02 | Pseudomonas phage AH02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39095 | 54.946 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ501270 | Pantoea phage AH07 | Pantoea phage AH07 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37859 | 52.22 | Unclassified | Unclassified | Myoviridae | Pantoea |
| MZ501269 | Pantoea phage AH01 | Pantoea phage AH01 unclassified Guernseyvirinae Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46062 | 52.232 | Unclassified | Guernseyvirinae | Siphoviridae | Pantoea |
| MZ501268 | Erwinia phage AH06 | Erwinia phage AH06 unclassified Agricanvirus Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 275293 | 48.066 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MZ501267 | Erwinia phage AH04 | Erwinia phage AH04 Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262639 | 43.324 | Petsuvirus | Unclassified | Myoviridae | Erwinia |
| MZ501266 | Erwinia phage AH03 | Erwinia phage AH03 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43866 | 43.774 | Unclassified | Unclassified | Siphoviridae | Erwinia |
| MZ501265 | Bacillus phage 278BB001 | Bacillus phage 278BB001 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158563 | 42.046 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MZ501264 | Bacillus phage 274BB002 | Bacillus phage 274BB002 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52372 | 42.509 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MZ501263 | Bacillus phage 043JT007 | Bacillus phage 043JT007 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161640 | 42.009 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MZ501262 | Bacillus phage 035JT004 | Bacillus phage 035JT004 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163318 | 41.88 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MZ501261 | Bacillus phage 010DV005 | Bacillus phage 010DV005 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161884 | 41.979 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MZ501260 | Bacillus phage 010DV004 | Bacillus phage 010DV004 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161885 | 41.979 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MZ099557 | Acinetobacter phage vB_AbaP_WU2001 | Acinetobacter phage vB_AbaP_WU2001 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40792 | 39.41 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ687409 | Pseudomonas phage vB_Pae_QDWS | Pseudomonas phage vB_Pae_QDWS unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43170 | 62.126 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MZ681930 | Enterobacteria phage phiC600P9 | Enterobacteria phage phiC600P9 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164663 | 35.624 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MZ648041 | Mycobacterium phage DaddyDaniels | Mycobacterium phage DaddyDaniels unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68743 | 66.381 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ648040 | Streptomyces phage Triste | Streptomyces phage Triste unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51568 | 65.688 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ648039 | Streptomyces phage RedBear | Streptomyces phage RedBear unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51271 | 65.579 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ648038 | Mycobacterium phage Tomlarah | Mycobacterium phage Tomlarah unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69188 | 66.468 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ648037 | Mycobacterium phage QuincyRose | Mycobacterium phage QuincyRose unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59719 | 66.473 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ648036 | Streptomyces phage Lizz | Streptomyces phage Lizz Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69287 | 64.246 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MZ648035 | Microbacterium phage Shotgun | Microbacterium phage Shotgun unclassified Quhwahvirus Quhwahvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52659 | 68.852 | Quhwahvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ648034 | Mycobacterium phage Ohno789 | Mycobacterium phage Ohno789 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52817 | 63.606 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ648033 | Streptomyces phage Rooney | Streptomyces phage Rooney Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69514 | 64.278 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MZ648032 | Streptomyces phage Chucky | Streptomyces phage Chucky unclassified Likavirus Likavirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51594 | 65.69 | Likavirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MZ568829 | Shewanella phage vB_SspM_M16-3 | Shewanella phage vB_SspM_M16-3 unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40166 | 46.798 | Unclassified | Peduovirinae | Myoviridae | Shewanella |
| MZ568828 | Shewanella phage vB_SspM_MuM16-2 | Shewanella phage vB_SspM_MuM16-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33702 | 46.849 | Unclassified | Unclassified | Myoviridae | Shewanella |
| MZ568827 | Shewanella phage vB_SspS_MuM16-1 | Shewanella phage vB_SspS_MuM16-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37883 | 47.834 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| MZ568826 | Shewanella phage vB_SspS_KASIA | Shewanella phage vB_SspS_KASIA Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91102 | 40.234 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| MZ592922 | Vibrio phage vB_VpS_C2 | Vibrio phage vB_VpS_C2 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77312 | 48.734 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MZ592921 | Vibrio phage vB_VpP_NS8 | Vibrio phage vB_VpP_NS8 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43252 | 49.244 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MZ592920 | Vibrio phage vB_VpP_G1 | Vibrio phage vB_VpP_G1 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43859 | 47.224 | Unclassified | Melnykvirinae | Autographiviridae | Vibrio |
| MZ474491 | Phage silverpheasant213 | Phage silverpheasant213 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7022 | 54.315 | Unclassified | Unclassified | Inoviridae | Unspecified |
| MZ474490 | Phage toucan80 | Phage toucan80 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6761 | 51.427 | Unclassified | Unclassified | Inoviridae | Unspecified |
| MZ474489 | Phage blackswan219-1 | Phage blackswan219-1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7703 | 56.549 | Unclassified | Unclassified | Inoviridae | Unspecified |
| MZ474488 | Phage blackswan219-6 | Phage blackswan219-6 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6732 | 56.447 | Unclassified | Unclassified | Inoviridae | Unspecified |
| MZ475898 | Erwinia phage pEa_SNUABM_25 | Erwinia phage pEa_SNUABM_25 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263017 | 48.981 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475897 | Erwinia phage pEa_SNUABM_21 | Erwinia phage pEa_SNUABM_21 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265102 | 48.967 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475896 | Erwinia phage pEa_SNUABM_7 | Erwinia phage pEa_SNUABM_7 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265635 | 48.954 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475895 | Erwinia phage pEa_SNUABM_14 | Erwinia phage pEa_SNUABM_14 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 264285 | 48.973 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475894 | Erwinia phage pEa_SNUABM_46 | Erwinia phage pEa_SNUABM_46 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263817 | 48.963 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475893 | Erwinia phage pEa_SNUABM_45 | Erwinia phage pEa_SNUABM_45 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263891 | 48.973 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475892 | Erwinia phage pEa_SNUABM_34 | Erwinia phage pEa_SNUABM_34 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 264306 | 48.955 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ475891 | Erwinia phage pEa_SNUABM_13 | Erwinia phage pEa_SNUABM_13 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 264714 | 48.925 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MZ440881 | Klebsiella phage ZCKP8 | Klebsiella phage ZCKP8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48490 | 47.509 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ369152 | Vibrio phage BUCT006 | Vibrio phage BUCT006 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42603 | 44.884 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MZ326868 | Stenotrophomonas phage Marzo | Stenotrophomonas phage Marzo unclassified Menderavirus Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159384 | 54.007 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MZ326864 | Alcaligenes phage Piluca | Alcaligenes phage Piluca Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35451 | 40.851 | Unclassified | Unclassified | Podoviridae | Alcaligenes |
| MZ326863 | Burkholderia phage Paku | Burkholderia phage Paku unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42727 | 61.863 | Unclassified | Okabevirinae | Autographiviridae | Burkholderia |
| MZ326857 | Stenotrophomonas phage Piffle | Stenotrophomonas phage Piffle unclassified Pokkenvirus Pokkenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76332 | 54.855 | Pokkenvirus | Unclassified | Schitoviridae | Stenotrophomonas |
| MZ171380 | Mycobacterium phage Pumbaa | Mycobacterium phage Pumbaa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51317 | 63.973 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ171379 | Microbacterium phage Stormbreaker8 | Microbacterium phage Stormbreaker8 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41751 | 63.359 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ171378 | Mycobacterium phage Eradicator | Mycobacterium phage Eradicator unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55730 | 61.324 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ171376 | Mycobacterium phage Hoot | Mycobacterium phage Hoot unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50006 | 61.531 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ171373 | Mycobacterium phage DrSeegs | Mycobacterium phage DrSeegs unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76652 | 58.93 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ171372 | Gordonia phage Madi | Gordonia phage Madi unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66439 | 59.525 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MZ170041 | Escherichia phage vB_EcoM_FN | Escherichia phage vB_EcoM_FN unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168411 | 35.356 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ170040 | Escherichia phage vB_EcoM_FJ1 | Escherichia phage vB_EcoM_FJ1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170053 | 35.444 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT732469 | Polaribacter phage Freya_7 | Polaribacter phage Freya_7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46194 | 28.831 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732468 | Polaribacter phage Freya_6 | Polaribacter phage Freya_6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46194 | 28.833 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732467 | Polaribacter phage Freya_5 | Polaribacter phage Freya_5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45722 | 28.883 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT995928 | Pseudomonas phage U5 | Pseudomonas phage U5 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 282684 | 36.858 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MT995927 | Pseudomonas phage U1B | Pseudomonas phage U1B unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 282825 | 36.857 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MT995926 | Pseudomonas phage T2P | Pseudomonas phage T2P unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 282684 | 36.858 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MZ576218 | Escherichia phage vB_EcoS-phiEc4 | Escherichia phage vB_EcoS-phiEc4 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44540 | 50.887 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MZ669808 | Enterobacter phage Phc | Enterobacter phage Phc unclassified Eclunavirus Eclunavirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52494 | 48.566 | Eclunavirus | Unclassified | Drexlerviridae | Enterobacter |
| OU343168 | Pseudomonas phage vB_PaeM_phiRU | Pseudomonas phage vB_PaeM_phiRU Elvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 211869 | 49.374 | Elvirus | Unclassified | Myoviridae | Pseudomonas |
| OU343167 | Pseudomonas phage vB_PaeM_phiLBG22 | Pseudomonas phage vB_PaeM_phiLBG22 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 302555 | 43.598 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| OU509537 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 58588 | 56.196 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OU509536 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 165752 | 39.597 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OU509535 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 347546 | 31.985 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OU509534 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 44964 | 54.214 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OU509533 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 43726 | 54.133 | Unclassified | Unclassified | Unclassified | Klebsiella |
| MZ424863 | Proteus phage vB_PmiM_ZX7 | Proteus phage vB_PmiM_ZX7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43433 | 38.259 | Unclassified | Unclassified | Myoviridae | Proteus |
| MZ292654 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.18 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MZ147816 | Enterococcus phage 113 | Enterococcus phage 113 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155715 | 37.12 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MW794192 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124410 | 37.089 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794191 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123241 | 37.108 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794190 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129632 | 37.115 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794189 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123244 | 37.101 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794188 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129336 | 37.12 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794187 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123239 | 37.115 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794186 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123240 | 37.105 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794185 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123239 | 37.106 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794184 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123217 | 37.107 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794183 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129629 | 37.116 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794182 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129607 | 37.116 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794181 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123895 | 37.113 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794180 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129265 | 37.12 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794179 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128568 | 37.152 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794178 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122800 | 37.163 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794177 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129607 | 37.125 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794176 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125994 | 37.135 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794175 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123219 | 37.101 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794174 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123808 | 37.086 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794173 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130453 | 37.133 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794172 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129620 | 37.125 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794171 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124984 | 37.178 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794170 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124984 | 37.17 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794169 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124926 | 37.172 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794168 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124926 | 37.177 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794167 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124926 | 37.173 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794166 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124987 | 37.183 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794165 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124926 | 37.184 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794164 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.178 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794163 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124985 | 37.175 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794162 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124968 | 37.186 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794161 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125101 | 37.182 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794160 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125101 | 37.184 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794159 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125106 | 37.193 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794158 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.177 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794157 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.186 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794156 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124968 | 37.19 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794155 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.179 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794154 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124986 | 37.183 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794153 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127275 | 37.223 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794152 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124926 | 37.178 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW794151 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125047 | 37.184 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW722082 | Klebsiella phage vB_KpnS_ZX4 | Klebsiella phage vB_KpnS_ZX4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45424 | 48.371 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MZ231001 | Vibrio phage NO16-like 87-9-116 | Vibrio phage NO16-like 87-9-116 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10593 | 47.324 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ231000 | Vibrio phage NO16-like 91-7-154 | Vibrio phage NO16-like 91-7-154 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.348 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230999 | Vibrio phage NO16-like 601_91 | Vibrio phage NO16-like 601_91 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.357 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230998 | Vibrio phage NO16-like 1178_90 | Vibrio phage NO16-like 1178_90 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.385 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230997 | Vibrio phage NO16-like 9014_8 | Vibrio phage NO16-like 9014_8 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.329 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230996 | Vibrio phage NO16-like NB10 | Vibrio phage NO16-like NB10 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.376 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230995 | Vibrio phage NO16-like VIB1 | Vibrio phage NO16-like VIB1 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10593 | 47.446 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230994 | Vibrio phage NO16-like VIB88 | Vibrio phage NO16-like VIB88 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10593 | 47.437 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230993 | Vibrio phage NO16-like VIB93 | Vibrio phage NO16-like VIB93 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10593 | 47.446 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MZ230992 | Vibrio phage NO16-like VIB134 | Vibrio phage NO16-like VIB134 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10597 | 47.24 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MT188666 | Vibrio phage vB_Vipa10 | Vibrio phage vB_Vipa10 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 9318 | 45.031 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT188665 | Vibrio phage vB_Vipa4291 | Vibrio phage vB_Vipa4291 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 9663 | 45.007 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT188664 | Vibrio phage vB_VipaK | Vibrio phage vB_VipaK unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10809 | 44.028 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT188663 | Vibrio phage vB_Vipa36 | Vibrio phage vB_Vipa36 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 9721 | 44.748 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT188662 | Vibrio phage vB_Vipa26 | Vibrio phage vB_Vipa26 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10893 | 44.35 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT887289 | Escherichia phage SME50 | Escherichia phage SME50 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90298 | 38.943 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN692672 | Pseudomonas phage vB_PaeP_Lx18 | Pseudomonas phage vB_PaeP_Lx18 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42735 | 62.162 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KY653117 | Staphylococcus phage IME1323_01 | Staphylococcus phage IME1323_01 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44282 | 34.179 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| MZ333458 | Enterococcus phage SEsuP-1 | Enterococcus phage SEsuP-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39319 | 33.518 | Unclassified | Unclassified | Podoviridae | Enterococcus |
| MZ333457 | Enterococcus phage SSsP-1 | Enterococcus phage SSsP-1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57270 | 39.848 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MZ333456 | Enterococcus phage VEsP-1 | Enterococcus phage VEsP-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39426 | 35.129 | Unclassified | Unclassified | Podoviridae | Enterococcus |
| CP060690 | Liberibacter phage P-Myan16-2 | Liberibacter phage P-Myan16-2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36303 | 39.382 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| MK203850 | Alteromonas phage ZP6 | Alteromonas phage ZP6 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38080 | 50.095 | Unclassified | Unclassified | Podoviridae | Alteromonas |
| LC644557 | Tetragenococcus phage phiWJ7 | Tetragenococcus phage phiWJ7 unclassified Rakietenvirinae Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17405 | 32.916 | Unclassified | Rakietenvirinae | Rountreeviridae | Tetragenococcus |
| MZ398248 | Stenotrophomonas phage BUCT608 | Stenotrophomonas phage BUCT608 unclassified Menderavirus Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160122 | 54.15 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MZ398247 | Enterobacteria phage IME177 | Enterobacteria phage IME177 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39224 | 50.008 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacteria |
| MZ398246 | Escherichia phage IME178 | Escherichia phage IME178 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108588 | 38.783 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MZ398244 | Klebsiella phage IME184 | Klebsiella phage IME184 unclassified Molineuxvirinae Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44598 | 50.253 | Unclassified | Molineuxvirinae | Autographiviridae | Klebsiella |
| MZ398242 | Klebsiella phage IME268 | Klebsiella phage IME268 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49552 | 50.45 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ398241 | Stenotrophomonas phage BUCT626 | Stenotrophomonas phage BUCT626 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61662 | 56.193 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ398240 | Stenotrophomonas phage BUCT627 | Stenotrophomonas phage BUCT627 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61860 | 56.279 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MZ388554 | Microbacterium phage EugeneKrabs | Microbacterium phage EugeneKrabs unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54754 | 58.418 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MZ388553 | Gordonia phage Leroy | Gordonia phage Leroy Horusvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53863 | 62.926 | Horusvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MZ274310 | Mycobacterium phage Boilgate | Mycobacterium phage Boilgate unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57889 | 65.354 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ274305 | Mycobacterium phage ShamWow | Mycobacterium phage ShamWow unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75933 | 63.02 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ274304 | Gordonia phage Agueybana | Gordonia phage Agueybana unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52807 | 66.775 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MZ456039 | Gordonia phage Samman98 | Gordonia phage Samman98 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47548 | 66.491 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ456037 | Microbacterium phage Pickles13 | Microbacterium phage Pickles13 unclassified Appavirus Appavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39237 | 67.844 | Appavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ456036 | Mycobacterium phage Allegro | Mycobacterium phage Allegro unclassified Rosebushvirus Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67439 | 68.923 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MZ456038 | Gordonia phage Shlim410 | Gordonia phage Shlim410 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53182 | 65.603 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ456035 | Microbacterium phage Welcome | Microbacterium phage Welcome Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63952 | 61.685 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MZ456034 | Mycobacterium phage Roksolana | Mycobacterium phage Roksolana unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52702 | 61.434 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ560703 | Salmonella phage vB_SalS_TU03 | Salmonella phage vB_SalS_TU03 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41756 | 47.057 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MZ560701 | Escherichia phage vB_EcoM_TU01 | Escherichia phage vB_EcoM_TU01 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169046 | 37.418 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ503613 | Solobacterium phage SMO_1P | Solobacterium phage SMO_1P Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35406 | 37.979 | Unclassified | Unclassified | Siphoviridae | Solobacterium |
| MZ503612 | Fusobacterium phage FNU_1P | Fusobacterium phage FNU_1P Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41888 | 27.378 | Unclassified | Unclassified | Siphoviridae | Fusobacterium |
| MZ285879 | Pseudomonas phage PA8 | Pseudomonas phage PA8 unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63654 | 63.555 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MZ285878 | Pseudomonas phage PA4 | Pseudomonas phage PA4 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66459 | 55.579 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MZ643272 | Staphylococcus phage vB_Sau-RP15 | Staphylococcus phage vB_Sau-RP15 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139486 | 29.889 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MZ618622 | Acinetobacter phage Abp95 | Acinetobacter phage Abp95 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43176 | 37.843 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MZ598515 | Klebsiella phage P1 | Klebsiella phage P1 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50675 | 50.875 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ593728 | Acinetobacter phage BUCT628 | Acinetobacter phage BUCT628 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44935 | 37.521 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MZ593174 | Acinetobacter phage vB_Api_3043-K38 | Acinetobacter phage vB_Api_3043-K38 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41882 | 39.308 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ570151 | Salmonella phage JNwz02 | Salmonella phage JNwz02 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114390 | 40.215 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MZ614453 | Salmonella phage MS2 LVA | Salmonella phage MS2 LVA Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39726 | 40.49 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW749007 | Bacillus phage vB_BspH_TimeGriffin | Bacillus phage vB_BspH_TimeGriffin unclassified Siophivirus Siophivirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148525 | 39.064 | Siophivirus | Bastillevirinae | Herelleviridae | Bacillus |
| MW748999 | Escherichia phage vB_EcoM_Skers | Escherichia phage vB_EcoM_Skers unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154769 | 40.326 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| GU595417 | Escherichia phage UAB_Phi78 | Escherichia phage UAB_Phi78 Escherichia virus UAB78 Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43984 | 47.449 | Zindervirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MT830620 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3569 | 52.059 | Levivirus | Unclassified | Leviviridae | Escherichia |
| MZ489634 | Salmonella phage D10 | Salmonella phage D10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45715 | 46.077 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MZ502380 | Escherichia phage vB_EcoM_Nami | Escherichia phage vB_EcoM_Nami unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165679 | 35.529 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ502379 | Escherichia phage vB_EcoM_Shinka | Escherichia phage vB_EcoM_Shinka unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162160 | 35.419 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ220766 | Edwardsiella phage PVN09 | Edwardsiella phage PVN09 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37945 | 48.889 | Teseptimavirus | Studiervirinae | Autographiviridae | Edwardsiella |
| MZ220765 | Edwardsiella phage PVN06 | Edwardsiella phage PVN06 unclassified Yokohamavirus Yokohamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44032 | 52.827 | Yokohamavirus | Unclassified | Myoviridae | Edwardsiella |
| MZ598487 | Pseudomonas phage FRS | Pseudomonas phage FRS unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40335 | 57.699 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MZ561532 | Shigella phage PSD9 | Shigella phage PSD9 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169768 | 39.459 | Dhakavirus | Tevenvirinae | Myoviridae | Shigella |
| MZ516827 | Vibrio phage vB_VpP_DE10 | Vibrio phage vB_VpP_DE10 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42871 | 49.192 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MZ394712 | Escherichia phage U1G | Escherichia phage U1G unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73275 | 42.953 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MZ348426 | Pseudomonas phage psageK4 | Pseudomonas phage psageK4 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98440 | 47.751 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MZ348425 | Pseudomonas phage psageB2 | Pseudomonas phage psageB2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50739 | 58.507 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW258709 | Kosakonia phage Kc166A | Kosakonia phage Kc166A unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41374 | 52.664 | Kayfunavirus | Studiervirinae | Autographiviridae | Kosakonia |
| MW258711 | Kosakonia phage Kc166B | Kosakonia phage Kc166B Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40785 | 57.421 | Unclassified | Unclassified | Podoviridae | Kosakonia |
| MW258710 | Kosakonia phage Kc237 | Kosakonia phage Kc237 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40897 | 59.427 | Unclassified | Unclassified | Podoviridae | Kosakonia |
| MW250276 | Kosakonia virus Kc318 | Kosakonia virus Kc318 Cronosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41938 | 55.47 | Cronosvirus | Melnykvirinae | Autographiviridae | Kosakonia |
| MW250275 | Kosakonia virus Kc261 | Kosakonia virus Kc261 unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42432 | 55.475 | Bonnellvirus | Unclassified | Autographiviridae | Kosakonia |
| MW021766 | Serratia phage vB_SmaS_Opt-148 | Serratia phage vB_SmaS_Opt-148 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41293 | 45.741 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MT783706 | Acinetobacter phage Aristophanes | Acinetobacter phage Aristophanes unclassified Beijerinckvirinae Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43505 | 42.528 | Unclassified | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT427400 | Escherichia phage phi2013 | Escherichia phage phi2013 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49833 | 45.556 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MW980074 | Rhizobium phage RHEph27 | Rhizobium phage RHEph27 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46040 | 56.249 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MW980073 | Rhizobium phage RHEph26 | Rhizobium phage RHEph26 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53859 | 56.421 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MW980072 | Rhizobium phage RHEph24 | Rhizobium phage RHEph24 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77615 | 45.999 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MW980071 | Rhizobium phage RHEph22 | Rhizobium phage RHEph22 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77655 | 45.912 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MW980070 | Rhizobium phage RHEph21 | Rhizobium phage RHEph21 Paadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40454 | 58.602 | Paadamvirus | Unclassified | Autographiviridae | Rhizobium |
| MW980069 | Rhizobium phage RHEph19 | Rhizobium phage RHEph19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67843 | 56.489 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW980068 | Rhizobium phage RHEph18 | Rhizobium phage RHEph18 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44197 | 59.095 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MW980067 | Rhizobium phage RHEph17 | Rhizobium phage RHEph17 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6053 | 58.946 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MW980066 | Rhizobium phage RHEph16 | Rhizobium phage RHEph16 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77178 | 45.478 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MW980065 | Rhizobium phage RHEph15 | Rhizobium phage RHEph15 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46206 | 56.291 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MW980064 | Rhizobium phage RHEph12 | Rhizobium phage RHEph12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193606 | 53.39 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MW980063 | Rhizobium phage RHph_N46 | Rhizobium phage RHph_N46 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192456 | 53.475 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MW980062 | Rhizobium phage RHph_I40 | Rhizobium phage RHph_I40 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46695 | 49.142 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| MW980061 | Rhizobium phage RHph_X2_30 | Rhizobium phage RHph_X2_30 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69699 | 57.034 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MW845760 | Erwinia phage pEa_SNUABM_37 | Erwinia phage pEa_SNUABM_37 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 294404 | 49.517 | Unclassified | Unclassified | Myoviridae | Erwinia |
| MW845759 | Erwinia phage pEa_SNUABM_38 | Erwinia phage pEa_SNUABM_38 unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224179 | 50.637 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| MW845758 | Erwinia phage pEa_SNUABM_11 | Erwinia phage pEa_SNUABM_11 Wellingtonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 251480 | 50.715 | Wellingtonvirus | Unclassified | Myoviridae | Erwinia |
| MZ005685 | Mycobacterium phage SmellyB | Mycobacterium phage SmellyB unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52154 | 61.537 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005684 | Microbacterium phage Strathdee | Microbacterium phage Strathdee unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41802 | 63.413 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ005681 | Microbacterium phage Jenos | Microbacterium phage Jenos unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42413 | 63.768 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MZ005674 | Mycobacterium phage Dole | Mycobacterium phage Dole unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60621 | 66.348 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005673 | Mycobacterium phage LochMonster | Mycobacterium phage LochMonster unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51608 | 63.977 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ005670 | Microbacterium phage Mandalorian | Microbacterium phage Mandalorian unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40407 | 63.756 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MW677529 | Rhodobacter phage RcZahn | Rhodobacter phage RcZahn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101599 | 60.715 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677527 | Rhodobacter phage RcTiptonus | Rhodobacter phage RcTiptonus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94091 | 57.872 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677526 | Rhodobacter phage RcThunderbird | Rhodobacter phage RcThunderbird Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43941 | 54.801 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| MW677525 | Rhodobacter phage RcSimone-Hastad | Rhodobacter phage RcSimone-Hastad Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63102 | 60.749 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677524 | Rhodobacter phage RcSalem | Rhodobacter phage RcSalem Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67718 | 60.034 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677523 | Rhodobacter phage RcRios | Rhodobacter phage RcRios Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68774 | 60.268 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677522 | Rhodobacter phage RcPutin | Rhodobacter phage RcPutin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67605 | 60.263 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677521 | Rhodobacter phage RcPescado | Rhodobacter phage RcPescado Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67494 | 60.42 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677520 | Rhodobacter phage RcOceanus | Rhodobacter phage RcOceanus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37609 | 64.163 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677519 | Rhodobacter phage RcMrWorf | Rhodobacter phage RcMrWorf Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67929 | 60.055 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677518 | Rhodobacter phage RcMcDreamy | Rhodobacter phage RcMcDreamy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68244 | 59.964 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677517 | Rhodobacter phage RcKemmy | Rhodobacter phage RcKemmy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41345 | 63.705 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677516 | Rhodobacter phage RcIroh | Rhodobacter phage RcIroh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68475 | 60.19 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677515 | Rhodobacter phage RcHotPocket | Rhodobacter phage RcHotPocket Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41765 | 64.111 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677514 | Rhodobacter phage RcHartney | Rhodobacter phage RcHartney Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43528 | 55.07 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| MW677513 | Rhodobacter phage RcGingersnap | Rhodobacter phage RcGingersnap Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68225 | 60.195 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677512 | Rhodobacter phage RcFrancesLouise | Rhodobacter phage RcFrancesLouise Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42073 | 64.039 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677511 | Rhodobacter phage RcDurkin | Rhodobacter phage RcDurkin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94639 | 57.789 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677510 | Rhodobacter phage RcDormio | Rhodobacter phage RcDormio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41640 | 64.083 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677509 | Rhodobacter phage RcBaka | Rhodobacter phage RcBaka Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41643 | 64.138 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MW677528 | Rhodobacter phage RcWaterboi | Rhodobacter phage RcWaterboi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38301 | 64.823 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| MZ399596 | Carnobacterium phage cd4 | Carnobacterium phage cd4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56713 | 38.718 | Unclassified | Unclassified | Siphoviridae | Carnobacterium |
| MZ398136 | Carnobacterium phage cd3 | Carnobacterium phage cd3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57171 | 38.904 | Unclassified | Unclassified | Siphoviridae | Carnobacterium |
| MZ398135 | Carnobacterium phage cd2 | Carnobacterium phage cd2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57220 | 38.955 | Unclassified | Unclassified | Siphoviridae | Carnobacterium |
| MZ089803 | Microvirus mar57 | Microvirus mar57 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6419 | 42.577 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089800 | Microvirus mar54 | Microvirus mar54 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6294 | 41.754 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089799 | Microvirus mar53 | Microvirus mar53 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6128 | 42.2 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089795 | Microvirus mar49 | Microvirus mar49 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6039 | 46.481 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089791 | Microvirus mar45 | Microvirus mar45 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5881 | 39.551 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089786 | Microvirus mar40 | Microvirus mar40 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5741 | 37.433 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089784 | Microvirus mar38 | Microvirus mar38 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5634 | 34.168 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089773 | Microvirus mar27 | Microvirus mar27 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5158 | 41.353 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089762 | Microvirus mar16 | Microvirus mar16 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4862 | 44.447 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089761 | Microvirus mar15 | Microvirus mar15 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4815 | 39.751 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089757 | Microvirus mar11 | Microvirus mar11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4725 | 38.624 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089754 | Microvirus mar8 | Microvirus mar8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 41.297 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ462995 | Stappia phage SI01 | Stappia phage SI01 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44247 | 58.298 | Unclassified | Unclassified | Autographiviridae | Stappia |
| MZ326168 | Salmonella phage NINP13076 | Salmonella phage NINP13076 unclassified Seunavirus Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161329 | 44.417 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| MZ209307 | Arthrobacter phage GoCrazy | Arthrobacter phage GoCrazy unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56809 | 45.433 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ209306 | Mycobacterium phage Murai | Mycobacterium phage Murai unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71538 | 65.346 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ209305 | Mycobacterium phage Trooper | Mycobacterium phage Trooper unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52519 | 63.013 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ209300 | Arthrobacter phage Kardesai | Arthrobacter phage Kardesai unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57866 | 44.871 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MZ209299 | Mycobacterium phage Latretium | Mycobacterium phage Latretium unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59282 | 61.187 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW879344 | Erwinia phage pEa_SNUABM_49 | Erwinia phage pEa_SNUABM_49 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 358049 | 34.368 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MW879343 | Erwinia phage pEa_SNUABM_44 | Erwinia phage pEa_SNUABM_44 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 356749 | 34.455 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MW879342 | Erwinia phage pEa_SNUABM_19 | Erwinia phage pEa_SNUABM_19 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 355277 | 34.455 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MW879341 | Erwinia phage pEa_SNUABM_54 | Erwinia phage pEa_SNUABM_54 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 272635 | 48.225 | Unclassified | Unclassified | Myoviridae | Erwinia |
| MW879340 | Erwinia phage pEa_SNUABM_48 | Erwinia phage pEa_SNUABM_48 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 294405 | 49.517 | Unclassified | Unclassified | Myoviridae | Erwinia |
| MW911615 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121570 | 37.124 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW911613 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121928 | 37.139 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW911614 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121671 | 37.14 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW911612 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121708 | 37.136 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MZ424864 | Escherichia phage vB_EcoP_ZX6 | Escherichia phage vB_EcoP_ZX6 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43510 | 53.68 | Drulisvirus | Slopekvirinae | Autographiviridae | Escherichia |
| MW176034 | Acinetobacter phage JeffCo | Acinetobacter phage JeffCo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43285 | 47.799 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MW176033 | Acinetobacter phage Effie | Acinetobacter phage Effie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53598 | 37.47 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MW176032 | Acinetobacter phage Barton | Acinetobacter phage Barton Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43041 | 47.72 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MZ152915 | Staphylococcus phage SeAlphi | Staphylococcus phage SeAlphi unclassified Andhravirus Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18292 | 29.909 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MZ152914 | Mycobacterium phage Maco7 | Mycobacterium phage Maco7 unclassified Mapvirus Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47820 | 58.749 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW822008 | Escherichia phage BF17 | Escherichia phage BF17 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 383493 | 35.445 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MW822007 | Escherichia phage BF15 | Escherichia phage BF15 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167801 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ089811 | Microvirus mar65 | Microvirus mar65 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7225 | 38.422 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089810 | Microvirus mar64 | Microvirus mar64 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6994 | 48.256 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089809 | Microvirus mar63 | Microvirus mar63 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6688 | 48.146 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089808 | Microvirus mar62 | Microvirus mar62 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6623 | 43.53 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089807 | Microvirus mar61 | Microvirus mar61 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6609 | 42.911 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089806 | Microvirus mar60 | Microvirus mar60 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6523 | 34.677 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089805 | Microvirus mar59 | Microvirus mar59 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6455 | 37.707 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089804 | Microvirus mar58 | Microvirus mar58 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6439 | 43.578 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089802 | Microvirus mar56 | Microvirus mar56 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6357 | 38.76 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089801 | Microvirus mar55 | Microvirus mar55 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6340 | 43.281 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089798 | Microvirus mar52 | Microvirus mar52 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6115 | 40.131 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089797 | Microvirus mar51 | Microvirus mar51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6091 | 41.75 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089796 | Microvirus mar50 | Microvirus mar50 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6058 | 41.499 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089794 | Microvirus mar48 | Microvirus mar48 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5939 | 40.377 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089793 | Microvirus mar47 | Microvirus mar47 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5933 | 40.384 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089792 | Microvirus mar46 | Microvirus mar46 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5909 | 40.244 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089790 | Microvirus mar44 | Microvirus mar44 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5873 | 42.891 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089789 | Microvirus mar43 | Microvirus mar43 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5863 | 38.496 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089788 | Microvirus mar42 | Microvirus mar42 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5785 | 42.662 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089787 | Microvirus mar41 | Microvirus mar41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5785 | 42.697 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089785 | Microvirus mar39 | Microvirus mar39 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 36.776 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089783 | Microvirus mar37 | Microvirus mar37 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5589 | 33.083 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089782 | Microvirus mar36 | Microvirus mar36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5497 | 45.097 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089781 | Microvirus mar35 | Microvirus mar35 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5453 | 43.554 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089780 | Microvirus mar34 | Microvirus mar34 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5368 | 37.891 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089779 | Microvirus mar33 | Microvirus mar33 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5328 | 49.869 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089778 | Microvirus mar32 | Microvirus mar32 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5259 | 41.282 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089777 | Microvirus mar31 | Microvirus mar31 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5235 | 42.865 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089776 | Microvirus mar30 | Microvirus mar30 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5204 | 37.394 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089775 | Microvirus mar29 | Microvirus mar29 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5167 | 51.693 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089774 | Microvirus mar28 | Microvirus mar28 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5162 | 44.944 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089772 | Microvirus mar26 | Microvirus mar26 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5144 | 36.645 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089771 | Microvirus mar25 | Microvirus mar25 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5129 | 40.359 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089770 | Microvirus mar24 | Microvirus mar24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5117 | 36.603 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089769 | Microvirus mar23 | Microvirus mar23 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5069 | 43.519 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089768 | Microvirus mar22 | Microvirus mar22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5030 | 37.913 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089767 | Microvirus mar21 | Microvirus mar21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4978 | 45.781 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089766 | Microvirus mar20 | Microvirus mar20 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4946 | 43.004 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089765 | Microvirus mar19 | Microvirus mar19 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4892 | 43.97 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089764 | Microvirus mar18 | Microvirus mar18 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4882 | 46.395 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089763 | Microvirus mar17 | Microvirus mar17 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4873 | 38.375 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089760 | Microvirus mar14 | Microvirus mar14 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4811 | 45.541 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089759 | Microvirus mar13 | Microvirus mar13 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4777 | 43.165 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089758 | Microvirus mar12 | Microvirus mar12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4765 | 47.513 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089756 | Microvirus mar10 | Microvirus mar10 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4713 | 34.5 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089755 | Microvirus mar9 | Microvirus mar9 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4696 | 37.585 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089753 | Microvirus mar7 | Microvirus mar7 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4646 | 45.609 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089752 | Microvirus mar6 | Microvirus mar6 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4626 | 45.893 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089751 | Microvirus mar5 | Microvirus mar5 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4605 | 35.331 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089750 | Microvirus mar4 | Microvirus mar4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4589 | 47.941 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089749 | Microvirus mar3 | Microvirus mar3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4586 | 42.826 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089748 | Microvirus mar2 | Microvirus mar2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4265 | 40.774 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ089747 | Microvirus mar1 | Microvirus mar1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4265 | 40.657 | Microvirus | Unclassified | Microviridae | Unspecified |
| MZ322028 | Gordonia phage Posh | Gordonia phage Posh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52295 | 66.11 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322023 | Mycobacterium Phage PetiteSangsue | Mycobacterium Phage PetiteSangsue unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 63.713 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322022 | Gordonia phage Float294 | Gordonia phage Float294 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67545 | 65.577 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322021 | Gordonia phage Cafasso | Gordonia phage Cafasso Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87157 | 67.576 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322016 | Mycobacterium phage SoSeph | Mycobacterium phage SoSeph unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61968 | 65.009 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ322010 | Gordonia phage Bonum | Gordonia phage Bonum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64476 | 65.452 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MZ322005 | Mycobacterium phage Phillis | Mycobacterium phage Phillis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51427 | 61.24 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW749009 | Bacillus phage vB_BspP_Dartukuta | Bacillus phage vB_BspP_Dartukuta Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60862 | 50.102 | Unclassified | Unclassified | Podoviridae | Bacillus |
| MW749006 | Bacillus phage vB_BspM_AgentSmith | Bacillus phage vB_BspM_AgentSmith Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 200233 | 37.033 | Unclassified | Unclassified | Myoviridae | Bacillus |
| MW749005 | Escherichia phage vB_EcoS_Sponge | Escherichia phage vB_EcoS_Sponge unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44887 | 54.526 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MW749003 | Bacillus phage vB_BspM_Internexus | Bacillus phage vB_BspM_Internexus unclassified Takahashivirus Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252262 | 27.666 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| MW749002 | Bacillus phage vB_BspH_Mawwa | Bacillus phage vB_BspH_Mawwa unclassified Siophivirus Siophivirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149014 | 38.985 | Siophivirus | Bastillevirinae | Herelleviridae | Bacillus |
| MW749001 | Escherichia phage vB_EcoM_Kelasse | Escherichia phage vB_EcoM_Kelasse unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167034 | 35.606 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MW749000 | Hafnia phage vB_HpaM_IsaacDaniel | Hafnia phage vB_HpaM_IsaacDaniel unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167265 | 35.624 | Tequatrovirus | Tevenvirinae | Myoviridae | Hafnia |
| MW748998 | Escherichia phage vB_EcoM_SophiaRose | Escherichia phage vB_EcoM_SophiaRose unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139943 | 43.468 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MZ444140 | Pseudomonas phage PA7 | Pseudomonas phage PA7 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 282378 | 36.818 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MZ042490 | Stenotrophomonas phage BUCT608 | Stenotrophomonas phage BUCT608 unclassified Menderavirus Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160122 | 54.15 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MW362867 | Salmonella phage SP2 SHa-2019 | Salmonella phage SP2 SHa-2019 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85702 | 38.987 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW311372 | Salmonella phage SEP1 | Salmonella phage SEP1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85703 | 38.986 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW321605 | Salmonella phage SP4 SHa-2019 | Salmonella phage SP4 SHa-2019 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85706 | 38.986 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW365948 | Microbacterium phage CrunchyBoi | Microbacterium phage CrunchyBoi unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53800 | 59.084 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW365947 | Microbacterium phage PineapplePluto | Microbacterium phage PineapplePluto unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53868 | 59.109 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW965453 | Ralstonia phage vB_RsoP_BMB50 | Ralstonia phage vB_RsoP_BMB50 Higashivirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43665 | 61.791 | Higashivirus | Okabevirinae | Autographiviridae | Ralstonia |
| MZ322024 | Mycobacterium phage JeTaime | Mycobacterium phage JeTaime unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75099 | 62.938 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ496294 | Vibrio phage ST2-2pr | Vibrio phage ST2-2pr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11617 | 44.237 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MZ496293 | Vibrio phage ST2-1pr | Vibrio phage ST2-1pr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33570 | 43.801 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MZ496292 | Pseudomonas phage PARCL1pr | Pseudomonas phage PARCL1pr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39565 | 62.965 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ171369 | Methanoculleus virus L4768 | Methanoculleus virus L4768 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37078 | 63.108 | Unclassified | Unclassified | Siphoviridae | Methanoculleus |
| MW366783 | Acinetobacter phage Pipo | Acinetobacter phage Pipo unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41956 | 39.16 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW366784 | Acinetobacter phage Paty | Acinetobacter phage Paty unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41264 | 39.419 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW822006 | Escherichia phage BF9 | Escherichia phage BF9 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44963 | 54.478 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| AB231700 | Microcystis virus Ma-LMM01 | Microcystis virus Ma-LMM01 Fukuivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162109 | 45.953 | Fukuivirus | Unclassified | Myoviridae | Microcystis |
| LC616079 | Stx2a-converting phage Stx2_KH16-043 | Stx2a-converting phage Stx2_KH16-043 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45156 | 52.491 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC616078 | Stx2a-converting phage Stx2_10E082 | Stx2a-converting phage Stx2_10E082 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65396 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616077 | Stx2a-converting phage Stx2_13518 | Stx2a-converting phage Stx2_13518 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65392 | 49.483 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616076 | Stx2a-converting phage Stx2_2935 | Stx2a-converting phage Stx2_2935 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65398 | 49.486 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616075 | Stx2a-converting phage Stx2_06E050 | Stx2a-converting phage Stx2_06E050 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65392 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616074 | Stx2a-converting phage Stx2_10902 | Stx2a-converting phage Stx2_10902 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65394 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616073 | Stx2a-converting phage Stx2_08E027 | Stx2a-converting phage Stx2_08E027 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616072 | Stx2a-converting phage Stx2_PV03-14 | Stx2a-converting phage Stx2_PV03-14 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616071 | Stx2a-converting phage Stx2_8048 | Stx2a-converting phage Stx2_8048 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616070 | Stx2a-converting phage Stx2_15436 | Stx2a-converting phage Stx2_15436 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616069 | Stx2a-converting phage Stx2_635 | Stx2a-converting phage Stx2_635 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65396 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616068 | Stx2a-converting phage Stx2_522 | Stx2a-converting phage Stx2_522 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65400 | 49.486 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616067 | Stx2a-converting phage Stx2_356 | Stx2a-converting phage Stx2_356 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65392 | 49.486 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616066 | Stx2a-converting phage Stx2_11106 | Stx2a-converting phage Stx2_11106 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66718 | 49.559 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616065 | Stx2a-converting phage Stx2_463 | Stx2a-converting phage Stx2_463 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66582 | 49.552 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616064 | Stx2a-converting phage Stx2_453 | Stx2a-converting phage Stx2_453 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66710 | 49.559 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616063 | Stx2a-converting phage Stx2_3418 | Stx2a-converting phage Stx2_3418 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65443 | 49.573 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616062 | Stx2a-converting phage Stx2_3896 | Stx2a-converting phage Stx2_3896 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66709 | 49.56 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616061 | Stx2a-converting phage Stx2_12874 | Stx2a-converting phage Stx2_12874 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66707 | 49.557 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616060 | Stx2a-converting phage Stx2_3417 | Stx2a-converting phage Stx2_3417 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68032 | 49.647 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616059 | Stx2a-converting phage Stx2_3105 | Stx2a-converting phage Stx2_3105 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65413 | 49.484 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616058 | Stx2a-converting phage Stx2_716 | Stx2a-converting phage Stx2_716 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65393 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616057 | Stx2a-converting phage Stx2_579 | Stx2a-converting phage Stx2_579 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616056 | Stx2a-converting phage Stx2_3772 | Stx2a-converting phage Stx2_3772 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65394 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616055 | Stx2a-converting phage Stx2_1603 | Stx2a-converting phage Stx2_1603 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67978 | 49.565 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616054 | Stx2a-converting phage Stx2_12E092 | Stx2a-converting phage Stx2_12E092 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67541 | 49.589 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616053 | Stx2a-converting phage Stx2_12849 | Stx2a-converting phage Stx2_12849 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.482 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616052 | Stx2a-converting phage Stx2_7982 | Stx2a-converting phage Stx2_7982 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65394 | 49.483 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616051 | Stx2a-converting phage Stx2_PV07-173 | Stx2a-converting phage Stx2_PV07-173 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66703 | 49.558 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616050 | Stx2a-converting phage Stx2_3350 | Stx2a-converting phage Stx2_3350 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66705 | 49.578 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616049 | Stx2a-converting phage Stx2_11Y11 | Stx2a-converting phage Stx2_11Y11 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66711 | 49.578 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616048 | Stx2a-converting phage Stx2_07Y06 | Stx2a-converting phage Stx2_07Y06 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66663 | 49.471 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616047 | Stx2a-converting phage Stx2_6804 | Stx2a-converting phage Stx2_6804 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65392 | 49.483 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616046 | Stx2a-converting phage Stx2_5122 | Stx2a-converting phage Stx2_5122 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66613 | 49.558 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616045 | Stx2a-converting phage Stx2_481 | Stx2a-converting phage Stx2_481 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66664 | 49.469 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616044 | Stx2a-converting phage Stx2_8243 | Stx2a-converting phage Stx2_8243 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65422 | 49.48 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616043 | Stx2a-converting phage Stx2_3725 | Stx2a-converting phage Stx2_3725 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65396 | 49.483 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616042 | Stx2a-converting phage Stx2_12E064 | Stx2a-converting phage Stx2_12E064 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65397 | 49.482 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616041 | Stx2a-converting phage Stx2_PV12-16 | Stx2a-converting phage Stx2_PV12-16 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65391 | 49.481 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616040 | Stx2a-converting phage Stx2_3993 | Stx2a-converting phage Stx2_3993 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66707 | 49.577 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616039 | Stx2a-converting phage Stx2_131033 | Stx2a-converting phage Stx2_131033 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66711 | 49.559 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616038 | Stx2a-converting phage Stx2_707 | Stx2a-converting phage Stx2_707 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67931 | 49.462 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616037 | Stx2a-converting phage Stx2_4151 | Stx2a-converting phage Stx2_4151 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65395 | 49.485 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616036 | Stx2a-converting phage Stx2_PV06-102 | Stx2a-converting phage Stx2_PV06-102 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66659 | 49.474 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616035 | Stx2a-converting phage Stx2_13E027 | Stx2a-converting phage Stx2_13E027 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67740 | 49.511 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616034 | Stx2a-converting phage Stx2_EH0337 | Stx2a-converting phage Stx2_EH0337 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66701 | 49.578 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616033 | Stx2a-converting phage Stx2_EH0787 | Stx2a-converting phage Stx2_EH0787 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66751 | 49.676 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616032 | Stx2a-converting phage Stx2_EH1965 | Stx2a-converting phage Stx2_EH1965 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65382 | 49.483 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| LC616031 | Stx2a-converting phage Stx2_EH0505 | Stx2a-converting phage Stx2_EH0505 Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65389 | 49.421 | Unclassified | Sepvirinae | Podoviridae | Unspecified |
| NC_048048 | Caulobacter phage CcrSC | Caulobacter phage CcrSC Caulobacter virus CcrSC Bertelyvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 317489 | 64.183 | Bertelyvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| MZ079855 | Klebsiella phage vB_Kpn_3 | Klebsiella phage vB_Kpn_3 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112003 | 41.361 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MW508890 | Phage PEA2-3 | Phage PEA2-3 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52770 | 45.992 | Rosemountvirus | Unclassified | Myoviridae | Unspecified |
| MT310859 | Mycobacterium phage MarkPhew | Mycobacterium phage MarkPhew unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62153 | 67.86 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN136198 | Escherichia phage vB_EcoM_4HA13 | Escherichia phage vB_EcoM_4HA13 Escherichia virus 4HA13 Sabourvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52401 | 42.824 | Sabourvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MZ308445 | Lactococcus phage CAP | Lactococcus phage CAP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38693 | 36.094 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MH588547 | Caulobacter phage CcrSC | Caulobacter phage CcrSC Caulobacter virus CcrSC Bertelyvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 317489 | 64.183 | Bertelyvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| MZ402608 | Rhodococcus phage WC1 | Rhodococcus phage WC1 unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46439 | 58.602 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MW790496 | Salmonella phage vB_SalP_ABTNLsp11242 | Salmonella phage vB_SalP_ABTNLsp11242 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40644 | 48.177 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW790495 | Salmonella phage vB_SalS_ABTNLsp11241 | Salmonella phage vB_SalS_ABTNLsp11241 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43556 | 49.649 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN935203 | Proteus phage VTCCBPA139 | Proteus phage VTCCBPA139 unclassified Novosibvirus Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97330 | 38.414 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MN564818 | Pseudomonas phage DRL-P1 | Pseudomonas phage DRL-P1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66243 | 54.934 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_049466 | Escherichia phage vB_EcoM_4HA13 | Escherichia phage vB_EcoM_4HA13 Escherichia virus 4HA13 Sabourvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52401 | 42.824 | Sabourvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MZ291552 | Escherichia phage vB_EcoM-UFV09 | Escherichia phage vB_EcoM-UFV09 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173384 | 35.427 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT580897 | Klebsiella phage KpnM6E1 | Klebsiella phage KpnM6E1 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167920 | 39.581 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| OU342755 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 49924 | 51.206 | Unclassified | Unclassified | Unclassified | Klebsiella |
| OU342754 | Klebsiella pneumoniae phage | Klebsiella pneumoniae phage unclassified bacterial viruses Viruses | 40697 | 53.191 | Unclassified | Unclassified | Unclassified | Klebsiella |
| MZ343158 | Mycobacterium phage RitSun | Mycobacterium phage RitSun unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55975 | 60.815 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ420230 | Yersinia phage HQ103 | Yersinia phage HQ103 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35113 | 51.468 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| MW788656 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29016 | 34.529 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MW788655 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11968 | 33.122 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MT596500 | Staphylococcus phage vB_SepP_BE03 | Staphylococcus phage vB_SepP_BE03 unclassified Andhravirus Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18271 | 29.966 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MN091625 | Staphylococcus phage PALS_1 | Staphylococcus phage PALS_1 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138776 | 30.105 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| KY290975 | Escherichia phage YUEEL01 | Escherichia phage YUEEL01 Escherichia virus YUEEL01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168266 | 35.36 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ150789 | Microbacterium phage Footloose | Microbacterium phage Footloose Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39738 | 61.656 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW712719 | Arthrobacter phage Persistence | Arthrobacter phage Persistence Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50433 | 62.15 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MZ417521 | Escherichia phage 132 | Escherichia phage 132 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166922 | 35.485 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ375754 | Burkholderia phage PK23 | Burkholderia phage PK23 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35343 | 65.122 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| MZ150790 | Microbacterium phage Badulia | Microbacterium phage Badulia unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 15460 | 54.631 | Unclassified | Unclassified | Tectiviridae | Microbacterium |
| MZ150787 | Microbacterium phage QuadZero | Microbacterium phage QuadZero unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40140 | 63.63 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ150785 | Gordonia phage Jalammah | Gordonia phage Jalammah Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51552 | 67.057 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MZ150782 | Microbacterium phage PermaG | Microbacterium phage PermaG unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42283 | 65.48 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MW862991 | Microbacterium phage StagePhright | Microbacterium phage StagePhright unclassified Akonivirus Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54442 | 59.702 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW862990 | Mycobacterium phage Padfoot | Mycobacterium phage Padfoot unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59905 | 66.555 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW862986 | Gordonia phage Phishy | Gordonia phage Phishy unclassified Nyceiraevirus Nyceiraevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43061 | 67.495 | Nyceiraevirus | Unclassified | Siphoviridae | Gordonia |
| MW862985 | Mycobacterium phage Jarvi | Mycobacterium phage Jarvi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.549 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW862984 | Mycobacterium phage MeaningOfLife | Mycobacterium phage MeaningOfLife unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60432 | 66.605 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW862979 | Mycobacterium phage Peel | Mycobacterium phage Peel unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59711 | 66.546 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712736 | Streptomyces phage TaidaOne | Streptomyces phage TaidaOne unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56183 | 59.488 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MW712734 | Mycobacterium phage Durfee | Mycobacterium phage Durfee unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59905 | 66.557 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712733 | Mycobacterium phage Blizzard | Mycobacterium phage Blizzard unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59905 | 66.557 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712730 | Mycobacterium phage BaghaKamala | Mycobacterium phage BaghaKamala unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59132 | 66.614 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712723 | Mycobacterium phage Asayake | Mycobacterium phage Asayake unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59905 | 66.555 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712722 | Mycobacterium phage Twitch | Mycobacterium phage Twitch unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59711 | 66.551 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712718 | Mycobacterium phage Scout | Mycobacterium phage Scout unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49605 | 63.921 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW712717 | Gordonia phage BoyNamedSue | Gordonia phage BoyNamedSue unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52698 | 66.786 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MZ182247 | Vibrio phage vB_VpP_DE18 | Vibrio phage vB_VpP_DE18 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43464 | 49.561 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MT596506 | Staphylococcus phage vB_SepM_BE09 | Staphylococcus phage vB_SepM_BE09 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140668 | 28.018 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596505 | Staphylococcus phage vB_SepM_BE08 | Staphylococcus phage vB_SepM_BE08 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140659 | 28.019 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596504 | Staphylococcus phage vB_SepM_BE07 | Staphylococcus phage vB_SepM_BE07 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140661 | 27.991 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596503 | Staphylococcus phage vB_SepM_BE06 | Staphylococcus phage vB_SepM_BE06 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140659 | 27.992 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596502 | Staphylococcus phage vB_SepM_BE05 | Staphylococcus phage vB_SepM_BE05 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140271 | 28.044 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596501 | Staphylococcus phage vB_SepM_BE04 | Staphylococcus phage vB_SepM_BE04 unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142331 | 27.934 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT596499 | Staphylococcus phage vB_SepS_BE02 | Staphylococcus phage vB_SepS_BE02 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95233 | 29.354 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MT596498 | Staphylococcus phage vB_SepS_BE01 | Staphylococcus phage vB_SepS_BE01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42718 | 34.835 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT596497 | Staphylococcus phage 456 | Staphylococcus phage 456 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43393 | 34.72 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MZ420150 | Enterococcus phage vB_EfaM_LG1 | Enterococcus phage vB_EfaM_LG1 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150025 | 35.885 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MZ359091 | Staphylococcus phage JPL-50 | Staphylococcus phage JPL-50 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16927 | 29.273 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MZ359670 | Klebsiella phage 066044 | Klebsiella phage 066044 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38704 | 50.922 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MZ342906 | Escherichia phage vB_EcoP_SU7 | Escherichia phage vB_EcoP_SU7 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76626 | 42.054 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MZ336020 | Aeromonas phage pAh6.2TG | Aeromonas phage pAh6.2TG unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51780 | 52.484 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| MZ208805 | Klebsiella phage SRD2021 | Klebsiella phage SRD2021 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45212 | 53.831 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MW630116 | Escherichia phage vB_EcoM-ZQ3 | Escherichia phage vB_EcoM-ZQ3 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170042 | 37.606 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MW630115 | Escherichia phage vB_EcoP-ZQ2 | Escherichia phage vB_EcoP-ZQ2 unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72231 | 42.849 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MG720309 | Vibrio phage Ares1 | Vibrio phage Ares1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80500 | 45.075 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG720308 | Vibrio phage Aphrodite1 | Vibrio phage Aphrodite1 Vibrio virus Aphrodite1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237722 | 43.36 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| MZ318363 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36942 | 48.473 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ318362 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36942 | 48.473 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ318361 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36942 | 48.476 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MZ189262 | Escherichia phage W143 | Escherichia phage W143 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166244 | 35.458 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ189261 | Escherichia phage S143_2 | Escherichia phage S143_2 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168769 | 37.68 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MW057920 | Clostridium phage CPQ1 | Clostridium phage CPQ1 unclassified Brucesealvirus Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18535 | 34.61 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| MZ274225 | Salmonella phage fmb-p1 | Salmonella phage fmb-p1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40731 | 49.832 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW590329 | Klebsiella phage Kp_Pokalde_001 | Klebsiella phage Kp_Pokalde_001 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44535 | 54.088 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MW831865 | Stenotrophomonas phage BUCT598 | Stenotrophomonas phage BUCT598 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43581 | 59.987 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| MZ221764 | Klebsiella phage ABTNL-2 | Klebsiella phage ABTNL-2 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49159 | 51.034 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MZ223858 | Enterobacter phage EV136_1 | Enterobacter phage EV136_1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39701 | 52.069 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| MW677132 | Enterococcus phage EFC1 | Enterococcus phage EFC1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56099 | 39.96 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MT732475 | Winogradskyella phage Peternella_1 | Winogradskyella phage Peternella_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39649 | 35.391 | Unclassified | Unclassified | Myoviridae | Winogradskyella |
| MT732474 | Tenacibaculum phage Gundel_1 | Tenacibaculum phage Gundel_1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78833 | 30.411 | Unclassified | Unclassified | Podoviridae | Tenacibaculum |
| MT732473 | Polaribacter phage Leef_1 | Polaribacter phage Leef_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37547 | 29.664 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732472 | Polaribacter phage Freya_10 | Polaribacter phage Freya_10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48920 | 28.688 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732471 | Polaribacter phage Freya_9 | Polaribacter phage Freya_9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48613 | 28.673 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732470 | Polaribacter phage Freya_8 | Polaribacter phage Freya_8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48018 | 28.745 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732466 | Polaribacter phage Freya_4 | Polaribacter phage Freya_4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44820 | 28.951 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732465 | Polaribacter phage Freya_3 | Polaribacter phage Freya_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44820 | 28.949 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732464 | Polaribacter phage Freya_2 | Polaribacter phage Freya_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44820 | 28.951 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732463 | Polaribacter phage Freya_1 | Polaribacter phage Freya_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43978 | 28.876 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732462 | Polaribacter phage Danklef_5 | Polaribacter phage Danklef_5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48177 | 28.96 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732461 | Polaribacter phage Danklef_4 | Polaribacter phage Danklef_4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47426 | 28.908 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732460 | Polaribacter phage Danklef_3 | Polaribacter phage Danklef_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47396 | 28.922 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732459 | Polaribacter phage Danklef_2 | Polaribacter phage Danklef_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47302 | 28.914 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732458 | Polaribacter phage Danklef_1 | Polaribacter phage Danklef_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47186 | 28.818 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| MT732457 | Olleya phage Harreka_1 | Olleya phage Harreka_1 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43175 | 32.002 | Unclassified | Unclassified | Unclassified | Olleya |
| MT732449 | Cellulophaga phage Omtje_5 | Cellulophaga phage Omtje_5 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6559 | 31.179 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| MT732448 | Cellulophaga phage Omtje_4 | Cellulophaga phage Omtje_4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6558 | 31.183 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| MT732447 | Cellulophaga phage Omtje_3 | Cellulophaga phage Omtje_3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6558 | 31.168 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| MT732446 | Cellulophaga phage Omtje_2 | Cellulophaga phage Omtje_2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6558 | 31.183 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| MT732445 | Cellulophaga phage Omtje_1 | Cellulophaga phage Omtje_1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6558 | 31.153 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| MT732442 | Cellulophaga phage Ingeline_8 | Cellulophaga phage Ingeline_8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42628 | 32.239 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732441 | Cellulophaga phage Ingeline_7 | Cellulophaga phage Ingeline_7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42625 | 32.242 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732440 | Cellulophaga phage Ingeline_6 | Cellulophaga phage Ingeline_6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42625 | 32.242 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732439 | Cellulophaga phage Ingeline_5 | Cellulophaga phage Ingeline_5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42625 | 32.239 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732438 | Cellulophaga phage Ingeline_4 | Cellulophaga phage Ingeline_4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42624 | 32.24 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732437 | Cellulophaga phage Ingeline_3 | Cellulophaga phage Ingeline_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42624 | 32.24 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732436 | Cellulophaga phage Ingeline_2 | Cellulophaga phage Ingeline_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42624 | 32.24 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732435 | Cellulophaga phage Ingeline_1 | Cellulophaga phage Ingeline_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42624 | 32.24 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732434 | Cellulophaga phage Calle_3 | Cellulophaga phage Calle_3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72980 | 38.112 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| MT732433 | Cellulophaga phage Calle_2 | Cellulophaga phage Calle_2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72979 | 38.107 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| MT732432 | Cellulophaga phage Calle_1 | Cellulophaga phage Calle_1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72979 | 38.108 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| MT732455 | Maribacter phage Molly_5 | Maribacter phage Molly_5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124898 | 36.19 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732454 | Maribacter phage Molly_4 | Maribacter phage Molly_4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125038 | 36.191 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732453 | Maribacter phage Molly_3 | Maribacter phage Molly_3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124695 | 36.23 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732452 | Maribacter phage Molly_2 | Maribacter phage Molly_2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124695 | 36.228 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732451 | Maribacter phage Molly_1 | Maribacter phage Molly_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124695 | 36.229 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732450 | Maribacter phage Colly_1 | Maribacter phage Colly_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124169 | 36.296 | Unclassified | Unclassified | Myoviridae | Maribacter |
| MT732444 | Cellulophaga phage Nekkels_2 | Cellulophaga phage Nekkels_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54332 | 31.451 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MT732443 | Cellulophaga phage Nekkels_1 | Cellulophaga phage Nekkels_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53385 | 31.516 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| MW009675 | Vibrio phage vB_VpP_BT-1011 | Vibrio phage vB_VpP_BT-1011 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40065 | 42.998 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MZ384014 | Bacillus phage DLn1 | Bacillus phage DLn1 unclassified Hemphillvirus Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25379 | 30.659 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| MW073100 | Microbacterium phage vB_MoxS-R1 | Microbacterium phage vB_MoxS-R1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42563 | 63.652 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW192778 | Staphylococcus phage B_UFSM5 | Staphylococcus phage B_UFSM5 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41732 | 33.976 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MW147366 | Staphylococcus phage B_UFSM4 | Staphylococcus phage B_UFSM4 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41290 | 33.972 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MW929175 | Pseudomonas phage K4 | Pseudomonas phage K4 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50358 | 44.632 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| MZ355727 | Microbacterium phage WilliamStrong | Microbacterium phage WilliamStrong unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40613 | 63.736 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MZ355726 | Mycobacterium phage Gyzlar | Mycobacterium phage Gyzlar unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49064 | 63.925 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ355725 | Mycobacterium Phage Sparxx | Mycobacterium Phage Sparxx unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.911 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ355723 | Mycobacterium Phage PeaceMeal1 | Mycobacterium Phage PeaceMeal1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49899 | 64.498 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MZ028625 | Gordonia phage DekHockey33 | Gordonia phage DekHockey33 unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52969 | 61.991 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MW269952 | Escherichia phage P479 | Escherichia phage P479 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172033 | 40.262 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MZ130500 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98182 | 32.57 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130499 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97706 | 31.399 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130498 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104752 | 33.211 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130497 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104085 | 36.217 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130496 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102100 | 33.149 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130495 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101130 | 32.952 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130494 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100255 | 32.526 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130493 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100650 | 33.317 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130492 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96892 | 31.394 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130491 | UAG-readthrough crAss clade sp. | UAG-readthrough crAss clade sp. UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98573 | 33.068 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130490 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92132 | 31.852 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130489 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95815 | 32.715 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130488 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91714 | 24.939 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130487 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94533 | 30.76 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130486 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92361 | 26.658 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130485 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104564 | 36.131 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130484 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97536 | 32.026 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130483 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93564 | 35.78 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130482 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99704 | 33.27 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130481 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98001 | 29.025 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130480 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93391 | 31.034 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130479 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96466 | 31.987 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130478 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100314 | 34.416 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130477 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100971 | 32.226 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130476 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92241 | 33.206 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130475 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99984 | 32.68 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MZ130474 | CrAss-like virus sp. | CrAss-like virus sp. crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92367 | 31.551 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW145539 | Streptomyces phage SscP1EGY | Streptomyces phage SscP1EGY unclassified Bingvirus Bingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51751 | 57.914 | Bingvirus | Unclassified | Siphoviridae | Streptomyces |
| MW514247 | Roseobacter phage CRP-13 | Roseobacter phage CRP-13 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55022 | 44.231 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MW514246 | Roseobacter phage CRP-9 | Roseobacter phage CRP-9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56157 | 41.612 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MT735630 | Vibrio phage vB_VpS_PG28 | Vibrio phage vB_VpS_PG28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82712 | 48.075 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MZ355721 | Mycobacterium phage Stinson | Mycobacterium phage Stinson unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59918 | 66.564 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW900438 | Vibrio phage vB_ValP_VA-RY-3 | Vibrio phage vB_ValP_VA-RY-3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40271 | 43.989 | Unclassified | Unclassified | Podoviridae | Vibrio |
| AB609718 | Enterococcus phage phiEF24C-P2 | Enterococcus phage phiEF24C-P2 Enterococcus virus EF24C Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142072 | 35.737 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MT345684 | Alteromonas phage vB_AcoS-R7M | Alteromonas phage vB_AcoS-R7M Alteromonas virus R7M Amoyvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56163 | 45.61 | Amoyvirus | Queuovirinae | Siphoviridae | Alteromonas |
| MZ089979 | Bacillus phage Sole | Bacillus phage Sole unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14444 | 39.58 | Unclassified | Unclassified | Tectiviridae | Bacillus |
| MZ089978 | Bacillus phage Sato | Bacillus phage Sato unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14852 | 39.16 | Unclassified | Unclassified | Tectiviridae | Bacillus |
| NC_055919 | Klebsiella phage vB_KpnP_P184 | Klebsiella phage vB_KpnP_P184 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76617 | 44.34 | Unclassified | Unclassified | Schitoviridae | Klebsiella |
| NC_055915 | Flavobacterium phage vB_FspM_immuto_2-6A | Flavobacterium phage vB_FspM_immuto_2-6A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160410 | 34.52 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| NC_055913 | Vibrio phage Gary | Vibrio phage Gary unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128602 | 40.195 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055910 | Providencia phage PSTCR5 | Providencia phage PSTCR5 Priunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109434 | 39.999 | Priunavirus | Unclassified | Siphoviridae | Providencia |
| NC_055907 | Enterococcus phage 9183 | Enterococcus phage 9183 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86301 | 33.003 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_055860 | Mycobacterium phage Heath | Mycobacterium phage Heath unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69258 | 68.727 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_055856 | Gordonia Phage Barsten | Gordonia Phage Barsten unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59501 | 67.666 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055851 | Vibrio phage Athena | Vibrio phage Athena unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127019 | 40.097 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055850 | Vibrio phage Quinn | Vibrio phage Quinn unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127684 | 40.094 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055849 | Vibrio phage Cody | Vibrio phage Cody unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126683 | 40.179 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055848 | Vibrio phage AG74 | Vibrio phage AG74 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126504 | 40.172 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055847 | Vibrio phage River4 | Vibrio phage River4 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128425 | 40.233 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055841 | Agrobacterium phage OLIVR5 | Agrobacterium phage OLIVR5 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151073 | 46.936 | Unclassified | Unclassified | Ackermannviridae | Agrobacterium |
| NC_055840 | Agrobacterium phage OLIVR1 | Agrobacterium phage OLIVR1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75576 | 45.389 | Unclassified | Unclassified | Schitoviridae | Agrobacterium |
| NC_055839 | Xanthomonas phage FoX4 | Xanthomonas phage FoX4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60418 | 61.758 | Unclassified | Unclassified | Siphoviridae | Xanthomonas |
| NC_055837 | Xanthomonas phage FoX3 | Xanthomonas phage FoX3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44153 | 59.346 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| NC_055836 | Xanthomonas phage FoX2 | Xanthomonas phage FoX2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44086 | 59.475 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| NC_055834 | Enterococus phage vipetofem | Enterococus phage vipetofem Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85371 | 30.291 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_055833 | Enterococcus phage nattely | Enterococcus phage nattely Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85669 | 30.205 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_055830 | Vibrio phage Chester | Vibrio phage Chester unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126550 | 40.237 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055829 | Vibrio phage Bennett | Vibrio phage Bennett unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126552 | 40.098 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055826 | Bacteriophage DSS3_VP1 | Bacteriophage DSS3_VP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75087 | 47.498 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_055824 | Vibrio phage phi50-12 | Vibrio phage phi50-12 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68059 | 38.452 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| NC_055823 | Acinetobacter phage VB_ApiP_XC38 | Acinetobacter phage VB_ApiP_XC38 unclassified Presleyvirus Presleyvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79328 | 39.579 | Presleyvirus | Unclassified | Schitoviridae | Acinetobacter |
| NC_055821 | Gordonia phage NadineRae | Gordonia phage NadineRae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64714 | 66.057 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055820 | Gordonia phage Sukkupi | Gordonia phage Sukkupi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66184 | 66.194 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055819 | Gordonia phage Keitabear | Gordonia phage Keitabear unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59759 | 67.299 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055818 | Gordonia phage Jabberwocky | Gordonia phage Jabberwocky unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59449 | 67.592 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055808 | Gordonia terrae phage McKinley | Gordonia terrae phage McKinley unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59022 | 67.605 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055807 | Corynebacterium phage Stickynote | Corynebacterium phage Stickynote unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67086 | 52.176 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_055806 | Gordonia Phage Lollipop1437 | Gordonia Phage Lollipop1437 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67311 | 65.731 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055801 | Vibrio phage Pontus | Vibrio phage Pontus unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128289 | 40.233 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055800 | Vibrio phage Brizo | Vibrio phage Brizo unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120106 | 40.128 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055794 | Vibrio phage Achelous | Vibrio phage Achelous unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121089 | 40.036 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_055789 | Gordonia phage IDyn | Gordonia phage IDyn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65218 | 66.173 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055788 | Gordonia phage NosilaM | Gordonia phage NosilaM Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67662 | 65.579 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055784 | Vibrio virus vB_VspP_SBP1 | Vibrio virus vB_VspP_SBP1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78071 | 44.645 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| NC_055779 | Gordonia phage Tangent | Gordonia phage Tangent unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58281 | 67.18 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055778 | Gordonia phage Stultus | Gordonia phage Stultus unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57916 | 67.738 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055776 | Gordonia phage Geodirt | Gordonia phage Geodirt unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59495 | 67.656 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055775 | Gordonia phage Fosterous | Gordonia phage Fosterous unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59136 | 67.225 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055774 | Gordonia phage Bibwit | Gordonia phage Bibwit unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57880 | 67.457 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055773 | Gordonia phage Ailee | Gordonia phage Ailee unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58474 | 67.505 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055772 | Corynebacterium phage Kimchi1738 | Corynebacterium phage Kimchi1738 unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66915 | 52.257 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_055771 | Gordonia phage Emianna | Gordonia phage Emianna Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68190 | 65.801 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055770 | Gordonia phage Buggaboo | Gordonia phage Buggaboo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68626 | 65.627 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055769 | Gordonia phage Turuncu | Gordonia phage Turuncu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66808 | 65.383 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055767 | Gordonia phage KidneyBean | Gordonia phage KidneyBean Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67802 | 65.824 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055766 | Gordonia phage Foxboro | Gordonia phage Foxboro Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67773 | 65.842 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055764 | Gordonia phage Skysand | Gordonia phage Skysand Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67359 | 65.527 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055763 | Gordonia phage Marietta | Gordonia phage Marietta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64370 | 66.13 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055759 | Mycobacterium phage Ryadel | Mycobacterium phage Ryadel unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72658 | 65.207 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_055758 | Gordonia phage Pleakley | Gordonia phage Pleakley Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64105 | 65.218 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055755 | Gordonia phage NatB6 | Gordonia phage NatB6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67081 | 65.749 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055753 | Gordonia phage Rofo | Gordonia phage Rofo unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58767 | 67.291 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_055750 | Mycobacterium phage KingTut | Mycobacterium phage KingTut unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64827 | 66.462 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_055747 | Aeromonas phage 60AhydR15PP | Aeromonas phage 60AhydR15PP unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165795 | 41.233 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_055746 | Aeromonas phage 50AhydR13PP | Aeromonas phage 50AhydR13PP unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164983 | 41.074 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_055738 | Gordonia phage Flapper | Gordonia phage Flapper Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67527 | 65.301 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055736 | Vibrio phage 1.245.O._10N.261.54.C7 | Vibrio phage 1.245.O._10N.261.54.C7 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71702 | 43.12 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| NC_055735 | Vibrio phage 1.238.A._10N.261.52.F10 | Vibrio phage 1.238.A._10N.261.52.F10 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70494 | 43.188 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| NC_055733 | Aeromonas phage Ah1 | Aeromonas phage Ah1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221116 | 42.05 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| NC_055732 | Gordonia phage Patio | Gordonia phage Patio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66251 | 65.557 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055731 | Gordonia phage Kabluna | Gordonia phage Kabluna Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65869 | 65.556 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_055717 | Vibrio phage pVco-5 | Vibrio phage pVco-5 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74325 | 38.716 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| NC_055714 | Mycobacterium phage Fortunato | Mycobacterium phage Fortunato unclassified Coopervirus Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70679 | 68.958 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_055712 | Escherichia phage phT4A | Escherichia phage phT4A unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171598 | 41.451 | Slopekvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_055711 | Pseudomonas phage PaMx41 | Pseudomonas phage PaMx41 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43490 | 45.215 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| NC_055052 | Ralstonia virus RSIBR1 | Ralstonia virus RSIBR1 unclassified Habenivirus Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6945 | 61.253 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| NC_055051 | Vibrio phage ND1-fs1 | Vibrio phage ND1-fs1 unclassified Fibrovirus Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6856 | 43.436 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| MZ020222 | Vibrio phage BUCT233 | Vibrio phage BUCT233 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43245 | 49.275 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| NC_055921 | Salmonella phage vB_SalP_TR2 | Salmonella phage vB_SalP_TR2 unclassified Ithacavirus Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70823 | 40.646 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| NC_055917 | Bacillus phage WhyPhy | Bacillus phage WhyPhy unclassified Bundooravirus Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18642 | 34.948 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| NC_055904 | crAssphage cr273_1 | crAssphage cr273_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97886 | 36.266 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055903 | crAssphage cr272_1 | crAssphage cr272_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98667 | 36.183 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055902 | crAssphage cr131_1 | crAssphage cr131_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96717 | 35.294 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055901 | crAssphage cr130_1 | crAssphage cr130_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94826 | 31.772 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055899 | crAssphage cr124_1 | crAssphage cr124_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101532 | 35.803 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055896 | crAssphage cr113_1 | crAssphage cr113_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98557 | 33.037 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055895 | crAssphage cr7_1 | crAssphage cr7_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103832 | 35.691 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055892 | crAssphage cr109_1 | crAssphage cr109_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99168 | 33.051 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055891 | crAssphage cr60_1 | crAssphage cr60_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100198 | 33.09 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055889 | crAssphage cr53_1 | crAssphage cr53_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101153 | 32.971 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055887 | crAssphage cr271_1 | crAssphage cr271_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92815 | 30.545 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055886 | crAssphage cr127_1 | crAssphage cr127_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83412 | 25.922 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055885 | crAssphage cr128_1 | crAssphage cr128_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95150 | 33.197 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055884 | crAssphage cr126_1 | crAssphage cr126_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99212 | 34.776 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055883 | crAssphage cr85_1 | crAssphage cr85_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98164 | 37.722 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055882 | crAssphage cr116_1 | crAssphage cr116_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101082 | 32.585 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055881 | crAssphage cr112_1 | crAssphage cr112_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102301 | 34.681 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055880 | crAssphage cr111_1 | crAssphage cr111_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102434 | 39.544 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055879 | crAssphage cr110_1 | crAssphage cr110_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89781 | 30.753 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055878 | crAssphage cr108_1 | crAssphage cr108_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101437 | 37.271 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055876 | crAssphage cr11_1 | crAssphage cr11_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93559 | 30.001 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055875 | crAssphage cr10_1 | crAssphage cr10_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99985 | 33.532 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055874 | crAssphage cr3_1 | crAssphage cr3_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91101 | 30.729 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055873 | crAssphage cr1_1 | crAssphage cr1_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99882 | 32.195 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055872 | crAssphage cr118_1 | crAssphage cr118_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92600 | 28.706 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055871 | crAssphage cr106_1 | crAssphage cr106_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101662 | 33.424 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055870 | crAssphage cr107_1 | crAssphage cr107_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93787 | 31.997 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055869 | crAssphage cr55_1 | crAssphage cr55_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98340 | 31.9 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055868 | crAssphage cr50_1 | crAssphage cr50_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98826 | 32.58 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_055866 | Enterococcus phage AE4_17 | Enterococcus phage AE4_17 unclassified Copernicusvirus Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18477 | 32.879 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_055855 | Flavobacterium phage vB_FspP_elemoD_13-5B | Flavobacterium phage vB_FspP_elemoD_13-5B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60393 | 31.651 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| NC_055854 | Flavobacterium phage vB_FspP_elemoE_6-9C | Flavobacterium phage vB_FspP_elemoE_6-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59979 | 31.953 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| NC_055853 | Flavobacterium phage vB_FspP_elemoF_6-3D | Flavobacterium phage vB_FspP_elemoF_6-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59846 | 31.899 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| NC_055852 | Flavobacterium phage vB_FspP_elemoA_7-9A | Flavobacterium phage vB_FspP_elemoA_7-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61691 | 31.64 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| NC_055832 | Bacteroides phage DAC15 | Bacteroides phage DAC15 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99494 | 37.042 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| NC_055828 | Bacteroides phage crAss002 | Bacteroides phage crAss002 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93030 | 31.929 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| NC_055059 | Streptomyces phage Forthebois | Streptomyces phage Forthebois Streptomyces virus Forthebois Deltatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 18251 | 53.586 | Deltatectivirus | Unclassified | Tectiviridae | Streptomyces |
| NC_055035 | Fusobacterium phage Fnu1 | Fusobacterium phage Fnu1 Fusobacterium virus FNU1 Latrobevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130914 | 24.95 | Latrobevirus | Unclassified | Siphoviridae | Fusobacterium |
| NC_055027 | Lactococcus phage CHPC971 | Lactococcus phage CHPC971 Lactococcus virus CHPC971 Fremauxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54381 | 34.062 | Fremauxvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_054963 | Ralstonia phage Gervaise | Ralstonia phage Gervaise Ralstonia virus Gervaise Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61164 | 64.721 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| NC_054956 | Ralstonia phage Cimandef | Ralstonia phage Cimandef Ralstonia virus Cimandef Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55171 | 64.822 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| NC_054898 | Vibrio virus 2019VC1 | Vibrio virus 2019VC1 Jhansiroadvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47778 | 44.947 | Jhansiroadvirus | Unclassified | Drexlerviridae | Vibrio |
| NC_054897 | Citrobacter virus HCF1 | Citrobacter virus HCF1 Hicfunavirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45806 | 44.509 | Hicfunavirus | Unclassified | Drexlerviridae | Citrobacter |
| NC_055706 | Azobacteroides phage ProJPt-Bp1 | Azobacteroides phage ProJPt-Bp1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99517 | 42.309 | Unclassified | Unclassified | Podoviridae | Azobacteroides |
| NC_055061 | Rhodococcus phage Toil | Rhodococcus phage Toil Rhodococcus virus Toil Epsilontectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 17253 | 54.472 | Epsilontectivirus | Unclassified | Tectiviridae | Rhodococcus |
| NC_055060 | Streptomyces phage WheeHeim | Streptomyces phage WheeHeim Streptomyces virus WheeHeim Deltatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 18266 | 54.621 | Deltatectivirus | Unclassified | Tectiviridae | Streptomyces |
| NC_055055 | Xanthomonas phage Xf409 | Xanthomonas phage Xf409 Xanthomonas virus Xf109 Xylivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8280 | 59.746 | Xylivirus | Unclassified | Inoviridae | Xanthomonas |
| NC_055054 | Ralstonia phage RS611 | Ralstonia phage RS611 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6386 | 62.12 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| NC_055053 | Ralstonia phage RSS-TH1 | Ralstonia phage RSS-TH1 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7273 | 62.12 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| NC_055050 | Lactococcus phage PLgW-1 | Lactococcus phage PLgW-1 Lactococcus virus PLgW1 Uwajimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30189 | 37.322 | Uwajimavirus | Unclassified | Siphoviridae | Lactococcus |
| NC_055034 | Arthrobacter phage Wheelbite | Arthrobacter phage Wheelbite Arthrobacter virus Wheelbite Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59879 | 64.766 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_055033 | Arthrobacter phage Salgado | Arthrobacter phage Salgado Arthrobacter virus Salgado Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59807 | 64.635 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_055032 | Arthrobacter phage LiSara | Arthrobacter phage LiSara Arthrobacter virus LiSara Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60137 | 64.745 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054966 | Klebsiella phage SopranoGao | Klebsiella phage SopranoGao Klebsiella virus SopranoGao Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61644 | 57.266 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| NC_054964 | Ralstonia phage GP4 | Ralstonia phage GP4 Ralstonia virus GP4 Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61129 | 64.014 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| NC_054952 | Stenotrophomonas phage IME-SM1 | Stenotrophomonas phage IME-SM1 Stenotrophomonas virus IMESM1 Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159514 | 54.142 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MW043865 | Alteromonas phage vB_AmeP_R8W | Alteromonas phage vB_AmeP_R8W Foturvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48825 | 40.551 | Foturvirus | Unclassified | Autographiviridae | Alteromonas |
| MW748992 | Vibrio phage vB_ValP_IME234 | Vibrio phage vB_ValP_IME234 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50337 | 41.675 | Unclassified | Unclassified | Podoviridae | Vibrio |
| NC_047853 | Enterococcus phage phiEF24C-P2 | Enterococcus phage phiEF24C-P2 Enterococcus virus EF24C Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142072 | 35.737 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MZ043613 | Acinetobacter phage TaPaz | Acinetobacter phage TaPaz Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93703 | 32.639 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MZ182246 | Enterococcus phage vB_EfaS_785CS | Enterococcus phage vB_EfaS_785CS unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40946 | 34.785 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MW345254 | Cronobacter phage vB_CtuP_B1 | Cronobacter phage vB_CtuP_B1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39460 | 51.488 | Kayfunavirus | Studiervirinae | Autographiviridae | Cronobacter |
| MW296877 | Klebsiella phage Kpn-11mx | Klebsiella phage Kpn-11mx unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38774 | 50.812 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT499898 | Salmonella phage vB_SentM_sal3 | Salmonella phage vB_SentM_sal3 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57988 | 56.505 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MT499897 | Salmonella phage vB_SentM_sal2 | Salmonella phage vB_SentM_sal2 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59515 | 56.308 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MT499896 | Salmonella phage vB_SentM_sal1 | Salmonella phage vB_SentM_sal1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157338 | 44.924 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT193890 | Vibrio phage vB_Vipa71 | Vibrio phage vB_Vipa71 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10295 | 44.517 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MN737551 | Salmonella phage vB_SalS-LPSTLL | Salmonella phage vB_SalS-LPSTLL Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47243 | 45.645 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MW812343 | Erwinia phage pEa_SNUABM_6 | Erwinia phage pEa_SNUABM_6 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240164 | 50.595 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW812342 | Erwinia phage pEa_SNUABM_10 | Erwinia phage pEa_SNUABM_10 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 238920 | 51.417 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW812341 | Erwinia phage pEa_SNUABM_9 | Erwinia phage pEa_SNUABM_9 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 247273 | 49.693 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW812340 | Erwinia phage pEa_SNUABM_42 | Erwinia phage pEa_SNUABM_42 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241904 | 51.339 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW812339 | Erwinia phage pEa_SNUABM_29 | Erwinia phage pEa_SNUABM_29 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 248836 | 50.291 | Seoulvirus | Unclassified | Myoviridae | Erwinia |
| MW812338 | Erwinia phage pEa_SNUABM_4 | Erwinia phage pEa_SNUABM_4 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 249193 | 51.362 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MZ047271 | Campylobacter phage PC10 | Campylobacter phage PC10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136144 | 31.154 | Unclassified | Unclassified | Myoviridae | Campylobacter |
| MZ043897 | Escherichia phage E212 | Escherichia phage E212 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38252 | 46.983 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MZ031013 | Escherichia phage vB_EcoA_fTuEco01 | Escherichia phage vB_EcoA_fTuEco01 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44320 | 44.903 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MZ004649 | Salmonella phage vB_STM-ZS | Salmonella phage vB_STM-ZS unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40542 | 51.13 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW934263 | Stenotrophomonas phage BUCT603 | Stenotrophomonas phage BUCT603 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44912 | 63.68 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MW924497 | Staphylococcus phage PHB21 | Staphylococcus phage PHB21 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45260 | 33.959 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MW862109 | Pseudomonas phage Medea1 | Pseudomonas phage Medea1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58919 | 58.059 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW787012 | Bacillus phage SWEP1 | Bacillus phage SWEP1 unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162461 | 37.794 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MW679410 | Escherichia phage C6 | Escherichia phage C6 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169530 | 39.569 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MW978786 | Aeromonas phage BUCT552 | Aeromonas phage BUCT552 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59685 | 60.032 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MW689258 | Hafnia phage Pocis76 | Hafnia phage Pocis76 unclassified Tempevirinae Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48795 | 44.867 | Unclassified | Tempevirinae | Drexlerviridae | Hafnia |
| MW725301 | Salmonella phage PSDA-2 | Salmonella phage PSDA-2 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40062 | 51.241 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW713578 | Klebsiella phage Shaphc-TDM-1124-4 | Klebsiella phage Shaphc-TDM-1124-4 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112841 | 45.538 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MW767161 | Cronobacter phage JC03 | Cronobacter phage JC03 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177089 | 44.672 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| MW650886 | Escherichia phage vB_EcoM-ZQ1 | Escherichia phage vB_EcoM-ZQ1 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149115 | 49.051 | Unclassified | Aglimvirinae | Ackermannviridae | Escherichia |
| MW926912 | Acinetobacter phage vB_AbaP_100 | Acinetobacter phage vB_AbaP_100 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41743 | 39.571 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MZ150758 | Salmonella phage S19cd | Salmonella phage S19cd unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85995 | 39.005 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MZ150757 | Salmonella phage Lv5cm | Salmonella phage Lv5cm unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169920 | 35.295 | Tequatrovirus | Tevenvirinae | Myoviridae | Salmonella |
| MG752970 | Ralstonia phage RsoM2USA | Ralstonia phage RsoM2USA Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 343806 | 41.436 | Unclassified | Unclassified | Myoviridae | Ralstonia |
| MW960043 | Stenotrophomonas phage BUCT609 | Stenotrophomonas phage BUCT609 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43145 | 58.285 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| MZ188970 | Phage CBW1004C-Prop1 | Phage CBW1004C-Prop1 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35132 | 58.084 | Unclassified | Unclassified | Unclassified | Unspecified |
| MW660583 | Bacteroides phage LoVEphage | Bacteroides phage LoVEphage unclassified bacterial viruses Viruses | 71495 | 39.029 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MZ078141 | Anabaena phage Elbi | Anabaena phage Elbi Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68626 | 36.69 | Unclassified | Unclassified | Myoviridae | Anabaena |
| MW715619 | Salmonella phage JD01 | Salmonella phage JD01 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44880 | 49.713 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW815121 | Clostridium phage vB_CpeP_HN02 | Clostridium phage vB_CpeP_HN02 unclassified Gregsiragusavirus Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17754 | 28.202 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| MW767996 | Yersinia phage vB_YpP-YepMm | Yersinia phage vB_YpP-YepMm unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38512 | 47.123 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| MH382198 | Salmonella phage 3A_8767 | Salmonella phage 3A_8767 Salmonella virus 3A8767 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38821 | 49.254 | Teseptimavirus | Studiervirinae | Autographiviridae | Salmonella |
| MZ188976 | Cronobacter phage JK004 | Cronobacter phage JK004 unclassified Punavirus Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93825 | 53.963 | Punavirus | Unclassified | Myoviridae | Cronobacter |
| MZ089742 | Pseudomonas phage PSA40 | Pseudomonas phage PSA40 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45506 | 52.477 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089741 | Pseudomonas phage PSA39 | Pseudomonas phage PSA39 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47030 | 63.889 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ089740 | Pseudomonas phage PSA37 | Pseudomonas phage PSA37 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45506 | 52.452 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089739 | Pseudomonas phage PSA34 | Pseudomonas phage PSA34 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43749 | 52.049 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089738 | Pseudomonas phage PSA31 | Pseudomonas phage PSA31 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45505 | 52.533 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089737 | Pseudomonas phage PSA28 | Pseudomonas phage PSA28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48440 | 58.322 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ089736 | Pseudomonas phage PSA25 | Pseudomonas phage PSA25 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64290 | 55.467 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MZ089734 | Pseudomonas phage PSA20 | Pseudomonas phage PSA20 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62296 | 64.418 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MZ089733 | Pseudomonas phage PSA16 | Pseudomonas phage PSA16 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45623 | 52.484 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089732 | Pseudomonas phage PSA13 | Pseudomonas phage PSA13 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45731 | 52.592 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MZ089731 | Pseudomonas phage PSA11 | Pseudomonas phage PSA11 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48815 | 44.824 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| MZ089730 | Pseudomonas phage PSA09 | Pseudomonas phage PSA09 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62015 | 55.53 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MZ089729 | Pseudomonas phage PSA07_PB1 | Pseudomonas phage PSA07_PB1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65891 | 54.942 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MZ089728 | Pseudomonas phage PSA04 | Pseudomonas phage PSA04 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48544 | 59.165 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MZ127829 | Sphingopyxis phage VSN-002 | Sphingopyxis phage VSN-002 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41771 | 61.71 | Unclassified | Unclassified | Unclassified | Sphingopyxis |
| MT135176 | Hafnia phage yong1 | Hafnia phage yong1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43329 | 47.649 | Unclassified | Unclassified | Myoviridae | Hafnia |
| MW316733 | Acinetobacter phage Bestia | Acinetobacter phage Bestia Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97874 | 37.111 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MW316732 | Acinetobacter phage Herod | Acinetobacter phage Herod Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97418 | 37.117 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MW316731 | Acinetobacter phage Mithridates | Acinetobacter phage Mithridates Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96720 | 37.17 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MW741821 | Escherichia phage vB_EcoP_DE5 | Escherichia phage vB_EcoP_DE5 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77305 | 42.088 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MW546076 | Staphylococcus virus Remus | Staphylococcus virus Remus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141985 | 29.839 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546072 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148660 | 30.355 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW791312 | Lactococcus phage Nocturne116 | Lactococcus phage Nocturne116 unclassified Questintvirus Questintvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25554 | 37.986 | Questintvirus | Unclassified | Siphoviridae | Lactococcus |
| MW825358 | Xanthomonas phage vB_XciM_LucasX | Xanthomonas phage vB_XciM_LucasX Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 305651 | 54.916 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MW862804 | Escherichia phage UAE_MI-01 | Escherichia phage UAE_MI-01 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44281 | 54.714 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MW812345 | Pseudomonas phage vB_Pci_PCMW57 | Pseudomonas phage vB_Pci_PCMW57 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40117 | 57.609 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MW296865 | Salmonella phage SP3_SHan-2021 | Salmonella phage SP3_SHan-2021 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88041 | 38.848 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KF981876 | Mycobacterium phage ADLER F1725 | Mycobacterium phage ADLER F1725 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95702 | 62.717 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW546077 | Staphylococcus phage Romulus | Staphylococcus phage Romulus unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136651 | 29.872 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546075 | Staphylococcus phage PM93 | Staphylococcus phage PM93 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144038 | 29.806 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546074 | Staphylococcus phage PM4 | Staphylococcus phage PM4 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148627 | 30.361 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546073 | Staphylococcus phage BT3 | Staphylococcus phage BT3 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146878 | 30.167 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546071 | Staphylococcus phage PM56 | Staphylococcus phage PM56 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136656 | 29.871 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546070 | Staphylococcus phage PM32 | Staphylococcus phage PM32 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148303 | 30.306 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546069 | Staphylococcus phage PM36 | Staphylococcus phage PM36 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143035 | 30.407 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546068 | Staphylococcus phage PM34 | Staphylococcus phage PM34 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143062 | 30.392 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546067 | Staphylococcus phage PM28 | Staphylococcus phage PM28 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142406 | 30.379 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546066 | Staphylococcus phage PM25 | Staphylococcus phage PM25 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142467 | 30.431 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546065 | Staphylococcus phage PM22 | Staphylococcus phage PM22 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140588 | 30.412 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW546064 | Staphylococcus phage PM9 | Staphylococcus phage PM9 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148495 | 30.361 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW459163 | Acinetobacter phage vB_AbaP_APK128 | Acinetobacter phage vB_AbaP_APK128 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42013 | 39.219 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| LC574321 | Staphylococcus phage SaGU1 | Staphylococcus phage SaGU1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140909 | 30.22 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| NC_054762 | Rhodococcus phage PhailMary | Rhodococcus phage PhailMary unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45614 | 58.857 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MN605466 | Erysiphe necator associated levi-like virus 1 | Erysiphe necator associated levi-like virus 1 unclassified Leviviridae Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3638 | 50.55 | Unclassified | Unclassified | Leviviridae | Unspecified |
| MW655991 | Klebsiella phage RAD2 | Klebsiella phage RAD2 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49276 | 50.388 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN812687 | Pectobacterium phage Possum | Pectobacterium phage Possum unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73752 | 48.471 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MW713383 | Mycobacterium phage SirSheldon | Mycobacterium phage SirSheldon unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81714 | 61.537 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MW411580 | Vibrio phage vB_VpaS_VP-RY-9 | Vibrio phage vB_VpaS_VP-RY-9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81604 | 45.751 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW001769 | Pectobacterium phage phiPccP-1 | Pectobacterium phage phiPccP-1 Unyawovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40387 | 47.775 | Unyawovirus | Studiervirinae | Autographiviridae | Pectobacterium |
| KC960489 | Mycobacterium phage Adler | Mycobacterium phage Adler Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95705 | 62.716 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054725 | Gordonia phage Skog | Gordonia phage Skog Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152435 | 65.764 | Unclassified | Unclassified | Myoviridae | Gordonia |
| NC_054724 | Rhodococcus phage Finch | Rhodococcus phage Finch Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138896 | 63.079 | Unclassified | Unclassified | Myoviridae | Rhodococcus |
| NC_054723 | Gordonia phage Pupper | Gordonia phage Pupper Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150830 | 66.106 | Unclassified | Unclassified | Myoviridae | Gordonia |
| NC_054722 | Mycobacterium phage Sauce | Mycobacterium phage Sauce unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156554 | 64.721 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054721 | Mycobacterium phage Quasimodo | Mycobacterium phage Quasimodo unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156354 | 64.624 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054720 | Mycobacterium phage QBert | Mycobacterium phage QBert unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153772 | 64.812 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054719 | Mycobacterium phage NoodleTree | Mycobacterium phage NoodleTree unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151100 | 64.746 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054718 | Mycobacterium phage Mangeria | Mycobacterium phage Mangeria unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156170 | 64.833 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054717 | Mycobacterium phage CharlieB | Mycobacterium phage CharlieB unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155886 | 64.688 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054716 | Mycobacterium phage Cane17 | Mycobacterium phage Cane17 unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160330 | 64.613 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_054715 | Mycobacterium phage Bigswole | Mycobacterium phage Bigswole unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156514 | 64.775 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| MW876471 | Escherichia phage P762 | Escherichia phage P762 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38954 | 50.046 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MW672041 | Enterococcus phage vb_GEC_Ef_S_9 | Enterococcus phage vb_GEC_Ef_S_9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33631 | 36.056 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MW250787 | Escherichia phage UPEC07 | Escherichia phage UPEC07 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170004 | 40.313 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MW250786 | Escherichia phage UPEC06 | Escherichia phage UPEC06 unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143140 | 41.224 | Certrevirus | Vequintavirinae | Myoviridae | Escherichia |
| MW250785 | Escherichia phage UPEC03 | Escherichia phage UPEC03 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178307 | 44.557 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| MW924889 | Staphylococcus phage vB_StaphS-IVBph354 | Staphylococcus phage vB_StaphS-IVBph354 unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41970 | 32.964 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MW883060 | Escherichia phage vB_EcoP_SYGE1 | Escherichia phage vB_EcoP_SYGE1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39713 | 48.438 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MW883059 | Escherichia phage vB_EcoM_SYGD1 | Escherichia phage vB_EcoM_SYGD1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171255 | 35.319 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN563785 | Pseudomonas phage vB_PA45_GUMS | Pseudomonas phage vB_PA45_GUMS unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92467 | 49.413 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN563784 | Pseudomonas phage vB_PA32_GUMS | Pseudomonas phage vB_PA32_GUMS unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 280208 | 36.904 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MN563783 | Pseudomonas phage vB_PA6_GUMS | Pseudomonas phage vB_PA6_GUMS unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66806 | 55.128 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_054466 | Aeromonas phage 4_D05 | Aeromonas phage 4_D05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42249 | 55.987 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| NC_054465 | Aeromonas phage 2_D05 | Aeromonas phage 2_D05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43233 | 56.36 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| NC_054463 | Ralstonia phage Bakoly | Ralstonia phage Bakoly Ralstonia virus Bakoly Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44222 | 62.324 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MW916539 | Bacteroides phage GEC_vB_Bfr_VA7 | Bacteroides phage GEC_vB_Bfr_VA7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47095 | 38.526 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| MW748993 | Pseudomonas phage BUCT566 | Pseudomonas phage BUCT566 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49154 | 59.436 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW748991 | Enterobacter phage IME278 | Enterobacter phage IME278 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40164 | 51.987 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| MW924650 | Gordonia phage Neobush | Gordonia phage Neobush unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50906 | 66.67 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MW924648 | Gordonia phage Trumpet | Gordonia phage Trumpet unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50338 | 66.661 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MW314138 | Bacteroides phage vB_BfraS_NCTC | Bacteroides phage vB_BfraS_NCTC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47160 | 38.827 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| MW883061 | Escherichia phage vB_EcoM_SYGMH1 | Escherichia phage vB_EcoM_SYGMH1 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137423 | 43.606 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MW654182 | Nocardia phage KYD2 | Nocardia phage KYD2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46380 | 67.025 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| NC_054461 | Xanthomonas virus phiXaf18 | Xanthomonas virus phiXaf18 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47407 | 62.97 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| NC_054460 | Xanthomonas phage KPhi1 | Xanthomonas phage KPhi1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46077 | 62.817 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MW627366 | Pseudomonas phage phiGM22-3 | Pseudomonas phage phiGM22-3 Tunggulvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42662 | 58.654 | Tunggulvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MW722521 | Salmonella phage S9-5 | Salmonella phage S9-5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39167 | 48.163 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW822539 | Xanthomonas phage Soumapaz | Xanthomonas phage Soumapaz Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43818 | 66.605 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW822538 | Xanthomonas phage Fontebon | Xanthomonas phage Fontebon Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43824 | 66.614 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW822537 | Xanthomonas phage Alcala | Xanthomonas phage Alcala Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43741 | 66.581 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW822536 | Xanthomonas phage Usaquen | Xanthomonas phage Usaquen Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43741 | 66.61 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW822535 | Xanthomonas phage Bolivar | Xanthomonas phage Bolivar Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43782 | 66.605 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW802488 | Xanthomonas phage Olaya | Xanthomonas phage Olaya Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43722 | 66.573 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MW822011 | Shigella phage TB004 | Shigella phage TB004 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169805 | 35.429 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| NC_054447 | Mycobacterium phage Pinnie | Mycobacterium phage Pinnie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44842 | 68.761 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054446 | Mycobacterium phage Lemuria | Mycobacterium phage Lemuria Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44998 | 68.85 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054445 | Mycobacterium phage Mercurio | Mycobacterium phage Mercurio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44877 | 69.029 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054444 | Mycobacterium phage Antsirabe | Mycobacterium phage Antsirabe Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44739 | 69.007 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054443 | Mycobacterium phage Avocado | Mycobacterium phage Avocado Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45389 | 68.739 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054442 | Mycobacterium phage Grizzly | Mycobacterium phage Grizzly unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41752 | 66.758 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054441 | Mycobacterium phage Rabbs | Mycobacterium phage Rabbs unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42430 | 66.432 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054440 | Mycobacterium phage Taheera | Mycobacterium phage Taheera unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41077 | 66.434 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_054438 | Gordonia phage TZGordon | Gordonia phage TZGordon unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44904 | 66.094 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_054437 | Gordonia phage TinaLin | Gordonia phage TinaLin unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43366 | 65.408 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_054436 | Gordonia phage Schmidt | Gordonia phage Schmidt unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43099 | 65.67 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_054435 | Gordonia phage Dardanus | Gordonia phage Dardanus unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43143 | 66.595 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_054434 | Gordonia phage Catfish | Gordonia phage Catfish unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46888 | 64.976 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MW367417 | Pectobacterium phage PcCB7V | Pectobacterium phage PcCB7V unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146054 | 50.387 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| CP059058 | Proteus phage ASh-2020a | Proteus phage ASh-2020a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59078 | 46.72 | Unclassified | Unclassified | Siphoviridae | Proteus |
| NC_054431 | Microbacterium phage Scamander | Microbacterium phage Scamander Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17452 | 68.731 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_054430 | Arthrobacter phage Wawa | Arthrobacter phage Wawa unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43850 | 60.899 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054429 | Arthrobacter phage Urla | Arthrobacter phage Urla unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43940 | 61.181 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054428 | Arthrobacter phage Sergei | Arthrobacter phage Sergei unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43950 | 61.863 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054427 | Arthrobacter phage Oxynfrius | Arthrobacter phage Oxynfrius unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44163 | 60.829 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054426 | Arthrobacter phage Nubia | Arthrobacter phage Nubia unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44045 | 60.695 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054425 | Arthrobacter phage MamaPearl | Arthrobacter phage MamaPearl unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43746 | 61.263 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054424 | Arthrobacter phage Litotes | Arthrobacter phage Litotes unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43496 | 61.081 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054423 | Arthrobacter phage Kittykat | Arthrobacter phage Kittykat unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43603 | 60.686 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054422 | Arthrobacter phage Huntingdon | Arthrobacter phage Huntingdon unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43891 | 60.65 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054421 | Arthrobacter phage Greenhouse | Arthrobacter phage Greenhouse unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43977 | 60.827 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054420 | Arthrobacter phage GreenHearts | Arthrobacter phage GreenHearts unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44251 | 60.753 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054419 | Arthrobacter phage Gisselle | Arthrobacter phage Gisselle unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43089 | 60.735 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054418 | Arthrobacter phage BigMack | Arthrobacter phage BigMack unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43357 | 61.266 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_054393 | Mycobacterium phage Typha | Mycobacterium phage Typha Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75687 | 65.953 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_054146 | Ralstonia phage phiRSP | Ralstonia phage phiRSP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44526 | 62.256 | Unclassified | Unclassified | Myoviridae | Ralstonia |
| NC_031253 | Mycobacterium phage Bipper | Mycobacterium phage Bipper Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77832 | 67.341 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_031028 | Gordonia phage Ghobes | Gordonia phage Ghobes Gordonia virus Ghobes Ghobesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45285 | 65.247 | Ghobesvirus | Unclassified | Siphoviridae | Gordonia |
| NC_054392 | Gordonia phage Archimedes | Gordonia phage Archimedes Ghobesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44938 | 62.818 | Ghobesvirus | Unclassified | Siphoviridae | Gordonia |
| MW380985 | Aeromonas phage PVN05 | Aeromonas phage PVN05 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51884 | 52.428 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| MW380984 | Aeromonas phage PVN04 | Aeromonas phage PVN04 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51721 | 52.443 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| MW380983 | Aeromonas phage PVN03 | Aeromonas phage PVN03 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50725 | 52.347 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| MW528836 | Staphylococcus phage vB_SauH_2002 | Staphylococcus phage vB_SauH_2002 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145076 | 30.358 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW672037 | Klebsiella phage B1 | Klebsiella phage B1 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50040 | 50.408 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW835180 | Pseudomonas phage vB_PaeS_PAJD-1 | Pseudomonas phage vB_PaeS_PAJD-1 unclassified Nipunavirus Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57919 | 58.323 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| MW822601 | Synechoccus phage S-SRP02 | Synechoccus phage S-SRP02 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42143 | 63.425 | Unclassified | Unclassified | Autographiviridae | Synechococcus |
| MW367418 | Pectobacterium phage PcCB251 | Pectobacterium phage PcCB251 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40557 | 48.603 | Berlinvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MT768060 | Escherichia phage vB_EcoS_swi2 | Escherichia phage vB_EcoS_swi2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47611 | 46.254 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MT768059 | Escherichia phage vB_EcoM_swi3 | Escherichia phage vB_EcoM_swi3 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52782 | 46.021 | Rosemountvirus | Unclassified | Myoviridae | Escherichia |
| MW451251 | Vibrio phage TY21 | Vibrio phage TY21 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76778 | 48.789 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MW451250 | Vibrio phage TY18 | Vibrio phage TY18 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191500 | 34.899 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW451249 | Vibrio phage GH32 | Vibrio phage GH32 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56078 | 51.184 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MW451248 | Vibrio phage VP506 | Vibrio phage VP506 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50577 | 41.63 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW451247 | Vibrio phage VP505 | Vibrio phage VP505 Trungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43009 | 43.586 | Trungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| MW451246 | Vibrio phage VP11 | Vibrio phage VP11 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74998 | 48.921 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MW629017 | Enterobacter phage PF-CE2 | Enterobacter phage PF-CE2 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178248 | 44.762 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| LC623721 | Enterococcus phage vB_EfaS-SRH2 | Enterococcus phage vB_EfaS-SRH2 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38749 | 34.778 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| LC483177 | Klebsiella phage 05F01 | Klebsiella phage 05F01 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147331 | 35.546 | Unclassified | Vequintavirinae | Myoviridae | Klebsiella |
| MW567727 | Salmonella phage STWB21 | Salmonella phage STWB21 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112834 | 40.373 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW735835 | Pseudomonas phage pPa_SNUABM_DT01 | Pseudomonas phage pPa_SNUABM_DT01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265520 | 56.081 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MW671054 | Aeromonas phage PZL-Ah152 | Aeromonas phage PZL-Ah152 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 51.744 | Unclassified | Studiervirinae | Autographiviridae | Aeromonas |
| MT234340 | Agrobacterium phage OLIVR3 | Agrobacterium phage OLIVR3 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75576 | 45.386 | Unclassified | Unclassified | Schitoviridae | Agrobacterium |
| MT234339 | Agrobacterium phage OLIVR2 | Agrobacterium phage OLIVR2 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75576 | 45.381 | Unclassified | Unclassified | Schitoviridae | Agrobacterium |
| MT234338 | Agrobacterium phage OLIVR1 | Agrobacterium phage OLIVR1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75576 | 45.389 | Unclassified | Unclassified | Schitoviridae | Agrobacterium |
| MW722081 | Klebsiella phage vB_KpnS_ZX2 | Klebsiella phage vB_KpnS_ZX2 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51632 | 51.826 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW495044 | Klebsiella phage vB_KpnP_P184 | Klebsiella phage vB_KpnP_P184 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76617 | 44.34 | Unclassified | Unclassified | Schitoviridae | Klebsiella |
| HG992588 | Vibrio phage B1_1 | Vibrio phage B1_1 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77004 | 48.856 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MT951568 | Xanthomonas phage Xaa_vB_phi31 | Xanthomonas phage Xaa_vB_phi31 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42825 | 60.371 | Unclassified | Okabevirinae | Autographiviridae | Xanthomonas |
| MT438396 | Klebsiella phage KPP-1 | Klebsiella phage KPP-1 unclassified Mydovirus Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143369 | 44.671 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| MT161385 | Xanthomonas phage FoX4 | Xanthomonas phage FoX4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60418 | 61.758 | Unclassified | Unclassified | Siphoviridae | Xanthomonas |
| MT161387 | Xanthomonas phage FoX7 | Xanthomonas phage FoX7 Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61010 | 52.262 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| MT161386 | Xanthomonas phage FoX6 | Xanthomonas phage FoX6 Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61077 | 52.249 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| MT161384 | Xanthomonas phage FoX5 | Xanthomonas phage FoX5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43872 | 59.466 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT161383 | Xanthomonas phage FoX3 | Xanthomonas phage FoX3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44153 | 59.346 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT161382 | Xanthomonas phage FoX2 | Xanthomonas phage FoX2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44086 | 59.475 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT161381 | Xanthomonas phage FoX1 | Xanthomonas phage FoX1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44763 | 59.429 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT234343 | Agrobacterium phage OLIVR6 | Agrobacterium phage OLIVR6 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152004 | 46.942 | Unclassified | Unclassified | Ackermannviridae | Agrobacterium |
| MT234342 | Agrobacterium phage OLIVR5 | Agrobacterium phage OLIVR5 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151073 | 46.936 | Unclassified | Unclassified | Ackermannviridae | Agrobacterium |
| MT234341 | Agrobacterium phage OLIVR4 | Agrobacterium phage OLIVR4 unclassified Schmittlotzvirus Schmittlotzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67981 | 51.722 | Schmittlotzvirus | Unclassified | Myoviridae | Agrobacterium |
| MW416012 | Salmonella phage vB Seyj3-1 | Salmonella phage vB Seyj3-1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39619 | 50.726 | Kayfunavirus | Studiervirinae | Autographiviridae | Salmonella |
| MW601222 | Mycobacterium phage Fefferhead | Mycobacterium phage Fefferhead unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61366 | 67.022 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW601220 | Gordonia phage Guey18 | Gordonia phage Guey18 unclassified Ronaldovirus Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68896 | 50.145 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MW601218 | Mycobacterium phage Juliette | Mycobacterium phage Juliette unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58071 | 67.461 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW600722 | Klebsiella phage KPP-5 | Klebsiella phage KPP-5 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38245 | 50.788 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW041639 | Lactococcus phage RH6 | Lactococcus phage RH6 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30299 | 35.067 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041638 | Lactococcus phage GL7 | Lactococcus phage GL7 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29705 | 34.25 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW602648 | Escherichia phage vB_EcoM_fHy-Eco03 | Escherichia phage vB_EcoM_fHy-Eco03 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57011 | 45.06 | Rosemountvirus | Unclassified | Myoviridae | Escherichia |
| MT872956 | Stenotrophomonas phage phiSHP3 | Stenotrophomonas phage phiSHP3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37611 | 65.305 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MT129655 | Escherichia phage MN05 | Escherichia phage MN05 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76899 | 42.153 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT129654 | Escherichia phage MN04 | Escherichia phage MN04 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169519 | 39.438 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MT129653 | Escherichia phage MN03 | Escherichia phage MN03 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77187 | 42.159 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT129652 | Escherichia phage MN01 | Escherichia phage MN01 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172584 | 40.423 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MW291508 | Stenotrophomonas phage BUCT555 | Stenotrophomonas phage BUCT555 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39440 | 61.433 | Unclassified | Unclassified | Podoviridae | Stenotrophomonas |
| MW072790 | Burkholderia phage phiE094 | Burkholderia phage phiE094 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37727 | 64.484 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| MT210152 | Bacillus phage vB_Bacillus_1020A | Bacillus phage vB_Bacillus_1020A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45486 | 37.515 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MW032452 | Enterobacter phage N5822 | Enterobacter phage N5822 unclassified Henuseptimavirus Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78128 | 48.765 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Enterobacter |
| NC_053512 | Mycobacterium phage Kumao | Mycobacterium phage Kumao Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70373 | 62.072 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW015081 | Synechococcus phage S-SRM01 | Synechococcus phage S-SRM01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240842 | 35.642 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MT375531 | Pelagibacter phage Unn EXVC019P | Pelagibacter phage Unn EXVC019P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41069 | 33.463 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MT375530 | Pelagibacter phage Ran EXVC014P | Pelagibacter phage Ran EXVC014P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41529 | 34.138 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MT375529 | Pelagibacter phage Kolga EXVC016S | Pelagibacter phage Kolga EXVC016S unclassified bacterial viruses Viruses | 48659 | 30.471 | Unclassified | Unclassified | Unclassified | Pelagibacter |
| MT375528 | Pelagibacter phage Jormungand EXVC012P | Pelagibacter phage Jormungand EXVC012P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41529 | 34.138 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MT375527 | Pelagibacter phage Hroenn EXVC015P | Pelagibacter phage Hroenn EXVC015P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41069 | 33.463 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MT375526 | Pelagibacter phage Himinglaeva EXVC011P | Pelagibacter phage Himinglaeva EXVC011P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41069 | 33.463 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MT375525 | Pelagibacter phage Greip EXVC021P | Pelagibacter phage Greip EXVC021P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34916 | 31.533 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MT375524 | Pelagibacter phage Gjalp EXVC020P | Pelagibacter phage Gjalp EXVC020P unclassified Fussvirus Fussvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37857 | 32.417 | Fussvirus | Unclassified | Autographiviridae | Pelagibacter |
| MT375523 | Pelagibacter phage Eyrgjafa EXVC018P | Pelagibacter phage Eyrgjafa EXVC018P unclassified Fussvirus Fussvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38005 | 32.556 | Fussvirus | Unclassified | Autographiviridae | Pelagibacter |
| MT375522 | Methylophilales phage Venkman EXVC282S | Methylophilales phage Venkman EXVC282S Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38624 | 34.403 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT375521 | Pelagibacter phage Eistla EXVC025P | Pelagibacter phage Eistla EXVC025P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39638 | 32.658 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MT375520 | Pelagibacter phage Bylgja EXVC010P | Pelagibacter phage Bylgja EXVC010P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41069 | 33.463 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MT375519 | Pelagibacter phage Aegir EXVC013S | Pelagibacter phage Aegir EXVC013S unclassified bacterial viruses Viruses | 18297 | 31.065 | Unclassified | Unclassified | Unclassified | Pelagibacter |
| MW015080 | Synechococcus phage S-SRP01 | Synechococcus phage S-SRP01 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45017 | 48.868 | Unclassified | Unclassified | Autographiviridae | Synechococcus |
| LC620505 | Escherichia phage NiC89 | Escherichia phage NiC89 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166727 | 35.417 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_053503 | Mycobacterium phage Laurie | Mycobacterium phage Laurie Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66507 | 68.955 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MT215167 | Vibrio phage JSF37 | Vibrio phage JSF37 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38451 | 42.738 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| MW584212 | Mycobacterium phage prophiGD13-2 | Mycobacterium phage prophiGD13-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51341 | 63.9 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584210 | Mycobacterium phage prophiGD34-2 | Mycobacterium phage prophiGD34-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40044 | 63.742 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584208 | Mycobacterium phage prophi52-1 | Mycobacterium phage prophi52-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49796 | 64.021 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584205 | Mycobacterium phage prophiGD21-1 | Mycobacterium phage prophiGD21-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40775 | 63.556 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584203 | Mycobacterium phage prophiGD33-1 | Mycobacterium phage prophiGD33-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51076 | 63.938 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584200 | Mycobacterium phage prophiGD42-2 | Mycobacterium phage prophiGD42-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40071 | 63.54 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584198 | Mycobacterium phage prophiGD43A-2 | Mycobacterium phage prophiGD43A-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40147 | 63.397 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584194 | Mycobacterium phage prophi62-1 | Mycobacterium phage prophi62-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39188 | 63.601 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584193 | Mycobacterium phage prophi89-1 | Mycobacterium phage prophi89-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40492 | 63.375 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584192 | Mycobacterium phage prophi91-1 | Mycobacterium phage prophi91-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49758 | 64.108 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584191 | Mycobacterium phage prophiGD05-2 | Mycobacterium phage prophiGD05-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43869 | 64.373 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584186 | Mycobacterium phage prophi108-1 | Mycobacterium phage prophi108-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42642 | 63.989 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584184 | Mycobacterium phage prophiGD08-2 | Mycobacterium phage prophiGD08-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40071 | 63.542 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584183 | Mycobacterium phage prophiGD53-3 | Mycobacterium phage prophiGD53-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41918 | 63.922 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584182 | Mycobacterium phage prophiGD43A-3 | Mycobacterium phage prophiGD43A-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50750 | 63.947 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584181 | Mycobacterium phage prophi57-1 | Mycobacterium phage prophi57-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52581 | 63.789 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584177 | Mycobacterium phage prophi62-3 | Mycobacterium phage prophi62-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42642 | 63.986 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584170 | Mycobacterium phage prophiGD36-2 | Mycobacterium phage prophiGD36-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43586 | 64.615 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584167 | Mycobacterium phage prophiGD43A-5 | Mycobacterium phage prophiGD43A-5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76541 | 64.33 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584163 | Mycobacterium phage prophiGD51-1 | Mycobacterium phage prophiGD51-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51204 | 64.286 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584162 | Mycobacterium phage prophi104-2 | Mycobacterium phage prophi104-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49556 | 64.311 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584156 | Mycobacterium phage prophiGD44-1 | Mycobacterium phage prophiGD44-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53061 | 64.079 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584154 | Mycobacterium phage prophiGD39-2 | Mycobacterium phage prophiGD39-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52581 | 63.808 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584151 | Mycobacterium phage prophiGD11-2 | Mycobacterium phage prophiGD11-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39951 | 63.535 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584150 | Mycobacterium phage prophi100A-2 | Mycobacterium phage prophi100A-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50828 | 63.914 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584149 | Mycobacterium phage prophiGD16-1 | Mycobacterium phage prophiGD16-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40056 | 63.159 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584148 | Mycobacterium phage prophiGD25-1 | Mycobacterium phage prophiGD25-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52581 | 63.789 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584176 | Mycobacterium phage prophiGD90-1 | Mycobacterium phage prophiGD90-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60552 | 59.549 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584211 | Mycobacterium phage prophiGD102-1 | Mycobacterium phage prophiGD102-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63235 | 59.25 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584209 | Mycobacterium phage prophiGD04-1 | Mycobacterium phage prophiGD04-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60432 | 62.474 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584207 | Mycobacterium phage prophiGD12-2 | Mycobacterium phage prophiGD12-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54460 | 62.501 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584206 | Mycobacterium phage prophi91-4 | Mycobacterium phage prophi91-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58498 | 63.214 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584204 | Mycobacterium phage prophiGD21-2 | Mycobacterium phage prophiGD21-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61985 | 59.251 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584201 | Mycobacterium phage prophiGD08-3 | Mycobacterium phage prophiGD08-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52601 | 64.449 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584199 | Mycobacterium phage prophiGD02-2 | Mycobacterium phage prophiGD02-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60552 | 59.544 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584197 | Mycobacterium phage prophiGD43A-4 | Mycobacterium phage prophiGD43A-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54433 | 57.518 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584196 | Mycobacterium phage prophiGD27-1 | Mycobacterium phage prophiGD27-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59545 | 59.639 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584195 | Mycobacterium phage prophiGD51-2 | Mycobacterium phage prophiGD51-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51838 | 69.191 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584190 | Mycobacterium phage prophiGD03-1 | Mycobacterium phage prophiGD03-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54821 | 63.986 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584189 | Mycobacterium phage prophiGD54-1 | Mycobacterium phage prophiGD54-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60696 | 62.456 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584188 | Mycobacterium phage prophiGD91-2 | Mycobacterium phage prophiGD91-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64208 | 59.471 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584185 | Mycobacterium phage prophiGD05-3 | Mycobacterium phage prophiGD05-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55492 | 63.916 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584180 | Mycobacterium phage prophiGD15-1 | Mycobacterium phage prophiGD15-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70582 | 59.604 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584179 | Mycobacterium phage prophiGD43A-1 | Mycobacterium phage prophiGD43A-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62303 | 59.556 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584178 | Mycobacterium phage prophiGD21-3 | Mycobacterium phage prophiGD21-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52874 | 63.697 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584175 | Mycobacterium phage prophi62-2 | Mycobacterium phage prophi62-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54310 | 64.43 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584174 | Mycobacterium phage prophi43-6 | Mycobacterium phage prophi43-6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66367 | 64.003 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584173 | Mycobacterium phage prophi102-2 | Mycobacterium phage prophi102-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60696 | 62.459 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584172 | Mycobacterium phage prophiGD24-2 | Mycobacterium phage prophiGD24-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52875 | 63.707 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584171 | Mycobacterium phage prophiGD22-1 | Mycobacterium phage prophiGD22-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60575 | 59.386 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584169 | Mycobacterium phage prophiGD05-1 | Mycobacterium phage prophiGD05-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60892 | 63.016 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584168 | Mycobacterium phage prophi58-1 | Mycobacterium phage prophi58-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56085 | 63.719 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584166 | Mycobacterium phage prophi88-1 | Mycobacterium phage prophi88-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66356 | 64.005 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584165 | Mycobacterium phage prophiGD17-1 | Mycobacterium phage prophiGD17-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51224 | 63.339 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584164 | Mycobacterium phage prophiGD57-2 | Mycobacterium phage prophiGD57-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60012 | 59.42 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584161 | Mycobacterium phage prophiGD17-2 | Mycobacterium phage prophiGD17-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61797 | 59.519 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584159 | Mycobacterium phage prophiGD24-3 | Mycobacterium phage prophiGD24-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54885 | 57.735 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584158 | Mycobacterium phage prophiGD20-1 | Mycobacterium phage prophiGD20-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69032 | 59.5 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584157 | Mycobacterium phage prophi68-1 | Mycobacterium phage prophi68-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60432 | 62.472 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584155 | Mycobacterium phage prophiGD11-3 | Mycobacterium phage prophiGD11-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54740 | 64.545 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584153 | Mycobacterium phage prophi79-1 | Mycobacterium phage prophi79-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76148 | 62.59 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584152 | Mycobacterium phage prophiGD11-1 | Mycobacterium phage prophiGD11-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62235 | 59.325 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW595221 | Pseudomonas phage vB_PaeM_V524 | Pseudomonas phage vB_PaeM_V524 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65447 | 55.535 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW595220 | Pseudomonas phage vB_PaeM_V523 | Pseudomonas phage vB_PaeM_V523 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66301 | 55.645 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW534378 | Gordonia phage Cashline | Gordonia phage Cashline Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52378 | 66.326 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN091626 | Staphylococcus phage PALS_2 | Staphylococcus phage PALS_2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 268748 | 25.133 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MW584187 | Mycobacterium phage prophi91-3 | Mycobacterium phage prophi91-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45607 | 63.602 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW584202 | Mycobacterium phage prophiGD54-2 | Mycobacterium phage prophiGD54-2 Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79047 | 58.229 | Unclassified | Mclasvirinae | Siphoviridae | Mycobacterium |
| MW584160 | Mycobacterium phage prophi86-1 | Mycobacterium phage prophi86-1 Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80793 | 57.468 | Unclassified | Mclasvirinae | Siphoviridae | Mycobacterium |
| MW590698 | Acinetobacter phage 53 | Acinetobacter phage 53 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42916 | 47.106 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MW578835 | Mycobacterium phage Netyap | Mycobacterium phage Netyap unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76366 | 58.908 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW554488 | Staphylococcus phage BUCT556 | Staphylococcus phage BUCT556 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43207 | 34.059 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MW560978 | Oceanospirillum phage vB_OliS_GJ44 | Oceanospirillum phage vB_OliS_GJ44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33786 | 48.766 | Unclassified | Unclassified | Siphoviridae | Oceanospirillum |
| MW460248 | Mycobacterium phage Maco2 | Mycobacterium phage Maco2 unclassified Mapvirus Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48239 | 58.672 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW512834 | Pseudomonas phage AUS391 | Pseudomonas phage AUS391 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65515 | 55.52 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW512833 | Pseudomonas phage AUS301 | Pseudomonas phage AUS301 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49434 | 55.824 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW512832 | Pseudomonas phage AUS260 | Pseudomonas phage AUS260 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42020 | 54.66 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MW512831 | Pseudomonas phage AUS034 | Pseudomonas phage AUS034 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44167 | 52.247 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MW512830 | Pseudomonas phage Qaehs | Pseudomonas phage Qaehs unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66283 | 55.578 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW512264 | Escherichia phage PTK | Escherichia phage PTK unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160320 | 37.653 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MW481326 | Escherichia phage vB_EcoM-Pr103Blw | Escherichia phage vB_EcoM-Pr103Blw unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88421 | 38.725 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| BK013347 | Thermocrinis Great Boiling Spring virus | Thermocrinis Great Boiling Spring virus unclassified archaeal viruses Viruses | 41208 | 36.918 | Unclassified | Unclassified | Unclassified | Thermocrinis |
| BK013346 | Aquificae Joseph’s Coat Spring virus | Aquificae Joseph’s Coat Spring virus unclassified archaeal viruses Viruses | 39372 | 33.968 | Unclassified | Unclassified | Unclassified | Unspecified |
| BK013345 | Aquificae Conch Spring virus | Aquificae Conch Spring virus unclassified archaeal viruses Viruses | 37258 | 35.515 | Unclassified | Unclassified | Unclassified | Unspecified |
| NC_053249 | Gordonia phage Verity | Gordonia phage Verity Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61248 | 67.418 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053248 | Gordonia phage Zipp | Gordonia phage Zipp Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61219 | 67.378 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053247 | Gordonia Phage Zitch | Gordonia Phage Zitch Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58928 | 67.71 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053246 | Gordonia phage Lilbeanie | Gordonia phage Lilbeanie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59210 | 67.715 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053245 | Gordonia phage Ali17 | Gordonia phage Ali17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58301 | 68.404 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053244 | Gordonia phage MelBins | Gordonia phage MelBins Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59761 | 68.337 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053243 | Gordonia phage EMoore | Gordonia phage EMoore Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58844 | 68.031 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053242 | Gordonia phage Leonard | Gordonia phage Leonard Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59268 | 68.435 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053236 | Gordonia phage YorkOnyx | Gordonia phage YorkOnyx unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57430 | 68.247 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053235 | Gordonia phage JKSyngboy | Gordonia phage JKSyngboy unclassified Vividuovirus Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58176 | 67.985 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_053211 | Gordonia phage Tiamoceli | Gordonia phage Tiamoceli Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55829 | 67.698 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_053210 | Gordonia phage Dexdert | Gordonia phage Dexdert Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55006 | 67.504 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW582532 | Nocardia phage P3.1 | Nocardia phage P3.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66621 | 57.033 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| MW582531 | Nocardia phage P69 | Nocardia phage P69 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68903 | 65.556 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| MW590223 | Cellulosimicrobium phage DS1 | Cellulosimicrobium phage DS1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46217 | 55.317 | Unclassified | Unclassified | Podoviridae | Cellulosimicrobium |
| MW495853 | Streptococcus phage A1 | Streptococcus phage A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37239 | 37.915 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MW589261 | Arcanobacterium phage vB-ApyS-JF1 | Arcanobacterium phage vB-ApyS-JF1 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90130 | 30.576 | Sextaecvirus | Unclassified | Siphoviridae | Arcanobacterium |
| MW544066 | Salmonella phage vB_SalP_TR2 | Salmonella phage vB_SalP_TR2 unclassified Ithacavirus Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70823 | 40.646 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| MW584871 | Aeromonas phage vB_AsM_ZHF | Aeromonas phage vB_AsM_ZHF unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161887 | 41.25 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| MW651859 | Pseudomonas phage vB_PstS-pAN | Pseudomonas phage vB_PstS-pAN Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39466 | 62.679 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW329986 | Pseudomonas phage vB_PaeP_fHoPae04 | Pseudomonas phage vB_PaeP_fHoPae04 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45491 | 52.23 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MW358880 | Mycoplasma phage MGyu-2021a | Mycoplasma phage MGyu-2021a unclassified bacterial viruses Viruses | 24404 | 22.07 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW355480 | Salmonella phage vB_SenP_ER25 | Salmonella phage vB_SenP_ER25 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41727 | 47.061 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW355479 | Salmonella phage vB_SenS_ER24 | Salmonella phage vB_SenS_ER24 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60438 | 56.58 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355478 | Salmonella phage vB_SenAc_BPS7 | Salmonella phage vB_SenAc_BPS7 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158348 | 44.435 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW355477 | Salmonella phage vB_SenAc_BPS5 | Salmonella phage vB_SenAc_BPS5 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157674 | 44.634 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW355476 | Salmonella phage vB_SenS_BPS4 | Salmonella phage vB_SenS_BPS4 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58529 | 56.623 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355475 | Salmonella phage vB_SenAc_BPS3 | Salmonella phage vB_SenAc_BPS3 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157578 | 44.64 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW355474 | Salmonella phage vB_SenS_BPS1 | Salmonella phage vB_SenS_BPS1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58852 | 56.591 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355473 | Salmonella phage vB_SenS_ER9 | Salmonella phage vB_SenS_ER9 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42664 | 49.843 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355472 | Salmonella phage vB_SenS_ER8 | Salmonella phage vB_SenS_ER8 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42863 | 49.796 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355471 | Salmonella phage vB_SenS_ER7 | Salmonella phage vB_SenS_ER7 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42664 | 49.855 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355470 | Salmonella phage vB_SenS_ER6 | Salmonella phage vB_SenS_ER6 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58872 | 56.405 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355469 | Salmonella phage vB_SenS_ER5 | Salmonella phage vB_SenS_ER5 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42663 | 49.844 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355468 | Salmonella phage vB_SenS_ER4 | Salmonella phage vB_SenS_ER4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42848 | 49.846 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355467 | Salmonella phage vB_SenS_ER3 | Salmonella phage vB_SenS_ER3 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58971 | 56.385 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355466 | Salmonella phage vB_SenS_ER2 | Salmonella phage vB_SenS_ER2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42848 | 49.841 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355465 | Salmonella phage vB_SenS_ER23 | Salmonella phage vB_SenS_ER23 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43125 | 49.637 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355464 | Salmonella phage vB_SenS_ER22 | Salmonella phage vB_SenS_ER22 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42948 | 49.786 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355463 | Salmonella phage vB_SenS_ER21 | Salmonella phage vB_SenS_ER21 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42503 | 49.804 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355462 | Salmonella phage vB_SenS_ER20 | Salmonella phage vB_SenS_ER20 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42844 | 49.818 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355461 | Salmonella phage vB_SenS_ER1 | Salmonella phage vB_SenS_ER1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42847 | 49.798 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355460 | Salmonella phage vB_SenS_ER19 | Salmonella phage vB_SenS_ER19 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58344 | 56.585 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MW355459 | Salmonella phage vB_SenS_ER18 | Salmonella phage vB_SenS_ER18 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42862 | 49.795 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355458 | Salmonella phage vB_SenS_ER17 | Salmonella phage vB_SenS_ER17 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42848 | 49.844 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355457 | Salmonella phage vB_SenS_ER16 | Salmonella phage vB_SenS_ER16 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43078 | 49.907 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355456 | Salmonella phage vB_SenS_ER15 | Salmonella phage vB_SenS_ER15 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42846 | 49.841 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355455 | Salmonella phage vB_SenS_ER14 | Salmonella phage vB_SenS_ER14 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42841 | 49.814 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355454 | Salmonella phage vB_SenS_ER13 | Salmonella phage vB_SenS_ER13 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43078 | 49.907 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355453 | Salmonella phage vB_SenS_ER12 | Salmonella phage vB_SenS_ER12 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42843 | 49.817 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355452 | Salmonella phage vB_SenS_ER11 | Salmonella phage vB_SenS_ER11 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42840 | 49.813 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355451 | Salmonella phage vB_SenS_ER10 | Salmonella phage vB_SenS_ER10 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42665 | 49.844 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MW355450 | Salmonella phage vB_SenAc_BPS6 | Salmonella phage vB_SenAc_BPS6 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157347 | 44.63 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW355449 | Salmonella phage vB_SenS_BPS2 | Salmonella phage vB_SenS_BPS2 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58526 | 56.623 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| HG994092 | Klebsiella phage TUN1 | Klebsiella phage TUN1 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41181 | 52.942 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW557846 | Pseudomonas phage zikora | Pseudomonas phage zikora unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65837 | 54.876 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT647606 | Pelagibacter phage Mosig EXVC030M | Pelagibacter phage Mosig EXVC030M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141461 | 33.008 | Unclassified | Unclassified | Myoviridae | Pelagibacter |
| MT647605 | Pelagibacter phage Lederberg EXVC029P | Pelagibacter phage Lederberg EXVC029P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33623 | 33.126 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MT161459 | Aeromonas phage Ahp1_CNU-2021 | Aeromonas phage Ahp1_CNU-2021 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218317 | 47.944 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| CP071049 | Staphylococcus phage WBG8381 | Staphylococcus phage WBG8381 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42493 | 34.909 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MW004545 | Enterococcus phage EFGrNG | Enterococcus phage EFGrNG unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145199 | 37.327 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MT920315 | Erwinia phage FBB1 | Erwinia phage FBB1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175889 | 36.454 | Unclassified | Tevenvirinae | Myoviridae | Erwinia |
| CP069347 | Clostridioides phage pCD5401_3 | Clostridioides phage pCD5401_3 unclassified Sherbrookevirus Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34243 | 29.381 | Sherbrookevirus | Unclassified | Myoviridae | Clostridioides |
| MT002873 | Tenacibaculum phage JQ | Tenacibaculum phage JQ Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40712 | 30.649 | Unclassified | Unclassified | Siphoviridae | Tenacibaculum |
| MN812722 | Vibrio phage NF | Vibrio phage NF Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44507 | 43.128 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MW462221 | Campylobacter phage CAM-P21 | Campylobacter phage CAM-P21 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12587 | 30.071 | Unclassified | Unclassified | Myoviridae | Campylobacter |
| MW495066 | Microcystis phage MaAM05 | Microcystis phage MaAM05 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273876 | 54.222 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MW598258 | Escherichia phage U136B | Escherichia phage U136B unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49223 | 42.919 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MT984581 | Escherichia phage T4 | Escherichia phage T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168908 | 35.29 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT899444 | Liberibacter phage P-PA19-1 | Liberibacter phage P-PA19-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37601 | 40.901 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| MW452941 | Methylophilales phage MEP301 | Methylophilales phage MEP301 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34774 | 40.355 | Unclassified | Unclassified | Unclassified | Unspecified |
| MW582634 | Corynebacterium phage CL31 | Corynebacterium phage CL31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45061 | 54.364 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| MT135464 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7064 | 43.545 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135463 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5677 | 38.541 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135462 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5714 | 40.48 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135461 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6006 | 37.829 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135460 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6327 | 38.723 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135459 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4575 | 37.224 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135458 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4648 | 30.594 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135457 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4955 | 39.374 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135456 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5132 | 40.179 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135455 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6441 | 37.448 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135454 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3752 | 36.061 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135453 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3815 | 38.716 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135452 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3977 | 43.626 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135451 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4090 | 42.372 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135450 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4201 | 41.466 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135449 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4287 | 32.54 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135448 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4558 | 36.003 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135447 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4587 | 42.097 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135446 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4607 | 36.141 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135445 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5428 | 37.841 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135444 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5752 | 41.255 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135443 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5753 | 40.97 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135442 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6093 | 40.653 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135441 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6251 | 39.05 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135440 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7046 | 39.781 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135439 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7081 | 36.619 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135438 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5205 | 38.002 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135437 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 36.552 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135436 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6172 | 43.195 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135435 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6342 | 41.895 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135434 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6538 | 41.465 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135433 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5722 | 39.584 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135432 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6209 | 41.923 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135431 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4093 | 38.236 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135430 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4422 | 37.313 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135429 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4512 | 37.168 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135428 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4920 | 39.309 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135427 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5417 | 43.05 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135426 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5001 | 39.832 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135425 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5053 | 40.412 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135424 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5118 | 39.097 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135423 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5944 | 40.999 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135422 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6202 | 41.857 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135421 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4826 | 39.847 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135420 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5856 | 37.961 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135419 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6195 | 37.369 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135418 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6527 | 36.479 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135417 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 36.352 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135416 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6259 | 39.255 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135415 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6771 | 42.283 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135414 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6221 | 39.238 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135413 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4788 | 37.469 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135412 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5137 | 40.841 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135411 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5354 | 38.364 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135410 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6495 | 39.045 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135409 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6042 | 42.304 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135408 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4688 | 43.558 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135407 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5807 | 37.145 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135406 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4543 | 38.059 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135405 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4636 | 42.148 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135404 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4921 | 40.134 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135403 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5028 | 41.408 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135402 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6349 | 36.62 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135401 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5724 | 38.871 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135400 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6187 | 37.094 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135399 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5929 | 38.86 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135398 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6865 | 36.839 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135397 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5781 | 37.191 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135396 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6120 | 39.608 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135395 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6767 | 41.318 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135394 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7030 | 38.45 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135393 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6171 | 38.794 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135392 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4595 | 43.917 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135391 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5588 | 39.817 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135390 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6103 | 35.999 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135389 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 2701 | 39.837 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135388 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5641 | 41.234 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135387 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5507 | 37.734 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135386 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6317 | 37.185 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135385 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5229 | 39.855 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135384 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6024 | 41.766 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135383 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5211 | 42.564 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135382 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5680 | 42.975 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135381 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5794 | 43.355 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135380 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5844 | 40.264 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135379 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6067 | 39.229 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135378 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6031 | 39.894 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135377 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4914 | 39.479 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135376 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6207 | 41.759 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135375 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5869 | 38.661 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135374 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6176 | 39.33 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135373 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6354 | 37.866 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135372 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5799 | 37.248 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135371 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6130 | 42.202 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135370 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5904 | 37.127 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135369 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6121 | 39.258 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135368 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6125 | 38.024 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135367 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5639 | 39.901 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135366 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5472 | 39.072 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135365 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5986 | 34.781 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135364 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4660 | 38.627 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135363 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4741 | 42.544 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135362 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7089 | 37.452 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135361 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6244 | 41.672 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135360 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5850 | 41.026 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135359 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6090 | 38.654 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135358 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4898 | 39.343 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135357 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5559 | 33.927 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135356 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6039 | 33.764 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135355 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5650 | 40.106 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135354 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4878 | 38.294 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135353 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6230 | 36.998 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135352 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6227 | 43.311 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135351 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5393 | 48.266 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135350 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4776 | 40.096 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135349 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4660 | 42.983 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135348 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5965 | 43.319 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135347 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5471 | 37.525 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135346 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5834 | 42.235 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135345 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5391 | 37.006 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135344 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4973 | 40.157 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135343 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6257 | 38.932 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135342 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5470 | 45.393 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135341 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6618 | 37.247 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135340 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6233 | 41.393 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135339 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6267 | 41.695 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135338 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4868 | 47.453 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135337 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5060 | 39.921 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135336 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7047 | 38.754 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135335 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6269 | 40.916 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135334 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5022 | 40.98 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135333 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6100 | 37.77 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135332 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6214 | 40.2 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135331 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 39.261 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135330 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4549 | 36.645 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135329 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4767 | 41.997 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135328 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4846 | 44.243 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135327 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6259 | 35.948 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135326 | Microvirus sp. | Microvirus sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4681 | 39.885 | Microvirus | Unclassified | Microviridae | Unspecified |
| MT135314 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7970 | 44.743 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135313 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4631 | 41.892 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135312 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4224 | 36.458 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135311 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5044 | 43.438 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135310 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5368 | 40.853 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135309 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5968 | 39.796 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135308 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4976 | 43.991 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135307 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5891 | 39.823 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135306 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4986 | 41.757 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135305 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4953 | 44.518 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135304 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5067 | 37.557 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135303 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4961 | 41.645 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135302 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4520 | 44.181 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135301 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5753 | 41.179 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135300 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4174 | 44.777 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135299 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5124 | 42.428 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135298 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5763 | 39.979 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135297 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4523 | 45.191 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135296 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5466 | 43.725 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135295 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5544 | 36.923 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135294 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5128 | 41.829 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135293 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4916 | 43.938 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135292 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4914 | 44.322 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135291 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4961 | 41.363 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135290 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5424 | 44.285 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135289 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4515 | 44.784 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135288 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5167 | 41.01 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135287 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5617 | 40.591 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135286 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4425 | 44.249 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135285 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4531 | 44.405 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135284 | Inovirus sp. | Inovirus sp. unclassified Inovirus Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5343 | 39.678 | Inovirus | Unclassified | Inoviridae | Unspecified |
| MT135282 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4857 | 37.039 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135281 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4869 | 40.049 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135280 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3671 | 40.861 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135279 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4347 | 42.19 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135278 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5035 | 45.879 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135277 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5565 | 44.241 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135276 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5445 | 40.826 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135275 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5341 | 42.183 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135274 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4935 | 39.433 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135273 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5038 | 45.494 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135272 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5308 | 43.915 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135271 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4811 | 39.597 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135270 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4855 | 41.895 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135269 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4408 | 37.024 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135268 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5539 | 45.405 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135267 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4784 | 39.883 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135266 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4270 | 40.07 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135265 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4717 | 40.449 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135264 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5025 | 41.791 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135263 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4896 | 44.914 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135262 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4906 | 39.421 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135261 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5054 | 43.906 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135260 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4920 | 45.366 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135259 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5642 | 45.604 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135258 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5381 | 48.114 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135257 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5202 | 45.483 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135256 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4387 | 40.141 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT135255 | Gokushovirinae sp. | Gokushovirinae sp. unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5316 | 42.4 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MW021764 | Klebsiella phage P528 | Klebsiella phage P528 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51895 | 51.344 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MT664721 | Escherichia phage vB_EcoM_APEC | Escherichia phage vB_EcoM_APEC Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35832 | 41.287 | Unclassified | Unclassified | Podoviridae | Escherichia |
| MW357609 | Phage vB_SabS_Sds2 | Phage vB_SabS_Sds2 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114770 | 40.261 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MN939540 | Listeria phage vB_Liva_VAfA18 | Listeria phage vB_Liva_VAfA18 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134508 | 35.967 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MN939539 | Listeria phage vB_Lino_VEfB7 | Listeria phage vB_Lino_VEfB7 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135461 | 36.114 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MN101224 | Klebsiella phage KMI10 | Klebsiella phage KMI10 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59583 | 41.948 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN101223 | Klebsiella phage KMI9 | Klebsiella phage KMI9 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168074 | 41.68 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN101222 | Klebsiella phage KMI8 | Klebsiella phage KMI8 unclassified Drexlerviridae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52904 | 49.295 | Unclassified | Unclassified | Drexlerviridae | Klebsiella |
| MN101221 | Klebsiella phage KMI7 | Klebsiella phage KMI7 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131200 | 41.947 | Unclassified | Tevenvirinae | Myoviridae | Klebsiella |
| MN101220 | Klebsiella phage KMI6 | Klebsiella phage KMI6 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43883 | 53.634 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN101219 | Klebsiella phage KMI5 | Klebsiella phage KMI5 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43735 | 54.023 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN101218 | Klebsiella phage KMI4 | Klebsiella phage KMI4 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40622 | 52.733 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN101217 | Klebsiella phage KMI3 | Klebsiella phage KMI3 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43746 | 54.019 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN101216 | Klebsiella phage KMI2 | Klebsiella phage KMI2 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41023 | 52.624 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN052874 | Klebsiella phage KMI1 | Klebsiella phage KMI1 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37414 | 52.812 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| NC_053015 | Pectobacterium phage MA12 | Pectobacterium phage MA12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58573 | 54.535 | Unclassified | Unclassified | Siphoviridae | Pectobacterium |
| NC_053014 | Pectobacterium phage MA11 | Pectobacterium phage MA11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55830 | 54.542 | Unclassified | Unclassified | Siphoviridae | Pectobacterium |
| NC_053012 | Serratia phage JS26 | Serratia phage JS26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63971 | 57.013 | Unclassified | Unclassified | Siphoviridae | Serratia |
| NC_053011 | Erwinia phage pEp_SNUABM_08 | Erwinia phage pEp_SNUABM_08 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62716 | 57.244 | Unclassified | Unclassified | Siphoviridae | Erwinia |
| NC_052989 | Cronobacter phage JC01 | Cronobacter phage JC01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61736 | 58.515 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| NC_052983 | Escherichia phage E21 | Escherichia phage E21 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58536 | 46.898 | Unclassified | Unclassified | Podoviridae | Escherichia |
| NC_052982 | Proteus phage P16-2532 | Proteus phage P16-2532 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58931 | 46.712 | Unclassified | Unclassified | Siphoviridae | Proteus |
| NC_052981 | Proteus phage PM87 | Proteus phage PM87 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59128 | 46.729 | Unclassified | Unclassified | Siphoviridae | Proteus |
| NC_052979 | Providencia phage Kokobel1 | Providencia phage Kokobel1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59837 | 48.916 | Unclassified | Unclassified | Siphoviridae | Providencia |
| NC_052977 | Bacteriophage Phobos | Bacteriophage Phobos Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56734 | 63.324 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_052976 | Pseudomonas phage PspYZU01 | Pseudomonas phage PspYZU01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58279 | 63.122 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_052975 | Klebsiella phage YMC16/01/N133_KPN_BP | Klebsiella phage YMC16/01/N133_KPN_BP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58387 | 58.88 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| NC_052973 | Xylella phage Bacata | Xylella phage Bacata unclassified Sanovirus Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56232 | 62.95 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| NC_052972 | Ruegeria phage vB_RpoS-V7 | Ruegeria phage vB_RpoS-V7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59573 | 64.123 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| NC_052971 | Ruegeria phage vB_RpoS-V11 | Ruegeria phage vB_RpoS-V11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59549 | 64.025 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| NC_052970 | Ruegeria phage vB_RpoS-V18 | Ruegeria phage vB_RpoS-V18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59111 | 64.029 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| NC_052969 | Ruegeria phage vB_RpoS-V16 | Ruegeria phage vB_RpoS-V16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61382 | 63.641 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| NC_052968 | Synechococcus virus S-ESS1 | Synechococcus virus S-ESS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60362 | 60.904 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_052965 | Pseudomonas phage ZC01 | Pseudomonas phage ZC01 unclassified Abidjanvirus Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57061 | 63.381 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_052964 | Pseudomonas phage Epa5 | Pseudomonas phage Epa5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64096 | 62.247 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW507129 | Gordonia phage Ennea | Gordonia phage Ennea Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67632 | 65.669 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MW353172 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 18095 | 24.548 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353171 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 23035 | 23.108 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353170 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 30949 | 23.878 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353169 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 59948 | 22.765 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353168 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 33493 | 23.596 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353167 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 31160 | 23.354 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW353166 | Mycoplasma phage sp. | Mycoplasma phage sp. unclassified bacterial viruses Viruses | 33469 | 23.586 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| MW462191 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 103248 | 60.622 | Unclassified | Unclassified | Unclassified | Unspecified |
| MW507132 | Gordonia phage Ekhein | Gordonia phage Ekhein unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76612 | 58.942 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MW507128 | Mycobacterium phage Phoebe | Mycobacterium phage Phoebe unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50876 | 64.005 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW507126 | Microbacterium phage Shocker | Microbacterium phage Shocker Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45431 | 63.043 | Unclassified | Unclassified | Podoviridae | Microbacterium |
| MW507124 | Microbacterium phage BlueRugrat | Microbacterium phage BlueRugrat unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54124 | 60.247 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW354667 | Bacillus phage BSTP3 | Bacillus phage BSTP3 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153753 | 42.104 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_052941 | Microbacterium phage Efeko | Microbacterium phage Efeko Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17491 | 68.59 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052940 | Microbacterium phage BonaeVitae | Microbacterium phage BonaeVitae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17451 | 68.22 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052939 | Microbacterium phage Quaker | Microbacterium phage Quaker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17450 | 68.636 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052938 | Microbacterium phage PoRanda | Microbacterium phage PoRanda Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17510 | 69.109 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052937 | Microbacterium phage Noelani | Microbacterium phage Noelani Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17349 | 68.154 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052936 | Microbacterium phage Nobel | Microbacterium phage Nobel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17049 | 68.989 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052935 | Microbacterium phage Kaijohn | Microbacterium phage Kaijohn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17426 | 68.759 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052934 | Microbacterium phage Bri160 | Microbacterium phage Bri160 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17442 | 68.559 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052933 | Microbacterium phage Azizam | Microbacterium phage Azizam Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17385 | 68.749 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| NC_052932 | Microbacterium phage PaoPu | Microbacterium phage PaoPu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17362 | 68.529 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MW570842 | Mycobacterium phage phiT45-1 | Mycobacterium phage phiT45-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43407 | 64.669 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MT104474 | Pseudomonas phage MR14 | Pseudomonas phage MR14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46593 | 58.668 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MW476489 | Lactobacillus phage vB_Lga_AB1 | Lactobacillus phage vB_Lga_AB1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40899 | 36.03 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| CP069348 | Clostridioides phage pCD1602_4 | Clostridioides phage pCD1602_4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132519 | 26.761 | Unclassified | Unclassified | Siphoviridae | Clostridioides |
| NC_052663 | Escherichia phage VEcB | Escherichia phage VEcB unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147702 | 37.556 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| NC_052662 | Escherichia phage Mt1B1_P17 | Escherichia phage Mt1B1_P17 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151202 | 38.917 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052661 | Escherichia phage ukendt | Escherichia phage ukendt unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150947 | 39.025 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052660 | Escherichia phage tuntematon | Escherichia phage tuntematon unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150473 | 39.105 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052659 | Escherichia phage nieznany | Escherichia phage nieznany unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144998 | 39.056 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052658 | Escherichia phage nepoznato | Escherichia phage nepoznato unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151514 | 38.877 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052657 | Escherichia phage muut | Escherichia phage muut unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146307 | 37.365 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| NC_052656 | Escherichia phage anhysbys | Escherichia phage anhysbys unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149335 | 39.116 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052655 | Escherichia phage alia | Escherichia phage alia unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147009 | 37.511 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| NC_052654 | Escherichia phage vB_EcoM-Ro121c4YLVW | Escherichia phage vB_EcoM-Ro121c4YLVW unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151671 | 39.099 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_052653 | Escherichia phage vB_vPM_PD06 | Escherichia phage vB_vPM_PD06 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149506 | 37.652 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| NC_052652 | Escherichia phage vB_EcoM_PHB05 | Escherichia phage vB_EcoM_PHB05 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147659 | 37.536 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| LC592711 | Burkholderia phage FLC6 | Burkholderia phage FLC6 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 227105 | 52.009 | Chiangmaivirus | Unclassified | Myoviridae | Burkholderia |
| MW477799 | Salmonella phage vB_Se_STGO-35-1 | Salmonella phage vB_Se_STGO-35-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47483 | 46.488 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MT880872 | Staphylococcus phage PhiSepi-HH3 | Staphylococcus phage PhiSepi-HH3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147057 | 30.811 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT880871 | Staphylococcus phage PI-Sepi-HH2 | Staphylococcus phage PI-Sepi-HH2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36164 | 29.314 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT880870 | Staphylococcus phage PhiSepi-HH1 | Staphylococcus phage PhiSepi-HH1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34053 | 28.435 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MN939408 | Enterococcus phage vB_EfaS_TV16 | Enterococcus phage vB_EfaS_TV16 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58127 | 40.062 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MH807812 | Dickeya phage Kamild | Dickeya phage Kamild unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152612 | 49.168 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MH807820 | Dickeya phage Coodle | Dickeya phage Coodle unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152515 | 49.132 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MH393889 | Clostridium phage susfortuna | Clostridium phage susfortuna Clostridium virus susfortuna Susfortunavirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19046 | 29.224 | Susfortunavirus | Unclassified | Guelinviridae | Clostridium |
| MH807817 | Dickeya phage Mysterion | Dickeya phage Mysterion Dickeya virus Mysterion Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40651 | 48.11 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH807815 | Dickeya phage Luksen | Dickeya phage Luksen Dickeya virus Luksen Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40729 | 48.214 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH807813 | Dickeya phage Katbat | Dickeya phage Katbat Dickeya virus Katbat Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40441 | 48.038 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH807810 | Dickeya phage Dagda_B1 | Dickeya phage Dagda_B1 unclassified Aarhusvirus Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40484 | 48.113 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH059632 | Dickeya phage Dagda | Dickeya phage Dagda Dickeya virus Dagda Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40484 | 48.113 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH059639 | Dickeya phage Ninurta | Dickeya phage Ninurta Dickeya virus Ninurta Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39699 | 50.986 | Ningirsuvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH059637 | Pectobacterium phage Jarilo | Pectobacterium phage Jarilo Pectobacterium virus Jarilo Jarilovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40552 | 50.094 | Jarilovirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_014459 | Mycobacterium phage CrimD | Mycobacterium phage CrimD Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59798 | 66.877 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028978 | Mycobacterium phage Murucutumbu | Mycobacterium phage Murucutumbu unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60609 | 66.693 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028922 | Mycobacterium phage Cambiare | Mycobacterium phage Cambiare Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45161 | 68.798 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_028861 | Mycobacterium phage FlagStaff | Mycobacterium phage FlagStaff Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44576 | 68.521 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_028791 | Mycobacterium phage MOOREtheMARYer | Mycobacterium phage MOOREtheMARYer Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 68.581 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_023747 | Mycobacterium phage BarrelRoll | Mycobacterium phage BarrelRoll unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59672 | 66.616 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023705 | Mycobacterium virus Liefie | Mycobacterium virus Liefie Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41650 | 66.783 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| EU861004 | Staphylococcus phage vB_SauS-phiIPLA88 | Staphylococcus phage vB_SauS-phiIPLA88 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42526 | 34.913 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MW452562 | Nocardia phage NC1 | Nocardia phage NC1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68913 | 65.557 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| MW460250 | Staphylococcus phage SAOMS1 | Staphylococcus phage SAOMS1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140135 | 30.366 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW460249 | Pseudomonas phage Itty13 | Pseudomonas phage Itty13 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61818 | 65.515 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MW460247 | Burkholderia phage BCSR129 | Burkholderia phage BCSR129 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66147 | 58.417 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MW460246 | Burkholderia phage BCSR52 | Burkholderia phage BCSR52 unclassified Bcepfunavirus Bcepfunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70038 | 51.451 | Bcepfunavirus | Unclassified | Myoviridae | Burkholderia |
| MW460245 | Burkholderia phage BCSR5 | Burkholderia phage BCSR5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 227351 | 54.737 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MW354670 | Bacillus phage BSTP6 | Bacillus phage BSTP6 unclassified Salasvirus Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19367 | 39.784 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| MW476490 | Lactobacillus phage vB_Lcr_AB1 | Lactobacillus phage vB_Lcr_AB1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47292 | 40.654 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| MW477480 | Bacillus phage Whiting18 | Bacillus phage Whiting18 unclassified Salasvirus Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19548 | 39.636 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| MT899443 | Liberibacter phage P-PA19-2 | Liberibacter phage P-PA19-2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31505 | 38.886 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| MN807247 | Mycobacterium phage Chupacabra | Mycobacterium phage Chupacabra unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50836 | 65.202 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586037 | Mycobacterium phage GooberAzure | Mycobacterium phage GooberAzure unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75129 | 62.941 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KX756439 | Mycobacterium phage PhancyPhin | Mycobacterium phage PhancyPhin unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42454 | 66.133 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MW344765 | Halorubrum virus Humcor1 | Halorubrum virus Humcor1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46474 | 62.452 | Unclassified | Unclassified | Siphoviridae | Halorubrum |
| MW218148 | Staphylococcus phage vB_StaM_SA1 | Staphylococcus phage vB_StaM_SA1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 260727 | 26.833 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MT627482 | Enterococcus phage vB_EFaS_TV217 | Enterococcus phage vB_EFaS_TV217 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41486 | 34.578 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT133563 | Pseudomonas phage billy | Pseudomonas phage billy unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65858 | 54.953 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT133562 | Pseudomonas phage willy | Pseudomonas phage willy unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65233 | 55.595 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT133561 | Pseudomonas phage goonie | Pseudomonas phage goonie unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65201 | 55.58 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT133560 | Pseudomonas phage fnug | Pseudomonas phage fnug unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 278899 | 36.856 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MT119378 | Pseudomonas phage datas | Pseudomonas phage datas unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60746 | 54.807 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119377 | Pseudomonas phage crassa | Pseudomonas phage crassa unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66295 | 55.218 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119376 | Pseudomonas phage chunk | Pseudomonas phage chunk unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66231 | 55.63 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119375 | Pseudomonas phage chumba | Pseudomonas phage chumba unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65966 | 54.98 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119374 | Pseudomonas phage antinowhere | Pseudomonas phage antinowhere unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65852 | 55.189 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119372 | Pseudomonas phage cory | Pseudomonas phage cory unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66395 | 55.204 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119371 | Pseudomonas phage oldone | Pseudomonas phage oldone unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45313 | 52.504 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT119370 | Pseudomonas phage steven | Pseudomonas phage steven unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65632 | 54.865 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119369 | Pseudomonas phage sortsol | Pseudomonas phage sortsol unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66445 | 55.672 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119368 | Pseudomonas phage shane | Pseudomonas phage shane unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66467 | 55.643 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119367 | Pseudomonas phage misfit | Pseudomonas phage misfit unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65887 | 54.933 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119366 | Pseudomonas phage jett | Pseudomonas phage jett unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66262 | 55.569 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119365 | Pseudomonas phage goodold | Pseudomonas phage goodold unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66226 | 55.61 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119364 | Pseudomonas phage elmo | Pseudomonas phage elmo unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65276 | 54.976 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119363 | Pseudomonas phage debbie | Pseudomonas phage debbie unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66259 | 55.666 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT119362 | Pseudomonas phage clash | Pseudomonas phage clash unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44912 | 52.089 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT119361 | Enterococus phage vipetofem | Enterococus phage vipetofem Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85371 | 30.291 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT119360 | Enterococcus phage nattely | Enterococcus phage nattely Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85669 | 30.205 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT119359 | Enterococcus phage heks | Enterococcus phage heks unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39708 | 34.983 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT074476 | Salmonella phage kage | Salmonella phage kage unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157658 | 44.482 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074460 | Salmonella phage bux | Salmonella phage bux unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112486 | 39.961 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074484 | Salmonella phage celemicas | Salmonella phage celemicas unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43193 | 49.767 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074483 | Salmonella phage skrot | Salmonella phage skrot unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42879 | 49.726 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074482 | Salmonella phage horsemountain | Salmonella phage horsemountain unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42375 | 49.638 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074481 | Salmonella phage sidste | Salmonella phage sidste unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42124 | 50 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074473 | Salmonella phage blauehaus | Salmonella phage blauehaus unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41993 | 49.651 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074468 | Salmonella phage rutana | Salmonella phage rutana unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114544 | 39.936 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074467 | Salmonella phage falkor | Salmonella phage falkor unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114211 | 39.765 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074464 | Salmonella phage bobsandoy | Salmonella phage bobsandoy unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113461 | 39.867 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074463 | Salmonella phage gmork | Salmonella phage gmork unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113259 | 39.89 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074462 | Salmonella phage templet | Salmonella phage templet unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41598 | 49.474 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074461 | Salmonella phage smaug | Salmonella phage smaug unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112590 | 39.756 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074456 | Salmonella phage polluks | Salmonella phage polluks unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111599 | 40.013 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074455 | Salmonella phage beppo | Salmonella phage beppo unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111322 | 40.099 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074454 | Salmonella phage ende | Salmonella phage ende unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111297 | 39.969 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074453 | Salmonella phage misterkot | Salmonella phage misterkot unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110937 | 39.778 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074452 | Salmonella phage vaffelhjerte | Salmonella phage vaffelhjerte unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110831 | 39.959 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074451 | Salmonella phage wast | Salmonella phage wast unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40748 | 49.811 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074447 | Salmonella phage phagemcphageface | Salmonella phage phagemcphageface unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 105405 | 39.926 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074446 | Salmonella phage triss | Salmonella phage triss unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53333 | 45.865 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074445 | Salmonella phage renfri | Salmonella phage renfri unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52952 | 45.894 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074444 | Salmonella phage emhyr | Salmonella phage emhyr unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52948 | 45.896 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074443 | Salmonella phage zoltan | Salmonella phage zoltan unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52933 | 45.866 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074442 | Salmonella phage ciri | Salmonella phage ciri unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52916 | 45.946 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074441 | Salmonella phage brunost | Salmonella phage brunost unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52821 | 46.012 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074438 | Salmonella phage rivia | Salmonella phage rivia unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52587 | 45.876 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074437 | Salmonella phage nenneke | Salmonella phage nenneke unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52576 | 45.914 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074435 | Salmonella phage brorfarstad | Salmonella phage brorfarstad unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52392 | 45.749 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074434 | Salmonella phage emiel | Salmonella phage emiel unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52333 | 45.9 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074432 | Salmonella phage dunkel | Salmonella phage dunkel unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43985 | 49.744 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074431 | Salmonella phage demigod | Salmonella phage demigod unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43605 | 50.146 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT074430 | Salmonella phage pink | Salmonella phage pink unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43203 | 49.714 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN850631 | Escherichia phage mio | Escherichia phage mio unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83259 | 39.059 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850628 | Escherichia phage bob | Escherichia phage bob unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45256 | 54.481 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MN850581 | Escherichia phage sortkaff | Escherichia phage sortkaff unclassified Sortsnevirus Sortsnevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42446 | 59.447 | Sortsnevirus | Unclassified | Podoviridae | Escherichia |
| MN850651 | Escherichia phage moskry | Escherichia phage moskry unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169410 | 37.637 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850650 | Escherichia phage jat | Escherichia phage jat unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44417 | 54.531 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MN850649 | Escherichia phage nomine | Escherichia phage nomine unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137991 | 43.65 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850648 | Escherichia phage anhysbys | Escherichia phage anhysbys unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149335 | 39.116 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MN850647 | Escherichia phage tootiki | Escherichia phage tootiki unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88257 | 39.004 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850646 | Escherichia phage nom | Escherichia phage nom unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136114 | 43.573 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850645 | Escherichia phage outra | Escherichia phage outra unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145482 | 37.448 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MN850644 | Escherichia phage ekra | Escherichia phage ekra unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87282 | 38.914 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850642 | Escherichia phage navn | Escherichia phage navn unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139427 | 43.582 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850639 | Escherichia phage radambza | Escherichia phage radambza unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86702 | 38.94 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850637 | Escherichia phage warpig | Escherichia phage warpig unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86106 | 38.985 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850636 | Escherichia phage dune | Escherichia phage dune unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88511 | 39.043 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850635 | Escherichia phage bumzen | Escherichia phage bumzen unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87360 | 39.075 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850634 | Escherichia phage tinuso | Escherichia phage tinuso unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50856 | 44.768 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850633 | Escherichia phage allfine | Escherichia phage allfine unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86963 | 39.035 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850632 | Escherichia phage alia | Escherichia phage alia unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147009 | 37.511 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MN850630 | Escherichia phage naam | Escherichia phage naam unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137129 | 43.699 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850629 | Escherichia phage mckay | Escherichia phage mckay unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44443 | 54.501 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MN850627 | Escherichia phage pangalan | Escherichia phage pangalan unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136944 | 43.687 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850626 | Escherichia phage nimi | Escherichia phage nimi unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137039 | 43.705 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850625 | Escherichia phage smaasur | Escherichia phage smaasur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41110 | 55.41 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850624 | Escherichia phage usur | Escherichia phage usur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41906 | 55.414 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850622 | Escherichia phage mobillu | Escherichia phage mobillu unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163063 | 37.681 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850621 | Escherichia phage dhabil | Escherichia phage dhabil unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165644 | 39.465 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850619 | Escherichia phage finno | Escherichia phage finno unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87544 | 38.935 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850618 | Escherichia phage tuntematon | Escherichia phage tuntematon unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150473 | 39.105 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MN850617 | Escherichia phage forsur | Escherichia phage forsur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42476 | 55.387 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850616 | Escherichia phage buks | Escherichia phage buks unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40308 | 49.749 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN850615 | Escherichia phage kvi | Escherichia phage kvi unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163673 | 40.47 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850614 | Escherichia phage adrianh | Escherichia phage adrianh unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88226 | 38.924 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850612 | Escherichia phage magaca | Escherichia phage magaca unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135826 | 43.644 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850611 | Escherichia phage nataliec | Escherichia phage nataliec unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89137 | 39.03 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850610 | Escherichia phage Andreotti | Escherichia phage Andreotti unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83391 | 39.212 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850609 | Escherichia phage dhaeg | Escherichia phage dhaeg unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170817 | 39.392 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850608 | Escherichia phage megetsur | Escherichia phage megetsur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42132 | 55.775 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850606 | Escherichia phage tuinn | Escherichia phage tuinn unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50505 | 44.718 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850605 | Escherichia phage fjerdesal | Escherichia phage fjerdesal unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87715 | 38.968 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850604 | Escherichia phage tunzivis | Escherichia phage tunzivis unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50596 | 44.573 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850603 | Escherichia phage pinkbiff | Escherichia phage pinkbiff unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88814 | 39.033 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850601 | Escherichia phage inny | Escherichia phage inny unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147483 | 37.394 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MN850599 | Escherichia phage lillemer | Escherichia phage lillemer unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5492 | 47.141 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| MN850598 | Escherichia phage nieznany | Escherichia phage nieznany unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144998 | 39.056 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MN850597 | Escherichia phage isim | Escherichia phage isim unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138289 | 43.644 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850595 | Escherichia phage naswa | Escherichia phage naswa unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138583 | 43.585 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850593 | Escherichia phage inoa | Escherichia phage inoa unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138710 | 43.593 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850592 | Escherichia phage aldrigsur | Escherichia phage aldrigsur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42379 | 55.726 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850590 | Escherichia phage moha | Escherichia phage moha unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168676 | 37.612 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850589 | Escherichia phage welsh | Escherichia phage welsh unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45207 | 54.598 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MN850587 | Escherichia phage mistaenkt | Escherichia phage mistaenkt unclassified Suspvirus Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86664 | 39.995 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| MN850586 | Escherichia phage orkinos | Escherichia phage orkinos unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49798 | 44.596 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850585 | Escherichia phage skuden | Escherichia phage skuden unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87263 | 38.885 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850584 | Escherichia phage arall | Escherichia phage arall unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145715 | 37.383 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MN850583 | Escherichia phage glasur | Escherichia phage glasur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42507 | 55.41 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850582 | Escherichia phage ityhuna | Escherichia phage ityhuna unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50768 | 44.658 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850580 | Escherichia phage momo | Escherichia phage momo unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88168 | 38.985 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850579 | Escherichia phage mogra | Escherichia phage mogra unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168724 | 37.686 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850578 | Escherichia phage nomo | Escherichia phage nomo unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137702 | 43.653 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850577 | Escherichia phage heid | Escherichia phage heid unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87590 | 39.042 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850576 | Escherichia phage ime | Escherichia phage ime unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137114 | 43.637 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MN850575 | Escherichia phage rolling | Escherichia phage rolling unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46017 | 54.197 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MN850574 | Escherichia phage kaaroe | Escherichia phage kaaroe unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163719 | 40.499 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN850573 | Escherichia phage muut | Escherichia phage muut unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146307 | 37.365 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MN850571 | Escherichia phage nepoznato | Escherichia phage nepoznato unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151514 | 38.877 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MN850570 | Escherichia phage mellemsur | Escherichia phage mellemsur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40770 | 55.816 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850568 | Escherichia phage altidsur | Escherichia phage altidsur unclassified Bonnellvirus Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42197 | 55.717 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MN850566 | Escherichia phage garuso | Escherichia phage garuso unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85798 | 38.944 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN850565 | Escherichia phage ukendt | Escherichia phage ukendt unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150947 | 39.025 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MN850564 | Escherichia phage humlepung | Escherichia phage humlepung unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85311 | 39.081 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN895436 | Escherichia phage teqsoen | Escherichia phage teqsoen unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166468 | 35.467 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN895435 | Escherichia phage teqhal | Escherichia phage teqhal unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168070 | 35.449 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK791318 | Escherichia phage Lilledu | Escherichia phage Lilledu unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5484 | 47.21 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| MK629526 | Escherichia phage Lilleen | Escherichia phage Lilleen unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5342 | 46.93 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| MK651787 | Escherichia phage Sortsne | Escherichia phage Sortsne Escherichia virus Sortsne Sortsnevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41912 | 59.973 | Sortsnevirus | Unclassified | Podoviridae | Escherichia |
| MK629529 | Escherichia phage Lilleto | Escherichia phage Lilleto unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5494 | 46.815 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| MK629528 | Escherichia phage Lidtsur | Escherichia phage Lidtsur Escherichia virus Lidtsur Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42291 | 54.633 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| MK629527 | Escherichia phage Lilleven | Escherichia phage Lilleven unclassified Alphatrevirus Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6090 | 44.45 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| MK629525 | Escherichia phage Lilleput | Escherichia phage Lilleput unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5490 | 47.049 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| MK599416 | Salmonella phage Akira | Salmonella phage Akira Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45367 | 46.022 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MK552105 | Escherichia phage Jahat_MG145 | Escherichia phage Jahat_MG145 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50984 | 45.718 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MK125140 | Enterococcus phage Nonaheksakonda | Enterococcus phage Nonaheksakonda unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41994 | 34.562 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK105855 | Escherichia phage Skarpretter | Escherichia phage Skarpretter Escherichia virus Skarpretter Skarprettervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42042 | 55.82 | Skarprettervirus | Unclassified | Podoviridae | Escherichia |
| MH362766 | Escherichia phage Halfdan | Escherichia phage Halfdan unclassified Lokivirus Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42858 | 53.666 | Lokivirus | Unclassified | Siphoviridae | Escherichia |
| MT829326 | Enterococcus phage FX417 | Enterococcus phage FX417 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41061 | 34.661 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT661598 | Enterococcus phage vB_EFaS_TV54 | Enterococcus phage vB_EFaS_TV54 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42438 | 34.568 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT661597 | Enterococcus phage vB_EFaS_TV51 | Enterococcus phage vB_EFaS_TV51 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41821 | 34.664 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT074440 | Salmonella phage assan | Salmonella phage assan unclassified Astrithrvirus Astrithrvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11578 | 39.826 | Astrithrvirus | Unclassified | Podoviridae | Salmonella |
| MT074433 | Salmonella phage slyngel | Salmonella phage slyngel Salmonella virus slyngel Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51048 | 44.012 | Hanrivervirus | Tempevirinae | Drexlerviridae | Salmonella |
| MN850643 | Escherichia phage tiwna | Escherichia phage tiwna Escherichia virus tiwna Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51014 | 44.631 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850641 | Escherichia phage tonijn | Escherichia phage tonijn Escherichia virus tonijn Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51627 | 44.637 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850640 | Escherichia phage herni | Escherichia phage herni Escherichia virus herni Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50971 | 44.088 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850638 | Escherichia phage tunus | Escherichia phage tunus Escherichia virus tunus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51111 | 44.799 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850623 | Escherichia phage sortsyn | Escherichia phage sortsyn unclassified Murrayvirus Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42116 | 59.011 | Murrayvirus | Unclassified | Siphoviridae | Escherichia |
| MN850620 | Escherichia phage atuna | Escherichia phage atuna Escherichia virus atuna Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50732 | 44.599 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850613 | Escherichia phage tonnikala | Escherichia phage tonnikala Escherichia virus tonnikala Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51277 | 44.788 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850607 | Escherichia phage egaa | Escherichia phage egaa Escherichia virus egaa Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51643 | 44.128 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850602 | Escherichia phage damhaus | Escherichia phage damhaus Escherichia virus damhaus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51154 | 44.145 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850600 | Escherichia phage haarsle | Escherichia phage haarsle Escherichia virus haarsle Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48613 | 44.038 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850596 | Escherichia phage tonn | Escherichia phage tonn Escherichia virus tonn Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51012 | 44.472 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850594 | Escherichia phage flopper | Escherichia phage flopper Escherichia virus flopper Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52092 | 44.203 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MN850591 | Escherichia phage aalborv | Escherichia phage aalborv Escherichia virus aalborv Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46660 | 43.89 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850588 | Escherichia phage sortregn | Escherichia phage sortregn unclassified Murrayvirus Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38200 | 59.322 | Murrayvirus | Unclassified | Siphoviridae | Escherichia |
| MN850572 | Escherichia phage aaroes | Escherichia phage aaroes Escherichia virus aaroes Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51662 | 44.125 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850569 | Escherichia phage vojen | Escherichia phage vojen Escherichia virus vojen Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50709 | 44.13 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN850567 | Escherichia phage grams | Escherichia phage grams Escherichia virus grams Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49530 | 44.099 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN895438 | Escherichia phage teqdroes | Escherichia phage teqdroes Escherichia virus teqdroes Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166833 | 35.438 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN895437 | Escherichia phage teqskov | Escherichia phage teqskov Escherichia virus teqskov Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165017 | 35.359 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN895434 | Escherichia phage teqhad | Escherichia phage teqhad Escherichia virus teqhad Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167892 | 35.327 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN029011 | Pseudomonas phage Iggy | Pseudomonas phage Iggy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60628 | 56.51 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK672798 | Escherichia phage Skure | Escherichia phage Skure unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59463 | 44.559 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MK125141 | Salmonella phage Lumpael | Salmonella phage Lumpael Salmonella virus Lumpael Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41126 | 59.466 | Murrayvirus | Unclassified | Siphoviridae | Salmonella |
| MK080312 | Salmonella phage Astrid | Salmonella phage Astrid unclassified Astrithrvirus Astrithrvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11713 | 39.751 | Astrithrvirus | Unclassified | Podoviridae | Salmonella |
| MW286268 | Pseudomonas phage Strit | Pseudomonas phage Strit Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41373 | 58.081 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MW286267 | Pseudomonas phage Misse | Pseudomonas phage Misse Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41342 | 58.122 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MW286266 | Pseudomonas phage Bertil | Pseudomonas phage Bertil Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41374 | 58.073 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MN954399 | Vibrio phage VPT02 | Vibrio phage VPT02 Vipunavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120547 | 39.894 | Vipunavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MW421582 | Flavobacterium phage FPSV-S8 | Flavobacterium phage FPSV-S8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35187 | 37.846 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW419910 | Enterobacter phage ATCEA23 | Enterobacter phage ATCEA23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43692 | 56.388 | Unclassified | Unclassified | Siphoviridae | Enterobacter |
| MW331437 | Escherichia phage phiWAO78-1 | Escherichia phage phiWAO78-1 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44551 | 50.829 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MW239157 | Klebsiella phage vB_KpnM_17-11 | Klebsiella phage vB_KpnM_17-11 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165894 | 39.53 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MT152698 | Halorubrum phage Hardycor1 | Halorubrum phage Hardycor1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45142 | 67.799 | Unclassified | Unclassified | Siphoviridae | Halorubrum |
| MW411578 | Salmonella phage SP_Thor | Salmonella phage SP_Thor unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107183 | 40.091 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW421584 | Flavobacterium phage FPSV-D3 | Flavobacterium phage FPSV-D3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35195 | 37.841 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW421583 | Flavobacterium phage FPSV-F8 | Flavobacterium phage FPSV-F8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35187 | 37.846 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW421581 | Flavobacterium phage FPSV-S4 | Flavobacterium phage FPSV-S4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35187 | 37.826 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW421580 | Flavobacterium phage FPSV-D7 | Flavobacterium phage FPSV-D7 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.801 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW421579 | Flavobacterium phage FPSV-D1 | Flavobacterium phage FPSV-D1 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.807 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW421578 | Flavobacterium phage FPSV-D41 | Flavobacterium phage FPSV-D41 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.806 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW421577 | Flavobacterium phage FPSV-S30 | Flavobacterium phage FPSV-S30 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89814 | 26.946 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW421576 | Flavobacterium phage FPSV-D17 | Flavobacterium phage FPSV-D17 unclassified Unahavirus Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47383 | 31.741 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| MW364975 | Staphylococcus virus vB_SepS_E72 | Staphylococcus virus vB_SepS_E72 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44592 | 34.54 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MW364974 | Staphylococcus virus vB_SepS_459 | Staphylococcus virus vB_SepS_459 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42498 | 34.719 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MW364973 | Staphylococcus virus vB_SepS_456 | Staphylococcus virus vB_SepS_456 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43266 | 34.699 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MW364972 | Staphylococcus virus vB_SepS_48 | Staphylococcus virus vB_SepS_48 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42460 | 34.769 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MW364971 | Staphylococcus virus vB_SepS_27 | Staphylococcus virus vB_SepS_27 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42935 | 34.587 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| MW435859 | Microbacterium phage WildNOut | Microbacterium phage WildNOut unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41796 | 63.355 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MW435854 | Mycobacterium phage Psycho | Mycobacterium phage Psycho unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62110 | 65.347 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW394391 | Klebsiella phage vB_KpM_FBKp24 | Klebsiella phage vB_KpM_FBKp24 unclassified bacterial viruses Viruses | 307210 | 45.131 | Unclassified | Unclassified | Unclassified | Klebsiella |
| MW394390 | Klebsiella phage vB_KpM_FBKp34 | Klebsiella phage vB_KpM_FBKp34 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141376 | 36.026 | Unclassified | Vequintavirinae | Myoviridae | Klebsiella |
| MW394389 | Klebsiella phage vB_KpP_FBKp16 | Klebsiella phage vB_KpP_FBKp16 unclassified Molineuxvirinae Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44010 | 51.872 | Unclassified | Molineuxvirinae | Autographiviridae | Klebsiella |
| MW394388 | Klebsiella phage vB_KpP_FBKp27 | Klebsiella phage vB_KpP_FBKp27 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76339 | 44.247 | Unclassified | Unclassified | Schitoviridae | Klebsiella |
| MW367898 | Proteus phage SJ_PmiM | Proteus phage SJ_PmiM unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164100 | 31.074 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| MW423798 | Salmonella phage vB STyj5-1 | Salmonella phage vB STyj5-1 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110592 | 39.873 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW423797 | Salmonella phage vB Seyj1-1 | Salmonella phage vB Seyj1-1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88795 | 38.733 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW385300 | Providencia phage PSTRCR_121 | Providencia phage PSTRCR_121 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165776 | 34.664 | Unclassified | Tevenvirinae | Myoviridae | Providencia |
| MW430345 | Staphylococcus phage ZCSS1 | Staphylococcus phage ZCSS1 unclassified Sciuriunavirus Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139743 | 31.775 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW349138 | Erwinia phage pEa_SNUABM_27 | Erwinia phage pEa_SNUABM_27 unclassified Loessnervirus Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53014 | 44.068 | Loessnervirus | Cleopatravirinae | Chaseviridae | Erwinia |
| MW344054 | Vibrio phage Gary | Vibrio phage Gary unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128602 | 40.195 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MW221967 | Staphylococcus phage PMBT9 | Staphylococcus phage PMBT9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41623 | 36.192 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MW175491 | Pseudomonas phage PN09 | Pseudomonas phage PN09 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99229 | 48.157 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| NC_049978 | IAS virus | IAS virus crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99915 | 31.948 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW419775 | Bacillus phage WhyPhy | Bacillus phage WhyPhy unclassified Bundooravirus Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18642 | 34.948 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| MT416090 | Pseudomonas phage vB_Pae_AM.P2 | Pseudomonas phage vB_Pae_AM.P2 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73308 | 53.448 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MN988502 | Rhizobium phage RHph_Y3_56_1 | Rhizobium phage RHph_Y3_56_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40181 | 58.508 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988561 | Rhizobium phage RHph_TM61 | Rhizobium phage RHph_TM61 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 202964 | 41.151 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988560 | Rhizobium phage RHph_TM21B | Rhizobium phage RHph_TM21B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153953 | 41.961 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988559 | Rhizobium phage RHph_TM39 | Rhizobium phage RHph_TM39 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189896 | 41.402 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988558 | Rhizobium phage RHph_Y1_11 | Rhizobium phage RHph_Y1_11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100469 | 59.087 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988557 | Rhizobium phage RHph_Y52 | Rhizobium phage RHph_Y52 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69157 | 56.837 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988556 | Rhizobium phage RHph_I1_23 | Rhizobium phage RHph_I1_23 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43424 | 58.293 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988555 | Rhizobium phage RHph_I1_6 | Rhizobium phage RHph_I1_6 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77174 | 46.277 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MN988554 | Rhizobium phage RHph_I72 | Rhizobium phage RHph_I72 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55716 | 58.768 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988553 | Rhizobium phage RHph_I34 | Rhizobium phage RHph_I34 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154143 | 48.494 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988552 | Rhizobium phage RHph_I4 | Rhizobium phage RHph_I4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108441 | 58.5 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988551 | Rhizobium phage RHph_N3_8 | Rhizobium phage RHph_N3_8 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18277 | 50.665 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988550 | Rhizobium phage RHph_N39 | Rhizobium phage RHph_N39 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4781 | 56.076 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988549 | Rhizobium phage RHph_Y2_17_2 | Rhizobium phage RHph_Y2_17_2 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54438 | 56.457 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988548 | Rhizobium phage RHph_Y2_17_1 | Rhizobium phage RHph_Y2_17_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 117010 | 59.871 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988547 | Rhizobium phage RHph_Y2_11 | Rhizobium phage RHph_Y2_11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 117010 | 59.869 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988546 | Rhizobium phage RHph_Y2_4 | Rhizobium phage RHph_Y2_4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69421 | 57.269 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988545 | Rhizobium phage RHph_Y1_1 | Rhizobium phage RHph_Y1_1 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54438 | 56.457 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988544 | Rhizobium phage RHph_Y21 | Rhizobium phage RHph_Y21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69214 | 56.851 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988543 | Rhizobium phage RHph_N2_6 | Rhizobium phage RHph_N2_6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116666 | 59.765 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988542 | Rhizobium phage RHph_I3_18 | Rhizobium phage RHph_I3_18 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43069 | 58.392 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988541 | Rhizobium phage RHph_I65 | Rhizobium phage RHph_I65 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54511 | 58.711 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988540 | Rhizobium phage RHph_I42 | Rhizobium phage RHph_I42 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193262 | 53.516 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988539 | Rhizobium phage RHph_I20 | Rhizobium phage RHph_I20 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41785 | 60.098 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988538 | Rhizobium phage RHph_Y25 | Rhizobium phage RHph_Y25 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43040 | 58.685 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988537 | Rhizobium phage RHph_TM3_3_10 | Rhizobium phage RHph_TM3_3_10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67842 | 56.49 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988536 | Rhizobium phage RHph_N65 | Rhizobium phage RHph_N65 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48682 | 60.753 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988535 | Rhizobium phage RHph_N42 | Rhizobium phage RHph_N42 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193452 | 53.439 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988534 | Rhizobium phage RHph_N34 | Rhizobium phage RHph_N34 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153811 | 48.957 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988533 | Rhizobium phage RHph_N2 | Rhizobium phage RHph_N2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44194 | 59.094 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988532 | Rhizobium phage RHph_I1_9 | Rhizobium phage RHph_I1_9 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154206 | 48.494 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988531 | Rhizobium phage RHph_Y5A | Rhizobium phage RHph_Y5A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69421 | 57.265 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988530 | Rhizobium phage RHph_N17 | Rhizobium phage RHph_N17 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54317 | 56.132 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988529 | Rhizobium phage RHph_I38 | Rhizobium phage RHph_I38 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46695 | 49.149 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| MN988528 | Rhizobium phage RHph_N37 | Rhizobium phage RHph_N37 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46450 | 49.277 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| MN988527 | Rhizobium phage RHph_N3_19 | Rhizobium phage RHph_N3_19 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68656 | 42.633 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988526 | Rhizobium phage RHph_N3_2 | Rhizobium phage RHph_N3_2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43061 | 57.927 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988525 | Rhizobium phage RHph_Y65 | Rhizobium phage RHph_Y65 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 206821 | 41.174 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988524 | Rhizobium phage RHph_TM3_3_6 | Rhizobium phage RHph_TM3_3_6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 206788 | 41.101 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988523 | Rhizobium phage RHph_TM2_3B | Rhizobium phage RHph_TM2_3B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 206253 | 41.113 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988522 | Rhizobium phage RHph_TM40 | Rhizobium phage RHph_TM40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 207431 | 41.045 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988521 | Rhizobium phage RHph_TM30 | Rhizobium phage RHph_TM30 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 207623 | 41.049 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988520 | Rhizobium phage RHph_I9 | Rhizobium phage RHph_I9 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154143 | 48.495 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988519 | Rhizobium phage RHph_I1_18 | Rhizobium phage RHph_I1_18 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170978 | 47.503 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988518 | Rhizobium phage RHph_N2_10 | Rhizobium phage RHph_N2_10 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6010 | 58.719 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988517 | Rhizobium phage RHph_N38 | Rhizobium phage RHph_N38 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78126 | 46.152 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MN988516 | Rhizobium phage RHph_N28_2 | Rhizobium phage RHph_N28_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52336 | 59.164 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988515 | Rhizobium phage RHph_N28_1 | Rhizobium phage RHph_N28_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193452 | 53.439 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988514 | Rhizobium phage RHph_Y2_7 | Rhizobium phage RHph_Y2_7 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46273 | 56.126 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988513 | Rhizobium phage RHph_Y86 | Rhizobium phage RHph_Y86 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46273 | 56.156 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988512 | Rhizobium phage RHph_Y48 | Rhizobium phage RHph_Y48 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46273 | 56.134 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988511 | Rhizobium phage RHph_Y2 | Rhizobium phage RHph_Y2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6006 | 58.708 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988510 | Rhizobium phage RHph_I3_11 | Rhizobium phage RHph_I3_11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68071 | 42.896 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988509 | Rhizobium phage RHph_I46 | Rhizobium phage RHph_I46 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154143 | 48.494 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988508 | Rhizobium phage RHph_I36 | Rhizobium phage RHph_I36 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54438 | 56.457 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988507 | Rhizobium phage RHph_N3_13 | Rhizobium phage RHph_N3_13 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68049 | 42.959 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988506 | Rhizobium phage RHph_N1_15 | Rhizobium phage RHph_N1_15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116665 | 59.765 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988505 | Rhizobium phage RHph_N1_10 | Rhizobium phage RHph_N1_10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115974 | 59.788 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988504 | Rhizobium phage RHph_N25 | Rhizobium phage RHph_N25 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6010 | 58.686 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988503 | Rhizobium phage RHph_Y3_56_2 | Rhizobium phage RHph_Y3_56_2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.469 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988501 | Rhizobium phage RHph_Y3_43 | Rhizobium phage RHph_Y3_43 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54023 | 56.489 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988500 | Rhizobium phage RHph_Y55 | Rhizobium phage RHph_Y55 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54496 | 56.002 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988499 | Rhizobium phage RHph_TM1_20A | Rhizobium phage RHph_TM1_20A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.552 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988498 | Rhizobium phage RHph_Y3_19 | Rhizobium phage RHph_Y3_19 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.469 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988497 | Rhizobium phage RHph_Y2_6 | Rhizobium phage RHph_Y2_6 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78314 | 45.427 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MN988496 | Rhizobium phage RHph_TM3_3_14B | Rhizobium phage RHph_TM3_3_14B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59457 | 58.893 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988495 | Rhizobium phage RHph_TM3_3_13 | Rhizobium phage RHph_TM3_3_13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67917 | 56.482 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988494 | Rhizobium phage RHph_TM3_3_12 | Rhizobium phage RHph_TM3_3_12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.452 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988493 | Rhizobium phage RHph_TM38 | Rhizobium phage RHph_TM38 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46454 | 49.188 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| MN988492 | Rhizobium phage RHph_TM33 | Rhizobium phage RHph_TM33 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46454 | 49.188 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| MN988491 | Rhizobium phage RHph_Y3_36 | Rhizobium phage RHph_Y3_36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.452 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988490 | Rhizobium phage RHph_Y3_1 | Rhizobium phage RHph_Y3_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40173 | 58.497 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988489 | Rhizobium phage RHph_Y2_19 | Rhizobium phage RHph_Y2_19 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.485 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988488 | Rhizobium phage RHph_Y1_20 | Rhizobium phage RHph_Y1_20 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45999 | 56.114 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988487 | Rhizobium phage RHph_Y1_10 | Rhizobium phage RHph_Y1_10 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43230 | 57.904 | Unclassified | Unclassified | Podoviridae | Rhizobium |
| MN988486 | Rhizobium phage RHph_Y68 | Rhizobium phage RHph_Y68 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152762 | 49.387 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN988485 | Rhizobium phage RHph_Y67 | Rhizobium phage RHph_Y67 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4801 | 56.697 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988484 | Rhizobium phage RHph_Y60 | Rhizobium phage RHph_Y60 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69093 | 56.505 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988483 | Rhizobium phage RHph_Y38 | Rhizobium phage RHph_Y38 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77869 | 45.869 | Unclassified | Unclassified | Schitoviridae | Rhizobium |
| MN988482 | Rhizobium phage RHph_Y17 | Rhizobium phage RHph_Y17 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54023 | 56.489 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| MN988481 | Rhizobium phage RHph_TM3_4A | Rhizobium phage RHph_TM3_4A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.402 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988480 | Rhizobium phage RHph_TM1_10A | Rhizobium phage RHph_TM1_10A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.402 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988479 | Rhizobium phage RHph_TM15 | Rhizobium phage RHph_TM15 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.436 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988478 | Rhizobium phage RHph_TM3_3_7A | Rhizobium phage RHph_TM3_3_7A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6037 | 58.489 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988477 | Rhizobium phage RHph_TM32 | Rhizobium phage RHph_TM32 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6210 | 58.857 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988476 | Rhizobium phage RHph_TM1_7A | Rhizobium phage RHph_TM1_7A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6204 | 58.865 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988475 | Rhizobium phage RHph_TM3_3_5B | Rhizobium phage RHph_TM3_3_5B unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.452 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988474 | Rhizobium phage RHph_TM3_3_4B | Rhizobium phage RHph_TM3_3_4B unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6037 | 58.456 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988473 | Rhizobium phage RHph_TM1_10B | Rhizobium phage RHph_TM1_10B unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6033 | 58.412 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988472 | Rhizobium phage RHph_TM3_14A | Rhizobium phage RHph_TM3_14A Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60889 | 56.252 | Unclassified | Unclassified | Unclassified | Rhizobium |
| MN988471 | Rhizobium phage RHph_TM3_3_14A | Rhizobium phage RHph_TM3_3_14A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6038 | 58.48 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988470 | Rhizobium phage RHph_TM3_3_9 | Rhizobium phage RHph_TM3_3_9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67842 | 56.49 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988469 | Rhizobium phage RHph_TM3_3_5A | Rhizobium phage RHph_TM3_3_5A unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6034 | 58.469 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988468 | Rhizobium phage RHph_TM3_3_3 | Rhizobium phage RHph_TM3_3_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53608 | 60.683 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN988467 | Rhizobium phage RHph_TM36 | Rhizobium phage RHph_TM36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6037 | 58.473 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988466 | Rhizobium phage RHph_TM34 | Rhizobium phage RHph_TM34 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46058 | 56.236 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MN988465 | Rhizobium phage RHph_TM26 | Rhizobium phage RHph_TM26 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54519 | 58.528 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| MN988464 | Rhizobium phage RHph_TM24 | Rhizobium phage RHph_TM24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6208 | 58.827 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988463 | Rhizobium phage RHph_TM29 | Rhizobium phage RHph_TM29 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60940 | 56.265 | Unclassified | Unclassified | Unclassified | Rhizobium |
| MN988462 | Rhizobium phage RHph_TM27B | Rhizobium phage RHph_TM27B Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60894 | 56.234 | Unclassified | Unclassified | Unclassified | Rhizobium |
| MN988461 | Rhizobium phage RHph_TM27A | Rhizobium phage RHph_TM27A Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60894 | 56.235 | Unclassified | Unclassified | Unclassified | Rhizobium |
| MN988460 | Rhizobium phage RHph_TM23 | Rhizobium phage RHph_TM23 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6046 | 58.468 | Unclassified | Unclassified | Microviridae | Rhizobium |
| MN988459 | Rhizobium phage RHph_TM16 | Rhizobium phage RHph_TM16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101119 | 58.773 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MT354569 | Pseudomonas phage phiK7B1 | Pseudomonas phage phiK7B1 unclassified Nickievirus Nickievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112269 | 56.468 | Nickievirus | Unclassified | Siphoviridae | Pseudomonas |
| MT310860 | Mycobacterium phage Chris | Mycobacterium phage Chris unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62067 | 67.226 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN908692 | Mycobacterium Phage BGlluviae | Mycobacterium Phage BGlluviae unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59308 | 66.5 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586025 | Mycobacterium phage Mdavu | Mycobacterium phage Mdavu unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56443 | 67.078 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369746 | Mycobacterium phage Inky | Mycobacterium phage Inky unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.549 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369742 | Mycobacterium phage Spock | Mycobacterium phage Spock unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59709 | 66.546 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234231 | Mycobacterium phage Rapunzel97 | Mycobacterium phage Rapunzel97 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59687 | 66.602 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234230 | Mycobacterium phage Atiba | Mycobacterium phage Atiba unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59556 | 66.544 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234226 | Mycobacterium phage Curiosium | Mycobacterium phage Curiosium unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61222 | 65.645 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234205 | Mycobacterium phage Veliki | Mycobacterium phage Veliki unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59734 | 66.595 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234201 | Mycobacterium phage Guanica15 | Mycobacterium phage Guanica15 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60974 | 66.576 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234198 | Mycobacterium phage Zavala | Mycobacterium phage Zavala unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59969 | 66.593 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234197 | Mycobacterium phage Ramen | Mycobacterium phage Ramen unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59462 | 66.589 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234182 | Mycobacterium phage Geraldini | Mycobacterium phage Geraldini unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59818 | 66.607 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234174 | Mycobacterium phage Efra2 | Mycobacterium phage Efra2 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61284 | 66.557 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234165 | Mycobacterium phage Yunkel11 | Mycobacterium phage Yunkel11 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60757 | 66.559 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494119 | Mycobacterium Phage Niklas | Mycobacterium Phage Niklas unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60989 | 68.509 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310138 | Mycobacterium phage Shaobing | Mycobacterium phage Shaobing unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61030 | 68.543 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK460247 | Mycobacterium phage YoureAdopted | Mycobacterium phage YoureAdopted unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59504 | 66.554 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK460246 | Mycobacterium phage Nibb | Mycobacterium phage Nibb unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62293 | 67.565 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK376955 | Mycobacterium phage KiSi | Mycobacterium phage KiSi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62558 | 68.749 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112537 | Mycobacterium phage Capricorn | Mycobacterium phage Capricorn unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.546 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH910042 | Mycobacterium phage Scarlett | Mycobacterium phage Scarlett unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62306 | 68.745 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH910039 | Mycobacterium phage Oscar | Mycobacterium phage Oscar unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62437 | 68.764 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH910038 | Mycobacterium phage LeMond | Mycobacterium phage LeMond unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62515 | 68.766 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016503 | Mycobacterium phage Prithvi | Mycobacterium phage Prithvi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60311 | 66.771 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727544 | Mycobacterium phage Dalmuri | Mycobacterium phage Dalmuri unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.546 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697578 | Mycobacterium phage Belladonna | Mycobacterium phage Belladonna unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.554 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513977 | Mycobacterium phage Mynx | Mycobacterium phage Mynx unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60055 | 66.594 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536822 | Mycobacterium phage Joy99 | Mycobacterium phage Joy99 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59837 | 66.551 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536821 | Mycobacterium phage Homura | Mycobacterium phage Homura unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.559 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH509444 | Mycobacterium phage BEEST | Mycobacterium phage BEEST unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59906 | 66.556 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399770 | Mycobacterium phage Biglebops | Mycobacterium phage Biglebops unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56454 | 67.086 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371113 | Mycobacterium phage Beezoo | Mycobacterium phage Beezoo unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60494 | 66.501 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077577 | Mycobacterium phage AlishaPH | Mycobacterium phage AlishaPH unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57034 | 66.606 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051246 | Mycobacterium phage ActinUp | Mycobacterium phage ActinUp unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59812 | 66.62 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020245 | Mycobacterium phage SgtBeansprout | Mycobacterium phage SgtBeansprout unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56439 | 67.087 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG962371 | Mycobacterium phage LaterM | Mycobacterium phage LaterM unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60143 | 66.55 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG962364 | Mycobacterium phage Deby | Mycobacterium phage Deby unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60463 | 66.499 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001453 | Mycobacterium phage Adonis | Mycobacterium phage Adonis unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60031 | 66.676 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919532 | Mycobacterium phage Sulley | Mycobacterium phage Sulley unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59873 | 66.37 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919518 | Mycobacterium phage LilPharaoh | Mycobacterium phage LilPharaoh unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56167 | 67.079 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF472892 | Mycobacterium phage TreyKay | Mycobacterium phage TreyKay unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60311 | 66.774 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668267 | Mycobacterium phage Apocalypse | Mycobacterium phage Apocalypse unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59947 | 66.37 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140430 | Mycobacterium phage Tachez | Mycobacterium phage Tachez unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59556 | 66.549 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140428 | Mycobacterium phage Slimphazie | Mycobacterium phage Slimphazie unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60143 | 66.616 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140416 | Mycobacterium phage LastHope | Mycobacterium phage LastHope unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60934 | 66.876 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140412 | Mycobacterium phage Jeckyll | Mycobacterium phage Jeckyll unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59708 | 66.551 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185722 | Mycobacterium phage Peanam | Mycobacterium phage Peanam unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61041 | 68.547 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF377440 | Mycobacterium phage Bella96 | Mycobacterium phage Bella96 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60746 | 66.143 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY380102 | Mycobacterium phage CREW | Mycobacterium phage CREW unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59707 | 66.555 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX808132 | Mycobacterium phage Amelie | Mycobacterium phage Amelie unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56439 | 67.087 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX657796 | Mycobacterium phage Urkel | Mycobacterium phage Urkel unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60526 | 66.224 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX657795 | Mycobacterium phage DrHayes | Mycobacterium phage DrHayes unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60526 | 66.226 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX657794 | Mycobacterium phage SamuelLPlaqson | Mycobacterium phage SamuelLPlaqson unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60526 | 66.224 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641264 | Mycobacterium phage LindNT | Mycobacterium phage LindNT unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60053 | 66.784 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX585253 | Mycobacterium phage HedwigODU | Mycobacterium phage HedwigODU unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59812 | 66.625 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX585251 | Mycobacterium phage Waterfoul | Mycobacterium phage Waterfoul unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61248 | 64.938 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KT281789 | Mycobacterium phage Enkosi | Mycobacterium phage Enkosi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59052 | 67.183 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM677211 | Mycobacterium phage Murucutumbu | Mycobacterium phage Murucutumbu unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60609 | 66.693 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ567045 | Mycobacterium phage Emerson | Mycobacterium phage Emerson unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60310 | 66.599 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF713486 | Mycobacterium phage Validus | Mycobacterium phage Validus unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62466 | 68.431 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN643714 | Mycobacterium phage BarrelRoll | Mycobacterium phage BarrelRoll unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59672 | 66.616 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704105 | Mycobacterium phage Adephagia | Mycobacterium phage Adephagia unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59646 | 66.606 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN185608 | Mycobacterium virus JAWS | Mycobacterium virus JAWS Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59749 | 66.617 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT151604 | Bacillus phage P59 | Bacillus phage P59 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159363 | 42.337 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MW345241 | Acinetobacter phage vB_AbaP_APK26 | Acinetobacter phage vB_AbaP_APK26 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41097 | 39.261 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW394467 | Bacillus phage 268TH004 | Bacillus phage 268TH004 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52834 | 42.452 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MW406974 | Pseudomonas phage TH15 | Pseudomonas phage TH15 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65936 | 55.574 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MW388003 | Acinetobacter phage Melin | Acinetobacter phage Melin unclassified Hadassahvirus Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167969 | 36.328 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MW388002 | Acinetobacter phage Mokit | Acinetobacter phage Mokit unclassified Hadassahvirus Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167638 | 36.324 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MW388001 | Acinetobacter phage Minot | Acinetobacter phage Minot unclassified Hadassahvirus Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167029 | 36.33 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MW349129 | Staphylococcus phage Madawaska | Staphylococcus phage Madawaska Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 265446 | 25.132 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MW349128 | Staphylococcus phage Machias | Staphylococcus phage Machias Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 274478 | 24.753 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MW353177 | Flavobacterium phage vB_FspM_immuto_13-6C | Flavobacterium phage vB_FspM_immuto_13-6C Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160366 | 34.502 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW353176 | Flavobacterium phage vB_FspM_immuto_3-5A | Flavobacterium phage vB_FspM_immuto_3-5A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160366 | 34.504 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW353175 | Flavobacterium phage vB_FspM_immuto_2-6A | Flavobacterium phage vB_FspM_immuto_2-6A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160410 | 34.52 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MW448170 | Klebsiella phage vB_KpnM_M1 | Klebsiella phage vB_KpnM_M1 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176329 | 41.852 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MW384823 | Erwinia phage pEa_SNUABM_43 | Erwinia phage pEa_SNUABM_43 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241904 | 51.338 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| MW366843 | Erwinia phage pEa_SNUABM_5 | Erwinia phage pEa_SNUABM_5 Yoloswagvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 267169 | 48.857 | Yoloswagvirus | Unclassified | Myoviridae | Erwinia |
| MW354669 | Bacillus phage BSTP5 | Bacillus phage BSTP5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55448 | 39.908 | Unclassified | Unclassified | Podoviridae | Bacillus |
| MW354668 | Bacillus phage BSTP4 | Bacillus phage BSTP4 unclassified Salasvirus Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19145 | 39.948 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| MW344056 | Vibrio phage ABurr | Vibrio phage ABurr unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126843 | 40.237 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MW344055 | Vibrio phage GRLPWR | Vibrio phage GRLPWR unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128336 | 40.205 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MW417503 | Klebsiella phage LF20 | Klebsiella phage LF20 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50107 | 50.741 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW390886 | Pseudomonas phage AIIMS-Pa-B1 | Pseudomonas phage AIIMS-Pa-B1 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37578 | 64.57 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MW419087 | Bacillus phage 015DV004 | Bacillus phage 015DV004 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156577 | 38.576 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MW419086 | Bacillus phage 015DV002 | Bacillus phage 015DV002 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153882 | 38.706 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MW419085 | Bacillus phage 000TH009 | Bacillus phage 000TH009 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155191 | 38.639 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MW419084 | Bacillus phage 000TH008 | Bacillus phage 000TH008 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155246 | 38.61 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MW442830 | Flavobacterium phage FPSV-D18 | Flavobacterium phage FPSV-D18 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90238 | 26.822 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW442829 | Flavobacterium phage FPSV-D16 | Flavobacterium phage FPSV-D16 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.815 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW442828 | Flavobacterium phage FPSV-D13 | Flavobacterium phage FPSV-D13 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.811 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW442827 | Flavobacterium phage FPSV-D12 | Flavobacterium phage FPSV-D12 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.816 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW442826 | Flavobacterium phage FPSV-D9 | Flavobacterium phage FPSV-D9 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.812 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW442825 | Flavobacterium phage FPSV-D5 | Flavobacterium phage FPSV-D5 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.797 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW387018 | Gardnerella phage vB_Gva_AB1 | Gardnerella phage vB_Gva_AB1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50268 | 39.546 | Unclassified | Unclassified | Siphoviridae | Gardnerella |
| MW413353 | Salmonella phage SPHG1 | Salmonella phage SPHG1 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47119 | 46.143 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MW388005 | Salmonella phage SPHG3 | Salmonella phage SPHG3 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149831 | 44.494 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW363799 | Staphylococcus phage LSA2366 | Staphylococcus phage LSA2366 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17056 | 29.397 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MW363798 | Staphylococcus phage LSA2308 | Staphylococcus phage LSA2308 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144592 | 29.886 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MW358931 | Providencia phage PSTRCR_128 | Providencia phage PSTRCR_128 Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38838 | 46.287 | Minipunavirus | Studiervirinae | Autographiviridae | Providencia |
| MW358930 | Providencia phage PSTRCR_114 | Providencia phage PSTRCR_114 Kakivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43106 | 42.702 | Kakivirus | Unclassified | Autographiviridae | Providencia |
| MW358929 | Providencia phage PSTRCR_117lys | Providencia phage PSTRCR_117lys Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41059 | 49.402 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW358928 | Providencia phage PSTRCR_120 | Providencia phage PSTRCR_120 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39396 | 42.085 | Unclassified | Studiervirinae | Autographiviridae | Providencia |
| MW358927 | Providencia phage PSTRCR_127 | Providencia phage PSTRCR_127 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167162 | 34.714 | Unclassified | Tevenvirinae | Myoviridae | Providencia |
| MW296032 | Salmonella phage pSal-SNUABM-01 | Salmonella phage pSal-SNUABM-01 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89500 | 44.96 | Unclassified | Unclassified | Podoviridae | Salmonella |
| MW247147 | Helicobacter phage COL 23-PUJ | Helicobacter phage COL 23-PUJ unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28622 | 37.293 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| MW247145 | Helicobacter phage COL 6-PUJ | Helicobacter phage COL 6-PUJ unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26487 | 37.698 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| MW247144 | Helicobacter phage COL 5-PUJ | Helicobacter phage COL 5-PUJ unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26486 | 37.699 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| MW247146 | Helicobacter phage COL 16-PUJ | Helicobacter phage COL 16-PUJ unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19749 | 36.62 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| MW247143 | Helicobacter phage COL 2-PUJ | Helicobacter phage COL 2-PUJ unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19298 | 36.579 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| MT966873 | Klebsiella phage P560 | Klebsiella phage P560 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40562 | 53.057 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT966872 | Klebsiella phage P510 | Klebsiella phage P510 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39383 | 53.124 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT782071 | Escherichia phage JN02 | Escherichia phage JN02 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169453 | 37.669 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MW286157 | Escherichia phage vB_EcoM-BECP11 | Escherichia phage vB_EcoM-BECP11 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165973 | 35.366 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MW286156 | Escherichia phage vB_EcoS-BECP10 | Escherichia phage vB_EcoS-BECP10 unclassified Rogunavirus Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47915 | 43.692 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| MW331544 | Vibrio phage vB_Vc_SrVc2 | Vibrio phage vB_Vc_SrVc2 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43157 | 49.169 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MW406976 | Pseudomonas phage HX1 | Pseudomonas phage HX1 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43113 | 62.239 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MW406975 | Pseudomonas phage MYY9 | Pseudomonas phage MYY9 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43005 | 62.207 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MW388004 | Acinetobacter phage Morttis | Acinetobacter phage Morttis unclassified Hadassahvirus Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169150 | 36.332 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MW393850 | Stenotrophomonas phage Salva | Stenotrophomonas phage Salva Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60789 | 56.377 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MW448541 | Flavobacterium phage FPSV-D27 | Flavobacterium phage FPSV-D27 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.803 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW448540 | Flavobacterium phage FPSV-D25 | Flavobacterium phage FPSV-D25 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.813 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW448539 | Flavobacterium phage FPSV-D24 | Flavobacterium phage FPSV-D24 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91725 | 26.811 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MW132713 | Gordonia phage TinaLin | Gordonia phage TinaLin unclassified Vendettavirus Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43366 | 65.408 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| MW348917 | Bacillus phage Thornton | Bacillus phage Thornton unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26319 | 30.487 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MW325771 | Salmonella phage KM16 | Salmonella phage KM16 unclassified Moonvirus Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170126 | 38.613 | Moonvirus | Tevenvirinae | Myoviridae | Salmonella |
| MW248466 | Staphylococcus phage MarsHill | Staphylococcus phage MarsHill Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266637 | 25.178 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| MW114771 | Vibrio phage pVco-14 | Vibrio phage pVco-14 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59428 | 46.998 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MW023914 | Rhizobium phage vB_RleA_TRX32-1 | Rhizobium phage vB_RleA_TRX32-1 Paadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42787 | 59.226 | Paadamvirus | Unclassified | Autographiviridae | Rhizobium |
| MT949699 | Acinetobacter phage Maestro | Acinetobacter phage Maestro unclassified Hadassahvirus Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169176 | 36.331 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MW423739 | Vibrio phage PH669 | Vibrio phage PH669 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16687 | 45.802 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MW423738 | Vibrio phage PH669 | Vibrio phage PH669 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18896 | 46.862 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MW423737 | Vibrio phage PH669 | Vibrio phage PH669 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23061 | 46.247 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MK610268 | Hafnia phage vB_HpaA_yong1 | Hafnia phage vB_HpaA_yong1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40286 | 50.146 | Kayfunavirus | Studiervirinae | Autographiviridae | Hafnia |
| MT380195 | Klebsiella phage vB_Kpn_B01 | Klebsiella phage vB_Kpn_B01 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113227 | 45.459 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MT002874 | Pseudoalteromonas phage XC | Pseudoalteromonas phage XC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46490 | 40.013 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MT740748 | Aeromonas phage T7-Ah | Aeromonas phage T7-Ah unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40153 | 51.824 | Unclassified | Studiervirinae | Autographiviridae | Aeromonas |
| MN087708 | Enterobacter phage vB_EhoM-IME523 | Enterobacter phage vB_EhoM-IME523 unclassified Kanagawavirus Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172763 | 39.97 | Kanagawavirus | Unclassified | Myoviridae | Enterobacter |
| MN929097 | Parabacteroides phage PDS1 | Parabacteroides phage PDS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44691 | 45.278 | Unclassified | Unclassified | Siphoviridae | Parabacteroides |
| MN917146 | Bacteroides phage crAss002 | Bacteroides phage crAss002 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93030 | 31.929 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| NC_028803 | Mycobacterium phage OSmaximus | Mycobacterium phage OSmaximus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69118 | 66.327 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028681 | Mycobacterium phage Pops | Mycobacterium phage Pops Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68367 | 66.601 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MW272540 | Aeromonas phage ZPAH1 | Aeromonas phage ZPAH1 unclassified Ishigurovirus Ishigurovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 230916 | 38.22 | Ishigurovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MW343794 | Cronobacter phage A24 | Cronobacter phage A24 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75106 | 44.051 | Unclassified | Unclassified | Podoviridae | Cronobacter |
| MW239124 | Citrobacter phage CkP1 | Citrobacter phage CkP1 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168463 | 36.889 | Karamvirus | Tevenvirinae | Myoviridae | Citrobacter |
| MW233709 | Escherichia phage UPEC01 | Escherichia phage UPEC01 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169801 | 40.429 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT975991 | Alteromonas phage phiAFP1 | Alteromonas phage phiAFP1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5859 | 40.587 | Unclassified | Unclassified | Inoviridae | Alteromonas |
| MT653137 | Salmonella phage S55 | Salmonella phage S55 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42781 | 49.728 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK455770 | Aeromonas phage HJG | Aeromonas phage HJG unclassified Uliginvirus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44965 | 50.268 | Uliginvirus | Colwellvirinae | Autographiviridae | Aeromonas |
| LC596379 | Enterococcus phage phi EF19G | Enterococcus phage phi EF19G unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143400 | 35.952 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| LC596378 | Enterococcus phage phi EF14H1 | Enterococcus phage phi EF14H1 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143280 | 35.949 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| LC596377 | Enterococcus phage phi EF7H | Enterococcus phage phi EF7H unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143399 | 35.953 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MW269554 | Achromobacter phage vB_AchrS_AchV4 | Achromobacter phage vB_AchrS_AchV4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59489 | 62.83 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| MT944117 | Escherichia phage PNJ1809-36 | Escherichia phage PNJ1809-36 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152343 | 39.106 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| AP019417 | Vibrio phage KIT05 | Vibrio phage KIT05 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50628 | 41.627 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MW021758 | Serratia phage vB_SmaM_Yaphecito | Serratia phage vB_SmaM_Yaphecito unclassified Moabitevirus Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 275554 | 46.764 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| MW021763 | Serratia phage vB_SmaS_Bigdog | Serratia phage vB_SmaS_Bigdog Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42495 | 45.699 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MW021761 | Serratia phage vB_SmaS_Rovert | Serratia phage vB_SmaS_Rovert Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38613 | 42.323 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MW021760 | Salmonella phage vB_SenM_SilasIsHot | Salmonella phage vB_SenM_SilasIsHot unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160559 | 45.134 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW021755 | Serratia phage vB_SmaS_Serratianator | Serratia phage vB_SmaS_Serratianator Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42052 | 45.967 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MW021750 | Citrobacter phage vB_CfrD_Supreme284 | Citrobacter phage vB_CfrD_Supreme284 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49439 | 42.851 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| MT901800 | Erwinia phage PEar6 | Erwinia phage PEar6 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6608 | 41.677 | Unclassified | Unclassified | Inoviridae | Erwinia |
| LC567842 | Stx2a-converting phage Stx2_12E129_yecE | Stx2a-converting phage Stx2_12E129_yecE Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46100 | 52.154 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567841 | Stx2a-converting phage Stx2_12E129_PPompW | Stx2a-converting phage Stx2_12E129_PPompW Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90031 | 51.581 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567840 | Stx2a-converting phage Stx2_EH1846 | Stx2a-converting phage Stx2_EH1846 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48297 | 52.417 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567839 | Enterobacteria phage PPompW_EH1846 | Enterobacteria phage PPompW_EH1846 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44438 | 51.229 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567838 | Stx2a-converting phage Stx2_95 | Stx2a-converting phage Stx2_95 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46305 | 52.051 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567837 | Stx2a-converting phage Stx2_EH2246 | Stx2a-converting phage Stx2_EH2246 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49002 | 52.341 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567836 | Enterobacteria phage PPompW_EH2246 | Enterobacteria phage PPompW_EH2246 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43789 | 51.659 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567835 | Enterobacteria phage PPyecE_EH1910 | Enterobacteria phage PPyecE_EH1910 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49808 | 52.277 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567834 | Stx2a-converting phage Stx2_EH1910 | Stx2a-converting phage Stx2_EH1910 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89439 | 52.666 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567833 | Stx1a-converting phage Stx1_EH1992 | Stx1a-converting phage Stx1_EH1992 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49929 | 51.503 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567832 | Enterobacteria phage PPompW_EH1992 | Enterobacteria phage PPompW_EH1992 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44479 | 51.278 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567831 | Stx1a-converting phage Stx1_EH1995 | Stx1a-converting phage Stx1_EH1995 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49937 | 51.499 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567830 | Stx2d-converting phage Stx2_112808 | Stx2d-converting phage Stx2_112808 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45909 | 52.166 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567829 | Stx2a-converting phage Stx2_EH2201 | Stx2a-converting phage Stx2_EH2201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48694 | 52.397 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567828 | Enterobacteria phage PPompW_EH2201 | Enterobacteria phage PPompW_EH2201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45811 | 51.322 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567827 | Stx1a-converting phage Stx1_699 | Stx1a-converting phage Stx1_699 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49396 | 51.205 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567826 | Enterobacteria phage PPompW_699 | Enterobacteria phage PPompW_699 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44349 | 51.235 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567825 | Stx1a-converting phage Stx1_499 | Stx1a-converting phage Stx1_499 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49243 | 51.23 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567824 | Stx2a-converting phage Stx2_499 | Stx2a-converting phage Stx2_499 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93786 | 51.757 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567823 | Stx1a-converting phage Stx1_132418 | Stx1a-converting phage Stx1_132418 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49399 | 51.26 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567822 | Enterobacteria phage PPompW_132418 | Enterobacteria phage PPompW_132418 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44350 | 51.236 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC567821 | Stx1a-converting phage Stx1_14744 | Stx1a-converting phage Stx1_14744 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49422 | 51.22 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567820 | Stx2a-converting phage Stx2_14744 | Stx2a-converting phage Stx2_14744 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89850 | 51.683 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567819 | Stx1a-converting phage Stx1_14040 | Stx1a-converting phage Stx1_14040 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49573 | 51.238 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC567818 | Stx2a-converting phage Stx2_14040 | Stx2a-converting phage Stx2_14040 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89855 | 51.691 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC545446 | Nitratiruptor phage NrS-5 | Nitratiruptor phage NrS-5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43030 | 38.994 | Unclassified | Unclassified | Siphoviridae | Nitratiruptor |
| LC545445 | Nitratiruptor phage NrS-4 | Nitratiruptor phage NrS-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43030 | 38.994 | Unclassified | Unclassified | Siphoviridae | Nitratiruptor |
| LC545444 | Nitratiruptor phage NrS-3 | Nitratiruptor phage NrS-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40036 | 39.202 | Unclassified | Unclassified | Siphoviridae | Nitratiruptor |
| LC545443 | Nitratiruptor phage NrS-2 | Nitratiruptor phage NrS-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40465 | 39.162 | Unclassified | Unclassified | Siphoviridae | Nitratiruptor |
| MW032477 | Lactococcus phage FB14 | Lactococcus phage FB14 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31042 | 34.492 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041640 | Lactococcus phage FB10 | Lactococcus phage FB10 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30272 | 35.062 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041637 | Lactococcus phage GP13 | Lactococcus phage GP13 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29126 | 34.883 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041636 | Lactococcus phage RH10 | Lactococcus phage RH10 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30860 | 34.112 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041635 | Lactococcus phage GP15 | Lactococcus phage GP15 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29166 | 34.636 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041634 | Lactococcus phage GP14 | Lactococcus phage GP14 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29200 | 34.575 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041633 | Lactococcus phage FB6 | Lactococcus phage FB6 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30766 | 34.499 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW041632 | Lactococcus phage FB3 | Lactococcus phage FB3 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28969 | 34.789 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MW291028 | Mycobacterium phage Peeb | Mycobacterium phage Peeb unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41876 | 66.582 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MW291026 | Microbacterium phage Erla | Microbacterium phage Erla unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41538 | 63.366 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MW291023 | Mycobacterium phage Harella | Mycobacterium phage Harella unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76859 | 62.976 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MW291019 | Mycobacterium phage BangNhom | Mycobacterium phage BangNhom unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51808 | 63.928 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT901799 | Erwinia phage PEar4 | Erwinia phage PEar4 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6801 | 41.656 | Unclassified | Unclassified | Inoviridae | Erwinia |
| MT901798 | Erwinia phage PEar2 | Erwinia phage PEar2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6651 | 41.693 | Unclassified | Unclassified | Inoviridae | Erwinia |
| MT901797 | Erwinia phage PEar1 | Erwinia phage PEar1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6646 | 41.679 | Unclassified | Unclassified | Inoviridae | Erwinia |
| NC_049472 | Klebsiella virus UPM 2146 | Klebsiella virus UPM 2146 unclassified Taipeivirus Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160795 | 46.306 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_049391 | Escherichia virus P88-4B11 | Escherichia virus P88-4B11 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37100 | 51.765 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049390 | Escherichia virus P2-4A7b | Escherichia virus P2-4A7b Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32848 | 50.932 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049389 | Escherichia virus P2-4E6b | Escherichia virus P2-4E6b Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31128 | 51.982 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049388 | Escherichia virus P2-4C9 | Escherichia virus P2-4C9 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32150 | 51.938 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049387 | Escherichia virus P2-4B2 | Escherichia virus P2-4B2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32118 | 51.389 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049386 | Escherichia virus P2-2H1 | Escherichia virus P2-2H1 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32662 | 51.111 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049385 | Escherichia virus P2-2H4 | Escherichia virus P2-2H4 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31342 | 52.131 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| NC_049384 | Enterobacter phage EspM4VN | Enterobacter phage EspM4VN unclassified Agtrevirus Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160766 | 50.701 | Agtrevirus | Aglimvirinae | Ackermannviridae | Enterobacter |
| NC_049380 | Vibrio phage JSF3 | Vibrio phage JSF3 unclassified Pacinivirus Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69396 | 34.573 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| NC_048755 | Stenotrophomonas phage YB07 | Stenotrophomonas phage YB07 unclassified Menderavirus Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159862 | 54.128 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| NC_047757 | Caulobacter phage Seuss | Caulobacter phage Seuss Caulobacter virus Seuss Seussvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85138 | 61.403 | Seussvirus | Unclassified | Siphoviridae | Caulobacter |
| NC_048207 | Gordonia phage SteveFrench | Gordonia phage SteveFrench Gordonia virus Stevefrench Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75687 | 59.182 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048205 | Escherichia phage vB_Eco_mar004NP2 | Escherichia phage vB_Eco_mar004NP2 Escherichia virus mar004NP2 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107601 | 38.944 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048203 | Escherichia phage vB_Eco_mar003J3 | Escherichia phage vB_Eco_mar003J3 Escherichia virus mar003J3 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115471 | 39.775 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048199 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 27548 | 29.81 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| NC_048196 | Escherichia phage vB_EcoM_Schickermooser | Escherichia phage vB_EcoM_Schickermooser Escherichia virus Schickermooser Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151194 | 39.008 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_048195 | Escherichia phage vB_EcoM_WFC | Escherichia phage vB_EcoM_WFC Escherichia virus WFC Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72472 | 46.011 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| NC_048194 | Escherichia phage vB_EcoM_WFH | Escherichia phage vB_EcoM_WFH Escherichia virus WFH Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71283 | 46.118 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| NC_048193 | Escherichia phage PTXU04 | Escherichia phage PTXU04 Escherichia virus PTXU04 Xuquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61600 | 52.844 | Xuquatrovirus | Unclassified | Podoviridae | Escherichia |
| NC_048190 | Rheinheimera phage vB_RspM_Barba19A | Rheinheimera phage vB_RspM_Barba19A Rheinheimera virus Barba19A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84608 | 38.247 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| NC_048189 | Rheinheimera phage vB_RspM_Barba18A | Rheinheimera phage vB_RspM_Barba18A Rheinheimera virus Barba18A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| NC_048186 | Escherichia phage vB_EcoM_KWBSE43-6 | Escherichia phage vB_EcoM_KWBSE43-6 Escherichia virus KWBSE43-6 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158607 | 46.091 | Taipeivirus | Unclassified | Ackermannviridae | Escherichia |
| NC_048182 | Edwardsiella virus pEtSU | Edwardsiella virus pEtSU Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 276734 | 43.173 | Petsuvirus | Unclassified | Myoviridae | Edwardsiella |
| NC_048173 | Erwinia phage Derbicus | Erwinia phage Derbicus Erwinia virus Derbicus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223950 | 50.564 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| NC_048166 | Pseudomonas phage Lana | Pseudomonas phage Lana Pseudomonas virus Lana Lanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88403 | 60.825 | Lanavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_048158 | Halorubrum pleomorphic virus 11 | Halorubrum pleomorphic virus 11 Betapleolipovirus HRPV11 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9368 | 55.209 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_048157 | Halorubrum pleomorphic virus 10 | Halorubrum pleomorphic virus 10 Betapleolipovirus HRPV10 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9296 | 55.669 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_048156 | Halorubrum pleomorphic virus 12 | Halorubrum pleomorphic virus 12 Betapleolipovirus HRPV12 Betapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 9944 | 55.31 | Betapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_048152 | Arthrobacter phage Sonali | Arthrobacter phage Sonali Arthrobacter virus Sonali Sonalivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61073 | 67.74 | Sonalivirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048151 | Vibrio phage VspSw_1 | Vibrio phage VspSw_1 Vibrio virus VspSw1 Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113760 | 43.836 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| NC_048149 | Salmonella phage 1-23 | Salmonella phage 1-23 Salmonella virus 123 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112496 | 39.969 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048148 | Providencia phage vB PstP PS3 | Providencia phage vB PstP PS3 Providencia virus PS3 Kakivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41024 | 42.992 | Kakivirus | Unclassified | Autographiviridae | Providencia |
| NC_048147 | Xanthomonas virus XcP1 | Xanthomonas virus XcP1 Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61830 | 52.496 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| NC_048145 | Salmonella phage 3-29 | Salmonella phage 3-29 Salmonella virus 329 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110377 | 40.045 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048144 | Microbacterium phage Schubert | Microbacterium phage Schubert Microbacterium virus Schubert Schubertvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38820 | 61.419 | Schubertvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048142 | Acinetobacter phage AbKT21phiIII | Acinetobacter phage AbKT21phiIII Acinetobacter virus AbKT21III Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40898 | 39.359 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_048139 | Streptomyces phage Hiyaa | Streptomyces phage Hiyaa Streptomyces virus Hiyaa Hiyaavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83219 | 66.661 | Hiyaavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048137 | Pantoea phage vB_PagS_AAS23 | Pantoea phage vB_PagS_AAS23 Pantoea virus AAS23 Sauletekiovirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51170 | 47.645 | Sauletekiovirus | Unclassified | Drexlerviridae | Pantoea |
| NC_048133 | Aeromonas phage ZPAH7 | Aeromonas phage ZPAH7 Aeromonas virus ZPAH7 Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30791 | 56.78 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_048123 | Vibrio phage 1.204.O._10N.222.46.F12 | Vibrio phage 1.204.O._10N.222.46.F12 Vibrio virus Cyclit Cyclitvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44168 | 43.516 | Cyclitvirus | Unclassified | Autographiviridae | Vibrio |
| NC_048120 | Pectobacterium phage Zenivior | Pectobacterium phage Zenivior Pectobacterium virus Zenivior Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43724 | 48.998 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048119 | Pectobacterium phage Phoria | Pectobacterium phage Phoria Pectobacterium virus Phoria Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43825 | 48.634 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048118 | Pectobacterium phage Koot | Pectobacterium phage Koot Pectobacterium virus Koot Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43402 | 49.023 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048116 | Pectobacterium phage Clickz | Pectobacterium phage Clickz Pectobacterium virus Clickz Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43310 | 49.192 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048115 | Arthrobacter phage Liebe | Arthrobacter phage Liebe Arthrobacter virus Liebe Liebevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45803 | 68.819 | Liebevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048111 | Pectobacterium phage Arno160 | Pectobacterium phage Arno160 Pectobacterium virus Arno160 Wanjuvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41823 | 51.455 | Wanjuvirus | Melnykvirinae | Autographiviridae | Pectobacterium |
| NC_048110 | Salmonella phage Seafire | Salmonella phage Seafire Salmonella virus Seafire Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111851 | 40.013 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048101 | Microbacterium phage Goodman | Microbacterium phage Goodman Microbacterium virus Goodman Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42363 | 66.31 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048100 | Arthrobacter phage Kepler | Arthrobacter phage Kepler Arthrobacter virus Kepler Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38449 | 66.735 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048099 | Arthrobacter phage Coral | Arthrobacter phage Coral Arthrobacter virus Coral Coralvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38317 | 66.66 | Coralvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048098 | Arthrobacter phage Andrew | Arthrobacter phage Andrew Arthrobacter virus Andrew Andrewvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38802 | 65.533 | Andrewvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_048090 | Microbacterium phage Armstrong | Microbacterium phage Armstrong Microbacterium virus Armstrong Armstrongvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39928 | 67.138 | Armstrongvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048089 | Salmonella phage STG2 | Salmonella phage STG2 Salmonella virus STG2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114275 | 40.118 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048086 | Proteus phage Stubb | Proteus phage Stubb Proteus virus Stubb Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104410 | 38.512 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| NC_048081 | Acinetobacter phage vB_AbaP_B09_Aci08 | Acinetobacter phage vB_AbaP_B09_Aci08 Acinetobacter virus Aci08 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42067 | 39.285 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_048080 | Acinetobacter phage vB_AbaM_B09_Aci05 | Acinetobacter phage vB_AbaM_B09_Aci05 Acinetobacter virus Aci05 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102789 | 37.207 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_048077 | Dickeya phage vB_DsoM_JA11 | Dickeya phage vB_DsoM_JA11 Dickeya virus JA11 Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 255356 | 44.47 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| NC_048076 | Acinetobacter phage vB_AbaP_46-62_Aci07 | Acinetobacter phage vB_AbaP_46-62_Aci07 Acinetobacter virus Aci07 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42330 | 39.135 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_048075 | Acinetobacter phage vB_AbaM_B09_Aci02-2 | Acinetobacter phage vB_AbaM_B09_Aci02-2 Acinetobacter virus Aci02-2 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104354 | 37.182 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_048074 | Acinetobacter phage vB_AbaM_B09_Aci01-1 | Acinetobacter phage vB_AbaM_B09_Aci01-1 Acinetobacter virus Aci01-1 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103628 | 37.186 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_048073 | Escherichia phage FEC19 | Escherichia phage FEC19 Escherichia virus FEC19 Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68472 | 46.117 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| NC_048068 | Microbacterium phage OneinaGillian | Microbacterium phage OneinaGillian Microbacterium virus OneinaGillian Gillianvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61703 | 67.071 | Gillianvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048067 | Escherichia phage C130_2 | Escherichia phage C130_2 Escherichia virus C1302 Giessenvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41775 | 55.43 | Giessenvirus | Unclassified | Podoviridae | Escherichia |
| NC_048062 | Salmonella phage Sw2 | Salmonella phage Sw2 Salmonella virus Sw2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114274 | 40.214 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048061 | Pectobacterium phage Nobby | Pectobacterium phage Nobby Pectobacterium virus Nobby Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42818 | 48.996 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048060 | Dickeya phage Mysterion | Dickeya phage Mysterion Dickeya virus Mysterion Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40651 | 48.11 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_048059 | Dickeya phage Luksen | Dickeya phage Luksen Dickeya virus Luksen Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40729 | 48.214 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_048058 | Pectobacterium phage Lelidair | Pectobacterium phage Lelidair Pectobacterium virus Lelidair Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43686 | 48.659 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048057 | Dickeya phage Katbat | Dickeya phage Katbat Dickeya virus Katbat Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40441 | 48.038 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_048056 | Pectobacterium phage Gaspode | Pectobacterium phage Gaspode Pectobacterium virus Gaspode Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44364 | 48.808 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_048054 | Dickeya phage vB_DsoM_AD1 | Dickeya phage vB_DsoM_AD1 Dickeya virus AD1 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261658 | 49.366 | Alexandravirus | Unclassified | Myoviridae | Dickeya |
| NC_048053 | Dickeya phage vB_DsoM_JA29 | Dickeya phage vB_DsoM_JA29 Dickeya virus JA29 Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253323 | 43.843 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| NC_048052 | Dickeya phage vB_DsoP_JA10 | Dickeya phage vB_DsoP_JA10 Dickeya virus JA10 Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40131 | 51.451 | Ningirsuvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_048047 | Caulobacter phage CcrBL9 | Caulobacter phage CcrBL9 Caulobacter virus CcrBL9 Bertelyvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 322272 | 63.679 | Bertelyvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_048046 | Caulobacter phage CcrPW | Caulobacter phage CcrPW Caulobacter virus CcrPW Colossusvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 308141 | 62.242 | Colossusvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_048045 | Caulobacter phage CcrBL10 | Caulobacter phage CcrBL10 Caulobacter virus CcrBL10 Poindextervirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 220934 | 65.687 | Poindextervirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_048041 | Vibrio phage vB_VpS_PG07 | Vibrio phage vB_VpS_PG07 Vibrio virus PG07 Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112106 | 43.65 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| NC_048038 | Microbacterium phage Neferthena | Microbacterium phage Neferthena Microbacterium virus Neferthena Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41706 | 64.386 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048036 | Vibrio phage vB_VpaP_KF2 | Vibrio phage vB_VpaP_KF2 Vibrio virus KF2 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43571 | 49.482 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| NC_048035 | Vibrio phage vB_VpaP_KF1 | Vibrio phage vB_VpaP_KF1 Vibrio virus KF1 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43237 | 49.49 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| NC_048029 | Sphingomonas phage Scott | Sphingomonas phage Scott Sphingomonas virus Scott Scottvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43077 | 57.43 | Scottvirus | Unclassified | Autographiviridae | Sphingomonas |
| NC_048028 | Gordonia phage Ronaldo | Gordonia phage Ronaldo Gordonia virus Ronaldo Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68389 | 50.163 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048027 | Gordonia phage Fryberger | Gordonia phage Fryberger Gordonia virus Fryberger Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67234 | 50.178 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048022 | Eggerthella phage PMBT5 | Eggerthella phage PMBT5 Eggerthella virus PMBT5 Lentavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30930 | 51.3 | Lentavirus | Unclassified | Siphoviridae | Eggerthella |
| NC_048021 | Gordonia phage Daredevil | Gordonia phage Daredevil Gordonia virus DareDevil Daredevilvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90490 | 64.843 | Daredevilvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048017 | Microbacterium phage Eden | Microbacterium phage Eden Microbacterium virus Eden Edenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40833 | 66.282 | Edenvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_048016 | Erwinia phage Wellington | Erwinia phage Wellington Erwinia virus Wellington Wellingtonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244950 | 52.115 | Wellingtonvirus | Unclassified | Myoviridae | Erwinia |
| NC_048013 | Salmonella phage S124 | Salmonella phage S124 Salmonella virus S124 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112564 | 40.119 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048012 | Salmonella phage S147 | Salmonella phage S147 Salmonella virus S147 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111447 | 39.985 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048011 | Salmonella phage S133 | Salmonella phage S133 Salmonella virus S133 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110926 | 40.061 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048010 | Salmonella phage S132 | Salmonella phage S132 Salmonella virus S132 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116832 | 39.944 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048009 | Salmonella phage S131 | Salmonella phage S131 Salmonella virus S131 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110091 | 39.216 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048008 | Salmonella phage S126 | Salmonella phage S126 Salmonella virus S126 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111999 | 40.1 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048007 | Salmonella phage S116 | Salmonella phage S116 Salmonella virus S116 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110664 | 40.085 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048006 | Salmonella phage S114 | Salmonella phage S114 Salmonella virus S114 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110926 | 40.062 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048005 | Salmonella phage S113 | Salmonella phage S113 Salmonella virus S113 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112582 | 39.96 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048001 | Salmonella phage SP3 | Salmonella phage SP3 Salmonella virus SP3 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109306 | 39.022 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048000 | Salmonella phage LVR16A | Salmonella phage LVR16A Salmonella virus LVR16A Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111601 | 40.078 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_047999 | Salmonella phage SH9 | Salmonella phage SH9 Salmonella virus SH9 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111607 | 40.145 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_047997 | Pseudomonas phage PFP1 | Pseudomonas phage PFP1 Pseudomonas virus PFP1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40914 | 55.815 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047995 | Erwinia phage vB_EamM_Alexandra | Erwinia phage vB_EamM_Alexandra Erwinia virus Alexandra Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266532 | 50.126 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| NC_047992 | Microbacterium phage Zeta1847 | Microbacterium phage Zeta1847 Microbacterium virus Zeta1847 Zetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47921 | 71.415 | Zetavirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047991 | Gordonia phage Trine | Gordonia phage Trine Gordonia virus Trine Trinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43758 | 67.816 | Trinevirus | Unclassified | Siphoviridae | Gordonia |
| NC_047990 | Gordonia phage Suzy | Gordonia phage Suzy Gordonia virus Suzy Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67118 | 59.094 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| NC_047986 | Microbacterium phage Krampus | Microbacterium phage Krampus Microbacterium virus Krampus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56708 | 63.799 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047982 | Ralstonia phage RPSC1 | Ralstonia phage RPSC1 Ralstonia virus RPSC1 Stompelvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39628 | 61.555 | Stompelvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_047981 | Pseudomonas phage PspYZU08 | Pseudomonas phage PspYZU08 Pseudomonas virus PspYZU08 Pijolavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40184 | 55.261 | Pijolavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047979 | Escherichia phage Gostya9 | Escherichia phage Gostya9 Escherichia virus Gostya9 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101665 | 39.371 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047977 | Microbacterium phage Hendrix | Microbacterium phage Hendrix Microbacterium virus Hendrix Rogerhendrixvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97757 | 66.075 | Rogerhendrixvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047975 | Microbacterium phage Squash | Microbacterium phage Squash Microbacterium virus Squash Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62312 | 66.788 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047973 | Microbacterium phage Hyperion | Microbacterium phage Hyperion Microbacterium virus Hyperion Squashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61769 | 67.009 | Squashvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_047966 | Aeromonas phage 25AhydR2PP | Aeromonas phage 25AhydR2PP Aeromonas virus 25AhydR2PP Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 54.897 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_047965 | Pseudomonas phage 22PfluR64PP | Pseudomonas phage 22PfluR64PP Pseudomonas virus 22PfluR64PP Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40822 | 55.806 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047964 | Dickeya phage Ninurta | Dickeya phage Ninurta Dickeya virus Ninurta Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39699 | 50.986 | Ningirsuvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_047963 | Pectobacterium phage Jarilo | Pectobacterium phage Jarilo Pectobacterium virus Jarilo Jarilovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40552 | 50.094 | Jarilovirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_047962 | Xanthomonas phage Carpasina | Xanthomonas phage Carpasina Xanthomonas virus Carpasina Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61939 | 52.356 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| NC_047961 | Dickeya phage Dagda | Dickeya phage Dagda Dickeya virus Dagda Aarhusvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40484 | 48.113 | Aarhusvirus | Studiervirinae | Autographiviridae | Dickeya |
| NC_047957 | Pseudomonas phage Alpheus | Pseudomonas phage Alpheus Pseudomonas virus Alpheus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45756 | 57.824 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| NC_047956 | Pseudomonas phage Achelous | Pseudomonas phage Achelous Pseudomonas virus Achelous Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46585 | 57.345 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| NC_047955 | Pseudomonas phage Nerthus | Pseudomonas phage Nerthus Pseudomonas virus Nerthus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45779 | 58.505 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| NC_047954 | Pseudomonas phage Njord | Pseudomonas phage Njord Pseudomonas virus Njord Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46644 | 56.62 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| NC_047950 | Aeromonas phage AhSzw-1 | Aeromonas phage AhSzw-1 Aeromonas virus AhSzw1 Shenzhenvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115739 | 43.82 | Shenzhenvirus | Unclassified | Demerecviridae | Aeromonas |
| NC_047949 | Aeromonas phage AhSzq-1 | Aeromonas phage AhSzq-1 Aeromonas virus AhSzq1 Shenzhenvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112558 | 43.861 | Shenzhenvirus | Unclassified | Demerecviridae | Aeromonas |
| NC_047941 | Salmonella phage vB_SpuP_Spp16 | Salmonella phage vB_SpuP_Spp16 Salmonella virus Spp16 Panjvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41832 | 39.458 | Panjvirus | Melnykvirinae | Autographiviridae | Salmonella |
| NC_047932 | Klebsiella phage vB_KpnM_KpS110 | Klebsiella phage vB_KpnM_KpS110 Klebsiella virus KpS110 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156801 | 46.143 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_047931 | Lactobacillus phage Maenad | Lactobacillus phage Maenad Lactobacillus virus Maenad Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78646 | 39.074 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_047930 | Ralstonia phage RsoP1IDN | Ralstonia phage RsoP1IDN Ralstonia virus RsoP1IDN Higashivirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41135 | 62.747 | Higashivirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_047921 | Gordonia phage Flakey | Gordonia phage Flakey Gordonia virus Flakey Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76889 | 58.924 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_047917 | Erwinia phage vB_EamP-S2 | Erwinia phage vB_EamP-S2 Erwinia virus S2 Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45495 | 49.792 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| NC_047903 | Vibrio phage VEN | Vibrio phage VEN Vibrio virus VEN Trungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44603 | 43.531 | Trungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| NC_047902 | Pseudomonas phage PMBT3 | Pseudomonas phage PMBT3 Pseudomonas virus PMBT3 Maxrubnervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87196 | 60.37 | Maxrubnervirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_047901 | Klebsiella phage Menlow | Klebsiella phage Menlow Klebsiella virus Menlow Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157281 | 46.422 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_047900 | Klebsiella phage May | Klebsiella phage May Klebsiella virus May Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159631 | 46.768 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_047894 | Pseudomonas phage uligo | Pseudomonas phage uligo Pseudomonas virus uligo Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47868 | 56.301 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| NC_047892 | Gordonia phage BirksAndSocks | Gordonia phage BirksAndSocks Gordonia virus Birksandsocks Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77354 | 58.881 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_047888 | Ralstonia phage DU_RP_I | Ralstonia phage DU_RP_I Ralstonia virus DURPI Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40657 | 58.789 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_047887 | Pectobacterium phage DU_PP_V | Pectobacterium phage DU_PP_V Pectobacterium virus DUPPV Hongcheonvirus Mccorquodalevirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106185 | 39.879 | Hongcheonvirus | Mccorquodalevirinae | Demerecviridae | Pectobacterium |
| NC_047885 | Escherichia phage chee24 | Escherichia phage chee24 Escherichia virus chee24 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120622 | 39.263 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047884 | Escherichia phage saus132 | Escherichia phage saus132 Escherichia virus saus132 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.745 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047882 | Vibrio phage JSF12 | Vibrio phage JSF12 Vibrio virus JSF12 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111671 | 42.713 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_047880 | Erwinia phage vB_EamM_Y3 | Erwinia phage vB_EamM_Y3 Erwinia virus Y3 Sasquatchvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261365 | 47.178 | Sasquatchvirus | Unclassified | Myoviridae | Erwinia |
| NC_047873 | Pseudomonas phage UNO-SLW1 | Pseudomonas phage UNO-SLW1 Pseudomonas virus UNOSLW1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39215 | 57.899 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_047870 | Rhizobium phage RHEph09 | Rhizobium phage RHEph09 Rhizobium virus RHEph09 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45962 | 49.304 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| NC_047869 | Rhizobium phage RHEph08 | Rhizobium phage RHEph08 Rhizobium virus RHEph08 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43619 | 49.435 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| NC_047868 | Rhizobium phage RHEph02 | Rhizobium phage RHEph02 Rhizobium virus RHEph02 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46486 | 49.35 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| NC_047865 | Aeromonas phage CF7 | Aeromonas phage CF7 Aeromonas virus CF7 Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42439 | 58.95 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_047860 | Caulobacter phage Lullwater | Caulobacter phage Lullwater Caulobacter virus Lullwater Lullwatervirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44638 | 54.653 | Lullwatervirus | Unclassified | Autographiviridae | Caulobacter |
| NC_047859 | Salmonella phage SP01 | Salmonella phage SP01 Salmonella virus SP01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 117842 | 38.982 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_047846 | Agrobacterium phage Atu_ph03 | Agrobacterium phage Atu_ph03 Agrobacterium virus Atuph03 Atuphduovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45175 | 54.734 | Atuphduovirus | Unclassified | Autographiviridae | Agrobacterium |
| NC_047845 | Agrobacterium phage Atu_ph02 | Agrobacterium phage Atu_ph02 Agrobacterium virus Atuph02 Atuphduovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45423 | 54.829 | Atuphduovirus | Unclassified | Autographiviridae | Agrobacterium |
| NC_047840 | Xanthomonas phage phi Xc10 | Xanthomonas phage phi Xc10 Xanthomonas virus Xc10 Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44597 | 62.435 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| NC_047836 | Marinomonas phage CB5A | Marinomonas phage CB5A Marinomonas virus CB5A Murciavirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45385 | 43.217 | Murciavirus | Colwellvirinae | Autographiviridae | Marinomonas |
| NC_047835 | Escherichia phage OSYSP | Escherichia phage OSYSP Escherichia virus OSYSP Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110901 | 39.162 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047834 | Alteromonas virus vB_AspP-H4/4 | Alteromonas virus vB_AspP-H4/4 Alteromonas virus H4-4 Foturvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47631 | 40.761 | Foturvirus | Unclassified | Autographiviridae | Alteromonas |
| NC_047831 | Pasteurella phage vB_PmuP_PHB02 | Pasteurella phage vB_PmuP_PHB02 Pasteurella virus PHB02 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38670 | 40.846 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| NC_047822 | Escherichia phage phiLLS | Escherichia phage phiLLS Escherichia virus phiLLS Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107263 | 38.989 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047821 | Marinomonas phage CPP1m | Marinomonas phage CPP1m Marinomonas virus CPP1m Murciavirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44245 | 43.363 | Murciavirus | Colwellvirinae | Autographiviridae | Marinomonas |
| NC_047819 | Pectobacterium phage PPWS4 | Pectobacterium phage PPWS4 Pectobacterium virus PPWS4 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39906 | 48.905 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_047815 | Erwinia phage vB_EamM_Yoloswag | Erwinia phage vB_EamM_Yoloswag Erwinia virus Yoloswag Yoloswagvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 259700 | 46.915 | Yoloswagvirus | Unclassified | Myoviridae | Erwinia |
| NC_047803 | Vibrio phage Vp670 | Vibrio phage Vp670 Vibrio virus Vp670 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43121 | 43.401 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| NC_047802 | Pectobacterium phage PP99 | Pectobacterium phage PP99 Pectobacterium virus PP99 Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 45.545 | Gajwadongvirus | Unclassified | Autographiviridae | Pectobacterium |
| NC_047778 | Pectobacterium phage PP2 | Pectobacterium phage PP2 Pectobacterium virus PP2 Wanjuvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41841 | 51.622 | Wanjuvirus | Melnykvirinae | Autographiviridae | Pectobacterium |
| NC_047776 | Escherichia phage ESCO5 | Escherichia phage ESCO5 Escherichia virus ESCO5 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149312 | 38.961 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_047770 | Escherichia phage ESCO13 | Escherichia phage ESCO13 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149813 | 39.125 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_047762 | Xanthomonas phage XAJ24 | Xanthomonas phage XAJ24 Xanthomonas virus XAJ24 Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44862 | 58.205 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| NC_047758 | Pectobacterium phage PPWS1 | Pectobacterium phage PPWS1 Pectobacterium virus PPWS1 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44539 | 51.063 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_047756 | Caulobacter phage Sansa | Caulobacter phage Sansa Caulobacter virus Sansa Sansavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80297 | 65.751 | Sansavirus | Unclassified | Siphoviridae | Caulobacter |
| NC_047754 | Escherichia phage phiAPCEc03 | Escherichia phage phiAPCEc03 Escherichia virus phiAPCEc03 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103737 | 38.933 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047751 | Ralstonia phage phiITL-1 | Ralstonia phage phiITL-1 Ralstonia virus ITL1 Serkorvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38731 | 62.751 | Serkorvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_047749 | Escherichia phage DT571/2 | Escherichia phage DT571/2 Escherichia virus DT5712 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108418 | 39.678 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_047745 | Listeria phage LP-124 | Listeria phage LP-124 Listeria virus LP064 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135764 | 35.874 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_047741 | Vibrio phage JSF7 | Vibrio phage JSF7 Vibrio virus JSF7 Tawavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46318 | 48.307 | Tawavirus | Unclassified | Autographiviridae | Vibrio |
| NC_047737 | Vibrio phage AS51 | Vibrio phage AS51 Vibrio virus AS51 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42542 | 43.47 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| NC_042564 | Escherichia virus BF23 | Escherichia virus BF23 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14504 | 40.306 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_042355 | Corynebacterium phage phi674 | Corynebacterium phage phi674 Corynebacterium virus phi674 Ikedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43193 | 50.749 | Ikedavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_042354 | Corynebacterium phage phi673 | Corynebacterium phage phi673 Corynebacterium virus phi673 Ikedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44530 | 51.087 | Ikedavirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_042353 | Salinibacter virus SRUTV1 | Salinibacter virus SRUTV1 Kairosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50672 | 59.445 | Kairosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_042352 | Salinibacter virus M31CR41-2 | Salinibacter virus M31CR41-2 Kairosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51889 | 59.658 | Kairosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_042350 | Salinibacter virus M8CR30-2 | Salinibacter virus M8CR30-2 Holosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35099 | 64.592 | Holosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_042349 | Salinibacter virus M8CC19 | Salinibacter virus M8CC19 Kryptosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53812 | 52.99 | Kryptosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_042348 | Salinibacter virus M1EM1 | Salinibacter virus M1EM1 Holosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35883 | 64.66 | Holosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| NC_042345 | Xylella phage Salvo | Xylella phage Salvo Xylella virus Salvo Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55601 | 63.04 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| NC_042344 | Xylella phage Sano | Xylella phage Sano Xylella virus Sano Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56147 | 62.361 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| NC_042338 | Mycobacterium virus MacnCheese | Mycobacterium virus MacnCheese Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61567 | 67.283 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042332 | Mycobacterium virus JAWS | Mycobacterium virus JAWS Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59749 | 66.617 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042325 | Mycobacterium virus Pixie | Mycobacterium virus Pixie Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61147 | 67.303 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042138 | Klebsiella phage PMBT1 | Klebsiella phage PMBT1 Klebsiella virus PMBT1 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175206 | 41.882 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_042137 | Acinetobacter phage Loki | Acinetobacter phage Loki Acinetobacter virus Loki Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41308 | 44.347 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| NC_042134 | Enterococcus phage AUEF3 | Enterococcus phage AUEF3 Enterococcus virus AUEF3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41257 | 34.452 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042132 | Gordonia phage Sour | Gordonia phage Sour Gordonia virus Sour Sourvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61670 | 68.009 | Sourvirus | Unclassified | Siphoviridae | Gordonia |
| NC_042131 | Pectobacterium phage vB_PatP_CB5 | Pectobacterium phage vB_PatP_CB5 Pectobacterium virus CB5 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44549 | 48.984 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_042127 | Enterococcus phage vB_EfaS_AL2 | Enterococcus phage vB_EfaS_AL2 Enterococcus virus AL2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40836 | 34.514 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042124 | Acinetobacter phage vB_AbaP_D2 | Acinetobacter phage vB_AbaP_D2 Acinetobacter virus D2 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39964 | 39.233 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_042120 | Pantoea phage vB_PagS_Vid5 | Pantoea phage vB_PagS_Vid5 Pantoea virus Vid5 Vidquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61437 | 48.821 | Vidquintavirus | Unclassified | Siphoviridae | Pantoea |
| NC_042119 | Microbacterium phage Pikmin | Microbacterium phage Pikmin Microbacterium virus Pikmin Pikminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39307 | 61.269 | Pikminvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042118 | Microbacterium phage Koji | Microbacterium phage Koji Microbacterium virus Koji Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39403 | 64.173 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042117 | Microbacterium phage Golden | Microbacterium phage Golden Microbacterium virus Golden Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39640 | 64.084 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042111 | Microbacterium phage Ilzat | Microbacterium phage Ilzat Microbacterium virus Ilzat Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41525 | 63.485 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042110 | Microbacterium phage Hamlet | Microbacterium phage Hamlet Microbacterium virus Hamlet Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41934 | 63.402 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042109 | Microbacterium phage Eleri | Microbacterium phage Eleri Microbacterium virus Eleri Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40366 | 61.975 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042108 | Gordonia phage Brandonk123 | Gordonia phage Brandonk123 Gordonia virus Brandonk123 Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58842 | 67.287 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_042105 | Streptomyces phage BillNye | Streptomyces phage BillNye Streptomyces virus BillNye Wilnyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127084 | 52.833 | Wilnyevirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042104 | Pseudomonas phage PollyC | Pseudomonas phage PollyC Pseudomonas virus PollyC Pollyceevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41651 | 57.595 | Pollyceevirus | Unclassified | Autographiviridae | Pseudomonas |
| NC_042102 | Gordonia phage Troje | Gordonia phage Troje Gordonia virus Troje Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45909 | 60.378 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| NC_042101 | Enterococcus phage PMBT2 | Enterococcus phage PMBT2 Enterococcus virus PMBT2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41489 | 34.691 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042099 | Microbacterium phage Dismas | Microbacterium phage Dismas Microbacterium virus Dismas Dismasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41593 | 69.632 | Dismasvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_042098 | Erwinia phage vB_EamM_Desertfox | Erwinia phage vB_EamM_Desertfox Erwinia virus Desertfox Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 272458 | 49.877 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_042095 | Vibrio phage Thalassa | Vibrio phage Thalassa Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128602 | 40.24 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_042094 | Vibrio phage Ceto | Vibrio phage Ceto Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128241 | 39.927 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_042093 | Klebsiella phage Sugarland | Klebsiella phage Sugarland Klebsiella virus Sugarland Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111103 | 44.958 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| NC_042092 | Pseudomonas phage tabernarius | Pseudomonas phage tabernarius Pseudomonas virus tabernarius Tabernariusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67498 | 52.296 | Tabernariusvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_042090 | Proteus phage PM135 | Proteus phage PM135 Proteus virus PM135 Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104329 | 38.369 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| NC_042089 | Gordonia phage Mahdia | Gordonia phage Mahdia Gordonia virus Mahdia Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42475 | 67.249 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| NC_042088 | Gordonia phage Gustav | Gordonia phage Gustav Gordonia virus Gustav Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44408 | 67.855 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| NC_042087 | Corynebacterium phage Zion | Corynebacterium phage Zion Corynebacterium virus Zion Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66392 | 52.298 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_042086 | Corynebacterium phage Darwin | Corynebacterium phage Darwin Corynebacterium virus Darwin Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67743 | 52.082 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_042085 | Corynebacterium phage C3PO | Corynebacterium phage C3PO Corynebacterium virus C3PO Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67383 | 52.305 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_042083 | Gluconobacter phage GC1 | Gluconobacter phage GC1 Gluconobacter virus GC1 Gammatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 16523 | 50.475 | Gammatectivirus | Unclassified | Tectiviridae | Gluconobacter |
| NC_042082 | Stenotrophomonas phage vB_SmaS_DLP_5 | Stenotrophomonas phage vB_SmaS_DLP_5 Stenotrophomonas virus DLP5 Delepquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96542 | 58.369 | Delepquintavirus | Unclassified | Siphoviridae | Stenotrophomonas |
| NC_042074 | Vibrio phage JSF10 | Vibrio phage JSF10 Vibrio virus JSF10 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111671 | 42.705 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_042050 | Mycobacterium phage Godines | Mycobacterium phage Godines Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67277 | 69.025 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_042049 | Rhodobacter phage RcCronus | Rhodobacter phage RcCronus Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35985 | 65.427 | Cronusvirus | Unclassified | Siphoviridae | Rhodobacter |
| NC_042047 | Serratia phage vB-Sru-IME250 | Serratia phage vB-Sru-IME250 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154938 | 47.351 | Taipeivirus | Unclassified | Ackermannviridae | Serratia |
| NC_042045 | Gordonia phage Lennon | Gordonia phage Lennon Gordonia virus Lennon Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60237 | 67.364 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_042040 | Rhodococcus phage Trina | Rhodococcus phage Trina Rhodococcus virus Trina Trinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139262 | 44.724 | Trinavirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_042035 | Mycobacterium phage Zemanar | Mycobacterium phage Zemanar Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71092 | 68.876 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_042034 | Mycobacterium phage ChrisnMich | Mycobacterium phage ChrisnMich Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70428 | 69.075 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_042033 | Mycobacterium phage Send513 | Mycobacterium phage Send513 Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71547 | 55.998 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042032 | Mycobacterium phage Marvin | Mycobacterium phage Marvin Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65100 | 63.424 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042031 | Mycobacterium phage Anaya | Mycobacterium phage Anaya Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60835 | 66.394 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_042023 | Enterococcus phage phiSHEF5 | Enterococcus phage phiSHEF5 Enterococcus virus phiSHEF5 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41598 | 34.747 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042022 | Enterococcus phage phiSHEF4 | Enterococcus phage phiSHEF4 Enterococcus virus phiSHEF4 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41081 | 34.724 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042021 | Enterococcus phage phiSHEF2 | Enterococcus phage phiSHEF2 Enterococcus virus phiSHEF2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41712 | 34.554 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_042020 | Arthrobacter phage Adat | Arthrobacter phage Adat Arthrobacter virus Adat Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45428 | 45.749 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| NC_042019 | Aeromonas phage AS-gz | Aeromonas phage AS-gz Aeromonas virus Asgz Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162422 | 41.127 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_042018 | Erwinia phage vB_EamM_RisingSun | Erwinia phage vB_EamM_RisingSun Erwinia virus Risingsun Risingsunvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235108 | 48.318 | Risingsunvirus | Unclassified | Myoviridae | Erwinia |
| NC_042016 | Streptomyces phage DrGrey | Streptomyces phage DrGrey Strepomyces virus Drgrey Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56076 | 59.48 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_042015 | Arthrobacter phage Shade | Arthrobacter phage Shade Arthrobacter virus Shade Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50930 | 61.082 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_042014 | Arthrobacter phage Piccoletto | Arthrobacter phage Piccoletto Arthrobacter virus Piccoletto Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49808 | 63.556 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_042007 | Acinetobacter phage vB_ApiP_P2 | Acinetobacter phage vB_ApiP_P2 Acinetobacter virus P2 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41514 | 39.334 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_042006 | Acinetobacter phage vB_ApiP_P1 | Acinetobacter phage vB_ApiP_P1 Acinetobacter virus P1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41208 | 39.196 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_042004 | Acinetobacter phage vB_AbaP_B3 | Acinetobacter phage vB_AbaP_B3 Acinetobacter virus B3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40598 | 39.275 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_042003 | Acinetobacter phage vB_AbaP_B1 | Acinetobacter phage vB_AbaP_B1 Acinetobacter virus B1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40879 | 39.145 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_042002 | Arthrobacter phage Abidatro | Arthrobacter phage Abidatro Arthrobacter virus Abidatro Galaxyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39122 | 68.545 | Galaxyvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_042001 | Arthrobacter phage Molivia | Arthrobacter phage Molivia Arthrobacteria virus Molivia Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58247 | 53.462 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041999 | Arthrobacter phage Beans | Arthrobacter phage Beans Arthrobacter virus Beans Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49797 | 63.566 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041997 | Acidovorax phage ACP17 | Acidovorax phage ACP17 Acidovorax virus ACP17 Busanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156281 | 58.738 | Busanvirus | Unclassified | Myoviridae | Acidovorax |
| NC_041994 | Pseudomonas phage Noxifer | Pseudomonas phage Noxifer Pseudomonas virus Noxifer Noxifervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 278136 | 53.526 | Noxifervirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041993 | Mycobacterium phage Sebata | Mycobacterium phage Sebata Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155286 | 64.782 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_041981 | Klebsiella phage Miro | Klebsiella phage Miro Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176055 | 41.779 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_041980 | Enterobacter phage phiEap-3 | Enterobacter phage phiEap-3 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175814 | 42.005 | Slopekvirus | Tevenvirinae | Myoviridae | Enterobacter |
| NC_041975 | Erwinia phage vB_EamM_Special G | Erwinia phage vB_EamM_Special G Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273224 | 49.791 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_041974 | Erwinia phage vB_EamM_Simmy50 | Erwinia phage vB_EamM_Simmy50 Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271088 | 49.855 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_041973 | Erwinia phage vB_EamM_RAY | Erwinia phage vB_EamM_RAY Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271182 | 49.872 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_041972 | Erwinia phage vB_EamM_Deimos-Minion | Erwinia phage vB_EamM_Deimos-Minion Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273501 | 49.861 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_041967 | Acinetobacter phage WCHABP5 | Acinetobacter phage WCHABP5 Acinetobacter virus WCHABP5 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40409 | 39.395 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_041964 | Pseudomonas phage phiR18 | Pseudomonas phage phiR18 Kochitakasuvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63560 | 60.349 | Kochitakasuvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_041963 | Rhodobacter phage RcSpartan | Rhodobacter phage RcSpartan Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44194 | 54.885 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| NC_041962 | Tsukamurella phage TIN4 | Tsukamurella phage TIN4 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76268 | 59.29 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| NC_041961 | Arthrobacter phage Tank | Arthrobacter phage Tank Tankvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67592 | 62.87 | Tankvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041960 | Enterococcus phage SAP6 | Enterococcus phage SAP6 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58619 | 39.996 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_041959 | Enterococcus phage IMEEF1 | Enterococcus phage IMEEF1 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57081 | 40.045 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_041952 | Arthrobacter phage Gordon | Arthrobacter phage Gordon Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58279 | 49.841 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041951 | Arthrobacter phage CaptnMurica | Arthrobacter phage CaptnMurica Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58159 | 49.588 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041950 | Gordonia phage Katyusha | Gordonia phage Katyusha Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75380 | 59.461 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| NC_041949 | Arthrobacter phage Amigo | Arthrobacter phage Amigo Amigovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59173 | 52.948 | Amigovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041948 | Arthrobacter phage Circum | Arthrobacter phage Circum Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58353 | 45.175 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041947 | Arthrobacter phage Laroye | Arthrobacter phage Laroye Laroyevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60005 | 64.788 | Laroyevirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041946 | Arthrobacter phage Wayne | Arthrobacter phage Wayne Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44371 | 61.139 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041945 | Arthrobacter phage Pumancara | Arthrobacter phage Pumancara Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42830 | 61.749 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041944 | Arthrobacter phage Preamble | Arthrobacter phage Preamble Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43374 | 60.691 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041943 | Arthrobacter phage Korra | Arthrobacter phage Korra Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43707 | 61.075 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041942 | Arthrobacter phage Joann | Arthrobacter phage Joann Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44183 | 60.725 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041941 | Arthrobacter phage HunterDalle | Arthrobacter phage HunterDalle Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43336 | 61.561 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041940 | Arthrobacter phage Glenn | Arthrobacter phage Glenn Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44389 | 60.819 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041939 | Arthrobacter phage DrRobert | Arthrobacter phage DrRobert Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42601 | 60.611 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041938 | Arthrobacter phage Bennie | Arthrobacter phage Bennie Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43075 | 61.416 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041934 | Pseudomonas phage LKO4 | Pseudomonas phage LKO4 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61818 | 64.426 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_041933 | Arthrobacter phage Sonny | Arthrobacter phage Sonny Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50909 | 61.091 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041932 | Arthrobacter phage Martha | Arthrobacter phage Martha Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51027 | 60.954 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041931 | Arthrobacter phage Jawnski | Arthrobacter phage Jawnski Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49419 | 63.439 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041930 | Arthrobacter phage Brent | Arthrobacter phage Brent Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49879 | 63.425 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041927 | Sphingobium phage Lacusarx | Sphingobium phage Lacusarx Sphingobium virus Lacusarx Lacusarxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130138 | 60.175 | Lacusarxvirus | Unclassified | Siphoviridae | Sphingobium |
| NC_041921 | Dinoroseobacter phage vB_DshS-R5C | Dinoroseobacter phage vB_DshS-R5C Dinoroseobacter virus D5C Nanhaivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77874 | 61.546 | Nanhaivirus | Unclassified | Siphoviridae | Dinoroseobacter |
| NC_041918 | Geobacillus phage TP-84 | Geobacillus phage TP-84 Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47703 | 45.542 | Saundersvirus | Unclassified | Siphoviridae | Geobacillus |
| NC_041917 | Serratia phage BF | Serratia phage BF Serratia virus BF Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 357154 | 34.423 | Eneladusvirus | Unclassified | Myoviridae | Serratia |
| NC_041915 | Acinetobacter phage vB_AbaP_AS11 | Acinetobacter phage vB_AbaP_AS11 Acinetobacter virus AS11 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41642 | 39.292 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_041914 | Acinetobacter phage vB_AbaP_AS12 | Acinetobacter phage vB_AbaP_AS12 Acinetobacter virus AS12 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41402 | 39.315 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_041913 | Providencia phage vB_PreS_PR1 | Providencia phage vB_PreS_PR1 Providencia virus PR1 Priunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118537 | 39.518 | Priunavirus | Unclassified | Siphoviridae | Providencia |
| NC_041912 | Aquamicrobium phage P14 | Aquamicrobium phage P14 Aquamicrobium virus P14 Aqualcavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40551 | 57.833 | Aqualcavirus | Unclassified | Autographiviridae | Aquamicrobium |
| NC_041911 | Ralstonia phage RP12 | Ralstonia phage RP12 Ralstonia virus RP12 Ripduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279845 | 53.404 | Ripduovirus | Unclassified | Myoviridae | Ralstonia |
| NC_041908 | Rhizobium phage RHEph04 | Rhizobium phage RHEph04 Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53018 | 56.422 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| NC_041905 | Acinetobacter phage SH-Ab 15519 | Acinetobacter phage SH-Ab 15519 Acinetobacter virus SH-Ab 15519 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40493 | 39.464 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_041901 | Mycobacterium phage Lukilu | Mycobacterium phage Lukilu Mycobacterium virus Lukilu Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157034 | 64.656 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_041899 | Klebsiella phage vB_Kpn_IME260 | Klebsiella phage vB_Kpn_IME260 Klebsiella virus IME260 Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123490 | 45.372 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| NC_041889 | Streptomyces phage Rima | Streptomyces phage Rima Strepomyces virus Rima Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56168 | 59.557 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| NC_041887 | Rhodococcus phage Weasels2 | Rhodococcus phage Weasels2 Rhodococcus virus Weasel Weaselvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134973 | 41.293 | Weaselvirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_041885 | Pseudomonas phage VSW-3 | Pseudomonas phage VSW-3 Pseudomonas virus VSW3 Napahaivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40556 | 57.54 | Napahaivirus | Unclassified | Autographiviridae | Pseudomonas |
| NC_041884 | Acinetobacter phage vB_AbaM_ME3 | Acinetobacter phage vB_AbaM_ME3 Acinetobacter virus ME3 Metrivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 234900 | 30.764 | Metrivirus | Unclassified | Myoviridae | Acinetobacter |
| NC_041881 | Pseudomonas phage pf16 | Pseudomonas phage pf16 Pseudomonas virus pf16 Chakrabartyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158136 | 52.653 | Chakrabartyvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041880 | Pseudomonas phage phiPMW | Pseudomonas phage phiPMW Pseudomonas virus PMW Plaisancevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103218 | 45.149 | Plaisancevirus | Unclassified | Myoviridae | Pseudomonas |
| NC_041879 | Bacillus phage Mgbh1 | Bacillus phage Mgbh1 Bacillus virus Mgbh1 Magadivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58951 | 45.414 | Magadivirus | Unclassified | Siphoviridae | Bacillus |
| NC_041877 | Pseudomonas phage SM1 | Pseudomonas phage SM1 Pseudomonas virus SM1 Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93191 | 55.24 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_041876 | Arthrobacter phage Galaxy | Arthrobacter phage Galaxy Arthrobacter virus Galaxy Galaxyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37809 | 68.433 | Galaxyvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_041875 | Arthrobacter phage Jasmine | Arthrobacter phage Jasmine Arthrobacter virus Jasmine Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46723 | 45.896 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| NC_041872 | Flavobacterium phage FpV4 | Flavobacterium phage FpV4 Flavobacterium virus Fpv4 Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88308 | 27.218 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| NC_041862 | Listeria phage LP-064 | Listeria phage LP-064 Listeria virus LP064 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135279 | 35.926 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_041861 | Enterococcus phage VD13 | Enterococcus phage VD13 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55073 | 40.003 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_041858 | Bacillus phage Taylor | Bacillus phage Taylor Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49492 | 42.294 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_030937 | Xanthomonas phage f30-Xaj | Xanthomonas phage f30-Xaj Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44262 | 59.882 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| NC_030928 | Xanthomonas phage f20-Xaj | Xanthomonas phage f20-Xaj Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43851 | 59.83 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| NC_029005 | Streptomyces phage phiSAJS1 | Streptomyces phage phiSAJS1 unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56451 | 68.33 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_011269 | Mycobacterium phage ScottMcG | Mycobacterium phage ScottMcG Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154017 | 64.829 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_031224 | Arthrobacter phage Mudcat | Arthrobacter phage Mudcat Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59443 | 45.104 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_028987 | Acinetobacter phage IME-200 | Acinetobacter phage IME-200 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41243 | 39.313 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_031045 | Salmonella phage GG32 | Salmonella phage GG32 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157855 | 44.545 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_031933 | Salmonella phage 118970_sal2 | Salmonella phage 118970_sal2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114180 | 40.278 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_031931 | Flavobacterium phage Fpv2 | Flavobacterium phage Fpv2 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89010 | 26.995 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| NC_031917 | Pseudoalteromonas phage BS5 | Pseudoalteromonas phage BS5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39949 | 40.597 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| NC_031914 | Flavobacterium phage Fpv1 | Flavobacterium phage Fpv1 Flavobacterium virus Fpv1 Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88994 | 26.992 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| NC_031912 | Flavobacterium phage Fpv20 | Flavobacterium phage Fpv20 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89937 | 26.944 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| NC_031908 | Pseudoalteromonas phage PH1 | Pseudoalteromonas phage PH1 unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42685 | 42.237 | Kafunavirus | Unclassified | Podoviridae | Pseudoalteromonas |
| NC_031904 | Flavobacterium phage Fpv3 | Flavobacterium phage Fpv3 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88421 | 27.222 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| NC_022331 | Mycobacterium phage Bane1 | Mycobacterium phage Bane1 Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69309 | 68.867 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_031252 | Arthrobacter phage BarretLemon | Arthrobacter phage BarretLemon Arthrobacter virus BarretLemon Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 60.903 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_031247 | Gordonia phage BetterKatz | Gordonia phage BetterKatz Gordonia virus BetterKatz Betterkatzvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50636 | 67.114 | Betterkatzvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031245 | Bacillus phage SP-15 | Bacillus phage SP-15 Bacillus virus SP15 Thornevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221908 | 38.608 | Thornevirus | Unclassified | Myoviridae | Bacillus |
| NC_031239 | Gordonia phage Vivi2 | Gordonia phage Vivi2 Gordonia virus Vivi2 Vividuovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59337 | 67.133 | Vividuovirus | Unclassified | Siphoviridae | Gordonia |
| NC_031230 | Gordonia phage Yvonnetastic | Gordonia phage Yvonnetastic Gordonia virus Yvonnetastic Yvonnevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98136 | 59.749 | Yvonnevirus | Unclassified | Siphoviridae | Gordonia |
| NC_031234 | Gordonia phage Emalyn | Gordonia phage Emalyn Gordonia virus Emalyn Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43982 | 61.214 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031231 | Arthrobacter phage KellEzio | Arthrobacter phage KellEzio Kelleziovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58871 | 63.344 | Kelleziovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_031280 | Acinetobacter phage vB_AbaM_phiAbaA1 | Acinetobacter phage vB_AbaM_phiAbaA1 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104906 | 37.774 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_031254 | Arthrobacter phage Kitkat | Arthrobacter phage Kitkat Kelleziovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58560 | 63.391 | Kelleziovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_031128 | Salmonella phage vB-SalM-PM10 | Salmonella phage vB-SalM-PM10 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158081 | 44.608 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_031112 | Gordonia phage Phinally | Gordonia phage Phinally Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59265 | 68.437 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_031109 | Gordonia phage Jumbo | Gordonia phage Jumbo Gordonia virus Jumbo Gorjumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78302 | 54.452 | Gorjumvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031102 | Mycobacterium phage Sneeze | Mycobacterium phage Sneeze unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42429 | 66.455 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031096 | Pectobacterium phage PP90 | Pectobacterium phage PP90 Pectobacterium virus PP90 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44570 | 48.889 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_031080 | Mycobacterium phage Tonenili | Mycobacterium phage Tonenili unclassified Bixzunavirus Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160985 | 64.09 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_031074 | Gordonia phage Bantam | Gordonia phage Bantam Gordonia virus Bantam Bantamvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92580 | 64.705 | Bantamvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031043 | Erwinia phage vB_EamM_Phobos | Erwinia phage vB_EamM_Phobos unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229501 | 49.137 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| NC_031042 | Salmonella phage NR01 | Salmonella phage NR01 Salmonella virus NR01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111325 | 38.838 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_031022 | Shigella phage SHSML-45 | Shigella phage SHSML-45 Shigella virus SHSML45 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108050 | 38.738 | Tequintavirus | Markadamsvirinae | Demerecviridae | Shigella |
| NC_031014 | Pseudomonas phage Andromeda | Pseudomonas phage Andromeda Pseudomonas virus Andromeda Bifseptvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40008 | 58.233 | Bifseptvirus | Unclassified | Autographiviridae | Pseudomonas |
| NC_031010 | Erwinia phage vB_EamM_Kwan | Erwinia phage vB_EamM_Kwan Wellingtonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246390 | 52.102 | Wellingtonvirus | Unclassified | Myoviridae | Erwinia |
| NC_031007 | Erwinia phage vB_EamM_EarlPhillipIV | Erwinia phage vB_EamM_EarlPhillipIV unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223935 | 50.589 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| NC_031001 | Gordonia phage Terapin | Gordonia phage Terapin Gordonia virus Terapin Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66611 | 59.552 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| NC_031035 | Mycobacterium phage Gengar | Mycobacterium phage Gengar unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61626 | 64.987 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_031086 | Acinetobacter phage phiAB6 | Acinetobacter phage phiAB6 Acinetobacter virus phiAB6 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40570 | 39.468 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_031039 | Bacillus phage AR9 | Bacillus phage AR9 unclassified Takahashivirus Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 251042 | 27.752 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| NC_030944 | Gordonia phage Demosthenes | Gordonia phage Demosthenes Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74073 | 59.255 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030941 | Gordonia phage Cozz | Gordonia phage Cozz Gordonia virus Cozz Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46600 | 60.043 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030936 | Gordonia phage Bachita | Gordonia phage Bachita Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93843 | 61.948 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| NC_030935 | Gordonia phage GRU3 | Gordonia phage GRU3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17727 | 66.509 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_030924 | Gordonia phage Monty | Gordonia phage Monty Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75680 | 58.948 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030923 | Pseudomonas phage YMC11/06/C171_PPU_BP | Pseudomonas phage YMC11/06/C171_PPU_BP Pseudomonas virus C171 Kantovirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45417 | 60.354 | Kantovirus | Corkvirinae | Autographiviridae | Pseudomonas |
| NC_030914 | Gordonia phage Kvothe | Gordonia phage Kvothe Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75462 | 59.48 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030906 | Gordonia phage GMA6 | Gordonia phage GMA6 Gordonia virus GMA6 Bendigovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83324 | 58.179 | Bendigovirus | Unclassified | Myoviridae | Gordonia |
| NC_030905 | Gordonia phage Woes | Gordonia phage Woes Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73752 | 59.125 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030901 | Gordonia phage ClubL | Gordonia phage ClubL Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92618 | 61.92 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| NC_030948 | Ralstonia phage RSP15 | Ralstonia phage RSP15 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167619 | 44.702 | Unclassified | Unclassified | Ackermannviridae | Ralstonia |
| NC_030932 | Gordonia phage Gsput1 | Gordonia phage Gsput1 Gordonia virus Gsput1 Gesputvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43505 | 62.832 | Gesputvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030927 | Gordonia phage Splinter | Gordonia phage Splinter Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45858 | 66.13 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_030915 | Gordonia phage Orchid | Gordonia phage Orchid Gordonia virus Orchid Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80650 | 47.007 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| NC_030911 | Gordonia phage Vendetta | Gordonia phage Vendetta Vendettavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45858 | 66.135 | Vendettavirus | Unclassified | Siphoviridae | Gordonia |
| NC_030907 | Gordonia phage GMA5 | Gordonia phage GMA5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17562 | 66.353 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_030848 | Haloarcula californiae icosahedral virus 1 | Haloarcula californiae icosahedral virus 1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 31314 | 68.126 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| NC_030696 | Gordonia phage Smoothie | Gordonia phage Smoothie Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93139 | 61.932 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| NC_029106 | Achromobacter phage phiAxp-2 | Achromobacter phage phiAxp-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62220 | 60.111 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| NC_029067 | Vibrio phage eugene 12A10 | Vibrio phage eugene 12A10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138234 | 37.954 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_029100 | Pseudoalteromonas phage Pq0 | Pseudoalteromonas phage Pq0 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33399 | 40.286 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| NC_029097 | Rhodobacter phage RcTitan | Rhodobacter phage RcTitan Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44496 | 55.14 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| NC_029092 | Caulobacter phage Percy | Caulobacter phage Percy Caulobacter virus Percy Percyvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44974 | 60.902 | Percyvirus | Unclassified | Autographiviridae | Caulobacter |
| NC_029081 | Pectobacterium phage Peat1 | Pectobacterium phage Peat1 Pectobacterium virus Peat1 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45633 | 48.862 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_029078 | Erysipelothrix phage SE-1 | Erysipelothrix phage SE-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34997 | 33.951 | Unclassified | Unclassified | Siphoviridae | Erysipelothrix |
| NC_029075 | Achromobacter phage JWF | Achromobacter phage JWF Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81541 | 60.138 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| NC_029074 | Gordonia phage GordTnk2 | Gordonia phage GordTnk2 Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75987 | 50.694 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| NC_029061 | Rhodoferax phage P26218 | Rhodoferax phage P26218 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36315 | 56.696 | Unclassified | Unclassified | Podoviridae | Rhodoferax |
| NC_029060 | Gordonia phage GordDuk1 | Gordonia phage GordDuk1 unclassified Gordtnkvirus Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76276 | 50.653 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| NC_029057 | Vibrio phage qdvp001 | Vibrio phage qdvp001 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134742 | 35.349 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_029047 | Verrucomicrobia phage P8625 | Verrucomicrobia phage P8625 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32894 | 50.967 | Unclassified | Unclassified | Siphoviridae | Verrucomicrobia |
| NC_029044 | Mycobacterium phage Breeniome | Mycobacterium phage Breeniome Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154434 | 64.78 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_029042 | Salmonella phage 38 | Salmonella phage 38 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156833 | 44.616 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_028777 | Bacillus phage Stills | Bacillus phage Stills Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80798 | 35.481 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| NC_029033 | Achromobacter phage phiAxp-1 | Achromobacter phage phiAxp-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45045 | 56.017 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| NC_028856 | Bacillus phage Stahl | Bacillus phage Stahl Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80148 | 35.257 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| NC_028840 | Escherichia phage slur09 | Escherichia phage slur09 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111751 | 39.015 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_029031 | Synechococcus phage S-CBP42 | Synechococcus phage S-CBP42 Synechococcus virus SCBP42 Aegirvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45218 | 54.551 | Aegirvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_029026 | Enterococcus phage EFLK1 | Enterococcus phage EFLK1 Enterococcus virus EFLK1 Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130952 | 35.888 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_029019 | Stenotrophomonas phage vB_SmaS-DLP_2 | Stenotrophomonas phage vB_SmaS-DLP_2 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42593 | 53.715 | Septimatrevirus | Unclassified | Siphoviridae | Stenotrophomonas |
| NC_029016 | Enterococcus phage vB_EfaS_IME198 | Enterococcus phage vB_EfaS_IME198 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58000 | 40.019 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_029011 | Sulfolobus monocaudavirus SMV4 | Sulfolobus monocaudavirus SMV4 unclassified Bicaudaviridae Bicaudaviridae Viruses | 51711 | 38.615 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| NC_029007 | Ralstonia phage RSJ5 | Ralstonia phage RSJ5 Ralstonia virus RSJ5 Risjevirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43745 | 60.661 | Risjevirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_029000 | Stenotrophomonas phage IME13 | Stenotrophomonas phage IME13 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162327 | 41.179 | Tulanevirus | Unclassified | Myoviridae | Stenotrophomonas |
| NC_028999 | Pseudomonas phage PhiPA3 | Pseudomonas phage PhiPA3 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 309208 | 47.725 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_028992 | Sulfolobales Virus YNP2 | Sulfolobales Virus YNP2 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 34783 | 42.653 | Unclassified | Unclassified | Unclassified | Unspecified |
| NC_028990 | Enterococcus phage vB_EfaS_IME196 | Enterococcus phage vB_EfaS_IME196 Enterococcus virus IME196 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38886 | 34.974 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_028988 | Ralstonia phage RSJ2 | Ralstonia phage RSJ2 Ralstonia virus RSJ2 Risjevirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44360 | 60.911 | Risjevirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_028986 | Mycobacterium phage Alice | Mycobacterium phage Alice Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153401 | 64.681 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_028983 | Bacillus phage Shanette | Bacillus phage Shanette Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138877 | 40.77 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| NC_028982 | Bacillus phage JL | Bacillus phage JL Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137918 | 40.802 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| NC_028979 | Mycobacterium phage Cheetobro | Mycobacterium phage Cheetobro unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57253 | 67.967 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028974 | Streptomyces phage YDN12 | Streptomyces phage YDN12 Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56528 | 69.21 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_028972 | Gordonia phage Gmala1 | Gordonia phage Gmala1 unclassified Gordtnkvirus Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75167 | 50.798 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| NC_028968 | Mycobacterium phage BrownCNA | Mycobacterium phage BrownCNA Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71214 | 68.878 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028966 | Tsukamurella phage TIN3 | Tsukamurella phage TIN3 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76269 | 59.291 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| NC_028954 | Rhodobacter phage RcRhea | Rhodobacter phage RcRhea Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36065 | 65.446 | Cronusvirus | Unclassified | Siphoviridae | Rhodobacter |
| NC_028950 | Ralstonia phage RSL2 | Ralstonia phage RSL2 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223932 | 52.057 | Chiangmaivirus | Unclassified | Myoviridae | Ralstonia |
| NC_028942 | Mycobacterium phage Phipps | Mycobacterium phage Phipps Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68293 | 66.461 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028938 | Acidianus bottle-shaped virus 2 strain ABV2 | Acidianus bottle-shaped virus 2 strain ABV2 Bottigliavirus ABV2 Bottigliavirus Ampullaviridae Viruses | 22613 | 33.33 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| NC_028934 | Mycobacterium phage Vincenzo | Mycobacterium phage Vincenzo Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72139 | 68.909 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028931 | Pseudomonas phage PaMx28 | Pseudomonas phage PaMx28 Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55108 | 66.406 | Pamexvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_028926 | Bacillus phage Palmer | Bacillus phage Palmer Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40000 | 40.668 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_028924 | Polaribacter phage P12002L | Polaribacter phage P12002L Incheonvrus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48689 | 28.935 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| NC_028907 | Mycobacterium phage Kikipoo | Mycobacterium phage Kikipoo Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68839 | 66.52 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028899 | Ralstonia phage RSF1 | Ralstonia phage RSF1 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222888 | 52.259 | Chiangmaivirus | Unclassified | Myoviridae | Ralstonia |
| NC_028895 | Vibrio phage phi 3 | Vibrio phage phi 3 Vibrio virus phi3 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116138 | 42.827 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_028879 | Pseudomonas phage PaMx42 | Pseudomonas phage PaMx42 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43225 | 54.635 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_028869 | Mycobacterium phage HyRo | Mycobacterium phage HyRo Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153714 | 64.686 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_028865 | Tsukamurella phage TIN2 | Tsukamurella phage TIN2 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76964 | 58.92 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| NC_028848 | Acinetobacter phage Fri1 | Acinetobacter phage Fri1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41805 | 39.287 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_028846 | Mycobacterium phage ShedlockHolmes | Mycobacterium phage ShedlockHolmes unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61081 | 67.301 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_028826 | Enterococcus phage IME-EFm5 | Enterococcus phage IME-EFm5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42265 | 35.509 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_028819 | Pseudoalteromonas phage H103 | Pseudoalteromonas phage H103 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43190 | 41.174 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| NC_028818 | Streptomyces phage TP1604 | Streptomyces phage TP1604 Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57168 | 69.208 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_028812 | Proteus phage pPM_01 | Proteus phage pPM_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58546 | 46.862 | Unclassified | Unclassified | Siphoviridae | Proteus |
| NC_028809 | Pseudomonas phage PaMx74 | Pseudomonas phage PaMx74 Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58637 | 68.382 | Pamexvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_028801 | Mycobacterium phage Vortex | Mycobacterium phage Vortex unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68346 | 66.45 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028787 | Acidianus bottle-shaped virus 3 strain ABV3 | Acidianus bottle-shaped virus 3 strain ABV3 Bottigliavirus ABV3 Bottigliavirus Ampullaviridae Viruses | 28489 | 31.654 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| NC_028782 | Bacillus phage Pavlov | Bacillus phage Pavlov unclassified Pagevirus Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40024 | 40.621 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_028773 | Cronobacter phage S13 | Cronobacter phage S13 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182145 | 40.199 | Unclassified | Tevenvirinae | Myoviridae | Cronobacter |
| NC_028770 | Pseudomonas phage PaMx11 | Pseudomonas phage PaMx11 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59878 | 64.453 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_028754 | Salmonella phage Shivani | Salmonella phage Shivani Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120098 | 38.839 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_028750 | Klebsiella phage Matisse | Klebsiella phage Matisse Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176081 | 41.774 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_028742 | Mycobacterium phage Baee | Mycobacterium phage Baee Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70270 | 67.645 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028672 | Cronobacter phage PBES 02 | Cronobacter phage PBES 02 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149732 | 50.725 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| NC_028702 | Delftia phage IME-DE1 | Delftia phage IME-DE1 Delftia virus IMEDE1 Piedvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38084 | 60.427 | Piedvirus | Unclassified | Autographiviridae | Delftia |
| NC_028692 | Arthrobacter phage Decurro | Arthrobacter phage Decurro Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15524 | 60.236 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_028691 | Mycobacterium phage Apizium | Mycobacterium phage Apizium Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68227 | 66.44 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028690 | Mycobacterium phage Eremos | Mycobacterium phage Eremos Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68472 | 66.531 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028689 | Mycobacterium phage Colbert | Mycobacterium phage Colbert Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67774 | 66.495 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028684 | Acinetobacter phage vB_AbaP_PD-6A3 | Acinetobacter phage vB_AbaP_PD-6A3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41563 | 39.477 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_028682 | Mycobacterium phage Badfish | Mycobacterium phage Badfish Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69030 | 66.522 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028679 | Acinetobacter phage vB_AbaP_PD-AB9 | Acinetobacter phage vB_AbaP_PD-AB9 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40938 | 39.338 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_028677 | Rhodococcus phage CosmicSans | Rhodococcus phage CosmicSans unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.548 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_028675 | Acinetobacter phage phiAB1 | Acinetobacter phage phiAB1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41526 | 39.086 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_028674 | Mycobacterium phage ShiVal | Mycobacterium phage ShiVal Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68355 | 66.498 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028673 | Gordonia phage GMA7 | Gordonia phage GMA7 Gordonia virus GMA7 Getseptimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73419 | 56.611 | Getseptimavirus | Unclassified | Siphoviridae | Gordonia |
| NC_028668 | Gordonia phage GMA3 | Gordonia phage GMA3 Gordonia virus GMA3 Gamtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77779 | 51.263 | Gamtrevirus | Unclassified | Siphoviridae | Gordonia |
| NC_028665 | Gordonia phage GTE6 | Gordonia phage GTE6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56982 | 67.797 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_028658 | Mycobacterium phage Swish | Mycobacterium phage Swish Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68735 | 66.48 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_028653 | Gordonia phage GTE8 | Gordonia phage GTE8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67617 | 65.957 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_028651 | Hydrogenobaculum phage 1 | Hydrogenobaculum phage 1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 19351 | 38.184 | Unclassified | Unclassified | Unclassified | Hydrogenobaculum |
| NC_026610 | Vibrio phage VpKK5 | Vibrio phage VpKK5 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56637 | 51.318 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| NC_011085 | Morganella phage MmP1 | Morganella phage MmP1 Morganella virus MmP1 Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38457 | 46.519 | Minipunavirus | Studiervirinae | Autographiviridae | Morganella |
| NC_008202 | Mycobacterium phage Halo | Mycobacterium phage Halo Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42289 | 66.722 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_003390 | Synechococcus phage P60 | Synechococcus phage P60 Tiilvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46675 | 53.285 | Tiilvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_027987 | Leuconostoc phage 1-A4 | Leuconostoc phage 1-A4 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29508 | 36.146 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_027985 | Mycobacterium phage UncleHowie | Mycobacterium phage UncleHowie Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68016 | 66.508 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_027395 | Escherichia phage vB_EcoP_SU10 | Escherichia phage vB_EcoP_SU10 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77327 | 42.063 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| NC_027365 | Mycobacterium virus Fionnbarth | Mycobacterium virus Fionnbarth Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58076 | 68.007 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_027363 | Proteus phage PM 75 | Proteus phage PM 75 Proteus virus PM75 Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41480 | 41.56 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| NC_027358 | Leuconostoc phage Ln-9 | Leuconostoc phage Ln-9 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28527 | 36.439 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_027356 | Escherichia virus DT57C | Escherichia virus DT57C Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108065 | 39.65 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_027342 | Proteus phage PM16 | Proteus phage PM16 Proteus virus PM16 Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41268 | 41.386 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| NC_027332 | Acinetobacter phage YMC13/03/R2096 | Acinetobacter phage YMC13/03/R2096 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98170 | 37.036 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| NC_027335 | Enterococcus phage ECP3 | Enterococcus phage ECP3 Enterococcus virus ECP3 Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145518 | 35.908 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_027330 | Escherichia phage ECBP5 | Escherichia phage ECBP5 Escherichia virus ECBP5 Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45403 | 45.891 | Gajwadongvirus | Unclassified | Autographiviridae | Escherichia |
| NC_027394 | Bacillus phage Pookie | Bacillus phage Pookie Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40214 | 40.573 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_027372 | Bacillus phage Pascal | Bacillus phage Pascal Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39639 | 39.87 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_027296 | Rhizobium phage RHEph06 | Rhizobium phage RHEph06 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53721 | 56.429 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| NC_027297 | Salmonella virus Stitch | Salmonella virus Stitch Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123475 | 40.307 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_027292 | Pseudomonas phage Pf-10 | Pseudomonas phage Pf-10 Pseudomonas virus Pf10 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39167 | 56.474 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_027119 | Salmonella phage Det7 | Salmonella phage Det7 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157498 | 44.598 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_026609 | Lactobacillus phage Ldl1 | Lactobacillus phage Ldl1 Lactobacillus virus Ldl1 Lidleunavirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74806 | 37.763 | Lidleunavirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| NC_026608 | Paracoccus phage vB_PmaS-R3 | Paracoccus phage vB_PmaS-R3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42093 | 56.368 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| NC_026603 | Mycobacterium phage Keshu | Mycobacterium phage Keshu unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61251 | 67.254 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_025822 | Vibrio phage phi-A318 | Vibrio phage phi-A318 Vibrio virus A318 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42544 | 43.47 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| NC_023693 | Enterobacteria phage phi92 | Enterobacteria phage phi92 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148612 | 37.434 | Unclassified | Vequintavirinae | Myoviridae | Enterobacteria |
| NC_025441 | Escherichia phage ECML-117 | Escherichia phage ECML-117 Escherichia virus ECML-117 Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66854 | 46.198 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| NC_025440 | Listeria phage WIL-1 | Listeria phage WIL-1 Listeria virus WIL1 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134369 | 35.988 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_025423 | Bacillus phage CP-51 | Bacillus phage CP-51 Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138658 | 40.86 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| NC_025422 | Caulobacter phage Cr30 | Caulobacter phage Cr30 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155997 | 37.842 | Unclassified | Unclassified | Myoviridae | Caulobacter |
| NC_025470 | Shewanella sp. phage 1/40 | Shewanella sp. phage 1/40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139004 | 36.888 | Unclassified | Unclassified | Myoviridae | Shewanella |
| NC_025452 | Dickeya phage RC-2014 | Dickeya phage RC-2014 Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155346 | 49.649 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| NC_025466 | Shewanella sp. phage 3/49 | Shewanella sp. phage 3/49 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40161 | 41.966 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| NC_025465 | Enterococcus phage EfaCPT1 | Enterococcus phage EfaCPT1 Enterococcus virus EfaCPT1 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40923 | 34.641 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| NC_025463 | Shewanella sp. phage 1/44 | Shewanella sp. phage 1/44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49640 | 39.823 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| NC_025462 | Acinetobacter phage vB_AbaM_Acibel004 | Acinetobacter phage vB_AbaM_Acibel004 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99730 | 37.27 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| NC_025458 | Shewanella sp. phage 1/41 | Shewanella sp. phage 1/41 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43510 | 42.703 | Unclassified | Unclassified | Myoviridae | Shewanella |
| NC_025457 | Acinetobacter phage vB_AbaP_Acibel007 | Acinetobacter phage vB_AbaP_Acibel007 Acinetobacter virus Acibel007 Daemvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42654 | 41.166 | Daemvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_025455 | Synechococcus phage S-CBP2 | Synechococcus phage S-CBP2 Synechococcus virus SCBP2 Kembevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46237 | 55.034 | Kembevirus | Unclassified | Autographiviridae | Synechococcus |
| NC_025439 | Idiomarinaceae phage 1N2-2 | Idiomarinaceae phage 1N2-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34773 | 51.989 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_025436 | Shewanella sp. phage 1/4 | Shewanella sp. phage 1/4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133824 | 36.855 | Unclassified | Unclassified | Myoviridae | Shewanella |
| NC_025429 | Rhizobium phage vB_RleM_P10VF | Rhizobium phage vB_RleM_P10VF unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156446 | 49.877 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| NC_025428 | Ruegeria phage DSS3-P1 | Ruegeria phage DSS3-P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59601 | 64.133 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| NC_025421 | Lactobacillus phage Ld3 | Lactobacillus phage Ld3 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29616 | 41.646 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_025420 | Lactobacillus phage Ld17 | Lactobacillus phage Ld17 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32975 | 41.498 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_025415 | Lactobacillus phage Ld25A | Lactobacillus phage Ld25A Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32799 | 41.663 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_024369 | Vibrio phage X29 | Vibrio phage X29 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41569 | 46.068 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_024783 | Escherichia phage EK99P-1 | Escherichia phage EK99P-1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44332 | 54.475 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_024392 | Listeria phage LP-114 | Listeria phage LP-114 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66676 | 36.434 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| NC_024389 | Leuconostoc phage phiLNTR2 | Leuconostoc phage phiLNTR2 unclassified Unaquatrovirus Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28336 | 36.014 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024388 | Leuconostoc phage LN34 | Leuconostoc phage LN34 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28022 | 35.972 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024383 | Listeria phage LP-083-2 | Listeria phage LP-083-2 Listeria virus LP083-2 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135831 | 35.868 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_024378 | Leuconostoc phage phiLNTR3 | Leuconostoc phage phiLNTR3 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28015 | 36.052 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| NC_024381 | Pseudomonas phage vB_PaeS_SCH_Ab26 | Pseudomonas phage vB_PaeS_SCH_Ab26 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43056 | 53.426 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_024375 | Listeria phage LP-026 | Listeria phage LP-026 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67150 | 36.258 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| NC_024366 | Mycobacterium phage OkiRoe | Mycobacterium phage OkiRoe unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62661 | 64.938 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024363 | Mycobacterium phage Manad | Mycobacterium phage Manad Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68807 | 66.431 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_024359 | Listeria phage LP-048 | Listeria phage LP-048 Listeria virus LP048 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133048 | 35.966 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_024354 | Cronobacter phage CR8 | Cronobacter phage CR8 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149162 | 50.832 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| NC_023863 | Vibrio phage CHOED | Vibrio phage CHOED Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66316 | 43.679 | Unclassified | Unclassified | Podoviridae | Vibrio |
| NC_024217 | Nitrincola phage 1M3-16 | Nitrincola phage 1M3-16 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82438 | 41.306 | Unclassified | Unclassified | Unclassified | Nitrincola |
| NC_024209 | Mycobacterium phage Hawkeye | Mycobacterium phage Hawkeye Hawkeyevirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67383 | 57.05 | Hawkeyevirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| NC_024212 | Enterococcus phage VD13 | Enterococcus phage VD13 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55726 | 40.008 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_024145 | Mycobacterium phage Hosp | Mycobacterium phage Hosp unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70667 | 69.886 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_024139 | Escherichia phage vB_EcoS_FFH_1 | Escherichia phage vB_EcoS_FFH_1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108483 | 39.24 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_024138 | Mycobacterium phage MosMoris | Mycobacterium phage MosMoris Marvinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65243 | 63.352 | Marvinvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024135 | Mycobacterium phage Bernal13 | Mycobacterium phage Bernal13 Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42392 | 66.199 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024122 | Salmonella phage vB-SalM-SJ3 | Salmonella phage vB-SalM-SJ3 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162910 | 44.377 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_021342 | Edwardsiella phage PEi21 | Edwardsiella phage PEi21 Yokohamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43378 | 52.57 | Yokohamavirus | Unclassified | Myoviridae | Edwardsiella |
| NC_023741 | Mycobacterium phage Stinger | Mycobacterium phage Stinger Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69641 | 68.557 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022774 | Bacillus phage Slash | Bacillus phage Slash Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80382 | 35.232 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| NC_023859 | Microbacterium phage vB_MoxS-ISF9 | Microbacterium phage vB_MoxS-ISF9 Microbacterium virus ISF9 Farahnazvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59254 | 62.759 | Farahnazvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_023841 | Dunaliella viridis virus SI2 | Dunaliella viridis virus SI2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37328 | 62.704 | Unclassified | Unclassified | Podoviridae | Dunaliella |
| NC_023740 | Mycobacterium phage JacAttac | Mycobacterium phage JacAttac Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68311 | 66.505 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023737 | Mycobacterium phage Pleione | Mycobacterium phage Pleione Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155586 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_023736 | Ralstonia phage RSB2 | Ralstonia phage RSB2 Ralstonia virus RSB2 Kelmasvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40411 | 61.751 | Kelmasvirus | Unclassified | Autographiviridae | Ralstonia |
| NC_023733 | Mycobacterium phage MoMoMixon | Mycobacterium phage MoMoMixon Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154573 | 64.767 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_023727 | Mycobacterium phage Vista | Mycobacterium phage Vista Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68494 | 66.48 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023725 | Mycobacterium phage Nappy | Mycobacterium phage Nappy Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156646 | 64.652 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_023722 | Rhodococcus phage ReqiPine5 | Rhodococcus phage ReqiPine5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59231 | 67.095 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_023717 | Cronobacter phage CR9 | Cronobacter phage CR9 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151924 | 50.556 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| NC_023714 | Mycobacterium phage LinStu | Mycobacterium phage LinStu Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153882 | 64.837 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_023712 | Mycobacterium virus Firecracker | Mycobacterium virus Firecracker Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71341 | 65.498 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023711 | Mycobacterium phage Oline | Mycobacterium phage Oline Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68720 | 66.388 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023706 | Rhodococcus phage ReqiDocB7 | Rhodococcus phage ReqiDocB7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75772 | 56.658 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_023701 | Mycobacterium phage Acadian | Mycobacterium phage Acadian Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69864 | 68.423 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023696 | Mycobacterium phage Dandelion | Mycobacterium phage Dandelion Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157568 | 64.726 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_023691 | Mycobacterium phage Patience | Mycobacterium phage Patience Patiencevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70506 | 50.304 | Patiencevirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023688 | Aeromonas phage PX29 | Aeromonas phage PX29 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222006 | 42.043 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| NC_023614 | Croceibacter phage P2559Y | Croceibacter phage P2559Y Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43153 | 38.922 | Unclassified | Unclassified | Siphoviridae | Croceibacter |
| NC_023612 | Geobacillus phage GBK2 | Geobacillus phage GBK2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39078 | 43.093 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| NC_023605 | Vibrio phage PVA1 | Vibrio phage PVA1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41529 | 43.697 | Unclassified | Unclassified | Podoviridae | Vibrio |
| NC_023604 | Mycobacterium phage Jolie2 | Mycobacterium phage Jolie2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44306 | 68.065 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_023603 | Mycobacterium phage 39HC | Mycobacterium phage 39HC unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71565 | 70.003 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023600 | Mycobacterium phage Jolie1 | Mycobacterium phage Jolie1 unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71058 | 69.877 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023549 | Clavibacter phage CN1A | Clavibacter phage CN1A Clavibacter virus CN1A Cinunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56789 | 62.088 | Cinunavirus | Unclassified | Siphoviridae | Clavibacter |
| NC_023590 | Acinetobacter phage IMEAB3 | Acinetobacter phage IMEAB3 Acinetobacter virus IMEAB3 Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43050 | 45.475 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| NC_023570 | Acinetobacter phage Petty | Acinetobacter phage Petty Acinetobacter virus Petty Pettyvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40739 | 42.186 | Pettyvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_023569 | Vibrio phage SHOU24 | Vibrio phage SHOU24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77837 | 46.048 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| NC_023566 | Rhizobium phage vB_RglS_P106B | Rhizobium phage vB_RglS_P106B Rhizobium virus P106B Rigallicvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56024 | 47.922 | Rigallicvirus | Unclassified | Siphoviridae | Rhizobium |
| NC_023563 | Mycobacterium phage Suffolk | Mycobacterium phage Suffolk Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68262 | 66.561 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023557 | Erwinia phage Ea35-70 | Erwinia phage Ea35-70 Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271084 | 49.881 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| NC_023554 | Mycobacterium phage JAMaL | Mycobacterium phage JAMaL Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70841 | 68.751 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_023610 | Erwinia phage PhiEaH1 | Erwinia phage PhiEaH1 Erwinia virus EaH1 Iapetusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218339 | 52.341 | Iapetusvirus | Unclassified | Myoviridae | Erwinia |
| NC_021781 | Listeria phage LP-125 | Listeria phage LP-125 Listeria virus LP083-2 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135281 | 35.93 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| NC_021787 | Listeria phage LP-037 | Listeria phage LP-037 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64756 | 36.559 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| NC_022764 | Bacillus phage Page | Bacillus phage Page Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39874 | 40.706 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_022762 | Lactobacillus phage phiLdb | Lactobacillus phage phiLdb Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33996 | 41.952 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| NC_022770 | Bacillus phage Pony | Bacillus phage Pony Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39844 | 40.699 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| NC_022767 | Bacillus phage Staley | Bacillus phage Staley Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81656 | 35.346 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| NC_022766 | Bacillus phage Glittering | Bacillus phage Glittering Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49246 | 42.054 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_022765 | Bacillus phage Riggi | Bacillus phage Riggi Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49836 | 41.46 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_012223 | Escherichia virus SSL2009a | Escherichia virus SSL2009a Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44899 | 54.667 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_023007 | Bacillus phage vB_BanS-Tsamsa | Bacillus phage vB_BanS-Tsamsa Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168876 | 34.323 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_023005 | Pseudomonas phage PPpW-4 | Pseudomonas phage PPpW-4 Pseudomonas virus PPpW4 Phutvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41386 | 56.775 | Phutvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_022987 | Xylella phage Prado | Xylella phage Prado Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43940 | 63.022 | Pradovirus | Unclassified | Autographiviridae | Xylella |
| NC_022986 | Pseudomonas phage PAK_P4 | Pseudomonas phage PAK_P4 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93147 | 49.267 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_022982 | Xylella phage Paz | Xylella phage Paz unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43869 | 60.168 | Unclassified | Okabevirinae | Autographiviridae | Xylella |
| NC_022917 | Ralstonia phage RSB3 | Ralstonia phage RSB3 Ralstonia virus RSB3 Jiaoyazivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44578 | 61.032 | Jiaoyazivirus | Unclassified | Autographiviridae | Ralstonia |
| NC_022916 | Burkholderia phage JG068 | Burkholderia phage JG068 Burkholderia virus JG068 Mguuvirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41604 | 60.689 | Mguuvirus | Okabevirinae | Autographiviridae | Burkholderia |
| NC_022761 | Bacillus phage CampHawk | Bacillus phage CampHawk Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146193 | 40.195 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| NC_022751 | Cyanophage PP | Cyanophage PP Podoviridae Caudovirales Viruses | 42480 | 46.408 | Unclassified | Unclassified | Podoviridae | Unspecified |
| NC_022343 | Klebsiella virus 0507KN21 | Klebsiella virus 0507KN21 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159991 | 46.705 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_022328 | Mycobacterium phage Adawi | Mycobacterium phage Adawi Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70236 | 68.792 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022325 | Mycobacterium phage Dylan | Mycobacterium phage Dylan unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69815 | 65.416 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022064 | Mycobacterium phage Reprobate | Mycobacterium phage Reprobate Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70120 | 68.269 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022063 | Mycobacterium phage Phelemich | Mycobacterium phage Phelemich unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70115 | 68.252 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022061 | Mycobacterium phage KayaCho | Mycobacterium phage KayaCho unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70838 | 70.006 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_022057 | Mycobacterium phage Catdawg | Mycobacterium phage Catdawg unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72108 | 65.394 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_022053 | Mycobacterium phage Papyrus | Mycobacterium phage Papyrus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70657 | 55.977 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021868 | Streptococcus phage SPQS1 | Streptococcus phage SPQS1 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58305 | 39.873 | Saphexavirus | Unclassified | Siphoviridae | Streptococcus |
| NC_021804 | Cellulophaga phage phi39:1 | Cellulophaga phage phi39:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28760 | 31.332 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| NC_021802 | Cellulophaga phage phi10:1 | Cellulophaga phage phi10:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53664 | 31.535 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| NC_021800 | Cellulophaga phage phi46:1 | Cellulophaga phage phi46:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34844 | 38.256 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| NC_021799 | Cellulophaga phage phi19:1 | Cellulophaga phage phi19:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57447 | 31.239 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| NC_021798 | Cellulophaga phage phi17:2 | Cellulophaga phage phi17:2 Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145343 | 32.651 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| NC_021795 | Cellulophaga phage phi17:1 | Cellulophaga phage phi17:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38776 | 36.476 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| NC_021791 | Cellulophaga phage phi12:1 | Cellulophaga phage phi12:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39148 | 36.587 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| NC_021790 | Cellulophaga phage phi18:1 | Cellulophaga phage phi18:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39189 | 36.515 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| NC_021788 | Cellulophaga phage phi4:1 | Cellulophaga phage phi4:1 Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145865 | 32.672 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| NC_021785 | Listeria phage LP-110 | Listeria phage LP-110 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65132 | 36.342 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| NC_021776 | Vibrio phage VPMS1 | Vibrio phage VPMS1 unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42313 | 44.669 | Kafunavirus | Unclassified | Podoviridae | Vibrio |
| NC_021563 | Serratia phage Eta | Serratia phage Eta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42724 | 49.913 | Unclassified | Unclassified | Siphoviridae | Serratia |
| NC_021556 | Mycobacterium phage Leo | Mycobacterium phage Leo unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39981 | 66.862 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021561 | Vibrio phage pYD38-B | Vibrio phage pYD38-B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37324 | 43.077 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| NC_021540 | Vibrio phage JA-1 | Vibrio phage JA-1 unclassified Pacinivirus Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69278 | 34.555 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| NC_021531 | Cronobacter phage CR5 | Cronobacter phage CR5 unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223989 | 50.057 | Derbicusvirus | Unclassified | Myoviridae | Cronobacter |
| NC_021471 | Halovirus HSTV-1 | Halovirus HSTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 32189 | 60.297 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021349 | Mycobacterium phage Astraea | Mycobacterium phage Astraea Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154872 | 64.683 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_021348 | Mycobacterium phage ArcherS7 | Mycobacterium phage ArcherS7 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156558 | 64.692 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_021347 | Rhodococcus phage E3 | Rhodococcus phage E3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142563 | 67.466 | Unclassified | Unclassified | Myoviridae | Rhodococcus |
| NC_021346 | Mycobacterium phage Gizmo | Mycobacterium phage Gizmo Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157482 | 64.642 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_021340 | Halovirus HHTV-2 | Halovirus HHTV-2 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 52643 | 66.592 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021337 | Acinetobacter phage AB3 | Acinetobacter phage AB3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31185 | 39.176 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_021330 | Halovirus HCTV-1 | Halovirus HCTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 103257 | 56.958 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021329 | Halovirus HRTV-4 | Halovirus HRTV-4 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 35722 | 59.482 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021328 | Halovirus HGTV-1 | Halovirus HGTV-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143855 | 50.363 | Halovirus | Unclassified | Myoviridae | Unspecified |
| NC_021327 | Halovirus HCTV-5 | Halovirus HCTV-5 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 102105 | 57.594 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021322 | Halovirus HHTV-1 | Halovirus HHTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 49107 | 56.46 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021319 | Halovirus HCTV-2 | Halovirus HCTV-2 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 54291 | 68.127 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_021316 | Acinetobacter phage Abp1 | Acinetobacter phage Abp1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42185 | 39.151 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_021310 | Mycobacterium phage Newman | Mycobacterium phage Newman Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68598 | 66.518 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_019933 | Xanthomonas virus CP1 | Xanthomonas virus CP1 Klementvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43870 | 53.31 | Klementvirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_021073 | Vibrio phage henriette 12B8 | Vibrio phage henriette 12B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107218 | 40.98 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_021068 | Vibrio phage douglas 12A4 | Vibrio phage douglas 12A4 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57611 | 38.956 | Unclassified | Unclassified | Podoviridae | Vibrio |
| NC_021067 | Vibrio phage helene 12B3 | Vibrio phage helene 12B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135982 | 37.584 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_021063 | Mycobacterium phage FF47 | Mycobacterium phage FF47 Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47724 | 58.637 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_021062 | Pseudomonas phage Phi-S1 | Pseudomonas phage Phi-S1 Pseudomonas virus PhiS1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40192 | 56.213 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_020998 | Haloarcula hispanica virus PH1 | Haloarcula hispanica virus PH1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 28072 | 67.64 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| NC_019419 | Escherichia virus JL1 | Escherichia virus JL1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43457 | 54.774 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_020882 | Sulfolobales Mexican fusellovirus 1 | Sulfolobales Mexican fusellovirus 1 unclassified Fuselloviridae Fuselloviridae Viruses | 14847 | 45.43 | Unclassified | Unclassified | Fuselloviridae | Unspecified |
| NC_020872 | Cyanophage SS120-1 | Cyanophage SS120-1 Prochlorococcus virus SS120-1 Banchanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46997 | 40.492 | Banchanvirus | Unclassified | Autographiviridae | Prochlorococcus |
| NC_020868 | Vibrio phage VBP32 | Vibrio phage VBP32 unclassified Stoningtonvirus Stoningtonvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76718 | 42.483 | Stoningtonvirus | Fuhrmanvirinae | Schitoviridae | Vibrio |
| NC_020863 | Vibrio phage PWH3a-P1 | Vibrio phage PWH3a-P1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129155 | 35.504 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_020860 | Cellulophaga phage phiSM | Cellulophaga phage phiSM Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44557 | 33.927 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| NC_020857 | Cyanophage MED4-117 | Cyanophage MED4-117 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38834 | 37.246 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| NC_020854 | Cyanophage KBS-S-2A | Cyanophage KBS-S-2A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40658 | 49.068 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_020853 | Loktanella phage pCB2051-A | Loktanella phage pCB2051-A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56958 | 54.976 | Unclassified | Unclassified | Siphoviridae | Loktanella |
| NC_020850 | Vibrio phage VBM1 | Vibrio phage VBM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38374 | 42.258 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_020847 | Prochlorococcus phage MED4-184 | Prochlorococcus phage MED4-184 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38327 | 37.157 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| NC_020846 | Vibrio phage pYD21-A | Vibrio phage pYD21-A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46917 | 43.897 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| NC_020843 | Vibrio phage 11895-B1 | Vibrio phage 11895-B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126434 | 35.529 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_020842 | Cellulophaga phage phiST | Cellulophaga phage phiST Cbastvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79114 | 30.201 | Cbastvirus | Unclassified | Siphoviridae | Cellulophaga |
| NC_020840 | Tetraselmis viridis virus S20 | Tetraselmis viridis virus S20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38987 | 61.22 | Unclassified | Unclassified | Siphoviridae | Tetraselmis |
| NC_020488 | Vibrio phage VvAW1 | Vibrio phage VvAW1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38682 | 49.113 | Unclassified | Unclassified | Podoviridae | Vibrio |
| NC_020484 | Pelagibacter phage HTVC008M | Pelagibacter phage HTVC008M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147284 | 33.448 | Unclassified | Unclassified | Myoviridae | Pelagibacter |
| NC_020482 | Pelagibacter phage HTVC011P | Pelagibacter phage HTVC011P Pelagibacter virus HTVC011P Stopavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39921 | 31.958 | Stopavirus | Unclassified | Autographiviridae | Pelagibacter |
| NC_020480 | Bacillus phage Finn | Bacillus phage Finn Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50161 | 41.69 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| NC_016989 | Haloarcula hispanica icosahedral virus 2 | Haloarcula hispanica icosahedral virus 2 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 30578 | 66.515 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| NC_020082 | Edwardsiella phage MSW-3 | Edwardsiella phage MSW-3 Yokohamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42746 | 53.1 | Yokohamavirus | Unclassified | Myoviridae | Edwardsiella |
| NC_020080 | Klebsiella phage KP27 | Klebsiella phage KP27 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174413 | 41.817 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_020079 | Escherichia phage phAPEC8 | Escherichia phage phAPEC8 Escherichia virus phAPEC8 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147737 | 39.148 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| NC_019925 | Dickeya virus Limestone | Dickeya virus Limestone Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152427 | 49.279 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| NC_020205 | Xanthomonas citri phage CP2 | Xanthomonas citri phage CP2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42963 | 67.018 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| NC_020158 | Halovirus HVTV-1 | Halovirus HVTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 102319 | 58.271 | Halovirus | Unclassified | Unclassified | Unspecified |
| NC_019521 | Sphingomonas phage PAU | Sphingomonas phage PAU Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219372 | 31.313 | Unclassified | Unclassified | Myoviridae | Sphingomonas |
| NC_019519 | Agrobacterium phage 7-7-1 | Agrobacterium phage 7-7-1 Agrobacterium virus 7-7-1 Schmittlotzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69391 | 52.377 | Schmittlotzvirus | Unclassified | Myoviridae | Agrobacterium |
| NC_019930 | Tetrasphaera phage TJE1 | Tetrasphaera phage TJE1 Tetrasphaera virus TJE1 Tijeunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49219 | 57.742 | Tijeunavirus | Unclassified | Myoviridae | Tetrasphaera |
| NC_019926 | Erwinia phage phiEa100 | Erwinia phage phiEa100 unclassified Eracentumvirus Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45554 | 49.677 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| NC_019924 | Clostridium phage phi8074-B1 | Clostridium phage phi8074-B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47595 | 35.382 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_019923 | Pseudomonas phage AF | Pseudomonas phage AF Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42689 | 58.439 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| NC_019913 | Pseudomonas phage PaP1 | Pseudomonas phage PaP1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91715 | 49.362 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| NC_019910 | Salmonella phage SKML-39 | Salmonella phage SKML-39 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159624 | 50.154 | Agtrevirus | Aglimvirinae | Ackermannviridae | Salmonella |
| NC_019725 | Escherichia phage ADB-2 | Escherichia phage ADB-2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.555 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| NC_019724 | Escherichia phage HK578 | Escherichia phage HK578 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43741 | 54.484 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| NC_019543 | Aeromonas phage Aes508 | Aeromonas phage Aes508 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160646 | 41.227 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_019542 | Pectobacterium phage PP1 | Pectobacterium phage PP1 Pectobacterium virus PP1 Axomammavirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44400 | 49.736 | Axomammavirus | Molineuxvirinae | Autographiviridae | Pectobacterium |
| NC_019530 | Salmonella phage Sh19 | Salmonella phage Sh19 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157785 | 44.685 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_019529 | Vibrio phage pVp-1 | Vibrio phage pVp-1 Vibrio virus pVp1 Vipunavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111506 | 39.707 | Vipunavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| NC_019528 | Aeromonas phage phiAS7 | Aeromonas phage phiAS7 Aeromonas virus AS7 Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41572 | 56.918 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_019525 | Bdellovibrio phage phi1422 | Bdellovibrio phage phi1422 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45354 | 44.294 | Unclassified | Unclassified | Myoviridae | Bdellovibrio |
| NC_019518 | Vibrio phage vB_VchM-138 | Vibrio phage vB_VchM-138 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44485 | 45.555 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_019510 | Erwinia phage vB_EamP-L1 | Erwinia phage vB_EamP-L1 Erwinia virus L1 Elunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39282 | 51.937 | Elunavirus | Studiervirinae | Autographiviridae | Erwinia |
| NC_019508 | Clostridium phage phiCP34O | Clostridium phage phiCP34O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38309 | 30.528 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_019506 | Clostridium phage phiCP13O | Clostridium phage phiCP13O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38329 | 30.363 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_019496 | Clostridium phage phiCP26F | Clostridium phage phiCP26F Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39188 | 30.262 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_019457 | Vibrio phage CP-T1 | Vibrio phage CP-T1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 45.415 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_019454 | Pantoea phage LIMElight | Pantoea phage LIMElight Limelightvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44546 | 53.991 | Limelightvirus | Unclassified | Autographiviridae | Pantoea |
| NC_019452 | Escherichia phage PhaxI | Escherichia phage PhaxI Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156628 | 44.533 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| NC_019420 | Edwardsiella phage KF-1 | Edwardsiella phage KF-1 Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41549 | 48.276 | Kafunavirus | Unclassified | Podoviridae | Edwardsiella |
| NC_019411 | Caulobacter virus Swift | Caulobacter virus Swift Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219216 | 66.073 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_019410 | Caulobacter virus Karma | Caulobacter virus Karma unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221828 | 66.207 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_019408 | Caulobacter virus Rogue | Caulobacter virus Rogue Phicbkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223720 | 66.105 | Phicbkvirus | Unclassified | Siphoviridae | Caulobacter |
| NC_019407 | Caulobacter virus Magneto | Caulobacter virus Magneto unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218929 | 66.15 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_019406 | Caulobacter phage CcrColossus | Caulobacter phage CcrColossus Caulobacter virus CcrColossus Colossusvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279967 | 62.173 | Colossusvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_019405 | Caulobacter virus phiCbK | Caulobacter virus phiCbK Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215710 | 66.153 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| NC_019402 | Cronobacter phage vB_CsaP_GAP52 | Cronobacter phage vB_CsaP_GAP52 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76631 | 44.221 | Unclassified | Unclassified | Podoviridae | Cronobacter |
| NC_018859 | Escherichia phage ECBP2 | Escherichia phage ECBP2 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77315 | 42.42 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| NC_018837 | Pectobacterium phage My1 | Pectobacterium phage My1 Pectobacterium virus My1 Myunavirus Mccorquodalevirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122024 | 40.614 | Myunavirus | Mccorquodalevirinae | Demerecviridae | Pectobacterium |
| NC_018835 | Escherichia phage NJ01 | Escherichia phage NJ01 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77448 | 42.049 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| NC_018831 | Listeria phage P70 | Listeria phage P70 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67170 | 36.457 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| NC_017971 | Pseudomonas phage tf | Pseudomonas phage tf Pseudomonas virus tf Krylovvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46271 | 53.202 | Krylovvirus | Unclassified | Podoviridae | Pseudomonas |
| NC_018283 | Burkholderia phage vB_BceS_AH2 | Burkholderia phage vB_BceS_AH2 Burkholderia virus AH2 Ahduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58065 | 61.314 | Ahduovirus | Unclassified | Siphoviridae | Burkholderia |
| NC_018278 | Burkholderia phage vB_BceS_KL1 | Burkholderia phage vB_BceS_KL1 Kilunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42832 | 54.606 | Kilunavirus | Unclassified | Siphoviridae | Burkholderia |
| NC_018276 | Croceibacter phage P2559S | Croceibacter phage P2559S Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44988 | 41.853 | Unclassified | Unclassified | Siphoviridae | Croceibacter |
| NC_018272 | Nonlabens phage P12024L | Nonlabens phage P12024L Inhavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35652 | 35.875 | Inhavirus | Unclassified | Siphoviridae | Nonlabens |
| NC_018271 | Nonlabens phage P12024S | Nonlabens phage P12024S Inhavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35700 | 35.535 | Inhavirus | Unclassified | Siphoviridae | Nonlabens |
| NC_018270 | Weissella phage phiYS61 | Weissella phage phiYS61 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33594 | 43.919 | Unclassified | Unclassified | Podoviridae | Weissella |
| NC_018269 | Marinomonas phage P12026 | Marinomonas phage P12026 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31766 | 46.603 | Unclassified | Unclassified | Siphoviridae | Marinomonas |
| NC_018088 | Colwellia phage 9A | Colwellia phage 9A Colwellia virus 9A Franklinbayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104936 | 36.611 | Franklinbayvirus | Unclassified | Siphoviridae | Colwellia |
| NC_018086 | Enterococcus phage BC611 | Enterococcus phage BC611 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53996 | 40.449 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| NC_017975 | Halocynthia phage JM-2012 | Halocynthia phage JM-2012 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167292 | 35.395 | Unclassified | Unclassified | Myoviridae | Unspecified |
| NC_017974 | Cronobacter phage CR3 | Cronobacter phage CR3 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149273 | 50.948 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| NC_017972 | Pseudomonas phage Lu11 | Pseudomonas phage Lu11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 280538 | 50.883 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| NC_017983 | Salisaeta icosahedral phage 1 | Salisaeta icosahedral phage 1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 43788 | 57.226 | Unclassified | Unclassified | Unclassified | Salisaeta |
| NC_017969 | Escherichia phage vB_EcoS_AKFV33 | Escherichia phage vB_EcoS_AKFV33 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108853 | 38.945 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_017864 | Pseudomonas phage vB_Pae-Kakheti25 | Pseudomonas phage vB_Pae-Kakheti25 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42844 | 53.802 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_016766 | Synechococcus phage S-CBS4 | Synechococcus phage S-CBS4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69420 | 50.828 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_016764 | Pseudomonas phage Bf7 | Pseudomonas phage Bf7 Pseudomonas virus Bf7 Bifseptvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40058 | 58.405 | Bifseptvirus | Unclassified | Autographiviridae | Pseudomonas |
| NC_016655 | Rhodococcus phage REQ1 | Rhodococcus phage REQ1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51342 | 66.324 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_016653 | Rhodococcus phage RER2 | Rhodococcus phage RER2 Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46586 | 58.571 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_016651 | Rhodococcus phage RRH1 | Rhodococcus phage RRH1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14270 | 68.381 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| NC_016569 | Nocardia phage NBR1 | Nocardia phage NBR1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46140 | 67.503 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| NC_016567 | Vibrio phage SIO-2 | Vibrio phage SIO-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81184 | 44.978 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| NC_016566 | Shigella phage EP23 | Shigella phage EP23 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44077 | 54.421 | Dhillonvirus | Unclassified | Siphoviridae | Shigella |
| NC_016435 | Gordonia phage GRU1 | Gordonia phage GRU1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65766 | 65.475 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_016434 | Gordonia phage GTE5 | Gordonia phage GTE5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65839 | 65.053 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_016166 | Gordonia phage GTE7 | Gordonia phage GTE7 Gordonia virus GTE7 Getseptimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74431 | 56.783 | Getseptimavirus | Unclassified | Siphoviridae | Gordonia |
| NC_016163 | Yersinia phage phiR1-37 | Yersinia phage phiR1-37 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262391 | 32.474 | Unclassified | Unclassified | Myoviridae | Yersinia |
| NC_016164 | Synechococcus phage S-CBS1 | Synechococcus phage S-CBS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30332 | 58.75 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_016073 | Salmonella phage SFP10 | Salmonella phage SFP10 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157950 | 44.534 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_015938 | Salmonella phage 7-11 | Salmonella phage 7-11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89916 | 44.128 | Unclassified | Unclassified | Podoviridae | Salmonella |
| NC_015937 | Thermus phage TMA | Thermus phage TMA Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151483 | 32.625 | Unclassified | Unclassified | Myoviridae | Thermus |
| NC_015721 | Bdellovibrio phage phi1402 | Bdellovibrio phage phi1402 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23931 | 50.349 | Unclassified | Unclassified | Myoviridae | Bdellovibrio |
| NC_015720 | Gordonia phage GTE2 | Gordonia phage GTE2 Gordonia virus GTE2 Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45530 | 60.343 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| NC_015465 | Synechococcus phage S-CBS3 | Synechococcus phage S-CBS3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33004 | 60.72 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_015463 | Synechococcus phage S-CBS2 | Synechococcus phage S-CBS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72332 | 54.474 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| NC_015293 | Pseudoalteromonas phage H105/1 | Pseudoalteromonas phage H105/1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30651 | 40.821 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| NC_015270 | Enterococcus phage EFRM31 | Enterococcus phage EFRM31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16945 | 34.577 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_015269 | Salmonella virus SPC35 | Salmonella virus SPC35 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118351 | 39.385 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_015264 | Pseudomonas phage phiIBB-PF7A | Pseudomonas phage phiIBB-PF7A Pseudomonas virus IBBPF7A Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40973 | 56.271 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_015210 | Tsukamurella phage TPA2 | Tsukamurella phage TPA2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61440 | 69.574 | Unclassified | Unclassified | Siphoviridae | Tsukamurella |
| NC_015208 | Pseudomonas phage phi15 | Pseudomonas phage phi15 Pseudomonas virus Phi15 Troedvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39562 | 58.164 | Troedvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_015159 | Vibrio phage ICP3 | Vibrio phage ICP3 Vibrio virus ICP3 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39162 | 42.858 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| NC_015157 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125956 | 37.073 | Unclassified | Unclassified | Myoviridae | Vibrio |
| NC_014636 | Aeromonas phage phiAS5 | Aeromonas phage phiAS5 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225268 | 42.997 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| NC_014635 | Aeromonas phage phiAS4 | Aeromonas phage phiAS4 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163875 | 41.297 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_014457 | Clostridium phage phiCTP1 | Clostridium phage phiCTP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59199 | 31.256 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| NC_014036 | Klebsiella phage KP15 | Klebsiella phage KP15 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174436 | 41.831 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| NC_013758 | Haloarcula hispanica pleomorphic virus 1 | Haloarcula hispanica pleomorphic virus 1 Alphapleolipovirus HHPV1 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 8082 | 55.778 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Haloarcula |
| NC_013693 | Shigella phage Ag3 | Shigella phage Ag3 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158006 | 50.398 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| NC_013697 | Delftia phage PhiW-14 | Delftia phage PhiW-14 Delftia virus PhiW14 Ionavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157486 | 56.277 | Ionavirus | Unclassified | Myoviridae | Delftia |
| NC_013698 | Clavibacter phage CMP1 | Clavibacter phage CMP1 Clavibacter virus CMP1 Cimpunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58652 | 57.007 | Cimpunavirus | Unclassified | Siphoviridae | Clavibacter |
| NC_013651 | Vibrio phage N4 | Vibrio phage N4 Vibrio virus N4 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38497 | 42.803 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| NC_010811 | Ralstonia phage phiRSL1 | Ralstonia phage phiRSL1 Mieseafarmvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231255 | 58.026 | Mieseafarmvirus | Unclassified | Myoviridae | Ralstonia |
| NC_013021 | Cyanophage PSS2 | Cyanophage PSS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107530 | 52.301 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| NC_012742 | Xanthomonas phage phiL7 | Xanthomonas phage phiL7 Xanthomonas virus PhiL7 Eisenstarkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44080 | 55.64 | Eisenstarkvirus | Unclassified | Siphoviridae | Xanthomonas |
| NC_012662 | Vibrio phage VP93 | Vibrio phage VP93 Vibrio virus VP93 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43931 | 49.109 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| NC_012558 | Halorubrum pleomorphic virus 1 | Halorubrum pleomorphic virus 1 Alphapleolipovirus HRPV1 Alphapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 7048 | 54.186 | Alphapleolipovirus | Unclassified | Pleolipoviridae | Halorubrum |
| NC_012419 | Enterococcus phage EFAP-1 | Enterococcus phage EFAP-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21115 | 36.718 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| NC_011646 | Abalone shriveling syndrome-associated virus | Abalone shriveling syndrome-associated virus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34952 | 39.52 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| NC_011523 | Bacillus phage AP50 | Bacillus phage AP50 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14398 | 38.651 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| NC_011421 | Bacillus phage SPO1 | Bacillus phage SPO1 Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132562 | 39.965 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| NC_011308 | Listeria phage P40 | Listeria phage P40 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35638 | 39.259 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_011271 | Mycobacterium phage Cali | Mycobacterium phage Cali Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155372 | 64.72 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_011270 | Mycobacterium phage Spud | Mycobacterium phage Spud Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154906 | 64.777 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_011272 | Mycobacterium phage Rizal | Mycobacterium phage Rizal Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153894 | 64.743 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_011222 | Bacteroides phage B40-8 | Bacteroides phage B40-8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44929 | 38.628 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| NC_011201 | Ralstonia phage RSB1 | Ralstonia phage RSB1 Ralstonia virus RSB1 Higashivirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43079 | 61.738 | Higashivirus | Okabevirinae | Autographiviridae | Ralstonia |
| NC_011044 | Mycobacterium phage Nigel | Mycobacterium phage Nigel Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69904 | 68.342 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_010821 | Pseudomonas phage 201phi2-1 | Pseudomonas phage 201phi2-1 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 316674 | 45.344 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| NC_010583 | Escherichia phage Eps7 | Escherichia phage Eps7 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111382 | 39.901 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_010363 | Lactococcus phage asccphi28 | Lactococcus phage asccphi28 Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18762 | 33.658 | Unclassified | Unclassified | Salasmaviridae | Lactococcus |
| NC_010116 | Pseudomonas virus Yua | Pseudomonas virus Yua Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58663 | 64.262 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_001421 | Enterobacteria phage PRD1 | Enterobacteria phage PRD1 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14927 | 48.101 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| NC_009936 | Pseudomonas phage LKA1 | Pseudomonas phage LKA1 Stubburvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41593 | 60.902 | Stubburvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_009904 | Enterococcus phage EF24C | Enterococcus phage EF24C Enterococcus virus EF24C Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142072 | 35.738 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| NC_009814 | Listeria phage P35 | Listeria phage P35 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35822 | 40.818 | Unclassified | Unclassified | Siphoviridae | Listeria |
| NC_009816 | Corynebacterium phage P1201 | Corynebacterium phage P1201 Corynebacterium virus P1201 Chunghsingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70579 | 50.919 | Chunghsingvirus | Unclassified | Siphoviridae | Corynebacterium |
| NC_009760 | Bacillus phage 0305phi8-36 | Bacillus phage 0305phi8-36 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218948 | 41.795 | Unclassified | Unclassified | Myoviridae | Bacillus |
| NC_009597 | Pyrococcus abyssi virus 1 | Pyrococcus abyssi virus 1 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 18098 | 47.16 | Unclassified | Unclassified | Unclassified | Pyrococcus |
| NC_009531 | Synechococcus phage Syn5 | Synechococcus phage Syn5 Voetvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46214 | 54.97 | Voetvirus | Unclassified | Autographiviridae | Synechococcus |
| NC_009447 | Burkholderia phage BcepGomr | Burkholderia phage BcepGomr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52414 | 56.294 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| NC_009014 | Erwinia phage Era103 | Erwinia phage Era103 Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45445 | 49.838 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| NC_008584 | Thermus phage phiYS40 | Thermus phage phiYS40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152372 | 32.586 | Unclassified | Unclassified | Myoviridae | Thermus |
| NC_008204 | Mycobacterium phage Qyrzula | Mycobacterium phage Qyrzula Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67188 | 68.963 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_008207 | Mycobacterium phage Catera | Mycobacterium phage Catera Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153766 | 64.735 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_008195 | Mycobacterium phage Cooper | Mycobacterium phage Cooper Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70654 | 69.106 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_008197 | Mycobacterium phage Orion | Mycobacterium phage Orion Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68427 | 66.51 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_008208 | Aeromonas phage 25 | Aeromonas phage 25 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161475 | 41.04 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| NC_005091 | Burkholderia phage BcepNazgul | Burkholderia phage BcepNazgul Burkholderia virus BcepNazgul Nazgulvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57455 | 60.639 | Nazgulvirus | Unclassified | Siphoviridae | Burkholderia |
| NC_007918 | His2 virus | His2 virus Gammapleolipovirus His2 Gammapleolipovirus Pleolipoviridae Haloruvirales Huolimaviricetes Saleviricota Trapavirae Monodnaviria Viruses | 16067 | 40.368 | Gammapleolipovirus | Unclassified | Pleolipoviridae | Unspecified |
| NC_007914 | His 1 virus | His 1 virus Salterprovirus His1 Salterprovirus Halspiviridae Viruses | 14462 | 38.854 | Salterprovirus | Unclassified | Halspiviridae | Unspecified |
| NC_007809 | Pseudomonas phage M6 | Pseudomonas phage M6 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59446 | 64.464 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_007807 | Pseudomonas phage 119X | Pseudomonas phage 119X Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43365 | 44.893 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| NC_007806 | Pseudomonas phage 73 | Pseudomonas phage 73 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42999 | 53.559 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_007451 | Enterobacteria phage PR4 | Enterobacteria phage PR4 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14954 | 48.322 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| NC_006356 | Flavobacterium phage 11b | Flavobacterium phage 11b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36012 | 30.593 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| NC_005886 | Burkholderia phage BcepB1A | Burkholderia phage BcepB1A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47399 | 54.453 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| NC_007217 | Haloarcula hispanica virus SH1 | Haloarcula hispanica virus SH1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 30889 | 68.384 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| NC_007149 | Vibrio phage VP4 | Vibrio phage VP4 Vibrio virus VP4 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39503 | 42.612 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| NC_007022 | Aeromonas virus 31 | Aeromonas virus 31 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172963 | 43.913 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| NC_006953 | Listonella phage phiHSIC | Listonella phage phiHSIC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37966 | 43.963 | Unclassified | Unclassified | Siphoviridae | Listonella |
| NC_005885 | Actinoplanes phage phiAsp2 | Actinoplanes phage phiAsp2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58638 | 70.388 | Unclassified | Unclassified | Siphoviridae | Actinoplanes |
| NC_005884 | Pseudomonas phage PaP2 | Pseudomonas phage PaP2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43783 | 45.365 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| NC_005882 | Burkholderia phage BcepMu | Burkholderia phage BcepMu Bcepmuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36748 | 62.861 | Bcepmuvirus | Unclassified | Myoviridae | Burkholderia |
| NC_005342 | Burkholderia phage Bcep43 | Burkholderia phage Bcep43 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48024 | 63.435 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| NC_004333 | Burkholderia phage Bcep781 | Burkholderia phage Bcep781 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48247 | 63.33 | Naesvirus | Unclassified | Myoviridae | Burkholderia |
| NC_005872 | Pyrobaculum spherical virus | Pyrobaculum spherical virus Globulovirus Globuloviridae Viruses | 28337 | 48.4 | Globulovirus | Unclassified | Globuloviridae | Pyrobaculum |
| NC_005859 | Escherichia phage T5 | Escherichia phage T5 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121750 | 39.27 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_005284 | Burkholderia phage phi1026b | Burkholderia phage phi1026b Stanholtvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54865 | 60.683 | Stanholtvirus | Unclassified | Siphoviridae | Burkholderia |
| NC_005260 | Aeromonas phage Aeh1 | Aeromonas phage Aeh1 Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 233234 | 42.779 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| NC_005259 | Mycobacterium phage PG1 | Mycobacterium phage PG1 Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68999 | 66.502 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_004814 | Streptococcus virus C1 | Streptococcus virus C1 Rosenblumvirus Picovirinae Podoviridae Caudovirales Viruses | 16687 | 34.626 | Rosenblumvirus | Picovirinae | Podoviridae | Streptococcus |
| NC_004735 | Rhodothermus phage RM378 | Rhodothermus phage RM378 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129908 | 42.494 | Unclassified | Unclassified | Myoviridae | Rhodothermus |
| NC_004689 | Mycobacterium phage Barnyard | Mycobacterium phage Barnyard Barnyardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70797 | 57.313 | Barnyardvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004687 | Mycobacterium phage Bxz1 | Mycobacterium phage Bxz1 Bixzunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156102 | 64.77 | Bixzunavirus | Unclassified | Myoviridae | Mycobacterium |
| NC_004685 | Mycobacterium phage Corndog | Mycobacterium phage Corndog Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69777 | 65.384 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_004684 | Mycobacterium phage Rosebush | Mycobacterium phage Rosebush Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67480 | 68.957 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_003387 | Mycobacterium phage TM4 | Mycobacterium phage TM4 Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52797 | 68.114 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_001447 | Acholeplasma virus L2 | Acholeplasma virus L2 Plasmavirus Plasmaviridae Viruses | 11965 | 31.96 | Plasmavirus | Unclassified | Plasmaviridae | Acholeplasma |
| NC_049972 | Bacillus phage vB_Bpu_PumA2 | Bacillus phage vB_Bpu_PumA2 Bacillus virus PumA2 Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18932 | 35.802 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| NC_049971 | Bacillus phage vB_Bpu_PumA1 | Bacillus phage vB_Bpu_PumA1 Bacillus virus PumA1 Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18466 | 34.285 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| NC_049970 | Bacillus phage Karezi | Bacillus phage Karezi Bacillus virus Karezi Karezivirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20083 | 37.305 | Karezivirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| NC_049950 | Escherichia phage 2H10 | Escherichia phage 2H10 Escherichia virus 2H10 Glaedevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46288 | 50.326 | Glaedevirus | Unclassified | Siphoviridae | Escherichia |
| NC_049940 | Enterococcus phage vB_EfaP_Ef7.4 | Enterococcus phage vB_EfaP_Ef7.4 Enterococcus virus Ef74 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18415 | 33.098 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049939 | Enterococcus phage vB_EfaP_Ef7.3 | Enterococcus phage vB_EfaP_Ef7.3 Enterococcus virus Ef73 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18818 | 32.952 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049938 | Enterococcus phage vB_EfaP_Efmus1 | Enterococcus phage vB_EfaP_Efmus1 Enterococcus virus Efmus1 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17927 | 33.542 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049936 | Enterococcus phage vB_EfaP_Efmus3 | Enterococcus phage vB_EfaP_Efmus3 Enterococcus virus Efmus3 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18286 | 33.337 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049935 | Enterococcus phage vB_EfaP_Efmus4 | Enterococcus phage vB_EfaP_Efmus4 Enterococcus virus Efmus4 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18186 | 33.427 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049934 | Enterococcus phage vB_EfaP_Ef7.2 | Enterococcus phage vB_EfaP_Ef7.2 Enterococcus virus Ef72 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18737 | 33.159 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049933 | Enterococcus phage vB_EfaP_Ef6.3 | Enterococcus phage vB_EfaP_Ef6.3 Enterococcus virus Ef63 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18136 | 32.637 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049932 | Enterococcus phage vB_EfaP_Ef6.2 | Enterococcus phage vB_EfaP_Ef6.2 Enterococcus virus Ef62 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17966 | 32.823 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049928 | Microbacterium phage MonChoix | Microbacterium phage MonChoix Microbacterium virus MonChoix Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41670 | 63.365 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_049927 | Microbacterium phage McGalleon | Microbacterium phage McGalleon Microbacterium virus McGalleon Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42562 | 63.658 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_049857 | Lactococcus phage P1048 | Lactococcus phage P1048 Lactococcus virus P1048 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123023 | 32.967 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049851 | Burkholderia phage BcepSauron | Burkholderia phage BcepSauron Burkholderia virus BcepSauron Sarumanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262653 | 58.097 | Sarumanvirus | Unclassified | Myoviridae | Burkholderia |
| NC_049850 | Burkholderia phage BcepSaruman | Burkholderia phage BcepSaruman Burkholderia virus BcepSaruman Sarumanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263735 | 58.139 | Sarumanvirus | Unclassified | Myoviridae | Burkholderia |
| NC_049849 | Achromobacter phage Motura | Achromobacter phage Motura Achromobacter virus Motura Moturavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221431 | 53.709 | Moturavirus | Unclassified | Myoviridae | Achromobacter |
| NC_049808 | Lactococcus phage P596 | Lactococcus phage P596 Lactococcus virus P596 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57909 | 34.364 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049512 | Salmonella phage SE14 | Salmonella phage SE14 Salmonella virus SE14 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152926 | 44.839 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049509 | Salmonella phage P46FS4 | Salmonella phage P46FS4 Salmonella virus P46FS4 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158706 | 50.444 | Agtrevirus | Aglimvirinae | Ackermannviridae | Salmonella |
| NC_049508 | Salmonella phage maane | Salmonella phage maane Salmonella virus maane Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159128 | 45.159 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049507 | Salmonella phage pertopsoe | Salmonella phage pertopsoe Salmonella virus pertopsoe Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158905 | 45.279 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049506 | Salmonella phage moki | Salmonella phage moki Salmonella virus moki Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158230 | 44.42 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049505 | Salmonella phage barely | Salmonella phage barely Salmonella virus barely Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157920 | 44.598 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049504 | Salmonella phage dinky | Salmonella phage dinky Salmonella virus dinky Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157254 | 44.768 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049503 | Salmonella phage aagejoakim | Salmonella phage aagejoakim Salmonella virus aagejoakim Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156801 | 44.491 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049502 | Salmonella phage bering | Salmonella phage bering Salmonella virus bering Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156130 | 44.703 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049501 | Salmonella phage allotria | Salmonella phage allotria Salmonella virus allotria Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151015 | 45.12 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049500 | Salmonella phage heyday | Salmonella phage heyday Salmonella virus heyday Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144232 | 44.47 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049499 | Salmonella phage rabagast | Salmonella phage rabagast Salmonella virus rabagast Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143249 | 44.529 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049498 | Arthrobacter phage Abba | Arthrobacter phage Abba Arthrobacter virus Abba Ayohtrevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46623 | 64.957 | Ayohtrevirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049489 | Arthrobacter phage DrYang | Arthrobacter phage DrYang Arthrobacter virus DrYang Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69061 | 62.173 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049473 | Arthrobacter phage Kuleana | Arthrobacter phage Kuleana Arthrobacter virus Kuleana Kuleanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38850 | 65.833 | Kuleanavirus | Unclassified | Siphoviridae | Arthrobacter |
| NC_049471 | Arthrobacter phage Lymara | Arthrobacter phage Lymara Arthrobacter virus Lymara Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68762 | 61.492 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049468 | Arthrobacter phage Zartrosa | Arthrobacter phage Zartrosa Arthrobacter virus Zartrosa Marthavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50782 | 60.953 | Marthavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049467 | Klebsiella phage KpCHEMY26 | Klebsiella phage KpCHEMY26 Klebsiella virus KpCHEMY26 Pylasvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70678 | 41.716 | Pylasvirus | Humphriesvirinae | Schitoviridae | Klebsiella |
| NC_049465 | Arthrobacter phage Vibaki | Arthrobacter phage Vibaki Arthrobacter virus Vibaki Vibakivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48201 | 66.7 | Vibakivirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049463 | Stenotrophomonas phage Pokken | Stenotrophomonas phage Pokken Stenotrophomonas virus Pokken Pokkenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76239 | 55.111 | Pokkenvirus | Unclassified | Schitoviridae | Stenotrophomonas |
| NC_049462 | Klebsiella phage Magnus | Klebsiella phage Magnus Klebsiella virus Magnus Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157741 | 46.261 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| NC_049458 | Salmonella phage SS9 | Salmonella phage SS9 Salmonella virus SS9 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156329 | 44.662 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049455 | Shigella phage MK-13 | Shigella phage MK-13 Shigella virus MK13 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158755 | 50.417 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| NC_048878 | Alteromonas phage vB_AcoS-R7M | Alteromonas phage vB_AcoS-R7M Alteromonas virus R7M Amoyvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56163 | 45.61 | Amoyvirus | Queuovirinae | Siphoviridae | Alteromonas |
| NC_048877 | Microbacterium phage Phedro | Microbacterium phage Phedro Microbacterium virus Phedro Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54442 | 59.713 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| NC_048875 | Pantoea phage vB_PagM_SSEM1 | Pantoea phage vB_PagM_SSEM1 Pantoea virus SSEM1 Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54982 | 44.182 | Loessnervirus | Cleopatravirinae | Chaseviridae | Pantoea |
| NC_048874 | Escherichia phage EC6098 | Escherichia phage EC6098 Enterogokushovirus EC6098 Enterogokushovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4526 | 48.387 | Enterogokushovirus | Gokushovirinae | Microviridae | Escherichia |
| NC_048872 | Salmonella phage atrejo | Salmonella phage atrejo Salmonella virus atrejo Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114059 | 39.971 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048871 | Salmonella phage oselot | Salmonella phage oselot Salmonella virus oselot Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113503 | 39.94 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048870 | Salmonella phage bastian | Salmonella phage bastian Salmonella virus bastian Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112477 | 39.961 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048869 | Salmonella phage fuchur | Salmonella phage fuchur Salmonella virus fuchur Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112117 | 40.017 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048868 | Salmonella phage rokbiter | Salmonella phage rokbiter Salmonella virus rokbiter Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112006 | 39.926 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048867 | Salmonella phage faergetype | Salmonella phage faergetype Salmonella virus faergetype Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110183 | 39.957 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048866 | Salmonella phage bombadil | Salmonella phage bombadil Salmonella virus bombadil Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109539 | 40.005 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048865 | Salmonella phage oldekolle | Salmonella phage oldekolle Salmonella virus oldekolle Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109372 | 39.285 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048862 | Salmonella phage astrithr | Salmonella phage astrithr Salmonella virus astrithr Astrithrvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11569 | 39.727 | Astrithrvirus | Unclassified | Podoviridae | Salmonella |
| NC_048861 | Proteus phage Privateer | Proteus phage Privateer Proteus virus Privateer Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90710 | 34.523 | Privateervirus | Unclassified | Podoviridae | Proteus |
| NC_048860 | Microbacterium phage Arete | Microbacterium phage Arete Microbacterium virus Arete Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53692 | 64.609 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| NC_048859 | Bifidobacterium phage BadAargau2 | Bifidobacterium phage BadAargau2 Bifidobacterium virus BadAargau2 Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18899 | 50.426 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| NC_048858 | Bifidobacterium phage BadAztec1 | Bifidobacterium phage BadAztec1 Bifidobacterium virus BadAztec1 Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18635 | 48.801 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| NC_048857 | Phage NBSal005 | Phage NBSal005 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121040 | 39.123 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| NC_048856 | Phage NBSal003 | Phage NBSal003 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 119416 | 39.561 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| NC_048855 | Phage NBEco002 | Phage NBEco002 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118118 | 39.077 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| NC_048854 | Streptomyces phage JXY1 | Streptomyces phage JXY1 Streptomyces virus JXY1 Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45385 | 60.253 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| NC_048853 | Phage NBEco001 | Phage NBEco001 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116606 | 39.489 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| NC_048852 | Salmonella phage L6jm | Salmonella phage L6jm Salmonella virus L6jm Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118452 | 39.249 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048851 | Gordonia phage John316 | Gordonia phage John316 Gordonia virus John316 Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77221 | 58.977 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048847 | Alteromonas phage vB_AmeM_PT11-V22 | Alteromonas phage vB_AmeM_PT11-V22 Alteromonas virus PT11-V22 Myoalterovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92760 | 38.387 | Myoalterovirus | Unclassified | Myoviridae | Alteromonas |
| NC_048845 | Escherichia phage flopper | Escherichia phage flopper Escherichia virus flopper Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52092 | 44.203 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| NC_048844 | Flavobacterium phage vB_FspS_tooticki6-1 | Flavobacterium phage vB_FspS_tooticki6-1 Flavobacterium virus Tooticki Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36912 | 29.283 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048843 | Flavobacterium phage vB_FspS_tant8-1 | Flavobacterium phage vB_FspS_tant8-1 Flavobacterium virus Tant Tantvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40542 | 34.406 | Tantvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048842 | Flavobacterium phage vB_FspS_stinky9-1 | Flavobacterium phage vB_FspS_stinky9-1 Flavobacterium virus Stinky Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38502 | 29.375 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048838 | Flavobacterium phage vB_FspS_mymlan6-1 | Flavobacterium phage vB_FspS_mymlan6-1 Flavobacterium virus Mymlan Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38501 | 29.158 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048837 | Flavobacterium phage vB_FspS_mumin9-1 | Flavobacterium phage vB_FspS_mumin9-1 Flavobacterium virus Mumin Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38527 | 29.159 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048833 | Flavobacterium phage vB_FspS_hemulen6-1 | Flavobacterium phage vB_FspS_hemulen6-1 Flavobacterium virus Hemulen Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39039 | 29.537 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048831 | Flavobacterium phage vB_FspS_filifjonk9-1 | Flavobacterium phage vB_FspS_filifjonk9-1 Flavobacterium virus Filifjonk Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38028 | 29.786 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| NC_048830 | Flavobacterium phage vB_FspM_pippi8-1 | Flavobacterium phage vB_FspM_pippi8-1 Flavobacterium virus Pippi Pippivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34711 | 37.582 | Pippivirus | Unclassified | Myoviridae | Flavobacterium |
| NC_048829 | Flavobacterium phage vB_FspM_lotta8-1 | Flavobacterium phage vB_FspM_lotta8-1 Flavobacterium virus Lotta Pippivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34751 | 37.55 | Pippivirus | Unclassified | Myoviridae | Flavobacterium |
| NC_048826 | Gordonia phage Yikes | Gordonia phage Yikes Gordonia virus Yikes Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68155 | 58.222 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048824 | Mycobacterium phage Indlulamithi | Mycobacterium phage Indlulamithi Mycobacterium virus Indlulamithi Indlulamithivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71538 | 52.689 | Indlulamithivirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048821 | Photobacterium phage PDCC-1 | Photobacterium phage PDCC-1 Photobacterium virus PDCC1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237509 | 43.252 | Aphroditevirus | Gorgonvirinae | Myoviridae | Photobacterium |
| NC_048820 | Gordonia phage Jellybones | Gordonia phage Jellybones Gordonia virus Jellybones Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77514 | 58.985 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048819 | Salmonella phage 1-19 | Salmonella phage 1-19 Salmonella virus 1-19 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110775 | 39.998 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048818 | Escherichia virus VEc33 | Escherichia virus VEc33 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108640 | 39.109 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048815 | Gordonia phage Sadboi | Gordonia phage Sadboi Gordonia virus Sadboi Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67485 | 58.078 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048814 | Gordonia phage Lambo | Gordonia phage Lambo Gordonia virus Lambo Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67813 | 58.296 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048813 | Gordonia phage Ranch | Gordonia phage Ranch Gordonia virus Ranch Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66892 | 58.346 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048812 | Gordonia phage Gibbin | Gordonia phage Gibbin Gordonia virus Gibbin Lambovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67785 | 58.267 | Lambovirus | Unclassified | Siphoviridae | Gordonia |
| NC_048811 | Salmonella phage 1-29 | Salmonella phage 1-29 Salmonella virus 1-29 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111636 | 40.075 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048809 | Salmonella phage 2-3 | Salmonella phage 2-3 Salmonella virus 2-3 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114986 | 39.647 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048808 | Salmonella phage OSY-STA | Salmonella phage OSY-STA Salmonella virus OSYSTA Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111039 | 40.048 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048804 | Stenotrophomonas phage Mendera | Stenotrophomonas phage Mendera Stenotrophomonas virus Mendera Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159961 | 54.015 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| NC_048803 | Proteus phage Myduc | Proteus phage Myduc Proteus virus Myduc Myducvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53392 | 39.274 | Myducvirus | Cleopatravirinae | Chaseviridae | Proteus |
| NC_048802 | Stenotrophomonas phage Moby | Stenotrophomonas phage Moby Stenotrophomonas virus Moby Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159365 | 54.137 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| NC_048801 | Microbacterium phage Araxxi | Microbacterium phage Araxxi Microbacterium virus Araxxi Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53981 | 64.836 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| NC_048796 | Panteoa phage Kyle | Panteoa phage Kyle Panteoa virus Kyle Kylevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73168 | 49.498 | Kylevirus | Unclassified | Myoviridae | Pantoea |
| NC_048795 | Salmonella phage Th1 | Salmonella phage Th1 Salmonella virus Th1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110466 | 39.208 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048794 | Salmonella virus VSe12 | Salmonella virus VSe12 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109654 | 39.087 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048792 | Serratia phage Moabite | Serratia phage Moabite Serratia virus Moabite Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273933 | 46.783 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| NC_048786 | Salmonella phage SE11 | Salmonella phage SE11 Salmonella virus SE11 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107763 | 39.195 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048783 | Gordonia phage Beaver | Gordonia phage Beaver Gordonia virus Beaver Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77394 | 58.884 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048781 | Salmonella phage vB_SenS_SB13 | Salmonella phage vB_SenS_SB13 Salmonella virus SB13 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112513 | 39.921 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048779 | Vibrio phage USC-1 | Vibrio phage USC-1 Vibrio virus USC1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 238099 | 43.243 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| NC_048778 | Microbacterium phage Alex44 | Microbacterium phage Alex44 Microbacterium virus Alex44 Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54396 | 59.949 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| NC_048777 | Microbacterium phage TinyTimothy | Microbacterium phage TinyTimothy Microbacterium virus TinyTimothy Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53932 | 56.751 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| NC_048776 | Aeromonas phage LAh_9 | Aeromonas phage LAh_9 Aeromonas virus LAh9 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97988 | 42.438 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| NC_048775 | Aeromonas phage LAh_8 | Aeromonas phage LAh_8 Aeromonas virus LAh8 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97408 | 42.233 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| NC_048774 | Aeromonas phage LAh_6 | Aeromonas phage LAh_6 Aeromonas virus LAh6 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101390 | 42.311 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| NC_048773 | Aeromonas phage 4_4572 | Aeromonas phage 4_4572 Aeromonas virus 4-4572 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102915 | 42.476 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| NC_048772 | Aeromonas phage 4L372XY | Aeromonas phage 4L372XY Aeromonas virus 4L372XY Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124279 | 35.644 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| NC_048771 | Aeromonas phage 4L372D | Aeromonas phage 4L372D Aeromonas virus 4L372D Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138690 | 35.062 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| NC_048770 | Aeromonas phage 2L372X | Aeromonas phage 2L372X Aeromonas virus 2L372X Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115893 | 35.558 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| NC_048769 | Aeromonas phage 2L372D | Aeromonas phage 2L372D Aeromonas virus 2L372D Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122963 | 35.294 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| NC_048768 | Gordonia phage Chelms | Gordonia phage Chelms Gordonia virus Chelms Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76261 | 58.583 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048767 | Pantoea phage vB_PagM_PSKM | Pantoea phage vB_PagM_PSKM Pantoea virus PSKM Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49935 | 52.356 | Dibbivirus | Unclassified | Myoviridae | Pantoea |
| NC_048766 | Pantoea phage vB_PagM_AAM37 | Pantoea phage vB_PagM_AAM37 Pantoea virus AAM37 Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49990 | 52.048 | Dibbivirus | Unclassified | Myoviridae | Pantoea |
| NC_048761 | Microbacterium phage Akoni | Microbacterium phage Akoni Microbacterium virus Akoni Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54307 | 60.14 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| NC_048760 | Salmonella phage Sepoy | Salmonella phage Sepoy Salmonella virus Sepoy Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114080 | 40.314 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048759 | Serratia phage MTx | Serratia phage MTx Serratia virus MTx Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68621 | 49.886 | Myosmarvirus | Unclassified | Myoviridae | Serratia |
| NC_048757 | Nodularia phage vB_NspS-kac68v161 | Nodularia phage vB_NspS-kac68v161 Nodularia virus kac68v161 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145400 | 40.235 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| NC_048754 | Lactobacillus phage SAC12B | Lactobacillus phage SAC12B Lactobacillus virus SAC12B Tybeckvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136608 | 32.411 | Tybeckvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048753 | Lactobacillus phage 3-521 | Lactobacillus phage 3-521 Lactobacillus virus 3-521 Watanabevirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140816 | 36.933 | Watanabevirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048752 | Lactobacillus phage 521B | Lactobacillus phage 521B Lactobacillus virus 521B Tybeckvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136442 | 32.268 | Tybeckvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048664 | Cronobacter phage vB_CsaP_009 | Cronobacter phage vB_CsaP_009 Cronobacter virus 009 Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92122 | 35.122 | Privateervirus | Unclassified | Podoviridae | Cronobacter |
| NC_048191 | Rheinheimera phage Barba21A | Rheinheimera phage Barba21A Rheinheimera virus Barba21A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80361 | 38.28 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| NC_048188 | Rheinheimera phage Barba8S | Rheinheimera phage Barba8S Rheinheimera virus Barba8S Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81920 | 38.315 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| NC_048187 | Rheinheimera phage Barba5S | Rheinheimera phage Barba5S Rheinheimera virus Barba5S Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82744 | 38.218 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| NC_042005 | Acinetobacter phage vB_AbaP_B5 | Acinetobacter phage vB_AbaP_B5 Acinetobacter virus B5 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41608 | 39.31 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW273926 | Pelagibacter phage HTVC204P | Pelagibacter phage HTVC204P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34069 | 31.028 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273925 | Pelagibacter phage HTVC203P | Pelagibacter phage HTVC203P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34938 | 32.097 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273924 | Pelagibacter phage HTVC100P | Pelagibacter phage HTVC100P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34606 | 31.847 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273923 | Pelagibacter phage HTVC035P | Pelagibacter phage HTVC035P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36066 | 31.908 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273922 | Pelagibacter phage HTVC034P | Pelagibacter phage HTVC034P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35450 | 32.623 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273921 | Pelagibacter phage HTVC028P | Pelagibacter phage HTVC028P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36388 | 33.126 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW273920 | Pelagibacter phage HTVC024P | Pelagibacter phage HTVC024P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35448 | 31.528 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MW281503 | Bacillus phage Anthos | Bacillus phage Anthos unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164822 | 37.75 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MW021756 | Cronobacter phage vB_CsaM_SemperBestia | Cronobacter phage vB_CsaM_SemperBestia unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177494 | 44.679 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| MW021748 | Citrobacter phage vB_CfrD_ZerotoHero | Citrobacter phage vB_CfrD_ZerotoHero unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49802 | 42.85 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| MW021765 | Klebsiella phage vB_KpnS_IMGroot | Klebsiella phage vB_KpnS_IMGroot Klebsiella virus IMGroot Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52866 | 51.347 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW021762 | Klebsiella phage vB_KpnS_SegesCirculi | Klebsiella phage vB_KpnS_SegesCirculi Klebsiella virus SegesCirculi Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50713 | 51.121 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW021759 | Serratia phage vB_SmaM_Hera | Serratia phage vB_SmaM_Hera unclassified Myosmarvirus Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67493 | 49.901 | Myosmarvirus | Unclassified | Myoviridae | Serratia |
| MW021757 | Citrobacter phage vB_CfrM_Sajous1 | Citrobacter phage vB_CfrM_Sajous1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157255 | 44.862 | Kuttervirus | Cvivirinae | Ackermannviridae | Citrobacter |
| MW021754 | Salmonella phage vB_SenM_FrontPhageNews | Salmonella phage vB_SenM_FrontPhageNews unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157832 | 44.612 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW021753 | Salmonella phage vB_SenM_AR2819 | Salmonella phage vB_SenM_AR2819 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156899 | 44.968 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW021752 | Klebsiella phage vB_KpnM_BovinicusUrsus | Klebsiella phage vB_KpnM_BovinicusUrsus unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166829 | 39.587 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MW021751 | Cronobacter phage vB_CsaM_Cronuts | Cronobacter phage vB_CsaM_Cronuts unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177882 | 44.54 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| MW021749 | Citrobacter phage vB_Cfr_Xman | Citrobacter phage vB_Cfr_Xman unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178331 | 43.174 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| MW001214 | Bacillus phage SP8 | Bacillus phage SP8 unclassified Okubovirus Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138741 | 40.297 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| MT863731 | Campylobacter phage F375 | Campylobacter phage F375 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131645 | 26.202 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863730 | Campylobacter phage F374 | Campylobacter phage F374 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131642 | 26.204 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863729 | Campylobacter phage F372 | Campylobacter phage F372 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131465 | 26.272 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863728 | Campylobacter phage F371 | Campylobacter phage F371 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132743 | 26.206 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863727 | Campylobacter phage F370 | Campylobacter phage F370 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132742 | 26.206 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863726 | Campylobacter phage F368 | Campylobacter phage F368 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131924 | 26.228 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863725 | Campylobacter phage F367 | Campylobacter phage F367 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131922 | 26.227 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863724 | Campylobacter phage F365 | Campylobacter phage F365 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131925 | 26.238 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863723 | Campylobacter phage F361 | Campylobacter phage F361 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131889 | 26.225 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863722 | Campylobacter phage F360 | Campylobacter phage F360 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131919 | 26.221 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863721 | Campylobacter phage F358 | Campylobacter phage F358 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131926 | 26.228 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863720 | Campylobacter phage F357 | Campylobacter phage F357 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131922 | 26.222 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863719 | Campylobacter phage F356 | Campylobacter phage F356 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133731 | 26.191 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863718 | Campylobacter phage F355 | Campylobacter phage F355 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133670 | 26.19 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863717 | Campylobacter phage F352 | Campylobacter phage F352 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131638 | 26.197 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863716 | Campylobacter phage F348 | Campylobacter phage F348 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131681 | 26.181 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863715 | Campylobacter phage F336 | Campylobacter phage F336 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131195 | 26.307 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT863714 | Campylobacter phage F207 | Campylobacter phage F207 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130779 | 26.254 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MW055915 | Mycobacterium phage Plumbus | Mycobacterium phage Plumbus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54468 | 61.133 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MW055914 | Microbacterium phage Xitlalli | Microbacterium phage Xitlalli unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53929 | 60.205 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MW055912 | Arthrobacter phage Truckee | Arthrobacter phage Truckee unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57724 | 50.547 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MW055911 | Microbacterium phage Thorongil | Microbacterium phage Thorongil unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41331 | 63.335 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MW055905 | Streptomyces phage Indigenous | Streptomyces phage Indigenous unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56119 | 59.5 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MW055901 | Gordonia phage Bunker | Gordonia phage Bunker Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45508 | 67.061 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| NC_049977 | Bacteroides phage crAss001 | Bacteroides phage crAss001 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102679 | 34.743 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| NC_049973 | Bacillus virus SRT01hs | Bacillus virus SRT01hs Gaunavirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20784 | 34.945 | Gaunavirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| NC_049961 | Bacillus phage BeachBum | Bacillus phage BeachBum Bacillus virus BeachBum Harambevirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21054 | 35.409 | Harambevirus | Unclassified | Salasmaviridae | Bacillus |
| NC_049960 | Bacillus phage Harambe | Bacillus phage Harambe Bacillus virus Harambe Harambevirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21684 | 35.293 | Harambevirus | Unclassified | Salasmaviridae | Bacillus |
| NC_049937 | Enterococcus phage Idefix | Enterococcus phage Idefix Enterococcus virus Idefix Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18168 | 33.201 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049931 | Enterococcus phage vB_EfaP_IME199 | Enterococcus phage vB_EfaP_IME199 Enterococcus virus IME199 Minhovirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18838 | 34.627 | Minhovirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049930 | Enterococcus phage vB_EfaP_Zip | Enterococcus phage vB_EfaP_Zip Enterococcus virus Zip Minhovirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18742 | 35.044 | Minhovirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_049929 | Microbacterium phage Teagan | Microbacterium phage Teagan Microbacterium virus Teagan Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41793 | 63.451 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| NC_049807 | Lactococcus phage AM6 | Lactococcus phage AM6 Lactococcus virus AM6 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62054 | 34.246 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049806 | Lactococcus phage LW31 | Lactococcus phage LW31 Lactococcus virus LW31 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60551 | 34.292 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049805 | Lactococcus phage phiQ1 | Lactococcus phage phiQ1 Lactococcus virus phiQ1 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59833 | 34.412 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| NC_049444 | Klebsiella phage Pylas | Klebsiella phage Pylas Klebsiella virus Pylas Pylasvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70408 | 41.213 | Pylasvirus | Humphriesvirinae | Schitoviridae | Klebsiella |
| NC_049443 | Salmonella phage SeSz-1 | Salmonella phage SeSz-1 Salmonella virus SeSz1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155297 | 44.768 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049442 | Arthrobacter phage KBurrousTX | Arthrobacter phage KBurrousTX Arthrobacter virus KBurrousTX Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71020 | 62.295 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_049440 | Salmonella phage SenALZ1 | Salmonella phage SenALZ1 Salmonella virus SenALZ1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154811 | 44.56 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049439 | Salmonella phage SenASZ3 | Salmonella phage SenASZ3 Salmonella virus SenASZ3 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157630 | 44.74 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049437 | Salmonella phage S118 | Salmonella phage S118 Salmonella virus S118 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157013 | 44.801 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049436 | Ruegeria phage vB_RpoP-V13 | Ruegeria phage vB_RpoP-V13 Ruegeria virus V13 Pomeroyivirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74830 | 50.771 | Pomeroyivirus | Rhodovirinae | Schitoviridae | Ruegeria |
| NC_049435 | Ruegeria phage vB_RpoP-V12 | Ruegeria phage vB_RpoP-V12 Ruegeria virus V12 Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74675 | 47.936 | Aorunvirus | Rhodovirinae | Schitoviridae | Ruegeria |
| NC_049433 | Escherichia phage EP75 | Escherichia phage EP75 Escherichia virus EP75 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158143 | 44.644 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| NC_049431 | Vibrio phage 1.097.O._10N.286.49.B3 | Vibrio phage 1.097.O._10N.286.49.B3 Vibrio virus 49B3 Dorisvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76918 | 42.596 | Dorisvirus | Pontosvirinae | Schitoviridae | Vibrio |
| NC_049430 | Vibrio phage 1.026.O._10N.222.49.C7 | Vibrio phage 1.026.O._10N.222.49.C7 Vibrio virus 49C7 Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75880 | 43.102 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| NC_049428 | Salmonella phage Mutine | Salmonella phage Mutine Salmonella virus Mutine Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161502 | 44.303 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049427 | Escherichia phage FEC14 | Escherichia phage FEC14 Escherichia virus FEC14 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158639 | 44.616 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| NC_049396 | Salmonella phage SP1 | Salmonella phage SP1 Salmonella virus SP1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156585 | 44.674 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049382 | Salmonella phage BSP101 | Salmonella phage BSP101 Salmonella virus BSP101 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157665 | 44.452 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049379 | Vibrio phage vB_VspP_pVa5 | Vibrio phage vB_VspP_pVa5 Vibrio virus PVA5 Galateavirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78145 | 43.158 | Galateavirus | Pontosvirinae | Schitoviridae | Vibrio |
| NC_049372 | Roseobacter phage RD-1410W1-01 | Roseobacter phage RD-1410W1-01 Roseobacter virus RD1410W1-01 Aoqinvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72674 | 49.527 | Aoqinvirus | Rhodovirinae | Schitoviridae | Roseobacter |
| NC_049371 | Dinoroseobacter phage DS-1410Ws-06 | Dinoroseobacter phage DS-1410Ws-06 Dinoroseobacter virus DS1410Ws06 Sanyabayvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76466 | 50.005 | Sanyabayvirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| NC_049350 | Vibrio phage VCO139 | Vibrio phage VCO139 Vibrio virus VCO139 Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68964 | 34.593 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| NC_049345 | Roseovarius Plymouth podovirus 1 | Roseovarius Plymouth podovirus 1 Plymouthvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74704 | 49.052 | Plymouthvirus | Rhodovirinae | Schitoviridae | Unspecified |
| NC_048629 | Clostridium phage CpV1 | Clostridium phage CpV1 Clostridium virus CpV1 Capvunavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16748 | 30.505 | Capvunavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| NC_048627 | Escherichia phage SP15 | Escherichia phage SP15 Escherichia virus SP15 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110964 | 39.134 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048650 | Streptomyces phage BRock | Streptomyces phage BRock Streptomyces virus BRock Borockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112523 | 52.316 | Borockvirus | Unclassified | Myoviridae | Streptomyces |
| NC_048785 | Salmonella phage SE24 | Salmonella phage SE24 Salmonella virus SE24 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114855 | 39.987 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_048751 | Pantoea phage vB_PagM_LIET2 | Pantoea phage vB_PagM_LIET2 Pantoea virus LIET2 Lietduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74710 | 53.989 | Lietduovirus | Unclassified | Myoviridae | Pantoea |
| NC_048749 | Gordonia phage Rickmore | Gordonia phage Rickmore Gordonia virus Rickmore Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60218 | 50.538 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| NC_048748 | Escherichia phage vB_EcoS_HdH2 | Escherichia phage vB_EcoS_HdH2 Escherichia virus HdH2 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120120 | 39.321 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048747 | Vibrio phage 2 TSL-2019 | Vibrio phage 2 TSL-2019 Vibrio phage 2TSL2019 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 242446 | 43.571 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| NC_048746 | Gordonia phage Sombrero | Gordonia phage Sombrero Gordonia virus Sombrero Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76485 | 59.018 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048743 | Mycobacterium phage DrLupo | Mycobacterium phage DrLupo Mycobacterium virus DrLupo Barnyardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70030 | 57.501 | Barnyardvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_048740 | Arthrobacter phage Mendel | Arthrobacter phage Mendel Arthobacter virus Mendel Anjalivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19428 | 59.826 | Anjalivirus | Unclassified | Podoviridae | Arthrobacter |
| NC_048739 | Arthrobacter phage Anjali | Arthrobacter phage Anjali Arthrobacter virus Anjali Anjalivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19679 | 59.373 | Anjalivirus | Unclassified | Podoviridae | Arthrobacter |
| NC_048738 | Clostridium phage CPD2 | Clostridium phage CPD2 Clostridium virus CPD2 Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18212 | 27.943 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| NC_048736 | Serratia phage vB_SmaA_3M | Serratia phage vB_SmaA_3M Serratia virus 3M Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159398 | 51.414 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| NC_048735 | Bacillus phage vB_BboS-125 | Bacillus phage vB_BboS-125 Bacillus virus 125 Elmenteitavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58528 | 48.568 | Elmenteitavirus | Unclassified | Myoviridae | Bacillus |
| NC_048733 | Microbacterium phage Burro | Microbacterium phage Burro Microbacterium virus Burro Burrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54473 | 64.338 | Burrovirus | Unclassified | Podoviridae | Microbacterium |
| NC_048731 | Escherichia phage fp01 | Escherichia phage fp01 Escherichia virus fp01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109515 | 39.003 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| NC_048728 | Streptomyces phage Comrade | Streptomyces phage Comrade Streptomyces virus Comrade Gilsonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129015 | 47.06 | Gilsonvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048725 | Bacillus phage Wes44 | Bacillus phage Wes44 Bacillus virus Wes44 Carmenvirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42248 | 42.028 | Carmenvirus | Gutmannvirinae | Siphoviridae | Bacillus |
| NC_048720 | Streptomyces phage Blueeyedbeauty | Streptomyces phage Blueeyedbeauty Streptomyces virus Blueeyedbeauty Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130473 | 47.88 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048719 | Streptomyces phage Annadreamy | Streptomyces phage Annadreamy Streptomyces virus Annadreamy Annadreamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125726 | 47.604 | Annadreamyvirus | Unclassified | Siphoviridae | Streptomyces |
| NC_048712 | Clostridium phage susfortuna | Clostridium phage susfortuna Clostridium virus susfortuna Susfortunavirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19046 | 29.224 | Susfortunavirus | Unclassified | Guelinviridae | Clostridium |
| NC_048709 | Vibrio phage YC | Vibrio phage YC Vibrio virus YC Campanilevirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147890 | 47.33 | Campanilevirus | Unclassified | Ackermannviridae | Vibrio |
| NC_048708 | Rhodococcus phage Takoda | Rhodococcus phage Takoda Rhodococcus virus Takoda Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46807 | 58.737 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_048707 | Clostridium phage CPS2 | Clostridium phage CPS2 Clostridium virus CPS2 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17961 | 33.3 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| NC_048706 | Streptomyces phage FlowerPower | Streptomyces phage FlowerPower Streptomyces virus FlowerPower Flowerpowervirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46133 | 60.681 | Flowerpowervirus | Beephvirinae | Podoviridae | Streptomyces |
| NC_048704 | Pectobacterium phage Nepra | Pectobacterium phage Nepra Pectobacterium virus Nepra Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74509 | 48.701 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| NC_048703 | Xanthomonas phage RiverRider | Xanthomonas phage RiverRider Xanthomonas virus RiverRider Riverridervirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76335 | 49.393 | Riverridervirus | Unclassified | Schitoviridae | Xanthomonas |
| NC_048702 | Erwinia phage vB_EamM-Bue1 | Erwinia phage vB_EamM-Bue1 Erwinia virus Bue1 Nezavisimistyvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164037 | 50.204 | Nezavisimistyvirus | Unclassified | Ackermannviridae | Erwinia |
| NC_048698 | Bacillus phage Carmen17 | Bacillus phage Carmen17 Bacillus virus Carmen17 Carmenvirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41820 | 42.037 | Carmenvirus | Gutmannvirinae | Siphoviridae | Bacillus |
| NC_048697 | Pseudomonas phage Littlefix | Pseudomonas phage Littlefix Pseudomonas virus Littlefix Littlefixvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74895 | 48.734 | Littlefixvirus | Unclassified | Schitoviridae | Pseudomonas |
| NC_048695 | Gordonia phage Adgers | Gordonia phage Adgers Gordonia virus Adgers Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76008 | 58.99 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_048687 | Pseudomonas phage PMBT14 | Pseudomonas phage PMBT14 Pseudomonas virus PMBT14 Knuthellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47820 | 54.548 | Knuthellervirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_048686 | Streptomyces phage Immanuel3 | Streptomyces phage Immanuel3 Streptomyces virus Immanuel3 Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46094 | 59.632 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| NC_048685 | Streptomyces phage Manuel | Streptomyces phage Manuel Streptomyces virus Manuel Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45177 | 60.095 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| NC_048684 | Streptomyces phage WRightOn | Streptomyces phage WRightOn Streptomyces virus WRightOn Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45221 | 60.313 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| NC_048679 | Leptospira phage LE4 | Leptospira phage LE4 Leptospira virus LE4 Nylescharonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47866 | 37.285 | Nylescharonvirus | Unclassified | Myoviridae | Leptospira |
| NC_048678 | Leptospira phage LE3 | Leptospira phage LE3 Leptospira virus LE3 Nylescharonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48288 | 37.312 | Nylescharonvirus | Unclassified | Myoviridae | Leptospira |
| NC_048677 | Lactobacillus phage LpeD | Lactobacillus phage LpeD Lactobacillus virus LpeD Elpedvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145162 | 34.112 | Elpedvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048674 | Aeromonas phage AS-sw | Aeromonas phage AS-sw Aeromonas virus ASsw Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 230024 | 38.657 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| NC_048673 | Aeromonas phage AS-zj | Aeromonas phage AS-zj Aeromonas virus ASzj Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229929 | 38.717 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| NC_048672 | Lentibacter virus ICBM1 | Lentibacter virus ICBM1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40163 | 47.023 | Siovirus | Cobavirinae | Zobellviridae | Lentibacter |
| NC_048671 | Lentibacter virus vB_LenP_ICBM2 | Lentibacter virus vB_LenP_ICBM2 unclassified Siovirus Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40907 | 47.816 | Siovirus | Cobavirinae | Zobellviridae | Lentibacter |
| NC_048669 | Rhodococcus phage Hiro | Rhodococcus phage Hiro Rhodococcus virus Hiro Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46854 | 58.719 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_048668 | Serratia phage vB_SmaM_ 2050HW | Serratia phage vB_SmaM_ 2050HW Serratia virus 2050HW Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 276025 | 46.793 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| NC_048661 | Clostridium phage CPS1 | Clostridium phage CPS1 Clostridium virus CPS1 Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19089 | 28.257 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| NC_048660 | Aeromonas phage phiA8-29 | Aeromonas phage phiA8-29 Aeromonas virus A8-29 Tedavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144974 | 48.18 | Tedavirus | Unclassified | Ackermannviridae | Aeromonas |
| NC_048659 | Pectobacterium phage phiA41 | Pectobacterium phage phiA41 Pectobacterium virus phiA41 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75764 | 48.701 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| NC_048654 | Pectobacterium phage vB_PatP_CB4 | Pectobacterium phage vB_PatP_CB4 Pectobacterium virus CB4 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76621 | 48.643 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| NC_048653 | Pectobacterium phage vB_PatP_CB1 | Pectobacterium phage vB_PatP_CB1 Pectobacterium virus CB1 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76041 | 48.691 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| NC_048646 | Cronobacter phage vB_CsaM_leE | Cronobacter phage vB_CsaM_leE Cronobacter virus leE Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181570 | 44.68 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| NC_048645 | Cronobacter phage vB_CsaM_leB | Cronobacter phage vB_CsaM_leB Cronobacter virus leB Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177907 | 44.871 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| NC_048639 | Pseudomonas phage ZC08 | Pseudomonas phage ZC08 Pseudomonas virus ZC08 Zicotriavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70774 | 43.074 | Zicotriavirus | Unclassified | Schitoviridae | Pseudomonas |
| NC_048638 | Pseudomonas phage ZC03 | Pseudomonas phage ZC03 Pseudomonas virus ZC03 Zicotriavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69844 | 42.601 | Zicotriavirus | Unclassified | Schitoviridae | Pseudomonas |
| NC_048134 | Lactobacillus phage Lpa804 | Lactobacillus phage Lpa804 Lactobacillus virus Lpa804 Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142567 | 36.623 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_048128 | Sulfolobales Beppu filamentous virus 2 | Sulfolobales Beppu filamentous virus 2 Alphalipothrixvirus SBFV2 Alphalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 38091 | 39.458 | Alphalipothrixvirus | Unclassified | Lipothrixviridae | Unspecified |
| NC_048127 | Sulfolobales Beppu filamentous virus 3 | Sulfolobales Beppu filamentous virus 3 Deltalipothrixvirus SBFV3 Deltalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 31324 | 37.926 | Deltalipothrixvirus | Unclassified | Lipothrixviridae | Unspecified |
| NC_048125 | Vibrio phage 1.261.O._10N.286.51.A7 | Vibrio phage 1.261.O._10N.286.51.A7 Vibrio virus 51A7 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71992 | 43.166 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| NC_048124 | Vibrio phage 1.224.A._10N.261.48.B1 | Vibrio phage 1.224.A._10N.261.48.B1 Vibrio virus 48B1 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71915 | 43.063 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| NC_048037 | Sulfolobus filamentous virus 1 | Sulfolobus filamentous virus 1 Alphalipothrixvirus SFV1 Alphalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 37311 | 35.802 | Alphalipothrixvirus | Unclassified | Lipothrixviridae | Sulfolobus |
| NC_048003 | Pseudomonas phage phCDa | Pseudomonas phage phCDa Pseudomonas virus phCDa Shizishanvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72821 | 53.268 | Shizishanvirus | Unclassified | Schitoviridae | Pseudomonas |
| NC_047978 | Erwinia phage Faunus | Erwinia phage Faunus Erwinia virus Faunus Faunusvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54065 | 43.649 | Faunusvirus | Cleopatravirinae | Chaseviridae | Erwinia |
| NC_047866 | Aeromonas phage Ahp1 | Aeromonas phage Ahp1 Aeromonas virus Ahp1 Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42167 | 58.864 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_047839 | Pseudoalteromonas phage J2-1 | Pseudoalteromonas phage J2-1 Pseudoalteromonas virus J2-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120295 | 35.844 | Unclassified | Unclassified | Myoviridae | Pseudoalteromonas |
| NC_047832 | Pasteurella phage PHB01 | Pasteurella phage PHB01 Pasteurella virus PHB01 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37287 | 40.725 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| NC_047824 | Shewanella phage SppYZU05 | Shewanella phage SppYZU05 Shewanella virus SppYZU05 Yushanvirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54319 | 50.64 | Yushanvirus | Nefertitivirinae | Chaseviridae | Shewanella |
| NC_047791 | Pectobacterium phage PP101 | Pectobacterium phage PP101 Pectobacterium virus PP101 Suwonvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53333 | 44.944 | Suwonvirus | Cleopatravirinae | Chaseviridae | Pectobacterium |
| NC_047746 | Vibrio phage Vc1 | Vibrio phage Vc1 Vibrio virus Vc1 Gutovirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44541 | 44.157 | Gutovirus | Colwellvirinae | Autographiviridae | Vibrio |
| NC_042136 | Vibrio phage VP4B | Vibrio phage VP4B Vibrio virus VP4B Tidunavirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 236053 | 43.256 | Tidunavirus | Gorgonvirinae | Myoviridae | Vibrio |
| NC_042112 | Campylobacter virus CP81 | Campylobacter virus CP81 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132454 | 26.042 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_042100 | Vibrio phage Aphrodite1 | Vibrio phage Aphrodite1 Vibrio virus Aphrodite1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237722 | 43.36 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| NC_042061 | Salmonella phage STML-13-1 | Salmonella phage STML-13-1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157235 | 44.782 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_042053 | Arthrobacter phage PrincessTrina | Arthrobacter phage PrincessTrina Arthrobacter virus PrincessTrina Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70265 | 61.584 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_042000 | Arthrobacter phage Colucci | Arthrobacter phage Colucci Arthrobacter virus Colucci Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70707 | 61.701 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_041953 | Pseudomonas phage PaMx25 | Pseudomonas phage PaMx25 Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57899 | 58.595 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| NC_041916 | Vibrio phage pTD1 | Vibrio phage pTD1 Vibrio virus pTD1 Tidunavirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239276 | 44.546 | Tidunavirus | Gorgonvirinae | Myoviridae | Vibrio |
| NC_041907 | Pseudomonas phage PA26 | Pseudomonas phage PA26 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72321 | 54.818 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_041896 | Campylobacter phage vB_CjeM_Los1 | Campylobacter phage vB_CjeM_Los1 Campylobacter virus Los1 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134073 | 26.196 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_028693 | Enterococcus phage vB_EfaP_IME195 | Enterococcus phage vB_EfaP_IME195 Enterococcus virus IME195 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18655 | 33.203 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_036595 | Aeropyrum globular virus 1 | Aeropyrum globular virus 1 unclassified archaeal viruses Viruses | 18222 | 55.801 | Unclassified | Unclassified | Unclassified | Aeropyrum |
| NC_035464 | Metallosphaera turreted icosahedral virus | Metallosphaera turreted icosahedral virus unclassified archaeal viruses Viruses | 9780 | 53.998 | Unclassified | Unclassified | Unclassified | Metallosphaera |
| NC_027348 | Delftia phage RG-2014 | Delftia phage RG-2014 Delftia virus RG2014 Dendoorenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73882 | 59.889 | Dendoorenvirus | Unclassified | Schitoviridae | Delftia |
| NC_031927 | Synechococcus phage S-CAM7 | Synechococcus phage S-CAM7 Synechococcus virus SCAM7 Mazuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216121 | 41.18 | Mazuvirus | Unclassified | Myoviridae | Synechococcus |
| NC_031909 | Campylobacter phage PC14 | Campylobacter phage PC14 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134927 | 26.16 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_031902 | Salmonella phage 100268_sal2 | Salmonella phage 100268_sal2 Salmonella virus 100268sal2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125114 | 40.218 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_031274 | Pseudomonas phage O4 | Pseudomonas phage O4 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50509 | 44.568 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| NC_031229 | Gordonia phage Hotorobo | Gordonia phage Hotorobo unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76972 | 58.859 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| NC_023594 | Shewanella phage Spp001 | Shewanella phage Spp001 Shewanella virus Spp001 Yushanvirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54791 | 49.417 | Yushanvirus | Nefertitivirinae | Chaseviridae | Shewanella |
| NC_031104 | Pseudomonas phage vB_PaeP_MAG4 | Pseudomonas phage vB_PaeP_MAG4 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72979 | 54.82 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_031063 | Pseudomonas phage PEV2 | Pseudomonas phage PEV2 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72697 | 54.863 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_031058 | Pseudomonas phage NP1 | Pseudomonas phage NP1 Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58566 | 58.399 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| NC_031057 | Citrobacter phage vB_CfrM_CfP1 | Citrobacter phage vB_CfrM_CfP1 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180219 | 43.119 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_029314 | Acidianus rod-shaped virus 2 | Acidianus rod-shaped virus 2 Hoswirudivirus ARV2 Hoswirudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 29763 | 32.349 | Hoswirudivirus | Unclassified | Rudiviridae | Acidianus |
| NC_029118 | Lactococcus phage GE1 | Lactococcus phage GE1 Lactococcus virus GE1 Chertseyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24847 | 37.872 | Chertseyvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_029101 | Pseudomonas phage YH30 | Pseudomonas phage YH30 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72192 | 54.916 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_029094 | Pseudoalteromonas phage H101 | Pseudoalteromonas phage H101 Pseudoalteromonas virus H101 Shandongvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131903 | 37.364 | Shandongvirus | Unclassified | Myoviridae | Pseudoalteromonas |
| NC_028755 | Citrobacter phage Margaery | Citrobacter phage Margaery Citrobacter virus Margaery Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178182 | 44.868 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_029017 | Pseudomonas phage KPP21 | Pseudomonas phage KPP21 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73420 | 53.499 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_029013 | Citrobacter phage IME-CF2 | Citrobacter phage IME-CF2 Citrobacter virus CF2 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177688 | 43.181 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_028885 | Pseudomonas phage DL64 | Pseudomonas phage DL64 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72378 | 54.953 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_028831 | Escherichia phage slur01 | Escherichia phage slur01 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61235 | 44.429 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| NC_028799 | Vibrio phage phi 1 | Vibrio phage phi 1 Vibrio virus phi1 Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66708 | 34.503 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| NC_028789 | Bacillus phage VMY22 | Bacillus phage VMY22 Bacillus virus VMY22 Mingyongvirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18609 | 36.364 | Mingyongvirus | Unclassified | Salasmaviridae | Bacillus |
| NC_028776 | Escherichia phage CAjan | Escherichia phage CAjan Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59670 | 44.708 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| NC_027988 | Citrobacter phage CVT22 | Citrobacter phage CVT22 Citrobacter virus CVT22 Citrovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47636 | 41.632 | Citrovirus | Unclassified | Zobellviridae | Citrobacter |
| NC_018087 | Acinetobacter phage ZZ1 | Acinetobacter phage ZZ1 Acinetobacter virus ZZ1 Zedzedvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166687 | 34.409 | Zedzedvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_027388 | Pseudomonas phage YH6 | Pseudomonas phage YH6 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73050 | 54.88 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_027381 | Escherichia phage Pollock | Escherichia phage Pollock Pollockvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68365 | 36.02 | Pollockvirus | Humphriesvirinae | Schitoviridae | Escherichia |
| NC_027341 | Lactococcus phage WRP3 | Lactococcus phage WRP3 Lactococcus virus WRP3 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130008 | 32.36 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_027345 | Pseudomonas phage Pa2 | Pseudomonas phage Pa2 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73008 | 54.857 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_027340 | Erwinia phage phiEa2809 | Erwinia phage phiEa2809 Nezavisimistyvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162160 | 50.28 | Nezavisimistyvirus | Unclassified | Ackermannviridae | Erwinia |
| NC_027378 | Escherichia phage Seurat | Escherichia phage Seurat Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56776 | 44.575 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| NC_027374 | Bacillus phage Moonbeam | Bacillus phage Moonbeam Bacillus virus Moonbeam Moonbeamvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161239 | 40.132 | Moonbeamvirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_027366 | Bacillus phage Mater | Bacillus phage Mater Bacillus virus Mater Matervirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164302 | 39.466 | Matervirus | Bastillevirinae | Herelleviridae | Bacillus |
| NC_027120 | Lactococcus phage 1358 | Lactococcus phage 1358 Lactococcus virus 1358 Whiteheadvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36892 | 51.038 | Whiteheadvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_026606 | Arthrobacter phage vB_ArtM-ArV1 | Arthrobacter phage vB_ArtM-ArV1 Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71200 | 61.621 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| NC_025446 | Escherichia phage ECML-4 | Escherichia phage ECML-4 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157308 | 45.032 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| NC_025414 | Citrobacter phage Miller | Citrobacter phage Miller Citrobacter virus Miller Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178171 | 43.13 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| NC_025467 | Enterococcus phage vB_Efae230P-4 | Enterococcus phage vB_Efae230P-4 Enterococcus virus Efae230P4 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17972 | 32.757 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| NC_025459 | Aeromonas phage pAh6-C | Aeromonas phage pAh6-C Aeromonas virus pAh6C Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53744 | 52.826 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| NC_024371 | Mycobacterium phage Damien | Mycobacterium phage Damien unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68386 | 57.604 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_024367 | Dinoroseobacter phage DFL12phi1 | Dinoroseobacter phage DFL12phi1 Baltimorevirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75028 | 49.26 | Baltimorevirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| NC_024358 | Anabaena phage A-4L | Anabaena phage A-4L Anabaena virus A4L Kozyakovvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41750 | 43.437 | Kozyakovvirus | Unclassified | Podoviridae | Anabaena |
| NC_024215 | Lactococcus phage P078 | Lactococcus phage P078 Lactococcus virus P078 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54452 | 33.516 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_024214 | Lactococcus phage P162 | Lactococcus phage P162 Lactococcus virus P162 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55584 | 33.333 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_024208 | Lactococcus phage P118 | Lactococcus phage P118 Lactococcus virus P118 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54173 | 33.495 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_024203 | Lactococcus phage P092 | Lactococcus phage P092 Lactococcus virus P092 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53513 | 33.528 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| NC_024149 | Lactococcus phage WP-2 | Lactococcus phage WP-2 Lactococcus virus WP2 Negarvirus Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18899 | 31.118 | Negarvirus | Unclassified | Rountreeviridae | Lactococcus |
| NC_024140 | Pseudomonas phage vB_PaeP_C2-10_Ab09 | Pseudomonas phage vB_PaeP_C2-10_Ab09 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72028 | 54.897 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_022768 | Salmonella phage Maynard | Salmonella phage Maynard Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154701 | 45.552 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_023865 | Pectobacterium phage PM1 | Pectobacterium phage PM1 Pectobacterium virus PM1 Suwonvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55098 | 44.925 | Suwonvirus | Cleopatravirinae | Chaseviridae | Pectobacterium |
| NC_023856 | Salmonella phage vB-SalM-SJ2 | Salmonella phage vB-SalM-SJ2 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152460 | 44.878 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_020862 | Sulfitobacter phage phiCB2047-B | Sulfitobacter phage phiCB2047-B Sulfitobacter virus phiCB2047B Raunefjordenvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74485 | 42.983 | Raunefjordenvirus | Rhodovirinae | Schitoviridae | Sulfitobacter |
| NC_023581 | Acinetobacter phage Presley | Acinetobacter phage Presley Acinetobacter virus Presley Presleyvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77792 | 37.766 | Presleyvirus | Unclassified | Schitoviridae | Acinetobacter |
| NC_023578 | Mycobacterium phage Oaker | Mycobacterium phage Oaker unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69099 | 57.546 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_023574 | Lactococcus phage phiL47 | Lactococcus phage phiL47 Lactococcus virus phiL47 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128546 | 32.532 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_022772 | Salmonella phage Marshall | Salmonella phage Marshall Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156338 | 45.587 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_021806 | Cellulophaga phage phi14:2 | Cellulophaga phage phi14:2 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100418 | 29.605 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021803 | Cellulophaga phage phi13:2 | Cellulophaga phage phi13:2 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72369 | 32.873 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021796 | Cellulophaga phage phi38:1 | Cellulophaga phage phi38:1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72534 | 38.055 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021794 | Cellulophaga phage phi18:3 | Cellulophaga phage phi18:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71443 | 32.884 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021792 | Cellulophaga phage phi46:3 | Cellulophaga phage phi46:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72961 | 32.696 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| NC_021772 | Salmonella phage FSL SP-058 | Salmonella phage FSL SP-058 Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72394 | 39.586 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| NC_021782 | Salmonella phage FSL SP-076 | Salmonella phage FSL SP-076 Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72098 | 39.535 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| NC_021344 | Escherichia phage Lw1 | Escherichia phage Lw1 Escherichia virus Lw1 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176227 | 43.491 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_021335 | Halorubrum Tailed Virus 7 | Halorubrum Tailed Virus 7 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69048 | 59.64 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| NC_021300 | Pseudoalteromonas phage RIO-1 | Pseudoalteromonas phage RIO-1 Pseudoalteromonas virus RIO1 Melvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43882 | 44.722 | Melvirus | Unclassified | Zobellviridae | Pseudoalteromonas |
| NC_020849 | Pseudoalteromonas phage pYD6-A | Pseudoalteromonas phage pYD6-A Pseudoalteromonas virus pYD6A Matsuvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76802 | 38.658 | Matsuvirus | Fuhrmanvirinae | Schitoviridae | Pseudoalteromonas |
| NC_020848 | Vibrio phage VBP47 | Vibrio phage VBP47 Vibrio virus VBP47 Stoningtonvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76705 | 42.456 | Stoningtonvirus | Fuhrmanvirinae | Schitoviridae | Vibrio |
| NC_020481 | Pelagibacter phage HTVC010P | Pelagibacter phage HTVC010P Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34892 | 31.967 | Unclassified | Unclassified | Siphoviridae | Pelagibacter |
| NC_020159 | Halovirus HSTV-2 | Halovirus HSTV-2 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68527 | 60.338 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| NC_020083 | Serratia phage phiMAM1 | Serratia phage phiMAM1 Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157834 | 51.984 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| NC_019540 | Salinivibrio phage CW02 | Salinivibrio phage CW02 Salinivibrio virus CW02 Salinovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49390 | 47.672 | Salinovirus | Unclassified | Zobellviridae | Salinivibrio |
| NC_019538 | Aeromonas phage CC2 | Aeromonas phage CC2 Aeromonas virus CC2 Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231743 | 38.821 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| NC_019523 | Clostridium phage phi24R | Clostridium phage phi24R Clostridium virus phi24R Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18919 | 27.803 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| NC_019514 | Erwinia phage vB_EamP-S6 | Erwinia phage vB_EamP-S6 Erwinia virus S6 Waedenswilvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74669 | 52.09 | Waedenswilvirus | Unclassified | Schitoviridae | Erwinia |
| NC_019507 | Campylobacter phage CP21 | Campylobacter phage CP21 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182833 | 27.151 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_019504 | Erwinia phage vB_EamM-Y2 | Erwinia phage vB_EamM-Y2 Erwinia virus Y2 Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56621 | 44.229 | Loessnervirus | Cleopatravirinae | Chaseviridae | Erwinia |
| NC_019485 | Enterobacter phage EcP1 | Enterobacter phage EcP1 Enterobacter virus EcP1 Eceepunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59080 | 36.875 | Eceepunavirus | Unclassified | Schitoviridae | Enterobacter |
| NC_019413 | Sulfolobales Mexican rudivirus 1 | Sulfolobales Mexican rudivirus 1 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 27431 | 46.597 | Rudivirus | Unclassified | Rudiviridae | Unspecified |
| NC_019398 | Cronobacter phage vB_CsaM_GAP161 | Cronobacter phage vB_CsaM_GAP161 Cronobacter virus GAP161 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178193 | 44.523 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| NC_018861 | Campylobacter phage CP30A | Campylobacter phage CP30A Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133572 | 26.127 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| NC_018280 | Celeribacter phage P12053L | Celeribacter phage P12053L Celeribacter virus P12053L Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38889 | 46.1 | Siovirus | Cobavirinae | Zobellviridae | Celeribacter |
| NC_018084 | Clostridium phage phiZP2 | Clostridium phage phiZP2 Clostridium virus phiZP2 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18078 | 34.478 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| NC_018083 | Clostridium phage phiCPV4 | Clostridium phage phiCPV4 Clostridium virus phiCPV4 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17972 | 34.487 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| NC_017981 | Xanthomonas phage vB_XveM_DIBBI | Xanthomonas phage vB_XveM_DIBBI Xanthomonas virus DIBBI Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49981 | 52.402 | Dibbivirus | Unclassified | Myoviridae | Xanthomonas |
| NC_017980 | Clostridium phage phiCP7R | Clostridium phage phiCP7R Clostridium virus phiCP7R Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18397 | 34.81 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| NC_017976 | Bacillus phage PBC1 | Bacillus phage PBC1 Bacillus virus PBC1 Pebcunavirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41164 | 41.684 | Pebcunavirus | Gutmannvirinae | Siphoviridae | Bacillus |
| NC_016650 | Rhodococcus virus RGL3 | Rhodococcus virus RGL3 Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48072 | 62.66 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| NC_015296 | Salmonella virus ViI | Salmonella virus ViI Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157061 | 45.22 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_014663 | Acinetobacter phage Acj9 | Acinetobacter phage Acj9 Acinetobacter virus Acj9 Acajnonavirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169947 | 40.028 | Acajnonavirus | Twarogvirinae | Myoviridae | Acinetobacter |
| NC_014321 | Hyperthermophilic Archaeal Virus 2 | Hyperthermophilic Archaeal Virus 2 unclassified archaeal viruses Viruses | 17666 | 52.083 | Unclassified | Unclassified | Unclassified | Unspecified |
| NC_013691 | Pseudomonas phage LUZ7 | Pseudomonas phage LUZ7 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74901 | 53.218 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_013692 | Pseudomonas phage LIT1 | Pseudomonas phage LIT1 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72544 | 55.019 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| NC_013638 | Pseudomonas phage phi-2 | Pseudomonas phage phi-2 Pseudomonas virus f2 Tunggulvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43144 | 58.914 | Tunggulvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| NC_012663 | Lactococcus phage P087 | Lactococcus phage P087 Lactococcus virus P087 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60074 | 34.384 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| NC_012530 | Lactobacillus phage Lb338-1 | Lactobacillus phage Lb338-1 Mooreparkvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142111 | 37.398 | Mooreparkvirus | Unclassified | Herelleviridae | Lactobacillus |
| NC_011292 | Mycobacterium phage Konstantine | Mycobacterium phage Konstantine Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68952 | 57.337 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_011290 | Mycobacterium phage Gumball | Mycobacterium phage Gumball Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64807 | 59.554 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| NC_051721 | Mycobacterium phage MooMoo | Mycobacterium phage MooMoo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55178 | 61.969 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| NC_051641 | Mycobacterium phage Byougenkin | Mycobacterium phage Byougenkin unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54685 | 61.425 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051631 | Mycobacterium phage Zavala | Mycobacterium phage Zavala unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59969 | 66.593 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051629 | Mycobacterium phage Yuna | Mycobacterium phage Yuna unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62192 | 67.441 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051628 | Mycobacterium phage Urkel | Mycobacterium phage Urkel unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60526 | 66.224 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051627 | Mycobacterium phage SirPhilip | Mycobacterium phage SirPhilip unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61882 | 66.701 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051626 | Mycobacterium phage Prithvi | Mycobacterium phage Prithvi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60311 | 66.771 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051624 | Mycobacterium phage Nibb | Mycobacterium phage Nibb unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62293 | 67.565 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051623 | Mycobacterium phage Marshawn | Mycobacterium phage Marshawn unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61464 | 68.146 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051622 | Mycobacterium phage MarkPhew | Mycobacterium phage MarkPhew unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62153 | 67.86 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051621 | Mycobacterium phage LaterM | Mycobacterium phage LaterM unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60143 | 66.55 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051620 | Mycobacterium phage LastHope | Mycobacterium phage LastHope unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60934 | 66.876 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051619 | Mycobacterium phage KiSi | Mycobacterium phage KiSi unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62558 | 68.749 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051617 | Mycobacterium phage Chris | Mycobacterium phage Chris unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62067 | 67.226 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051616 | Mycobacterium phage Bella96 | Mycobacterium phage Bella96 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60746 | 66.143 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051615 | Mycobacterium phage Beezoo | Mycobacterium phage Beezoo unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60494 | 66.501 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051614 | Mycobacterium phage Apocalypse | Mycobacterium phage Apocalypse unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59947 | 66.37 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051613 | Mycobacterium phage Amelie | Mycobacterium phage Amelie unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56439 | 67.087 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051612 | Mycobacterium phage AlishaPH | Mycobacterium phage AlishaPH unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57034 | 66.606 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051611 | Mycobacterium phage Adonis | Mycobacterium phage Adonis unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60031 | 66.676 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051610 | Mycobacterium phage Adephagia | Mycobacterium phage Adephagia unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59646 | 66.606 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051609 | Mycobacterium phage Ellie | Mycobacterium phage Ellie unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61945 | 67.042 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051608 | Mycobacterium phage Ekdilam | Mycobacterium phage Ekdilam unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 67.432 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051607 | Mycobacterium phage DarthP | Mycobacterium phage DarthP unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61594 | 67.244 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051606 | Mycobacterium phage Amohnition | Mycobacterium phage Amohnition unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61761 | 67.246 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051605 | Mycobacterium phage Amgine | Mycobacterium phage Amgine unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62236 | 66.447 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051604 | Mycobacterium phage Hurricane | Mycobacterium phage Hurricane unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61318 | 67.14 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051603 | Mycobacterium phage Ximenita | Mycobacterium phage Ximenita unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61027 | 66.741 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051601 | Mycobacterium phage Krueger | Mycobacterium phage Krueger unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60321 | 66.541 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051600 | Mycobacterium phage Bryler | Mycobacterium phage Bryler unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57666 | 66.098 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051599 | Mycobacterium phage Unicorn | Mycobacterium phage Unicorn unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61208 | 66.184 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051596 | Mycobacterium phage Rando14 | Mycobacterium phage Rando14 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59925 | 64.295 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051595 | Mycobacterium phage Paola | Mycobacterium phage Paola unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61535 | 65.01 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051594 | Mycobacterium phage Leston | Mycobacterium phage Leston unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61808 | 64.95 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051592 | Mycobacterium phage Patt | Mycobacterium phage Patt unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54611 | 67.604 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051591 | Mycobacterium phage Taquito | Mycobacterium phage Taquito unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58390 | 67.467 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051590 | Mycobacterium phage Eponine | Mycobacterium phage Eponine unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58678 | 67.419 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051589 | Mycobacterium phage Henu3 | Mycobacterium phage Henu3 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58899 | 67.612 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_051587 | Mycobacterium phage Serendipitous | Mycobacterium phage Serendipitous unclassified Acadianvirus Acadianvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69867 | 68.008 | Acadianvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051586 | Mycobacterium phage Imvubu | Mycobacterium phage Imvubu unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69897 | 70.497 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051585 | Mycobacterium phage CRB2 | Mycobacterium phage CRB2 unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72217 | 69.655 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051584 | Mycobacterium phage Quesadilla | Mycobacterium phage Quesadilla unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70992 | 69.86 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051583 | Mycobacterium phage Saguaro | Mycobacterium phage Saguaro unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69448 | 69.538 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051582 | Mycobacterium phage LilMcDreamy | Mycobacterium phage LilMcDreamy unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71552 | 69.347 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051581 | Mycobacterium phage BirdsNest | Mycobacterium phage BirdsNest unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70668 | 70.081 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_051580 | Mycobacterium phage Thonko | Mycobacterium phage Thonko unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69471 | 68.633 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| NC_047896 | Acinetobacter phage SWH-Ab-1 | Acinetobacter phage SWH-Ab-1 Acinetobacter virus SWHAb1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41567 | 39.416 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| NC_043027 | Bacillus phage PBS1 | Bacillus phage PBS1 Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252197 | 27.71 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| NC_042115 | Pseudomonas phage vB_PaeS_PAO1_Ab19 | Pseudomonas phage vB_PaeS_PAO1_Ab19 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58139 | 63.303 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| NC_042013 | Agrobacterium phage Atu_ph07 | Agrobacterium phage Atu_ph07 Agrobacterium virus Atuph07 Polybotosvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 490380 | 37.06 | Polybotosvirus | Unclassified | Myoviridae | Agrobacterium |
| NC_015251 | Aeromonas phage 65 | Aeromonas phage 65 Ishigurovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235229 | 37.196 | Ishigurovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| NC_014467 | Escherichia virus RB16 | Escherichia virus RB16 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176788 | 43.521 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_012697 | Silicibacter phage DSS3phi2 | Silicibacter phage DSS3phi2 unclassified Aorunvirus Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74611 | 47.915 | Aorunvirus | Rhodovirinae | Schitoviridae | Silicibacter |
| NC_012696 | Sulfitobacter phage EE36phi1 | Sulfitobacter phage EE36phi1 Sulfitobacter virus EE36phi1 Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73325 | 47.04 | Aorunvirus | Rhodovirinae | Schitoviridae | Sulfitobacter |
| NC_010576 | Lactococcus phage 1706 | Lactococcus phage 1706 Lactococcus virus 1706 Fremauxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55597 | 33.689 | Fremauxvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_010537 | Acidianus filamentous virus 9 | Acidianus filamentous virus 9 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 41172 | 34.645 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_010154 | Acidianus filamentous virus 8 | Acidianus filamentous virus 8 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 38179 | 36.803 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_010153 | Acidianus filamentous virus 7 | Acidianus filamentous virus 7 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36895 | 37.973 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_010152 | Acidianus filamentous virus 6 | Acidianus filamentous virus 6 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 39577 | 36.87 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_010155 | Acidianus filamentous virus 3 | Acidianus filamentous virus 3 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 40449 | 36.854 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_003214 | Sulfolobus islandicus filamentous virus | Sulfolobus islandicus filamentous virus Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 40900 | 33.352 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Sulfolobus |
| NC_009884 | Acidianus filamentous virus 2 | Acidianus filamentous virus 2 Deltalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 31787 | 35.656 | Deltalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_009643 | Actinomyces virus Av1 | Actinomyces virus Av1 Dybvigvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17171 | 49.502 | Dybvigvirus | Unclassified | Podoviridae | Actinomyces |
| NC_009551 | Phormidium virus WMP3 | Phormidium virus WMP3 Wumptrevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43249 | 46.48 | Wumptrevirus | Unclassified | Podoviridae | Phormidium |
| NC_009452 | Acidianus bottle-shaped virus | Acidianus bottle-shaped virus Bottigliavirus ABV Bottigliavirus Ampullaviridae Viruses | 23814 | 34.564 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| NC_009015 | Burkholderia phage BcepF1 | Burkholderia phage BcepF1 Bcepfunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72415 | 55.893 | Bcepfunavirus | Unclassified | Myoviridae | Burkholderia |
| NC_008367 | Phormidium phage Pf-WMP4 | Phormidium phage Pf-WMP4 Wumpquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40938 | 51.803 | Wumpquatrovirus | Unclassified | Podoviridae | Phormidium |
| NC_008364 | Lactococcus phage Q54 | Lactococcus phage Q54 Lactococcus virus Q54 Questintvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26537 | 37.099 | Questintvirus | Unclassified | Siphoviridae | Lactococcus |
| NC_008198 | Mycobacterium phage PBI1 | Mycobacterium phage PBI1 Mycobacterium virus PBI1 Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64494 | 59.711 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| NC_008200 | Mycobacterium phage PLot | Mycobacterium phage PLot Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64787 | 59.663 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| NC_007808 | Pseudomonas phage PA11 | Pseudomonas phage PA11 Pseudomonas virus PA11 Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49639 | 44.783 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| NC_007023 | Escherichia virus RB43 | Escherichia virus RB43 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180500 | 43.199 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| NC_006556 | Thermoproteus tenax spherical virus 1 | Thermoproteus tenax spherical virus 1 Alphaglobulovirus TTSV1 Alphaglobulovirus Globuloviridae Viruses | 20933 | 49.529 | Alphaglobulovirus | Unclassified | Globuloviridae | Thermoproteus |
| NC_005830 | Acidianus filamentous virus 1 | Acidianus filamentous virus 1 Gammalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 20869 | 36.94 | Gammalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| NC_002649 | Bacillus phage GA1 | Bacillus phage GA1 Gaunavirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21129 | 34.654 | Gaunavirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| NC_002519 | Roseobacter phage SIO1 | Roseobacter phage SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39898 | 46.16 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| NC_002515 | Mycoplasma virus P1 | Mycoplasma virus P1 Delislevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11660 | 26.792 | Delislevirus | Unclassified | Podoviridae | Mycoplasma |
| NC_001825 | Streptococcus phage Cp1 | Streptococcus phage Cp1 Cepunavirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19343 | 38.831 | Cepunavirus | Unclassified | Salasmaviridae | Streptococcus |
| NC_047801 | Pectobacterium phage PP47 | Pectobacterium phage PP47 Pectobacterium virus PP47 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40995 | 48.889 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_047797 | Pectobacterium phage PP81 | Pectobacterium phage PP81 Pectobacterium virus PP81 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40751 | 48.917 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| NC_049497 | Salmonella phage Se-J | Salmonella phage Se-J Salmonella virus SeJ Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160400 | 44.694 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049496 | Salmonella phage Se-G | Salmonella phage Se-G Salmonella virus SeG Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158892 | 44.71 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_049495 | Salmonella phage Se-B | Salmonella phage Se-B Salmonella virus SeB Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158087 | 44.522 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| NC_048630 | Pseudoalteromonas phage HP1 | Pseudoalteromonas phage HP1 Pseudoalteromonas virus HP1 Melvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45035 | 44.665 | Melvirus | Unclassified | Zobellviridae | Pseudoalteromonas |
| NC_043767 | Mycobacterium virus TA17a | Mycobacterium virus TA17a Rosebushvirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67324 | 68.984 | Rosebushvirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| MG835568 | Klebsiella phage SH-Kp 152410 | Klebsiella phage SH-Kp 152410 Klebsiella virus SHKp152410 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40945 | 52.346 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN854830 | Enterococcus phage PBEF129 | Enterococcus phage PBEF129 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141520 | 35.907 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MT497273 | Rheinheimera phage vB_RspM_barba_12-9G | Rheinheimera phage vB_RspM_barba_12-9G unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79770 | 38.192 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497272 | Rheinheimera phage vB_RspM_barba_12-9F | Rheinheimera phage vB_RspM_barba_12-9F unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.298 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497271 | Rheinheimera phage vB_RspM_barba_12-9E | Rheinheimera phage vB_RspM_barba_12-9E unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80423 | 38.256 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497270 | Rheinheimera phage vB_RspM_barba_12-9D | Rheinheimera phage vB_RspM_barba_12-9D unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.304 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497269 | Rheinheimera phage vB_RspM_barba_12-9C | Rheinheimera phage vB_RspM_barba_12-9C unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.291 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497268 | Rheinheimera phage vB_RspM_barba_12-9B | Rheinheimera phage vB_RspM_barba_12-9B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497267 | Rheinheimera phage vB_RspM_barba_12-9A | Rheinheimera phage vB_RspM_barba_12-9A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.262 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497266 | Rheinheimera phage vB_RspM_barba_12-8I | Rheinheimera phage vB_RspM_barba_12-8I unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497265 | Rheinheimera phage vB_RspM_barba_12-8H | Rheinheimera phage vB_RspM_barba_12-8H unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497264 | Rheinheimera phage vB_RspM_barba_12-8G | Rheinheimera phage vB_RspM_barba_12-8G unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497263 | Rheinheimera phage vB_RspM_barba_12-8F | Rheinheimera phage vB_RspM_barba_12-8F unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80435 | 38.246 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497262 | Rheinheimera phage vB_RspM_barba_12-8E | Rheinheimera phage vB_RspM_barba_12-8E unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80333 | 38.161 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497261 | Rheinheimera phage vB_RspM_barba_12-8C | Rheinheimera phage vB_RspM_barba_12-8C unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80394 | 38.244 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497260 | Rheinheimera phage vB_RspM_barba_12-8B | Rheinheimera phage vB_RspM_barba_12-8B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497259 | Rheinheimera phage vB_RspM_barba_12-8A | Rheinheimera phage vB_RspM_barba_12-8A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497258 | Rheinheimera phage vB_RspM_barba_12-7J | Rheinheimera phage vB_RspM_barba_12-7J unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80378 | 38.237 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497257 | Rheinheimera phage vB_RspM_barba_12-7I | Rheinheimera phage vB_RspM_barba_12-7I unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497256 | Rheinheimera phage vB_RspM_barba_12-7H | Rheinheimera phage vB_RspM_barba_12-7H unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80347 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497255 | Rheinheimera phage vB_RspM_barba_12-7G | Rheinheimera phage vB_RspM_barba_12-7G unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.246 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497254 | Rheinheimera phage vB_RspM_barba_12-7F | Rheinheimera phage vB_RspM_barba_12-7F unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80616 | 38.317 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497253 | Rheinheimera phage vB_RspM_barba_12-7E | Rheinheimera phage vB_RspM_barba_12-7E unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497252 | Rheinheimera phage vB_RspM_barba_12-7D | Rheinheimera phage vB_RspM_barba_12-7D unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.22 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497251 | Rheinheimera phage vB_RspM_barba_12-7C | Rheinheimera phage vB_RspM_barba_12-7C unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80446 | 38.247 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497250 | Rheinheimera phage vB_RspM_barba_12-7B | Rheinheimera phage vB_RspM_barba_12-7B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.295 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497249 | Rheinheimera phage vB_RspM_barba_12-7A | Rheinheimera phage vB_RspM_barba_12-7A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.248 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497248 | Rheinheimera phage vB_RspM_barba_12-6J | Rheinheimera phage vB_RspM_barba_12-6J unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.246 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497247 | Rheinheimera phage vB_RspM_barba_12-6I | Rheinheimera phage vB_RspM_barba_12-6I unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497246 | Rheinheimera phage vB_RspM_barba_12-6H | Rheinheimera phage vB_RspM_barba_12-6H unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497245 | Rheinheimera phage vB_RspM_barba_12-6G | Rheinheimera phage vB_RspM_barba_12-6G unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497244 | Rheinheimera phage vB_RspM_barba_12-6F | Rheinheimera phage vB_RspM_barba_12-6F unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497243 | Rheinheimera phage vB_RspM_barba_12-6E | Rheinheimera phage vB_RspM_barba_12-6E unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80347 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497242 | Rheinheimera phage vB_RspM_barba_12-6D | Rheinheimera phage vB_RspM_barba_12-6D unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497241 | Rheinheimera phage vB_RspM_barba_12-6C | Rheinheimera phage vB_RspM_barba_12-6C unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497240 | Rheinheimera phage vB_RspM_barba_12-6B | Rheinheimera phage vB_RspM_barba_12-6B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497239 | Rheinheimera phage vB_RspM_barba_12-6A | Rheinheimera phage vB_RspM_barba_12-6A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497238 | Rheinheimera phage vB_RspM_barba_11-8A | Rheinheimera phage vB_RspM_barba_11-8A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.248 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497237 | Rheinheimera phage vB_RspM_barba_10-9J | Rheinheimera phage vB_RspM_barba_10-9J unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80561 | 38.291 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497236 | Rheinheimera phage vB_RspM_barba_10-9I | Rheinheimera phage vB_RspM_barba_10-9I unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80562 | 38.295 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497235 | Rheinheimera phage vB_RspM_barba_10-9H | Rheinheimera phage vB_RspM_barba_10-9H unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497234 | Rheinheimera phage vB_RspM_barba_10-9G | Rheinheimera phage vB_RspM_barba_10-9G unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497233 | Rheinheimera phage vB_RspM_barba_10-9F | Rheinheimera phage vB_RspM_barba_10-9F unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497232 | Rheinheimera phage vB_RspM_barba_10-9E | Rheinheimera phage vB_RspM_barba_10-9E unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.294 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497231 | Rheinheimera phage vB_RspM_barba_10-9D | Rheinheimera phage vB_RspM_barba_10-9D unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497230 | Rheinheimera phage vB_RspM_barba_10-9C | Rheinheimera phage vB_RspM_barba_10-9C unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80280 | 38.295 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497229 | Rheinheimera phage vB_RspM_barba_10-9B | Rheinheimera phage vB_RspM_barba_10-9B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.295 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MW341595 | Shigella phage Sfk20 | Shigella phage Sfk20 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164878 | 35.615 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MW117144 | Pseudomonas phage AIIMS-Pa-A1 | Pseudomonas phage AIIMS-Pa-A1 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40977 | 62.293 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MW250641 | Vibrio phage vB_VpP_DE17 | Vibrio phage vB_VpP_DE17 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43394 | 49.235 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MT740730 | Ralstonia phage Cimandef | Ralstonia phage Cimandef Ralstonia virus Cimandef Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55171 | 64.822 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| CP063968 | Clostridium phage CWou-2020b | Clostridium phage CWou-2020b unclassified bacterial viruses Viruses | 52215 | 30.823 | Unclassified | Unclassified | Unclassified | Clostridium |
| CP063964 | Clostridium phage CWou-2020a | Clostridium phage CWou-2020a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35258 | 27.971 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MW353181 | Mycobacterium phage phiT46-1 | Mycobacterium phage phiT46-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52849 | 63.708 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314859 | Mycobacterium phage phiGD17-1 | Mycobacterium phage phiGD17-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51185 | 63.347 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314858 | Mycobacterium phage phiGD20-1 | Mycobacterium phage phiGD20-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59055 | 59.365 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314857 | Mycobacterium phage phiGD21-1 | Mycobacterium phage phiGD21-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40735 | 63.565 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314856 | Mycobacterium phage phiGD22-1 | Mycobacterium phage phiGD22-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60512 | 59.383 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314855 | Mycobacterium phage phiGD23-1 | Mycobacterium phage phiGD23-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60535 | 59.387 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314854 | Mycobacterium phage phiGD24-3 | Mycobacterium phage phiGD24-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54877 | 57.74 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314853 | Mycobacterium phage phiGD34-2 | Mycobacterium phage phiGD34-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40005 | 63.75 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314852 | Mycobacterium phage phiGD57-1 | Mycobacterium phage phiGD57-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52563 | 63.798 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314851 | Mycobacterium phage phiGD89-1 | Mycobacterium phage phiGD89-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40450 | 63.392 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MW314848 | Arthrobacter phage Kittykat | Arthrobacter phage Kittykat unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43603 | 60.686 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| MW314846 | Mycobacterium phage KashFlow | Mycobacterium phage KashFlow unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111628 | 61.021 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MW238767 | Staphylococcus phage IME1364_02 | Staphylococcus phage IME1364_02 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42338 | 35.224 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MW175414 | Escherichia virus VEc25 | Escherichia virus VEc25 unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71429 | 41.287 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MW291016 | Gordonia phage Blino | Gordonia phage Blino Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54168 | 66.903 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT497279 | Rheinheimera phage vB_RspM_barba_5-7A | Rheinheimera phage vB_RspM_barba_5-7A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.248 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497278 | Rheinheimera phage vB_RspM_barba_13-8A | Rheinheimera phage vB_RspM_barba_13-8A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81257 | 38.292 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497277 | Rheinheimera phage vB_RspM_barba_13-6A | Rheinheimera phage vB_RspM_barba_13-6A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497276 | Rheinheimera phage vB_RspM_barba_13-4A | Rheinheimera phage vB_RspM_barba_13-4A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.288 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497275 | Rheinheimera phage vB_RspM_barba_12-9J | Rheinheimera phage vB_RspM_barba_12-9J unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80447 | 38.251 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497274 | Rheinheimera phage vB_RspM_barba_12-9I | Rheinheimera phage vB_RspM_barba_12-9I unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80655 | 38.288 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497228 | Rheinheimera phage vB_RspM_barba_10-9A | Rheinheimera phage vB_RspM_barba_10-9A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80556 | 38.293 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497227 | Rheinheimera phage vB_RspM_barba_10-8D | Rheinheimera phage vB_RspM_barba_10-8D unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497226 | Rheinheimera phage vB_RspM_barba_10-8B | Rheinheimera phage vB_RspM_barba_10-8B unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80168 | 38.162 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497225 | Rheinheimera phage vB_RspM_barba_10-8A | Rheinheimera phage vB_RspM_barba_10-8A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80348 | 38.224 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497224 | Rheinheimera phage vB_RspM_barba_1-3A | Rheinheimera phage vB_RspM_barba_1-3A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80441 | 38.248 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MT497125 | Flavobacterium phage vB_FspP_elemoA_9-1A | Flavobacterium phage vB_FspP_elemoA_9-1A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497124 | Flavobacterium phage vB_FspP_elemoA_8-9B | Flavobacterium phage vB_FspP_elemoA_8-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497123 | Flavobacterium phage vB_FspP_elemoA_8-9A | Flavobacterium phage vB_FspP_elemoA_8-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60873 | 31.643 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497122 | Flavobacterium phage vB_FspP_elemoA_8-3A | Flavobacterium phage vB_FspP_elemoA_8-3A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497121 | Flavobacterium phage vB_FspP_elemoA_8-1D | Flavobacterium phage vB_FspP_elemoA_8-1D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497120 | Flavobacterium phage vB_FspP_elemoA_8-1B | Flavobacterium phage vB_FspP_elemoA_8-1B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497119 | Flavobacterium phage vB_FspP_elemoA_8-1A | Flavobacterium phage vB_FspP_elemoA_8-1A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497118 | Flavobacterium phage vB_FspP_elemoA_7-9C | Flavobacterium phage vB_FspP_elemoA_7-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60104 | 31.75 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497117 | Flavobacterium phage vB_FspP_elemoA_7-9B | Flavobacterium phage vB_FspP_elemoA_7-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497116 | Flavobacterium phage vB_FspP_elemoA_7-5A | Flavobacterium phage vB_FspP_elemoA_7-5A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60873 | 31.651 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497115 | Flavobacterium phage vB_FspP_elemoA_7-3B | Flavobacterium phage vB_FspP_elemoA_7-3B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.766 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497114 | Flavobacterium phage vB_FspP_elemoA_7-3A | Flavobacterium phage vB_FspP_elemoA_7-3A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60064 | 31.761 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497113 | Flavobacterium phage vB_FspP_elemoA_6-9A | Flavobacterium phage vB_FspP_elemoA_6-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.766 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497112 | Flavobacterium phage vB_FspP_elemoA_6-5A | Flavobacterium phage vB_FspP_elemoA_6-5A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60086 | 31.791 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497111 | Flavobacterium phage vB_FspP_elemoA_5-9B | Flavobacterium phage vB_FspP_elemoA_5-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60085 | 31.782 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497110 | Flavobacterium phage vB_FspP_elemoA_5-9A | Flavobacterium phage vB_FspP_elemoA_5-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497109 | Flavobacterium phage vB_FspP_elemoA_5-3D | Flavobacterium phage vB_FspP_elemoA_5-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497108 | Flavobacterium phage vB_FspP_elemoA_4-9A | Flavobacterium phage vB_FspP_elemoA_4-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497107 | Flavobacterium phage vB_FspP_elemoC_3-9B | Flavobacterium phage vB_FspP_elemoC_3-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60198 | 31.764 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497106 | Flavobacterium phage vB_FspP_elemoA_3-9A | Flavobacterium phage vB_FspP_elemoA_3-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497105 | Flavobacterium phage vB_FspP_elemoA_3-5A | Flavobacterium phage vB_FspP_elemoA_3-5A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60466 | 31.786 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497104 | Flavobacterium phage vB_FspP_elemoA_2-9A | Flavobacterium phage vB_FspP_elemoA_2-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497103 | Flavobacterium phage vB_FspP_elemoA_2-5G | Flavobacterium phage vB_FspP_elemoA_2-5G crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497102 | Flavobacterium phage vB_FspP_elemoA_2-5F | Flavobacterium phage vB_FspP_elemoA_2-5F crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.766 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497101 | Flavobacterium phage vB_FspP_elemoA_2-5B | Flavobacterium phage vB_FspP_elemoA_2-5B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.77 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497100 | Flavobacterium phage vB_FspP_elemoA_2-5A | Flavobacterium phage vB_FspP_elemoA_2-5A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497099 | Flavobacterium phage vB_FspP_elemoA_15-9C | Flavobacterium phage vB_FspP_elemoA_15-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497098 | Flavobacterium phage vB_FspP_elemoC_15-9A | Flavobacterium phage vB_FspP_elemoC_15-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60162 | 31.749 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497097 | Flavobacterium phage vB_FspP_elemoA_15-3D | Flavobacterium phage vB_FspP_elemoA_15-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61110 | 31.797 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497096 | Flavobacterium phage vB_FspP_elemoA_15-3B | Flavobacterium phage vB_FspP_elemoA_15-3B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497095 | Flavobacterium phage vB_FspP_elemoA_14-9B | Flavobacterium phage vB_FspP_elemoA_14-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497094 | Flavobacterium phage vB_FspP_elemoA_14-9A | Flavobacterium phage vB_FspP_elemoA_14-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497093 | Flavobacterium phage vB_FspP_elemoA_14-3D | Flavobacterium phage vB_FspP_elemoA_14-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.77 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497092 | Flavobacterium phage vB_FspP_elemoA_13-9B | Flavobacterium phage vB_FspP_elemoA_13-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497091 | Flavobacterium phage vB_FspP_elemoD_13-5B | Flavobacterium phage vB_FspP_elemoD_13-5B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60393 | 31.651 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497090 | Flavobacterium phage vB_FspP_elemoA_13-3D | Flavobacterium phage vB_FspP_elemoA_13-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497089 | Flavobacterium phage vB_FspP_elemoA_13-3B | Flavobacterium phage vB_FspP_elemoA_13-3B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497088 | Flavobacterium phage vB_FspP_elemoA_13-1E | Flavobacterium phage vB_FspP_elemoA_13-1E crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497087 | Flavobacterium phage vB_FspP_elemoA_13-1B | Flavobacterium phage vB_FspP_elemoA_13-1B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60071 | 31.759 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497086 | Flavobacterium phage vB_FspP_elemoA_12-9A | Flavobacterium phage vB_FspP_elemoA_12-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.77 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497085 | Flavobacterium phage vB_FspP_elemoA_12-3A | Flavobacterium phage vB_FspP_elemoA_12-3A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497084 | Flavobacterium phage vB_FspP_elemoA_12-1B | Flavobacterium phage vB_FspP_elemoA_12-1B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497083 | Flavobacterium phage vB_FspP_elemoA_11-9C | Flavobacterium phage vB_FspP_elemoA_11-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60377 | 31.805 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497082 | Flavobacterium phage vB_FspP_elemoA_11-9B | Flavobacterium phage vB_FspP_elemoA_11-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497081 | Flavobacterium phage vB_FspP_elemoA_11-5C | Flavobacterium phage vB_FspP_elemoA_11-5C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60093 | 31.781 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497080 | Flavobacterium phage vB_FspP_elemoA_10-9C | Flavobacterium phage vB_FspP_elemoA_10-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60768 | 31.706 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497079 | Flavobacterium phage vB_FspP_elemoA_10-9A | Flavobacterium phage vB_FspP_elemoA_10-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497078 | Flavobacterium phage vB_FspP_elemoE_10-3D | Flavobacterium phage vB_FspP_elemoE_10-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59743 | 31.932 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497077 | Flavobacterium phage vB_FspP_elemoA_1-9C | Flavobacterium phage vB_FspP_elemoA_1-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.766 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497076 | Flavobacterium phage vB_FspP_elemoA_1-9B | Flavobacterium phage vB_FspP_elemoA_1-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497075 | Flavobacterium phage vB_FspP_elemoA_1-9A | Flavobacterium phage vB_FspP_elemoA_1-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60061 | 31.776 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497074 | Flavobacterium phage vB_FspP_elemoA_1-5B | Flavobacterium phage vB_FspP_elemoA_1-5B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.77 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497073 | Flavobacterium phage vB_FspP_elemoA_8-9C | Flavobacterium phage vB_FspP_elemoA_8-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60950 | 31.708 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497072 | Flavobacterium phage vB_FspP_elemoE_6-9C | Flavobacterium phage vB_FspP_elemoE_6-9C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59979 | 31.953 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497071 | Flavobacterium phage vB_FspP_elemoF_6-3D | Flavobacterium phage vB_FspP_elemoF_6-3D crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59846 | 31.899 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497070 | Flavobacterium phage vB_FspP_elemoA_2-5C | Flavobacterium phage vB_FspP_elemoA_2-5C crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60601 | 31.764 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497069 | Flavobacterium phage vB_FspP_elemoA_15-9B | Flavobacterium phage vB_FspP_elemoA_15-9B crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60073 | 31.768 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497068 | Flavobacterium phage vB_FspP_elemoB_14-3B | Flavobacterium phage vB_FspP_elemoB_14-3B Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60925 | 31.636 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497067 | Flavobacterium phage vB_FspP_elemoC_14-1A | Flavobacterium phage vB_FspP_elemoC_14-1A Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61048 | 31.616 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497066 | Flavobacterium phage vB_FspP_elemoC_13-1C | Flavobacterium phage vB_FspP_elemoC_13-1C Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60458 | 31.728 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497065 | Flavobacterium phage vB_FspP_elemoA_13-1A | Flavobacterium phage vB_FspP_elemoA_13-1A Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61110 | 31.793 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT497017 | Flavobacterium phage vB_FspP_elemoA_7-9A | Flavobacterium phage vB_FspP_elemoA_7-9A crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61691 | 31.64 | Unclassified | Unclassified | Podoviridae | Flavobacterium |
| MT225101 | Escherichia phage Lys19259Vzw | Escherichia phage Lys19259Vzw unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61072 | 49.203 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| MH973725 | Pseudomonas phage SaPL | Pseudomonas phage SaPL unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45796 | 52.564 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MH979674 | Pseudomonas phage RLP | Pseudomonas phage RLP Pseudomonas virus RLP Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43446 | 62.093 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MK170160 | Acinetobacter phage TAC1 | Acinetobacter phage TAC1 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101770 | 37.493 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MW057861 | Providencia phage PSTCR7 | Providencia phage PSTCR7 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57986 | 36.88 | Unclassified | Queuovirinae | Siphoviridae | Providencia |
| MT680615 | Proteus phage PmP19 | Proteus phage PmP19 unclassified Molineuxvirinae Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44305 | 51.931 | Unclassified | Molineuxvirinae | Autographiviridae | Proteus |
| MW006482 | Salmonella phage GEC_vB_NS7 | Salmonella phage GEC_vB_NS7 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86886 | 39.03 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW006480 | Salmonella phage GEC_vB_N6 | Salmonella phage GEC_vB_N6 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158116 | 44.909 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW006479 | Salmonella phage GEC_vB_N5 | Salmonella phage GEC_vB_N5 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110015 | 39.251 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW006478 | Salmonella phage GEC_vB_N3 | Salmonella phage GEC_vB_N3 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109645 | 40.085 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW006477 | Salmonella phage GEC_vB_MG | Salmonella phage GEC_vB_MG unclassified Seunavirus Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141829 | 46.187 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| MW006476 | Salmonella phage GEC_vB_GOT | Salmonella phage GEC_vB_GOT unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157200 | 44.894 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW006475 | Salmonella phage GEC_vB_BS | Salmonella phage GEC_vB_BS unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85840 | 39.014 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW006474 | Salmonella phage GEC_vB_B1 | Salmonella phage GEC_vB_B1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87369 | 39.064 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MW082584 | Serratia phage 4S | Serratia phage 4S unclassified Muldoonvirus Muldoonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173061 | 39.87 | Muldoonvirus | Unclassified | Myoviridae | Serratia |
| MW082583 | Serratia phage Tsm2 | Serratia phage Tsm2 unclassified Guernseyvirinae Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39799 | 56.979 | Unclassified | Guernseyvirinae | Siphoviridae | Serratia |
| MW149275 | Salmonella phage vB_SalS_ABTNLsp9 | Salmonella phage vB_SalS_ABTNLsp9 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109695 | 39.248 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW149274 | Salmonella phage vB_SalM_ABTNLsp5 | Salmonella phage vB_SalM_ABTNLsp5 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172418 | 39.67 | Dhakavirus | Tevenvirinae | Myoviridae | Salmonella |
| MW149273 | Salmonella phage vB_SalS_ABTNLsp4 | Salmonella phage vB_SalS_ABTNLsp4 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106178 | 39.115 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW149272 | Salmonella phage vB_SalS_ABTNLsp1 | Salmonella phage vB_SalS_ABTNLsp1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58922 | 56.324 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MN508615 | Escherichia phage ES17 | Escherichia phage ES17 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75007 | 42.101 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MW057862 | Providencia phage PSTCR7lys | Providencia phage PSTCR7lys Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22938 | 41.695 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MT708544 | Rhizobium phage Palo | Rhizobium phage Palo unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46322 | 51.725 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MT366761 | Vibrio phage vB_VcorM_GR11A | Vibrio phage vB_VcorM_GR11A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194800 | 44.867 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MT366760 | Vibrio phage vB_VcorM_GR7B | Vibrio phage vB_VcorM_GR7B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 207758 | 42.654 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MT413450 | Pseudomonas phage Epa15 | Pseudomonas phage Epa15 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66002 | 55.63 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN812691 | Pectobacterium phage Horatius | Pectobacterium phage Horatius unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73737 | 48.47 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MT986025 | Gordonia phage GrootJr | Gordonia phage GrootJr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67176 | 65.861 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT889393 | Gordonia phage NovumRegina | Gordonia phage NovumRegina Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67175 | 65.862 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT366762 | Vibrio phage vB_VcorM_GR28A | Vibrio phage vB_VcorM_GR28A unclassified Campanilevirus Campanilevirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154046 | 47.671 | Campanilevirus | Unclassified | Ackermannviridae | Vibrio |
| KY742649 | Proteus phage VB_PmiS-Isfahan | Proteus phage VB_PmiS-Isfahan Proteus virus Isfahan Gorganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54836 | 36.093 | Gorganvirus | Unclassified | Siphoviridae | Proteus |
| NC_051726 | Mycobacterium phage Mendokysei | Mycobacterium phage Mendokysei unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43511 | 66.358 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| MW206381 | Salmonella phage SP76 | Salmonella phage SP76 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111639 | 39.91 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MW006481 | Salmonella phage GEC_vB_N7 | Salmonella phage GEC_vB_N7 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111684 | 39.926 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| NC_047883 | Acinetobacter phage SWH-Ab-3 | Acinetobacter phage SWH-Ab-3 Acinetobacter virus SWHAb3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41730 | 39.382 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT899504 | Staphylococcus phage JD419 | Staphylococcus phage JD419 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45509 | 33.536 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MT682391 | Aeromonas phage Atoyac23 | Aeromonas phage Atoyac23 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41506 | 59.196 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| MT682390 | Aeromonas phage Atoyac15 | Aeromonas phage Atoyac15 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42010 | 58.912 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| MT682389 | Aeromonas phage Atoyac14 | Aeromonas phage Atoyac14 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41707 | 59.302 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| MT682388 | Aeromonas phage Atoyac13 | Aeromonas phage Atoyac13 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41639 | 59.267 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| MT682387 | Aeromonas phage Atoyac10 | Aeromonas phage Atoyac10 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41676 | 59.231 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| MT682386 | Aeromonas phage Atoyac1 | Aeromonas phage Atoyac1 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41721 | 59.294 | Unclassified | Melnykvirinae | Autographiviridae | Aeromonas |
| NC_048167 | Vibrio phage vB_VpaS_OWB | Vibrio phage vB_VpaS_OWB Vibrio virus OWB Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43264 | 48.74 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MH172264 | Klebsiella phage KP32_isolate 196 | Klebsiella phage KP32_isolate 196 Klebsiella virus KP32i196 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40337 | 53.023 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MH172263 | Klebsiella phage KP32_isolate 195 | Klebsiella phage KP32_isolate 195 Klebsiella virus KP32i195 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40540 | 52.753 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MH172262 | Klebsiella phage KP32_isolate 194 | Klebsiella phage KP32_isolate 194 Klebsiella virus KP32i194 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41161 | 52.836 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MH172261 | Klebsiella phage KP32_isolate 192 | Klebsiella phage KP32_isolate 192 Klebsiella virus KP32i192 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40635 | 53.203 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX349323 | Cyanophage S-RIM12 isolate W1_08_0910 | Cyanophage S-RIM12 isolate W1_08_0910 Synechococcus virus SRIM12-08 Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175559 | 39.519 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349313 | Cyanophage S-RIM12 isolate RW_06_0310 | Cyanophage S-RIM12 isolate RW_06_0310 Synechococcus virus SRIM12-06 Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175557 | 39.526 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349310 | Cyanophage S-RIM12 isolate RW_01_0310 | Cyanophage S-RIM12 isolate RW_01_0310 Synechococcus virus SRIM12-01 Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174670 | 39.513 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KJ019091 | Synechococcus phage ACG-2014f_Syn7803US26 | Synechococcus phage ACG-2014f_Syn7803US26 Synechococcus virus ACG2014fSyn7803US26 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 196842 | 41.536 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019058 | Synechococcus phage ACG-2014f_Syn7803C8 | Synechococcus phage ACG-2014f_Syn7803C8 Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223270 | 41.607 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019052 | Synechococcus phage ACG-2014f_Syn7803C7 | Synechococcus phage ACG-2014f_Syn7803C7 Synechococcus virus ACG2014f Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216628 | 41.575 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| MW057860 | Providencia phage PSTCR9 | Providencia phage PSTCR9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58873 | 49.474 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW057859 | Providencia phage PSTCR8lys | Providencia phage PSTCR8lys Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40280 | 41.889 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW057858 | Providencia phage PSTCR6 | Providencia phage PSTCR6 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155737 | 35.914 | Unclassified | Tevenvirinae | Myoviridae | Providencia |
| MW057857 | Providencia phage PSTCR5 | Providencia phage PSTCR5 Priunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109434 | 39.999 | Priunavirus | Unclassified | Siphoviridae | Providencia |
| MW057856 | Providencia phage PSTCR4 | Providencia phage PSTCR4 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57214 | 36.886 | Unclassified | Queuovirinae | Siphoviridae | Providencia |
| MW057855 | Providencia phage PSTCR3 | Providencia phage PSTCR3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39447 | 41.879 | Unclassified | Unclassified | Myoviridae | Providencia |
| MW057854 | Providencia phage PSTCR2 | Providencia phage PSTCR2 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40200 | 42.271 | Unclassified | Studiervirinae | Autographiviridae | Providencia |
| MW161468 | Cutibacterium phage PAVL45 | Cutibacterium phage PAVL45 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29772 | 54.283 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161467 | Cutibacterium phage PAVL34 | Cutibacterium phage PAVL34 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29535 | 54.288 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161466 | Cutibacterium phage PAVL33 | Cutibacterium phage PAVL33 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29627 | 54.363 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161465 | Cutibacterium phage PAVL21 | Cutibacterium phage PAVL21 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30034 | 54.302 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161464 | Cutibacterium phage PAVL20 | Cutibacterium phage PAVL20 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29800 | 54.279 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161463 | Cutibacterium phage FD3 | Cutibacterium phage FD3 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29638 | 54.187 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161462 | Cutibacterium phage FD2 | Cutibacterium phage FD2 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29768 | 54.256 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW161461 | Cutibacterium phage FD1 | Cutibacterium phage FD1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29774 | 54.265 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MW145139 | Providencia phage Kokobel1 | Providencia phage Kokobel1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59837 | 48.916 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW145138 | Providencia phage Kokobel2 | Providencia phage Kokobel2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45880 | 47.52 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW145137 | Providencia phage PSTNGR2lys | Providencia phage PSTNGR2lys Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50958 | 41.624 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MW145136 | Providencia phage PSTNGR1 | Providencia phage PSTNGR1 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41033 | 58.526 | Unclassified | Okabevirinae | Autographiviridae | Providencia |
| MT701598 | Streptomyces phage Sitrop | Streptomyces phage Sitrop unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48481 | 65.597 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT701597 | Streptomyces phage Sentinel | Streptomyces phage Sentinel unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50272 | 66.862 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT701596 | Streptomyces phage Shady | Streptomyces phage Shady Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45128 | 62.252 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT701595 | Streptomyces phage Shaeky | Streptomyces phage Shaeky Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45617 | 62.602 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT701594 | Streptomyces phage Spernnie | Streptomyces phage Spernnie unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50834 | 65.694 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT701593 | Streptomyces phage Sycamore | Streptomyces phage Sycamore Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44694 | 62.68 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT701592 | Klebsiella phage Solomon | Klebsiella phage Solomon unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51775 | 47.925 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MT701591 | Burkholderia phage Mana | Burkholderia phage Mana Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38038 | 64.315 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| MT701590 | Klebsiella phage Miami | Klebsiella phage Miami Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253383 | 43.947 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MT701589 | Klebsiella phage Pone | Klebsiella phage Pone unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44346 | 53.847 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT701588 | Klebsiella phage Metamorpho | Klebsiella phage Metamorpho unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171475 | 38.512 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MT701587 | Burkholderia phage Magia | Burkholderia phage Magia Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44942 | 65.062 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MT701586 | Burkholderia phage Mica | Burkholderia phage Mica Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43707 | 62.153 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MW073017 | Vibrio phage Va2 | Vibrio phage Va2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128360 | 40.714 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MW147367 | Synechococcus phage S-H9-2 | Synechococcus phage S-H9-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 187320 | 40.32 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MW117966 | Synechococcus phage S-H9-1 | Synechococcus phage S-H9-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192454 | 40.994 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MW117965 | Synechococcus phage S-H38 | Synechococcus phage S-H38 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180224 | 42.424 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MT941682 | Pseudomonas phage BUCT553 | Pseudomonas phage BUCT553 unclassified Uliginvirus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47148 | 56.605 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MT740315 | Escherichia phage JEP4 | Escherichia phage JEP4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39195 | 47.054 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MT740314 | Escherichia phage JEP1 | Escherichia phage JEP1 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143610 | 43.538 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT764208 | Escherichia phage JEP8 | Escherichia phage JEP8 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165295 | 40.475 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MT764207 | Escherichia phage JEP7 | Escherichia phage JEP7 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52936 | 45.94 | Rosemountvirus | Unclassified | Myoviridae | Escherichia |
| MT764206 | Escherichia phage JEP6 | Escherichia phage JEP6 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170340 | 35.311 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT710307 | Salmonella phage pSal-SNUABM-04 | Salmonella phage pSal-SNUABM-04 unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239626 | 51.209 | Machinavirus | Unclassified | Myoviridae | Salmonella |
| MT310882 | Mycobacterium phage JF1 | Mycobacterium phage JF1 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57990 | 67.884 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN945904 | Mycobacterium phage Eponine | Mycobacterium phage Eponine unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58678 | 67.419 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN945901 | Mycobacterium phage Ximenita | Mycobacterium phage Ximenita unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61027 | 66.741 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK290739 | Pectobacterium phage Q19 | Pectobacterium phage Q19 unclassified Pektosvirus Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40227 | 49.651 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MN892485 | Gordonia phage Kuwabara | Gordonia phage Kuwabara Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54102 | 63.273 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN813694 | Gordonia phage Opie | Gordonia phage Opie unclassified Schnabeltiervirus Schnabeltiervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47713 | 66.95 | Schnabeltiervirus | Unclassified | Siphoviridae | Gordonia |
| MN703410 | Mycobacterium phage Itos | Mycobacterium phage Itos unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74937 | 58.897 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN703406 | Mycobacterium phage Gabriela | Mycobacterium phage Gabriela unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75383 | 58.755 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586052 | Mycobacterium phage Kahlid | Mycobacterium phage Kahlid unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75952 | 58.885 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586030 | Mycobacterium phage BobsGarage | Mycobacterium phage BobsGarage unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68102 | 58.89 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586029 | Mycobacterium phage Hannaconda | Mycobacterium phage Hannaconda unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111740 | 60.955 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586024 | Mycobacterium phage TyDawg | Mycobacterium phage TyDawg unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81039 | 61.618 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MN369766 | Gordonia phage Yndexa | Gordonia phage Yndexa Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66184 | 66.193 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN369762 | Mycobacterium phage Bryler | Mycobacterium phage Bryler unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57666 | 66.098 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369761 | Mycobacterium phage Malthus | Mycobacterium phage Malthus unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57802 | 67.904 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369753 | Gordonia phage Sukkupi | Gordonia phage Sukkupi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66184 | 66.194 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN428063 | Gordonia phage NadineRae | Gordonia phage NadineRae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64714 | 66.057 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN428055 | Gordonia phage Mcklovin | Gordonia phage Mcklovin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53155 | 65.785 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN428048 | Gordonia phage DobbysSock | Gordonia phage DobbysSock Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48198 | 66.293 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN284895 | Mycobacterium phage Marshawn | Mycobacterium phage Marshawn unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61464 | 68.146 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234236 | Mycobacterium phage QueenHazel | Mycobacterium phage QueenHazel unclassified Brujitavirus Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48053 | 66.614 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| MN234199 | Mycobacterium phage Ekdilam | Mycobacterium phage Ekdilam unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 67.432 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234195 | Mycobacterium phage Xula | Mycobacterium phage Xula unclassified Brujitavirus Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48540 | 66.745 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| MN234176 | Mycobacterium phage Yuna | Mycobacterium phage Yuna unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62192 | 67.441 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062715 | Gordonia phage Marteena | Gordonia phage Marteena Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50531 | 66.583 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH479927 | Gordonia phage TimTam | Gordonia phage TimTam unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53431 | 66.553 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MH479912 | Gordonia phage Eviarto | Gordonia phage Eviarto unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53407 | 66.549 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MN096377 | Mycobacterium phage Finnry | Mycobacterium phage Finnry unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75632 | 59.294 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096374 | Microbacterium phage Roman | Microbacterium phage Roman Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64259 | 62.754 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN096370 | Gordonia phage Squiddly | Gordonia phage Squiddly Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57508 | 63.083 | Unclassified | Langleyhallvirinae | Siphoviridae | Gordonia |
| MN096363 | Gordonia phage Gibbles | Gordonia phage Gibbles unclassified Orchidvirus Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81488 | 46.977 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| MN096355 | Mycobacterium phage Purky | Mycobacterium phage Purky unclassified Pclasvirinae Pclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50513 | 66.385 | Unclassified | Pclasvirinae | Siphoviridae | Mycobacterium |
| MK937609 | Gordonia phage Sekhmet | Gordonia phage Sekhmet Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47735 | 66.079 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK937599 | Gordonia phage Dmitri | Gordonia phage Dmitri Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45736 | 66.807 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK937597 | Gordonia phage Spooky | Gordonia phage Spooky Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56875 | 63.33 | Unclassified | Langleyhallvirinae | Siphoviridae | Gordonia |
| MK919483 | Gordonia phage Suscepit | Gordonia phage Suscepit unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50338 | 66.661 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK919481 | Gordonia phage Newt | Gordonia phage Newt unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73414 | 59.026 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK919477 | Gordonia phage Luker | Gordonia phage Luker unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73453 | 58.99 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK977708 | Mycobacterium phage Fulbright | Mycobacterium phage Fulbright unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42396 | 66.275 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK977699 | Gordonia Phage Lollipop1437 | Gordonia Phage Lollipop1437 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67311 | 65.731 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK801729 | Gordonia phage Clark | Gordonia phage Clark Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47716 | 66.531 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK878903 | Gordonia phage Teal | Gordonia phage Teal unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73626 | 59.024 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK878901 | Gordonia phage HannahD | Gordonia phage HannahD unclassified Bowservirus Bowservirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46095 | 66.853 | Bowservirus | Unclassified | Siphoviridae | Gordonia |
| MK878900 | Mycobacterium phage MsGreen | Mycobacterium phage MsGreen unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75617 | 59.279 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK878898 | Gordonia phage Zameen | Gordonia phage Zameen unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50337 | 66.661 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK820642 | Gordonia phage Polly | Gordonia phage Polly unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49780 | 66.609 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK820641 | Gordonia phage EnalisNailo | Gordonia phage EnalisNailo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51094 | 67.112 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK875797 | Gordonia phage Haley23 | Gordonia phage Haley23 unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46221 | 66.621 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MK875796 | Gordonia phage Tayonia | Gordonia phage Tayonia unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50320 | 66.653 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK814755 | Gordonia phage Antonio | Gordonia phage Antonio unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50337 | 66.663 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MK814750 | Gordonia phage Ewald | Gordonia phage Ewald unclassified Nyceiraevirus Nyceiraevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41454 | 67.67 | Nyceiraevirus | Unclassified | Siphoviridae | Gordonia |
| MK757446 | Mycobacterium phage Candle | Mycobacterium phage Candle unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71390 | 55.98 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494121 | Gordonia phage Nimi13 | Gordonia phage Nimi13 unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73594 | 59.071 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK494109 | Gordonia phage Lahirium | Gordonia phage Lahirium unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73885 | 59.116 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK494106 | Gordonia phage Jormungandr | Gordonia phage Jormungandr unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73380 | 58.941 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK494098 | Gordonia phage Tredge | Gordonia phage Tredge unclassified Demosthenesvirus Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75345 | 59.62 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| MK494089 | Mycobacterium phage Ibrahim | Mycobacterium phage Ibrahim unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42596 | 66.131 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524528 | Mycobacterium phage ShrimpFriedEgg | Mycobacterium phage ShrimpFriedEgg unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42594 | 66.127 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524522 | Mycobacterium phage Purgamenstris | Mycobacterium phage Purgamenstris unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42595 | 66.139 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524520 | Mycobacterium phage Nenae | Mycobacterium phage Nenae unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42597 | 66.143 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524518 | Mycobacterium phage Smurph | Mycobacterium phage Smurph unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43700 | 66.27 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524515 | Mycobacterium phage Parmesanjohn | Mycobacterium phage Parmesanjohn unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43700 | 66.27 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524509 | Mycobacterium phage SpongeBob | Mycobacterium phage SpongeBob unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41278 | 66.244 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524505 | Gordonia phage Dogfish | Gordonia phage Dogfish unclassified Nyceiraevirus Nyceiraevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41907 | 67.461 | Nyceiraevirus | Unclassified | Siphoviridae | Gordonia |
| MK524488 | Mycobacterium phage Patt | Mycobacterium phage Patt unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54611 | 67.604 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524485 | Mycobacterium phage MissDaisy | Mycobacterium phage MissDaisy unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54464 | 67.582 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK433275 | Gordonia phage EMsquaredA | Gordonia phage EMsquaredA Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50531 | 66.583 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK433274 | Gordonia phage Bradissa | Gordonia phage Bradissa Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51954 | 67.169 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK433272 | Gordonia phage IDyn | Gordonia phage IDyn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65218 | 66.173 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK392367 | Mycobacterium phage BabeRuth | Mycobacterium phage BabeRuth unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42595 | 66.146 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK450436 | Gordonia phage Neoevie | Gordonia phage Neoevie unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73353 | 59.124 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK450435 | Gordonia phage Hello | Gordonia phage Hello unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73453 | 59.188 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK450430 | Gordonia phage Harambe | Gordonia phage Harambe unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73511 | 59.171 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK450428 | Gordonia phage Guillaume | Gordonia phage Guillaume unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73583 | 59.092 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MK376962 | Gordonia phage NosilaM | Gordonia phage NosilaM Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67662 | 65.579 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK376960 | Gordonia phage BiPauneto | Gordonia phage BiPauneto Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66935 | 66.222 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK376959 | Gordonia phage Aphelion | Gordonia phage Aphelion unclassified Smoothievirus Smoothievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93276 | 61.93 | Smoothievirus | Unclassified | Siphoviridae | Gordonia |
| MK376954 | Gordonia phage WhoseManz | Gordonia phage WhoseManz Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64214 | 66.051 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK284520 | Gordonia phage Lilas | Gordonia phage Lilas Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52632 | 66.914 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK279848 | Gordonia phage Dorito | Gordonia phage Dorito Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46784 | 66.514 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK202161 | Streptococcus phage CHPC1282 | Streptococcus phage CHPC1282 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34785 | 38.051 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK202160 | Streptococcus phage CHPC1232 | Streptococcus phage CHPC1232 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31974 | 38.281 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK202159 | Streptococcus phage CHPC1198 | Streptococcus phage CHPC1198 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33245 | 38.069 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK061410 | Gordonia phage Eyes | Gordonia phage Eyes Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46772 | 67.094 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK016492 | Gordonia phage Bialota | Gordonia phage Bialota unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52089 | 66.667 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MH926059 | Mycobacterium phage Riparian | Mycobacterium phage Riparian unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71199 | 55.976 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH926058 | Mycobacterium phage Reptar3000 | Mycobacterium phage Reptar3000 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54601 | 67.579 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH926055 | Mycobacterium phage Chewbacca | Mycobacterium phage Chewbacca unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43575 | 66.194 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH834601 | Mycobacterium phage BoostSeason | Mycobacterium phage BoostSeason unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58078 | 68.213 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH834600 | Mycobacterium phage BigCheese | Mycobacterium phage BigCheese unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72810 | 58.905 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779510 | Gordonia phage Kurt | Gordonia phage Kurt Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68205 | 65.803 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH779508 | Gordonia phage Jifall16 | Gordonia phage Jifall16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67470 | 65.804 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH779501 | Gordonia phage Emianna | Gordonia phage Emianna Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68190 | 65.801 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH779499 | Gordonia phage Buggaboo | Gordonia phage Buggaboo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68626 | 65.627 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH744425 | Mycobacterium phage Wamburgrxpress | Mycobacterium phage Wamburgrxpress unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74392 | 58.771 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH744424 | Gordonia phage Turuncu | Gordonia phage Turuncu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66808 | 65.383 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH727559 | Mycobacterium phage Samty | Mycobacterium phage Samty unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75307 | 59.272 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727552 | Gordonia phage KidneyBean | Gordonia phage KidneyBean Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67802 | 65.824 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH727547 | Gordonia phage Foxboro | Gordonia phage Foxboro Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67773 | 65.842 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH727545 | Mycobacterium phage DismalStressor | Mycobacterium phage DismalStressor unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58129 | 68.253 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651185 | Gordonia phage Phistory | Gordonia phage Phistory Gordonia virus Phistory Phistoryvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53085 | 62.978 | Phistoryvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MH651176 | Gordonia phage Horus | Gordonia phage Horus Gordonia virus Horus Horusvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55668 | 62.957 | Horusvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MH651171 | Mycobacterium phage Collard | Mycobacterium phage Collard unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61395 | 65.556 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669013 | Gordonia phage Skysand | Gordonia phage Skysand Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67359 | 65.527 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH669007 | Gordonia phage Marietta | Gordonia phage Marietta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64370 | 66.13 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH697593 | Mycobacterium phage Tapioca | Mycobacterium phage Tapioca unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44205 | 66.137 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH697592 | Mycobacterium phage Rando14 | Mycobacterium phage Rando14 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59925 | 64.295 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697587 | Mycobacterium phage IPhane7 | Mycobacterium phage IPhane7 unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81036 | 61.616 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MH697585 | Mycobacterium phage Gex | Mycobacterium phage Gex unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43697 | 66.268 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH697576 | Mycobacterium phage Aggie | Mycobacterium phage Aggie unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44333 | 66.537 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH576976 | Gordonia phage Teatealatte | Gordonia phage Teatealatte unclassified Demosthenesvirus Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75345 | 59.618 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| MH576973 | Mycobacterium phage Bromden | Mycobacterium phage Bromden unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70183 | 58.228 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576966 | Mycobacterium phage Thyatira | Mycobacterium phage Thyatira unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63874 | 64.574 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576960 | Gordonia phage Pleakley | Gordonia phage Pleakley Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64105 | 65.218 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH513979 | Gordonia phage Pollux | Gordonia phage Pollux Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52488 | 66.791 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH536824 | Gordonia phage NatB6 | Gordonia phage NatB6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67081 | 65.749 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH536819 | Gordonia phage Fury | Gordonia phage Fury Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64104 | 65.219 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH536812 | Gordonia phage Apricot | Gordonia phage Apricot Gordonia virus Apricot Apricotvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52195 | 63.06 | Apricotvirus | Unclassified | Siphoviridae | Gordonia |
| MH479923 | Gordonia phage RobinSparkles | Gordonia phage RobinSparkles unclassified Orchidvirus Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81834 | 46.881 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| MH271318 | Gordonia phage Trine | Gordonia phage Trine Gordonia virus Trine Trinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43758 | 67.816 | Trinevirus | Unclassified | Siphoviridae | Gordonia |
| MH271305 | Gordonia phage Morrissey | Gordonia phage Morrissey unclassified Gustavvirus Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44924 | 65.845 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| MH155867 | Gordonia phage Easley | Gordonia phage Easley Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46717 | 66.614 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| JN699628 | Mycobacterium phage Bongo | Mycobacterium phage Bongo Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80228 | 61.621 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| MH051258 | Mycobacterium phage SamScheppers | Mycobacterium phage SamScheppers unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58351 | 67.627 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051255 | Mycobacterium phage Leston | Mycobacterium phage Leston unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61808 | 64.95 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH025889 | Gordonia phage Hail2Pitt | Gordonia phage Hail2Pitt unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73659 | 59.116 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MG962374 | Mycobacterium phage Paola | Mycobacterium phage Paola unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61535 | 65.01 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001447 | Mycobacterium phage Nilo | Mycobacterium phage Nilo unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71752 | 55.976 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG872834 | Gordonia phage Confidence | Gordonia phage Confidence Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50646 | 66.357 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MG757166 | Gordonia phage SuperSulley | Gordonia phage SuperSulley Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64124 | 65.442 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MG757157 | Gordonia phage Flapper | Gordonia phage Flapper Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67527 | 65.301 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KT361920 | Mycobacterium phage Slarp | Mycobacterium phage Slarp unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57256 | 67.965 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM400683 | Mycobacterium phage Ariel | Mycobacterium phage Ariel unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109801 | 61.018 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MG845393 | Gordonia phage Beenie | Gordonia phage Beenie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47953 | 66.265 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MG770214 | Gordonia phage SteveFrench | Gordonia phage SteveFrench Gordonia virus Stevefrench Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75687 | 59.182 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MG770211 | Gordonia phage Flakey | Gordonia phage Flakey Gordonia virus Flakey Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76889 | 58.924 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MG099940 | Gordonia phage BirksAndSocks | Gordonia phage BirksAndSocks Gordonia virus Birksandsocks Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77354 | 58.881 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MG099935 | Gordonia phage Anamika | Gordonia phage Anamika unclassified Woesvirus Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73851 | 59.154 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| MG099948 | Mycobacterium phage Philonius | Mycobacterium phage Philonius unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43886 | 66.541 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MG198784 | Gordonia phage Gustav | Gordonia phage Gustav Gordonia virus Gustav Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44408 | 67.855 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| MG198783 | Gordonia phage Mahdia | Gordonia phage Mahdia Gordonia virus Mahdia Gustavvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42475 | 67.249 | Gustavvirus | Unclassified | Siphoviridae | Gordonia |
| MF919542 | Gordonia phage Patio | Gordonia phage Patio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66251 | 65.557 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MF919512 | Mycobacterium phage Klein | Mycobacterium phage Klein unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112329 | 60.757 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919510 | Gordonia phage Kabluna | Gordonia phage Kabluna Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65869 | 65.556 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MF185728 | Mycobacterium phage Finemlucis | Mycobacterium phage Finemlucis unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77031 | 58.859 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668284 | Mycobacterium phage Squint | Mycobacterium phage Squint unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110240 | 60.953 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140406 | Mycobacterium phage DarthP | Mycobacterium phage DarthP unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61594 | 67.244 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140402 | Mycobacterium phage Chancellor | Mycobacterium phage Chancellor unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57697 | 67.972 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140435 | Mycobacterium phage Wintermute | Mycobacterium phage Wintermute unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58046 | 68.015 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140421 | Mycobacterium phage Nicholas | Mycobacterium phage Nicholas unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70413 | 59.184 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140414 | Mycobacterium phage Krypton555 | Mycobacterium phage Krypton555 unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76075 | 59.268 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140413 | Mycobacterium phage Kingsolomon | Mycobacterium phage Kingsolomon unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69537 | 59.127 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140411 | Mycobacterium phage Findley | Mycobacterium phage Findley unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58150 | 68.339 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140408 | Mycobacterium phage DismalFunk | Mycobacterium phage DismalFunk unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58129 | 68.255 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140398 | Mycobacterium phage Amohnition | Mycobacterium phage Amohnition unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61761 | 67.246 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185720 | Mycobacterium phage Guillsminger | Mycobacterium phage Guillsminger unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63153 | 65.039 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185719 | Mycobacterium phage Edugator | Mycobacterium phage Edugator unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63344 | 65.326 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324915 | Mycobacterium phage Amgine | Mycobacterium phage Amgine unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62236 | 66.447 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324914 | Mycobacterium phage Krueger | Mycobacterium phage Krueger unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60321 | 66.541 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324913 | Mycobacterium phage Cain | Mycobacterium phage Cain unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60813 | 66.27 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324912 | Mycobacterium phage Phrank | Mycobacterium phage Phrank unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61109 | 66.246 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324911 | Mycobacterium phage SirPhilip | Mycobacterium phage SirPhilip unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61882 | 66.701 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324909 | Mycobacterium phage PhelpsODU | Mycobacterium phage PhelpsODU unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56580 | 66.099 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324908 | Mycobacterium phage Unicorn | Mycobacterium phage Unicorn unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61208 | 66.184 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF373841 | Mycobacterium phage Hurricane | Mycobacterium phage Hurricane unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61318 | 67.14 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140405 | Mycobacterium phage Clautastrophe | Mycobacterium phage Clautastrophe unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72055 | 59.191 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185717 | Mycobacterium phage AlleyCat | Mycobacterium phage AlleyCat unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62112 | 65.35 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF133445 | Mycobacterium phage Lucky2013 | Mycobacterium phage Lucky2013 unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108627 | 61.146 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| MF072690 | Mycobacterium phage Porcelain | Mycobacterium phage Porcelain unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109575 | 61.183 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| KY969628 | Mycobacterium phage Zenon | Mycobacterium phage Zenon unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71726 | 55.988 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY783914 | Mycobacterium phage GardenSalsa | Mycobacterium phage GardenSalsa unclassified Reyvirus Reyvirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80309 | 60.923 | Reyvirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| KY087992 | Mycobacterium phage Mitti | Mycobacterium phage Mitti unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57895 | 67.977 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY087993 | Mycobacterium phage Hammy | Mycobacterium phage Hammy unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61812 | 67.23 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX688047 | Mycobacterium phage Marcoliusprime | Mycobacterium phage Marcoliusprime unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58129 | 68.25 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX683428 | Mycobacterium phage TBond007 | Mycobacterium phage TBond007 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61145 | 67.32 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX621007 | Mycobacterium phage Taquito | Mycobacterium phage Taquito unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58390 | 67.467 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX557282 | Gordonia phage Nyceirae | Gordonia phage Nyceirae Gordonia virus Nyceirae Nyceiraevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41857 | 67.513 | Nyceiraevirus | Unclassified | Siphoviridae | Gordonia |
| KX636165 | Mycobacterium phage Gengar | Mycobacterium phage Gengar unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61626 | 64.987 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX557287 | Gordonia phage Zirinka | Gordonia phage Zirinka Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52077 | 66.667 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KX557273 | Gordonia phage BatStarr | Gordonia phage BatStarr unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53432 | 66.554 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KU998254 | Gordonia phage Kampe | Gordonia phage Kampe Gordonia virus Orchid Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80649 | 47.006 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| KU998253 | Gordonia phage Orchid | Gordonia phage Orchid Gordonia virus Orchid Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80650 | 47.007 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| KU998252 | Gordonia phage PatrickStar | Gordonia phage PatrickStar Gordonia virus Orchid Orchidvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80729 | 46.994 | Orchidvirus | Unclassified | Siphoviridae | Gordonia |
| KU998243 | Gordonia phage Kvothe | Gordonia phage Kvothe Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75462 | 59.48 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| KU998242 | Gordonia phage Demosthenes | Gordonia phage Demosthenes Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74073 | 59.255 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| KU998241 | Gordonia phage Monty | Gordonia phage Monty Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75680 | 58.948 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| KU998240 | Gordonia phage Woes | Gordonia phage Woes Woesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73752 | 59.125 | Woesvirus | Unclassified | Siphoviridae | Gordonia |
| KU963262 | Gordonia phage Benczkowski14 | Gordonia phage Benczkowski14 Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75380 | 59.463 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| KU963258 | Gordonia phage Katyusha | Gordonia phage Katyusha Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75380 | 59.461 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| KU963257 | Gordonia phage Kita | Gordonia phage Kita Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 66.665 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KU963255 | Gordonia phage Nymphadora | Gordonia phage Nymphadora Gordonia virus Nymphadora Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53431 | 66.555 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| KU935731 | Mycobacterium phage Phrann | Mycobacterium phage Phrann Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44872 | 66.28 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KU935728 | Mycobacterium phage Xeno | Mycobacterium phage Xeno Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42395 | 66.803 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KT591076 | Mycobacterium phage Weiss13 | Mycobacterium phage Weiss13 unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71436 | 55.921 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT591491 | Mycobacterium phage Bricole | Mycobacterium phage Bricole unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81128 | 61.627 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| KT591490 | Mycobacterium phage Mufasa | Mycobacterium phage Mufasa unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58065 | 68.232 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KR080206 | Mycobacterium phage ShedlockHolmes | Mycobacterium phage ShedlockHolmes unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61081 | 67.301 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM923971 | Mycobacterium phage Kratio | Mycobacterium phage Kratio unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62738 | 65.67 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027199 | Mycobacterium phage Keshu | Mycobacterium phage Keshu unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61251 | 67.254 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027206 | Mycobacterium phage Milly | Mycobacterium phage Milly unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58211 | 68.312 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM925136 | Mycobacterium phage MiaZeal | Mycobacterium phage MiaZeal unclassified Omegavirus Omegavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110764 | 61.153 | Omegavirus | Unclassified | Siphoviridae | Mycobacterium |
| KM588359 | Mycobacterium phage Carcharodon | Mycobacterium phage Carcharodon unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43680 | 66.241 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KM363596 | Mycobacterium phage Omnicron | Mycobacterium phage Omnicron unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61511 | 64.034 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ944841 | Mycobacterium phage Cheetobro | Mycobacterium phage Cheetobro unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57253 | 67.967 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ567042 | Mycobacterium phage OkiRoe | Mycobacterium phage OkiRoe unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62661 | 64.938 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ510412 | Mycobacterium phage ZoeJ | Mycobacterium phage ZoeJ unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57315 | 68.525 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KF986246 | Mycobacterium phage MichelleMyBell | Mycobacterium phage MichelleMyBell Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42240 | 66.03 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KF416342 | Mycobacterium phage Papyrus | Mycobacterium phage Papyrus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70657 | 55.977 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC900379 | Mycobacterium phage PegLeg | Mycobacterium phage PegLeg unclassified Bongovirus Bongovirus Mclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80955 | 61.549 | Bongovirus | Mclasvirinae | Siphoviridae | Mycobacterium |
| JX042579 | Mycobacterium virus MacnCheese | Mycobacterium virus MacnCheese Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61567 | 67.283 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN831653 | Mycobacterium virus Fionnbarth | Mycobacterium virus Fionnbarth Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58076 | 68.007 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN624851 | Mycobacterium phage Redi | Mycobacterium phage Redi Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42594 | 66.145 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| JN243855 | Mycobacterium virus Larva | Mycobacterium virus Larva Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62991 | 65.295 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF937104 | Mycobacterium phage Pixie | Mycobacterium phage Pixie Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61147 | 67.303 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704112 | Mycobacterium phage Send513 | Mycobacterium phage Send513 Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71547 | 55.998 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| HM152765 | Mycobacterium phage Island3 | Mycobacterium phage Island3 Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47287 | 66.794 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| AF068845 | Mycobacterium phage TM4 | Mycobacterium phage TM4 Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52797 | 68.114 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY250035 | Pectobacterium phage PP47 | Pectobacterium phage PP47 Pectobacterium virus PP47 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40995 | 48.889 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| KY124276 | Pectobacterium phage PP81 | Pectobacterium phage PP81 Pectobacterium virus PP81 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40751 | 48.917 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MT951623 | Escherichia phage vB_EcoS_bov15_1 | Escherichia phage vB_EcoS_bov15_1 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44700 | 54.485 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT884015 | Escherichia phage vb_EcoS_bov25_1D | Escherichia phage vb_EcoS_bov25_1D unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44670 | 54.484 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT884013 | Escherichia phage vb_EcoS_bov16_1 | Escherichia phage vb_EcoS_bov16_1 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44700 | 54.485 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT884012 | Escherichia phage vb_EcoS_bov11C2 | Escherichia phage vb_EcoS_bov11C2 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44700 | 54.488 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT884011 | Escherichia phage vb_EcoM_bov25_3 | Escherichia phage vb_EcoM_bov25_3 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135883 | 43.685 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT884010 | Escherichia phage vb_EcoM_bov22_2 | Escherichia phage vb_EcoM_bov22_2 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135884 | 43.684 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT884009 | Escherichia phage vb_EcoM_bov11CS3 | Escherichia phage vb_EcoM_bov11CS3 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135883 | 43.685 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT884008 | Escherichia phage vb_EcoM_bov10K2 | Escherichia phage vb_EcoM_bov10K2 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135883 | 43.685 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT884007 | Escherichia phage vb_EcoM_bov10K1 | Escherichia phage vb_EcoM_bov10K1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166364 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT884006 | Escherichia phage vb_EcoM_bov9_1 | Escherichia phage vb_EcoM_bov9_1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166363 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MW151244 | Acinetobacter phage Ab_SZ3 | Acinetobacter phage Ab_SZ3 unclassified Lokivirus Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43070 | 45.621 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| MW119570 | Mycobacterium phage Lang | Mycobacterium phage Lang unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41487 | 66.831 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MT857002 | Salmonella phage pSal-SNUABM-02 | Salmonella phage pSal-SNUABM-02 unclassified Agtrevirus Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161337 | 50.102 | Agtrevirus | Aglimvirinae | Ackermannviridae | Salmonella |
| MT553346 | Mycobacterium phage Scamp | Mycobacterium phage Scamp unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.874 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT012729 | Salmonella phage vB_SenTO17 | Salmonella phage vB_SenTO17 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41658 | 50.78 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT311645 | Salmonella phage vB_Sen-E22 | Salmonella phage vB_Sen-E22 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108987 | 39.218 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT114158 | Mycobacterium phage Polymorphads | Mycobacterium phage Polymorphads unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51393 | 63.836 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN945905 | Gordonia phage John316 | Gordonia phage John316 Gordonia virus John316 Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77221 | 58.977 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MN813697 | Mycobacterium phage Noelle | Mycobacterium phage Noelle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51483 | 63.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN813692 | Cutibacterium phage P106C | Cutibacterium phage P106C unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29641 | 54.185 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813691 | Cutibacterium phage P106A | Cutibacterium phage P106A unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29554 | 54.257 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813690 | Cutibacterium phage P106M | Cutibacterium phage P106M unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29641 | 54.185 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813689 | Cutibacterium phage P106L | Cutibacterium phage P106L unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29641 | 54.188 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813688 | Cutibacterium phage P106I | Cutibacterium phage P106I unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29470 | 54.099 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813683 | Cutibacterium phage P108C | Cutibacterium phage P108C unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29494 | 54.16 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813679 | Cutibacterium phage P107A | Cutibacterium phage P107A unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29465 | 53.861 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813677 | Cutibacterium phage P107C | Cutibacterium phage P107C unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29516 | 53.994 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN813675 | Cutibacterium phage P104B | Cutibacterium phage P104B unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29330 | 54.183 | Pahexavirus | Unclassified | Siphoviridae | Cutibacterium |
| MN728684 | Mycobacterium phage Ohfah | Mycobacterium phage Ohfah unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51412 | 63.835 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586031 | Gordonia phage Gambino | Gordonia phage Gambino Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53622 | 67.092 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN585963 | Gordonia phage Wocket | Gordonia phage Wocket Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49767 | 67.113 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN369740 | Gordonia phage Delian | Gordonia phage Delian Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47896 | 67.367 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MN444874 | Gordonia phage Jellybones | Gordonia phage Jellybones Gordonia virus Jellybones Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77514 | 58.985 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MN602268 | Bacteriophage DSS3_MAL1 | Bacteriophage DSS3_MAL1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149582 | 59.589 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MN602267 | Bacteriophage DSS3_PM1 | Bacteriophage DSS3_PM1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70044 | 47.339 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MN602266 | Bacteriophage DSS3_VP1 | Bacteriophage DSS3_VP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75087 | 47.498 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MN428057 | Gordonia phage Suerte | Gordonia phage Suerte Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47306 | 66.518 | Unclassified | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MN284905 | Mycobacterium phage AbbysRanger | Mycobacterium phage AbbysRanger unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51281 | 63.679 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN284894 | Streptomyces phage Fabian | Streptomyces phage Fabian unclassified Flowerpowervirus Flowerpowervirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46505 | 60.669 | Flowerpowervirus | Beephvirinae | Podoviridae | Streptomyces |
| MN329678 | Propionibacterium phage Cota | Propionibacterium phage Cota unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29549 | 54.005 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MN234218 | Gordonia phage Powerball | Gordonia phage Powerball Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47680 | 66.265 | Unclassified | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MN234179 | Gordonia phage Pipp | Gordonia phage Pipp Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50870 | 67.275 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK967390 | Gordonia phage Beaver | Gordonia phage Beaver Gordonia virus Beaver Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77394 | 58.884 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK919470 | Gordonia phage Mellie | Gordonia phage Mellie Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48015 | 67.491 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK801728 | Gordonia phage Gorko | Gordonia phage Gorko unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75158 | 58.986 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK878905 | Mycobacterium phage TygerBlood | Mycobacterium phage TygerBlood unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.888 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK878895 | Mycobacterium phage SheldonCooper | Mycobacterium phage SheldonCooper unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51809 | 63.925 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494126 | Mycobacterium phage PeterPeter | Mycobacterium phage PeterPeter unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 63.92 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494125 | Mycobacterium phage Kyee | Mycobacterium phage Kyee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51367 | 63.973 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494117 | Mycobacterium phage Miramae | Mycobacterium phage Miramae unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51300 | 63.871 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494097 | Mycobacterium phage Lorenzo | Mycobacterium phage Lorenzo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.884 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494093 | Mycobacterium phage Phighter1804 | Mycobacterium phage Phighter1804 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51470 | 64.014 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494088 | Mycobacterium phage Mazhar510 | Mycobacterium phage Mazhar510 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51371 | 63.941 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK433267 | Gordonia phage Butterball | Gordonia phage Butterball unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77511 | 58.86 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK433265 | Gordonia phage Exiguo | Gordonia phage Exiguo unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75285 | 58.939 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK433263 | Gordonia phage FelixAlejandro | Gordonia phage FelixAlejandro unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77553 | 58.911 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK433261 | Gordonia phage Msay19 | Gordonia phage Msay19 unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77513 | 59.004 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK359356 | Mycobacterium phage Nebs | Mycobacterium phage Nebs unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51412 | 63.831 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359340 | Mycobacterium phage Phontbonne | Mycobacterium phage Phontbonne unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51263 | 63.937 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359310 | Mycobacterium phage Katalie136 | Mycobacterium phage Katalie136 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51274 | 63.676 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359302 | Gordonia phage Sombrero | Gordonia phage Sombrero Gordonia virus Sombrero Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76485 | 59.018 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK359301 | Mycobacterium phage Kingmustik0402 | Mycobacterium phage Kingmustik0402 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51365 | 63.923 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310145 | Mycobacterium phage Lemur | Mycobacterium phage Lemur unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.899 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825710 | Mycobacterium phage Ruin | Mycobacterium phage Ruin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51365 | 63.88 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727563 | Mycobacterium phage Wizard007 | Mycobacterium phage Wizard007 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51029 | 63.85 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH697588 | Mycobacterium phage Jaykayelowell | Mycobacterium phage Jaykayelowell unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51367 | 63.905 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576969 | Gordonia phage Zarbodnamra | Gordonia phage Zarbodnamra Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49729 | 67.327 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MH536818 | Gordonia phage Frokostdame | Gordonia phage Frokostdame Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52531 | 66.919 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MH536817 | Mycobacterium phage Druantia | Mycobacterium phage Druantia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51343 | 63.841 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH509443 | Mycobacterium phage Avle17 | Mycobacterium phage Avle17 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 63.918 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450114 | Arthrobacter phage Brad | Arthrobacter phage Brad unclassified Jasminevirus Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45418 | 45.731 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| MH399785 | Mycobacterium phage NotAPhaseMom | Mycobacterium phage NotAPhaseMom unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.985 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399777 | Mycobacterium phage Houdini22 | Mycobacterium phage Houdini22 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.985 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH400309 | Escherichia phage Vec13 | Escherichia phage Vec13 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39747 | 50.17 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MH191395 | Xanthomonas phage XcP1 | Xanthomonas phage XcP1 Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61830 | 52.496 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| MH155879 | Mycobacterium phage Relief | Mycobacterium phage Relief unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51368 | 63.892 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH155875 | Mycobacterium phage Pipcraft | Mycobacterium phage Pipcraft unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.913 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH230877 | Mycobacterium phage NorthStar | Mycobacterium phage NorthStar unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.921 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077581 | Mycobacterium phage Nemo27 | Mycobacterium phage Nemo27 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.943 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051247 | Mycobacterium phage Annyong | Mycobacterium phage Annyong unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51418 | 63.933 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020249 | Mycobacterium phage Skipitt | Mycobacterium phage Skipitt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51365 | 63.929 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020241 | Gordonia phage Fenry | Gordonia phage Fenry Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49965 | 67.205 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MG962373 | Mycobacterium phage Morrow | Mycobacterium phage Morrow unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51411 | 63.827 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG944224 | Mycobacterium phage Wander | Mycobacterium phage Wander unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 63.929 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG944217 | Mycobacterium phage Phacado | Mycobacterium phage Phacado unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.938 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG757155 | Gordonia phage Boneham | Gordonia phage Boneham unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77497 | 58.863 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MG757152 | Gordonia phage Adgers | Gordonia phage Adgers Gordonia virus Adgers Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76008 | 58.99 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| KX619651 | Mycobacterium phage Roosevelt | Mycobacterium phage Roosevelt unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51502 | 63.856 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG251390 | Escherichia phage VEc3 | Escherichia phage VEc3 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44895 | 44.972 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MG004687 | Escherichia phage mutPK1A2 | Escherichia phage mutPK1A2 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44784 | 45.067 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MF919533 | Propionibacterium phage Supernova | Propionibacterium phage Supernova unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29217 | 54.379 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF919522 | Propionibacterium phage MEAK | Propionibacterium phage MEAK unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29223 | 53.954 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF919521 | Gordonia phage Lysidious | Gordonia phage Lysidious Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50948 | 67.002 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MF919516 | Propionibacterium phage LilBandit | Propionibacterium phage LilBandit unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29041 | 53.975 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF919515 | Propionibacterium phage Leviosa | Propionibacterium phage Leviosa unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29451 | 53.655 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF919509 | Mycobacterium phage JetBlade | Mycobacterium phage JetBlade unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.884 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919505 | Propionibacterium phage DrParker | Propionibacterium phage DrParker unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29742 | 54.559 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF919502 | Mycobacterium phage Demsculpinboyz | Mycobacterium phage Demsculpinboyz unclassified Avanivirus Avanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57437 | 60.865 | Avanivirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919491 | Propionibacterium phage Aquarius | Propionibacterium phage Aquarius unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30112 | 54.477 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF668279 | Arthrobacter phage Nellie | Arthrobacter phage Nellie unclassified Jasminevirus Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45428 | 45.752 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| MF668278 | Mycobacterium phage Morpher26 | Mycobacterium phage Morpher26 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51294 | 63.709 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF668273 | Arthrobacter phage GurgleFerb | Arthrobacter phage GurgleFerb unclassified Jasminevirus Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45426 | 45.74 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| MF668266 | Arthrobacter phage Adat | Arthrobacter phage Adat Arthrobacter virus Adat Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45428 | 45.749 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| KY798121 | Pseudomonas phage KNP | Pseudomonas phage KNP Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40491 | 57.294 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KY798120 | Pseudomonas phage WRT | Pseudomonas phage WRT Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40214 | 57.42 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KY471266 | Mycobacterium phage TinaFeyge | Mycobacterium phage TinaFeyge unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51367 | 63.915 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY549155 | Mycobacterium phage Tinybot | Mycobacterium phage Tinybot unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51402 | 64.033 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204251 | Mycobacterium phage Thanksgivukkah | Mycobacterium phage Thanksgivukkah unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.93 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204250 | Mycobacterium phage Kratark | Mycobacterium phage Kratark unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51407 | 63.91 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204249 | Mycobacterium phage KFPoly | Mycobacterium phage KFPoly unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51365 | 63.927 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204248 | Mycobacterium phage JoongJeon | Mycobacterium phage JoongJeon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 63.947 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083062 | Mycobacterium phage Millski | Mycobacterium phage Millski unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.939 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083061 | Mycobacterium phage Palestino | Mycobacterium phage Palestino unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51369 | 63.904 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX817174 | Mycobacterium phage LittleB | Mycobacterium phage LittleB unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.876 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX550442 | Mycobacterium phage Badger | Mycobacterium phage Badger unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51274 | 63.676 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX550441 | Mycobacterium phage LittleGuy | Mycobacterium phage LittleGuy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51178 | 63.899 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX648373 | Mycobacterium phage Cocoaberry | Mycobacterium phage Cocoaberry unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.983 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX641260 | Mycobacterium phage Stasia | Mycobacterium phage Stasia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51591 | 63.591 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX579975 | Mycobacterium phage Mundrea | Mycobacterium phage Mundrea unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51257 | 63.952 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX557274 | Gordonia phage CaptainKirk2 | Gordonia phage CaptainKirk2 Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47898 | 67.368 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KX458236 | Mycobacterium phage Wilbur | Mycobacterium phage Wilbur Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.886 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU998236 | Gordonia phage Blueberry | Gordonia phage Blueberry Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54990 | 67.03 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KU985094 | Mycobacterium phage Burger | Mycobacterium phage Burger Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51371 | 63.9 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU985092 | Mycobacterium phage Holli | Mycobacterium phage Holli Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51162 | 63.885 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU963259 | Gordonia phage Guacamole | Gordonia phage Guacamole Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 67.186 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KU963245 | Gordonia phage Hotorobo | Gordonia phage Hotorobo unclassified Montyvirus Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76972 | 58.859 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| KU867906 | Mycobacterium phage Romney | Mycobacterium phage Romney Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.886 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU160650 | Arthrobacter phage Jasmine | Arthrobacter phage Jasmine Arthrobacter virus Jasmine Jasminevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46723 | 45.896 | Jasminevirus | Unclassified | Podoviridae | Arthrobacter |
| KU055616 | Mycobacterium phage Iracema64 | Mycobacterium phage Iracema64 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51637 | 64.014 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT716493 | Mycobacterium phage Caelakin | Mycobacterium phage Caelakin Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.925 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT365397 | Mycobacterium phage Maverick | Mycobacterium phage Maverick unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.943 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP057620 | Mycobacterium phage HamSlice | Mycobacterium phage HamSlice unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.923 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ510414 | Mycobacterium phage Kampy | Mycobacterium phage Kampy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51378 | 63.891 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF954506 | Mycobacterium phage Nyxis | Mycobacterium phage Nyxis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51250 | 63.924 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF841476 | Mycobacterium phage Melvin | Mycobacterium phage Melvin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51369 | 63.887 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF024733 | Mycobacterium phage Medusa | Mycobacterium phage Medusa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51384 | 63.932 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661271 | Mycobacterium phage Dhanush | Mycobacterium phage Dhanush Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51373 | 63.917 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX307703 | Mycobacterium phage Sabertooth | Mycobacterium phage Sabertooth Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51377 | 63.898 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX262216 | Propionibacterium phage P14 | Propionibacterium phage P14 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29729 | 54.082 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX262215 | Propionibacterium phage P9.1 | Propionibacterium phage P9.1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29214 | 54.121 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JQ896627 | Mycobacterium phage ICleared | Mycobacterium phage ICleared unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51440 | 63.861 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN699015 | Mycobacterium virus LHTSCC | Mycobacterium virus LHTSCC Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51813 | 63.932 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN561150 | Mycobacterium phage TiroTheta9 | Mycobacterium phage TiroTheta9 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51367 | 63.891 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN243857 | Mycobacterium phage Wile | Mycobacterium phage Wile unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51308 | 63.69 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN243856 | Mycobacterium phage MeeZee | Mycobacterium phage MeeZee Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51368 | 63.906 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF792674 | Mycobacterium phage Shaka | Mycobacterium phage Shaka Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51369 | 63.877 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| HQ224500 | Vibrio phage CTXphi | Vibrio phage CTXphi Affertcholeramvirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10638 | 44.623 | Affertcholeramvirus | Unclassified | Inoviridae | Vibrio |
| HM152766 | Mycobacterium phage Eagle | Mycobacterium phage Eagle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51436 | 63.874 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| DQ840344 | Bacillus phage 1 | Bacillus phage 1 Svunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35055 | 44.85 | Svunavirus | Unclassified | Myoviridae | Bacillus |
| EF506570 | Spiroplasma phage SkV1CR23x | Spiroplasma phage SkV1CR23x Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7870 | 22.211 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| MT740731 | Ralstonia phage Claudette | Ralstonia phage Claudette Ralstonia virus Claudettte Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57085 | 64.367 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MN119376 | Streptomyces phage Geostin | Streptomyces phage Geostin unclassified Flowerpowervirus Flowerpowervirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46175 | 60.695 | Flowerpowervirus | Beephvirinae | Podoviridae | Streptomyces |
| MH155868 | Streptomyces phage FlowerPower | Streptomyces phage FlowerPower Streptomyces virus FlowerPower Flowerpowervirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46133 | 60.681 | Flowerpowervirus | Beephvirinae | Podoviridae | Streptomyces |
| MG518520 | Streptomyces phage Immanuel3 | Streptomyces phage Immanuel3 Streptomyces virus Immanuel3 Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46094 | 59.632 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| LR811969 | Salmonella phage Se_EM3 | Salmonella phage Se_EM3 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157099 | 44.546 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| LR811968 | Salmonella phage Se_EM4 | Salmonella phage Se_EM4 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157810 | 44.743 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| LR811967 | Salmonella phage Se_EM1 | Salmonella phage Se_EM1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159312 | 44.488 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| LR800396 | Salmonella phage Se_AO1 | Salmonella phage Se_AO1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157543 | 44.63 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MW091529 | Bacteriophage sp. 103231 | Bacteriophage sp. 103231 unclassified bacterial viruses Viruses | 18183 | 59.121 | Unclassified | Unclassified | Unclassified | Unspecified |
| MW091528 | Bacteriophage sp. 438212 | Bacteriophage sp. 438212 unclassified bacterial viruses Viruses | 31712 | 65.187 | Unclassified | Unclassified | Unclassified | Unspecified |
| KJ183193 | Podovirus Lau218 | Podovirus Lau218 Lauvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38756 | 34.707 | Lauvirus | Unclassified | Autographiviridae | Unspecified |
| KJ183192 | Podovirus Lau218 | Podovirus Lau218 Lauvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39311 | 34.69 | Lauvirus | Unclassified | Autographiviridae | Unspecified |
| KJ183191 | Podophage Lau218 | Podophage Lau218 Lauvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39425 | 34.699 | Lauvirus | Unclassified | Autographiviridae | Unspecified |
| MW084976 | Bacillus phage Kirov | Bacillus phage Kirov Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165667 | 35.48 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MW057919 | Pseudomonas phage vB_PaeM_kmuB | Pseudomonas phage vB_PaeM_kmuB unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266592 | 36.996 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MW056503 | Acinetobacter phage vB_AbaA_fBenAci003 | Acinetobacter phage vB_AbaA_fBenAci003 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40556 | 39.23 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW056502 | Acinetobacter phage vB_AbaA_fBenAci002 | Acinetobacter phage vB_AbaA_fBenAci002 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41953 | 39.218 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW056501 | Acinetobacter phage vB_AbaA_fBenAci001 | Acinetobacter phage vB_AbaA_fBenAci001 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40535 | 39.238 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MW042812 | Klebsiella phage 066134 | Klebsiella phage 066134 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.028 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042811 | Klebsiella phage 066121 | Klebsiella phage 066121 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.038 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042810 | Klebsiella phage 066059 | Klebsiella phage 066059 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40523 | 53.012 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042809 | Klebsiella phage 066058 | Klebsiella phage 066058 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40522 | 52.998 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042808 | Klebsiella phage 066056 | Klebsiella phage 066056 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40952 | 52.627 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042807 | Klebsiella phage 066053 | Klebsiella phage 066053 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40185 | 52.87 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042806 | Klebsiella phage 066046 | Klebsiella phage 066046 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39887 | 53.032 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042805 | Klebsiella phage 066045 | Klebsiella phage 066045 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39865 | 53.087 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042804 | Klebsiella phage 066042 | Klebsiella phage 066042 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39833 | 53.067 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042803 | Klebsiella phage 066041 | Klebsiella phage 066041 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40613 | 53.114 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042802 | Klebsiella phage 066039 | Klebsiella phage 066039 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51633 | 51.556 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MW042801 | Klebsiella phage 066038 | Klebsiella phage 066038 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.043 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042800 | Klebsiella phage 066037 | Klebsiella phage 066037 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40174 | 53.099 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042799 | Klebsiella phage 066036 | Klebsiella phage 066036 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.028 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042798 | Klebsiella phage 066032 | Klebsiella phage 066032 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40605 | 53.149 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042797 | Klebsiella phage 066031 | Klebsiella phage 066031 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.028 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042796 | Klebsiella phage 066028 | Klebsiella phage 066028 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40605 | 53.154 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042795 | Klebsiella phage 066027 | Klebsiella phage 066027 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40526 | 53.181 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042794 | Klebsiella phage 066026 | Klebsiella phage 066026 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.035 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042793 | Klebsiella phage 066025 | Klebsiella phage 066025 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.028 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042792 | Klebsiella phage 066024 | Klebsiella phage 066024 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40770 | 53.186 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042791 | Klebsiella phage 066023 | Klebsiella phage 066023 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40516 | 52.977 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042790 | Klebsiella phage 066022 | Klebsiella phage 066022 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40243 | 53.02 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042789 | Klebsiella phage 066019 | Klebsiella phage 066019 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40606 | 53.071 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042788 | Klebsiella phage 066015 | Klebsiella phage 066015 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40967 | 52.982 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MW042787 | Klebsiella phage 066013 | Klebsiella phage 066013 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40967 | 52.974 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT708548 | Streptomyces phage Salutena | Streptomyces phage Salutena unclassified Arequatrovirus Arequatrovirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51956 | 67.501 | Arequatrovirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT708545 | Rhizobium phage Pasto | Rhizobium phage Pasto unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42407 | 58.641 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MW013503 | Salmonella virus L cII-101 | Salmonella virus L cII-101 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40664 | 47.514 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW013502 | Salmonella virus L cI-40 13-am43 | Salmonella virus L cI-40 13-am43 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40663 | 47.5 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MW004544 | Enterococcus phage EFGrKN | Enterococcus phage EFGrKN unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147532 | 37.311 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| LC589952 | Enterobacter phage vB_EkoM5VN | Enterobacter phage vB_EkoM5VN unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178408 | 44.87 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MW074885 | Escherichia phage JB01 | Escherichia phage JB01 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39355 | 48.591 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT708549 | Burkholderia phage Maja | Burkholderia phage Maja unclassified Bcepfunavirus Bcepfunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68393 | 54.503 | Bcepfunavirus | Unclassified | Myoviridae | Burkholderia |
| MW012634 | Bacillus phage DLc1 | Bacillus phage DLc1 Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28950 | 31.095 | Unclassified | Northropvirinae | Salasmaviridae | Bacillus |
| MN539721 | Sulfolobus super-elliptical virus | Sulfolobus super-elliptical virus unclassified Fuselloviridae Fuselloviridae Viruses | 15365 | 33.433 | Unclassified | Unclassified | Fuselloviridae | Sulfolobus |
| CP025907 | Escherichia phage 204576 | Escherichia phage 204576 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31521 | 52.378 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MT774405 | crAssphage cr115_1 | crAssphage cr115_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99848 | 32.553 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774404 | crAssphage cr114_1 | crAssphage cr114_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98308 | 32.79 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774394 | crAssphage cr271_1 | crAssphage cr271_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92815 | 30.545 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774393 | crAssphage cr127_1 | crAssphage cr127_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83412 | 25.922 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774392 | crAssphage cr128_1 | crAssphage cr128_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95150 | 33.197 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774391 | crAssphage cr126_1 | crAssphage cr126_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99212 | 34.776 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774390 | crAssphage cr85_1 | crAssphage cr85_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98164 | 37.722 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774389 | crAssphage cr116_1 | crAssphage cr116_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101082 | 32.585 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774388 | crAssphage cr112_1 | crAssphage cr112_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102301 | 34.681 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774387 | crAssphage cr111_1 | crAssphage cr111_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102434 | 39.544 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774386 | crAssphage cr110_1 | crAssphage cr110_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89781 | 30.753 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774385 | crAssphage cr108_1 | crAssphage cr108_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101437 | 37.271 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774384 | crAssphage cr8_1 | crAssphage cr8_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92668 | 30.185 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774383 | crAssphage cr11_1 | crAssphage cr11_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93559 | 30.001 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774382 | crAssphage cr10_1 | crAssphage cr10_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99985 | 33.532 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774381 | crAssphage cr3_1 | crAssphage cr3_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91101 | 30.729 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774380 | crAssphage cr1_1 | crAssphage cr1_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99882 | 32.195 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774379 | crAssphage cr118_1 | crAssphage cr118_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92600 | 28.706 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774378 | crAssphage cr106_1 | crAssphage cr106_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101662 | 33.424 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774377 | crAssphage cr107_1 | crAssphage cr107_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93787 | 31.997 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774376 | crAssphage cr55_1 | crAssphage cr55_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98340 | 31.9 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774375 | crAssphage cr50_1 | crAssphage cr50_1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98826 | 32.58 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774411 | crAssphage cr273_1 | crAssphage cr273_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97886 | 36.266 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774410 | crAssphage cr272_1 | crAssphage cr272_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98667 | 36.183 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774409 | crAssphage cr131_1 | crAssphage cr131_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96717 | 35.294 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774408 | crAssphage cr130_1 | crAssphage cr130_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94826 | 31.772 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774407 | crAssphage cr125_1 | crAssphage cr125_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98095 | 32.751 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774406 | crAssphage cr124_1 | crAssphage cr124_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101532 | 35.803 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774403 | crAssphage cr113_1 | crAssphage cr113_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98557 | 33.037 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774402 | crAssphage cr7_1 | crAssphage cr7_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103832 | 35.691 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774401 | crAssphage cr6_1 | crAssphage cr6_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97422 | 30.93 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774400 | crAssphage cr4_1 | crAssphage cr4_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96055 | 33.495 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774399 | crAssphage cr109_1 | crAssphage cr109_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99168 | 33.051 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774398 | crAssphage cr60_1 | crAssphage cr60_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100198 | 33.09 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774397 | crAssphage cr56_1 | crAssphage cr56_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97290 | 33.008 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774396 | crAssphage cr53_1 | crAssphage cr53_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101153 | 32.971 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT774395 | crAssphage cr52_1 | crAssphage cr52_1 UAG-readthrough crAss clade crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103665 | 34.658 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW067003 | CrAssphage sp. C0531BW4 | CrAssphage sp. C0531BW4 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99127 | 29.014 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW067002 | CrAssphage sp. C0526BW15 | CrAssphage sp. C0526BW15 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99121 | 29.219 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW067001 | CrAssphage sp. C0521BW15 | CrAssphage sp. C0521BW15 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98005 | 29.142 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MW067000 | CrAssphage sp. C0521BD4 | CrAssphage sp. C0521BD4 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98005 | 29.134 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT986027 | Microbacterium phage Winzigespinne | Microbacterium phage Winzigespinne unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41556 | 63.442 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT986026 | Microbacterium phage Gubbabump | Microbacterium phage Gubbabump unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41492 | 63.405 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| CP062992 | Klebsiella phage AShe-2020a | Klebsiella phage AShe-2020a unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168979 | 39.318 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| CP058330 | Klebsiella phage vB_Kpn_1825-KPC53 | Klebsiella phage vB_Kpn_1825-KPC53 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 119621 | 50.285 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MT894005 | Klebsiella phage MMBB | Klebsiella phage MMBB unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50346 | 50.246 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MT894004 | Klebsiella phage MEW1 | Klebsiella phage MEW1 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47129 | 49.252 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MT968995 | Escherichia phage vB_EcoM_WL-3 | Escherichia phage vB_EcoM_WL-3 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165369 | 35.48 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT932329 | Campylobacter phage F379 | Campylobacter phage F379 unclassified Firehammervirus Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 183102 | 27.218 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MT366580 | Vibrio phage HY01 | Vibrio phage HY01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41772 | 47.446 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KC109329 | Escherichia phage CLB_P1 | Escherichia phage CLB_P1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40218 | 50.052 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT889398 | Gordonia phage Herod | Gordonia phage Herod unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53826 | 66.462 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MT889397 | Mycobacterium phage OfUltron | Mycobacterium phage OfUltron unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58617 | 61.091 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889396 | Mycobacterium phage Seabastian | Mycobacterium phage Seabastian unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58618 | 61.096 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889390 | Mycobacterium phage Bartimeaus | Mycobacterium phage Bartimeaus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.867 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889387 | Arthrobacter phage DanielleIgnace | Arthrobacter phage DanielleIgnace Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61275 | 62.936 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT889386 | Mycobacterium phage CentreCat | Mycobacterium phage CentreCat unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51412 | 63.87 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889385 | Mycobacterium phage Thoth | Mycobacterium phage Thoth unclassified Plotvirus Plotvirus Dclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64606 | 59.688 | Plotvirus | Dclasvirinae | Siphoviridae | Mycobacterium |
| MT889382 | Mycobacterium phage TroyPia | Mycobacterium phage TroyPia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51407 | 63.912 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889380 | Mycobacterium phage Coco12 | Mycobacterium phage Coco12 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57693 | 61.136 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889377 | Microbacterium phage Zayuliv | Microbacterium phage Zayuliv unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40457 | 63.865 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MT889374 | Mycobacterium phage Firehouse51 | Mycobacterium phage Firehouse51 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58166 | 61.476 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT889364 | Arthrobacter phage Yavru | Arthrobacter phage Yavru Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15193 | 64.253 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT423824 | Staphylococcus virus pSp_SNUABM-S | Staphylococcus virus pSp_SNUABM-S Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40159 | 35.733 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT162468 | Synechococcus phage S-H25 | Synechococcus phage S-H25 unclassified Tefnutvirus Tefnutvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173659 | 41.171 | Tefnutvirus | Unclassified | Myoviridae | Synechococcus |
| MT740738 | Ralstonia phage Gamede | Ralstonia phage Gamede Ralstonia virus Gamede Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58603 | 64.602 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| LR877332 | Shigella phage vB_SsoM_JK08 | Shigella phage vB_SsoM_JK08 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169207 | 35.377 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| LR877331 | Klebsiella phage vB_KpnM_311F | Klebsiella phage vB_KpnM_311F unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166710 | 39.595 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| LR757892 | Klebsiella phage vB_KpnS_2811 | Klebsiella phage vB_KpnS_2811 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49131 | 51.002 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| LR746311 | Shigella phage vB_SsoM_113 | Shigella phage vB_SsoM_113 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169243 | 35.366 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| LR746310 | Klebsiella phage vB_KpnM_05F | Klebsiella phage vB_KpnM_05F unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174231 | 41.975 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| LR746137 | Escherichia phage vB_EcoM_UP17 | Escherichia phage vB_EcoM_UP17 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152148 | 39.165 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MT833283 | Escherichia coli phage vB_EcoS_Ace | Escherichia coli phage vB_EcoS_Ace unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112791 | 39.962 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MT939488 | Erwinia phage pEa_SNUABM_50 | Erwinia phage pEa_SNUABM_50 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 356948 | 34.447 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MT939487 | Erwinia phage pEa_SNUABM_47 | Erwinia phage pEa_SNUABM_47 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 355376 | 34.482 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MT939486 | Erwinia phage pEa_SNUABM_12 | Erwinia phage pEa_SNUABM_12 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 358115 | 34.41 | Eneladusvirus | Unclassified | Myoviridae | Erwinia |
| MT900488 | Streptococcus phage SA01 | Streptococcus phage SA01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36088 | 37.536 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MT900487 | Streptococcus phage 23TH | Streptococcus phage 23TH Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32272 | 39.774 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MT992243 | Acinetobacter phage DMU1 | Acinetobacter phage DMU1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43482 | 47.834 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MT952005 | Aeromonas phage BUCT551 | Aeromonas phage BUCT551 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61382 | 61.71 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MT933737 | Pseudomonas phage PASA16 | Pseudomonas phage PASA16 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66127 | 55.563 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT926124 | Staphylococcus phage vB_SauP_EBHT | Staphylococcus phage vB_SauP_EBHT unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17471 | 29.569 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MT939242 | Enterococcus phage 9184 | Enterococcus phage 9184 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44108 | 35.817 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT939253 | Klebsiella phage vB_KpnM_Seu621 | Klebsiella phage vB_KpnM_Seu621 unclassified Mydovirus Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142896 | 44.638 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| MT939252 | Klebsiella phage vB_KpnP_Dlv622 | Klebsiella phage vB_KpnP_Dlv622 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44687 | 54.063 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT932212 | Salmonella phage VSe13 | Salmonella phage VSe13 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39265 | 47.332 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MT932213 | Escherichia phage VEc74 | Escherichia phage VEc74 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166100 | 35.43 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT932211 | Escherichia phage VEcB | Escherichia phage VEcB unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147702 | 37.556 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MT952846 | Microbacterium phage MaeLinda | Microbacterium phage MaeLinda unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41809 | 63.565 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT952844 | Gordonia phage Bosnia | Gordonia phage Bosnia unclassified Nymphadoravirus Nymphadoravirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54546 | 66.188 | Nymphadoravirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MT862763 | Escherichia phage vB_EcoP-U8 | Escherichia phage vB_EcoP-U8 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40294 | 50.139 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT857001 | Enterococcus phage EFA1 | Enterococcus phage EFA1 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40454 | 34.812 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT833387 | Escherichia phage HF4s | Escherichia phage HF4s Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39390 | 47.469 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MN788369 | Dickeya phage Ds3CZ | Dickeya phage Ds3CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155285 | 49.11 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813053 | Dickeya phage Ds25CZ | Dickeya phage Ds25CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151710 | 49.108 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813052 | Dickeya phage Ds23CZ | Dickeya phage Ds23CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149364 | 49.393 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813051 | Dickeya phage Ds20CZ | Dickeya phage Ds20CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154720 | 49.104 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813050 | Dickeya phage Ds16CZ | Dickeya phage Ds16CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152837 | 49.211 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813049 | Dickeya phage Ds9CZ | Dickeya phage Ds9CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154716 | 49.107 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MN813048 | Dickeya phage Ds5CZ | Dickeya phage Ds5CZ unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154722 | 49.112 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MT708550 | Achromobacter phage Mano | Achromobacter phage Mano Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42452 | 64.303 | Unclassified | Unclassified | Myoviridae | Achromobacter |
| MT708547 | Klebsiella phage Muenster | Klebsiella phage Muenster unclassified Alcyoneusvirus Alcyoneusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 346937 | 31.865 | Alcyoneusvirus | Unclassified | Myoviridae | Klebsiella |
| MT708546 | Rhizobium phage Paso | Rhizobium phage Paso unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45055 | 55.022 | Unclassified | Unclassified | Autographiviridae | Rhizobium |
| MT192033 | Chlorobiaceae phage CV-1-33 | Chlorobiaceae phage CV-1-33 unclassified bacterial viruses Viruses | 34612 | 44.944 | Unclassified | Unclassified | Unclassified | Unspecified |
| MT974436 | Salmonella phage ISTP3 | Salmonella phage ISTP3 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156121 | 44.779 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN715356 | Escherichia phage vB_EcoP_EcoN5 | Escherichia phage vB_EcoP_EcoN5 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76083 | 42.087 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| KY053532 | Siphovirus 29632 | Siphovirus 29632 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29632 | 59.638 | Siphovirus | Unclassified | Siphoviridae | Unspecified |
| KY083726 | Pectobacterium phage A38 | Pectobacterium phage A38 unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75764 | 48.701 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MT813197 | Escherichia phage vB_EcoP_Kapi1 | Escherichia phage vB_EcoP_Kapi1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39436 | 47.14 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MT818420 | Microbacterium phage Gilda | Microbacterium phage Gilda unclassified Krampusvirus Krampusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56701 | 63.763 | Krampusvirus | Unclassified | Siphoviridae | Microbacterium |
| MT947439 | Protaetiibacter phage SSC1 | Protaetiibacter phage SSC1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Unclassified | Unclassified | Microviridae | Protaetiibacter |
| MT771346 | Gordonia phage MichaelScott | Gordonia phage MichaelScott Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48397 | 66.019 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT740245 | Proteus phage M4H10_20 | Proteus phage M4H10_20 unclassified Novosibvirus Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104575 | 38.286 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MT740244 | Proteus phage 3H10_20 | Proteus phage 3H10_20 unclassified Privateervirus Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90382 | 34.548 | Privateervirus | Unclassified | Podoviridae | Proteus |
| MN857473 | Teseptimavirus S2B | Teseptimavirus S2B unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45682 | 66.766 | Teseptimavirus | Unclassified | Autographiviridae | Unspecified |
| MT787017 | Staphylococcus phage WV | Staphylococcus phage WV unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141342 | 30.574 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT818425 | Mycobacterium phage NiebruSaylor | Mycobacterium phage NiebruSaylor unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70777 | 65.376 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT818418 | Mycobacterium phage Shida | Mycobacterium phage Shida unclassified Corndogvirus Corndogvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71343 | 65.411 | Corndogvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT818417 | Mycobacterium phage Maxo | Mycobacterium phage Maxo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51378 | 63.891 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT822288 | Erwinia phage pEp_SNUABM_12 | Erwinia phage pEp_SNUABM_12 unclassified Ningirsuvirus Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39980 | 51.198 | Ningirsuvirus | Studiervirinae | Autographiviridae | Erwinia |
| MT822287 | Erwinia phage pEp_SNUABM_11 | Erwinia phage pEp_SNUABM_11 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39626 | 52.097 | Unclassified | Studiervirinae | Autographiviridae | Erwinia |
| MT822286 | Erwinia phage pEp_SNUABM_10 | Erwinia phage pEp_SNUABM_10 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39879 | 52.118 | Unclassified | Studiervirinae | Autographiviridae | Erwinia |
| MT822285 | Erwinia phage pEp_SNUABM_04 | Erwinia phage pEp_SNUABM_04 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39649 | 52.193 | Unclassified | Studiervirinae | Autographiviridae | Erwinia |
| MT822284 | Erwinia phage pEp_SNUABM_03 | Erwinia phage pEp_SNUABM_03 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39879 | 52.125 | Unclassified | Studiervirinae | Autographiviridae | Erwinia |
| MT939241 | Enterococcus phage 9183 | Enterococcus phage 9183 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86301 | 33.003 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT939240 | Enterococcus phage 9181 | Enterococcus phage 9181 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71854 | 40.595 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT777452 | Bacillus phage Baseball_field | Bacillus phage Baseball_field unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26863 | 30.436 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MT771350 | Microbacterium phage TatarkaPM | Microbacterium phage TatarkaPM unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41813 | 63.519 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT771348 | Gordonia phage BiteSize | Gordonia phage BiteSize unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66696 | 59.549 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MT771345 | Gordonia phage Sienna | Gordonia phage Sienna unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66807 | 59.591 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MT771344 | Microbacterium phage Zhengyi | Microbacterium phage Zhengyi unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54803 | 58.539 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT771342 | Gordonia phage Matteo | Gordonia phage Matteo unclassified Attisvirus Attisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44079 | 66.635 | Attisvirus | Unclassified | Siphoviridae | Gordonia |
| MT771341 | Gordonia phage DirtyBoi | Gordonia phage DirtyBoi unclassified Hedwigvirus Hedwigvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43833 | 67.093 | Hedwigvirus | Unclassified | Siphoviridae | Gordonia |
| MT771340 | Mycobacterium phage Jorgensen | Mycobacterium phage Jorgensen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53771 | 63.638 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT771339 | Gordonia phage Archimedes | Gordonia phage Archimedes Ghobesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44938 | 62.818 | Ghobesvirus | Unclassified | Siphoviridae | Gordonia |
| MT757399 | Enterococcus phage AE4_17 | Enterococcus phage AE4_17 unclassified Copernicusvirus Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18477 | 32.879 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MN539738 | Staphylococcus phage vB_SauM-D | Staphylococcus phage vB_SauM-D unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139031 | 30.451 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MN539737 | Staphylococcus phage vB_SauM-C | Staphylococcus phage vB_SauM-C unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140086 | 30.426 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MN539736 | Staphylococcus phage vB_SauM-A | Staphylococcus phage vB_SauM-A unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139088 | 30.458 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT811961 | Vibrio phage vB_pir03 | Vibrio phage vB_pir03 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 286284 | 43.62 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MT233525 | Salmonella phage vB_Sen_H9 | Salmonella phage vB_Sen_H9 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34894 | 38.594 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT233524 | Salmonella phage vB_Sen_I1 | Salmonella phage vB_Sen_I1 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112111 | 38.993 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT241605 | Achromobacter phage AMA1 | Achromobacter phage AMA1 unclassified Steinhofvirus Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46328 | 56.277 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| MT002875 | Pseudoalteromonas phage AL | Pseudoalteromonas phage AL Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33582 | 40.075 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KX147229 | Bacillus phage Belinda | Bacillus phage Belinda unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162308 | 38.784 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX156152 | Bacillus phage Smudge | Bacillus phage Smudge unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162040 | 38.755 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489402 | Bacillus phage Riley | Bacillus phage Riley Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162816 | 37.784 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489401 | Bacillus phage Megatron | Bacillus phage Megatron Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158750 | 38.803 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489400 | Bacillus phage Hoody T | Bacillus phage Hoody T Bacillus virus HoodyT Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159837 | 38.014 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489399 | Bacillus phage Hakuna | Bacillus phage Hakuna Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158100 | 38.696 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489398 | Bacillus phage Evoli | Bacillus phage Evoli Bacillus virus Evoli Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159656 | 38.059 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ489397 | Bacillus phage CAM003 | Bacillus phage CAM003 Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160541 | 38.031 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MT225100 | Escherichia phage Lys8385Vzw | Escherichia phage Lys8385Vzw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50953 | 51.553 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MT585315 | Flavobacterium phage V189 | Flavobacterium phage V189 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.74 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585314 | Flavobacterium phage V188 | Flavobacterium phage V188 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.738 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585313 | Flavobacterium phage V186 | Flavobacterium phage V186 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46498 | 29.926 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585312 | Flavobacterium phage V184 | Flavobacterium phage V184 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.738 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585311 | Flavobacterium phage V183 | Flavobacterium phage V183 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.738 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585310 | Flavobacterium phage FCOV-F56 | Flavobacterium phage FCOV-F56 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46494 | 29.924 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585309 | Flavobacterium phage FCOV-F48 | Flavobacterium phage FCOV-F48 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49063 | 29.733 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585308 | Flavobacterium phage FCOV-F47 | Flavobacterium phage FCOV-F47 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49066 | 29.735 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585307 | Flavobacterium phage FCOV-F44 | Flavobacterium phage FCOV-F44 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49043 | 29.739 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585306 | Flavobacterium phage FCOV-F43 | Flavobacterium phage FCOV-F43 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49080 | 29.741 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585305 | Flavobacterium phage FCOV-F42 | Flavobacterium phage FCOV-F42 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49066 | 29.737 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585304 | Flavobacterium phage FCOV-F41 | Flavobacterium phage FCOV-F41 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.742 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585303 | Flavobacterium phage FCOV-F40 | Flavobacterium phage FCOV-F40 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49063 | 29.739 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585302 | Flavobacterium phage FCOV-F39 | Flavobacterium phage FCOV-F39 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49082 | 29.744 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585301 | Flavobacterium phage FCOV-F38 | Flavobacterium phage FCOV-F38 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49055 | 29.738 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585300 | Flavobacterium phage FCOV-F36 | Flavobacterium phage FCOV-F36 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49072 | 29.742 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585299 | Flavobacterium phage FCOV-F35 | Flavobacterium phage FCOV-F35 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49053 | 29.739 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585298 | Flavobacterium phage FCOV-F34 | Flavobacterium phage FCOV-F34 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49060 | 29.739 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585297 | Flavobacterium phage FCOV-F33 | Flavobacterium phage FCOV-F33 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49075 | 29.742 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585296 | Flavobacterium phage FCOV-F32 | Flavobacterium phage FCOV-F32 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49068 | 29.726 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585295 | Flavobacterium phage FCOV-F31 | Flavobacterium phage FCOV-F31 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49073 | 29.727 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585294 | Flavobacterium phage FCOV-F30 | Flavobacterium phage FCOV-F30 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.733 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585293 | Flavobacterium phage FCOV-F29 | Flavobacterium phage FCOV-F29 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49063 | 29.723 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585292 | Flavobacterium phage FCOV-F28 | Flavobacterium phage FCOV-F28 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49064 | 29.735 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585291 | Flavobacterium phage FCOV-F26 | Flavobacterium phage FCOV-F26 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49073 | 29.725 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585290 | Flavobacterium phage FCOV-F24 | Flavobacterium phage FCOV-F24 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49073 | 29.727 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585289 | Flavobacterium phage FCOV-F23 | Flavobacterium phage FCOV-F23 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.736 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585288 | Flavobacterium phage FCOV-F22 | Flavobacterium phage FCOV-F22 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49060 | 29.739 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585287 | Flavobacterium phage FCOV-F21 | Flavobacterium phage FCOV-F21 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49072 | 29.74 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585286 | Flavobacterium phage FCOV-F20 | Flavobacterium phage FCOV-F20 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.74 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585285 | Flavobacterium phage FCOV-F19 | Flavobacterium phage FCOV-F19 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49036 | 29.737 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585284 | Flavobacterium phage FCOV-F18 | Flavobacterium phage FCOV-F18 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.736 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585283 | Flavobacterium phage FCOV-F17 | Flavobacterium phage FCOV-F17 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49076 | 29.736 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585282 | Flavobacterium phage FCOV-F15 | Flavobacterium phage FCOV-F15 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47224 | 30.127 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585281 | Flavobacterium phage FCOV-F14 | Flavobacterium phage FCOV-F14 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47224 | 30.122 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585280 | Flavobacterium phage FCOV-F12 | Flavobacterium phage FCOV-F12 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49081 | 29.745 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585279 | Flavobacterium phage FCOV-F11 | Flavobacterium phage FCOV-F11 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49078 | 29.73 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585278 | Flavobacterium phage FCOV-F10 | Flavobacterium phage FCOV-F10 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49078 | 29.726 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585277 | Flavobacterium phage FCOV-F8 | Flavobacterium phage FCOV-F8 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49063 | 29.733 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585276 | Flavobacterium phage FCOV-F7 | Flavobacterium phage FCOV-F7 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49058 | 29.724 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585275 | Flavobacterium phage FCOV-F4 | Flavobacterium phage FCOV-F4 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49052 | 29.722 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585274 | Flavobacterium phage FCOV-F3 | Flavobacterium phage FCOV-F3 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49071 | 29.726 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT585273 | Flavobacterium phage FCOV-F1 | Flavobacterium phage FCOV-F1 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49077 | 29.729 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT457475 | Arthronema virus TR020 | Arthronema virus TR020 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44805 | 45.843 | Unclassified | Unclassified | Podoviridae | Arthronema |
| MK756096 | Flavobacterium phage V186 | Flavobacterium phage V186 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46511 | 29.931 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756095 | Flavobacterium phage FCOV-S2 | Flavobacterium phage FCOV-S2 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46386 | 29.908 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756094 | Flavobacterium phage FCOV-S1 | Flavobacterium phage FCOV-S1 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46512 | 29.93 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756093 | Flavobacterium phage FCOV-F54 | Flavobacterium phage FCOV-F54 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47204 | 30.118 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756092 | Flavobacterium phage FCOV-F46 | Flavobacterium phage FCOV-F46 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47214 | 30.129 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756091 | Flavobacterium phage FCOV-F45 | Flavobacterium phage FCOV-F45 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47227 | 30.131 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756090 | Flavobacterium phage FCOV-F37 | Flavobacterium phage FCOV-F37 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49098 | 29.757 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756089 | Flavobacterium phage FCOV-F27 | Flavobacterium phage FCOV-F27 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49168 | 29.757 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756088 | Flavobacterium phage FCOV-F25 | Flavobacterium phage FCOV-F25 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49080 | 29.735 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756087 | Flavobacterium phage FCOV-F16 | Flavobacterium phage FCOV-F16 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47232 | 30.13 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756086 | Flavobacterium phage FCOV-F13 | Flavobacterium phage FCOV-F13 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47223 | 30.123 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756085 | Flavobacterium phage FCOV-F9 | Flavobacterium phage FCOV-F9 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49115 | 29.749 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756084 | Flavobacterium phage FCOV-F6 | Flavobacterium phage FCOV-F6 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49115 | 29.742 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MK756083 | Flavobacterium phage FCOV-F2 | Flavobacterium phage FCOV-F2 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49073 | 29.729 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MH598512 | Bacillus phage Wes44 | Bacillus phage Wes44 Bacillus virus Wes44 Carmenvirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42248 | 42.028 | Carmenvirus | Gutmannvirinae | Siphoviridae | Bacillus |
| MG784342 | Bacillus phage Carmen17 | Bacillus phage Carmen17 Bacillus virus Carmen17 Carmenvirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41820 | 42.037 | Carmenvirus | Gutmannvirinae | Siphoviridae | Bacillus |
| MT522007 | Gordonia phage Epsocamisio | Gordonia phage Epsocamisio unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52314 | 62.01 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MT522004 | Mycobacterium phage Gail | Mycobacterium phage Gail unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53702 | 62.387 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT498056 | Mycobacterium phage Chaph | Mycobacterium phage Chaph unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51383 | 63.899 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT553347 | Mycobacterium phage Awesomesauce | Mycobacterium phage Awesomesauce unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57054 | 62.897 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT553338 | Mycobacterium phage Deano | Mycobacterium phage Deano unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51379 | 63.876 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT361972 | Acinetobacter phage Ab11510-phi | Acinetobacter phage Ab11510-phi unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50916 | 38.864 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| MT310885 | Mycobacterium phage AN9 | Mycobacterium phage AN9 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51141 | 63.118 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310883 | Mycobacterium phage D32 | Mycobacterium phage D32 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49127 | 63.543 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN945902 | Rhodococcus phage Dinger | Rhodococcus phage Dinger unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46617 | 58.803 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MN908690 | Microbacterium phage Hubbs | Microbacterium phage Hubbs Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63862 | 62.763 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MN735432 | Mycobacteriophage Whitty | Mycobacteriophage Whitty unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42393 | 66.197 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN735431 | Mycobacterium phage Hegedechwinu | Mycobacterium phage Hegedechwinu unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57346 | 61.5 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN813698 | Gordonia phage Volt | Gordonia phage Volt unclassified Ronaldovirus Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68553 | 50.18 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MN813696 | Mycobacterium phage Cintron | Mycobacterium phage Cintron unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51679 | 64.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586056 | Streptomyces phage FidgetOrca | Streptomyces phage FidgetOrca unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56115 | 59.515 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN586054 | Streptomyces phage FrodoSwaggins | Streptomyces phage FrodoSwaggins unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56472 | 59.412 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN586051 | Mycobacterium phage Myrale | Mycobacterium phage Myrale unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76419 | 62.834 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586050 | Mycobacterium phage Lilpickle | Mycobacterium phage Lilpickle unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74969 | 62.979 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586044 | Mycobacterium phage Rimmer | Mycobacterium phage Rimmer unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75660 | 62.967 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586043 | Mycobacterium phage Buck | Mycobacterium phage Buck unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76688 | 62.972 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586041 | Mycobacterium phage Elite2014 | Mycobacterium phage Elite2014 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74969 | 62.981 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586035 | Mycobacterium phage ChosenOne | Mycobacterium phage ChosenOne unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74970 | 62.981 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586034 | Mycobacterium phage Hoonter | Mycobacterium phage Hoonter unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75267 | 63.041 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586032 | Mycobacterium phage Command613 | Mycobacterium phage Command613 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76025 | 62.921 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586019 | Mycobacterium phage Stark | Mycobacterium phage Stark unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74731 | 63.046 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586017 | Mycobacterium phage OrionPax | Mycobacterium phage OrionPax unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75101 | 63.018 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586014 | Gordonia phage Keelan | Gordonia phage Keelan unclassified Ronaldovirus Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67573 | 51.12 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MN586013 | Mycobacterium phage Traaww1 | Mycobacterium phage Traaww1 unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75423 | 62.873 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586002 | Mycobacterium phage BiancaTri92 | Mycobacterium phage BiancaTri92 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52181 | 63.431 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585984 | Mycobacterium phage AgronaGT15 | Mycobacterium phage AgronaGT15 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48850 | 64.219 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585969 | Mycobacterium phage Dieselweasel | Mycobacterium phage Dieselweasel unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50770 | 64.028 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369756 | Gordonia phage Anon | Gordonia phage Anon unclassified Soupsvirus Soupsvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52661 | 62.509 | Soupsvirus | Unclassified | Siphoviridae | Gordonia |
| MN369752 | Mycobacterium phage Flathead | Mycobacterium phage Flathead unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57663 | 61.181 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369739 | Mycobacterium phage Kenuha5 | Mycobacterium phage Kenuha5 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56502 | 61.693 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN497415 | Vibrio phage vB_VhaM_VH-8 | Vibrio phage vB_VhaM_VH-8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87255 | 36.077 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN497414 | Vibrio phage vB_VhaP_VH-5 | Vibrio phage vB_VhaP_VH-5 unclassified Beijerinckvirinae Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42485 | 44.479 | Unclassified | Beijerinckvirinae | Autographiviridae | Vibrio |
| MN369751 | Mycobacterium phage TDanisky | Mycobacterium phage TDanisky unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56275 | 61.233 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN428059 | Mycobacterium phage Kanye | Mycobacterium phage Kanye unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75453 | 63.067 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN284902 | Arthrobacter phage Makai | Arthrobacter phage Makai unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57972 | 50.293 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN284897 | Arthrobacter phage NapoleonB | Arthrobacter phage NapoleonB unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57846 | 45.267 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234229 | Arthrobacter phage StarLord | Arthrobacter phage StarLord Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54504 | 51.653 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234213 | Gordonia phage Schiebs | Gordonia phage Schiebs Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14646 | 70.606 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MN234210 | Arthrobacter phage Michelle | Arthrobacter phage Michelle Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54509 | 51.692 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234202 | Arthrobacter phage Salk | Arthrobacter phage Salk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54492 | 51.753 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234194 | Arthrobacter phage Loretta | Arthrobacter phage Loretta unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56953 | 50.742 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234188 | Mycobacterium phage Anthony | Mycobacterium phage Anthony unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52202 | 61.367 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234186 | Arthrobacter phage Egad | Arthrobacter phage Egad Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54517 | 51.635 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234185 | Mycobacterium phage Lemuria | Mycobacterium phage Lemuria Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44998 | 68.85 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN234183 | Mycobacterium phage Antsirabe | Mycobacterium phage Antsirabe Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44739 | 69.007 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN234175 | Arthrobacter phage Stayer | Arthrobacter phage Stayer Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54507 | 51.713 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234173 | Arthrobacter phage Sporto | Arthrobacter phage Sporto Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54780 | 51.581 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MN234168 | Arthrobacter phage Ingrid | Arthrobacter phage Ingrid unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56953 | 50.74 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234166 | Gordonia phage ASerpRocky | Gordonia phage ASerpRocky unclassified Demosthenesvirus Demosthenesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74355 | 59.211 | Demosthenesvirus | Unclassified | Siphoviridae | Gordonia |
| MN224565 | Streptomyces phage Intolerant | Streptomyces phage Intolerant Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53909 | 68.367 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MN224564 | Streptomyces phage Araceli | Streptomyces phage Araceli Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55045 | 68.211 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MN234219 | Mycobacterium phage Mercurio | Mycobacterium phage Mercurio Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44877 | 69.029 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN310544 | Mycobacterium phage Doug | Mycobacterium phage Doug unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58397 | 61.145 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967398 | Arthrobacter phage JEGGS | Arthrobacter phage JEGGS unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58287 | 45.091 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MK967397 | Mycobacterium phage Mahavrat | Mycobacterium phage Mahavrat unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55945 | 61.285 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967386 | Streptomyces phage Esketit | Streptomyces phage Esketit unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54664 | 59.493 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MN096364 | Mycobacterium phage Tomaszewski | Mycobacterium phage Tomaszewski unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74707 | 63.01 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096361 | Mycobacterium phage Gator | Mycobacterium phage Gator unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76351 | 62.936 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK937596 | Microbacterium phage PhillyPhilly | Microbacterium phage PhillyPhilly Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62869 | 62.711 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK937593 | Mycobacterium phage Flypotenuse | Mycobacterium phage Flypotenuse unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75248 | 62.925 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| MK937592 | Mycobacterium phage Phrappuccino | Mycobacterium phage Phrappuccino Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136361 | 67.378 | Unclassified | Unclassified | Myoviridae | Mycobacterium |
| MK919478 | Gordonia phage Ziko | Gordonia phage Ziko unclassified Ronaldovirus Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68860 | 50.125 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MK919474 | Microbacterium phage Lupine | Microbacterium phage Lupine Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62533 | 62.668 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK919473 | Arthrobacter phage Xenomorph | Arthrobacter phage Xenomorph unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58690 | 45.372 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MK977714 | Corynebacterium phage Bran | Corynebacterium phage Bran Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44484 | 67.64 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| MK977712 | Corynebacterium phage Lederberg | Corynebacterium phage Lederberg unclassified Samwavirus Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45644 | 68.072 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MK977701 | Mycobacterium phage Cracklewink | Mycobacterium phage Cracklewink Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76991 | 67.351 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK801723 | Gordonia phage Coeur | Gordonia phage Coeur Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16223 | 67.91 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK737944 | Microbacterium phage Warren | Microbacterium phage Warren unclassified Appavirus Appavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38493 | 67.94 | Appavirus | Unclassified | Siphoviridae | Microbacterium |
| MK737942 | Microbacterium phage Sucha | Microbacterium phage Sucha unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41826 | 66.148 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MK814759 | Gordonia phage Reyja | Gordonia phage Reyja Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41500 | 67.373 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK814756 | Microbacterium phage DizzyRudy | Microbacterium phage DizzyRudy Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55815 | 56.442 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK686070 | Streptomyces phage PherryCruz | Streptomyces phage PherryCruz unclassified Scapunavirus Scapunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43736 | 60.984 | Scapunavirus | Unclassified | Siphoviridae | Streptomyces |
| MK494129 | Mycobacterium phage AvatarAhPeg | Mycobacterium phage AvatarAhPeg unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51637 | 63.842 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494120 | Mycobacterium phage Piper2020 | Mycobacterium phage Piper2020 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57703 | 62.676 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494118 | Mycobacterium phage Cornucopia | Mycobacterium phage Cornucopia unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55422 | 61.394 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494115 | Mycobacterium phage Bumblebee11 | Mycobacterium phage Bumblebee11 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51273 | 63.704 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494101 | Mycobacterium phage Connomayer | Mycobacterium phage Connomayer unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51371 | 63.89 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494094 | Mycobacterium phage DirtyDunning | Mycobacterium phage DirtyDunning unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51418 | 63.976 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524500 | Mycobacterium phage Kevin1 | Mycobacterium phage Kevin1 unclassified Buttersvirus Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41988 | 66.231 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MK524495 | Rhodococcus phage Belenaria | Rhodococcus phage Belenaria unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46538 | 58.591 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MK524487 | Rhodococcus phage Espica | Rhodococcus phage Espica unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46537 | 58.577 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MK433279 | Mycobacterium phage Kareem | Mycobacterium phage Kareem unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.562 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK433277 | Mycobacterium phage Renaissance | Mycobacterium phage Renaissance unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41879 | 66.566 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK433273 | Mycobacterium phage Daishi | Mycobacterium phage Daishi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49750 | 64.249 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359325 | Mycobacterium phage Eros | Mycobacterium phage Eros unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.861 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359323 | Mycobacterium phage Baby16 | Mycobacterium phage Baby16 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51383 | 63.86 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359322 | Mycobacterium phage Datway | Mycobacterium phage Datway unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51303 | 63.675 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK376963 | Microbacterium phage Alleb | Microbacterium phage Alleb Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62684 | 64.567 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MK376961 | Gordonia phage Asapag | Gordonia phage Asapag Gordonia virus Asapag Getalongvirus Langleyhallvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55119 | 63.071 | Getalongvirus | Langleyhallvirinae | Siphoviridae | Gordonia |
| MK279902 | Mycobacterium phage Citius | Mycobacterium phage Citius unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51393 | 63.886 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279898 | Mycobacterium phage Cici | Mycobacterium phage Cici unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51287 | 63.706 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH910041 | Arthrobacter phage Seahorse | Arthrobacter phage Seahorse Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56481 | 62.697 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MK016501 | Mycobacterium phage Nebkiss | Mycobacterium phage Nebkiss unclassified Gaiavirus Gaiavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85613 | 56.49 | Gaiavirus | Unclassified | Siphoviridae | Mycobacterium |
| MH926061 | Corynebacterium phage Troy | Corynebacterium phage Troy unclassified Samwavirus Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 68.143 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MH834628 | Arthrobacter phage Sputnik | Arthrobacter phage Sputnik Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14927 | 58.042 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834625 | Arthrobacter phage Richie | Arthrobacter phage Richie Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53867 | 62.706 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834623 | Arthrobacter phage Peas | Arthrobacter phage Peas Arthrobacter virus Peas Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44103 | 64.712 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834614 | Arthrobacter phage Judy | Arthrobacter phage Judy Arthrobacter virus Judy Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43440 | 65.168 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834611 | Arthrobacter phage Eileen | Arthrobacter phage Eileen Arthrobacter virus Eileen Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41165 | 65.126 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834605 | Arthrobacter phage Constance | Arthrobacter phage Constance Arthrobacter virus Constance Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43706 | 65.128 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834603 | Arthrobacter phage Bridgette | Arthrobacter phage Bridgette Arthrobacter virus Bridgette Bridgettevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43113 | 65.089 | Bridgettevirus | Unclassified | Siphoviridae | Arthrobacter |
| MH834597 | Arthrobacter phage Atraxa | Arthrobacter phage Atraxa Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14927 | 58.036 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834594 | Arthrobacter phage Adaia | Arthrobacter phage Adaia Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15840 | 56.111 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH834598 | Arthrobacter phage Auxilium | Arthrobacter phage Auxilium Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49447 | 62.584 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MH727560 | Corynebacterium phage SamW | Corynebacterium phage SamW Corynebacterium virus SamW Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 68.134 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MH651170 | Mycobacterium phage ChampagnePapi | Mycobacterium phage ChampagnePapi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51811 | 63.903 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH479925 | Gordonia phage Ronaldo | Gordonia phage Ronaldo Gordonia virus Ronaldo Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68389 | 50.163 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MH479913 | Gordonia phage Fryberger | Gordonia phage Fryberger Gordonia virus Fryberger Ronaldovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67234 | 50.178 | Ronaldovirus | Unclassified | Siphoviridae | Gordonia |
| MH576949 | Mycobacterium phage BreSam8 | Mycobacterium phage BreSam8 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50843 | 64.023 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536823 | Microbacterium phage Musetta | Microbacterium phage Musetta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63604 | 61.701 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH536813 | Mycobacterium phage AugsMagnumOpus | Mycobacterium phage AugsMagnumOpus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48545 | 64.173 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH509441 | Mycobacterium phage Acolyte | Mycobacterium phage Acolyte unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52668 | 63.346 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH479911 | Mycobacterium phage Eapen | Mycobacterium phage Eapen unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 63.889 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH479909 | Mycobacterium phage Commander | Mycobacterium phage Commander unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51369 | 63.879 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450120 | Arthrobacter phage KeaneyLin | Arthrobacter phage KeaneyLin unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57847 | 45.394 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MH399787 | Mycobacterium phage Rubeelu | Mycobacterium phage Rubeelu unclassified Buttersvirus Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41491 | 65.822 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH399779 | Microbacterium phage Jacko | Microbacterium phage Jacko Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61421 | 64.989 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH371108 | Microbacterium phage Fork | Microbacterium phage Fork Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62090 | 61.804 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH371109 | Microbacterium phage Lyell | Microbacterium phage Lyell Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62716 | 61.595 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH316570 | Mycobacterium phage Silvafighter | Mycobacterium phage Silvafighter unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43243 | 66.26 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MH316569 | Rhodococcus phage Shuman | Rhodococcus phage Shuman unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46544 | 58.648 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271319 | Rhodococcus phage UhSalsa | Rhodococcus phage UhSalsa unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46539 | 58.577 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271316 | Microbacterium phage Tandem | Microbacterium phage Tandem Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63128 | 64.703 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271314 | Rhodococcus phage Swann | Rhodococcus phage Swann unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.614 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271311 | Rhodococcus phage Rasputin | Rhodococcus phage Rasputin unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46568 | 58.798 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271310 | Microbacterium phage Pioneer3 | Microbacterium phage Pioneer3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62954 | 64.641 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271309 | Rhodococcus phage Phrankenstein | Rhodococcus phage Phrankenstein unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46540 | 58.573 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271307 | Microbacterium phage OlinDD | Microbacterium phage OlinDD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63123 | 64.601 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271306 | Rhodococcus phage Nancinator | Rhodococcus phage Nancinator unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45936 | 58.638 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271300 | Microbacterium phage Hortus1 | Microbacterium phage Hortus1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63119 | 64.592 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH271299 | Rhodococcus phage Gollum | Rhodococcus phage Gollum unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46535 | 58.603 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271297 | Rhodococcus phage Erik | Rhodococcus phage Erik unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46429 | 58.55 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271294 | Rhodococcus phage Bryce | Rhodococcus phage Bryce unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46347 | 58.789 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271293 | Rhodococcus phage Bradshaw | Rhodococcus phage Bradshaw unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46606 | 58.561 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH271291 | Rhodococcus phage Alpacados | Rhodococcus phage Alpacados unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46493 | 58.852 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MH153801 | Microbacterium phage Count | Microbacterium phage Count Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78922 | 51.375 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH153800 | Microbacterium phage Camille | Microbacterium phage Camille Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53097 | 56.308 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH153799 | Microbacterium phage Appa | Microbacterium phage Appa Microbacterium virus Appa Appavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38684 | 68.041 | Appavirus | Unclassified | Siphoviridae | Microbacterium |
| MH001451 | Mycobacterium phage Nairb | Mycobacterium phage Nairb unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42393 | 66.2 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925349 | Mycobacterium phage Mendokysei | Mycobacterium phage Mendokysei unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43511 | 66.358 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF279418 | Mycobacterium phage Anubis | Mycobacterium phage Anubis unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50854 | 64.003 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU647628 | Arthrobacter phage Mudcat | Arthrobacter phage Mudcat Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59443 | 45.104 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| KU963252 | Gordonia phage Schnabeltier | Gordonia phage Schnabeltier Gordonia virus Schnabeltier Schnabeltiervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46895 | 67.037 | Schnabeltiervirus | Unclassified | Siphoviridae | Gordonia |
| MG099936 | Mycobacterium phage Andies | Mycobacterium phage Andies unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43779 | 66.171 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MG099942 | Mycobacterium phage Blackmoor | Mycobacterium phage Blackmoor unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.863 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919498 | Mycobacterium phage Cindaradix | Mycobacterium phage Cindaradix unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51575 | 64.012 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF919496 | Mycobacterium phage Cerulean | Mycobacterium phage Cerulean unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51809 | 63.927 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF324903 | Rhodococcus phage AppleCloud | Rhodococcus phage AppleCloud unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46389 | 58.663 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF324906 | Arthrobacter phage Cheesy | Arthrobacter phage Cheesy unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58739 | 45.248 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF324905 | Rhodococcus phage Alatin | Rhodococcus phage Alatin unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46673 | 58.651 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF324904 | Rhodococcus phage RexFury | Rhodococcus phage RexFury unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46627 | 58.558 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF324902 | Rhodococcus phage Krishelle | Rhodococcus phage Krishelle unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46985 | 58.542 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF324901 | Rhodococcus phage Naiad | Rhodococcus phage Naiad unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46619 | 58.646 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF324900 | Rhodococcus phage StCroix | Rhodococcus phage StCroix unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46619 | 58.646 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF189179 | Rhodococcus phage Yoncess | Rhodococcus phage Yoncess unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46353 | 58.799 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF189175 | Arthrobacter phage Tribby | Arthrobacter phage Tribby unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59084 | 45.225 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189174 | Arthrobacter phage Nason | Arthrobacter phage Nason unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58059 | 45.149 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189173 | Arthrobacter phage Heisenberger | Arthrobacter phage Heisenberger unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58208 | 45.126 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189172 | Arthrobacter phage Elsa | Arthrobacter phage Elsa unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58059 | 45.152 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189171 | Arthrobacter phage Correa | Arthrobacter phage Correa unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57401 | 45.233 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF189170 | Arthrobacter phage Arcadia | Arthrobacter phage Arcadia unclassified Mudcatvirus Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58059 | 45.152 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF537628 | Rhodococcus phage Bonanza | Rhodococcus phage Bonanza unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46932 | 58.847 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MF155936 | Mycobacterium phage ZenTime222 | Mycobacterium phage ZenTime222 unclassified Bernalvirus Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43344 | 66.014 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF373842 | Rhodococcus phage Jester | Rhodococcus phage Jester unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46314 | 58.695 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KY549154 | Rhodococcus phage BobbyDazzler | Rhodococcus phage BobbyDazzler unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46641 | 58.809 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KY549153 | Rhodococcus phage AngryOrchard | Rhodococcus phage AngryOrchard unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46597 | 58.813 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KY204247 | Mycobacterium phage Funston | Mycobacterium phage Funston unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.992 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204246 | Mycobacterium phage Clarenza | Mycobacterium phage Clarenza unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.914 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204245 | Mycobacterium phage Camperdownii | Mycobacterium phage Camperdownii unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51131 | 63.944 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY204244 | Mycobacterium phage BubbleTrouble | Mycobacterium phage BubbleTrouble unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51397 | 63.912 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083063 | Mycobacterium phage Eris | Mycobacterium phage Eris unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51386 | 63.977 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083060 | Mycobacterium phage Bruiser | Mycobacterium phage Bruiser unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.902 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083059 | Mycobacterium phage Achebe | Mycobacterium phage Achebe unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51433 | 63.673 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY083058 | Mycobacterium phage Abdiel | Mycobacterium phage Abdiel unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51381 | 63.854 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX817175 | Mycobacterium phage Albee | Mycobacterium phage Albee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.906 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX808130 | Mycobacterium phage Broseidon | Mycobacterium phage Broseidon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51374 | 63.859 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX712237 | Rhodococcus phage Partridge | Rhodococcus phage Partridge unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46962 | 58.811 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KX712236 | Rhodococcus phage Yogi | Rhodococcus phage Yogi unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46930 | 58.847 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KX620752 | Propionibacterium phage E1 | Propionibacterium phage E1 unclassified Anatolevirus Anatolevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35209 | 64.421 | Anatolevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX620749 | Propionibacterium phage B3 | Propionibacterium phage B3 Anatolevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35948 | 64.424 | Anatolevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX620748 | Propionibacterium phage Anatole | Propionibacterium phage Anatole Anatolevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35284 | 64.431 | Anatolevirus | Unclassified | Siphoviridae | Propionibacterium |
| KX611788 | Rhodococcus phage Harlequin | Rhodococcus phage Harlequin unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46383 | 58.759 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KX557234 | Mycobacterium phage Florean | Mycobacterium phage Florean unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51371 | 63.863 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX550082 | Rhodococcus phage Natosaleda | Rhodococcus phage Natosaleda unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46527 | 58.579 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KX557279 | Gordonia phage Hedwig | Gordonia phage Hedwig Gordonia virus Hedwig Hedwigvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44536 | 67.193 | Hedwigvirus | Unclassified | Siphoviridae | Gordonia |
| KX523125 | Mycobacterium phage BuzzBuzz | Mycobacterium phage BuzzBuzz Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50897 | 64.214 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU935730 | Mycobacterium phage Pipsqueaks | Mycobacterium phage Pipsqueaks Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43679 | 66.277 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KU935727 | Mycobacterium phage Panchino | Mycobacterium phage Panchino Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43516 | 65.948 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KU935726 | Mycobacterium phage Xerxes | Mycobacterium phage Xerxes unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43698 | 66.262 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| KU160671 | Arthrobacter phage Vulture | Arthrobacter phage Vulture unclassified Korravirus Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43336 | 61.563 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160648 | Arthrobacter phage HunterDalle | Arthrobacter phage HunterDalle Korravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43336 | 61.561 | Korravirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160642 | Arthrobacter phage Circum | Arthrobacter phage Circum Mudcatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58353 | 45.175 | Mudcatvirus | Unclassified | Siphoviridae | Arthrobacter |
| KT959213 | Rhodococcus phage TWAMP | Rhodococcus phage TWAMP unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.546 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KP027198 | Mycobacterium phage Gadost | Mycobacterium phage Gadost unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51376 | 63.966 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM233455 | Mycobacterium phage Farber | Mycobacterium phage Farber unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49190 | 64.151 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ510413 | Mycobacterium phage Bernal13 | Mycobacterium phage Bernal13 Bernalvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42392 | 66.199 | Bernalvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF841475 | Mycobacterium phage BellusTerra | Mycobacterium phage BellusTerra unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51236 | 63.857 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC576783 | Mycobacterium phage Butters | Mycobacterium phage Butters Buttersvirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41491 | 65.838 | Buttersvirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| JX042578 | Mycobacteriophage ElTiger69 | Mycobacteriophage ElTiger69 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51505 | 59.784 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ809701 | Mycobacterium phage Flux | Mycobacterium phage Flux Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51370 | 63.915 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN256079 | Mycobacterium phage Charlie | Mycobacterium phage Charlie Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43036 | 66.331 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| JN408459 | Mycobacterium virus Cuco | Mycobacterium virus Cuco Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50965 | 60.857 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN083853 | Mycobacterium phage Airmid | Mycobacterium phage Airmid unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51241 | 59.96 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT521996 | Mycobacterium phage Beckerton | Mycobacterium phage Beckerton unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68882 | 57.366 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| MT024859 | Gordonia phage DumpsterDude | Gordonia phage DumpsterDude Gordonia virus DumpsterDude Ruthyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52018 | 66.61 | Ruthyvirus | Unclassified | Siphoviridae | Gordonia |
| MN908688 | Mycobacterium phage Cornie | Mycobacterium phage Cornie Mycobacterium virus Cornie Cornievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56391 | 61.386 | Cornievirus | Unclassified | Siphoviridae | Mycobacterium |
| MN369747 | Microbacterium phage Wesak | Microbacterium phage Wesak unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53789 | 56.748 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK967377 | Gordonia phage MagicMan | Gordonia phage MagicMan unclassified Schnabeltiervirus Schnabeltiervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47598 | 67.135 | Schnabeltiervirus | Unclassified | Siphoviridae | Gordonia |
| MK977715 | Corynebacterium phage Adelaide | Corynebacterium phage Adelaide Corynebacterium virus Adelaide Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44104 | 67.982 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MK937613 | Corynebacterium phage StAB | Corynebacterium phage StAB Corynebacterium virus StAB Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45751 | 67.861 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MK977710 | Corynebacterium phage Stiles | Corynebacterium phage Stiles Corynebacterium virus Stiles Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42846 | 67.663 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MK977706 | Corynebacterium phage Dina | Corynebacterium phage Dina Corynebacterium virus Dina Samwavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44381 | 67.95 | Samwavirus | Unclassified | Siphoviridae | Corynebacterium |
| MK894438 | Microbacterium phage LilyLou | Microbacterium phage LilyLou unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54388 | 59.995 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK757449 | Microbacterium phage Akoni | Microbacterium phage Akoni Microbacterium virus Akoni Akonivirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54307 | 60.14 | Akonivirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK308638 | Microbacterium phage ArMaWen | Microbacterium phage ArMaWen unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53939 | 59.936 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MK284521 | Gordonia phage Kiko | Gordonia phage Kiko Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44268 | 66.493 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MK016500 | Arthrobacter phage Mendel | Arthrobacter phage Mendel Arthobacter virus Mendel Anjalivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19428 | 59.826 | Anjalivirus | Unclassified | Podoviridae | Arthrobacter |
| MK016490 | Arthrobacter phage Anjali | Arthrobacter phage Anjali Arthrobacter virus Anjali Anjalivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19679 | 59.373 | Anjalivirus | Unclassified | Podoviridae | Arthrobacter |
| MG962376 | Mycobacterium phage Thumb | Mycobacterium phage Thumb unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68408 | 57.502 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234161 | Gordonia phage Toast | Gordonia phage Toast Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49604 | 66.785 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MK801733 | Gordonia phage Chelms | Gordonia phage Chelms Gordonia virus Chelms Montyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76261 | 58.583 | Montyvirus | Unclassified | Siphoviridae | Gordonia |
| MK112550 | Streptomyces phage Olicious | Streptomyces phage Olicious unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46139 | 59.668 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MH271315 | Rhodococcus phage Takoda | Rhodococcus phage Takoda Rhodococcus virus Takoda Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46807 | 58.737 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| MG757156 | Mycobacterium phage Cuke | Mycobacterium phage Cuke Mycobacterium virus Cuke Cukevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68869 | 49.08 | Cukevirus | Unclassified | Siphoviridae | Mycobacterium |
| MG663586 | Streptomyces phage ZooBear | Streptomyces phage ZooBear unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46135 | 59.653 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MG663585 | Streptomyces phage ToriToki | Streptomyces phage ToriToki unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46077 | 59.674 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MG663584 | Streptomyces phage Romero | Streptomyces phage Romero unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46079 | 59.739 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MG663583 | Streptomyces phage Percastrophe | Streptomyces phage Percastrophe unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45999 | 59.695 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MG663582 | Streptomyces phage HaugeAnator | Streptomyces phage HaugeAnator unclassified Immanueltrevirus Immanueltrevirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46135 | 59.629 | Immanueltrevirus | Beephvirinae | Podoviridae | Streptomyces |
| MG518519 | Streptomyces phage Manuel | Streptomyces phage Manuel Streptomyces virus Manuel Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45177 | 60.095 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| MG515223 | Streptomyces phage WRightOn | Streptomyces phage WRightOn Streptomyces virus WRightOn Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45221 | 60.313 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| MF324898 | Rhodococcus phage Hiro | Rhodococcus phage Hiro Rhodococcus virus Hiro Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46854 | 58.719 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KX557275 | Gordonia phage CarolAnn | Gordonia phage CarolAnn Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54167 | 66.902 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KU998248 | Gordonia phage Utz | Gordonia phage Utz Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49768 | 67.652 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| KU963254 | Gordonia phage Obliviate | Gordonia phage Obliviate Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49286 | 67.53 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT740307 | Pseudomonas phage phiK7A1 | Pseudomonas phage phiK7A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98780 | 48.79 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MT884014 | Escherichia phage vb_EcoS_bov22_1 | Escherichia phage vb_EcoS_bov22_1 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44612 | 54.463 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT496971 | Escherichia phage Mt1B1_P10 | Escherichia phage Mt1B1_P10 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45359 | 45.061 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MT747435 | Enterococcus phage AE4_2 | Enterococcus phage AE4_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37561 | 37.621 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT747434 | Enterococcus phage ZEF1 | Enterococcus phage ZEF1 unclassified Copernicusvirus Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18454 | 32.86 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MT833282 | Escherichia coli phage vB_EcoM_Shy | Escherichia coli phage vB_EcoM_Shy unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88760 | 38.994 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MT936332 | Streptomyces phage phiRKBJ001 | Streptomyces phage phiRKBJ001 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69373 | 63.718 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT939492 | Xanthomonas phage Xoo-sp14 | Xanthomonas phage Xoo-sp14 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 232104 | 58.013 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT366946 | Bacillus phage Hyb1phi3Ts-SPbeta | Bacillus phage Hyb1phi3Ts-SPbeta unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131755 | 34.746 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| MT241607 | Achromobacter phage AMA2 | Achromobacter phage AMA2 unclassified Steinhofvirus Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45901 | 54.48 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| MN604053 | Escherichia phage E21 | Escherichia phage E21 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58536 | 46.898 | Unclassified | Unclassified | Podoviridae | Escherichia |
| MT152150 | Pseudomonas phage 8P | Pseudomonas phage 8P Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63753 | 64.355 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT711528 | Escherichia phage vB_EcoS_fTaEco03 | Escherichia phage vB_EcoS_fTaEco03 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41972 | 50.865 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711527 | Escherichia phage vB_EcoS_fTaEco01 | Escherichia phage vB_EcoS_fTaEco01 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41273 | 50.893 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711526 | Escherichia phage vB_EcoS_fPoEco01 | Escherichia phage vB_EcoS_fPoEco01 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44391 | 50.657 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711525 | Escherichia phage vB_EcoS_fKuEco01 | Escherichia phage vB_EcoS_fKuEco01 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40825 | 50.957 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711524 | Escherichia phage vB_EcoS_fFiEco03 | Escherichia phage vB_EcoS_fFiEco03 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43947 | 50.488 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711523 | Escherichia phage vB_EcoS_fFiEco02 | Escherichia phage vB_EcoS_fFiEco02 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44723 | 50.869 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MT711370 | Methanocaldococcus fervens tailed virus 1 | Methanocaldococcus fervens tailed virus 1 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 31202 | 33.28 | Unclassified | Unclassified | Unclassified | Methanocaldococcus |
| MT741944 | Acinetobacter phage vB_AbaP_APK81 | Acinetobacter phage vB_AbaP_APK81 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42233 | 39.325 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT684587 | Microbacterium phage Fede | Microbacterium phage Fede Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54911 | 59.988 | Unclassified | Eekayvirinae | Podoviridae | Microbacterium |
| MT843276 | Serratia phage vB_SspS_OS31 | Serratia phage vB_SspS_OS31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42280 | 51.154 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MT843275 | Serratia phage vB_SspM_BZS1 | Serratia phage vB_SspM_BZS1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44995 | 51.59 | Unclassified | Unclassified | Myoviridae | Serratia |
| MT664984 | Xanthomonas phage Xp12 | Xanthomonas phage Xp12 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63783 | 68.17 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| MT856905 | Enterococcus phage vB_EfaM_A2 | Enterococcus phage vB_EfaM_A2 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149431 | 37.061 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MT657344 | Microbacterium phage Kauala | Microbacterium phage Kauala unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39378 | 64.419 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MT657343 | Microbacterium phage GaeCeo | Microbacterium phage GaeCeo unclassified Schubertvirus Schubertvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40168 | 63.426 | Schubertvirus | Unclassified | Siphoviridae | Microbacterium |
| MT902335 | Pasteurella phage vB_PmuP_Pa7 | Pasteurella phage vB_PmuP_Pa7 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37402 | 40.877 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| MT855965 | Microcystis phage vB_MaeS-yong1 | Microcystis phage vB_MaeS-yong1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43665 | 66.735 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MT542512 | Escherichia phage PhiV-1 | Escherichia phage PhiV-1 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39461 | 48.686 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| MT520979 | Enterococcus phage vB_EfaS-271 | Enterococcus phage vB_EfaS-271 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40197 | 34.943 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MN270279 | Streptococcus phage phi-SsuZKB4_rum | Streptococcus phage phi-SsuZKB4_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110533 | 38.903 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270278 | Streptococcus phage phi-SsuYZDH5_rum | Streptococcus phage phi-SsuYZDH5_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47914 | 39.884 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270277 | Streptococcus phage phi-SsuYTJ2_rum | Streptococcus phage phi-SsuYTJ2_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53191 | 39.943 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270276 | Streptococcus phage phi-SsuSSJ28_rum | Streptococcus phage phi-SsuSSJ28_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62100 | 39.027 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270275 | Streptococcus phage phi-SsuSSJ27_rum | Streptococcus phage phi-SsuSSJ27_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47240 | 40.538 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270274 | Streptococcus phage phi-SsuSH0918_comEC | Streptococcus phage phi-SsuSH0918_comEC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49639 | 39.906 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270273 | Streptococcus phage phi-SsuSC05017_rum | Streptococcus phage phi-SsuSC05017_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57591 | 39.636 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270272 | Streptococcus phage phi-SsuNJ5_rum | Streptococcus phage phi-SsuNJ5_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53801 | 39.711 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270271 | Streptococcus phage phi-SsuNJ2_rum | Streptococcus phage phi-SsuNJ2_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53896 | 39.741 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270270 | Streptococcus phage phi-SsuJS7_SSU0237 | Streptococcus phage phi-SsuJS7_SSU0237 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47090 | 40.529 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270269 | Streptococcus phage phi-SsuHCJ3_rum | Streptococcus phage phi-SsuHCJ3_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56061 | 39.229 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270268 | Streptococcus phage phi-SsuHCJ31_comEC | Streptococcus phage phi-SsuHCJ31_comEC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58334 | 39.452 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270267 | Streptococcus phage phi-SsuFJZZ39_rum | Streptococcus phage phi-SsuFJZZ39_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73944 | 38.789 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270266 | Streptococcus phage phi-SsuFJZZ32_rum | Streptococcus phage phi-SsuFJZZ32_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73944 | 38.789 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270265 | Streptococcus phage phi-SsuFJSM8_rum | Streptococcus phage phi-SsuFJSM8_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33641 | 39.951 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270264 | Streptococcus phage phi-SsuFJSM7_rum | Streptococcus phage phi-SsuFJSM7_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49769 | 40.198 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270263 | Streptococcus phage phi-SsuFJSM5_rum | Streptococcus phage phi-SsuFJSM5_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49769 | 40.198 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270262 | Streptococcus phage phi-SsuFJNP9_rum | Streptococcus phage phi-SsuFJNP9_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56838 | 39.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270261 | Streptococcus phage phi-SsuFJNP8_rum | Streptococcus phage phi-SsuFJNP8_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77200 | 39.383 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270260 | Streptococcus phage phi-SsuFJNP3_rum | Streptococcus phage phi-SsuFJNP3_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56838 | 39.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270259 | Streptococcus phage phi-SgaBSJ31_rum | Streptococcus phage phi-SgaBSJ31_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106491 | 38.702 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN270258 | Streptococcus phage phi-SgaBSJ27_rum | Streptococcus phage phi-SgaBSJ27_rum Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110666 | 38.618 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MF374379 | Escherichia phage DN1 | Escherichia phage DN1 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41559 | 53.081 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MT806187 | Bacteroides phage F2 | Bacteroides phage F2 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10891 | 51.997 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT806186 | Bacteroides phage F2 | Bacteroides phage F2 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29762 | 51.65 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT806185 | Bacteroides phage F4 | Bacteroides phage F4 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40112 | 51.802 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT897907 | Mycobacterium phage Grif | Mycobacterium phage Grif unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50957 | 63.979 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT897904 | Mycobacterium phage Royals2015 | Mycobacterium phage Royals2015 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57356 | 61.708 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT741943 | Acinetobacter phage Meroveus | Acinetobacter phage Meroveus unclassified Lazarusvirus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167683 | 36.719 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MT684602 | Mycobacterium phage Rope | Mycobacterium phage Rope unclassified Papyrusvirus Papyrusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71003 | 56.038 | Papyrusvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684600 | Streptomyces phage TieDye | Streptomyces phage TieDye unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55031 | 59.516 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MT684597 | Mycobacterium phage Mandlovu | Mycobacterium phage Mandlovu unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55413 | 61.44 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684596 | Streptomyces phage Zeigle | Streptomyces phage Zeigle unclassified Manuelvirus Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44859 | 60.318 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| MT684595 | Mycobacterium phage Mova | Mycobacterium phage Mova unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57599 | 61.253 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684593 | Mycobacterium phage Moonbeam | Mycobacterium phage Moonbeam unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57984 | 61.1 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT684589 | Streptomyces phage Maya | Streptomyces phage Maya unclassified Rimavirus Rimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55713 | 59.501 | Rimavirus | Unclassified | Siphoviridae | Streptomyces |
| MT684588 | Mycobacterium phage Snekmaggedon | Mycobacterium phage Snekmaggedon unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41278 | 66.246 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MT684586 | Mycobacterium phage Gandalph | Mycobacterium phage Gandalph unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54974 | 61.369 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT758689 | Lactobacillus phage Lbab1 | Lactobacillus phage Lbab1 unclassified Herelleviridae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140488 | 37.351 | Unclassified | Unclassified | Herelleviridae | Lactobacillus |
| MT758688 | Mycobacterium phage CELFI | Mycobacterium phage CELFI unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77086 | 59.033 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT670422 | Pseudomonas phage phiPsa397 | Pseudomonas phage phiPsa397 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98951 | 47.725 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT670421 | Pseudomonas phage phiPsa381 | Pseudomonas phage phiPsa381 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98800 | 47.809 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT670420 | Pseudomonas phage phiPsa347 | Pseudomonas phage phiPsa347 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99689 | 47.702 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT670419 | Pseudomonas phage phiPsa315 | Pseudomonas phage phiPsa315 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98739 | 47.954 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT670418 | Pseudomonas phage phiPsa300 | Pseudomonas phage phiPsa300 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99272 | 47.682 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT670417 | Pseudomonas phage phiPsa267 | Pseudomonas phage phiPsa267 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100180 | 47.714 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MT764845 | Ruegeria phage Tedan | Ruegeria phage Tedan Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64084 | 54.425 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MT679222 | Proteus phage 2 | Proteus phage 2 unclassified Novosibvirus Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102041 | 38.328 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MT679223 | Proteus phage 1 | Proteus phage 1 unclassified Novosibvirus Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102298 | 38.313 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MT679221 | Proteus phage 7 | Proteus phage 7 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 242444 | 48.503 | Seoulvirus | Unclassified | Myoviridae | Proteus |
| MT419366 | Serratia phage SALSA | Serratia phage SALSA unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39933 | 50.542 | Teetrevirus | Studiervirinae | Autographiviridae | Serratia |
| MT813142 | Klebsiella phage NL_ZS_3 | Klebsiella phage NL_ZS_3 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38741 | 50.732 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT813141 | Klebsiella phage NL_ZS_2 | Klebsiella phage NL_ZS_2 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40222 | 52.792 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT813140 | Klebsiella phage NL_ZS_1 | Klebsiella phage NL_ZS_1 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40428 | 52.968 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT366948 | Bacillus phage Hyb3phi3Ts-SPbeta | Bacillus phage Hyb3phi3Ts-SPbeta unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127428 | 35.06 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| MT366947 | Bacillus phage Hyb2phi3Ts-SPbeta | Bacillus phage Hyb2phi3Ts-SPbeta unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128701 | 35.032 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| MT366945 | Bacillus phage phi3Ts | Bacillus phage phi3Ts unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127835 | 34.963 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| MN715150 | Citrobacter phage NS1 | Citrobacter phage NS1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38900 | 50.18 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| BK013338 | Haloterrigena jeotgali icosahedral virus 1 | Haloterrigena jeotgali icosahedral virus 1 Yingchengvirus HJIV1 Yingchengvirus Simuloviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 17189 | 63.989 | Yingchengvirus | Unclassified | Simuloviridae | Haloterrigena |
| BK013337 | Natrinema versiforme icosahedral virus 1 | Natrinema versiforme icosahedral virus 1 Yingchengvirus NVIV1 Yingchengvirus Simuloviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 18925 | 63.53 | Yingchengvirus | Unclassified | Simuloviridae | Natrinema |
| MT360682 | Vibrio virus 2019VC1 | Vibrio virus 2019VC1 Jhansiroadvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47778 | 44.947 | Jhansiroadvirus | Unclassified | Drexlerviridae | Vibrio |
| KX198613 | Pseudomonas phage AN14 | Pseudomonas phage AN14 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60973 | 64.514 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MT668624 | Listeria phage LP-Mix_6.2 | Listeria phage LP-Mix_6.2 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138470 | 35.876 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MT668623 | Listeria phage LP-Mix_6.1 | Listeria phage LP-Mix_6.1 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136804 | 35.929 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MT657334 | Microbacterium phage Stormbreaker | Microbacterium phage Stormbreaker unclassified Tinytimothyvirus Tinytimothyvirus Eekayvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54050 | 60.035 | Tinytimothyvirus | Eekayvirinae | Podoviridae | Microbacterium |
| MT767883 | Vibrio phage Saratov-15 | Vibrio phage Saratov-15 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51082 | 42.755 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MT909815 | Enterococcus phage iF6 | Enterococcus phage iF6 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156592 | 37.061 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MT012730 | Salmonella phage vB_SenM-1 | Salmonella phage vB_SenM-1 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52471 | 45.995 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT004791 | Salmonella phage vB_SenS-3 | Salmonella phage vB_SenS-3 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 119586 | 39.969 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT723940 | Mycobacterium phage Ellie | Mycobacterium phage Ellie unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61945 | 67.042 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MT723939 | Mycobacterium phage Bombshell | Mycobacterium phage Bombshell unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51372 | 63.893 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT723932 | Mycobacterium phage Schnauzer | Mycobacterium phage Schnauzer unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43700 | 66.256 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MT711890 | Pseudomonas phage PlaquesPlease | Pseudomonas phage PlaquesPlease unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41456 | 58.071 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MT711889 | Pseudomonas phage Waldo5 | Pseudomonas phage Waldo5 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41195 | 57.728 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MT711888 | Pseudomonas phage Dolphis | Pseudomonas phage Dolphis Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93507 | 63.705 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MT711887 | Pseudomonas phage Stalingrad | Pseudomonas phage Stalingrad unclassified Troedvirus Troedvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40471 | 57.859 | Troedvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MT639654 | Mycobacterium phage Jerm2 | Mycobacterium phage Jerm2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53215 | 63.567 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT639650 | Gordonia phage Ohgeesy | Gordonia phage Ohgeesy unclassified Baxtervirus Baxtervirus Nymbaxtervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53473 | 66.267 | Baxtervirus | Nymbaxtervirinae | Siphoviridae | Gordonia |
| MT639644 | Mycobacterium phage MaryBeth | Mycobacterium phage MaryBeth unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51854 | 63.534 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT639639 | Gordonia phage LittleFella | Gordonia phage LittleFella unclassified Terapinvirus Terapinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64351 | 60.184 | Terapinvirus | Unclassified | Siphoviridae | Gordonia |
| MT612989 | Vibrio phage vB_ValM_R11Z | Vibrio phage vB_ValM_R11Z unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246831 | 41.332 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MT612988 | Vibrio phage vB_ValM_R10Z | Vibrio phage vB_ValM_R10Z unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 247167 | 41.295 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MT559528 | Klebsiella phage vB_KpnM_cmc20193 | Klebsiella phage vB_KpnM_cmc20193 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48861 | 48.593 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MT559527 | Klebsiella phage vB_KpnP_cmc20192 | Klebsiella phage vB_KpnP_cmc20192 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44314 | 53.875 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT559526 | Klebsiella phage vB_KpnP_cmc20191 | Klebsiella phage vB_KpnP_cmc20191 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44066 | 53.971 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT496970 | Escherichia phage Mt1B1_P17 | Escherichia phage Mt1B1_P17 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151202 | 38.917 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MT496969 | Escherichia phage Mt1B1_P3 | Escherichia phage Mt1B1_P3 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40313 | 49.699 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT711979 | Streptomyces phage Endor2 | Streptomyces phage Endor2 unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48829 | 65.08 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT711978 | Streptomyces phage Endor1 | Streptomyces phage Endor1 unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49395 | 65.79 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| MT711975 | Streptomyces phage Alderaan | Streptomyces phage Alderaan unclassified Austintatiousvirus Austintatiousvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34411 | 72.128 | Austintatiousvirus | Unclassified | Siphoviridae | Streptomyces |
| MT711976 | Streptomyces phage Coruscant | Streptomyces phage Coruscant Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133697 | 48.387 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MT767754 | Microcystis phage Mwe-Yong908-1 | Microcystis phage Mwe-Yong908-1 unclassified bacterial viruses Viruses | 22207 | 35.471 | Unclassified | Unclassified | Unclassified | Microcystis |
| MT776806 | Mycobacterium phage MrMiyagi | Mycobacterium phage MrMiyagi unclassified Fowlmouthvirus Fowlmouthvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70048 | 48.751 | Fowlmouthvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT735629 | Vibrio phage vB_ValS_PJ32 | Vibrio phage vB_ValS_PJ32 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80294 | 45.541 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MT723942 | Gordonia phage Archis | Gordonia phage Archis Fairfaxidumvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49000 | 67.327 | Fairfaxidumvirus | Unclassified | Siphoviridae | Gordonia |
| MT711977 | Streptomyces phage Dagobah | Streptomyces phage Dagobah unclassified bacterial viruses Viruses | 47301 | 68.965 | Unclassified | Unclassified | Unclassified | Streptomyces |
| MK940809 | Staphylococcus phage Sa2wa_st8 | Staphylococcus phage Sa2wa_st8 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45914 | 33.121 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029518 | Staphylococcus phage phiSa2wa_st121mssa | Staphylococcus phage phiSa2wa_st121mssa Staphylococcus virus st121mssa Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45621 | 33.057 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029517 | Staphylococcus phage phiSa2wa_st93 | Staphylococcus phage phiSa2wa_st93 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45913 | 33.117 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029516 | Staphylococcus phage phiSa2wa_st93mssa | Staphylococcus phage phiSa2wa_st93mssa unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45913 | 33.113 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029515 | Staphylococcus phage phiSa2wa_st80 | Staphylococcus phage phiSa2wa_st80 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45164 | 33.257 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029514 | Staphylococcus phage phiSa2wa_st78 | Staphylococcus phage phiSa2wa_st78 Staphylococcus virus st78 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45878 | 33.216 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029513 | Staphylococcus phage phiSa2wa_st72 | Staphylococcus phage phiSa2wa_st72 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47213 | 33.137 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029512 | Staphylococcus phage phiSa2wa_st59 | Staphylococcus phage phiSa2wa_st59 unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42133 | 33.276 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MG029511 | Staphylococcus phage phiSa2wa_st30 | Staphylococcus phage phiSa2wa_st30 Staphylococcus virus st30 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45780 | 33.438 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG029510 | Staphylococcus phage phiSa2wa_st22 | Staphylococcus phage phiSa2wa_st22 Staphylococcus virus st22 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38576 | 33.241 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MG029509 | Staphylococcus phage phiSa2wa_st5 | Staphylococcus phage phiSa2wa_st5 Staphylococcus virus st5 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44823 | 33.407 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MF580410 | Staphylococcus phage phiSa2wa_st1 | Staphylococcus phage phiSa2wa_st1 Staphylococcus virus st1 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45585 | 33.281 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MT653145 | Salmonella phage 8sent65 | Salmonella phage 8sent65 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112133 | 39.971 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT653144 | Salmonella phage 5sent1 | Salmonella phage 5sent1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43760 | 49.904 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT653143 | Salmonella phage 3sent1 | Salmonella phage 3sent1 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109746 | 38.959 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT653146 | Salmonella phage 8sent1748 | Salmonella phage 8sent1748 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110720 | 39.288 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT799840 | Salmonella phage JN03 | Salmonella phage JN03 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241047 | 48.43 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| MT740291 | Salmonella phage SHWT1 | Salmonella phage SHWT1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40629 | 49.831 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT700412 | Bacillus phage 1_ICo-2020 | Bacillus phage 1_ICo-2020 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51227 | 41.79 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MT431535 | Flavobacterium phage FCV-1.01 | Flavobacterium phage FCV-1.01 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46364 | 29.767 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| MT808983 | Escherichia phage DY1 | Escherichia phage DY1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39817 | 49.841 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN746332 | Rauchvirus SR18 | Rauchvirus SR18 unclassified Rauchvirus Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45260 | 64.255 | Rauchvirus | Unclassified | Podoviridae | Unspecified |
| MN820898 | Sphingomonas phage vB_StuS_MMDA13 | Sphingomonas phage vB_StuS_MMDA13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63743 | 59.266 | Unclassified | Unclassified | Siphoviridae | Sphingomonas |
| MN393473 | Shigella phage Sfin-6 | Shigella phage Sfin-6 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50523 | 45.195 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MN585195 | Pseudomonas phage AUS531phi | Pseudomonas phage AUS531phi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48430 | 63.029 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MN047438 | Staphylococcus phage vB_SauM-515A1 | Staphylococcus phage vB_SauM-515A1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148511 | 30.236 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| LC576631 | Staphylococcus phage vB_SauH_DELF3 | Staphylococcus phage vB_SauH_DELF3 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136569 | 33.779 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT778840 | Rhizobium phage P11VFA | Rhizobium phage P11VFA Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109652 | 59.019 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MT778839 | Rhizobium phage P9VFCI | Rhizobium phage P9VFCI unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156502 | 49.814 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MT778838 | Rhizobium phage V1VFA-S | Rhizobium phage V1VFA-S Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118231 | 59.277 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MT778837 | Rhizobium phage AF3 | Rhizobium phage AF3 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156186 | 50.093 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MT786458 | Staphylococcus phage vB_SauH_SAP1 | Staphylococcus phage vB_SauH_SAP1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143375 | 30.161 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT770738 | Rhizobium phage B1VFA | Rhizobium phage B1VFA Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122345 | 59.351 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| MN563793 | Vibrio phage Valm-yong1 | Vibrio phage Valm-yong1 Vibrio virus Yong1 Yongunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33851 | 45.564 | Yongunavirus | Peduovirinae | Myoviridae | Vibrio |
| MT740741 | Ralstonia phage Hennie | Ralstonia phage Hennie unclassified Firingavirus Firingavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39476 | 59.439 | Firingavirus | Unclassified | Podoviridae | Ralstonia |
| MT740733 | Ralstonia phage Dimitile | Ralstonia phage Dimitile unclassified Cimandefvirus Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57009 | 64.581 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| MT740732 | Ralstonia phage Darius | Ralstonia phage Darius unclassified Gervaisevirus Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60734 | 64.827 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MT740727 | Ralstonia phage Alix | Ralstonia phage Alix unclassified Gervaisevirus Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57681 | 64.318 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MN131137 | Aeromonas phage D6 | Aeromonas phage D6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 259831 | 47.603 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MT675126 | Providencia phage vB_PreS-PatoteraRojo | Providencia phage vB_PreS-PatoteraRojo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60728 | 49.42 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MT740747 | Ralstonia phage Simangalove | Ralstonia phage Simangalove Ralstonia virus Simangalove Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44834 | 62.252 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MT740746 | Ralstonia phage Sarlave | Ralstonia phage Sarlave unclassified Bakolyvirus Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44858 | 62.256 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MT740745 | Ralstonia phage Raharianne | Ralstonia phage Raharianne Ralstonia virus Raharianne Rahariannevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46321 | 60.547 | Rahariannevirus | Unclassified | Myoviridae | Ralstonia |
| MT740744 | Ralstonia phage Jenny | Ralstonia phage Jenny unclassified Bakolyvirus Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43921 | 62.312 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MT740743 | Ralstonia phage Hyacinthe | Ralstonia phage Hyacinthe unclassified Rahariannevirus Rahariannevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46075 | 60.504 | Rahariannevirus | Unclassified | Myoviridae | Ralstonia |
| MT740742 | Ralstonia phage Heva | Ralstonia phage Heva Ralstonia virus Heva Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58352 | 64.62 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| MT740740 | Ralstonia phage Gervaise | Ralstonia phage Gervaise Ralstonia virus Gervaise Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61164 | 64.721 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MT740739 | Ralstonia phage Gerry | Ralstonia phage Gerry Ralstonia virus Gerry Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60898 | 64.73 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| MT740737 | Ralstonia phage Firinga | Ralstonia phage Firinga Ralstonia virus Firinga Firingavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39744 | 59.692 | Firingavirus | Unclassified | Podoviridae | Ralstonia |
| MT740736 | Ralstonia phage Eline | Ralstonia phage Eline Ralstonia virus Eline Cimandefvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57352 | 64.622 | Cimandefvirus | Unclassified | Podoviridae | Ralstonia |
| MT740735 | Ralstonia phage Elie | Ralstonia phage Elie unclassified Bakolyvirus Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44966 | 62.332 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MT740734 | Ralstonia phage Dina | Ralstonia phage Dina Ralstonia virus Dina Dinavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39161 | 65.274 | Dinavirus | Unclassified | Siphoviridae | Ralstonia |
| MT740729 | Ralstonia phage Bakoly | Ralstonia phage Bakoly Ralstonia virus Bakoly Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44222 | 62.324 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MT740728 | Ralstonia phage Anchaing | Ralstonia phage Anchaing Ralstonia virus Anchaing Anchaingvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43597 | 63.087 | Anchaingvirus | Unclassified | Autographiviridae | Ralstonia |
| MT740726 | Ralstonia phage Albius | Ralstonia phage Albius unclassified Rahariannevirus Rahariannevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46541 | 60.624 | Rahariannevirus | Unclassified | Myoviridae | Ralstonia |
| MT740725 | Ralstonia phage Adzire | Ralstonia phage Adzire unclassified Bakolyvirus Bakolyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44732 | 62.228 | Bakolyvirus | Unclassified | Myoviridae | Ralstonia |
| MN102098 | Aeromonas phage D3 | Aeromonas phage D3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262372 | 47.521 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MT664722 | Proteus phage Vb_PmiP-P59 | Proteus phage Vb_PmiP-P59 unclassified Privateervirus Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90187 | 34.651 | Privateervirus | Unclassified | Podoviridae | Proteus |
| MT675125 | Providencia phage vB_PreS-Stilesk | Providencia phage vB_PreS-Stilesk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60924 | 49.47 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MT675124 | Providencia phage vB_PreS-PibeRecoleta | Providencia phage vB_PreS-PibeRecoleta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60727 | 49.27 | Unclassified | Unclassified | Siphoviridae | Providencia |
| MT601274 | Bacillus phage vB_BsuS-Goe13 | Bacillus phage vB_BsuS-Goe13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126848 | 34.788 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MT601273 | Bacillus phage vB_BsuS-Goe12 | Bacillus phage vB_BsuS-Goe12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124287 | 34.797 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MT601272 | Bacillus phage vB_BsuS-Goe11 | Bacillus phage vB_BsuS-Goe11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126288 | 34.913 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MT601271 | Bacillus phage vB_BsuM-Goe10 | Bacillus phage vB_BsuM-Goe10 unclassified Okubovirus Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149539 | 40.351 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| MT601270 | Bacillus phage vB_BsuM-Goe9 | Bacillus phage vB_BsuM-Goe9 unclassified Okubovirus Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150961 | 40.355 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| MT625440 | Enterobacter phage PZJ0206 | Enterobacter phage PZJ0206 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40526 | 48.623 | Berlinvirus | Studiervirinae | Autographiviridae | Enterobacter |
| MT795651 | Vibrio phage vB_VnaS-AQKL99 | Vibrio phage vB_VnaS-AQKL99 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58464 | 45.866 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MK257721 | Acinetobacter phage vB_AbaP_APK93 | Acinetobacter phage vB_AbaP_APK93 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41668 | 39.376 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT663719 | Salmonella virus S144 | Salmonella virus S144 unclassified Loughboroughvirus Loughboroughvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53628 | 45.836 | Loughboroughvirus | Unclassified | Myoviridae | Salmonella |
| MT713136 | Aeromonas phage AP1 | Aeromonas phage AP1 unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 254490 | 40.333 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MT635598 | Bacteroides phage Bacuni_F1 | Bacteroides phage Bacuni_F1 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40421 | 51.75 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT514532 | Bacillus phage Bolokhovo | Bacillus phage Bolokhovo unclassified Andromedavirus Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49683 | 41.9 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| MT478993 | Escherichia phage vB_EcoS_011D5 | Escherichia phage vB_EcoS_011D5 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45204 | 54.491 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT478992 | Escherichia phage vB_EcoS_011D2 | Escherichia phage vB_EcoS_011D2 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50633 | 42.917 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MT580117 | Salmonella phage 66FD | Salmonella phage 66FD unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40964 | 50.261 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| MT580116 | Salmonella phage 65FD | Salmonella phage 65FD unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41328 | 50.278 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| MT677933 | Salmonella phage NBSal006 | Salmonella phage NBSal006 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41850 | 49.795 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT774487 | Salmonella phage MG40 | Salmonella phage MG40 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40315 | 47.156 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MT677934 | Salmonella phage NBSal007 | Salmonella phage NBSal007 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43747 | 49.809 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MT681669 | Aeromonas phage PZL-Ah1 | Aeromonas phage PZL-Ah1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38641 | 53.684 | Teseptimavirus | Studiervirinae | Autographiviridae | Aeromonas |
| MT662115 | Vibrio phage ValB1MD-2 | Vibrio phage ValB1MD-2 unclassified Peduovirinae Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36616 | 43.224 | Unclassified | Peduovirinae | Myoviridae | Vibrio |
| MT630433 | Bacteroides phage vB_BfrS_23 | Bacteroides phage vB_BfrS_23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48011 | 38.566 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| MT633129 | Acinetobacter phage Ab124 | Acinetobacter phage Ab124 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40471 | 39.505 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT591577 | Yersinia phage vB_YpM_Tongde | Yersinia phage vB_YpM_Tongde unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39492 | 51.783 | Muvirus | Unclassified | Myoviridae | Yersinia |
| MT227925 | Vibrio phage phiV141 | Vibrio phage phiV141 unclassified Tawavirus Tawavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43313 | 48.17 | Tawavirus | Unclassified | Autographiviridae | Vibrio |
| MN996301 | Ralstonia phage RpY1 | Ralstonia phage RpY1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43284 | 61.385 | Unclassified | Unclassified | Podoviridae | Ralstonia |
| MT611523 | Escherichia phage DK-13 | Escherichia phage DK-13 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172275 | 40.178 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MN729599 | Pseudomonas phage CHF33 | Pseudomonas phage CHF33 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40999 | 57.299 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN729598 | Pseudomonas phage CHF21 | Pseudomonas phage CHF21 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40557 | 57.364 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN729595 | Pseudomonas phage CHF1 | Pseudomonas phage CHF1 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40999 | 57.294 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| LR026998 | Salmonella phage YSD1 | Salmonella phage YSD1 Salmonella virus YSD1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58916 | 56.531 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MG250485 | Pseudomonas phage Pf1 ERZ-2017 | Pseudomonas phage Pf1 ERZ-2017 Pseudomonas virus Pf1ERZ2017 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39195 | 57.199 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KY514263 | Pectobacterium phage vB_PatM_CB7 | Pectobacterium phage vB_PatM_CB7 unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142778 | 50.11 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| MN342151 | Escherichia phage vB_EcoM_3HA14 | Escherichia phage vB_EcoM_3HA14 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88582 | 38.949 | Mooglevirus | Ounavirinae | Myoviridae | Escherichia |
| MN342150 | Escherichia phage vB_EcoM_3HA11 | Escherichia phage vB_EcoM_3HA11 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158429 | 44.862 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| MT491208 | Pseudomonas phage vB_PaeM_USP_25 | Pseudomonas phage vB_PaeM_USP_25 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62537 | 63.598 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MT491207 | Pseudomonas phage vB_PaeM_USP_18 | Pseudomonas phage vB_PaeM_USP_18 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63030 | 63.711 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MT491206 | Pseudomonas phage vB_PaeM_USP_3 | Pseudomonas phage vB_PaeM_USP_3 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65567 | 55.522 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT491205 | Pseudomonas phage vB_PaeM_USP_2 | Pseudomonas phage vB_PaeM_USP_2 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65606 | 55.452 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT491204 | Pseudomonas phage vB_PaeM_USP_1 | Pseudomonas phage vB_PaeM_USP_1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65918 | 55.578 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT330372 | Cronobacter phage JC01 | Cronobacter phage JC01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61736 | 58.515 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| MN263053 | Xanthomonas phage phiLF | Xanthomonas phage phiLF unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6009 | 59.943 | Unclassified | Unclassified | Inoviridae | Xanthomonas |
| MN445185 | Escherichia phage vB_EcoM_EC001 | Escherichia phage vB_EcoM_EC001 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240200 | 48.508 | Seoulvirus | Unclassified | Myoviridae | Escherichia |
| MN445184 | Escherichia phage vB_EcoA_4HA11 | Escherichia phage vB_EcoA_4HA11 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157406 | 45.259 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| MN445183 | Salmonella phage vB_SalM_SA002 | Salmonella phage vB_SalM_SA002 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 288012 | 48.837 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MN445182 | Salmonella phage vB_SalS_SA001 | Salmonella phage vB_SalS_SA001 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110965 | 38.972 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| LC554890 | Ralstonia phage RP13 | Ralstonia phage RP13 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170942 | 39.241 | Unclassified | Unclassified | Myoviridae | Ralstonia |
| MT682065 | Klebsiella virus KpV2883 | Klebsiella virus KpV2883 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43508 | 54.116 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT682064 | Klebsiella virus KpV2811 | Klebsiella virus KpV2811 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46391 | 48.208 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MT661596 | Proteus phage 10 | Proteus phage 10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223209 | 39.936 | Unclassified | Unclassified | Myoviridae | Proteus |
| MT354570 | Pseudomonas phage phiB1_1 | Pseudomonas phage phiB1_1 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41714 | 56.626 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT478991 | Escherichia phage vB_EcoM_011D4 | Escherichia phage vB_EcoM_011D4 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163764 | 40.505 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MT720689 | Vibrio phage XM1 | Vibrio phage XM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37633 | 42.096 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MT724048 | Staphylococcus phage SAP3 | Staphylococcus phage SAP3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41949 | 35.355 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT663720 | Pseudomonas phage DDSR119 | Pseudomonas phage DDSR119 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52905 | 60.391 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT613935 | Pseudomonas phage Persinger | Pseudomonas phage Persinger Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44033 | 62.86 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MT623546 | Acinetobacter phage Ab_121 | Acinetobacter phage Ab_121 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102499 | 37.203 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MT533174 | Escherichia phage CJ20 | Escherichia phage CJ20 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169884 | 40.439 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MN200778 | Vibrio phage VAI1 | Vibrio phage VAI1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6117 | 42.521 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MN200777 | Vibrio phage VAI2 | Vibrio phage VAI2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6117 | 42.472 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MT334653 | Buttiauxella phage vB_ButM_GuL6 | Buttiauxella phage vB_ButM_GuL6 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178039 | 43.383 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Buttiauxella |
| MT261384 | Salmonella virus PAT1 | Salmonella virus PAT1 unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41328 | 50.269 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| NC_049947 | Escherichia virus Lambda_1H12 | Escherichia virus Lambda_1H12 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46703 | 50.027 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MT162467 | Synechococcus phage S-H34 | Synechococcus phage S-H34 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167040 | 50.076 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MT162466 | Synechococcus phage S-N03 | Synechococcus phage S-N03 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167069 | 50.053 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MF939898 | Oenococcus phage vB_OeS_unk162 | Oenococcus phage vB_OeS_unk162 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36172 | 35.539 | Unclassified | Unclassified | Siphoviridae | Oenococcus |
| CP058906 | Micromonospora phage pMLP1 | Micromonospora phage pMLP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43352 | 71.515 | Unclassified | Unclassified | Siphoviridae | Micromonospora |
| MT623578 | Aeromonas phage PVN02 | Aeromonas phage PVN02 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51668 | 52.452 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| MT597419 | Pseudomonas phage vB_PsyP_3MF5 | Pseudomonas phage vB_PsyP_3MF5 Pseudomonas virus 3MF5 Warsawvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41608 | 56.528 | Warsawvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN937349 | Stenotrophomonas phage vB_SmaS_BUCT548 | Stenotrophomonas phage vB_SmaS_BUCT548 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62354 | 56.272 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MT536174 | Stenotrophomonas phage vB_SmaS-AXL_3 | Stenotrophomonas phage vB_SmaS-AXL_3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47545 | 63.348 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| MK820013 | Vibrio phage vB_VpaP_MGD2 | Vibrio phage vB_VpaP_MGD2 unclassified Kaohsiungvirus Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45105 | 43.867 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| MT385367 | Acinetobacter phage Abraxas | Acinetobacter phage Abraxas unclassified Lazarusvirus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166559 | 36.779 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MT409116 | Acinetobacter phage Octan | Acinetobacter phage Octan unclassified Lazarusvirus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166760 | 36.717 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN782535 | Acinetobacter phage vB_AbaM_Lazarus | Acinetobacter phage vB_AbaM_Lazarus Acinetobacter virus Lazarus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165151 | 36.763 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN732883 | Acinetobacter phage vB_AbaM_Kimel | Acinetobacter phage vB_AbaM_Kimel Acinetobacter virus Kimel Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167922 | 36.729 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN729600 | Pseudomonas phage CHF17 | Pseudomonas phage CHF17 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40882 | 57.302 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN729597 | Pseudomonas phage CHF19 | Pseudomonas phage CHF19 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40882 | 57.302 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN729596 | Pseudomonas phage CHF7 | Pseudomonas phage CHF7 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40557 | 57.376 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MN723850 | Acinetobacter phage vB_AbaM_Apostate | Acinetobacter phage vB_AbaM_Apostate Acinetobacter virus Apostate Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164746 | 36.752 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN709128 | Acinetobacter phage vB_AbaM_Berthold | Acinetobacter phage vB_AbaM_Berthold Acinetobacter virus Berthold Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165927 | 36.714 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN662249 | Acinetobacter phage Stupor | Acinetobacter phage Stupor unclassified Lazarusvirus Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164309 | 36.826 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN648195 | Acinetobacter phage vB_AbaM_Konradin | Acinetobacter phage vB_AbaM_Konradin Acinetobacter virus Konradin Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166210 | 36.746 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| CAJDKA010000002 | Enterococcus phage 163 | Enterococcus phage 163 unclassified Brockvirinae Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150836 | 37.036 | Unclassified | Brockvirinae | Herelleviridae | Enterococcus |
| CAJDJZ010000002 | Enterococcus phage vB_EfaH_149 | Enterococcus phage vB_EfaH_149 unclassified Brockvirinae Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142215 | 35.965 | Unclassified | Brockvirinae | Herelleviridae | Enterococcus |
| CAJDJY010000002 | Enterococcus phage 156 | Enterococcus phage 156 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141133 | 35.789 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| CAJCJZ010000002 | Enterococcus phage vB_EfaS_140 | Enterococcus phage vB_EfaS_140 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85454 | 30.185 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| CAJDJX010000002 | Enterococcus phage Q69 | Enterococcus phage Q69 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42141 | 34.567 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| CAJDKF010000002 | Enterococcus phage vB_EfaS_159 | Enterococcus phage vB_EfaS_159 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41718 | 34.359 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| CAJDKB010000002 | Enterococcus phage vB_EhiS_268 | Enterococcus phage vB_EhiS_268 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40542 | 34.87 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT521999 | Gordonia Phage Jablanski | Gordonia Phage Jablanski Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53521 | 66.475 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MT521997 | Gordonia phage Upyo | Gordonia phage Upyo unclassified Trinevirus Trinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45732 | 67.935 | Trinevirus | Unclassified | Siphoviridae | Gordonia |
| MT521994 | Gordonia phage Magel | Gordonia phage Magel unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59797 | 51.033 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MT647607 | Propionibacterium phage phiPA50S | Propionibacterium phage phiPA50S unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29590 | 54.387 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MN518894 | Escherichia virus LS2 | Escherichia virus LS2 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39554 | 50.018 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN518893 | Escherichia virus LS3 | Escherichia virus LS3 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40230 | 49.978 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT586120 | Synechococcus phage S-CAM7 | Synechococcus phage S-CAM7 Synechococcus virus SCAM7 Mazuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221362 | 41.177 | Mazuvirus | Unclassified | Myoviridae | Synechococcus |
| MT584805 | Bacillus phage Tomato | Bacillus phage Tomato unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146370 | 38.832 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MT500539 | Salmonella virus STSR3 | Salmonella virus STSR3 unclassified Drexlerviridae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49503 | 45.369 | Unclassified | Unclassified | Drexlerviridae | Salmonella |
| MT511058 | Escherichia phage vB_EcoD_SU57 | Escherichia phage vB_EcoD_SU57 unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46150 | 43.961 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MT522002 | Mycobacterium phage Topanga | Mycobacterium phage Topanga unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47881 | 65.059 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT682716 | Escherichia phage vB_EcoM_Gotham | Escherichia phage vB_EcoM_Gotham unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137025 | 43.683 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MT682715 | Escherichia phage vB_EcoS_Chapo | Escherichia phage vB_EcoS_Chapo unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51099 | 45.529 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MT682714 | Escherichia phage vB_EcoM_Lutter | Escherichia phage vB_EcoM_Lutter Escherichia virus Lutter Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170054 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682713 | Escherichia phage vB_EcoM_Ozark | Escherichia phage vB_EcoM_Ozark Escherichia virus Ozark Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167600 | 35.404 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682712 | Escherichia phage vB_EcoM_F1 | Escherichia phage vB_EcoM_F1 Escherichia virus EcoMF1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168410 | 35.355 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682711 | Escherichia phage vB_EcoM_FB | Escherichia phage vB_EcoM_FB unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171555 | 39.434 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682710 | Escherichia phage vB_EcoM_FT | Escherichia phage vB_EcoM_FT unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167431 | 35.32 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682709 | Escherichia phage vB_EcoM_SP1 | Escherichia phage vB_EcoM_SP1 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165416 | 35.579 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT682708 | Escherichia phage vB_EcoP_SP5M | Escherichia phage vB_EcoP_SP5M unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72896 | 43.181 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MT682707 | Escherichia phage vB_EcoP_SP7 | Escherichia phage vB_EcoP_SP7 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39305 | 50.073 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT682706 | Escherichia phage vB_EcoS_FP | Escherichia phage vB_EcoS_FP unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43757 | 44.091 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MT682705 | Escherichia phage vB_EcoS_SP8 | Escherichia phage vB_EcoS_SP8 unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57585 | 43.258 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MT495635 | Ralstonia phage RSBg | Ralstonia phage RSBg unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7540 | 61.711 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| LR794124 | Vibrio phage vB_Vc_SrVc9 | Vibrio phage vB_Vc_SrVc9 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43156 | 49.17 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| KJ409772 | Pseudomonas phage phiPsa374 | Pseudomonas phage phiPsa374 Pseudomonas virus Psa374 Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98287 | 47.716 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| MK863032 | Pseudomonas phage Zuri | Pseudomonas phage Zuri Pseudomonas virus Zuri Zurivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75873 | 53.528 | Zurivirus | Unclassified | Schitoviridae | Pseudomonas |
| MT584811 | Sulfitobacter phage phiGT1 | Sulfitobacter phage phiGT1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40019 | 56.386 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| MN402506 | Vibrio phage VMJ710 | Vibrio phage VMJ710 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121402 | 37.163 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MT498052 | Gordonia phage Jodelie19 | Gordonia phage Jodelie19 unclassified Kenoshavirus Kenoshavirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62331 | 51.417 | Kenoshavirus | Deejayvirinae | Siphoviridae | Gordonia |
| MT498039 | Gordonia phage Gill | Gordonia phage Gill unclassified Tanisvirus Tanisvirus Deejayvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59882 | 51.019 | Tanisvirus | Deejayvirinae | Siphoviridae | Gordonia |
| MT361768 | Ralstonia phage RPZH6 | Ralstonia phage RPZH6 unclassified Gervaisevirus Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64657 | 64.844 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MT457553 | Shewanella phage Thanatos-2 | Shewanella phage Thanatos-2 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155580 | 34.77 | Unclassified | Tevenvirinae | Myoviridae | Shewanella |
| MT074160 | Bacteroides phage SJC25 | Bacteroides phage SJC25 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38175 | 46.19 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074159 | Bacteroides phage SJC23 | Bacteroides phage SJC23 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38546 | 46.197 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074158 | Bacteroides phage SJC22 | Bacteroides phage SJC22 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38120 | 46.102 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074157 | Bacteroides phage SJC20 | Bacteroides phage SJC20 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37449 | 46.103 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074156 | Bacteroides phage SJC18 | Bacteroides phage SJC18 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37398 | 46.109 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074155 | Bacteroides phage SJC17 | Bacteroides phage SJC17 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38127 | 46.159 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074154 | Bacteroides phage SJC16 | Bacteroides phage SJC16 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38138 | 46.164 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074153 | Bacteroides phage SJC15 | Bacteroides phage SJC15 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38150 | 46.178 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074152 | Bacteroides phage SJC14 | Bacteroides phage SJC14 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38202 | 46.053 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074151 | Bacteroides phage SJC13 | Bacteroides phage SJC13 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38497 | 46.209 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074150 | Bacteroides phage SJC12 | Bacteroides phage SJC12 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38328 | 46.569 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074149 | Bacteroides phage SJC11 | Bacteroides phage SJC11 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38137 | 46.118 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074148 | Bacteroides phage SJC10 | Bacteroides phage SJC10 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37392 | 46.435 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074147 | Bacteroides phage SJC09 | Bacteroides phage SJC09 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38149 | 46.177 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074146 | Bacteroides phage SJC03 | Bacteroides phage SJC03 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166827 | 23.424 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074145 | Bacteroides phage SJC01 | Bacteroides phage SJC01 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38129 | 46.141 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074144 | Bacteroides phage HNL35 | Bacteroides phage HNL35 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37928 | 46.085 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074143 | Bacteroides phage HNL05 | Bacteroides phage HNL05 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37887 | 46.061 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074142 | Bacteroides phage DAC23 | Bacteroides phage DAC23 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179161 | 23.594 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074141 | Bacteroides phage DAC22 | Bacteroides phage DAC22 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179283 | 23.708 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074140 | Bacteroides phage DAC20 | Bacteroides phage DAC20 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178920 | 23.732 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074139 | Bacteroides phage DAC19 | Bacteroides phage DAC19 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178921 | 23.732 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074137 | Bacteroides phage DAC16 | Bacteroides phage DAC16 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178147 | 23.641 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074135 | Bacteroides phage ARB25 | Bacteroides phage ARB25 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37389 | 46.559 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MT074134 | Bacteroides phage ARB14 | Bacteroides phage ARB14 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37476 | 46.603 | Unclassified | Unclassified | Unclassified | Bacteroides |
| MN813681 | Mycobacterium phage Roary | Mycobacterium phage Roary unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52521 | 61.44 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| LR597660 | Escherichia phage T4_ev151 | Escherichia phage T4_ev151 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167097 | 35.291 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LR597659 | Escherichia phage T5_ev212 | Escherichia phage T5_ev212 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108546 | 39.025 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| LR597657 | Escherichia phage T4_ev240 | Escherichia phage T4_ev240 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166954 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LR597655 | Escherichia phage T5_ev219 | Escherichia phage T5_ev219 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110393 | 39.066 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| LR597653 | Escherichia phage Lambda_ev066 | Escherichia phage Lambda_ev066 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46723 | 50.016 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597652 | Escherichia phage Lambda_ev110 | Escherichia phage Lambda_ev110 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46770 | 50.015 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597651 | Escherichia phage Lambda_ev058 | Escherichia phage Lambda_ev058 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47678 | 49.927 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597650 | Escherichia phage Lambda_ev203 | Escherichia phage Lambda_ev203 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46680 | 50.03 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597648 | Escherichia phage Lambda_ev245 | Escherichia phage Lambda_ev245 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46704 | 50.03 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597646 | Escherichia phage Gluttony_ev152 | Escherichia phage Gluttony_ev152 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44825 | 54.608 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LR597644 | Escherichia phage Lambda_ev087 | Escherichia phage Lambda_ev087 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46710 | 50.032 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597643 | Escherichia phage Lambda_ev017 | Escherichia phage Lambda_ev017 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50126 | 49.689 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597639 | Escherichia phage Lambda_ev243 | Escherichia phage Lambda_ev243 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45560 | 50.316 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597636 | Escherichia phage Lambda_ev207 | Escherichia phage Lambda_ev207 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46685 | 50.288 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR597635 | Escherichia phage Lambda_ev099 | Escherichia phage Lambda_ev099 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47224 | 50.43 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MN234235 | Mycobacterium phage BogosyJay | Mycobacterium phage BogosyJay unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53217 | 62.768 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234209 | Mycobacterium phage Toaka | Mycobacterium phage Toaka unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53055 | 62.484 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234200 | Mycobacterium phage Maminiaina | Mycobacterium phage Maminiaina unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53199 | 62.781 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN234167 | Mycobacterium phage Vanisoa | Mycobacterium phage Vanisoa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53370 | 62.486 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN310542 | Mycobacterium phage Keziacharles14 | Mycobacterium phage Keziacharles14 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53549 | 61.646 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096376 | Mycobacterium phage Lucyedi | Mycobacterium phage Lucyedi unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52987 | 61.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK937605 | Mycobacterium phage Stephig9 | Mycobacterium phage Stephig9 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52669 | 61.362 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494124 | Mycobacterium phage Kristoff | Mycobacterium phage Kristoff unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51236 | 65.048 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494113 | Mycobacterium phage Refuge | Mycobacterium phage Refuge unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53594 | 63.548 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK468591 | Mycobacterium phage Arissanae | Mycobacterium phage Arissanae unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53365 | 62.599 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH825708 | Mycobacterium phage NearlyHeadless | Mycobacterium phage NearlyHeadless unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52383 | 61.409 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513984 | Mycobacterium phage Steamy | Mycobacterium phage Steamy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53420 | 63.401 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001459 | Mycobacterium phage KittenMittens | Mycobacterium phage KittenMittens unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 64.477 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF773750 | Mycobacterium phage OKCentral2016 | Mycobacterium phage OKCentral2016 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50072 | 65.13 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT990218 | Rhodococcus phage Lillie | Rhodococcus phage Lillie unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.55 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KT372002 | Rhodococcus phage CosmicSans | Rhodococcus phage CosmicSans unclassified Rerduovirus Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46596 | 58.548 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| KT184694 | Mycobacterium phage Smeadley | Mycobacterium phage Smeadley unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52392 | 61.414 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KR080193 | Mycobacterium phage Luchador | Mycobacterium phage Luchador unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53387 | 62.111 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101120 | Mycobacterium phage Trike | Mycobacterium phage Trike unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44716 | 64.498 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ156985 | Mycobacterium phage RhynO | Mycobacterium phage RhynO unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46739 | 65.262 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661279 | Mycobacterium phage Severus | Mycobacterium phage Severus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49894 | 64.449 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661275 | Mycobacterium phage HINdeR | Mycobacterium phage HINdeR unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52617 | 62.839 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX307704 | Mycobacterium virus Goose | Mycobacterium virus Goose Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50645 | 65.138 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX411619 | Mycobacterium virus Rebeuca | Mycobacterium virus Rebeuca Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51235 | 65.057 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ512844 | Mycobacterium phage Twister | Mycobacterium phage Twister Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51094 | 64.957 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN831654 | Mycobacterium virus Saintus | Mycobacterium virus Saintus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49228 | 61.235 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF957060 | Mycobacterium phage Timshel | Mycobacterium phage Timshel Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53278 | 63.144 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| NC_010463 | Salmonella virus Fels2 | Salmonella virus Fels2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33693 | 52.489 | Felsduovirus | Peduovirinae | Myoviridae | Salmonella |
| NC_004740 | Staphylococcus phage phiN315 | Staphylococcus phage phiN315 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44082 | 32.778 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| NC_004589 | Streptococcus phage 315.6 | Streptococcus phage 315.6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40014 | 39.714 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004588 | Streptococcus phage 315.5 | Streptococcus phage 315.5 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38206 | 38.083 | Unclassified | Unclassified | Unclassified | Streptococcus |
| NC_004587 | Streptococcus phage 315.4 | Streptococcus phage 315.4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41796 | 38.573 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004585 | Streptococcus phage 315.2 | Streptococcus phage 315.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41072 | 38.255 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004584 | Streptococcus phage 315.1 | Streptococcus phage 315.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39538 | 37.802 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MT498063 | Microbacterium phage Mercedes | Microbacterium phage Mercedes unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40230 | 63.271 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MT498062 | Microbacterium phage Leafus | Microbacterium phage Leafus unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42000 | 63.421 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT498049 | Microbacterium phage BigRedClifford | Microbacterium phage BigRedClifford unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41797 | 63.318 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT498048 | Mycobacterium phage Raymond7 | Mycobacterium phage Raymond7 unclassified Redivirus Redivirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42382 | 65.964 | Redivirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MT498044 | Mycobacterium phage Jamie19 | Mycobacterium phage Jamie19 unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41278 | 66.244 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MT498042 | Mycobacterium phage Rebel | Mycobacterium phage Rebel unclassified Charlievirus Charlievirus Nclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40580 | 66.518 | Charlievirus | Nclasvirinae | Siphoviridae | Mycobacterium |
| MT542697 | Klebsiella phage P509 | Klebsiella phage P509 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40954 | 53.206 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN122072 | Escherichia phage vB_EcoM_Bp10 | Escherichia phage vB_EcoM_Bp10 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52288 | 44.161 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MN067430 | Escherichia phage vB_EcoP-Ro103C3lw | Escherichia phage vB_EcoP-Ro103C3lw unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39389 | 52.84 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN117721 | Escherichia phage vB_EcoP_Bp7 | Escherichia phage vB_EcoP_Bp7 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52615 | 44.193 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT074138 | Bacteroides phage DAC17 | Bacteroides phage DAC17 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98900 | 37.034 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| MT074136 | Bacteroides phage DAC15 | Bacteroides phage DAC15 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99494 | 37.042 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| MN864865 | Acinetobacter phage vB_AbaM_PhT2 | Acinetobacter phage vB_AbaM_PhT2 Acinetobacter virus PhT2 Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166330 | 36.372 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MH523359 | Salmonella phage vB_SalM-LPST94 | Salmonella phage vB_SalM-LPST94 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156548 | 44.628 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH181876 | Salmonella phage vB_SalM-LPSEYT | Salmonella phage vB_SalM-LPSEYT unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53387 | 46.006 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| LR797923 | Bacteriophage APSE-7 | Bacteriophage APSE-7 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37531 | 43.964 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR813619 | Aeromonas phage PVN02 | Aeromonas phage PVN02 unclassified Pahsextavirus Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51668 | 52.452 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| NC_010393 | Phage Gifsy-2 | Phage Gifsy-2 unclassified bacterial viruses Viruses | 45840 | 51.095 | Unclassified | Unclassified | Unclassified | Unspecified |
| MT490373 | Mycobacterium phage OKaNui | Mycobacterium phage OKaNui unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51424 | 63.852 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT500540 | Listeria phage LP-HM00113468 | Listeria phage LP-HM00113468 Listeria virus LPHM00113468 Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40298 | 35.488 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| MT349888 | Pseudomonas phage PHW2 | Pseudomonas phage PHW2 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65984 | 55.692 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC552830 | Pseudomonas phage vB_PaeS-Yazdi-M | Pseudomonas phage vB_PaeS-Yazdi-M unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42439 | 54.448 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MT446423 | Escherichia virus TH58 | Escherichia virus TH58 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170748 | 37.537 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446422 | Escherichia virus TH57 | Escherichia virus TH57 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171230 | 40.375 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446421 | Escherichia virus TH55 | Escherichia virus TH55 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170389 | 40.302 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446420 | Escherichia virus TH54 | Escherichia virus TH54 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170988 | 40.369 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446418 | Escherichia virus TH50 | Escherichia virus TH50 unclassified Eapunavirus Eapunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41778 | 53.076 | Eapunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT446417 | Escherichia virus TH49 | Escherichia virus TH49 unclassified Eapunavirus Eapunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41043 | 53.022 | Eapunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT446416 | Escherichia virus TH47 | Escherichia virus TH47 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 352450 | 35.055 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MT446415 | Escherichia virus TH44 | Escherichia virus TH44 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171075 | 40.607 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446414 | Escherichia virus TH43 | Escherichia virus TH43 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77980 | 42.363 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT446413 | Escherichia virus TH42 | Escherichia virus TH42 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77284 | 42.343 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT446412 | Escherichia virus TH41 | Escherichia virus TH41 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170556 | 40.459 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446411 | Escherichia virus TH40 | Escherichia virus TH40 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171726 | 40.258 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446410 | Escherichia virus TH38 | Escherichia virus TH38 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81552 | 42.195 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT446408 | Escherichia virus TH35 | Escherichia virus TH35 unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69093 | 46.015 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MT446407 | Escherichia virus TH34 | Escherichia virus TH34 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77944 | 42.269 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT446405 | Escherichia virus TH32 | Escherichia virus TH32 unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69233 | 45.951 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MT446396 | Escherichia virus TH22 | Escherichia virus TH22 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170090 | 40.34 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446392 | Escherichia virus TH15 | Escherichia virus TH15 unclassified Gaprivervirus Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171591 | 40.259 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446391 | Escherichia virus TH13 | Escherichia virus TH13 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45377 | 54.442 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT446390 | Escherichia virus TH12 | Escherichia virus TH12 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169545 | 37.597 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446389 | Escherichia virus TH11 | Escherichia virus TH11 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45305 | 54.447 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT446388 | Escherichia virus TH10 | Escherichia virus TH10 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89140 | 38.875 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MT446387 | Escherichia virus TH09 | Escherichia virus TH09 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167656 | 35.465 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT446386 | Escherichia virus TH06 | Escherichia virus TH06 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77678 | 42.121 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT442039 | Vibrio phage vB_ValS_X1 | Vibrio phage vB_ValS_X1 unclassified Pogseptimavirus Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116033 | 43.861 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MT490303 | Prochlorococcus phage P-HS2 | Prochlorococcus phage P-HS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38327 | 37.157 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| MT422786 | Bacillus phage Novomoskovsk | Bacillus phage Novomoskovsk unclassified Andromedavirus Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49258 | 41.737 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| MT451985 | Microbacterium phage Dothraki | Microbacterium phage Dothraki unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41809 | 63.558 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT451983 | Microbacterium phage Ixel | Microbacterium phage Ixel unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40556 | 63.283 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MT451984 | Microbacterium phage Nebulous | Microbacterium phage Nebulous unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41419 | 64.256 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MT460517 | Vibrio phage Athena | Vibrio phage Athena unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127019 | 40.097 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT460518 | Vibrio phage Direpillow8 | Vibrio phage Direpillow8 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129165 | 40.091 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT460516 | Vibrio phage Quinn | Vibrio phage Quinn unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127684 | 40.094 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT460515 | Vibrio phage Cody | Vibrio phage Cody unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126683 | 40.179 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT460514 | Vibrio phage AG74 | Vibrio phage AG74 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126504 | 40.172 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT448617 | Vibrio phage River4 | Vibrio phage River4 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128425 | 40.233 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT448616 | Vibrio phage BBMuffin | Vibrio phage BBMuffin unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128290 | 40.153 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT448615 | Vibrio phage Marilyn | Vibrio phage Marilyn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42407 | 45.179 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MT349887 | Acinetobacter phage 5W | Acinetobacter phage 5W Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43032 | 39.842 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MK907285 | Salmonella phage vB_SalM-LPST153 | Salmonella phage vB_SalM-LPST153 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39176 | 49.094 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| MN047793 | Xanthomonas phage Xoo-sp13 | Xanthomonas phage Xoo-sp13 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 309023 | 40.934 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MT234670 | Pseudanabaena phage PA-SR01 | Pseudanabaena phage PA-SR01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137012 | 39.517 | Unclassified | Unclassified | Siphoviridae | Pseudanabaena |
| MT554104 | Staphylococcus phage ESa1 | Staphylococcus phage ESa1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153106 | 30.315 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG727701 | Paenibacillus phage Leyra | Paenibacillus phage Leyra Paenibacillus virus Leyra Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42276 | 41.395 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MT553343 | Microbacterium Phage ParleG | Microbacterium Phage ParleG unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41810 | 63.564 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT553341 | Microbacterium phage Kurt1 | Microbacterium phage Kurt1 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41803 | 63.55 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT553340 | Microbacterium phage Glamour | Microbacterium phage Glamour unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40224 | 62.08 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MT553339 | Microbacterium phage Lovelyunicorn | Microbacterium phage Lovelyunicorn unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41798 | 63.367 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT553337 | Mycobacterium phage DroogsArmy | Mycobacterium phage DroogsArmy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53254 | 63.015 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT303952 | Streptococcus phage phi29862 | Streptococcus phage phi29862 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54735 | 39.163 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN586062 | Mycobacterium phage Lev2 | Mycobacterium phage Lev2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50671 | 60.599 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586060 | Mycobacterium phage PetterN | Mycobacterium phage PetterN unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51121 | 59.745 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586016 | Mycobacterium phage MarysWell | Mycobacterium phage MarysWell unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51469 | 60.922 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN656993 | Enterobacter phage ATCEA85 | Enterobacter phage ATCEA85 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47484 | 56.154 | Unclassified | Unclassified | Siphoviridae | Enterobacter |
| LC373201 | Enterobacter phage EspM4VN | Enterobacter phage EspM4VN unclassified Agtrevirus Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160766 | 50.701 | Agtrevirus | Aglimvirinae | Ackermannviridae | Enterobacter |
| MK305887 | Mycobacterium phage HuhtaEnerson15 | Mycobacterium phage HuhtaEnerson15 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51106 | 60.967 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051256 | Mycobacterium phage Midas2 | Mycobacterium phage Midas2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51215 | 60.98 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051254 | Mycobacterium phage Jabiru | Mycobacterium phage Jabiru unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50530 | 59.986 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051250 | Mycobacterium phage Coog | Mycobacterium phage Coog unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51215 | 60.98 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020239 | Mycobacterium phage Naca | Mycobacterium phage Naca unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51594 | 59.831 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT438501 | Mycobacterium phage Theia | Mycobacterium phage Theia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51543 | 60.802 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT004677 | Mycobacterium phage UnionJack | Mycobacterium phage UnionJack unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49158 | 59.982 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF560330 | Mycobacterium phage Conspiracy | Mycobacterium phage Conspiracy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50755 | 60.615 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF017001 | Mycobacterium phage LittleCherry | Mycobacterium phage LittleCherry unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50690 | 60.933 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JQ684677 | Mycobacterium phage Tiger | Mycobacterium phage Tiger Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50332 | 60.665 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT446419 | Escherichia virus TH53 | Escherichia virus TH53 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52329 | 44.195 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446409 | Escherichia virus TH37 | Escherichia virus TH37 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53453 | 43.85 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446406 | Escherichia virus TH33 | Escherichia virus TH33 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52851 | 43.98 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446404 | Escherichia virus TH30 | Escherichia virus TH30 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51754 | 47.249 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| MT446403 | Escherichia virus TH29 | Escherichia virus TH29 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53648 | 43.877 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446402 | Escherichia virus TH28 | Escherichia virus TH28 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55824 | 44.135 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446401 | Escherichia virus TH27 | Escherichia virus TH27 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52344 | 46.619 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| MT446400 | Escherichia virus TH26 | Escherichia virus TH26 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51200 | 43.93 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446399 | Escherichia virus TH25 | Escherichia virus TH25 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51729 | 44.035 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446398 | Escherichia virus TH24 | Escherichia virus TH24 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52806 | 44.111 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446397 | Escherichia virus TH23 | Escherichia virus TH23 unclassified Carltongylesvirus Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52433 | 43.955 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MT446395 | Escherichia virus TH21 | Escherichia virus TH21 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52087 | 47.277 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| MT446394 | Escherichia virus TH20 | Escherichia virus TH20 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40001 | 50.216 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT446393 | Escherichia virus TH16 | Escherichia virus TH16 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40612 | 50.148 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MT459794 | Bacillus phage Gxv1 | Bacillus phage Gxv1 Bacillus virus Gxv1 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21781 | 39.686 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| MT375042 | Acinetobacter phage vB_AbaP_Alexa | Acinetobacter phage vB_AbaP_Alexa Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40237 | 49.067 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MT423823 | Staphylococcus virus pSp_SNUABM-J | Staphylococcus virus pSp_SNUABM-J Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40224 | 35.74 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MT438761 | Listeria virus P61 | Listeria virus P61 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136485 | 35.922 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MT465573 | Escherichia phage P1723 | Escherichia phage P1723 unclassified Foetvirus Foetvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40282 | 50.397 | Foetvirus | Studiervirinae | Autographiviridae | Escherichia |
| MT127620 | Escherichia phage PGN6866 | Escherichia phage PGN6866 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78549 | 42.265 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MT127619 | Escherichia phage PGN590 | Escherichia phage PGN590 Escherichia virus PGN590 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49043 | 43.806 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MT501516 | Vibrio phage vB_VpaP_MGD1 | Vibrio phage vB_VpaP_MGD1 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43290 | 49.522 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MN807295 | Acinetobacter phage vB_AbaP_APK116 | Acinetobacter phage vB_AbaP_APK116 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41765 | 39.081 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN692199 | Pectobacterium phage MA12 | Pectobacterium phage MA12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58573 | 54.535 | Unclassified | Unclassified | Siphoviridae | Pectobacterium |
| MN078104 | Bacteroides phage Barc2635 | Bacteroides phage Barc2635 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45990 | 38.878 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| MK932884 | Bifidobacterium phage PMBT6 | Bifidobacterium phage PMBT6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36561 | 61.653 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| CP053388 | Escherichia phage CMS-2020a | Escherichia phage CMS-2020a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107648 | 46.42 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| CP054387 | Escherichia phage CMS-2020a | Escherichia phage CMS-2020a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109272 | 45.309 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MH853356 | Streptococcus phage LF2 | Streptococcus phage LF2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44768 | 36.718 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN862347 | Sedum sarmentosum microvirus | Sedum sarmentosum microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5378 | 38.267 | Unclassified | Unclassified | Microviridae | Unspecified |
| MN862346 | Kummerowia striata microvirus | Kummerowia striata microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4367 | 56.103 | Unclassified | Unclassified | Microviridae | Unspecified |
| MN862345 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4489 | 48.429 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862344 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4547 | 49.791 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862343 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4596 | 48.825 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862342 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4577 | 60.454 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862341 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4764 | 53.988 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862340 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4602 | 54.628 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862339 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4562 | 42.153 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862338 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4615 | 47.974 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862337 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4770 | 59.455 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862336 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4497 | 54.481 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862335 | Kummerowia striata gokushovirus | Kummerowia striata gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4398 | 49.613 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862334 | Trichosanthes kirilowii gokushovirus | Trichosanthes kirilowii gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4756 | 42.704 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MN862333 | Trichosanthes kirilowii gokushovirus | Trichosanthes kirilowii gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4668 | 55.141 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MT311968 | Streptococcus phage phi29961 | Streptococcus phage phi29961 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59372 | 38.338 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MT311967 | Streptococcus phage phi29854 | Streptococcus phage phi29854 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54779 | 39.128 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MT430910 | Streptococcus phage smHBZ8 | Streptococcus phage smHBZ8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32219 | 38.775 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| LC553736 | Salmonella phage SAP012 | Salmonella phage SAP012 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59618 | 54.056 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| LC553735 | Escherichia phage O18-011 | Escherichia phage O18-011 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75646 | 42.1 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| LC553734 | Escherichia phage EK010 | Escherichia phage EK010 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78078 | 42.112 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MN116494 | Klebsiella phage vB_KpnP_Bp5 | Klebsiella phage vB_KpnP_Bp5 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43872 | 53.909 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN023191 | Brevibacterium phage AGM16 | Brevibacterium phage AGM16 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35637 | 60.743 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023190 | Brevibacterium phage AGM15 | Brevibacterium phage AGM15 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36825 | 60.861 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023189 | Brevibacterium phage AGM14 | Brevibacterium phage AGM14 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36678 | 60.84 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023188 | Brevibacterium phage AGM13 | Brevibacterium phage AGM13 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36480 | 60.811 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023187 | Brevibacterium phage AGM12 | Brevibacterium phage AGM12 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36429 | 60.814 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023186 | Brevibacterium phage AGM11 | Brevibacterium phage AGM11 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36341 | 60.805 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023185 | Brevibacterium phage AGM10 | Brevibacterium phage AGM10 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36341 | 60.802 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023184 | Brevibacterium phage AGM9 | Brevibacterium phage AGM9 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36282 | 60.799 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023183 | Brevibacterium phage AGM8 | Brevibacterium phage AGM8 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36232 | 60.792 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023182 | Brevibacterium phage AGM7 | Brevibacterium phage AGM7 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36232 | 60.789 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023181 | Brevibacterium phage AGM6 | Brevibacterium phage AGM6 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36231 | 60.799 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023180 | Brevibacterium phage AGM5 | Brevibacterium phage AGM5 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36231 | 60.796 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023179 | Brevibacterium phage AGM4 | Brevibacterium phage AGM4 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36083 | 60.782 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023178 | Brevibacterium phage AGM3 | Brevibacterium phage AGM3 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36034 | 60.77 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023177 | Brevibacterium phage AGM2 | Brevibacterium phage AGM2 unclassified Agmunavirus Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36032 | 60.771 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MN023176 | Brevibacterium phage AGM1 | Brevibacterium phage AGM1 Brevibacterium virus AGM1 Agmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36032 | 60.774 | Agmunavirus | Unclassified | Siphoviridae | Brevibacterium |
| MT322327 | Escherichia phage SSBS18_WS_10_728 | Escherichia phage SSBS18_WS_10_728 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86157 | 38.995 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MT006214 | CrAssphage LMMB | CrAssphage LMMB crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98458 | 28.952 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MT459144 | Vibrio phage Dax | Vibrio phage Dax unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127856 | 40.133 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT451873 | Vibrio phage vB_VhaS-VHB1 | Vibrio phage vB_VhaS-VHB1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80965 | 44.96 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MT425185 | Klebsiella phage Kp_Pokalde_002 | Klebsiella phage Kp_Pokalde_002 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41816 | 53.011 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MT416612 | Bacillus phage YungSlug | Bacillus phage YungSlug unclassified Spounavirinae Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150033 | 37.564 | Unclassified | Spounavirinae | Herelleviridae | Bacillus |
| MT411892 | Staphylococcus phage Metroid | Staphylococcus phage Metroid unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150935 | 30.4 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT374859 | Yersinia phage vB_YpP_T3 | Yersinia phage vB_YpP_T3 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38419 | 47.24 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| MT374858 | Yersinia phage vB_YpM_50 | Yersinia phage vB_YpM_50 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31096 | 52.126 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| MT374857 | Yersinia phage vB_YpM_46 | Yersinia phage vB_YpM_46 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32336 | 51.661 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| MT374856 | Yersinia phage vB_YpM_23 | Yersinia phage vB_YpM_23 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38732 | 51.787 | Muvirus | Unclassified | Myoviridae | Yersinia |
| MT374855 | Yersinia phage vB_YpM_22 | Yersinia phage vB_YpM_22 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31809 | 51.467 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| MT374854 | Yersinia phage vB_YpM_6 | Yersinia phage vB_YpM_6 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37277 | 51.879 | Muvirus | Unclassified | Myoviridae | Yersinia |
| MT374853 | Yersinia phage vB_YpM_5 | Yersinia phage vB_YpM_5 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37228 | 51.883 | Muvirus | Unclassified | Myoviridae | Yersinia |
| MT374852 | Yersinia phage vB_YpM_3 | Yersinia phage vB_YpM_3 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39387 | 51.733 | Muvirus | Unclassified | Myoviridae | Yersinia |
| MK922548 | Staphylococcus phage SA46-CTH4 | Staphylococcus phage SA46-CTH4 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17473 | 28.764 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK922547 | Staphylococcus phage SA30-CTH2 | Staphylococcus phage SA30-CTH2 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17545 | 28.755 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK922546 | Staphylococcus phage SA1-CTA1 | Staphylococcus phage SA1-CTA1 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17527 | 28.773 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK936476 | Staphylococcus phage SA46-CL1 | Staphylococcus phage SA46-CL1 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17508 | 28.81 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK936475 | Staphylococcus phage SA03-CTH2 | Staphylococcus phage SA03-CTH2 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17511 | 28.793 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MN716856 | Yersinia phage vB_YepM_ZN18 | Yersinia phage vB_YepM_ZN18 Yersinia virus ZN18 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167948 | 35.353 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MN830259 | Lactobacillus phage JNU_P10 | Lactobacillus phage JNU_P10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41955 | 45.065 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MN830258 | Lactobacillus phage JNU_P9 | Lactobacillus phage JNU_P9 unclassified Sukhumvitvirus Sukhumvitvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41609 | 43.829 | Sukhumvitvirus | Unclassified | Siphoviridae | Lactobacillus |
| MN830257 | Lactobacillus phage JNU_P11 | Lactobacillus phage JNU_P11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36721 | 36.35 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MN830256 | Lactobacillus phage JNU_P7 | Lactobacillus phage JNU_P7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38688 | 37.05 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MN830255 | Lactobacillus phage JNU_P4 | Lactobacillus phage JNU_P4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49020 | 43.684 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| MN830254 | Lactobacillus phage JNU_P2 | Lactobacillus phage JNU_P2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48969 | 43.683 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| MN830253 | Lactobacillus phage JNU_P5 | Lactobacillus phage JNU_P5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58246 | 43.677 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MN830252 | Lactobacillus phage JNU_P1 | Lactobacillus phage JNU_P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52315 | 44.958 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MT075871 | Klebsiella phage vB_KleS-HSE3 | Klebsiella phage vB_KleS-HSE3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46747 | 56.472 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MN433707 | Acinetobacter phage vB_AbaP_PMK34 | Acinetobacter phage vB_AbaP_PMK34 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41847 | 39.343 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MH673672 | Thermus phage phiFa | Thermus phage phiFa unclassified Oshimavirus Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77664 | 59.197 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| MT410774 | Psychrobacillus phage Spoks | Psychrobacillus phage Spoks Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36472 | 35.83 | Unclassified | Unclassified | Siphoviridae | Psychrobacillus |
| KF147891 | Pseudomonas phage PaBG | Pseudomonas phage PaBG Pseudomonas virus PaBG Baikalvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 258139 | 55.815 | Baikalvirus | Unclassified | Myoviridae | Pseudomonas |
| CP051288 | Salmonella phage ST-29 | Salmonella phage ST-29 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40874 | 47.032 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| CP051285 | Salmonella phage ST-32 | Salmonella phage ST-32 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40826 | 46.96 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| CP051279 | Salmonella phage ST-35 | Salmonella phage ST-35 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39033 | 46.922 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| CP051278 | Salmonella phage ST-87 | Salmonella phage ST-87 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40825 | 46.952 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| CP051275 | Salmonella phage SW-37 | Salmonella phage SW-37 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41358 | 49.911 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| CP051272 | Salmonella phage SW-70 | Salmonella phage SW-70 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42381 | 47.2 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MT408532 | Synechococcus phage SynMITS9220M01 | Synechococcus phage SynMITS9220M01 unclassified Bellamyvirus Bellamyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 190237 | 38.838 | Bellamyvirus | Unclassified | Myoviridae | Synechococcus |
| MT344105 | Acinetobacter phage fLi-Aba03 | Acinetobacter phage fLi-Aba03 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34931 | 40.09 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MT344104 | Acinetobacter phage fLi-Aba02 | Acinetobacter phage fLi-Aba02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35093 | 40.159 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MT344103 | Acinetobacter phage fEg-Aba01 | Acinetobacter phage fEg-Aba01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33779 | 40.102 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| AP018815 | Pseudomonas phage PaGU11 | Pseudomonas phage PaGU11 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65554 | 55.551 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| AP018814 | Enterobacteria phage T6 | Enterobacteria phage T6 Enterobacteria virus T6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168697 | 35.211 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| AP018813 | Escherichia phage T2 | Escherichia phage T2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163826 | 35.316 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT457552 | Shewanella phage Thanatos-1 | Shewanella phage Thanatos-1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160584 | 34.512 | Unclassified | Tevenvirinae | Myoviridae | Shewanella |
| MT366568 | Staphylococcus phage R4 | Staphylococcus phage R4 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45168 | 33.287 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MT360681 | Shigella virus 2019SD1 | Shigella virus 2019SD1 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53145 | 44.512 | Hanrivervirus | Tempevirinae | Drexlerviridae | Shigella |
| MT360680 | Klebsiella virus 2019KP1 | Klebsiella virus 2019KP1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48372 | 53.7 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT362618 | Microcystis phage MaeS | Microcystis phage MaeS Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79995 | 37.055 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MT350293 | Enterococcus phage EFA-2 | Enterococcus phage EFA-2 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39964 | 48.554 | Teseptimavirus | Studiervirinae | Autographiviridae | Enterococcus |
| MT350292 | Enterococcus phage EFA-1 | Enterococcus phage EFA-1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40712 | 50.145 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterococcus |
| MK387869 | Providencia phage vB PstP PS3 | Providencia phage vB PstP PS3 Providencia virus PS3 Kakivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41024 | 42.992 | Kakivirus | Unclassified | Autographiviridae | Providencia |
| MN651570 | Acinetobacter phage vB_AbaP_APK89 | Acinetobacter phage vB_AbaP_APK89 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41198 | 39.398 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN103533 | Mycobacterium phage Weirdo19 | Mycobacterium phage Weirdo19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52581 | 70.195 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MT104475 | Pseudomonas phage MR15 | Pseudomonas phage MR15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51347 | 58.054 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT104473 | Pseudomonas phage MR13 | Pseudomonas phage MR13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47456 | 57.946 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| NC_047800 | Ralstonia phage phiAp1 | Ralstonia phage phiAp1 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44793 | 60.967 | Unclassified | Krylovirinae | Autographiviridae | Ralstonia |
| NC_047765 | Ralstonia phage Rs551 | Ralstonia phage Rs551 Ralstonia virus RS551 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7929 | 60.827 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| MT104476 | Pseudomonas phage MR16 | Pseudomonas phage MR16 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42827 | 57.55 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104477 | Pseudomonas phage MR18 | Pseudomonas phage MR18 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41420 | 57.542 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104472 | Pseudomonas phage MR12 | Pseudomonas phage MR12 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41368 | 57.508 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104471 | Pseudomonas phage MR8 | Pseudomonas phage MR8 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41371 | 57.533 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104470 | Pseudomonas phage MR7 | Pseudomonas phage MR7 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41705 | 57.537 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104469 | Pseudomonas phage MR6 | Pseudomonas phage MR6 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41300 | 57.574 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104468 | Pseudomonas phage MR5 | Pseudomonas phage MR5 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41270 | 57.587 | Unclassified | Krylovirinae | Autographiviridae | Pseudomonas |
| MT104467 | Pseudomonas phage MR4 | Pseudomonas phage MR4 unclassified Gajwadongvirus Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44816 | 45.736 | Gajwadongvirus | Unclassified | Autographiviridae | Pseudomonas |
| MT104466 | Pseudomonas phage MR2 | Pseudomonas phage MR2 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40986 | 57.576 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MT104465 | Pseudomonas phage MR1 | Pseudomonas phage MR1 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39033 | 56.165 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| NC_005949 | Vibrio phage Vf12 | Vibrio phage Vf12 Villovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7965 | 45.675 | Villovirus | Unclassified | Inoviridae | Vibrio |
| LC547238 | Enterococcus phage phiM1EF2 | Enterococcus phage phiM1EF2 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56582 | 40.02 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| LC492752 | Staphylococcus phage KSAP11 | Staphylococcus phage KSAP11 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138307 | 29.878 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| LC492751 | Staphylococcus phage KSAP7 | Staphylococcus phage KSAP7 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137950 | 29.874 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT197176 | Klebsiella phage VLC6 | Klebsiella phage VLC6 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44294 | 54.159 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MK931297 | Escherichia phage vB_EcoP_3HA13 | Escherichia phage vB_EcoP_3HA13 unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70142 | 41.268 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MN958086 | Vibrio phage Bennett | Vibrio phage Bennett unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126552 | 40.098 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT424636 | Synechococcus phage S-SBP1 | Synechococcus phage S-SBP1 Ashivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45519 | 46.846 | Ashivirus | Unclassified | Autographiviridae | Synechococcus |
| MT108726 | Pseudomonas phage Epa6 | Pseudomonas phage Epa6 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66031 | 55.139 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK903033 | Staphylococcus phage SA44-CTH7 | Staphylococcus phage SA44-CTH7 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17522 | 28.867 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MN336266 | Salmonella phage PIZ SAE-01E2 | Salmonella phage PIZ SAE-01E2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43095 | 49.639 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN478376 | Bacteriophage Reminis | Bacteriophage Reminis unclassified Gajwadongvirus Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45228 | 42.746 | Gajwadongvirus | Unclassified | Autographiviridae | Unspecified |
| MK450538 | Staphylococcus phage vB_SpsS_QT1 | Staphylococcus phage vB_SpsS_QT1 Staphylococcus virus QT1 Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43029 | 36.882 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG264739 | Enterococcus phage phiNASRA1 | Enterococcus phage phiNASRA1 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40139 | 34.742 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MT341500 | Enterobacter phage EBPL | Enterobacter phage EBPL unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179574 | 44.829 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MT331608 | Escherichia phage SECphi4 | Escherichia phage SECphi4 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44569 | 54.28 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MT325768 | Psychrobacillus phage Perkons | Psychrobacillus phage Perkons Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136811 | 30.589 | Unclassified | Unclassified | Siphoviridae | Psychrobacillus |
| CP052843 | Bacillus phage Hybphi3Ts | Bacillus phage Hybphi3Ts unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31849 | 35.624 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| LC413195 | Klebsiella phage KN4-1 | Klebsiella phage KN4-1 Klebsiella virus KN4-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41219 | 52.944 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| LC413194 | Klebsiella phage KN3-1 | Klebsiella phage KN3-1 Klebsiella virus KN3-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41059 | 53.491 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| LC413193 | Klebsiella phage KN1-1 | Klebsiella phage KN1-1 Klebsiella virus KN1-1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40236 | 52.843 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| LT615366 | Enterococcus phage VPE25 | Enterococcus phage VPE25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86524 | 30.342 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MN876845 | Pyrobaculum spherical virus 2 | Pyrobaculum spherical virus 2 Alphaglobulovirus PSV2 Alphaglobulovirus Globuloviridae Viruses | 18212 | 45.058 | Alphaglobulovirus | Unclassified | Globuloviridae | Pyrobaculum |
| MN876843 | Metallosphaera rod-shaped virus 1 | Metallosphaera rod-shaped virus 1 Hoswirudivirus MRV1 Hoswirudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 20269 | 34.126 | Hoswirudivirus | Unclassified | Rudiviridae | Metallosphaera |
| MN876842 | Acidianus rod-shaped virus 3 | Acidianus rod-shaped virus 3 Hoswirudivirus ARV3 Hoswirudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 23666 | 32.025 | Hoswirudivirus | Unclassified | Rudiviridae | Acidianus |
| MN876841 | Saccharolobus solfataricus rod-shaped virus 1 | Saccharolobus solfataricus rod-shaped virus 1 Hoswirudivirus SSRV1 Hoswirudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 26097 | 32.314 | Hoswirudivirus | Unclassified | Rudiviridae | Saccharolobus |
| MK982307 | Enterococcus phage MSF2 | Enterococcus phage MSF2 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40880 | 34.447 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MN161573 | Salmonella virus P22 | Salmonella virus P22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41725 | 47.065 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MN417334 | Clostridium phage CPAS-15 | Clostridium phage CPAS-15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51829 | 34.182 | Unclassified | Unclassified | Myoviridae | Clostridium |
| MK867354 | Synechococcus phage S-SCSM1 | Synechococcus phage S-SCSM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 228827 | 36.842 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MK764450 | Flavobacterium phage FPSV-D35 | Flavobacterium phage FPSV-D35 unclassified Unahavirus Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42339 | 31.824 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| MK764449 | Flavobacterium phage FPSV-S29 | Flavobacterium phage FPSV-S29 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91708 | 26.808 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MK764448 | Flavobacterium phage FPSV-S27 | Flavobacterium phage FPSV-S27 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23990 | 38.303 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MK764447 | Flavobacterium phage FPSV-S20 | Flavobacterium phage FPSV-S20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.979 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764446 | Flavobacterium phage FPSV-S18 | Flavobacterium phage FPSV-S18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.974 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764445 | Flavobacterium phage FPSV-S8 | Flavobacterium phage FPSV-S8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22503 | 37.364 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MK764444 | Flavobacterium phage FPSV-S3 | Flavobacterium phage FPSV-S3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.979 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764443 | Flavobacterium phage FPSV-S2 | Flavobacterium phage FPSV-S2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42658 | 31.903 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764442 | Flavobacterium phage FPSV-S1 | Flavobacterium phage FPSV-S1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35187 | 37.841 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MK764441 | Flavobacterium phage FPSV-F12 | Flavobacterium phage FPSV-F12 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89814 | 26.957 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| MK764440 | Flavobacterium phage FPSV-F7 | Flavobacterium phage FPSV-F7 unclassified Unahavirus Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39282 | 32.132 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| MK764439 | Flavobacterium phage FPSV-D22 | Flavobacterium phage FPSV-D22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42699 | 31.975 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764438 | Flavobacterium phage FPSV-D19 | Flavobacterium phage FPSV-D19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42756 | 31.972 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MK764437 | Flavobacterium phage FPSV-D15 | Flavobacterium phage FPSV-D15 unclassified Unahavirus Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47249 | 31.764 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| MK764436 | Flavobacterium phage FPSV-D2 | Flavobacterium phage FPSV-D2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42762 | 32.007 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MT310897 | Mycobacterium phage Manatee | Mycobacterium phage Manatee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51040 | 63.791 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310881 | Mycobacterium phage JF2 | Mycobacterium phage JF2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45969 | 64.08 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310880 | Mycobacterium phage JF4 | Mycobacterium phage JF4 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45969 | 64.089 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310879 | Mycobacterium phage MA5 | Mycobacterium phage MA5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47895 | 64.092 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310878 | Mycobacterium phage MK4 | Mycobacterium phage MK4 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46535 | 64.345 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310876 | Mycobacterium phage VC3 | Mycobacterium phage VC3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51141 | 63.159 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MT310873 | Microbacterium phage Convict | Microbacterium phage Convict unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41798 | 63.359 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310866 | Microbacterium phage Pherferi | Microbacterium phage Pherferi unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41803 | 63.553 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310861 | Microbacterium phage BeautPeep30 | Microbacterium phage BeautPeep30 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41809 | 63.517 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310856 | Microbacterium phage Zada | Microbacterium phage Zada unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41814 | 63.481 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310855 | Microbacterium phage Rasovi | Microbacterium phage Rasovi unclassified Goodmanvirus Goodmanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42917 | 65.226 | Goodmanvirus | Unclassified | Siphoviridae | Microbacterium |
| MT310851 | Microbacterium phage Sedgewig | Microbacterium phage Sedgewig unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41505 | 63.438 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN922296 | Vibrio phage phiV039C | Vibrio phage phiV039C Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43405 | 42.954 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN734439 | Sphingomonas phage Kharn | Sphingomonas phage Kharn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43408 | 55.879 | Unclassified | Unclassified | Siphoviridae | Sphingomonas |
| MN734438 | Sphingomonas phage Lucius | Sphingomonas phage Lucius Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44200 | 55.966 | Unclassified | Unclassified | Siphoviridae | Sphingomonas |
| MN734437 | Sphingomonas phage Eidolon | Sphingomonas phage Eidolon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43318 | 54.996 | Unclassified | Unclassified | Siphoviridae | Sphingomonas |
| MN734436 | Aminobacter phage Erebus | Aminobacter phage Erebus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52229 | 57.424 | Unclassified | Unclassified | Myoviridae | Aminobacter |
| LT603684 | Pseudomonas phage phiC725A | Pseudomonas phage phiC725A unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64149 | 63.452 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MN604239 | Acinetobacter phage vB_AbaP_APK87 | Acinetobacter phage vB_AbaP_APK87 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42402 | 39.055 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN604238 | Acinetobacter phage vB_AbaP_APK44 | Acinetobacter phage vB_AbaP_APK44 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41461 | 39.07 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK053931 | Pectobacterium phage Arno160 | Pectobacterium phage Arno160 Pectobacterium virus Arno160 Wanjuvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41823 | 51.455 | Wanjuvirus | Melnykvirinae | Autographiviridae | Pectobacterium |
| LR794152 | Bacteriophage APSE-2 | Bacteriophage APSE-2 Hamiltonella virus APSE2 Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38420 | 42.902 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR794151 | Bacteriophage APSE-7 | Bacteriophage APSE-7 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36931 | 43.86 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR794150 | Bacteriophage APSE-3 | Bacteriophage APSE-3 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41343 | 44.37 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR794149 | Bacteriophage APSE-7 | Bacteriophage APSE-7 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36094 | 43.877 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR794148 | Hamiltonella phage APSE-8 | Hamiltonella phage APSE-8 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33476 | 44.465 | Sendosyvirus | Unclassified | Podoviridae | Hamiltonella |
| LR794147 | Bacteriophage APSE-7 | Bacteriophage APSE-7 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37264 | 43.935 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| LR794146 | Bacteriophage APSE-7 | Bacteriophage APSE-7 unclassified Sendosyvirus Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35191 | 43.838 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| MN938931 | Streptococcus phage D4446 | Streptococcus phage D4446 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33656 | 37.004 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MN070121 | Salmonella phage Shemara | Salmonella phage Shemara unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44342 | 51.143 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN095772 | Stenotrophomonas phage Moby | Stenotrophomonas phage Moby Stenotrophomonas virus Moby Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159365 | 54.137 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MN095770 | Serratia phage Slocum | Serratia phage Slocum unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112436 | 44.826 | Unclassified | Unclassified | Demerecviridae | Serratia |
| MN098328 | Stenotrophomonas phage Mendera | Stenotrophomonas phage Mendera Stenotrophomonas virus Mendera Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159961 | 54.015 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MN095771 | Serratia phage Muldoon | Serratia phage Muldoon Serratia virus Muldoon Muldoonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167547 | 41.802 | Muldoonvirus | Unclassified | Myoviridae | Serratia |
| MN098329 | Serratia phage Pila | Serratia phage Pila Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38678 | 48.767 | Teseptimavirus | Studiervirinae | Autographiviridae | Serratia |
| MN098327 | Klebsiella phage Mulock | Klebsiella phage Mulock Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43727 | 52.636 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MN098326 | Proteus phage Myduc | Proteus phage Myduc Proteus virus Myduc Myducvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53392 | 39.274 | Myducvirus | Cleopatravirinae | Chaseviridae | Proteus |
| MN098325 | Staphylococcus phage Portland | Staphylococcus phage Portland unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17711 | 29.293 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK050846 | Salmonella phage Seafire | Salmonella phage Seafire Salmonella virus Seafire Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111851 | 40.013 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN045228 | Staphylococcus phage Maine | Staphylococcus phage Maine unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141712 | 30.409 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK903282 | Escherichia phage Mansfield | Escherichia phage Mansfield unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68120 | 46.141 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MK903281 | Escherichia phage Penshu1 | Escherichia phage Penshu1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39263 | 50.559 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK903280 | Stenotrophomonas phage Ponderosa | Stenotrophomonas phage Ponderosa unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42612 | 59.976 | Unclassified | Okabevirinae | Autographiviridae | Stenotrophomonas |
| MK903279 | Escherichia phage Peacock | Escherichia phage Peacock unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39233 | 50.149 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK903278 | Xanthomonas phage Pagan | Xanthomonas phage Pagan unclassified Pradovirus Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44448 | 62.331 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| MK903277 | Escherichia phage Pisces | Escherichia phage Pisces unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39497 | 53.207 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN062189 | Serratia phage MyoSmar | Serratia phage MyoSmar Serratia virus MyoSmar Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68745 | 49.151 | Myosmarvirus | Unclassified | Myoviridae | Serratia |
| MN062188 | Proteus phage Saba | Proteus phage Saba Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60056 | 48.818 | Unclassified | Unclassified | Siphoviridae | Proteus |
| MN062187 | Xanthomonas phage Samson | Xanthomonas phage Samson unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43314 | 54.571 | Septimatrevirus | Unclassified | Siphoviridae | Xanthomonas |
| MN062186 | Stenotrophomonas phage Pokken | Stenotrophomonas phage Pokken Stenotrophomonas virus Pokken Pokkenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76239 | 55.111 | Pokkenvirus | Unclassified | Schitoviridae | Stenotrophomonas |
| MN062185 | Vibrio phage Phriendly | Vibrio phage Phriendly Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50218 | 41.386 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MN045231 | Escherichia phage Paul | Escherichia phage Paul unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79429 | 41.993 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MN045229 | Escherichia phage Mangalitsa | Escherichia phage Mangalitsa Escherichia virus Mangalitsa Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52329 | 44.012 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MK931446 | Salmonella phage Shelanagig | Salmonella phage Shelanagig unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42541 | 49.757 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK931445 | Klebsiella phage Shelby | Klebsiella phage Shelby Klebsiella virus Shelby Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49045 | 50.407 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK931444 | Klebsiella phage Skenny | Klebsiella phage Skenny Klebsiella virus Skenny Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49935 | 50.794 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK931443 | Klebsiella phage Sweeny | Klebsiella phage Sweeny Klebsiella virus Sweeny Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50241 | 50.672 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK931442 | Klebsiella phage Sin4 | Klebsiella phage Sin4 Klebsiella virus Sin4 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49916 | 50.288 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK931441 | Escherichia phage Snoke | Escherichia phage Snoke unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44454 | 50.191 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK931439 | Escherichia phage Sciku | Escherichia phage Sciku unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43130 | 50.1 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK931438 | Escherichia phage Schulenburg | Escherichia phage Schulenburg unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44748 | 49.996 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN066127 | Salmonella phage Matapan | Salmonella phage Matapan unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157408 | 44.937 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN044033 | Klebsiella phage Marfa | Klebsiella phage Marfa Klebsiella virus Marfa Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168532 | 40.993 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MK994515 | Serratia phage Moabite | Serratia phage Moabite Serratia virus Moabite Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273933 | 46.783 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| MK728825 | Salmonella phage Sepoy | Salmonella phage Sepoy Salmonella virus Sepoy Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114080 | 40.314 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK728824 | Salmonella phage Seabear | Salmonella phage Seabear unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112472 | 40.382 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK618657 | Klebsiella phage Sanco | Klebsiella phage Sanco unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48790 | 50.785 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MG459987 | Klebsiella phage Sugarland | Klebsiella phage Sugarland Klebsiella virus Sugarland Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111103 | 44.958 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MK608335 | Klebsiella phage Patroon | Klebsiella phage Patroon unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39442 | 50.54 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK618717 | Serratia phage MTx | Serratia phage MTx Serratia virus MTx Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68621 | 49.886 | Myosmarvirus | Unclassified | Myoviridae | Serratia |
| MK618716 | Staphylococcus phage Sebago | Staphylococcus phage Sebago Staphylococcus virus Sebago Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43878 | 35.314 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK618715 | Serratia phage Parlo | Serratia phage Parlo Serratia virus Parlo Parlovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62853 | 58.042 | Parlovirus | Unclassified | Podoviridae | Serratia |
| MK637516 | Agrobacterium phage Milano | Agrobacterium phage Milano unclassified Schmittlotzvirus Schmittlotzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68451 | 52.544 | Schmittlotzvirus | Unclassified | Myoviridae | Agrobacterium |
| MK598851 | Escherichia phage Minorna | Escherichia phage Minorna Escherichia virus Minorna Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43264 | 51.456 | Drulisvirus | Slopekvirinae | Autographiviridae | Escherichia |
| MK024806 | Proteus phage Mydo | Proteus phage Mydo Proteus virus Mydo Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145127 | 44.809 | Mydovirus | Vequintavirinae | Myoviridae | Proteus |
| MH920362 | Citrobacter phage Maleficent | Citrobacter phage Maleficent unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89570 | 38.875 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| MH899585 | Klebsiella phage Pylas | Klebsiella phage Pylas Klebsiella virus Pylas Pylasvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70408 | 41.213 | Pylasvirus | Humphriesvirinae | Schitoviridae | Klebsiella |
| MH830339 | Proteus phage Stubb | Proteus phage Stubb Proteus virus Stubb Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104410 | 38.512 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MH823906 | Citrobacter phage Maroon | Citrobacter phage Maroon unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178830 | 44.855 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| MH817999 | Klebsiella phage Seifer | Klebsiella phage Seifer Klebsiella virus Seifer Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58197 | 56.106 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MH586731 | Salmonella phage Meda | Salmonella phage Meda unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84668 | 38.846 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MH586730 | Salmonella phage Solent | Salmonella phage Solent unclassified Sashavirus Sashavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55978 | 43.469 | Sashavirus | Unclassified | Siphoviridae | Salmonella |
| MH729819 | Citrobacter phage Sazh | Citrobacter phage Sazh Citrobacter virus Sazh Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49665 | 42.811 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| MH729818 | Escherichia phage LL11 | Escherichia phage LL11 Escherichia virus LL11 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45185 | 45.072 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MH688040 | Salmonella phage Mooltan | Salmonella phage Mooltan unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156882 | 44.947 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH631453 | Salmonella phage Siskin | Salmonella phage Siskin Salmonella virus Siskin Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58476 | 56.604 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MH542234 | Staphylococcus phage Terranova | Staphylococcus phage Terranova unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141288 | 27.98 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH553517 | Serratia phage Scapp | Serratia phage Scapp Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42969 | 55.854 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MH321491 | Staphylococcus phage Twillingate | Staphylococcus phage Twillingate unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142592 | 27.987 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH321490 | Staphylococcus phage Quidividi | Staphylococcus phage Quidividi unclassified Sepunavirus Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141446 | 27.992 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH321493 | Salmonella phage Skate | Salmonella phage Skate Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47393 | 45.804 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH321492 | Staphylococcus phage Lorac | Staphylococcus phage Lorac unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43147 | 34.575 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MH491969 | Escherichia phage LL12 | Escherichia phage LL12 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136026 | 43.614 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MH333064 | Klebsiella phage Mineola | Klebsiella phage Mineola unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166130 | 39.513 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MH491968 | Escherichia phage LL5 | Escherichia phage LL5 Escherichia virus LL5 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49788 | 42.464 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MG596799 | Pseudomonas phage PMBT3 | Pseudomonas phage PMBT3 Pseudomonas virus PMBT3 Maxrubnervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87196 | 60.37 | Maxrubnervirus | Unclassified | Siphoviridae | Pseudomonas |
| MG428992 | Salmonella phage Mutine | Salmonella phage Mutine Salmonella virus Mutine Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161502 | 44.303 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MG428991 | Klebsiella phage May | Klebsiella phage May Klebsiella virus May Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159631 | 46.768 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MG428990 | Klebsiella phage Menlow | Klebsiella phage Menlow Klebsiella virus Menlow Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157281 | 46.422 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MF957259 | Salmonella phage Melville | Salmonella phage Melville Salmonella virus Melville Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159323 | 36.947 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| KY654690 | Citrobacter phage Mijalis | Citrobacter phage Mijalis unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87998 | 38.984 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| KY002061 | Salmonella phage vB_SenS_Sergei | Salmonella phage vB_SenS_Sergei unclassified Sashavirus Sashavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56051 | 43.493 | Sashavirus | Unclassified | Siphoviridae | Salmonella |
| KX987158 | Salmonella phage vB_SenS_Sasha | Salmonella phage vB_SenS_Sasha Salmonella virus Sasha Sashavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53263 | 43.732 | Sashavirus | Unclassified | Siphoviridae | Salmonella |
| KM236243 | Escherichia phage Seurat | Escherichia phage Seurat Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56776 | 44.575 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| MT178767 | Listeria virus P200 | Listeria virus P200 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135029 | 35.906 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MT178766 | Listeria virus P100plus | Listeria virus P100plus unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135028 | 35.91 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MT270409 | Phaeobacter phage MD18 | Phaeobacter phage MD18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149262 | 58.311 | Unclassified | Unclassified | Siphoviridae | Phaeobacter |
| MT150133 | Enterobacteria phage vB_EcoM_IME540 | Enterobacteria phage vB_EcoM_IME540 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170237 | 39.462 | Dhakavirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MK911750 | Streptococcus phage P738 | Streptococcus phage P738 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34037 | 37.113 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH791408 | Pseudomonas phage PaZq-1 | Pseudomonas phage PaZq-1 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94315 | 49.364 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT276994 | Aeromonas phage AhyVDH1 | Aeromonas phage AhyVDH1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39175 | 58.047 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MN692673 | Salmonella phage VB_StyS_BS5 | Salmonella phage VB_StyS_BS5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47604 | 45.526 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH791399 | Pseudomonas phage PaGz-1 | Pseudomonas phage PaGz-1 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93975 | 49.444 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK693005 | Klebsiella phage KPR2 | Klebsiella phage KPR2 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44067 | 53.743 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| LC541428 | Staphylococcus phage vB_SauS_JS02 | Staphylococcus phage vB_SauS_JS02 Staphylococcus virus JS02 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46435 | 33.143 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MT188223 | Acinetobacter phage vB_AbaM_D22 | Acinetobacter phage vB_AbaM_D22 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103539 | 37.446 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MT179807 | Escherichia phage vB_EcoM_IME537 | Escherichia phage vB_EcoM_IME537 Escherichia virus IME537 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168642 | 35.425 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MT135026 | Vibrio phage V09 | Vibrio phage V09 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 243881 | 42.636 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MT135025 | Vibrio phage V07 | Vibrio phage V07 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 349331 | 43.139 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MT135024 | Vibrio phage V05 | Vibrio phage V05 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157241 | 42.68 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MT121968 | Pseudomonas phage SC_10_H1H2H8_2017 | Pseudomonas phage SC_10_H1H2H8_2017 unclassified bacterial viruses Viruses | 11903 | 54.289 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| MT121967 | Pseudomonas phage SC_9_H1H8_2017 | Pseudomonas phage SC_9_H1H8_2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20910 | 59.445 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT121966 | Pseudomonas phage SC_8_H2H8_2017 | Pseudomonas phage SC_8_H2H8_2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47543 | 58.097 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT121964 | Phage DP SC_6_H4_2017 | Phage DP SC_6_H4_2017 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75500 | 35.189 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MT121960 | Phage DP SC_2_H4_2017 | Phage DP SC_2_H4_2017 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41288 | 33.516 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MT259468 | Aeromonas phage PS | Aeromonas phage PS unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41082 | 48.527 | Unclassified | Unclassified | Autographiviridae | Aeromonas |
| MT254578 | Bacillus phage Izhevsk | Bacillus phage Izhevsk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168638 | 34.319 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MT210154 | Xanthomonas phage PPDBI | Xanthomonas phage PPDBI Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39980 | 55.268 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MT151386 | Staphylococcus phage Twort | Staphylococcus phage Twort Twortvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139739 | 30.127 | Twortvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT227924 | Vibrio phage phiV208 | Vibrio phage phiV208 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45521 | 43.16 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MT176427 | Escherichia phage CJ19 | Escherichia phage CJ19 unclassified Nouzillyvirus Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49567 | 46.874 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| MT178448 | Vibrio phage vB_VpP_FE11 | Vibrio phage vB_VpP_FE11 unclassified Maculvirus Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43397 | 49.238 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MT251348 | Klebsiella phage SJM3 | Klebsiella phage SJM3 unclassified Lastavirus Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62420 | 56.235 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| MT251347 | Klebsiella phage LASTA | Klebsiella phage LASTA Klebsiella virus LASTA Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62420 | 56.237 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| MT197175 | Klebsiella phage VLC5 | Klebsiella phage VLC5 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44932 | 54.175 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MT263719 | Acinetobacter virus vB_AbaP_AGC01 | Acinetobacter virus vB_AbaP_AGC01 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41231 | 39.485 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT157285 | Klebsiella phage P-KP2 | Klebsiella phage P-KP2 unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172138 | 41.884 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN882554 | Butyrivibrio virus Idris | Butyrivibrio virus Idris Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31128 | 46.919 | Unclassified | Unclassified | Siphoviridae | Butyrivibrio |
| MN882553 | Butyrivibrio virus Ceridwen | Butyrivibrio virus Ceridwen Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39745 | 40.828 | Unclassified | Unclassified | Siphoviridae | Butyrivibrio |
| MN882552 | Butyrivibrio virus Bo-Finn | Butyrivibrio virus Bo-Finn Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33227 | 39.697 | Unclassified | Unclassified | Siphoviridae | Butyrivibrio |
| MN882551 | Butyrivibrio virus Arian | Butyrivibrio virus Arian Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33499 | 39.738 | Unclassified | Unclassified | Siphoviridae | Butyrivibrio |
| MN882550 | Butyrivibrio virus Arawn | Butyrivibrio virus Arawn Arawnvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31118 | 46.944 | Arawnvirus | Unclassified | Siphoviridae | Butyrivibrio |
| MT188704 | Mesorhizobium phage Cp1R7A-A1 | Mesorhizobium phage Cp1R7A-A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131434 | 60.339 | Unclassified | Unclassified | Siphoviridae | Mesorhizobium |
| MK241539 | Alteromonas phage P24 | Alteromonas phage P24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46945 | 43.796 | Unclassified | Unclassified | Siphoviridae | Alteromonas |
| MN901521 | Halorubrum virus Serpecor1 | Halorubrum virus Serpecor1 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74196 | 57.027 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MN901520 | Halorubrum virus Hardycor2 | Halorubrum virus Hardycor2 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77342 | 55.641 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MN241318 | Enterococcus phage PEf771 | Enterococcus phage PEf771 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151052 | 36.967 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MK249308 | Salmonella phage TS13 | Salmonella phage TS13 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59168 | 56.356 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MK249126 | Salmonella phage TS3 | Salmonella phage TS3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41568 | 49.887 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK214385 | Salmonella phage TS6 | Salmonella phage TS6 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41515 | 51.131 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK819239 | Pseudomonas phage vB_PaeP_YA3 | Pseudomonas phage vB_PaeP_YA3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45253 | 57.176 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MT230534 | Pantoea phage vB_PagM_SSEM1 | Pantoea phage vB_PagM_SSEM1 Pantoea virus SSEM1 Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54982 | 44.182 | Loessnervirus | Cleopatravirinae | Chaseviridae | Pantoea |
| MT094431 | Pseudomonas phage BIM BV-46 | Pseudomonas phage BIM BV-46 unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38860 | 56.413 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MT094430 | Pseudomonas phage BIM BV-45 | Pseudomonas phage BIM BV-45 unclassified Bifseptvirus Bifseptvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40565 | 58.146 | Bifseptvirus | Unclassified | Autographiviridae | Pseudomonas |
| MN379651 | Chicken microvirus mg8_109 | Chicken microvirus mg8_109 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4743 | 39.996 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379650 | Chicken microvirus mg8_102 | Chicken microvirus mg8_102 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4853 | 42.654 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379649 | Chicken microvirus mg8_95 | Chicken microvirus mg8_95 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4992 | 49.259 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379648 | Chicken microvirus mg8_48 | Chicken microvirus mg8_48 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6328 | 39.87 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379647 | Chicken microvirus mg8_45 | Chicken microvirus mg8_45 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6390 | 40.141 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379646 | Chicken microvirus mg8_41 | Chicken microvirus mg8_41 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6710 | 39.21 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379645 | Chicken microvirus mg7_25 | Chicken microvirus mg7_25 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4253 | 41.923 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379644 | Chicken microvirus mg7_20 | Chicken microvirus mg7_20 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4722 | 48.052 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379643 | Chicken microvirus mg7_19 | Chicken microvirus mg7_19 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4877 | 42.198 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379642 | Chicken microvirus mg7_16 | Chicken microvirus mg7_16 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4939 | 50.375 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379641 | Chicken microvirus mg7_14 | Chicken microvirus mg7_14 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5037 | 44.65 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379640 | Chicken microvirus mg7_10 | Chicken microvirus mg7_10 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5777 | 50.649 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379639 | Chicken microvirus mg7_8 | Chicken microvirus mg7_8 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6110 | 39.722 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379638 | Chicken microvirus mg7_7 | Chicken microvirus mg7_7 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6308 | 42.074 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379637 | Chicken microvirus mg7_6 | Chicken microvirus mg7_6 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6385 | 39.561 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379636 | Chicken microvirus mg7_1 | Chicken microvirus mg7_1 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6553 | 39.753 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379635 | Chicken microvirus mg6_218 | Chicken microvirus mg6_218 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4510 | 45.831 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379634 | Chicken microvirus mg6_154 | Chicken microvirus mg6_154 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5058 | 46.105 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379633 | Chicken microvirus mg5_146 | Chicken microvirus mg5_146 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4831 | 48.706 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379632 | Chicken microvirus mg4_207 | Chicken microvirus mg4_207 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4728 | 51.671 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379631 | Chicken microvirus mg4_164 | Chicken microvirus mg4_164 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5148 | 49.204 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379630 | Chicken microvirus mg4_115 | Chicken microvirus mg4_115 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5765 | 51.622 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN379629 | Chicken microvirus mg4_76 | Chicken microvirus mg4_76 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6541 | 39.749 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN106245 | Klebsiella phage EI | Klebsiella phage EI unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170423 | 40.656 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MN045230 | Klebsiella phage Magnus | Klebsiella phage Magnus Klebsiella virus Magnus Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157741 | 46.261 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MK618658 | Klebsiella phage Pharr | Klebsiella phage Pharr Klebsiella virus Pharr Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40599 | 53.267 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK630230 | Klebsiella phage Spivey | Klebsiella phage Spivey unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110659 | 44.911 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MK268344 | Salmonella phage Munch | Salmonella phage Munch Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 350103 | 35.549 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KT363872 | Citrobacter phage Mordin | Citrobacter phage Mordin Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89596 | 38.808 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| KT001920 | Klebsiella phage Sushi | Klebsiella phage Sushi Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48754 | 50.769 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KT001911 | Bacillus phage Pavlov | Bacillus phage Pavlov unclassified Pagevirus Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40024 | 40.621 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KP411017 | Bacillus phage Palmer | Bacillus phage Palmer Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40000 | 40.668 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KP143762 | Salmonella phage Mushroom | Salmonella phage Mushroom Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87709 | 39.035 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KP143763 | Salmonella phage Shivani | Salmonella phage Shivani Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120098 | 38.839 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KM236242 | Escherichia phage Pollock | Escherichia phage Pollock Pollockvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68365 | 36.02 | Pollockvirus | Humphriesvirinae | Schitoviridae | Escherichia |
| KM236240 | Citrobacter phage Moon | Citrobacter phage Moon Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170341 | 38.876 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| MN994499 | Phage NBEco005 | Phage NBEco005 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88272 | 38.841 | Felixounavirus | Ounavirinae | Myoviridae | Unspecified |
| MT176424 | Citrobacter phage PhiZZ6 | Citrobacter phage PhiZZ6 Citrobacter virus PhiZZ6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165795 | 35.432 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| MT136606 | Bacillus phage vB_BceS_KLEB30-3S | Bacillus phage vB_BceS_KLEB30-3S unclassified Lwoffvirus Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37134 | 38.251 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| MT119373 | Pseudomonas phage otherone | Pseudomonas phage otherone unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44930 | 52.081 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT118305 | Pseudomonas phage Epa41 | Pseudomonas phage Epa41 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43258 | 53.209 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MT118304 | Pseudomonas phage Epa40 | Pseudomonas phage Epa40 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42788 | 53.195 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MT118303 | Pseudomonas phage Epa39 | Pseudomonas phage Epa39 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66708 | 54.939 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118302 | Pseudomonas phage Epa38 | Pseudomonas phage Epa38 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61775 | 64.282 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MT118301 | Pseudomonas phage Epa33 | Pseudomonas phage Epa33 unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64021 | 63.482 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MT118300 | Pseudomonas phage Epa26 | Pseudomonas phage Epa26 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88805 | 54.809 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MT118299 | Pseudomonas phage Epa25 | Pseudomonas phage Epa25 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66811 | 55.561 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118298 | Pseudomonas phage Epa21 | Pseudomonas phage Epa21 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66764 | 55.632 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118297 | Pseudomonas phage Epa20 | Pseudomonas phage Epa20 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66505 | 55.617 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118296 | Pseudomonas phage Epa19 | Pseudomonas phage Epa19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32090 | 61.421 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT118295 | Pseudomonas phage Epa18 | Pseudomonas phage Epa18 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83260 | 54.726 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MT118294 | Pseudomonas phage Epa16 | Pseudomonas phage Epa16 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88727 | 54.77 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MT118293 | Pseudomonas phage Epa14 | Pseudomonas phage Epa14 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65797 | 55.282 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118292 | Pseudomonas phage Epa13 | Pseudomonas phage Epa13 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65680 | 55.504 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118291 | Pseudomonas phage Epa12 | Pseudomonas phage Epa12 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66520 | 55.666 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118290 | Pseudomonas phage Epa10 | Pseudomonas phage Epa10 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66774 | 55.697 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118289 | Pseudomonas phage Epa7 | Pseudomonas phage Epa7 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65629 | 55.457 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT118288 | Pseudomonas phage Epa4 | Pseudomonas phage Epa4 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45439 | 52.499 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT108723 | Pseudomonas phage Epa1 | Pseudomonas phage Epa1 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45230 | 52.145 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT108724 | Pseudomonas phage Epa2 | Pseudomonas phage Epa2 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43299 | 52.334 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MT108725 | Pseudomonas phage Epa5 | Pseudomonas phage Epa5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64096 | 62.247 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT108727 | Pseudomonas phage Epa11 | Pseudomonas phage Epa11 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66800 | 55.717 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT108728 | Pseudomonas phage Epa17 | Pseudomonas phage Epa17 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88859 | 54.765 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MT108729 | Pseudomonas phage Epa22 | Pseudomonas phage Epa22 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65897 | 55.426 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MT108730 | Pseudomonas phage Epa24 | Pseudomonas phage Epa24 unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88728 | 54.771 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MT108731 | Pseudomonas phage Epa43 | Pseudomonas phage Epa43 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64323 | 62.032 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MT084350 | Xylella phage Xfas53 | Xylella phage Xfas53 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35839 | 56.868 | Unclassified | Unclassified | Podoviridae | Xylella |
| MT074478 | Salmonella phage moki | Salmonella phage moki Salmonella virus moki Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158230 | 44.42 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074472 | Salmonella phage bering | Salmonella phage bering Salmonella virus bering Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156130 | 44.703 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074469 | Salmonella phage rabagast | Salmonella phage rabagast Salmonella virus rabagast Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143249 | 44.529 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074450 | Salmonella phage faergetype | Salmonella phage faergetype Salmonella virus faergetype Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110183 | 39.957 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN956515 | Vibrio phage V039C | Vibrio phage V039C Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43405 | 42.954 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN604698 | Bacillus phage vB_BcM_Sam46 | Bacillus phage vB_BcM_Sam46 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45419 | 41.65 | Unclassified | Unclassified | Myoviridae | Bacillus |
| MT185428 | Escherichia phage EC6098 | Escherichia phage EC6098 Enterogokushovirus EC6098 Enterogokushovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4526 | 48.387 | Enterogokushovirus | Gokushovirinae | Microviridae | Escherichia |
| MT090077 | Klebsiella phage CX1 | Klebsiella phage CX1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45013 | 53.882 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN882610 | Enterobacter phage vB_EclM_CIP9 | Enterobacter phage vB_EclM_CIP9 Enterobacter virus CIP9 Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174924 | 39.934 | Kanagawavirus | Unclassified | Myoviridae | Enterobacter |
| MN013084 | Klebsiella phage vB_KaeM_KaAlpha | Klebsiella phage vB_KaeM_KaAlpha unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172334 | 39.846 | Karamvirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN013077 | Klebsiella phage vB_KaeM_KaOmega | Klebsiella phage vB_KaeM_KaOmega unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149489 | 50.454 | Certrevirus | Vequintavirinae | Myoviridae | Klebsiella |
| MK473373 | Pseudomonas phage Lana | Pseudomonas phage Lana Pseudomonas virus Lana Lanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88403 | 60.825 | Lanavirus | Unclassified | Siphoviridae | Pseudomonas |
| JX080302 | Staphylococcus phage 676Z | Staphylococcus phage 676Z Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148564 | 30.405 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX080301 | Staphylococcus phage A3R | Staphylococcus phage A3R Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141018 | 30.498 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT074480 | Salmonella phage maane | Salmonella phage maane Salmonella virus maane Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159128 | 45.159 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074479 | Salmonella phage pertopsoe | Salmonella phage pertopsoe Salmonella virus pertopsoe Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158905 | 45.279 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074477 | Salmonella phage barely | Salmonella phage barely Salmonella virus barely Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157920 | 44.598 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074475 | Salmonella phage dinky | Salmonella phage dinky Salmonella virus dinky Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157254 | 44.768 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074474 | Salmonella phage aagejoakim | Salmonella phage aagejoakim Salmonella virus aagejoakim Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156801 | 44.491 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074471 | Salmonella phage allotria | Salmonella phage allotria Salmonella virus allotria Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151015 | 45.12 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074470 | Salmonella phage heyday | Salmonella phage heyday Salmonella virus heyday Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144232 | 44.47 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT074466 | Salmonella phage atrejo | Salmonella phage atrejo Salmonella virus atrejo Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114059 | 39.971 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074465 | Salmonella phage oselot | Salmonella phage oselot Salmonella virus oselot Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113503 | 39.94 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074459 | Salmonella phage bastian | Salmonella phage bastian Salmonella virus bastian Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112477 | 39.961 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074458 | Salmonella phage fuchur | Salmonella phage fuchur Salmonella virus fuchur Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112117 | 40.017 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074457 | Salmonella phage rokbiter | Salmonella phage rokbiter Salmonella virus rokbiter Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112006 | 39.926 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074449 | Salmonella phage bombadil | Salmonella phage bombadil Salmonella virus bombadil Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109539 | 40.005 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074448 | Salmonella phage oldekolle | Salmonella phage oldekolle Salmonella virus oldekolle Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109372 | 39.285 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MT074439 | Salmonella phage birk | Salmonella phage birk Salmonella virus birk Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52767 | 45.845 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074436 | Salmonella phage yarpen | Salmonella phage yarpen Salmonella virus yarpen Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52464 | 45.961 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MT074429 | Salmonella phage astrithr | Salmonella phage astrithr Salmonella virus astrithr Astrithrvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11569 | 39.727 | Astrithrvirus | Unclassified | Podoviridae | Salmonella |
| MT176426 | Serratia phage PhiZZ30 | Serratia phage PhiZZ30 Serratia virus PhiZZ30 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167484 | 35.345 | Tequatrovirus | Tevenvirinae | Myoviridae | Serratia |
| MT176425 | Citrobacter phage PhiZZ23 | Citrobacter phage PhiZZ23 Citrobacter virus PhiZZ23 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167516 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| MN966732 | Vibrio phage Chester | Vibrio phage Chester unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126550 | 40.237 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MT119766 | Xanthomonas phage PBR31 | Xanthomonas phage PBR31 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39980 | 55.268 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| MT110073 | Stenotrophomonas phage vB_SmaS_DLP_3 | Stenotrophomonas phage vB_SmaS_DLP_3 unclassified Delepquintavirus Delepquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96852 | 58.304 | Delepquintavirus | Unclassified | Siphoviridae | Stenotrophomonas |
| MT028297 | Proteus phage Privateer | Proteus phage Privateer Proteus virus Privateer Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90710 | 34.523 | Privateervirus | Unclassified | Podoviridae | Proteus |
| MT024867 | Arthrobacter phage BlueFeather | Arthrobacter phage BlueFeather Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16302 | 64.293 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MT024864 | Microbacterium phage Oxtober96 | Microbacterium phage Oxtober96 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41798 | 63.376 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MT024862 | Microbacterium phage Alakazam | Microbacterium phage Alakazam unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41269 | 64.378 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MN994504 | Phage NBSal005 | Phage NBSal005 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121040 | 39.123 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MN994503 | Phage NBSal004 | Phage NBSal004 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86841 | 38.812 | Felixounavirus | Ounavirinae | Myoviridae | Unspecified |
| MN994502 | Phage NBSal003 | Phage NBSal003 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 119416 | 39.561 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MN994501 | Phage NBSal002 | Phage NBSal002 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120250 | 40.087 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MN994500 | Phage NBSal001 | Phage NBSal001 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50922 | 42.178 | Tlsvirus | Tempevirinae | Drexlerviridae | Unspecified |
| MN994498 | Phage NBEco004 | Phage NBEco004 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85516 | 38.904 | Felixounavirus | Ounavirinae | Myoviridae | Unspecified |
| MN994497 | Phage NBEco003 | Phage NBEco003 Escherichia virus NBEco003 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166579 | 35.515 | Tequatrovirus | Tevenvirinae | Myoviridae | Unspecified |
| MN994496 | Phage NBEco002 | Phage NBEco002 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118118 | 39.077 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MN970211 | Phage NBEco001 | Phage NBEco001 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116606 | 39.489 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MT047590 | Thermoproteus spherical piliferous virus 1 | Thermoproteus spherical piliferous virus 1 Alphaglobulovirus TSPV1 Alphaglobulovirus Globuloviridae Viruses | 18655 | 56.467 | Alphaglobulovirus | Unclassified | Globuloviridae | Thermoproteus |
| MG983742 | Bacillus phage Anath | Bacillus phage Anath Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52369 | 41.139 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MG676223 | Vibrio phage ValSw3-3 | Vibrio phage ValSw3-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39846 | 43.154 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MN732896 | Elizabethkingia phage TCUEAP1 | Elizabethkingia phage TCUEAP1 unclassified bacterial viruses Viruses | 49816 | 39.343 | Unclassified | Unclassified | Unclassified | Elizabethkingia |
| MG922974 | Klebsiella phage KP8 | Klebsiella phage KP8 Klebsiella virus KP8 Kaypoctavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73679 | 44.201 | Kaypoctavirus | Enquatrovirinae | Schitoviridae | Klebsiella |
| CP025874 | Escherichia phage 503458 | Escherichia phage 503458 Escherichia virus 503458 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31194 | 53.446 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| CP025887 | Escherichia phage 503046 | Escherichia phage 503046 unclassified bacterial viruses Viruses | 23083 | 50.786 | Unclassified | Unclassified | Unclassified | Escherichia |
| CP025902 | Escherichia phage 500465-3 | Escherichia phage 500465-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13549 | 55.023 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| CP025901 | Escherichia phage 500465-2 | Escherichia phage 500465-2 Escherichia virus 500465-2 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33041 | 52.441 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| CP025900 | Escherichia phage 500465-1 | Escherichia phage 500465-1 Escherichia virus 500465-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39179 | 52.651 | Felsduovirus | Peduovirinae | Myoviridae | Escherichia |
| CP025923 | Escherichia phage 520873 | Escherichia phage 520873 Escherichia virus 520873 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37583 | 52.005 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| LR778216 | Xylella phage Cota | Xylella phage Cota unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42307 | 58.848 | Unclassified | Okabevirinae | Autographiviridae | Xylella |
| MN966731 | Vibrio phage Chazly21 | Vibrio phage Chazly21 unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127243 | 40.092 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MN966730 | Vibrio phage Cilsick | Vibrio phage Cilsick unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126526 | 40.232 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MN974282 | Vibrio phage OWB | Vibrio phage OWB Vibrio virus OWB Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43264 | 48.74 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MT080603 | Staphylococcus phage vB_SauM_EW9 | Staphylococcus phage vB_SauM_EW9 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141953 | 30.781 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080602 | Staphylococcus phage vB_SauM_EW74 | Staphylococcus phage vB_SauM_EW74 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139896 | 30.435 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080601 | Staphylococcus phage vB_SauM_EW72 | Staphylococcus phage vB_SauM_EW72 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143240 | 30.278 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080600 | Staphylococcus phage vB_SauM_EW71 | Staphylococcus phage vB_SauM_EW71 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139939 | 30.427 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080599 | Staphylococcus phage vB_SauM_EW7 | Staphylococcus phage vB_SauM_EW7 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143288 | 30.778 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080598 | Staphylococcus phage vB_SauM_EW6 | Staphylococcus phage vB_SauM_EW6 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143283 | 30.785 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080597 | Staphylococcus phage vB_SauM_EW5 | Staphylococcus phage vB_SauM_EW5 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143283 | 30.783 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080596 | Staphylococcus phage vB_SauM_EW42 | Staphylococcus phage vB_SauM_EW42 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139874 | 31.395 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080595 | Staphylococcus phage vB_SauM_EW41 | Staphylococcus phage vB_SauM_EW41 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132999 | 29.973 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080594 | Staphylococcus phage vB_SauM_EW4 | Staphylococcus phage vB_SauM_EW4 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140906 | 30.833 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080593 | Staphylococcus phage vB_SauM_EW36 | Staphylococcus phage vB_SauM_EW36 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139881 | 31.397 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080592 | Staphylococcus phage vB_SauM_EW3 | Staphylococcus phage vB_SauM_EW3 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143283 | 30.781 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080591 | Staphylococcus phage vB_SauM_EW29 | Staphylococcus phage vB_SauM_EW29 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143240 | 30.269 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080590 | Staphylococcus phage vB_SauM_EW27 | Staphylococcus phage vB_SauM_EW27 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135801 | 29.889 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080589 | Staphylococcus phage vB_SauM_EW26 | Staphylococcus phage vB_SauM_EW26 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143240 | 30.279 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080588 | Staphylococcus phage vB_SauM_EW22 | Staphylococcus phage vB_SauM_EW22 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135820 | 29.873 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080587 | Staphylococcus phage vB_SauM_EW20 | Staphylococcus phage vB_SauM_EW20 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135784 | 29.875 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080586 | Staphylococcus phage vB_SauM_EW2 | Staphylococcus phage vB_SauM_EW2 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140906 | 30.831 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080585 | Staphylococcus phage vB_SauM_EW18 | Staphylococcus phage vB_SauM_EW18 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143247 | 30.278 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080584 | Staphylococcus phage vB_SauM_EW15 | Staphylococcus phage vB_SauM_EW15 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135802 | 29.888 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080583 | Staphylococcus phage vB_SauM_EW13 | Staphylococcus phage vB_SauM_EW13 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145736 | 30.726 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT080582 | Staphylococcus phage vB_SauM_EW1 | Staphylococcus phage vB_SauM_EW1 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140906 | 30.833 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MT028492 | Ochrobactrum phage vB_OspP_OH | Ochrobactrum phage vB_OspP_OH Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41227 | 55.197 | Unclassified | Unclassified | Podoviridae | Ochrobactrum |
| MT028491 | Ochrobactrum phage vB_OspM_OC | Ochrobactrum phage vB_OspM_OC Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 227654 | 37.678 | Unclassified | Unclassified | Myoviridae | Ochrobactrum |
| MT001829 | Salmonella phage SE_PL | Salmonella phage SE_PL Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348657 | 35.582 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MT104122 | Sporosarcina phage Lietuvens | Sporosarcina phage Lietuvens Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64517 | 49.854 | Unclassified | Unclassified | Myoviridae | Sporosarcina |
| MT066160 | Vibrio phage Saratov-12 | Vibrio phage Saratov-12 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48368 | 42.849 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MN995824 | Enterococcus phage EFP1 | Enterococcus phage EFP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37561 | 37.621 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MT078988 | Salmonella phage P46FS4 | Salmonella phage P46FS4 Salmonella virus P46FS4 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158706 | 50.444 | Agtrevirus | Aglimvirinae | Ackermannviridae | Salmonella |
| MT023084 | Citrobacter phage IME-JL8 | Citrobacter phage IME-JL8 unclassified Sertoctavirus Sertoctavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49838 | 47.963 | Sertoctavirus | Tunavirinae | Drexlerviridae | Citrobacter |
| MN956514 | Salmonella phage L6jm | Salmonella phage L6jm Salmonella virus L6jm Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118452 | 39.249 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN953776 | Salmonella phage Chennai | Salmonella phage Chennai unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157462 | 44.587 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN927226 | Escherichia phage vB_EcoS_XY2 | Escherichia phage vB_EcoS_XY2 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44138 | 50.646 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN927225 | Escherichia phage vB_EcoS_XY1 | Escherichia phage vB_EcoS_XY1 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45566 | 50.509 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN908686 | Microbacterium phage Zepp | Microbacterium phage Zepp unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40189 | 63.59 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MN864145 | Escherichia phage F2 | Escherichia phage F2 Escherichia virus F2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168241 | 37.324 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN722429 | Salmonella phage vB_SpuP_Spp11 | Salmonella phage vB_SpuP_Spp11 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42372 | 49.589 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN552148 | Lactococcus phage P1532 | Lactococcus phage P1532 Lactococcus virus P1532 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29075 | 35.384 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN536026 | Pseudomonas phage vB_Pae-SS2019XI | Pseudomonas phage vB_Pae-SS2019XI Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57567 | 60.132 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KT803877 | Burkholderia phage PE067 | Burkholderia phage PE067 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43649 | 64.483 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MN176217 | Escherichia phage vB_Eco4M-7 | Escherichia phage vB_Eco4M-7 unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68084 | 46.181 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MN336263 | Staphylococcus phage Sb1M_9832 | Staphylococcus phage Sb1M_9832 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138231 | 30.425 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MN336262 | Staphylococcus phage Sb1M_6168 | Staphylococcus phage Sb1M_6168 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138231 | 30.425 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MN336261 | Staphylococcus phage Sb1_8383 | Staphylococcus phage Sb1_8383 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139606 | 30.399 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH745368 | Escherichia phage fp01 | Escherichia phage fp01 Escherichia virus fp01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109515 | 39.003 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MH779524 | Lactococcus phage vB_Llc_bIBBD4 | Lactococcus phage vB_Llc_bIBBD4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21907 | 35.998 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779527 | Lactococcus phage vB_Llc_bIBBp6/4 | Lactococcus phage vB_Llc_bIBBp6/4 Lactococcus virus bIBBp6-4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22220 | 35.482 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779526 | Lactococcus phage vB_Llc_bIBBL81 | Lactococcus phage vB_Llc_bIBBL81 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21915 | 36.003 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779525 | Lactococcus phage vB_Llc_bIBBE1 | Lactococcus phage vB_Llc_bIBBE1 Lactococcus virus bIBBE1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21749 | 36.043 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779523 | Lactococcus phage vB_Llc_bIBBAm4 | Lactococcus phage vB_Llc_bIBBAm4 Lactococcus virus bIBBAm4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22703 | 35.872 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779522 | Lactococcus phage vB_Llc_bIBBA3 | Lactococcus phage vB_Llc_bIBBA3 Lactococcus virus bIBBA3 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21906 | 36.159 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779521 | Lactococcus phage vB_Llc_bIBB94p4 | Lactococcus phage vB_Llc_bIBB94p4 Lactococcus virus blBB94p4 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21834 | 36.077 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779520 | Lactococcus phage vB_Llc_bIBB27 | Lactococcus phage vB_Llc_bIBB27 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21713 | 36.103 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779519 | Lactococcus phage vB_LLc_bIBB14 | Lactococcus phage vB_LLc_bIBB14 Lactococcus virus bIBB14 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21710 | 36.039 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH779518 | Lactococcus phage vB_Llc_bIBB12 | Lactococcus phage vB_Llc_bIBB12 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21458 | 36.206 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX438380 | Salmonella phage vB-SalM-PM10 | Salmonella phage vB-SalM-PM10 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158081 | 44.608 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MT006240 | Bifidobacterium phage BadAargau2 | Bifidobacterium phage BadAargau2 Bifidobacterium virus BadAargau2 Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18899 | 50.426 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| MT006239 | Bifidobacterium phage BadAztec2 | Bifidobacterium phage BadAztec2 unclassified Badaztecvirus Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18689 | 48.729 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| MT006238 | Bifidobacterium phage BadAztec4 | Bifidobacterium phage BadAztec4 unclassified Badaztecvirus Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19802 | 48.889 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| MT006237 | Bifidobacterium phage BigBern1 | Bifidobacterium phage BigBern1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39992 | 61.502 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| MT006236 | Bifidobacterium phage BitterVaud1 | Bifidobacterium phage BitterVaud1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51699 | 49.485 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| MT006235 | Bifidobacterium phage BlindBasel1 | Bifidobacterium phage BlindBasel1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31857 | 51.289 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| MT006234 | Bifidobacterium phage BlindUri1 | Bifidobacterium phage BlindUri1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32266 | 51.472 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| MT006233 | Bifidobacterium phage BadAztec1 | Bifidobacterium phage BadAztec1 Bifidobacterium virus BadAztec1 Badaztecvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18635 | 48.801 | Badaztecvirus | Unclassified | Podoviridae | Bifidobacterium |
| MN512538 | Vibrio phage Seahorse | Vibrio phage Seahorse Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45171 | 42.591 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK574078 | Pseudomonas phage KP1 | Pseudomonas phage KP1 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43470 | 46.648 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MN414250 | Escherichia phage RDN8.1 | Escherichia phage RDN8.1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39508 | 51.643 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN227145 | Salmonella phage BPSELC-1 | Salmonella phage BPSELC-1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86996 | 38.814 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MN813695 | Arthrobacter phage LouisXIV | Arthrobacter phage LouisXIV unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.118 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN703411 | Arthrobacter phage DrYang | Arthrobacter phage DrYang Arthrobacter virus DrYang Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69061 | 62.173 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MN617842 | Arthrobacter phage Saphira | Arthrobacter phage Saphira unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15549 | 59.985 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN234234 | Arthrobacter phage Lymara | Arthrobacter phage Lymara Arthrobacter virus Lymara Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68762 | 61.492 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MN204503 | Arthrobacter phage Linus | Arthrobacter phage Linus unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70526 | 61.657 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MN204499 | Arthrobacter phage Mordred | Arthrobacter phage Mordred unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70074 | 61.671 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MK820640 | Arthrobacter phage Laila | Arthrobacter phage Laila unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 59.63 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MK737943 | Arthrobacter phage Arby | Arthrobacter phage Arby unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MK494111 | Arthrobacter phage StewieGriff | Arthrobacter phage StewieGriff unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 59.643 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MK494107 | Arthrobacter phage Chipper1996 | Arthrobacter phage Chipper1996 unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70090 | 61.638 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MK494103 | Arthrobacter phage Blair | Arthrobacter phage Blair unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 59.643 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MK574011 | Edwardsiella phage ETP-1 | Edwardsiella phage ETP-1 unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41785 | 48.266 | Kafunavirus | Unclassified | Podoviridae | Edwardsiella |
| MH779506 | Arthrobacter phage Guntur | Arthrobacter phage Guntur unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.118 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH744419 | Arthrobacter phage KBurrousTX | Arthrobacter phage KBurrousTX Arthrobacter virus KBurrousTX Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71020 | 62.295 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MH576961 | Arthrobacter phage Ronnie | Arthrobacter phage Ronnie unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576957 | Arthrobacter phage Inspire2 | Arthrobacter phage Inspire2 unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576955 | Arthrobacter phage Hunnie | Arthrobacter phage Hunnie unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.112 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576952 | Arthrobacter phage Dewayne | Arthrobacter phage Dewayne unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.118 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576951 | Arthrobacter phage Copper | Arthrobacter phage Copper unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.118 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576950 | Arthrobacter phage CGermain | Arthrobacter phage CGermain unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.118 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MH576948 | Arthrobacter phage Azathoth | Arthrobacter phage Azathoth unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF668274 | Arthrobacter phage JayCookie | Arthrobacter phage JayCookie unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70362 | 61.672 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF185725 | Arthrobacter phage Tophat | Arthrobacter phage Tophat unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70091 | 61.64 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF185724 | Arthrobacter phage Scavito | Arthrobacter phage Scavito unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70123 | 61.636 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF185723 | Arthrobacter phage RosiePosie | Arthrobacter phage RosiePosie unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70396 | 61.68 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF185721 | Arthrobacter phage Kabreeze | Arthrobacter phage Kabreeze unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70035 | 61.693 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF185718 | Arthrobacter phage Colucci | Arthrobacter phage Colucci Arthrobacter virus Colucci Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70707 | 61.701 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| MF140429 | Arthrobacter phage Swenson | Arthrobacter phage Swenson unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15680 | 59.885 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140426 | Arthrobacter phage Seume | Arthrobacter phage Seume unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15319 | 60.272 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140420 | Arthrobacter phage Mariposa | Arthrobacter phage Mariposa unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.112 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140419 | Arthrobacter phage Lore | Arthrobacter phage Lore unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 59.643 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140417 | Arthrobacter phage Link | Arthrobacter phage Link unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15521 | 60.177 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140415 | Arthrobacter phage KylieMac | Arthrobacter phage KylieMac unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15540 | 59.762 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140409 | Arthrobacter phage Elkhorn | Arthrobacter phage Elkhorn unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 59.643 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140433 | Arthrobacter phage TinoCrisci | Arthrobacter phage TinoCrisci unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MF140431 | Arthrobacter phage Taj14 | Arthrobacter phage Taj14 unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15546 | 59.9 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KY434670 | Arthrobacter phage Chestnut | Arthrobacter phage Chestnut unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KX855962 | Arthrobacter phage HumptyDumpty | Arthrobacter phage HumptyDumpty unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69978 | 61.689 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KX855961 | Arthrobacter phage EdgarPoe | Arthrobacter phage EdgarPoe unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70176 | 61.639 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KX670787 | Arthrobacter phage Chocolat | Arthrobacter phage Chocolat unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69798 | 61.701 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KX670786 | Arthrobacter phage Chubster | Arthrobacter phage Chubster unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70258 | 61.731 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KX610765 | Arthrobacter phage Prospero | Arthrobacter phage Prospero unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.099 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KX576642 | Arthrobacter phage Massimo | Arthrobacter phage Massimo unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.112 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KX443695 | Arthrobacter phage Courtney3 | Arthrobacter phage Courtney3 unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KX522650 | Arthrobacter phage Conboy | Arthrobacter phage Conboy unclassified Klausavirus Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70096 | 61.687 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160674 | Arthrobacter phage Yank | Arthrobacter phage Yank unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15524 | 60.229 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160670 | Arthrobacter phage Toulouse | Arthrobacter phage Toulouse unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15319 | 60.272 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160667 | Arthrobacter phage Stratus | Arthrobacter phage Stratus unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15630 | 60.019 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160660 | Arthrobacter phage PrincessTrina | Arthrobacter phage PrincessTrina Arthrobacter virus PrincessTrina Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70265 | 61.584 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KU160658 | Arthrobacter phage Muttlie | Arthrobacter phage Muttlie unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15524 | 60.229 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160657 | Arthrobacter phage Moloch | Arthrobacter phage Moloch unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15630 | 60.026 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KU160655 | Arthrobacter phage Maggie | Arthrobacter phage Maggie unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.105 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KT783672 | Arthrobacter phage TymAbreu | Arthrobacter phage TymAbreu unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.112 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KT355475 | Arthrobacter phage Sandman | Arthrobacter phage Sandman unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15630 | 60.032 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KT355473 | Arthrobacter phage Jessica | Arthrobacter phage Jessica unclassified Decurrovirus Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15556 | 60.112 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| KT355471 | Arthrobacter phage Decurro | Arthrobacter phage Decurro Decurrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15524 | 60.236 | Decurrovirus | Unclassified | Siphoviridae | Arthrobacter |
| MN553585 | Pseudomonas phage 4phiC20-1 | Pseudomonas phage 4phiC20-1 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42232 | 62.251 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN553584 | Pseudomonas phage 4phiC20-2 | Pseudomonas phage 4phiC20-2 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72474 | 54.894 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MN553583 | Pseudomonas phage 4phiC20-2 | Pseudomonas phage 4phiC20-2 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55941 | 54.718 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KX925554 | Streptomyces phage BRock | Streptomyces phage BRock Streptomyces virus BRock Borockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112523 | 52.316 | Borockvirus | Unclassified | Myoviridae | Streptomyces |
| MN935200 | Staphylococcus phage VTCCBPA116 | Staphylococcus phage VTCCBPA116 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110012 | 30.464 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| LR595850 | Escherichia virus Lambda_1H12 | Escherichia virus Lambda_1H12 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46703 | 50.027 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR699804 | Escherichia phage rV5_ev146 | Escherichia phage rV5_ev146 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131919 | 43.671 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LR699048 | Escherichia phage VpaE1_ev035 | Escherichia phage VpaE1_ev035 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88002 | 38.957 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| LR694611 | Escherichia phage rV5_ev158 | Escherichia phage rV5_ev158 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138177 | 43.695 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LR694610 | Escherichia phage rV5_ev168 | Escherichia phage rV5_ev168 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138171 | 43.694 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LR694165 | Escherichia phage rV5_ev156 | Escherichia phage rV5_ev156 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136385 | 43.653 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LR597663 | Escherichia phage VpaE1_ev78 | Escherichia phage VpaE1_ev78 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87688 | 38.911 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| LR597658 | Escherichia phage VpaE1_ev108 | Escherichia phage VpaE1_ev108 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87749 | 38.92 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| LR597656 | Escherichia phage Stevie_ev116 | Escherichia phage Stevie_ev116 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48646 | 42.91 | Tlsvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LR597654 | Escherichia phage phAPEC8_ev052 | Escherichia phage phAPEC8_ev052 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150152 | 39.004 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| LR597649 | Escherichia phage ESSI2_ev015 | Escherichia phage ESSI2_ev015 Escherichia virus ev015 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30584 | 50.876 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| LR597647 | Escherichia phage rV5_ev147 | Escherichia phage rV5_ev147 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136384 | 43.653 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LR597641 | Escherichia phage mEp460_ev081 | Escherichia phage mEp460_ev081 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45865 | 49.34 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| LR597640 | Escherichia phage ESSI2_ev129 | Escherichia phage ESSI2_ev129 Escherichia virus ev129 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30927 | 51.483 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| LR597638 | Escherichia phage ESSI2_ev040 | Escherichia phage ESSI2_ev040 Escherichia virus ev040 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30597 | 50.822 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| LR597637 | Escherichia phage ESSI2_ev239 | Escherichia phage ESSI2_ev239 Escherichia virus ev239 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29203 | 51.378 | Seongnamvirus | Peduovirinae | Myoviridae | Escherichia |
| MT002975 | Brucella phage EF4 | Brucella phage EF4 unclassified Perisivirus Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38321 | 48.196 | Perisivirus | Unclassified | Podoviridae | Brucella |
| NC_045425 | Thermus phage phiOH3 | Thermus phage phiOH3 Thermus virus OH3 Thomixvirus Paulinoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5688 | 58.034 | Thomixvirus | Unclassified | Paulinoviridae | Thermus |
| MN553589 | Pseudomonas phage phiKMVC5-121 | Pseudomonas phage phiKMVC5-121 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42231 | 62.253 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN553587 | Pseudomonas phage phiKMVC5-121 | Pseudomonas phage phiKMVC5-121 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42231 | 62.232 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN508624 | Enterobacter phage EC-F2 | Enterobacter phage EC-F2 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178230 | 44.732 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MN508623 | Enterobacter phage EC-F1 | Enterobacter phage EC-F1 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178070 | 44.741 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MN508622 | Enterobacter phage EC-W2 | Enterobacter phage EC-W2 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176610 | 44.715 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MN508621 | Enterobacter phage EC-W1 | Enterobacter phage EC-W1 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178607 | 44.789 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MN508620 | Pseudomonas phage E2005-24-39 | Pseudomonas phage E2005-24-39 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66412 | 55.24 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN508619 | Pseudomonas phage Paer4 | Pseudomonas phage Paer4 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45446 | 52.473 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MN508618 | Escherichia phage ES26 | Escherichia phage ES26 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166756 | 35.388 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN508617 | Escherichia phage ES21 | Escherichia phage ES21 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166790 | 35.378 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN508616 | Escherichia phage ES19 | Escherichia phage ES19 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166781 | 35.376 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MN508614 | Escherichia phage ES12 | Escherichia phage ES12 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166240 | 35.365 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| EU307292 | Burkholderia phage Bups phi1 | Burkholderia phage Bups phi1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19870 | 64.6 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MN228696 | Sinorhizobium phage ort11 | Sinorhizobium phage ort11 Sinorhizobium virus ort11 Huelvavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75239 | 44.246 | Huelvavirus | Unclassified | Schitoviridae | Sinorhizobium |
| MN994275 | Streptomyces phage JXY1 | Streptomyces phage JXY1 Streptomyces virus JXY1 Manuelvirus Beephvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45385 | 60.253 | Manuelvirus | Beephvirinae | Podoviridae | Streptomyces |
| MN940411 | Staphylococcus phage f2b1 | Staphylococcus phage f2b1 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161551 | 40.065 | Unclassified | Bastillevirinae | Herelleviridae | Staphylococcus |
| MN904510 | Staphylococcus phage vB_SauS-SAP27 | Staphylococcus phage vB_SauS-SAP27 Staphylococcus virus SAP27 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43444 | 34.946 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MN877442 | Alteromonas phage vB_AmeM_PT11-V22 | Alteromonas phage vB_AmeM_PT11-V22 Alteromonas virus PT11-V22 Myoalterovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92760 | 38.387 | Myoalterovirus | Unclassified | Myoviridae | Alteromonas |
| MN901924 | Pseudomonas phage vB_PaeP_PE3 | Pseudomonas phage vB_PaeP_PE3 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43583 | 62.3 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN906996 | Pseudomonas phage LY218 | Pseudomonas phage LY218 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73083 | 54.873 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MN904502 | Listeria phage LP-018 | Listeria phage LP-018 unclassified Homburgvirus Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65284 | 36.395 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| MN871511 | Vibrio phage vspsw1 | Vibrio phage vspsw1 unclassified Pogseptimavirus Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113898 | 43.829 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MN871510 | Vibrio phage valsw3 | Vibrio phage valsw3 unclassified Pogseptimavirus Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114030 | 43.832 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MN871509 | Pseudomonas phage pasz5 | Pseudomonas phage pasz5 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66612 | 55.631 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871508 | Pseudomonas phage pasz2 | Pseudomonas phage pasz2 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66518 | 55.588 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871507 | Aeromonas phage asfd_1h | Aeromonas phage asfd_1h unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168967 | 42.231 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| MN871506 | Vibrio phage VspDsh_2 | Vibrio phage VspDsh_2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86939 | 45.703 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MN871505 | Vibrio phage ValDsh2 | Vibrio phage ValDsh2 unclassified Pogseptimavirus Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113924 | 43.831 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MN871504 | Salmonella phage SeTs-2 | Salmonella phage SeTs-2 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155121 | 44.554 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871503 | Salmonella virus SeLz-2 | Salmonella virus SeLz-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40176 | 45.841 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MN871502 | Salmonella phage SeHz-1 | Salmonella phage SeHz-1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156987 | 44.622 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871501 | Salmonella virus Se-U | Salmonella virus Se-U unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157965 | 44.547 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871500 | Salmonella phage Se-S | Salmonella phage Se-S unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157948 | 44.688 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871499 | Salmonella phage Se-N | Salmonella phage Se-N unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160400 | 44.699 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871498 | Salmonella phage Se-J | Salmonella phage Se-J Salmonella virus SeJ Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160400 | 44.694 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871497 | Salmonella phage Se-I | Salmonella phage Se-I unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160401 | 44.695 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871496 | Salmonella phage Se-H | Salmonella phage Se-H unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160400 | 44.695 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871495 | Salmonella phage Se-G | Salmonella phage Se-G Salmonella virus SeG Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158892 | 44.71 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871494 | Salmonella phage Se-F | Salmonella phage Se-F unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158844 | 44.712 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871493 | Salmonella phage Se-E | Salmonella phage Se-E unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157872 | 44.469 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871492 | Salmonella phage Se-D | Salmonella phage Se-D unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158087 | 44.522 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871491 | Salmonella phage Se-B | Salmonella phage Se-B Salmonella virus SeB Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158087 | 44.522 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MN871490 | Salmonella phage SEzh_1 | Salmonella phage SEzh_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49492 | 46.222 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MN871489 | Pseudomonas phage PaZh_1 | Pseudomonas phage PaZh_1 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 278407 | 36.878 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MN871488 | Pseudomonas phage PaYy-2 | Pseudomonas phage PaYy-2 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92348 | 49.333 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871487 | Pseudomonas phage PaSz-9_45_65k | Pseudomonas phage PaSz-9_45_65k unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65880 | 55.619 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871486 | Pseudomonas phage PaSz-9_45_61k | Pseudomonas phage PaSz-9_45_61k unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61157 | 64.377 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871485 | Pseudomonas phage PaSz-8_45_66k | Pseudomonas phage PaSz-8_45_66k unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66266 | 55.614 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871484 | Pseudomonas phage PaSz-8_45_45k | Pseudomonas phage PaSz-8_45_45k unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45455 | 52.439 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MN871483 | Pseudomonas phage PaSz-8_45_42k | Pseudomonas phage PaSz-8_45_42k unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42819 | 62.281 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN871482 | Pseudomonas virus PaSz-6 | Pseudomonas virus PaSz-6 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54656 | 64.668 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871481 | Pseudomonas phage PaSz-1_45_92k | Pseudomonas phage PaSz-1_45_92k unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92096 | 49.366 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871480 | Pseudomonas phage PaSz-1_45_270k | Pseudomonas phage PaSz-1_45_270k unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279520 | 36.913 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MN871479 | Pseudomonas phage PaSt-2_45_65k | Pseudomonas phage PaSt-2_45_65k unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65556 | 55.647 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871478 | Pseudomonas phage PaSt-2_45_61k | Pseudomonas phage PaSt-2_45_61k unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61865 | 64.586 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871477 | Pseudomonas virus PaLz-2-43 | Pseudomonas virus PaLz-2-43 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17024 | 51.333 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MN871476 | Pseudomonas phage PaLz-1-45 | Pseudomonas phage PaLz-1-45 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43890 | 52.126 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MN871475 | Pseudomonas virus Pa-sanjiao | Pseudomonas virus Pa-sanjiao unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158013 | 44.551 | Kuttervirus | Cvivirinae | Ackermannviridae | Pseudomonas |
| MN871474 | Pseudomonas virus Pa-oumiga | Pseudomonas virus Pa-oumiga Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60591 | 56.352 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MN871473 | Pseudomonas virus Pa-Z | Pseudomonas virus Pa-Z unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66201 | 55.632 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871472 | Pseudomonas virus Pa-X | Pseudomonas virus Pa-X Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60237 | 56.339 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MN871471 | Pseudomonas virus Pa-V | Pseudomonas virus Pa-V Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60591 | 56.352 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MN871470 | Pseudomonas phage Pa-U | Pseudomonas phage Pa-U unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92533 | 49.252 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871469 | Pseudomonas phage Pa-S | Pseudomonas phage Pa-S unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61639 | 64.467 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871468 | Pseudomonas phage Pa-Q | Pseudomonas phage Pa-Q unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41213 | 55.844 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871467 | Pseudomonas phage Pa-P | Pseudomonas phage Pa-P Pseudomonas virus NH4 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56994 | 55.737 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871466 | Pseudomonas phage Pa-O | Pseudomonas phage Pa-O unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66045 | 55.651 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871465 | Pseudomonas virus Pa-N | Pseudomonas virus Pa-N unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66272 | 55.665 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871464 | Pseudomonas phage Pa-M | Pseudomonas phage Pa-M unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66261 | 55.662 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871463 | Pseudomonas phage Pa-L | Pseudomonas phage Pa-L unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61641 | 64.47 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871462 | Pseudomonas virus Pa-K | Pseudomonas virus Pa-K unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51820 | 55.432 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871461 | Pseudomonas phage Pa-J | Pseudomonas phage Pa-J unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66167 | 55.661 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871460 | Pseudomonas phage Pa-I | Pseudomonas phage Pa-I unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65292 | 55.612 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871459 | Pseudomonas phage Pa-H | Pseudomonas phage Pa-H unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66166 | 55.655 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871458 | Pseudomonas phage Pa-G | Pseudomonas phage Pa-G unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43105 | 55.379 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871457 | Pseudomonas phage Pa-F | Pseudomonas phage Pa-F unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64807 | 55.634 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871456 | Pseudomonas phage Pa-C | Pseudomonas phage Pa-C unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60633 | 64.391 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MN871455 | Pseudomonas phage Pa-B | Pseudomonas phage Pa-B unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34243 | 62.162 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN871454 | Pseudomonas phage Pa-A | Pseudomonas phage Pa-A unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65924 | 55.06 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN871453 | Serratia phage KpZh_LPS_45 | Serratia phage KpZh_LPS_45 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 299008 | 45.683 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871452 | Serratia phage KpZh_1 | Serratia phage KpZh_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 298898 | 45.685 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871451 | Serratia phage KpYy_2_45 | Serratia phage KpYy_2_45 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 302439 | 45.71 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871450 | Serratia phage KpYy_1_41 | Serratia phage KpYy_1_41 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54417 | 56.899 | Chivirus | Unclassified | Siphoviridae | Serratia |
| MN871449 | Serratia phage KpXa_1 | Serratia phage KpXa_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 304264 | 45.737 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871448 | Serratia phage KpSz_1 | Serratia phage KpSz_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 299003 | 45.684 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871447 | Serratia phage KpLz_1_41 | Serratia phage KpLz_1_41 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 305006 | 45.7 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871445 | Serratia phage KpHz_2 | Serratia phage KpHz_2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 270429 | 45.072 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871444 | Serratia phage KpGz_1 | Serratia phage KpGz_1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 269351 | 45.892 | Unclassified | Unclassified | Myoviridae | Serratia |
| MN871443 | Enterococcus phage EfsWh_1 | Enterococcus phage EfsWh_1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58036 | 39.849 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MN871442 | Aeromonas phage Aszh-1 | Aeromonas phage Aszh-1 unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223245 | 38.527 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MN871441 | Aeromonas phage AsSzw2 | Aeromonas phage AsSzw2 unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 228419 | 37.81 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MN871440 | Aeromonas phage AsFcp_1 | Aeromonas phage AsFcp_1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221312 | 41.954 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| LR595891 | Escherichia virus P88-4B11 | Escherichia virus P88-4B11 Xuanwuvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37100 | 51.765 | Xuanwuvirus | Peduovirinae | Myoviridae | Escherichia |
| LR595887 | Escherichia virus P2-4A7b | Escherichia virus P2-4A7b Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32848 | 50.932 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595885 | Escherichia virus P2-4E6b | Escherichia virus P2-4E6b Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31128 | 51.982 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595883 | Escherichia virus P2-4C9 | Escherichia virus P2-4C9 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32150 | 51.938 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595873 | Escherichia virus P2-4B2 | Escherichia virus P2-4B2 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32118 | 51.389 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595870 | Escherichia virus P2-2H1 | Escherichia virus P2-2H1 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32662 | 51.111 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595869 | Escherichia virus P2-2H4 | Escherichia virus P2-2H4 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31342 | 52.131 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| LR595868 | Escherichia virus mEp460_4F5 | Escherichia virus mEp460_4F5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42927 | 50.115 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| LR595867 | Escherichia virus Mu_3F6 | Escherichia virus Mu_3F6 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37126 | 51.791 | Muvirus | Unclassified | Myoviridae | Escherichia |
| LR595866 | Escherichia virus Lambda_2G7b | Escherichia virus Lambda_2G7b unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47142 | 49.879 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595865 | Escherichia virus Lambda_2G7a | Escherichia virus Lambda_2G7a unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46701 | 50.027 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595864 | Escherichia virus Lambda_4A7 | Escherichia virus Lambda_4A7 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49168 | 49.77 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595863 | Escherichia virus Lambda_4B5 | Escherichia virus Lambda_4B5 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48141 | 50.13 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595862 | Escherichia phage 2H10 | Escherichia phage 2H10 Escherichia virus 2H10 Glaedevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46288 | 50.326 | Glaedevirus | Unclassified | Siphoviridae | Escherichia |
| LR595861 | Escherichia virus Lambda_4C10 | Escherichia virus Lambda_4C10 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49207 | 49.769 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595860 | Escherichia virus Lambda_2E9 | Escherichia virus Lambda_2E9 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46702 | 50.026 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| LR595859 | Escherichia virus Lambda_2B8 | Escherichia virus Lambda_2B8 unclassified Lambdavirus Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48763 | 50.188 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MN294712 | Acinetobacter phage vB_AbaP_APK48 | Acinetobacter phage vB_AbaP_APK48 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41105 | 39.258 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MT013111 | Staphylococcus phage SA75 | Staphylococcus phage SA75 Staphylococcus virus SA75 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43134 | 34.437 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK658834 | Staphylococcus virus Baq_Sau1 | Staphylococcus virus Baq_Sau1 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44384 | 34.046 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| CP019719 | Paenibacillus phage phiERICV | Paenibacillus phage phiERICV Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45618 | 42.115 | Unclassified | Unclassified | Myoviridae | Paenibacillus |
| MN379740 | Salmonella phage 8-19 | Salmonella phage 8-19 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52648 | 46.025 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MN379739 | Salmonella phage 1-19 | Salmonella phage 1-19 Salmonella virus 1-19 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110775 | 39.998 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN197465 | Salmonella phage 2-3 | Salmonella phage 2-3 Salmonella virus 2-3 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114986 | 39.647 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN218191 | Salmonella phage 1-29 | Salmonella phage 1-29 Salmonella virus 1-29 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111636 | 40.075 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN218190 | Salmonella phage 9-29 | Salmonella phage 9-29 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110776 | 39.998 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK370036 | Salmonella phage 1-23 | Salmonella phage 1-23 Salmonella virus 123 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112496 | 39.969 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK393882 | Salmonella phage 3-29 | Salmonella phage 3-29 Salmonella virus 329 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110377 | 40.045 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| LC519489 | Escherichia phage SP22 | Escherichia phage SP22 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87251 | 38.857 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MN692200 | Pectobacterium phage MA14 | Pectobacterium phage MA14 unclassified Cystovirus Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 10019 | 53.977 | Cystovirus | Unclassified | Cystoviridae | Pectobacterium |
| MN518139 | Pectobacterium phage MA11 | Pectobacterium phage MA11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55830 | 54.542 | Unclassified | Unclassified | Siphoviridae | Pectobacterium |
| MN509793 | Pectobacterium phage MA13 | Pectobacterium phage MA13 unclassified Melnykvirinae Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42464 | 53.433 | Unclassified | Melnykvirinae | Autographiviridae | Pectobacterium |
| MN688132 | Enterobacter phage LAU1 | Enterobacter phage LAU1 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30772 | 51.859 | Unclassified | Hendrixvirinae | Siphoviridae | Enterobacter |
| MN812241 | Flavobacterium phage vB_FspS_tooticki9-1 | Flavobacterium phage vB_FspS_tooticki9-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36912 | 29.28 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| MN812240 | Flavobacterium phage vB_FspS_tooticki6-1 | Flavobacterium phage vB_FspS_tooticki6-1 Flavobacterium virus Tooticki Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36912 | 29.283 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812239 | Flavobacterium phage vB_FspS_tant8-1 | Flavobacterium phage vB_FspS_tant8-1 Flavobacterium virus Tant Tantvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40542 | 34.406 | Tantvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812238 | Flavobacterium phage vB_FspS_stinky9-1 | Flavobacterium phage vB_FspS_stinky9-1 Flavobacterium virus Stinky Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38502 | 29.375 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812237 | Flavobacterium phage vB_FspS_snusmum9-1 | Flavobacterium phage vB_FspS_snusmum9-1 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38743 | 29.032 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812236 | Flavobacterium phage vB_FspS_snusmum6-3 | Flavobacterium phage vB_FspS_snusmum6-3 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39365 | 29.054 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812235 | Flavobacterium phage vB_FspS_snusmum6-2 | Flavobacterium phage vB_FspS_snusmum6-2 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39365 | 29.049 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812234 | Flavobacterium phage vB_FspS_snusmum6-1 | Flavobacterium phage vB_FspS_snusmum6-1 Flavobacterium virus Snusmum Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39365 | 29.051 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812233 | Flavobacterium phage vB_FspS_snork9-1 | Flavobacterium phage vB_FspS_snork9-1 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38084 | 29.493 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812232 | Flavobacterium phage vB_FspS_snork6-2 | Flavobacterium phage vB_FspS_snork6-2 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37732 | 29.5 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812231 | Flavobacterium phage vB_FspS_snork6-1 | Flavobacterium phage vB_FspS_snork6-1 Flavobacterium virus Snork Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38214 | 29.45 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812230 | Flavobacterium phage vB_FspS_sniff9-2 | Flavobacterium phage vB_FspS_sniff9-2 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37907 | 29.398 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812229 | Flavobacterium phage vB_FspS_sniff9-1 | Flavobacterium phage vB_FspS_sniff9-1 Flavobacterium virus Sniff Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38134 | 29.404 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812228 | Flavobacterium phage vB_FspS_mymlan6-2 | Flavobacterium phage vB_FspS_mymlan6-2 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38372 | 29.157 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812227 | Flavobacterium phage vB_FspS_mymlan6-1 | Flavobacterium phage vB_FspS_mymlan6-1 Flavobacterium virus Mymlan Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38501 | 29.158 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812226 | Flavobacterium phage vB_FspS_mumin9-3 | Flavobacterium phage vB_FspS_mumin9-3 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38398 | 29.153 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812225 | Flavobacterium phage vB_FspS_mumin9-2 | Flavobacterium phage vB_FspS_mumin9-2 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38527 | 29.156 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812224 | Flavobacterium phage vB_FspS_mumin9-1 | Flavobacterium phage vB_FspS_mumin9-1 Flavobacterium virus Mumin Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38527 | 29.159 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812223 | Flavobacterium phage vB_FspS_mumin6-4 | Flavobacterium phage vB_FspS_mumin6-4 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37532 | 29.042 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812222 | Flavobacterium phage vB_FspS_mumin6-3 | Flavobacterium phage vB_FspS_mumin6-3 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37682 | 29.144 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812221 | Flavobacterium phage vB_FspS_mumin6-2 | Flavobacterium phage vB_FspS_mumin6-2 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38185 | 29.116 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812220 | Flavobacterium phage vB_FspS_mumin6-1 | Flavobacterium phage vB_FspS_mumin6-1 unclassified Muminvirus Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38527 | 29.164 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812219 | Flavobacterium phage vB_FspS_morran9-1 | Flavobacterium phage vB_FspS_morran9-1 Flavobacterium virus Morran Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38385 | 29.462 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812218 | Flavobacterium phage vB_FspS_lillamy9-7 | Flavobacterium phage vB_FspS_lillamy9-7 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37531 | 29.44 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812217 | Flavobacterium phage vB_FspS_lillamy9-6 | Flavobacterium phage vB_FspS_lillamy9-6 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38007 | 29.452 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812216 | Flavobacterium phage vB_FspS_lillamy9-5 | Flavobacterium phage vB_FspS_lillamy9-5 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.361 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812215 | Flavobacterium phage vB_FspS_lillamy9-4 | Flavobacterium phage vB_FspS_lillamy9-4 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.338 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812214 | Flavobacterium phage vB_FspS_lillamy9-3 | Flavobacterium phage vB_FspS_lillamy9-3 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.304 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812213 | Flavobacterium phage vB_FspS_lillamy9-2 | Flavobacterium phage vB_FspS_lillamy9-2 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.367 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812212 | Flavobacterium phage vB_FspS_lillamy9-1 | Flavobacterium phage vB_FspS_lillamy9-1 Flavobacterium virus Lillamy Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38268 | 29.364 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812211 | Flavobacterium phage vB_FspS_laban6-1 | Flavobacterium phage vB_FspS_laban6-1 Flavobacterium virus Laban Labanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43236 | 31.141 | Labanvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812210 | Flavobacterium phage vB_FspS_hemulen9-1 | Flavobacterium phage vB_FspS_hemulen9-1 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38901 | 29.555 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812209 | Flavobacterium phage vB_FspS_hemulen6-2 | Flavobacterium phage vB_FspS_hemulen6-2 unclassified Lillamyvirus Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39030 | 29.544 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812208 | Flavobacterium phage vB_FspS_hemulen6-1 | Flavobacterium phage vB_FspS_hemulen6-1 Flavobacterium virus Hemulen Lillamyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39039 | 29.537 | Lillamyvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812207 | Flavobacterium phage vB_FspS_hattifnatt9-1 | Flavobacterium phage vB_FspS_hattifnatt9-1 Flavobacterium virus Hattifnatt Hattifnattvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39262 | 29.196 | Hattifnattvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812206 | Flavobacterium phage vB_FspS_filifjonk9-1 | Flavobacterium phage vB_FspS_filifjonk9-1 Flavobacterium virus Filifjonk Muminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38028 | 29.786 | Muminvirus | Unclassified | Siphoviridae | Flavobacterium |
| MN812205 | Flavobacterium phage vB_FspM_pippi8-1 | Flavobacterium phage vB_FspM_pippi8-1 Flavobacterium virus Pippi Pippivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34711 | 37.582 | Pippivirus | Unclassified | Myoviridae | Flavobacterium |
| MN812204 | Flavobacterium phage vB_FspM_lotta8-2 | Flavobacterium phage vB_FspM_lotta8-2 unclassified Pippivirus Pippivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34627 | 37.552 | Pippivirus | Unclassified | Myoviridae | Flavobacterium |
| MN812203 | Flavobacterium phage vB_FspM_lotta8-1 | Flavobacterium phage vB_FspM_lotta8-1 Flavobacterium virus Lotta Pippivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34751 | 37.55 | Pippivirus | Unclassified | Myoviridae | Flavobacterium |
| MN894885 | Escherichia virus Ec_Makalu_001 | Escherichia virus Ec_Makalu_001 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163752 | 40.602 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN882349 | Escherichia virus Ec_Makalu_003 | Escherichia virus Ec_Makalu_003 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162966 | 40.678 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK473382 | Clostridium phage JD032 | Clostridium phage JD032 unclassified Sherbrookevirus Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35109 | 29.944 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| MN794238 | Vibrio phage VH2_2019 | Vibrio phage VH2_2019 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81721 | 48.011 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MN642089 | Klebsiella phage N1M2 | Klebsiella phage N1M2 Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253367 | 40.862 | Petsuvirus | Unclassified | Myoviridae | Klebsiella |
| MN892483 | Microbacterium phage SonOfLevi | Microbacterium phage SonOfLevi unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41573 | 63.363 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN180253 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18100 | 35.735 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN180252 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16547 | 38.224 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN180251 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87722 | 34.6 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN180250 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16327 | 38.372 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN180249 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30054 | 36.737 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN180248 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25122 | 35.666 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN844877 | Sphaerotilus phage vB_SnaP-R1 | Sphaerotilus phage vB_SnaP-R1 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41507 | 62.474 | Unclassified | Krylovirinae | Autographiviridae | Sphaerotilus |
| MN862068 | Brevundimonas phage vB_BsubS-Delta | Brevundimonas phage vB_BsubS-Delta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87225 | 62.498 | Unclassified | Unclassified | Siphoviridae | Brevundimonas |
| MN794232 | Vibrio phage VH1_2019 | Vibrio phage VH1_2019 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246059 | 42.585 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MN850656 | Flavobacterium phage fF4 | Flavobacterium phage fF4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31958 | 36.035 | Unclassified | Unclassified | Myoviridae | Flavobacterium |
| MN840487 | Proteus phage 2207-N35 | Proteus phage 2207-N35 unclassified Gorganvirus Gorganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55645 | 35.892 | Gorganvirus | Unclassified | Siphoviridae | Proteus |
| MN840486 | Proteus phage P16-2532 | Proteus phage P16-2532 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58931 | 46.712 | Unclassified | Unclassified | Siphoviridae | Proteus |
| MN840485 | Escherichia phage 2725-N35 | Escherichia phage 2725-N35 unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45917 | 44.043 | Unclassified | Braunvirinae | Drexlerviridae | Escherichia |
| MN882542 | Escherichia phage JN01 | Escherichia phage JN01 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88360 | 38.755 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MH673671 | Thermus phage phiKo | Thermus phage phiKo unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 11129 | 61.677 | Unclassified | Unclassified | Tectiviridae | Thermus |
| MK559459 | Vibrio phage vB_VcaS_HC | Vibrio phage vB_VcaS_HC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81566 | 47.559 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| LC519601 | Cronobacter phage vB_CsaP_009 | Cronobacter phage vB_CsaP_009 Cronobacter virus 009 Privateervirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92122 | 35.122 | Privateervirus | Unclassified | Podoviridae | Cronobacter |
| MN553588 | Pseudomonas phage phiKMVC5-12 | Pseudomonas phage phiKMVC5-12 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42232 | 62.244 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN553586 | Pseudomonas phage phiKMVC5-4 | Pseudomonas phage phiKMVC5-4 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45446 | 52.48 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| LR745710 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39778 | 48.384 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| LR745709 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40210 | 48.421 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| LR745708 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40606 | 48.633 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| LR745707 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40532 | 48.633 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| LR745706 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38449 | 48.41 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN481366 | Escherichia phage vB_EcoP_PHB20 | Escherichia phage vB_EcoP_PHB20 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38552 | 49.064 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN481365 | Escherichia phage vB_EcoP_PHB19 | Escherichia phage vB_EcoP_PHB19 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40037 | 48.113 | Unclassified | Studiervirinae | Autographiviridae | Escherichia |
| MK759884 | Salmonella phage vB_SenM_SB18 | Salmonella phage vB_SenM_SB18 unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85311 | 43.776 | Kolesnikvirus | Ounavirinae | Myoviridae | Salmonella |
| MK759883 | Salmonella phage vB_SenS_SB15 | Salmonella phage vB_SenS_SB15 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41441 | 49.994 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN166083 | Acinetobacter phage Abp9 | Acinetobacter phage Abp9 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44820 | 37.691 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| AP019415 | Pseudomonas phage HU1 | Pseudomonas phage HU1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42551 | 56.377 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| LC472884 | Pseudomonas phage S50 | Pseudomonas phage S50 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66313 | 55.618 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC472883 | Pseudomonas phage S12-3 | Pseudomonas phage S12-3 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66257 | 55.579 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC472882 | Pseudomonas phage R26 | Pseudomonas phage R26 Pseudomonas virus R26 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65737 | 55.071 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC472881 | Pseudomonas phage R12 | Pseudomonas phage R12 Pseudomonas virus R12 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65415 | 55.397 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN604407 | Salmonella phage ST-3 | Salmonella phage ST-3 unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85180 | 43.805 | Kolesnikvirus | Ounavirinae | Myoviridae | Salmonella |
| MN453779 | Aeromonas phage PS2 | Aeromonas phage PS2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240447 | 38.819 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| LR743532 | Xanthomonas phage Bosa | Xanthomonas phage Bosa unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63828 | 64.837 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| LR743531 | Xanthomonas phage Tenjo | Xanthomonas phage Tenjo Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43850 | 67.241 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| LR743530 | Xanthomonas phage Suba | Xanthomonas phage Suba Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45510 | 52.338 | Unclassified | Unclassified | Siphoviridae | Xanthomonas |
| LR743529 | Xanthomonas phage Sopo | Xanthomonas phage Sopo Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43861 | 66.902 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| LR743528 | Xanthomonas phage Tabio | Xanthomonas phage Tabio Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43458 | 66.931 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| LR743524 | Xylella phage Bacata | Xylella phage Bacata unclassified Sanovirus Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56232 | 62.95 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| LR743523 | Xylella phage Usme | Xylella phage Usme Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43721 | 66.6 | Unclassified | Unclassified | Podoviridae | Xylella |
| MN395286 | Klebsiella phage PhiKpNIH-2 | Klebsiella phage PhiKpNIH-2 Klebsiella virus PhiKpNIH2 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49477 | 50.419 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN395285 | Klebsiella phage PhiKpNIH-10 | Klebsiella phage PhiKpNIH-10 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49472 | 50.97 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN395284 | Klebsiella phage PhiKpNIH-6 | Klebsiella phage PhiKpNIH-6 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171860 | 40.664 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MN562749 | Pseudomonas phage U47 | Pseudomonas phage U47 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43444 | 52.495 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MN655999 | Shigella phage Z31 | Shigella phage Z31 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89355 | 38.875 | Felixounavirus | Ounavirinae | Myoviridae | Shigella |
| MN655998 | Escherichia phage E26 | Escherichia phage E26 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164572 | 40.482 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN727882 | Geobacillus phage GBK1 | Geobacillus phage GBK1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45439 | 43.562 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| MN735435 | Microbacterium phage Renzie | Microbacterium phage Renzie unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41872 | 63.408 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN735430 | Microbacterium phage Ioannes | Microbacterium phage Ioannes unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41879 | 63.442 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN783016 | Klebsiella phage KP1801 | Klebsiella phage KP1801 Klebsiella virus KP1801 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49835 | 50.137 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN781580 | Shigella virus KRT47 | Shigella virus KRT47 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167485 | 35.557 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MN781674 | Escherichia phage vB_EcoS_XY3 | Escherichia phage vB_EcoS_XY3 unclassified Warwickvirus Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51345 | 44.756 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN807240 | Vibrio phage VP-HS15 | Vibrio phage VP-HS15 unclassified Kaohsiungvirus Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46759 | 43.867 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| MN807246 | Microbacterium phage Chamuel | Microbacterium phage Chamuel unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41568 | 63.349 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN813699 | Mycobacterium phage P28Green | Mycobacterium phage P28Green unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50886 | 63.953 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN813687 | Mycobacterium phage Onyinye | Mycobacterium phage Onyinye Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71419 | 58.808 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN813685 | Gordonia phage Gudmit | Gordonia phage Gudmit Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42205 | 65.912 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| LC516895 | Escherichia phage vB_EcoS-DELF2 | Escherichia phage vB_EcoS-DELF2 Escherichia virus DELF2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50168 | 45.505 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| AY190604 | Haloferax virus HF1 | Haloferax virus HF1 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75898 | 55.793 | Haloferacalesvirus | Unclassified | Myoviridae | Haloferax |
| AF222060 | Halorubrum phage HF2 | Halorubrum phage HF2 Halorubrum virus HF2 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77672 | 55.849 | Haloferacalesvirus | Unclassified | Myoviridae | Halorubrum |
| MN102376 | Vibrio phage vB_VpS_CA8 | Vibrio phage vB_VpS_CA8 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58480 | 46.423 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MN175679 | Vibrio phage vB_VpS_BA3 | Vibrio phage vB_VpS_BA3 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58648 | 46.3 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MK764384 | Staphylococcus phage SA46-CTH2 | Staphylococcus phage SA46-CTH2 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17505 | 28.866 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK318076 | Pseudomonas phage vB_PaeM_fHoPae01 | Pseudomonas phage vB_PaeM_fHoPae01 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66418 | 55.724 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| JX871397 | Enterobacteria phage phi80 | Enterobacteria phage phi80 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46150 | 52.132 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| LC487410 | Vibrio phage KIT04 | Vibrio phage KIT04 Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114933 | 40.243 | Unclassified | Ermolyevavirinae | Demerecviridae | Vibrio |
| MN032614 | Aeromonas phage PS1 | Aeromonas phage PS1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237367 | 38.792 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MF580961 | Salinibacter phage M31CR41-2 | Salinibacter phage M31CR41-2 Kairosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51889 | 59.658 | Kairosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580959 | Salinibacter phage M8CRM-1 | Salinibacter phage M8CRM-1 Salinibacter virus M8CRM1 Kryptosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53197 | 53.381 | Kryptosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580957 | Salinibacter phage M8CR30-2 | Salinibacter phage M8CR30-2 Holosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35099 | 64.592 | Holosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580956 | Salinibacter phage M8CC-19 | Salinibacter phage M8CC-19 Salinibacter virus M8CC19 Kryptosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53812 | 52.99 | Kryptosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580955 | Salinibacter phage M1EM-1 | Salinibacter phage M1EM-1 Salinibacter virus M1EM1 Holosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35883 | 64.66 | Holosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF629150 | Salinibacter phage SRUTV-1 | Salinibacter phage SRUTV-1 Salinibacter virus SRUTV1 Kairosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50672 | 59.445 | Kairosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| FJ750948 | Escherichia phage SSL-2009a | Escherichia phage SSL-2009a Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44899 | 54.667 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| JX865427 | Escherichia phage JL1 | Escherichia phage JL1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43457 | 54.774 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| JN051154 | Pediococcus phage cIP1 | Pediococcus phage cIP1 Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38013 | 47.607 | Coetzeevirus | Unclassified | Siphoviridae | Pediococcus |
| AF424781 | Staphylococcus phage phi 11 | Staphylococcus phage phi 11 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43604 | 34.497 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AF158600 | Streptococcus phage Sfi11 | Streptococcus phage Sfi11 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39807 | 38.556 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| U88974 | Streptococcus phage 01205 | Streptococcus phage 01205 Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43075 | 38.261 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| C2PVCG | Lactobacillus phage c2 | Lactobacillus phage c2 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22172 | 36.307 | Ceduovirus | Unclassified | Siphoviridae | Lactobacillus |
| LAMCG | Escherichia phage Lambda | Escherichia phage Lambda Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48502 | 49.858 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MN061582 | Acinetobacter phage vB_AbaP_IME546 | Acinetobacter phage vB_AbaP_IME546 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40770 | 39.144 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN580668 | Salmonella phage pSe_SNUABM_01 | Salmonella phage pSe_SNUABM_01 Salmonella virus SNUABM-01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172360 | 35.36 | Tequatrovirus | Tevenvirinae | Myoviridae | Salmonella |
| MK290738 | Pectobacterium phage Arno18 | Pectobacterium phage Arno18 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91660 | 44.511 | Unclassified | Ounavirinae | Myoviridae | Pectobacterium |
| MK290737 | Pectobacterium phage Arno162 | Pectobacterium phage Arno162 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91663 | 44.537 | Unclassified | Ounavirinae | Myoviridae | Pectobacterium |
| KX765865 | Salmonella phage ST-W77 | Salmonella phage ST-W77 Salmonella virus STW77 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157458 | 44.482 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KX765864 | Salmonella phage ST-374 | Salmonella phage ST-374 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59934 | 56.602 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KY769270 | Pectobacterium phage phiA41 | Pectobacterium phage phiA41 Pectobacterium virus phiA41 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75764 | 48.701 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| KX649889 | Salmonella phage SE-W109 | Salmonella phage SE-W109 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42147 | 49.6 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN709127 | Escherichia virus Ec_Makalu_002 | Escherichia virus Ec_Makalu_002 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164674 | 40.579 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MN698248 | Pelagibacter phage HTVC202P | Pelagibacter phage HTVC202P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32226 | 31.295 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698247 | Pelagibacter phage HTVC115P | Pelagibacter phage HTVC115P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54819 | 32.866 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698246 | Pelagibacter phage HTVC112P | Pelagibacter phage HTVC112P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32478 | 30.448 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698245 | Pelagibacter phage HTVC111P | Pelagibacter phage HTVC111P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31577 | 30.51 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698244 | Pelagibacter phage HTVC106P | Pelagibacter phage HTVC106P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36945 | 32.121 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698243 | Pelagibacter phage HTVC104P | Pelagibacter phage HTVC104P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54359 | 30.9 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698242 | Pelagibacter phage HTVC103P | Pelagibacter phage HTVC103P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54103 | 31.048 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698241 | Pelagibacter phage HTVC027P | Pelagibacter phage HTVC027P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57595 | 34.767 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698240 | Pelagibacter phage HTVC026P | Pelagibacter phage HTVC026P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32480 | 31.318 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN698239 | Pelagibacter phage HTVC023P | Pelagibacter phage HTVC023P Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60878 | 35.014 | Unclassified | Unclassified | Podoviridae | Pelagibacter |
| MN732867 | Erwinia phage Hena1 | Erwinia phage Hena1 Erwinia virus Hena1 Henunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148842 | 48.404 | Henunavirus | Vequintavirinae | Myoviridae | Erwinia |
| MN718199 | Vibrio phage vB_VchM_Kuja | Vibrio phage vB_VchM_Kuja Vibrio virus Kuja Kujavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148180 | 36.378 | Kujavirus | Unclassified | Ackermannviridae | Vibrio |
| MN710383 | Pseudomonas phage pf8_ST274-AUS411 | Pseudomonas phage pf8_ST274-AUS411 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10061 | 58.145 | Unclassified | Unclassified | Inoviridae | Pseudomonas |
| MN781108 | Klebsiella phage vB_Kpn_P545 | Klebsiella phage vB_Kpn_P545 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169275 | 40.822 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MN737622 | Methanobacterium virus PhiF3 | Methanobacterium virus PhiF3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32524 | 49.36 | Unclassified | Unclassified | Siphoviridae | Methanobacterium |
| MN562221 | Photobacterium phage PDCC-1 | Photobacterium phage PDCC-1 Photobacterium virus PDCC1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237509 | 43.252 | Aphroditevirus | Gorgonvirinae | Myoviridae | Photobacterium |
| MN549361 | Rhizobium phage RL2RES | Rhizobium phage RL2RES unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156878 | 49.971 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN549360 | Rhizobium phage RL38J1 | Rhizobium phage RL38J1 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158577 | 49.816 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| MN689779 | Klebsiella phage vB_KpnS_15-38_KLPPOU149 | Klebsiella phage vB_KpnS_15-38_KLPPOU149 Klebsiella virus KLPPOU149 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49316 | 51.032 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN689778 | Klebsiella phage vB_KpnM_15-38_KLPPOU148 | Klebsiella phage vB_KpnM_15-38_KLPPOU148 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49169 | 48.437 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MN718463 | Clostridium phage phiCDKH01 | Clostridium phage phiCDKH01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45089 | 28.719 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MN592896 | Alteromonas phage XX1924 | Alteromonas phage XX1924 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40580 | 43.659 | Unclassified | Unclassified | Siphoviridae | Alteromonas |
| MN794003 | Klebsiella phage VLC4 | Klebsiella phage VLC4 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44656 | 53.762 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN794002 | Klebsiella phage VLC3 | Klebsiella phage VLC3 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43351 | 53.904 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN794001 | Klebsiella phage VLC2 | Klebsiella phage VLC2 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43784 | 53.832 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN794000 | Klebsiella phage VLC1 | Klebsiella phage VLC1 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43411 | 53.827 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MK774614 | Aeromonas phage CF8 | Aeromonas phage CF8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 238150 | 42.198 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MK799832 | Synechococcus phage S-B05 | Synechococcus phage S-B05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 208857 | 39.865 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MK308677 | Vibrio phage vB_VmeM-Yong MS32 | Vibrio phage vB_VmeM-Yong MS32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 278116 | 45.868 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK308676 | Vibrio phage vB_VmeM-Yong MS31 | Vibrio phage vB_VmeM-Yong MS31 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 290537 | 45.872 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK308675 | Vibrio phage vB_VmeM-Yong XC32 | Vibrio phage vB_VmeM-Yong XC32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 290533 | 45.873 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK308674 | Vibrio phage vB_VmeM-Yong XC31 | Vibrio phage vB_VmeM-Yong XC31 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 290532 | 45.873 | Unclassified | Unclassified | Myoviridae | Vibrio |
| LC513943 | Enterococcus phage vB_EfaS-DELF1 | Enterococcus phage vB_EfaS-DELF1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40248 | 34.931 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KM051843 | Bacillus phage Bobb | Bacillus phage Bobb Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160281 | 41.331 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN648445 | Escherichia phage FP43 | Escherichia phage FP43 unclassified bacterial viruses Viruses | 169248 | 37.684 | Unclassified | Unclassified | Unclassified | Escherichia |
| MN695335 | Synechococcus phage B23 | Synechococcus phage B23 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 243633 | 35.393 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MN695334 | Synechococcus phage B3 | Synechococcus phage B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244930 | 35.363 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MN719123 | Vibrio phage VALG_phi6 | Vibrio phage VALG_phi6 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8529 | 44.343 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MN690600 | Vibrio phage VALG_phi8 | Vibrio phage VALG_phi8 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7311 | 46.341 | Unclassified | Unclassified | Inoviridae | Vibrio |
| MN689533 | Lactococcus phage PC_S1 | Lactococcus phage PC_S1 Lactococcus virus PCS1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21629 | 36.104 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689532 | Lactococcus phage PC_B3 | Lactococcus phage PC_B3 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21987 | 36.117 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689531 | Lactococcus phage PC_B1 | Lactococcus phage PC_B1 Lactococcus virus PCB1 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22099 | 36.092 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689530 | Lactococcus phage CHPC974 | Lactococcus phage CHPC974 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33796 | 35.016 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN689529 | Lactococcus phage CHPC973 | Lactococcus phage CHPC973 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22306 | 35.937 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689528 | Lactococcus phage CHPC972 | Lactococcus phage CHPC972 Lactococcus virus CHPC972 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23281 | 35.51 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689527 | Lactococcus phage CHPC967 | Lactococcus phage CHPC967 Lactococcus virus CHPC967 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22405 | 35.729 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689526 | Lactococcus phage CHPC966 | Lactococcus phage CHPC966 Lactococcus virus CHPC966 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21749 | 36.264 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689525 | Lactococcus phage CHPC965 | Lactococcus phage CHPC965 Lactococcus virus CHPC965 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25327 | 34.876 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689524 | Lactococcus phage CHPC964 | Lactococcus phage CHPC964 Lactococcus virus CHPC964 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29925 | 34.399 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689523 | Lactococcus phage CHPC959 | Lactococcus phage CHPC959 Lactococcus virus CHPC959 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29340 | 34.315 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689522 | Lactococcus phage CHPC958 | Lactococcus phage CHPC958 Lactococcus virus CHPC958 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32653 | 34.478 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689521 | Lactococcus phage CHPC836 | Lactococcus phage CHPC836 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36482 | 34.74 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN689520 | Lactococcus phage CHPC781 | Lactococcus phage CHPC781 Lactococcus virus CHPC781 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29218 | 34.475 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689519 | Lactococcus phage CHPC52 | Lactococcus phage CHPC52 Lactococcus virus CHPC52 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29652 | 34.149 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689518 | Lactococcus phage CHPC362 | Lactococcus phage CHPC362 Lactococcus virus CHPC362 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27642 | 35.428 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689517 | Lactococcus phage CHPC361 | Lactococcus phage CHPC361 Lactococcus virus CHPC361 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30162 | 34.451 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689516 | Lactococcus phage CHPC148 | Lactococcus phage CHPC148 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33553 | 34.897 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN689515 | Lactococcus phage CHPC134 | Lactococcus phage CHPC134 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21987 | 36.144 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689514 | Lactococcus phage CHPC129 | Lactococcus phage CHPC129 Lactococcus virus CHPC129 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30832 | 34.561 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN689513 | Lactococcus phage CHPC1242 | Lactococcus phage CHPC1242 Lactococcus virus CHPC1242 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21144 | 35.873 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689512 | Lactococcus phage CHPC122 | Lactococcus phage CHPC122 Lactococcus virus CHPC122 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22109 | 35.474 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689511 | Lactococcus phage CHPC1183 | Lactococcus phage CHPC1183 Lactococcus virus CHPC1183 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20030 | 36.216 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689510 | Lactococcus phage CHPC1182 | Lactococcus phage CHPC1182 Lactococcus virus CHPC1182 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20751 | 35.96 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689509 | Lactococcus phage CHPC1175 | Lactococcus phage CHPC1175 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36339 | 35.326 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN689508 | Lactococcus phage CHPC1170 | Lactococcus phage CHPC1170 Lactococcus virus CHPC1170 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21746 | 35.616 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689507 | Lactococcus phage CHPC116 | Lactococcus phage CHPC116 Lactococcus virus CHPC116 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21869 | 35.731 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689506 | Lactococcus phage CHPC1161 | Lactococcus phage CHPC1161 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21344 | 36.174 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689505 | Lactococcus phage CHPC1020 | Lactococcus phage CHPC1020 Lactococcus virus CHPC1020 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22410 | 35.573 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689504 | Lactococcus phage 5205F | Lactococcus phage 5205F Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21214 | 36.306 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN689503 | Lactococcus phage 5171F | Lactococcus phage 5171F Lactococcus virus 5171F Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21078 | 36.218 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN336265 | Salmonella phage vB_SenS-EnJE6 | Salmonella phage vB_SenS-EnJE6 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43129 | 49.575 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN592897 | Vibrio phage MZH0603 | Vibrio phage MZH0603 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43006 | 43.919 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MN013080 | Klebsiella phage vB_KpnS_SegesCirculi | Klebsiella phage vB_KpnS_SegesCirculi Klebsiella virus SegesCirculi Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50713 | 51.121 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013079 | Klebsiella phage vB_KpnS_Call | Klebsiella phage vB_KpnS_Call Klebsiella virus Call Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51487 | 51.549 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013076 | Klebsiella phage vB_KpnS_IMGroot | Klebsiella phage vB_KpnS_IMGroot Klebsiella virus IMGroot Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52866 | 51.347 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK238400 | CrAssphage ZA | CrAssphage ZA crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97757 | 29.164 | Unclassified | Unclassified | Podoviridae | Unspecified |
| KM979354 | Escherichia phage DT57C | Escherichia phage DT57C Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108065 | 39.65 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KC618326 | Escherichia phage P2 | Escherichia phage P2 Escherichia virus magyaro Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31200 | 52.603 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| HQ665011 | Escherichia phage vB_EcoS_AKFV33 | Escherichia phage vB_EcoS_AKFV33 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108853 | 38.945 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| HM134276 | Escherichia phage RB16 | Escherichia phage RB16 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176788 | 43.521 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| FJ839692 | Escherichia phage RB14 | Escherichia phage RB14 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165429 | 35.325 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| DQ087285 | Burkholderia phage phiE52237 | Burkholderia phage phiE52237 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37639 | 64.818 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| DQ904452 | Escherichia phage RB32 | Escherichia phage RB32 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165890 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| DQ426904 | Mannheimia phage PHL101 | Mannheimia phage PHL101 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34525 | 41.567 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| AY370674 | Escherichia phage K1-5 | Escherichia phage K1-5 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44385 | 45.247 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| AY135739 | Escherichia phage Wphi | Escherichia phage Wphi Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32684 | 51.723 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| AF158101 | Escherichia phage T4 | Escherichia phage T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168903 | 35.298 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| B1U32222 | Escherichia phage 186 | Escherichia phage 186 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30624 | 53.086 | Eganvirus | Peduovirinae | Myoviridae | Escherichia |
| AP019525 | Tenacibaculum phage PTm5 | Tenacibaculum phage PTm5 unclassified Shirahamavirus Shirahamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 226876 | 29.719 | Shirahamavirus | Unclassified | Myoviridae | Tenacibaculum |
| AP019524 | Tenacibaculum phage PTm1 | Tenacibaculum phage PTm1 Tenacibaculum virus PTm1 Shirahamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224680 | 29.744 | Shirahamavirus | Unclassified | Myoviridae | Tenacibaculum |
| MN096267 | Pseudomonas phage vB_PaS_IME307 | Pseudomonas phage vB_PaS_IME307 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51545 | 56.081 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK479295 | Salmonella phage vB_SenS_SE1 | Salmonella phage vB_SenS_SE1 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40987 | 51.267 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN496308 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11519 | 38.311 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MN496307 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11607 | 38.149 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MN496306 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11729 | 38.341 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MN496305 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 13206 | 38.755 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| LR597645 | Escherichia phage Fraca | Escherichia phage Fraca Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43664 | 51.599 | Unclassified | Hendrixvirinae | Siphoviridae | Escherichia |
| LR597642 | Escherichia phage Evi | Escherichia phage Evi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40019 | 50.556 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MN536023 | Vibrio phage VP24-2_Ke | Vibrio phage VP24-2_Ke unclassified Fibrovirus Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7180 | 42.549 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| MK613349 | Roseobacter phage CRP-7 | Roseobacter phage CRP-7 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58106 | 40.294 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MK613348 | Roseobacter phage CRP-6 | Roseobacter phage CRP-6 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44927 | 47.047 | Unclassified | Unclassified | Unclassified | Roseobacter |
| MK613347 | Roseobacter phage CRP-5 | Roseobacter phage CRP-5 unclassified Siovirus Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39600 | 45.634 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| MK613346 | Roseobacter phage CRP-4 | Roseobacter phage CRP-4 unclassified Siovirus Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40768 | 45.197 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| MK613345 | Roseobacter phage CRP-3 | Roseobacter phage CRP-3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52963 | 49.665 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MK613344 | Roseobacter phage CRP-2 | Roseobacter phage CRP-2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54148 | 42.929 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MK613343 | Roseobacter phage CRP-1 | Roseobacter phage CRP-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54045 | 42.433 | Unclassified | Unclassified | Podoviridae | Roseobacter |
| MN026740 | Salmonella phage C2 | Salmonella phage C2 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38554 | 48.908 | Teseptimavirus | Studiervirinae | Autographiviridae | Salmonella |
| MN530981 | Campylobacter phage DA10 | Campylobacter phage DA10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35379 | 27.104 | Unclassified | Unclassified | Myoviridae | Campylobacter |
| MN395291 | Acinetobacter phage vB_AbaP_IME546 | Acinetobacter phage vB_AbaP_IME546 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40540 | 39.122 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN586063 | Mycobacterium phage Isca | Mycobacterium phage Isca unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49968 | 64.177 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586048 | Mycobacterium phage Fernando | Mycobacterium phage Fernando unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN586005 | Mycobacterium phage Lorde | Mycobacterium phage Lorde unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53852 | 61.203 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN585987 | Mycobacterium phage Enby | Mycobacterium phage Enby unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54687 | 61.276 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN615702 | Pseudomonas phage vB_PaeM_SMS29 | Pseudomonas phage vB_PaeM_SMS29 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66348 | 55.675 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN615701 | Pseudomonas phage vB_PaeM_SMS21 | Pseudomonas phage vB_PaeM_SMS21 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66362 | 55.622 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN615700 | Pseudomonas phage vB_PaeM_SMS12 | Pseudomonas phage vB_PaeM_SMS12 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66362 | 55.648 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN615699 | Pseudomonas phage vB_PaeP_SPCG | Pseudomonas phage vB_PaeP_SPCG unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42410 | 62.254 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN615698 | Pseudomonas phage vB_PaeP_SPCB | Pseudomonas phage vB_PaeP_SPCB unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43125 | 62.354 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN552150 | Lactococcus phage P4565 | Lactococcus phage P4565 Lactococcus virus P4565 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32697 | 34.236 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN552149 | Lactococcus phage P656 | Lactococcus phage P656 Lactococcus virus P656 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28396 | 35.075 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MN552147 | Leuconostoc phage P974 | Leuconostoc phage P974 unclassified Unaquatrovirus Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27942 | 36.032 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| MN552146 | Leuconostoc phage P965 | Leuconostoc phage P965 Leuconostoc virus P965 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26526 | 36.504 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| MN552145 | Lactococcus phage P1048 | Lactococcus phage P1048 Lactococcus virus P1048 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123023 | 32.967 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| MN552144 | Lactococcus phage P1045 | Lactococcus phage P1045 Lactococcus virus P1045 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31050 | 36.135 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| MN552143 | Lactococcus phage P1046 | Lactococcus phage P1046 unclassified Fremauxvirus Fremauxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54289 | 34.105 | Fremauxvirus | Unclassified | Siphoviridae | Lactococcus |
| MN534320 | Lactococcus phage proPhi7 | Lactococcus phage proPhi7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38158 | 35.243 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN534319 | Lactococcus phage proPhi6 | Lactococcus phage proPhi6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37410 | 35.405 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN534318 | Lactococcus phage proPhi5 | Lactococcus phage proPhi5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41249 | 35.916 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN534317 | Lactococcus phage proPhi4 | Lactococcus phage proPhi4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36976 | 35.607 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN534316 | Lactococcus phage proPhi2 | Lactococcus phage proPhi2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36976 | 35.607 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN534315 | Lactococcus phage proPhi1 | Lactococcus phage proPhi1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41249 | 35.916 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| MN336264 | Salmonella phage vB_SenS-EnJE1 | Salmonella phage vB_SenS-EnJE1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42894 | 49.758 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN313256 | Pseudoalteromonas phage XCL1123 | Pseudoalteromonas phage XCL1123 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45773 | 40.655 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MN497953 | Mycobacterium phage GtownJaz | Mycobacterium phage GtownJaz unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50849 | 64.017 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN617840 | Microbacterium phage Klimt | Microbacterium phage Klimt unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41573 | 63.368 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN379461 | Vibrio phage Rostov M3 | Vibrio phage Rostov M3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12308 | 45.734 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN379460 | Vibrio phage Rostov M3 | Vibrio phage Rostov M3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31129 | 45.797 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN524845 | Bacillus phage vB_Bpu_PumA2 | Bacillus phage vB_Bpu_PumA2 Bacillus virus PumA2 Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18932 | 35.802 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| MN524844 | Bacillus phage vB_Bpu_PumA1 | Bacillus phage vB_Bpu_PumA1 Bacillus virus PumA1 Bundooravirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18466 | 34.285 | Bundooravirus | Unclassified | Salasmaviridae | Bacillus |
| MN516422 | Acinetobacter phage Bphi-R1888 | Acinetobacter phage Bphi-R1888 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44590 | 37.874 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MN516421 | Acinetobacter phage Bphi-R2919 | Acinetobacter phage Bphi-R2919 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44227 | 37.767 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MN149904 | Klebsiella phage 31 | Klebsiella phage 31 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39600 | 50.434 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN149903 | Klebsiella phage 117 | Klebsiella phage 117 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41092 | 53.127 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN617835 | Enterobacter phage prasa_myo | Enterobacter phage prasa_myo unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172614 | 39.883 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MN614471 | Acinetobacter phage vB_AbaP_APK48-3 | Acinetobacter phage vB_AbaP_APK48-3 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41196 | 39.227 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MN369737 | Microbacterium phage Benjalauren | Microbacterium phage Benjalauren unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41561 | 63.502 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN617773 | Yersinia phage vB_Yen_X1 | Yersinia phage vB_Yen_X1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49948 | 46.967 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MK994522 | Methanobacterium virus PhiF1 | Methanobacterium virus PhiF1 unclassified bacterial viruses Viruses | 73979 | 45.551 | Unclassified | Unclassified | Unclassified | Methanobacterium |
| MK907263 | Escherichia phage vB_EcoS-26174I | Escherichia phage vB_EcoS-26174I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44091 | 44.422 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907276 | Escherichia phage vB_EcoS-2862V | Escherichia phage vB_EcoS-2862V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.397 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907275 | Escherichia phage vB_EcoS-2862IV | Escherichia phage vB_EcoS-2862IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.451 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907274 | Escherichia phage vB_EcoS-2862III | Escherichia phage vB_EcoS-2862III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.44 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907273 | Escherichia phage vB_EcoS-2862II | Escherichia phage vB_EcoS-2862II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.427 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907272 | Escherichia phage vB_EcoS-2862I | Escherichia phage vB_EcoS-2862I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.404 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907271 | Escherichia phage vB_EcoS-26175V | Escherichia phage vB_EcoS-26175V unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114646 | 39.948 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK907270 | Escherichia phage vB_EcoS-26175IV | Escherichia phage vB_EcoS-26175IV unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114646 | 39.952 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK907269 | Escherichia phage vB_EcoS-26175III | Escherichia phage vB_EcoS-26175III unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114647 | 39.95 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK907268 | Escherichia phage vB_EcoS-26175II | Escherichia phage vB_EcoS-26175II unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114646 | 39.961 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK907267 | Escherichia phage vB_EcoS-26175I | Escherichia phage vB_EcoS-26175I unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114649 | 39.952 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK907264 | Escherichia phage vB_EcoS-26174II | Escherichia phage vB_EcoS-26174II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.395 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907262 | Escherichia phage vB_EcoS-26047II | Escherichia phage vB_EcoS-26047II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.438 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907261 | Escherichia phage vB_EcoS-26047I | Escherichia phage vB_EcoS-26047I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.402 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907260 | Escherichia phage vB_EcoS-26046V | Escherichia phage vB_EcoS-26046V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.39 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907259 | Escherichia phage vB_EcoS-26046IV | Escherichia phage vB_EcoS-26046IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.399 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907257 | Escherichia phage vB_EcoS-26046II | Escherichia phage vB_EcoS-26046II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.402 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907255 | Escherichia phage vB_EcoS-26020V | Escherichia phage vB_EcoS-26020V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.427 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907254 | Escherichia phage vB_EcoS-26020IV | Escherichia phage vB_EcoS-26020IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.451 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907253 | Escherichia phage vB_EcoS-26020III | Escherichia phage vB_EcoS-26020III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.422 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907252 | Escherichia phage vB_EcoS-26020II | Escherichia phage vB_EcoS-26020II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.458 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907251 | Escherichia phage vB_EcoS-26020I | Escherichia phage vB_EcoS-26020I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.413 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907249 | Escherichia phage vB_EcoS-25988IV | Escherichia phage vB_EcoS-25988IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.44 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907248 | Escherichia phage vB_EcoS-25988I | Escherichia phage vB_EcoS-25988I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.431 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907247 | Escherichia phage vB_EcoS-2006V | Escherichia phage vB_EcoS-2006V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.438 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907246 | Escherichia phage vB_EcoS-2006IV | Escherichia phage vB_EcoS-2006IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.413 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907245 | Escherichia phage vB_EcoS-2006III | Escherichia phage vB_EcoS-2006III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.424 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907244 | Escherichia phage vB_EcoS-2005IV | Escherichia phage vB_EcoS-2005IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.44 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907243 | Escherichia phage vB_EcoS-2005III | Escherichia phage vB_EcoS-2005III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.395 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907241 | Escherichia phage vB_EcoS-2004IV | Escherichia phage vB_EcoS-2004IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.422 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907240 | Escherichia phage vB_EcoS-2004III | Escherichia phage vB_EcoS-2004III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.386 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907239 | Escherichia phage vB_EcoM-12474V | Escherichia phage vB_EcoM-12474V unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33714 | 50.911 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| MK907238 | Escherichia phage vB_EcoM-12474IV | Escherichia phage vB_EcoM-12474IV unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33731 | 50.903 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| MK907237 | Escherichia phage vB_EcoM-12474III | Escherichia phage vB_EcoM-12474III Escherichia virus 12474III Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33807 | 50.918 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| MK907236 | Escherichia phage vB_EcoM-12474II | Escherichia phage vB_EcoM-12474II unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33688 | 50.896 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| MK907235 | Escherichia phage vB_EcoS-12469III | Escherichia phage vB_EcoS-12469III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.409 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907233 | Escherichia phage vB_EcoS-12469I | Escherichia phage vB_EcoS-12469I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.422 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907231 | Escherichia phage vB_EcoS-12397IV | Escherichia phage vB_EcoS-12397IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.386 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907230 | Escherichia phage vB_EcoS-12397III | Escherichia phage vB_EcoS-12397III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.406 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907229 | Escherichia phage vB_EcoS-12397II | Escherichia phage vB_EcoS-12397II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.424 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907228 | Escherichia phage vB_EcoS-12397I | Escherichia phage vB_EcoS-12397I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.409 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907227 | Escherichia phage vB_EcoS-12210III | Escherichia phage vB_EcoS-12210III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.433 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907226 | Escherichia phage vB_EcoS-12210I | Escherichia phage vB_EcoS-12210I Escherichia virus 12210I Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.413 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907266 | Escherichia phage vB_EcoS-26174IV | Escherichia phage vB_EcoS-26174IV unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.431 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907265 | Escherichia phage vB_EcoS-26174III | Escherichia phage vB_EcoS-26174III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.427 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907258 | Escherichia phage vB_EcoS-26046III | Escherichia phage vB_EcoS-26046III unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.384 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907256 | Escherichia phage vB_EcoS-26046I | Escherichia phage vB_EcoS-26046I unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.42 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907250 | Escherichia phage vB_EcoS-25988V | Escherichia phage vB_EcoS-25988V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.393 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907242 | Escherichia phage vB_EcoS-2004V | Escherichia phage vB_EcoS-2004V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.395 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907234 | Escherichia phage vB_EcoS-12469II | Escherichia phage vB_EcoS-12469II unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.402 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK907232 | Escherichia phage vB_EcoS-12397V | Escherichia phage vB_EcoS-12397V unclassified Veterinaerplatzvirus Veterinaerplatzvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44219 | 44.445 | Veterinaerplatzvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MN308080 | Pectobacterium phage MA1A | Pectobacterium phage MA1A unclassified Pektosvirus Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39781 | 48.77 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MN478377 | Bacteriophage Eos | Bacteriophage Eos unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109015 | 41.908 | Sugarlandvirus | Unclassified | Demerecviridae | Unspecified |
| MN478375 | Bacteriophage Titan-X | Bacteriophage Titan-X unclassified Pradovirus Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44062 | 61.084 | Pradovirus | Unclassified | Autographiviridae | Unspecified |
| MN478374 | Bacteriophage Phobos | Bacteriophage Phobos Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56734 | 63.324 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MK268686 | Enterococcus phage vB_EfaH_EF1TV | Enterococcus phage vB_EfaH_EF1TV unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143507 | 35.843 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MK170447 | Methanosarcina virus MetMV | Methanosarcina virus MetMV unclassified bacterial viruses Viruses | 67826 | 42.373 | Unclassified | Unclassified | Unclassified | Methanosarcina |
| MK170446 | Klebsiella phage 13 | Klebsiella phage 13 Klebsiella virus 13 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43094 | 50.629 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK135468 | Klebsiella phage 1 TK-2018 | Klebsiella phage 1 TK-2018 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43584 | 53.969 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MH853788 | Acinetobacter phage vB_AbaM_IME512 | Acinetobacter phage vB_AbaM_IME512 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46499 | 37.558 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MH853787 | Acinetobacter phage vB_AbaM_IME284 | Acinetobacter phage vB_AbaM_IME284 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43557 | 38.302 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MH853786 | Acinetobacter phage vB_AbaM_IME285 | Acinetobacter phage vB_AbaM_IME285 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45063 | 37.869 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| MN311492 | Antarctic microvirus TYR_006_V_25 | Antarctic microvirus TYR_006_V_25 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4281 | 38.192 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN311491 | Antarctic microvirus COCH21_V_SP_16 | Antarctic microvirus COCH21_V_SP_16 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4201 | 42.561 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN311490 | Antarctic microvirus COCH21_V_SP_13 | Antarctic microvirus COCH21_V_SP_13 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4317 | 47.047 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN311489 | Antarctic microvirus CAA_003_V_9 | Antarctic microvirus CAA_003_V_9 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4254 | 39.539 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN602881 | Erwinia phage Midgardsormr38 | Erwinia phage Midgardsormr38 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50485 | 50.859 | Unclassified | Unclassified | Siphoviridae | Erwinia |
| MN604230 | Bacillus phage vB_BcM_Sam112 | Bacillus phage vB_BcM_Sam112 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45037 | 41.621 | Unclassified | Unclassified | Myoviridae | Bacillus |
| MN434095 | Klebsiella phage JIPh_Kp122 | Klebsiella phage JIPh_Kp122 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166475 | 39.576 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN585130 | Klebsiella phage vB_Kp168 | Klebsiella phage vB_Kp168 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40114 | 53.283 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN536027 | Pseudomonas phage vB_Pae-SS2019XII | Pseudomonas phage vB_Pae-SS2019XII unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39701 | 63.608 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MN528768 | Lactococcus phage P596 | Lactococcus phage P596 Lactococcus virus P596 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57909 | 34.364 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| MN505213 | Serratia phage JS26 | Serratia phage JS26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63971 | 57.013 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MN481367 | Salmonella phage D1-2 | Salmonella phage D1-2 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86878 | 38.734 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MN602045 | Pseudomonas phage vB_PaeP_ASP23 | Pseudomonas phage vB_PaeP_ASP23 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42735 | 62.162 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MN342247 | Shigella phage Sfin-5 | Shigella phage Sfin-5 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50411 | 45.196 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MN340231 | Escherichia phage phiv205-1 | Escherichia phage phiv205-1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44616 | 47.543 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MN337573 | Shigella phage Sfin-4 | Shigella phage Sfin-4 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50407 | 45.202 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MN327636 | Pectobacterium phage MA6 | Pectobacterium phage MA6 unclassified Pektosvirus Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38553 | 49.083 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MN296515 | Shigella virus Sfin-3 | Shigella virus Sfin-3 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50309 | 45.352 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MN271656 | Pectobacterium phage MA2 | Pectobacterium phage MA2 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41857 | 51.774 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MN187550 | Escherichia phage phiv142-3 | Escherichia phage phiv142-3 unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39293 | 48.965 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| MN311493 | Antarctic microvirus TYR_006_V_SP_13 | Antarctic microvirus TYR_006_V_SP_13 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5286 | 39.009 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN311488 | Antarctic microvirus CAA_003_V_4 | Antarctic microvirus CAA_003_V_4 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4135 | 43.071 | Microvirus | Unclassified | Microviridae | Unspecified |
| MN311487 | Antarctic microvirus CAA_003_V_1 | Antarctic microvirus CAA_003_V_1 unclassified Microvirus Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4288 | 38.619 | Microvirus | Unclassified | Microviridae | Unspecified |
| LC507823 | Salmonella phage vB_StyS-sam | Salmonella phage vB_StyS-sam unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43221 | 49.765 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN106244 | Klebsiella phage Soft | Klebsiella phage Soft Klebsiella virus Soft Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57805 | 55.871 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MH631454 | Salmonella phage Sw2 | Salmonella phage Sw2 Salmonella virus Sw2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114274 | 40.214 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK608336 | Serratia phage Serbin | Serratia phage Serbin Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42882 | 51.618 | Unclassified | Unclassified | Siphoviridae | Serratia |
| MK737937 | Raoultella phage RP180 | Raoultella phage RP180 Raoultella virus RP180 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44851 | 50.342 | Kagunavirus | Guernseyvirinae | Siphoviridae | Raoultella |
| MG030347 | Proteus phage PM135 | Proteus phage PM135 Proteus virus PM135 Novosibvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104329 | 38.369 | Novosibvirus | Unclassified | Demerecviridae | Proteus |
| MG030346 | Proteus phage PM87 | Proteus phage PM87 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59128 | 46.729 | Unclassified | Unclassified | Siphoviridae | Proteus |
| KX965989 | Aeribacillus phage AP45 | Aeribacillus phage AP45 Aeribacillus virus AP45 Kamchatkavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51606 | 38.273 | Kamchatkavirus | Unclassified | Siphoviridae | Aeribacillus |
| KU946962 | Proteus phage PM 116 | Proteus phage PM 116 Proteus virus PM116 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44601 | 39.217 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| KM819696 | Proteus phage PM 93 | Proteus phage PM 93 Proteus virus PM93 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45169 | 39.363 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| KM819695 | Proteus phage PM 85 | Proteus phage PM 85 Proteus virus PM85 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43642 | 39.322 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| KM819694 | Proteus phage PM 75 | Proteus phage PM 75 Proteus virus PM75 Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41480 | 41.56 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| KF319020 | Proteus phage PM16 | Proteus phage PM16 Proteus virus PM16 Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41268 | 41.386 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| LC494302 | Escherichia phage SP27 | Escherichia phage SP27 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 347152 | 34.17 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MH356729 | Pseudomonas phage LP14 | Pseudomonas phage LP14 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73080 | 54.982 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MK903728 | Klebsiella phage SH-KP152226 | Klebsiella phage SH-KP152226 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41420 | 52.704 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK972831 | Shigella phage Sfin-2 | Shigella phage Sfin-2 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50390 | 45.213 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MN016939 | Bacteriophage R18C | Bacteriophage R18C Citrobacter virus R18C Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31834 | 51.602 | Peduovirus | Peduovirinae | Myoviridae | Unspecified |
| MK837012 | Pseudomonas virus Pa223 | Pseudomonas virus Pa223 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45703 | 52.491 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MK837011 | Pseudomonas virus Pa222 | Pseudomonas virus Pa222 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45770 | 52.609 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MK837010 | Pseudomonas virus Pa204 | Pseudomonas virus Pa204 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65924 | 55.579 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK837009 | Pseudomonas virus Pa193 | Pseudomonas virus Pa193 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66657 | 55.672 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN584918 | Vibrio phage phi50-12 | Vibrio phage phi50-12 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68059 | 38.452 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MN508356 | Acinetobacter phage VB_ApiP_XC38 | Acinetobacter phage VB_ApiP_XC38 unclassified Presleyvirus Presleyvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79328 | 39.579 | Presleyvirus | Unclassified | Schitoviridae | Acinetobacter |
| MN504636 | Pseudomonas phage Quinobequin-P09 | Pseudomonas phage Quinobequin-P09 unclassified Nipunavirus Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58277 | 58.342 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| MN461279 | Xanthomonas virus phiXaf18 | Xanthomonas virus phiXaf18 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47407 | 62.97 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MN419153 | Vibrio phage ICP1_2017_F_Mathbaria | Vibrio phage ICP1_2017_F_Mathbaria Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129621 | 37.117 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MN393079 | Salmonella virus VSiA | Salmonella virus VSiA unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42865 | 51.21 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN393078 | Salmonella virus VSe101 | Salmonella virus VSe101 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41757 | 50.056 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MN434096 | Klebsiella phage JIPh_Kp127 | Klebsiella phage JIPh_Kp127 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113671 | 45.267 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MN434094 | Klebsiella phage AmPh_EK80 | Klebsiella phage AmPh_EK80 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112215 | 44.816 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MN434093 | Klebsiella phage AmPh_EK52 | Klebsiella phage AmPh_EK52 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40713 | 52.936 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN434092 | Klebsiella phage AmPh_EK29 | Klebsiella phage AmPh_EK29 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169353 | 40.773 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MN310548 | Mycobacterium phage Juice456 | Mycobacterium phage Juice456 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56726 | 61.612 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MN176230 | Bacillus phage 056SW001B | Bacillus phage 056SW001B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52692 | 42.547 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176228 | Bacillus phage 049ML003 | Bacillus phage 049ML003 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44817 | 43.671 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176227 | Bacillus phage 049ML001 | Bacillus phage 049ML001 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45238 | 43.746 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176219 | Bacillus phage 000TH010 | Bacillus phage 000TH010 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46276 | 43.843 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN508817 | Yersinia phage JC221 | Yersinia phage JC221 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174931 | 41.231 | Unclassified | Tevenvirinae | Myoviridae | Yersinia |
| MN450150 | Pantoea phage vB_PagP-SK1 | Pantoea phage vB_PagP-SK1 unclassified Elunavirus Elunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39938 | 52.324 | Elunavirus | Studiervirinae | Autographiviridae | Pantoea |
| MN380459 | Klebsiella phage vB_KpnP_fHeKpn01 | Klebsiella phage vB_KpnP_fHeKpn01 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43329 | 53.599 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MF033351 | Acinetobacter phage vB_ApiP_P2 | Acinetobacter phage vB_ApiP_P2 Acinetobacter virus P2 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41514 | 39.334 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MF033350 | Acinetobacter phage vB_ApiP_P1 | Acinetobacter phage vB_ApiP_P1 Acinetobacter virus P1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41208 | 39.196 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MF033349 | Acinetobacter phage vB_AbaP_B5 | Acinetobacter phage vB_AbaP_B5 Acinetobacter virus B5 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41608 | 39.31 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MF033348 | Acinetobacter phage vB_AbaP_B3 | Acinetobacter phage vB_AbaP_B3 Acinetobacter virus B3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40598 | 39.275 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MF033347 | Acinetobacter phage vB_AbaP_B1 | Acinetobacter phage vB_AbaP_B1 Acinetobacter virus B1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40879 | 39.145 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MG983743 | Xanthomonas phage RiverRider | Xanthomonas phage RiverRider Xanthomonas virus RiverRider Riverridervirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76335 | 49.393 | Riverridervirus | Unclassified | Schitoviridae | Xanthomonas |
| MN065183 | Bacillus phage vB_BthS-HD29phi | Bacillus phage vB_BthS-HD29phi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32181 | 34.856 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN379832 | Klebsiella phage KL | Klebsiella phage KL Klebsiella virus KL Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47844 | 50.907 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN284901 | Microbacterium phage YuuY | Microbacterium phage YuuY unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40996 | 63.238 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MN284900 | Mycobacterium phage JeppNRM | Mycobacterium phage JeppNRM unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50575 | 63.96 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN270892 | Pectobacterium phage Wc4-1 | Pectobacterium phage Wc4-1 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92039 | 44.703 | Unclassified | Ounavirinae | Myoviridae | Pectobacterium |
| MN270891 | Pectobacterium phage Wc4 | Pectobacterium phage Wc4 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92039 | 44.704 | Unclassified | Ounavirinae | Myoviridae | Pectobacterium |
| MN270890 | Ralstonia phage P-PSG-11-1 | Ralstonia phage P-PSG-11-1 unclassified Gyeongsanvirus Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40313 | 58.807 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MN270889 | Ralstonia phage P-PSG-11 | Ralstonia phage P-PSG-11 unclassified Gyeongsanvirus Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40313 | 58.805 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MN270888 | Pectobacterium phage CX5-1 | Pectobacterium phage CX5-1 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43885 | 51.036 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MN270887 | Pectobacterium phage CX5 | Pectobacterium phage CX5 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43885 | 51.033 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MN399336 | Edwardsiella phage vB_EtaM_ET-ABTNL-9 | Edwardsiella phage vB_EtaM_ET-ABTNL-9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89762 | 37.256 | Unclassified | Unclassified | Myoviridae | Edwardsiella |
| MN334766 | Serratia phage PCH45 | Serratia phage PCH45 unclassified Iapetusvirus Iapetusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 212807 | 47.281 | Iapetusvirus | Unclassified | Myoviridae | Serratia |
| MN164484 | Escherichia virus ECH1 | Escherichia virus ECH1 unclassified Guelphvirus Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49553 | 43.565 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MN510331 | Escherichia phage vB_EcoP_PR_Kaz2018 | Escherichia phage vB_EcoP_PR_Kaz2018 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39704 | 49.869 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK061416 | Rothia phage Spartoi | Rothia phage Spartoi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35189 | 52.38 | Unclassified | Unclassified | Siphoviridae | Rothia |
| MH165274 | Acinetobacter phage AM101 | Acinetobacter phage AM101 Acinetobacter virus AM101 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166487 | 36.708 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MN172529 | Listeria phage LP-039 | Listeria phage LP-039 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136234 | 35.914 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MN128594 | Listeria phage LP-066 | Listeria phage LP-066 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138918 | 35.833 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MK416017 | Klebsiella phage ST101-KPC2phi6.3 | Klebsiella phage ST101-KPC2phi6.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43942 | 52.237 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK455769 | Aeromonas phage MJG | Aeromonas phage MJG unclassified Uliginvirus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45057 | 50.105 | Uliginvirus | Colwellvirinae | Autographiviridae | Aeromonas |
| MG432137 | Pectinobacterium phage PEAT2 | Pectinobacterium phage PEAT2 Pectinobacterium virus PEAT2 Peatvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48659 | 49.126 | Peatvirus | Unclassified | Myoviridae | Pectobacterium |
| MN234222 | Microbacterium phage Christoph | Microbacterium phage Christoph unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41554 | 63.421 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN234204 | Microbacterium phage ManRay | Microbacterium phage ManRay unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.436 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN234181 | Microbacterium phage Pocket | Microbacterium phage Pocket unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41583 | 63.382 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN234177 | Microbacterium phage Gershwin | Microbacterium phage Gershwin unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.393 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN234170 | Mycobacterium phage MalagasyRose | Mycobacterium phage MalagasyRose Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54301 | 65.264 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MN234163 | Microbacterium phage Calix | Microbacterium phage Calix unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.355 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN212906 | Acinetobacter phage vB_AbaP_D2M | Acinetobacter phage vB_AbaP_D2M unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39963 | 39.236 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK813940 | Aeromonas phage 4_L372X | Aeromonas phage 4_L372X Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45698 | 48.744 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MK947460 | Salmonella phage vB_SenS_SB28 | Salmonella phage vB_SenS_SB28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45126 | 46.164 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MK947459 | Salmonella phage vB_SenS_SB13 | Salmonella phage vB_SenS_SB13 Salmonella virus SB13 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112513 | 39.921 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK947458 | Salmonella phage vB_SenS_SB10 | Salmonella phage vB_SenS_SB10 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111347 | 40.057 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN218775 | Escherichia virus ECG4 | Escherichia virus ECG4 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40792 | 49.953 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MN266305 | Shigella phage SGF3 | Shigella phage SGF3 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.838 | Sinsheimervirus | Bullavirinae | Microviridae | Shigella |
| MN176232 | Bacillus phage 280BB001 | Bacillus phage 280BB001 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52371 | 42.508 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176231 | Bacillus phage 276BB001 | Bacillus phage 276BB001 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52370 | 42.507 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176229 | Bacillus phage 055SW001 | Bacillus phage 055SW001 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55382 | 44.188 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176226 | Bacillus phage 035JT001 | Bacillus phage 035JT001 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55312 | 44.367 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176225 | Bacillus phage 031MP004 | Bacillus phage 031MP004 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55687 | 44.215 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176224 | Bacillus phage 031MP002 | Bacillus phage 031MP002 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55901 | 44.205 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176223 | Bacillus phage 031MP003 | Bacillus phage 031MP003 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56127 | 44.193 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176222 | Bacillus phage 022DV001 | Bacillus phage 022DV001 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55479 | 44.163 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MN176221 | Bacillus phage 019DV004 | Bacillus phage 019DV004 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52749 | 42.643 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN176220 | Bacillus phage 019DV002 | Bacillus phage 019DV002 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52749 | 42.64 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN310543 | Microbacterium phage ChickenKing | Microbacterium phage ChickenKing unclassified Schubertvirus Schubertvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39924 | 63.353 | Schubertvirus | Unclassified | Siphoviridae | Microbacterium |
| MN317029 | Aeromonas phage vB_AhyS-A18P4 | Aeromonas phage vB_AhyS-A18P4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60975 | 62.021 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MK714353 | Klebsiella phage ST13-OXA48phi12.5 | Klebsiella phage ST13-OXA48phi12.5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44913 | 51.791 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK533143 | Paenibacillus phage vB_PlaP_API480 | Paenibacillus phage vB_PlaP_API480 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45026 | 39.24 | Unclassified | Unclassified | Podoviridae | Paenibacillus |
| MK416013 | Klebsiella phage ST16-OXA48phi5.1 | Klebsiella phage ST16-OXA48phi5.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57025 | 53.552 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK416018 | Klebsiella phage ST147-VIM1phi7.1 | Klebsiella phage ST147-VIM1phi7.1 Klebsiella virus ST147VIM1phi7-1 Vimunumvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34141 | 52.969 | Vimunumvirus | Peduovirinae | Myoviridae | Klebsiella |
| MK448235 | Klebsiella phage ST512-KPC3phi13.1 | Klebsiella phage ST512-KPC3phi13.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42666 | 52.862 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448233 | Klebsiella phage ST11-VIM1phi8.1 | Klebsiella phage ST11-VIM1phi8.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42666 | 52.862 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448237 | Klebsiella phage ST974-OXA48phi18.2 | Klebsiella phage ST974-OXA48phi18.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51967 | 52.851 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448234 | Klebsiella phage ST11-VIM1phi8.2 | Klebsiella phage ST11-VIM1phi8.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48230 | 52.65 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448232 | Klebsiella phage ST147-VIM1phi7.2 | Klebsiella phage ST147-VIM1phi7.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34200 | 51.743 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448231 | Klebsiella phage ST101-KPC2phi6.1 | Klebsiella phage ST101-KPC2phi6.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48131 | 52.602 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK448230 | Klebsiella phage ST16-OXA48phi5.2 | Klebsiella phage ST16-OXA48phi5.2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47305 | 53.791 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK433583 | Klebsiella phage ST899-OXA48phi17.2 | Klebsiella phage ST899-OXA48phi17.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17998 | 50.606 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK433579 | Klebsiella phage ST11-OXA48phi15.1 | Klebsiella phage ST11-OXA48phi15.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36137 | 51.748 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK422455 | Klebsiella phage ST340-VIM1phi10.1 | Klebsiella phage ST340-VIM1phi10.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36124 | 51.747 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK422451 | Klebsiella phage ST13-OXA48phi12.3 | Klebsiella phage ST13-OXA48phi12.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84199 | 49.912 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK416007 | Klebsiella phage ST405-OXA48phi1.2 | Klebsiella phage ST405-OXA48phi1.2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40495 | 52.135 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK416014 | Klebsiella phage ST16-OXA48phi5.3 | Klebsiella phage ST16-OXA48phi5.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29301 | 50.179 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK416015 | Klebsiella phage ST16-OXA48phi5.4 | Klebsiella phage ST16-OXA48phi5.4 Klebsiella virus ST16OXA48phi5-4 Reipivirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38322 | 50.06 | Reipivirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416019 | Klebsiella phage ST11-VIM1phi8.3 | Klebsiella phage ST11-VIM1phi8.3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39953 | 51.936 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK416022 | Klebsiella phage ST846-OXA48phi9.2 | Klebsiella phage ST846-OXA48phi9.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57402 | 51.646 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KX765863 | Salmonella phage ST-118 | Salmonella phage ST-118 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59106 | 56.698 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KX765862 | Salmonella phage ST-101 | Salmonella phage ST-101 Salmonella virus ST101 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59132 | 56.702 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MK257723 | Acinetobacter phage vB_AbaP_APK37 | Acinetobacter phage vB_AbaP_APK37 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41981 | 39.156 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK257722 | Acinetobacter phage vB_AbaP_APK32 | Acinetobacter phage vB_AbaP_APK32 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 39.313 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK257720 | Acinetobacter phage vB_AbaP_APK2-2 | Acinetobacter phage vB_AbaP_APK2-2 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41426 | 39.379 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK257719 | Acinetobacter phage vB_AbaP_APK2 | Acinetobacter phage vB_AbaP_APK2 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41476 | 39.24 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK660712 | Gordonia phage EpicDab | Gordonia phage EpicDab Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16658 | 69.126 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH707430 | Bacillus phage BSP7 | Bacillus phage BSP7 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55455 | 39.915 | Unclassified | Unclassified | Podoviridae | Bacillus |
| KY087898 | Pectobacterium phage PP101 | Pectobacterium phage PP101 Pectobacterium virus PP101 Suwonvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53333 | 44.944 | Suwonvirus | Cleopatravirinae | Chaseviridae | Pectobacterium |
| MK733264 | Yersinia phage YeP6 | Yersinia phage YeP6 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34969 | 50.342 | Unclassified | Unclassified | Podoviridae | Yersinia |
| MK733263 | Yersinia phage YeP5 | Yersinia phage YeP5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34969 | 50.342 | Unclassified | Unclassified | Podoviridae | Yersinia |
| MK733261 | Yersinia phage YeP3 | Yersinia phage YeP3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39751 | 48.039 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MK733260 | Yersinia phage YeP2 | Yersinia phage YeP2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39751 | 48.044 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MK733259 | Yersinia phage YeP1 | Yersinia phage YeP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39492 | 48.111 | Unclassified | Unclassified | Myoviridae | Yersinia |
| MK733262 | Yersinia phage YeP4 | Yersinia phage YeP4 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35029 | 50.373 | Unclassified | Unclassified | Podoviridae | Yersinia |
| MK557849 | Caudovirales GX15bay | Caudovirales GX15bay Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52012 | 63.416 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MK017820 | Clostridium phage CPD7 | Clostridium phage CPD7 unclassified Susfortunavirus Susfortunavirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18958 | 29.122 | Susfortunavirus | Unclassified | Guelinviridae | Clostridium |
| MK017819 | Clostridium phage CPD4 | Clostridium phage CPD4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52658 | 34.09 | Unclassified | Unclassified | Myoviridae | Clostridium |
| MH999280 | Clostridium phage CPD1 | Clostridium phage CPD1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39531 | 30.255 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| MH999279 | Clostridium phage CPD2 | Clostridium phage CPD2 Clostridium virus CPD2 Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18212 | 27.943 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| MH746817 | Achromobacter phage vB_Ade_ART | Achromobacter phage vB_Ade_ART Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95343 | 55.036 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| MK931440 | Escherichia phage Shashou | Escherichia phage Shashou unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44155 | 50.087 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MN166823 | Klebsiella phage ST512-KPC3phi13.5 | Klebsiella phage ST512-KPC3phi13.5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25624 | 51.998 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MN184887 | Erwinia phage pEp_SNUABM_01 | Erwinia phage pEp_SNUABM_01 Erwinia virus SNUABM01 Henunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147321 | 48.706 | Henunavirus | Vequintavirinae | Myoviridae | Erwinia |
| MN184886 | Erwinia phage pEp_SNUABM_08 | Erwinia phage pEp_SNUABM_08 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62716 | 57.244 | Unclassified | Unclassified | Siphoviridae | Erwinia |
| MN184885 | Erwinia phage pEp_SNUABM_09 | Erwinia phage pEp_SNUABM_09 unclassified Studiervirinae Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39848 | 52.166 | Unclassified | Studiervirinae | Autographiviridae | Erwinia |
| MN304941 | Staphylococcus phage vB_SauH_IME522 | Staphylococcus phage vB_SauH_IME522 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140246 | 30.192 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MN176574 | Klebsiella phage vB_KpnP_IME335 | Klebsiella phage vB_KpnP_IME335 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40361 | 53.133 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN176573 | Klebsiella phage vB_KpnP_IME337 | Klebsiella phage vB_KpnP_IME337 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44266 | 53.666 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MN176572 | Klebsiella phage vB_KpnP_IME308 | Klebsiella phage vB_KpnP_IME308 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43091 | 53.888 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MH766655 | Curvibacter phage TJ1 | Curvibacter phage TJ1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37136 | 58.151 | Unclassified | Unclassified | Myoviridae | Curvibacter |
| KX452700 | Enterobacter phage KNP7 | Enterobacter phage KNP7 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58058 | 56.48 | Chivirus | Unclassified | Siphoviridae | Enterobacter |
| KX452699 | Salmonella phage KNP6 | Salmonella phage KNP6 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59230 | 55.756 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KX452698 | Shigella phage KNP5 | Shigella phage KNP5 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193624 | 30.374 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KX452697 | Serratia phage KNP4 | Serratia phage KNP4 unclassified Miltonvirus Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160268 | 48.84 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| KX452696 | Enterobacter phage KNP3 | Enterobacter phage KNP3 Enterobacter virus KPN3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44911 | 39.193 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| KX452695 | Klebsiella phage KNP2 | Klebsiella phage KNP2 Klebsiella virus KNP2 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146203 | 38.548 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| MN252582 | Salmonella phage LPST144 | Salmonella phage LPST144 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39050 | 49.078 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| MN215888 | Vibrio phage vB_VpaS_HCMJ | Vibrio phage vB_VpaS_HCMJ unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76200 | 48.85 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MN385565 | Escherichia virus phiX174 | Escherichia virus phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| MN276049 | Acinetobacter phage BS46 | Acinetobacter phage BS46 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94068 | 33.453 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MN191861 | Burkholderia phage AMP1 | Burkholderia phage AMP1 unclassified Ampunavirus Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42798 | 61.751 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| MN337887 | Lactococcus phage vB_LacS_15 | Lactococcus phage vB_LacS_15 Lactococcus virus LacS15 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21006 | 36.142 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MN159079 | Aeromonas phage D9 | Aeromonas phage D9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 260857 | 47.479 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MN163281 | Klebsiella phage KpCHEMY26 | Klebsiella phage KpCHEMY26 Klebsiella virus KpCHEMY26 Pylasvirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70678 | 41.716 | Pylasvirus | Humphriesvirinae | Schitoviridae | Klebsiella |
| MH729379 | Aerococcus phage vB_AviM_AVP | Aerococcus phage vB_AviM_AVP unclassified Herelleviridae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133806 | 34.505 | Unclassified | Unclassified | Herelleviridae | Aerococcus |
| MN187969 | Salmonella phage OSY-STA | Salmonella phage OSY-STA Salmonella virus OSYSTA Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111039 | 40.048 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN131141 | Pseudomonas virus PaP1_EPu-2019 | Pseudomonas virus PaP1_EPu-2019 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66612 | 55.76 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK972696 | Salmonella phage Si3 KFo-2019 | Salmonella phage Si3 KFo-2019 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40012 | 46.833 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| NC_040375 | Alces alces faeces associated microvirus MP3 6497 | Alces alces faeces associated microvirus MP3 6497 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6434 | 36.509 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040374 | Alces alces faeces associated microvirus MP15 5067 | Alces alces faeces associated microvirus MP15 5067 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5004 | 36.771 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040373 | Alces alces faeces associated microvirus MP18 4940 | Alces alces faeces associated microvirus MP18 4940 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4892 | 36.958 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040350 | Alces alces faeces associated microvirus MP10 5560 | Alces alces faeces associated microvirus MP10 5560 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5497 | 38.039 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040349 | Alces alces faeces associated microvirus MP21 4718 | Alces alces faeces associated microvirus MP21 4718 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4662 | 48.263 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040342 | Alces alces faeces associated microvirus MP11 5517 | Alces alces faeces associated microvirus MP11 5517 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5398 | 32.29 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040341 | Lepus americanus faeces associated microvirus SHP1 6472 | Lepus americanus faeces associated microvirus SHP1 6472 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6357 | 37.124 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040329 | Lynx canadensis associated microvirus CLP 9413 | Lynx canadensis associated microvirus CLP 9413 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5312 | 49.812 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_040328 | Alces alces faeces associated microvirus MP12 5423 | Alces alces faeces associated microvirus MP12 5423 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5360 | 37.91 | Unclassified | Unclassified | Microviridae | Unspecified |
| LR588166 | Pseudomonas phage vB_PaeM_MIJ3 | Pseudomonas phage vB_PaeM_MIJ3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 288170 | 33.349 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MN153503 | Bacillus phage Chotacabras | Bacillus phage Chotacabras unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165032 | 37.584 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN131143 | Pseudomonas virus PA11P1 | Pseudomonas virus PA11P1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66049 | 55.739 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN131142 | Pseudomonas virus PA8P1 | Pseudomonas virus PA8P1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65690 | 55.695 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MN148435 | Shigella phage SGF2 | Shigella phage SGF2 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76964 | 42.298 | Kuravirus | Unclassified | Podoviridae | Shigella |
| MN103543 | Pseudomonas phage vB_PaeM_PS119XW | Pseudomonas phage vB_PaeM_PS119XW unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 301543 | 43.62 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MN103542 | Enterococcus phage vB_EfaS_EF1c55 | Enterococcus phage vB_EfaS_EF1c55 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55876 | 39.783 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MK799650 | Pseudomonas phage PAP-JP | Pseudomonas phage PAP-JP unclassified Nankokuvirus Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87872 | 54.765 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| MK359990 | Streptococcus phage phi-SC181 | Streptococcus phage phi-SC181 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59334 | 39.099 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| LR535913 | Salmonella phage SPFM15 | Salmonella phage SPFM15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239951 | 48.632 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535912 | Salmonella phage SPFM14 | Salmonella phage SPFM14 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.612 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535904 | Salmonella phage SPFM7 | Salmonella phage SPFM7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.621 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MK016663 | Synechococcus phage S-H68 | Synechococcus phage S-H68 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79639 | 40.366 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MN335165 | Jodiemicrovirus-1 | Jodiemicrovirus-1 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4768 | 33.851 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| MK972717 | Salmonella phage SE18 | Salmonella phage SE18 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110795 | 40.095 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972716 | Salmonella phage SE8 | Salmonella phage SE8 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107763 | 39.196 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972715 | Salmonella phage SE11 | Salmonella phage SE11 Salmonella virus SE11 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107763 | 39.195 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972714 | Salmonella phage SW3 | Salmonella phage SW3 unclassified Irrigatiovirus Irrigatiovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30037 | 54.353 | Irrigatiovirus | Peduovirinae | Myoviridae | Salmonella |
| MK972713 | Salmonella phage SW5 | Salmonella phage SW5 unclassified Irrigatiovirus Irrigatiovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30046 | 54.357 | Irrigatiovirus | Peduovirinae | Myoviridae | Salmonella |
| MK972712 | Salmonella phage SI7 | Salmonella phage SI7 Salmonella virus SI7 Irrigatiovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30164 | 54.363 | Irrigatiovirus | Peduovirinae | Myoviridae | Salmonella |
| MK972711 | Salmonella phage SW9 | Salmonella phage SW9 Salmonella virus SW9 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31123 | 53.006 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| MK972710 | Salmonella phage SI22 | Salmonella phage SI22 unclassified Eganvirus Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30920 | 52.956 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| MK972709 | Salmonella phage SE20 | Salmonella phage SE20 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113437 | 40.022 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972708 | Salmonella phage SE7 | Salmonella phage SE7 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114355 | 39.869 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972707 | Salmonella phage SE24 | Salmonella phage SE24 Salmonella virus SE24 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114855 | 39.987 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972706 | Salmonella phage SS8 | Salmonella phage SS8 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41772 | 49.672 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972705 | Salmonella phage SF2 | Salmonella phage SF2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41523 | 49.693 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972704 | Salmonella phage SE19 | Salmonella phage SE19 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112879 | 39.976 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK972703 | Salmonella phage SS7 | Salmonella phage SS7 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41795 | 49.609 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972702 | Salmonella phage SS5 | Salmonella phage SS5 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41392 | 49.715 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972701 | Salmonella phage SS6 | Salmonella phage SS6 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41731 | 49.623 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972700 | Salmonella phage SS1 | Salmonella phage SS1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31600 | 49.712 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972699 | Salmonella phage SS9 | Salmonella phage SS9 Salmonella virus SS9 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156329 | 44.662 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MK972698 | Salmonella phage SS3 | Salmonella phage SS3 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158539 | 44.613 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MK972697 | Salmonella phage SF10 | Salmonella phage SF10 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40139 | 46.84 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972695 | Salmonella phage SI5 | Salmonella phage SI5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40139 | 46.83 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972694 | Salmonella phage SE22 | Salmonella phage SE22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42143 | 46.622 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972693 | Salmonella phage SI23 | Salmonella phage SI23 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42016 | 46.611 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972692 | Salmonella phage SE21 | Salmonella phage SE21 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42916 | 46.621 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972691 | Salmonella phage SI1 | Salmonella phage SI1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41396 | 49.729 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK972690 | Salmonella phage SE14 | Salmonella phage SE14 Salmonella virus SE14 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152926 | 44.839 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MK972689 | Salmonella phage SF6 | Salmonella phage SF6 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40115 | 46.838 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972688 | Salmonella phage SI8 | Salmonella phage SI8 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39988 | 46.819 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972687 | Salmonella phage SE1 (in:P22virus) | Salmonella phage SE1 (in:P22virus) Salmonella virus SE1Spa Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42001 | 46.618 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972686 | Salmonella phage SF3 | Salmonella phage SF3 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42016 | 46.613 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK972685 | Salmonella phage SE10 | Salmonella phage SE10 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42000 | 46.619 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MH299806 | Listeria phage LP-KV022 | Listeria phage LP-KV022 unclassified Homburgvirus Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65128 | 36.338 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| MK962758 | Shigella phage JK55 | Shigella phage JK55 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86219 | 38.961 | Felixounavirus | Ounavirinae | Myoviridae | Shigella |
| MK962757 | Shigella phage JK45 | Shigella phage JK45 Shigella virus JK45 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170740 | 37.639 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MK962756 | Shigella phage JK42 | Shigella phage JK42 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168306 | 37.577 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| MK962755 | Shigella phage JK38 | Shigella phage JK38 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167852 | 35.478 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MK962754 | Shigella phage JK36 | Shigella phage JK36 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168893 | 37.729 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| MK962753 | Shigella phage JK32 | Shigella phage JK32 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176009 | 40.396 | Krischvirus | Tevenvirinae | Myoviridae | Shigella |
| MK962752 | Shigella phage JK23 | Shigella phage JK23 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168349 | 35.323 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MK962751 | Shigella phage JK16 | Shigella phage JK16 Shigella virus JK16 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51854 | 44.546 | Warwickvirus | Tempevirinae | Drexlerviridae | Shigella |
| MK962750 | Shigella phage CM8 | Shigella phage CM8 Shigella virus CM8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167247 | 35.28 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MK962749 | Shigella phage CM1 | Shigella phage CM1 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139598 | 43.535 | Vequintavirus | Vequintavirinae | Myoviridae | Shigella |
| KY250034 | Pectobacterium phage PP99 | Pectobacterium phage PP99 Pectobacterium virus PP99 Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44609 | 45.545 | Gajwadongvirus | Unclassified | Autographiviridae | Pectobacterium |
| MK421971 | Klebsiella phage vB_KpnM_GF | Klebsiella phage vB_KpnM_GF unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161728 | 39.584 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN013090 | Erwinia phage vB_EamM_TropicalSun | Erwinia phage vB_EamM_TropicalSun unclassified Myosmarvirus Myosmarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68439 | 49.127 | Myosmarvirus | Unclassified | Myoviridae | Erwinia |
| MN013089 | Bacillus phage vB_BspM_MarvelLand | Bacillus phage vB_BspM_MarvelLand unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156945 | 40.193 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN013088 | Bacillus phage vB_BspS_SplendidRed | Bacillus phage vB_BspS_SplendidRed Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42859 | 44.583 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MN013087 | Klebsiella phage vB_KpnS_Penguinator | Klebsiella phage vB_KpnS_Penguinator unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51678 | 51.505 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013085 | Klebsiella phage vB_KpnP_NahiliMali | Klebsiella phage vB_KpnP_NahiliMali unclassified Ningirsuvirus Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39556 | 51.206 | Ningirsuvirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN013083 | Klebsiella phage vB_KpnS_Alina | Klebsiella phage vB_KpnS_Alina Klebsiella virus Alina Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51780 | 51.613 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013082 | Klebsiella phage vB_KpnP_Sibilus | Klebsiella phage vB_KpnP_Sibilus unclassified Ningirsuvirus Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40171 | 51.226 | Ningirsuvirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN013081 | Klebsiella phage vB_KpnM_Potts1 | Klebsiella phage vB_KpnM_Potts1 unclassified Marfavirus Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169384 | 40.701 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MN013078 | Klebsiella phage vB_KpnS_KingDDD | Klebsiella phage vB_KpnS_KingDDD unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51562 | 51.581 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013075 | Klebsiella phage vB_KpnS_Domnhall | Klebsiella phage vB_KpnS_Domnhall Klebsiella virus Domnhall Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54438 | 51.587 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MN013074 | Klebsiella phage vB_KpnP_Emp27 | Klebsiella phage vB_KpnP_Emp27 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38603 | 50.68 | Teetrevirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN101230 | Klebsiella phage KPN6 | Klebsiella phage KPN6 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164270 | 39.548 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MN101228 | Klebsiella phage KPN4 | Klebsiella phage KPN4 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108916 | 45.584 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MN101227 | Klebsiella phage KPN3 | Klebsiella phage KPN3 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38503 | 52.5 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK813943 | Aeromonas phage 4_4572 | Aeromonas phage 4_4572 Aeromonas virus 4-4572 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102915 | 42.476 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| MK813942 | Aeromonas phage 4_4512 | Aeromonas phage 4_4512 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41963 | 52.291 | Unclassified | Unclassified | Podoviridae | Aeromonas |
| MK813941 | Aeromonas phage 4L372XY | Aeromonas phage 4L372XY Aeromonas virus 4L372XY Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124279 | 35.644 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| MK813939 | Aeromonas phage 4L372D | Aeromonas phage 4L372D Aeromonas virus 4L372D Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138690 | 35.062 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| MK813938 | Aeromonas phage 2L372X | Aeromonas phage 2L372X Aeromonas virus 2L372X Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115893 | 35.558 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| MK770413 | Salmonella phage SE4 | Salmonella phage SE4 unclassified Loughboroughvirus Loughboroughvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53494 | 45.478 | Loughboroughvirus | Unclassified | Myoviridae | Salmonella |
| MK770412 | Salmonella phage SE5 | Salmonella phage SE5 unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84567 | 43.674 | Kolesnikvirus | Ounavirinae | Myoviridae | Salmonella |
| MK770410 | Salmonella phage SE3 | Salmonella phage SE3 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114437 | 39.871 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN013086 | Klebsiella phage vB_Kpn_Chronis | Klebsiella phage vB_Kpn_Chronis Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45702 | 52.252 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MN153797 | Escherichia virus T1 | Escherichia virus T1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48822 | 45.461 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MN150686 | Bacillus phage BC-T25 | Bacillus phage BC-T25 unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157977 | 40.026 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MK728541 | Escherichia phage CEC_Kaz_2018 | Escherichia phage CEC_Kaz_2018 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44283 | 54.461 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MH460829 | Acinetobacter phage vB_ApiM_fHyAci03 | Acinetobacter phage vB_ApiM_fHyAci03 Acinetobacter virus fHyAci03 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165975 | 36.758 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MK512531 | Xanthomonas phage Cf2 | Xanthomonas phage Cf2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6454 | 58.181 | Unclassified | Unclassified | Inoviridae | Xanthomonas |
| KR347355 | Propionibacterium phage QueenBey | Propionibacterium phage QueenBey unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29338 | 54.145 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337653 | Propionibacterium phage Pirate | Propionibacterium phage Pirate Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29324 | 54.085 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MH363700 | Vibrio phage VP-1 | Vibrio phage VP-1 unclassified Vapseptimavirus Vapseptimavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150764 | 41.873 | Vapseptimavirus | Unclassified | Ackermannviridae | Vibrio |
| EU770222 | Mycobacterium phage Predator | Mycobacterium phage Predator Predatorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70110 | 56.286 | Predatorvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN026741 | Salmonella phage Th1 | Salmonella phage Th1 Salmonella virus Th1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110466 | 39.208 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN103401 | Bordetella phage vB_BbrS_PHB09 | Bordetella phage vB_BbrS_PHB09 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42129 | 62.79 | Unclassified | Unclassified | Siphoviridae | Bordetella |
| MN055691 | Escherichia phage Henu8 | Escherichia phage Henu8 Escherichia virus Henu8 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49890 | 44.171 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MN028536 | Staphylococcus phage vB_SpsM_WIS42 | Staphylococcus phage vB_SpsM_WIS42 Staphylococcus virus WIS42 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40093 | 35.695 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MN022787 | Salmonella virus VSe12 | Salmonella virus VSe12 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109654 | 39.087 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MN019128 | Escherichia phage Henu7 | Escherichia phage Henu7 Escherichia virus Henu7 Henuseptimavirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49526 | 48.205 | Henuseptimavirus | Tempevirinae | Drexlerviridae | Escherichia |
| MK976036 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17223 | 36.654 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| MN163280 | Klebsiella phage KpGranit | Klebsiella phage KpGranit unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122710 | 45.216 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MN150710 | Staphylococcus phage vB_SauP-436A1 | Staphylococcus phage vB_SauP-436A1 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18028 | 29.343 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK443266 | Pseudomonas phage RSP | Pseudomonas phage RSP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63016 | 56.051 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK443264 | Pseudomonas phage ITTPL | Pseudomonas phage ITTPL unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92730 | 49.424 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK443265 | Pseudomonas phage JHP | Pseudomonas phage JHP unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48656 | 49.562 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK089780 | Acinetobacter phage vB_AbaP_APK14 | Acinetobacter phage vB_AbaP_APK14 unclassified Friunavirus Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41767 | 39.196 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MH719195 | Pseudomonas phage Ps59 | Pseudomonas phage Ps59 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39020 | 61.738 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH719194 | Pseudomonas phage Ps60 | Pseudomonas phage Ps60 unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39676 | 63.152 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MH719193 | Pseudomonas phage H72 | Pseudomonas phage H72 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38579 | 62.661 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH719192 | Pseudomonas phage Ps56 | Pseudomonas phage Ps56 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39816 | 62.621 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH719191 | Pseudomonas phage Fc22 | Pseudomonas phage Fc22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38255 | 62.747 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH719190 | Pseudomonas phage H71 | Pseudomonas phage H71 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38223 | 62.653 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH719189 | Pseudomonas phage Fc02 | Pseudomonas phage Fc02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38122 | 62.814 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH885472 | Phage PBKP05 | Phage PBKP05 unclassified Drulisvirus Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30240 | 52.983 | Drulisvirus | Slopekvirinae | Autographiviridae | Unspecified |
| MG575420 | Proteus phage vB_PmiP_RS10pmA | Proteus phage vB_PmiP_RS10pmA unclassified Gorganvirus Gorganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53343 | 35.778 | Gorganvirus | Unclassified | Siphoviridae | Proteus |
| MK509462 | Shigella phage MK-13 | Shigella phage MK-13 Shigella virus MK13 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158755 | 50.417 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| MK886800 | Escherichia phage EcNP1 | Escherichia phage EcNP1 Escherichia virus EcNP1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166176 | 35.497 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK562503 | Shigella phage Buco | Shigella phage Buco Shigella virus Buco Bucovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43630 | 54.082 | Bucovirus | Slopekvirinae | Autographiviridae | Shigella |
| MN062718 | Mycobacterium phage SoilDragon | Mycobacterium phage SoilDragon unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50293 | 64.019 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN062713 | Microbacterium phage Alyxandracam | Microbacterium phage Alyxandracam unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41770 | 63.438 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN062703 | Microbacterium phage McGalleon | Microbacterium phage McGalleon Microbacterium virus McGalleon Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42562 | 63.658 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN062702 | Microbacterium phage Den3 | Microbacterium phage Den3 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41800 | 63.371 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MN043730 | Bacillus phage vB_BveM-Goe7 | Bacillus phage vB_BveM-Goe7 unclassified Bastillevirinae Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158674 | 41.842 | Unclassified | Bastillevirinae | Herelleviridae | Bacillus |
| MN043729 | Bacillus phage vB_BmeM-Goe8 | Bacillus phage vB_BmeM-Goe8 Bacillus virus vBBmeMGoe8 Goettingenvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161583 | 38.872 | Goettingenvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN062709 | Microbacterium phage Stanktossa | Microbacterium phage Stanktossa unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41806 | 63.555 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG383452 | Escherichia phage FEC14 | Escherichia phage FEC14 Escherichia virus FEC14 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158639 | 44.616 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| MK967400 | Microbacterium Phage Clancy | Microbacterium Phage Clancy unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.432 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK967399 | Mycobacterium phage Groupthink | Mycobacterium phage Groupthink unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50574 | 64.011 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK967394 | Microbacterium phage HanSolo | Microbacterium phage HanSolo unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41516 | 63.443 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK967391 | Microbacterium phage BonesMcCoy | Microbacterium phage BonesMcCoy unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41797 | 63.361 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK967388 | Microbacterium phage Velene | Microbacterium phage Velene unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41798 | 63.403 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK967380 | Rhodococcus phage Sleepyhead | Rhodococcus phage Sleepyhead Rhodococcus virus Sleepyhead Sleepyheadvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43943 | 61.022 | Sleepyheadvirus | Unclassified | Siphoviridae | Rhodococcus |
| MH976516 | Corynebacterium phage PeteyPab | Corynebacterium phage PeteyPab unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66432 | 52.3 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MH590606 | Microbacterium phage Andromedas | Microbacterium phage Andromedas unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40494 | 62.046 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MG198780 | Corynebacterium phage Zion | Corynebacterium phage Zion Corynebacterium virus Zion Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66392 | 52.298 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MG198778 | Corynebacterium phage PotatoChip | Corynebacterium phage PotatoChip unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66359 | 52.288 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MG198777 | Corynebacterium phage Darwin | Corynebacterium phage Darwin Corynebacterium virus Darwin Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67743 | 52.082 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MK797984 | Pseudomonas phage vB_PaeM_PA5oct | Pseudomonas phage vB_PaeM_PA5oct Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 286783 | 33.329 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MN096378 | Microbacterium phage MCubed | Microbacterium phage MCubed unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40381 | 62.032 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MN096368 | Mycobacterium phage Soshari | Mycobacterium phage Soshari unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50891 | 64.202 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MN096367 | Microbacterium phage Nucci | Microbacterium phage Nucci unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40373 | 63.713 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MN128593 | Listeria phage LP-031 | Listeria phage LP-031 unclassified Homburgvirus Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65468 | 36.41 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| MN114083 | Listeria phage LP-013 | Listeria phage LP-013 unclassified Homburgvirus Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65660 | 36.346 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| MN114082 | Listeria phage LP-010 | Listeria phage LP-010 unclassified Homburgvirus Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64629 | 36.409 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| MN022785 | Escherichia virus VEc20 | Escherichia virus VEc20 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167532 | 35.462 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK804892 | Aeromonas phage 4_D05 | Aeromonas phage 4_D05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42249 | 55.987 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MK570057 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 13425 | 31.352 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| MK570056 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 14659 | 27.089 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| MK570055 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 28967 | 29.851 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| MK570054 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 27570 | 29.797 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| MK570053 | Nitrosopumilus spindle-shaped virus | Nitrosopumilus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 27548 | 29.81 | Alphafusellovirus | Unclassified | Fuselloviridae | Nitrosopumilus |
| MN090155 | Escherichia phage vB_EcoS_PHB17 | Escherichia phage vB_EcoS_PHB17 Escherichia virus PHB17 Changchunvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48939 | 46.897 | Changchunvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MK937602 | Microbacterium phage Sinatra | Microbacterium phage Sinatra unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39183 | 64.405 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MG575421 | Proteus phage vB_PmiP_RS51pmB | Proteus phage vB_PmiP_RS51pmB Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42396 | 46.983 | Unclassified | Unclassified | Myoviridae | Proteus |
| MG575419 | Proteus phage vB_PmiP_RS8pmA | Proteus phage vB_PmiP_RS8pmA unclassified Novosibovirus Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 41.353 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| MG575418 | Proteus phage vB_PmiP_RS1pmA | Proteus phage vB_PmiP_RS1pmA unclassified Novosibovirus Novosibovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42112 | 41.076 | Novosibovirus | Slopekvirinae | Autographiviridae | Proteus |
| MN010756 | Mycobacterium phage Phreeze | Mycobacterium phage Phreeze unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67906 | 57.555 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| MK962641 | Achromobacter phage vB_AxyP_19-32_Axy24 | Achromobacter phage vB_AxyP_19-32_Axy24 Achromobacter virus Axy24 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74744 | 54.918 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962640 | Achromobacter phage vB_AxyP_19-32_Axy23 | Achromobacter phage vB_AxyP_19-32_Axy23 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43773 | 64.51 | Unclassified | Krylovirinae | Autographiviridae | Achromobacter |
| MK962639 | Achromobacter phage vB_AxyP_19-32_Axy22 | Achromobacter phage vB_AxyP_19-32_Axy22 unclassified Pourcelvirus Pourcelvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71710 | 54.373 | Pourcelvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962638 | Achromobacter phage vB_AxyP_19-32_Axy21 | Achromobacter phage vB_AxyP_19-32_Axy21 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43049 | 64.324 | Unclassified | Krylovirinae | Autographiviridae | Achromobacter |
| MK962637 | Achromobacter phage vB_AxyS_19-32_Axy20 | Achromobacter phage vB_AxyS_19-32_Axy20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46352 | 56.138 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| MK962636 | Achromobacter phage vB_AxyS_19-32_Axy19 | Achromobacter phage vB_AxyS_19-32_Axy19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46036 | 56.045 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| MK962635 | Achromobacter phage vB_AxyS_19-32_Axy18 | Achromobacter phage vB_AxyS_19-32_Axy18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45500 | 56.218 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| MK962634 | Achromobacter phage vB_AxyS_19-32_Axy16 | Achromobacter phage vB_AxyS_19-32_Axy16 unclassified Steinhofvirus Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46178 | 56.206 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| MK962633 | Achromobacter phage vB_AxyS_19-32_Axy14 | Achromobacter phage vB_AxyS_19-32_Axy14 unclassified Steinhofvirus Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46703 | 54.973 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| MK962632 | Achromobacter phage vB_AxyP_19-32_Axy13 | Achromobacter phage vB_AxyP_19-32_Axy13 unclassified Inbricusvirus Inbricusvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70103 | 55.839 | Inbricusvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962631 | Achromobacter phage vB_AxyP_19-32_Axy12 | Achromobacter phage vB_AxyP_19-32_Axy12 Achromobacter virus Axy12 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74096 | 55.038 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962630 | Achromobacter phage vB_AxyP_19-32_Axy11 | Achromobacter phage vB_AxyP_19-32_Axy11 Achromobacter virus Axy11 Pourcelvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73413 | 54.324 | Pourcelvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962629 | Achromobacter phage vB_AxyP_19-32_Axy10 | Achromobacter phage vB_AxyP_19-32_Axy10 Achromobacter virus Axy10 Pourcelvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73898 | 54.269 | Pourcelvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MK962628 | Achromobacter phage vB_AxyP_19-32_Axy09 | Achromobacter phage vB_AxyP_19-32_Axy09 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43287 | 64.322 | Unclassified | Krylovirinae | Autographiviridae | Achromobacter |
| MK962627 | Achromobacter phage vB_AxyS_19-32_Axy06 | Achromobacter phage vB_AxyS_19-32_Axy06 unclassified Steinhofvirus Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45830 | 56.295 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| MK962626 | Achromobacter phage vB_AxyP_19-32_Axy04 | Achromobacter phage vB_AxyP_19-32_Axy04 Achromobacter virus Axy04 Dongdastvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73834 | 54.934 | Dongdastvirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| MN094788 | Achromobacter phage Motura | Achromobacter phage Motura Achromobacter virus Motura Moturavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221431 | 53.709 | Moturavirus | Unclassified | Myoviridae | Achromobacter |
| MK358448 | Vibrio phage 5 TSL-2019 | Vibrio phage 5 TSL-2019 unclassified Aphroditevirus Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 238053 | 43.237 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| MK672805 | Vibrio phage Va_PF430-3_p42 | Vibrio phage Va_PF430-3_p42 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51907 | 44.848 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK672804 | Vibrio phage Va_178/90_p41 | Vibrio phage Va_178/90_p41 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54138 | 43.064 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK672803 | Vibrio phage Va_91-7-154_p41 | Vibrio phage Va_91-7-154_p41 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54138 | 43.066 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK672802 | Vibrio phage Va_90-11-286_p16 | Vibrio phage Va_90-11-286_p16 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41427 | 43.979 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK672801 | Vibrio phage Va_90-11-287_p41_T265 | Vibrio phage Va_90-11-287_p41_T265 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54432 | 43.061 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK672800 | Vibrio phage Va_90-11-287_p41_Ba35 | Vibrio phage Va_90-11-287_p41_Ba35 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54432 | 43.061 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK672799 | Vibrio phage Va_90-11-287_p41 | Vibrio phage Va_90-11-287_p41 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54432 | 43.061 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK368614 | Vibrio phage 2 TSL-2019 | Vibrio phage 2 TSL-2019 Vibrio phage 2TSL2019 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 242446 | 43.571 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| MH618488 | Enterococcus phage vB_EfaS_HEf13 | Enterococcus phage vB_EfaS_HEf13 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57811 | 40.03 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MN018232 | Synechococcus phage S-B43 | Synechococcus phage S-B43 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 213993 | 39.431 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MK929790 | Caulobacter phage RW | Caulobacter phage RW Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62362 | 63.771 | Unclassified | Unclassified | Podoviridae | Caulobacter |
| MN082625 | Bacillus phage Karezi | Bacillus phage Karezi Bacillus virus Karezi Karezivirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20083 | 37.305 | Karezivirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| MN082624 | Bacillus phage VioletteMad | Bacillus phage VioletteMad unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26061 | 30.125 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MN038179 | Bacillus phage ALPS | Bacillus phage ALPS unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161842 | 38.697 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN038178 | Bacillus phage Beyonphe | Bacillus phage Beyonphe unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163540 | 37.738 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN038177 | Panteoa phage Kyle | Panteoa phage Kyle Panteoa virus Kyle Kylevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73168 | 49.498 | Kylevirus | Unclassified | Myoviridae | Pantoea |
| MN038176 | Bacillus phage Phireball | Bacillus phage Phireball unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162042 | 38.69 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MN038175 | Panteoa phage Phynn | Panteoa phage Phynn unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173720 | 44.654 | Unclassified | Tevenvirinae | Myoviridae | Pantoea |
| MN080711 | Escherichia phage vB_Eco_D226 | Escherichia phage vB_Eco_D226 unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40050 | 48.409 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| MK919475 | Mycobacterium phage Camri | Mycobacterium phage Camri unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.571 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK919472 | Mycobacterium phage Benvolio | Mycobacterium phage Benvolio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53136 | 63.735 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK905543 | Vibrio phage USC-1 | Vibrio phage USC-1 Vibrio virus USC1 Aphroditevirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 238099 | 43.243 | Aphroditevirus | Gorgonvirinae | Myoviridae | Vibrio |
| MK867835 | Salmonella phage vB_SenS_SB9 | Salmonella phage vB_SenS_SB9 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113979 | 39.911 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK838116 | Aeromonas phage LAh10 | Aeromonas phage LAh10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 260310 | 47.484 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MK838115 | Aeromonas phage LAh_9 | Aeromonas phage LAh_9 Aeromonas virus LAh9 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97988 | 42.438 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| MK838114 | Aeromonas phage LAh_8 | Aeromonas phage LAh_8 Aeromonas virus LAh8 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97408 | 42.233 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| MK838113 | Aeromonas phage LAh_7 | Aeromonas phage LAh_7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61426 | 61.972 | Unclassified | Unclassified | Siphoviridae | Aeromonas |
| MK838112 | Aeromonas phage LAh_6 | Aeromonas phage LAh_6 Aeromonas virus LAh6 Lahexavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101390 | 42.311 | Lahexavirus | Unclassified | Podoviridae | Aeromonas |
| MK838111 | Aeromonas phage LAh5 | Aeromonas phage LAh5 unclassified Ahphunavirus Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41985 | 59.274 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK838110 | Aeromonas phage LAh4 | Aeromonas phage LAh4 unclassified Ahphunavirus Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42002 | 59.271 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK838109 | Aeromonas phage LAh3 | Aeromonas phage LAh3 unclassified Ahphunavirus Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42002 | 59.276 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK838108 | Aeromonas phage LAh2 | Aeromonas phage LAh2 unclassified Ahphunavirus Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42008 | 59.27 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK838107 | Aeromonas phage LAh1 | Aeromonas phage LAh1 unclassified Ahphunavirus Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42002 | 59.273 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK798144 | Pantoea phage vB_PagM_PSKM | Pantoea phage vB_PagM_PSKM Pantoea virus PSKM Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49935 | 52.356 | Dibbivirus | Unclassified | Myoviridae | Pantoea |
| MK798143 | Pantoea phage vB_PagM_AAM37 | Pantoea phage vB_PagM_AAM37 Pantoea virus AAM37 Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49990 | 52.048 | Dibbivirus | Unclassified | Myoviridae | Pantoea |
| MK798142 | Pantoea phage vB_PagM_AAM22 | Pantoea phage vB_PagM_AAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49744 | 48.436 | Unclassified | Unclassified | Myoviridae | Pantoea |
| MK801683 | Staphylococcus phage SAP40 | Staphylococcus phage SAP40 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42911 | 34.017 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK801682 | Staphylococcus phage SAP33 | Staphylococcus phage SAP33 Staphylococcus virus SAP33 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42414 | 33.963 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK801681 | Staphylococcus phage SAP11 | Staphylococcus phage SAP11 Staphylococcus virus SAP11 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45346 | 33.419 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MK801680 | Staphylococcus phage SAP8 | Staphylococcus phage SAP8 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45533 | 33.411 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MK770415 | Salmonella phage SF11 | Salmonella phage SF11 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42016 | 46.618 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK770414 | Salmonella phage SE16 | Salmonella phage SE16 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45914 | 46.409 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MK770411 | Salmonella phage SE13 | Salmonella phage SE13 Salmonella virus SE13 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52438 | 45.78 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MK770409 | Salmonella phage SF1 | Salmonella phage SF1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41396 | 49.691 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK761199 | Salmonella phage SI2 | Salmonella phage SI2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42013 | 49.599 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK761198 | Salmonella phage SS10 | Salmonella phage SS10 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41830 | 49.598 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK761197 | Salmonella phage SS4 | Salmonella phage SS4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41396 | 49.691 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK761196 | Salmonella phage SF5 | Salmonella phage SF5 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42199 | 49.674 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK761195 | Salmonella phage SF4 | Salmonella phage SF4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43514 | 49.694 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK084630 | Bacillus phage StevenHerd11 | Bacillus phage StevenHerd11 unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23953 | 30.635 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MK795384 | Vibrio phage VAP7 | Vibrio phage VAP7 Vibrio virus VAP7 Vapseptimavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144685 | 41.864 | Vapseptimavirus | Unclassified | Ackermannviridae | Vibrio |
| MH809528 | Lactobacillus phage Sabazios | Lactobacillus phage Sabazios unclassified Coetzeevirus Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38843 | 42.664 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| MG765278 | Lactobacillus phage Silenus | Lactobacillus phage Silenus unclassified Coetzeevirus Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38716 | 42.484 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| MG765275 | Lactobacillus phage Bassarid | Lactobacillus phage Bassarid unclassified Coetzeevirus Coetzeevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37921 | 42.72 | Coetzeevirus | Unclassified | Siphoviridae | Lactobacillus |
| LR535917 | Salmonella phage SPFM20 | Salmonella phage SPFM20 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 236956 | 48.623 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535911 | Salmonella phage SPFM12 | Salmonella phage SPFM12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.621 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535908 | Salmonella phage SPFM10 | Salmonella phage SPFM10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.617 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535907 | Salmonella phage SPFM9 | Salmonella phage SPFM9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.621 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535906 | Salmonella phage SPFM8 | Salmonella phage SPFM8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.619 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535905 | Salmonella phage SPFM6 | Salmonella phage SPFM6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240198 | 48.617 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535903 | Salmonella phage SPFM5 | Salmonella phage SPFM5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240194 | 48.613 | Unclassified | Unclassified | Myoviridae | Salmonella |
| LR535901 | Salmonella phage SPFM1 | Salmonella phage SPFM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 242624 | 48.525 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MK318083 | Aeromonas phage Akh-2 | Aeromonas phage Akh-2 Shenzhenvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114901 | 40.97 | Shenzhenvirus | Unclassified | Demerecviridae | Aeromonas |
| MK977707 | Mycobacterium phage LilSpotty | Mycobacterium phage LilSpotty Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49798 | 64.693 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MK977703 | Arthrobacter phage Kumotta | Arthrobacter phage Kumotta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40317 | 60.763 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MK977700 | Corynebacterium phage Stickynote | Corynebacterium phage Stickynote unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67086 | 52.176 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MK977697 | Microbacterium phage Etta | Microbacterium phage Etta unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41542 | 63.331 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK894437 | Microbacterium phage MonChoix | Microbacterium phage MonChoix Microbacterium virus MonChoix Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41670 | 63.365 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK894432 | Microbacterium phage Finny | Microbacterium phage Finny unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40313 | 62.144 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MN013356 | Pseudomonas phage vB_PaM_EPA1 | Pseudomonas phage vB_PaM_EPA1 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91394 | 49.27 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK934841 | Pseudomonas phage UMP151 | Pseudomonas phage UMP151 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41303 | 62.816 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK907780 | Vibrio phage Pontus | Vibrio phage Pontus unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128289 | 40.233 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MK895508 | Vibrio phage Brizo | Vibrio phage Brizo unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120106 | 40.128 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MK893987 | Staphylococcus phage PMBT8 | Staphylococcus phage PMBT8 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88129 | 31.617 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MK882925 | Dinoroseobacter phage vB_DshS-R4C | Dinoroseobacter phage vB_DshS-R4C Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36291 | 66.755 | Cronusvirus | Unclassified | Siphoviridae | Dinoroseobacter |
| MK817115 | Escherichia phage vB_EcoM_phAPEC6 | Escherichia phage vB_EcoM_phAPEC6 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 352598 | 34.111 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| NC_043427 | Aeropyrum coil-shaped virus | Aeropyrum coil-shaped virus Alphaspiravirus Spiraviridae Viruses | 24893 | 46.668 | Alphaspiravirus | Unclassified | Spiraviridae | Aeropyrum |
| NC_043029 | Stenotrophomonas phage SMA6 | Stenotrophomonas phage SMA6 Scuticavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7648 | 62.631 | Scuticavirus | Unclassified | Inoviridae | Stenotrophomonas |
| NC_043028 | Xanthomonas phage Xf109 | Xanthomonas phage Xf109 Xanthomonas virus Xf109 Xylivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7190 | 59.597 | Xylivirus | Unclassified | Inoviridae | Xanthomonas |
| NC_007823 | Escherichia phage NC28 | Escherichia phage NC28 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6065 | 44.831 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007827 | Escherichia phage NC29 | Escherichia phage NC29 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6259 | 44.352 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007825 | Escherichia phage ID52 | Escherichia phage ID52 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5538 | 45.865 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| NC_007824 | Escherichia phage ID62 | Escherichia phage ID62 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6051 | 44.935 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007822 | Escherichia phage WA45 | Escherichia phage WA45 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 44.941 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007820 | Escherichia phage NC35 | Escherichia phage NC35 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6039 | 44.726 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007819 | Escherichia phage ID32 | Escherichia phage ID32 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6071 | 45.034 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_007818 | Escherichia phage ID21 | Escherichia phage ID21 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 45.27 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| LR596903 | Roseburia phage Shimadzu | Roseburia phage Shimadzu Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44991 | 52.068 | Unclassified | Unclassified | Myoviridae | Roseburia |
| LR596902 | Roseburia phage Jekyll | Roseburia phage Jekyll unclassified bacterial viruses Viruses | 45629 | 40.906 | Unclassified | Unclassified | Unclassified | Roseburia |
| MN067333 | Escherichia phage Lyz12581Vzw | Escherichia phage Lyz12581Vzw Eschericia virus Lyz12581Vzw Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62668 | 50.137 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| MK977694 | Escherichia phage vB_EcoM-Sa45lw | Escherichia phage vB_EcoM-Sa45lw unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167353 | 35.35 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK112901 | Salmonella virus KFS-SE2 | Salmonella virus KFS-SE2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48608 | 45.801 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH598805 | Pelagibacter phage HTVC109P | Pelagibacter phage HTVC109P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41323 | 35.491 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598798 | Pelagibacter phage HTVC022P | Pelagibacter phage HTVC022P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42010 | 34.161 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598799 | Pelagibacter phage HTVC025P | Pelagibacter phage HTVC025P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37251 | 32.466 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MH598800 | Pelagibacter phage HTVC031P | Pelagibacter phage HTVC031P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41046 | 33.406 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598801 | Pelagibacter phage HTVC200P | Pelagibacter phage HTVC200P unclassified Votkovvirus Votkovvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42221 | 33.156 | Votkovvirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598802 | Pelagibacter phage HTVC201P | Pelagibacter phage HTVC201P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41415 | 33.092 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598803 | Pelagibacter phage HTVC121P | Pelagibacter phage HTVC121P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42600 | 33.521 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598804 | Pelagibacter phage HTVC105P | Pelagibacter phage HTVC105P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42385 | 33.531 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598806 | Pelagibacter phage HTVC119P | Pelagibacter phage HTVC119P unclassified Votkovvirus Votkovvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38357 | 32.007 | Votkovvirus | Unclassified | Autographiviridae | Pelagibacter |
| MH598807 | Pelagibacter phage HTVC120P | Pelagibacter phage HTVC120P unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42622 | 33.279 | Unclassified | Unclassified | Autographiviridae | Pelagibacter |
| MH590595 | Mycobacterium phage NEHalo | Mycobacterium phage NEHalo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50535 | 63.819 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH579717 | Pelagibacter phage HTVC021P | Pelagibacter phage HTVC021P unclassified Pelagivirus Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42809 | 33.551 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| MH576975 | Mycobacterium phage Target | Mycobacterium phage Target unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49097 | 63.597 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG812495 | Mycobacterium phage Pippin | Mycobacterium phage Pippin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52034 | 63.753 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG812489 | Mycobacterium phage Greg | Mycobacterium phage Greg unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51083 | 63.412 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| LR596615 | Yersinia phage fHe-Yen9-04 | Yersinia phage fHe-Yen9-04 Yersinia virus Yen9-04 Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 354378 | 31.646 | Eneladusvirus | Unclassified | Myoviridae | Yersinia |
| LR596614 | Escherichia phage vB_Eco_SLUR29 | Escherichia phage vB_Eco_SLUR29 Escherichia virus SLUR29 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48466 | 44.739 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LC487997 | Stx2-converting phage Stx2a F578 | Stx2-converting phage Stx2a F578 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24274 | 47.001 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LC487996 | Stx2-converting phage Stx2a 981509 | Stx2-converting phage Stx2a 981509 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25488 | 46.379 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MK804893 | Aeromonas phage 2L372D | Aeromonas phage 2L372D Aeromonas virus 2L372D Plateaulakevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122963 | 35.294 | Plateaulakevirus | Unclassified | Myoviridae | Aeromonas |
| MK804891 | Aeromonas phage 2_D05 | Aeromonas phage 2_D05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43233 | 56.36 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MK883718 | Escherichia phage vB_EcoM-ECP32 | Escherichia phage vB_EcoM-ECP32 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133317 | 43.687 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MK883717 | Escherichia phage vB_EcoM-ECP26 | Escherichia phage vB_EcoM-ECP26 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136993 | 43.585 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MK795385 | Klebsiella phage IME304 | Klebsiella phage IME304 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40746 | 52.393 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MN027503 | Enterococcus phage vB_OCPT_Ben | Enterococcus phage vB_OCPT_Ben unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151985 | 37.089 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MN013189 | Microcystis phage Mic1 | Microcystis phage Mic1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92627 | 35.17 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MK984681 | Bordetella phage vB_BbrP_BB8 | Bordetella phage vB_BbrP_BB8 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41593 | 58.806 | Unclassified | Unclassified | Autographiviridae | Bordetella |
| MK816297 | Pseudomonas virus PBPA162 | Pseudomonas virus PBPA162 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60626 | 56.388 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK809530 | Salmonella phage vB_SenS_SB6 | Salmonella phage vB_SenS_SB6 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112311 | 39.944 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK803322 | Pseudomonas phage RSP | Pseudomonas phage RSP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63016 | 56.051 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK800154 | Enterococcus phage Entf1 | Enterococcus phage Entf1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58938 | 39.93 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MK792752 | Listeria phage vB-LmoM-SH3-3 | Listeria phage vB-LmoM-SH3-3 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136801 | 36.044 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| MH809531 | Lactobacillus phage Bromius | Lactobacillus phage Bromius Lactobacillus virus Bromius Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140527 | 36.525 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MH809530 | Lactobacillus phage Dionysus | Lactobacillus phage Dionysus unclassified Harbinvirus Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141344 | 36.64 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MH809529 | Lactobacillus phage Iacchus | Lactobacillus phage Iacchus Lactobacillus virus Iacchus Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137973 | 36.505 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MG765279 | Lactobacillus phage Semele | Lactobacillus phage Semele Lactobacillus virus Semele Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139450 | 36.331 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MG765277 | Lactobacillus phage Bacchae | Lactobacillus phage Bacchae Lactobacillus virus Bacchae Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141124 | 36.346 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MG944235 | Xanthomonas phage XPV2 | Xanthomonas phage XPV2 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45969 | 61.189 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944232 | Xanthomonas phage XPP8 | Xanthomonas phage XPP8 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46278 | 61.038 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944230 | Xanthomonas phage XPP4 | Xanthomonas phage XPP4 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47397 | 61.196 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944227 | Xanthomonas phage XPP1 | Xanthomonas phage XPP1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46195 | 60.989 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MH443101 | Erwinia phage vB_EhrS_59 | Erwinia phage vB_EhrS_59 Erwinia virus Eho59 Feofaniavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47116 | 50.405 | Feofaniavirus | Unclassified | Siphoviridae | Erwinia |
| MH443100 | Erwinia phage vB_EhrS_49 | Erwinia phage vB_EhrS_49 Erwinia virus Eho49 Feofaniavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46835 | 50.844 | Feofaniavirus | Unclassified | Siphoviridae | Erwinia |
| KY006473 | Mycobacterium phage LeoAvram | Mycobacterium phage LeoAvram unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47079 | 63.827 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG852086 | Escherichia phage vB_EcoS-Ro145clw | Escherichia phage vB_EcoS-Ro145clw unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42031 | 50.575 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK504446 | Lactobacillus phage SAC12B | Lactobacillus phage SAC12B Lactobacillus virus SAC12B Tybeckvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136608 | 32.411 | Tybeckvirus | Unclassified | Herelleviridae | Lactobacillus |
| MK504445 | Lactobacillus phage ATCCB | Lactobacillus phage ATCCB Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80538 | 30.799 | Unclassified | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MK504444 | Lactobacillus phage 3-521 | Lactobacillus phage 3-521 Lactobacillus virus 3-521 Watanabevirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140816 | 36.933 | Watanabevirus | Unclassified | Herelleviridae | Lactobacillus |
| MK504443 | Lactobacillus phage 521B | Lactobacillus phage 521B Lactobacillus virus 521B Tybeckvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136442 | 32.268 | Tybeckvirus | Unclassified | Herelleviridae | Lactobacillus |
| MK504442 | Lactobacillus phage 3-SAC12 | Lactobacillus phage 3-SAC12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41292 | 40.008 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| MK215646 | Bacillus phage vB_BthM-Goe5 | Bacillus phage vB_BthM-Goe5 unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157804 | 37.948 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH203383 | Enterococcus phage vB_EfaS_AL3 | Enterococcus phage vB_EfaS_AL3 Enterococcus virus AL3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40789 | 34.838 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MG747435 | Ralstonia phage RsoM1USA | Ralstonia phage RsoM1USA Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39309 | 65.153 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| MK875794 | Microbacterium phage Chepli | Microbacterium phage Chepli unclassified Neferthenavirus Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40332 | 62.99 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MK770119 | Pantoea phage vB_PagS_AAS21 | Pantoea phage vB_PagS_AAS21 unclassified Demerecviridae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116649 | 39.007 | Unclassified | Unclassified | Demerecviridae | Pantoea |
| MK942064 | Caulobacter phage Jess A | Caulobacter phage Jess A unclassified Rauchvirus Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45011 | 64.329 | Rauchvirus | Unclassified | Podoviridae | Caulobacter |
| MK843319 | Bacillus phage vB_BthS-TP21T | Bacillus phage vB_BthS-TP21T unclassified Waukeshavirus Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51949 | 35.496 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| MK965970 | Salmonella phage DaR-2019b | Salmonella phage DaR-2019b unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88966 | 39.108 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MK965969 | Salmonella phage DaR-2019a | Salmonella phage DaR-2019a unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89091 | 39.062 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MK773491 | Salmonella phage 7t3 | Salmonella phage 7t3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348718 | 35.584 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MK778457 | Escherichia phage vB_EcoS_W011D | Escherichia phage vB_EcoS_W011D Escherichia virus W011D Changchunvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49847 | 46.241 | Changchunvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MK779875 | Lactococcus phage CHPC971 | Lactococcus phage CHPC971 Lactococcus virus CHPC971 Fremauxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54381 | 34.062 | Fremauxvirus | Unclassified | Siphoviridae | Lactococcus |
| MK496826 | Capybara microvirus Cap3_SP_668 | Capybara microvirus Cap3_SP_668 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4148 | 38.163 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496825 | Capybara microvirus Cap3_SP_649 | Capybara microvirus Cap3_SP_649 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4289 | 40.196 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496824 | Capybara microvirus Cap3_SP_646 | Capybara microvirus Cap3_SP_646 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4304 | 37.198 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496823 | Capybara microvirus Cap3_SP_645 | Capybara microvirus Cap3_SP_645 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4307 | 40.098 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496822 | Capybara microvirus Cap3_SP_641 | Capybara microvirus Cap3_SP_641 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4323 | 42.285 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496821 | Capybara microvirus Cap3_SP_632 | Capybara microvirus Cap3_SP_632 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4381 | 32.413 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496820 | Capybara microvirus Cap3_SP_613 | Capybara microvirus Cap3_SP_613 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4485 | 27.625 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496819 | Capybara microvirus Cap3_SP_612 | Capybara microvirus Cap3_SP_612 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4492 | 38.624 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496818 | Capybara microvirus Cap3_SP_603 | Capybara microvirus Cap3_SP_603 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4526 | 47.437 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496817 | Capybara microvirus Cap3_SP_588 | Capybara microvirus Cap3_SP_588 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4588 | 34.895 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496816 | Capybara microvirus Cap3_SP_586 | Capybara microvirus Cap3_SP_586 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4591 | 35.57 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496815 | Capybara microvirus Cap3_SP_581 | Capybara microvirus Cap3_SP_581 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4602 | 34.811 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496814 | Capybara microvirus Cap3_SP_578 | Capybara microvirus Cap3_SP_578 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4609 | 34.216 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496813 | Capybara microvirus Cap3_SP_562 | Capybara microvirus Cap3_SP_562 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4684 | 31.298 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496812 | Capybara microvirus Cap3_SP_554 | Capybara microvirus Cap3_SP_554 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4723 | 30.002 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496811 | Capybara microvirus Cap3_SP_550 | Capybara microvirus Cap3_SP_550 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4736 | 32.517 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496810 | Capybara microvirus Cap3_SP_546 | Capybara microvirus Cap3_SP_546 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4750 | 35.789 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496809 | Capybara microvirus Cap3_SP_541 | Capybara microvirus Cap3_SP_541 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4765 | 41.595 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496808 | Capybara microvirus Cap3_SP_539 | Capybara microvirus Cap3_SP_539 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4776 | 35.448 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496807 | Capybara microvirus Cap3_SP_535 | Capybara microvirus Cap3_SP_535 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4780 | 33.724 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496806 | Capybara microvirus Cap3_SP_481 | Capybara microvirus Cap3_SP_481 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5060 | 37.945 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496805 | Capybara microvirus Cap3_SP_478 | Capybara microvirus Cap3_SP_478 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5076 | 31.738 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496804 | Capybara microvirus Cap3_SP_475 | Capybara microvirus Cap3_SP_475 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5089 | 29.122 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496803 | Capybara microvirus Cap3_SP_472 | Capybara microvirus Cap3_SP_472 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5108 | 37.294 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496802 | Capybara microvirus Cap3_SP_470 | Capybara microvirus Cap3_SP_470 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5111 | 36.157 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496801 | Capybara microvirus Cap3_SP_469 | Capybara microvirus Cap3_SP_469 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5117 | 30.877 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496800 | Capybara microvirus Cap3_SP_468 | Capybara microvirus Cap3_SP_468 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5119 | 33.737 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496799 | Capybara microvirus Cap3_SP_465 | Capybara microvirus Cap3_SP_465 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5129 | 33.203 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496798 | Capybara microvirus Cap3_SP_457 | Capybara microvirus Cap3_SP_457 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5171 | 41.385 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496797 | Capybara microvirus Cap3_SP_450 | Capybara microvirus Cap3_SP_450 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5204 | 36.222 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496796 | Capybara microvirus Cap3_SP_449 | Capybara microvirus Cap3_SP_449 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5206 | 46.907 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496795 | Capybara microvirus Cap3_SP_446 | Capybara microvirus Cap3_SP_446 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5225 | 36.804 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496794 | Capybara microvirus Cap3_SP_445 | Capybara microvirus Cap3_SP_445 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5232 | 31.479 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496793 | Capybara microvirus Cap3_SP_444 | Capybara microvirus Cap3_SP_444 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5235 | 31.251 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496792 | Capybara microvirus Cap3_SP_443 | Capybara microvirus Cap3_SP_443 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5247 | 46.388 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496791 | Capybara microvirus Cap3_SP_442 | Capybara microvirus Cap3_SP_442 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5250 | 37.943 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496790 | Capybara microvirus Cap3_SP_441 | Capybara microvirus Cap3_SP_441 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5251 | 36.812 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496789 | Capybara microvirus Cap3_SP_437 | Capybara microvirus Cap3_SP_437 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5271 | 36.691 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496788 | Capybara microvirus Cap3_SP_433 | Capybara microvirus Cap3_SP_433 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5278 | 37.817 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496787 | Capybara microvirus Cap3_SP_431 | Capybara microvirus Cap3_SP_431 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5291 | 39.18 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496786 | Capybara microvirus Cap3_SP_423 | Capybara microvirus Cap3_SP_423 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5319 | 37.319 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496785 | Capybara microvirus Cap3_SP_416 | Capybara microvirus Cap3_SP_416 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5375 | 38.958 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496784 | Capybara microvirus Cap3_SP_414 | Capybara microvirus Cap3_SP_414 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5380 | 27.9 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496783 | Capybara microvirus Cap3_SP_410 | Capybara microvirus Cap3_SP_410 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5402 | 35.172 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496782 | Capybara microvirus Cap3_SP_407 | Capybara microvirus Cap3_SP_407 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5415 | 42.124 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496781 | Capybara microvirus Cap3_SP_394 | Capybara microvirus Cap3_SP_394 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5518 | 47.644 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496780 | Capybara microvirus Cap3_SP_393 | Capybara microvirus Cap3_SP_393 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5529 | 29.083 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496779 | Capybara microvirus Cap3_SP_391 | Capybara microvirus Cap3_SP_391 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5540 | 51.101 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496778 | Capybara microvirus Cap3_SP_389 | Capybara microvirus Cap3_SP_389 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5547 | 41.013 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496777 | Capybara microvirus Cap3_SP_386 | Capybara microvirus Cap3_SP_386 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5559 | 36.212 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496776 | Capybara microvirus Cap3_SP_383 | Capybara microvirus Cap3_SP_383 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5576 | 36.209 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496775 | Capybara microvirus Cap3_SP_379 | Capybara microvirus Cap3_SP_379 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5595 | 36.05 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496774 | Capybara microvirus Cap3_SP_374 | Capybara microvirus Cap3_SP_374 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5630 | 42.735 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496773 | Capybara microvirus Cap3_SP_367 | Capybara microvirus Cap3_SP_367 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 31.85 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496772 | Capybara microvirus Cap3_SP_363 | Capybara microvirus Cap3_SP_363 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5694 | 32.332 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496771 | Capybara microvirus Cap3_SP_352 | Capybara microvirus Cap3_SP_352 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5809 | 32.467 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496770 | Capybara microvirus Cap3_SP_347 | Capybara microvirus Cap3_SP_347 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5869 | 33.157 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496769 | Capybara microvirus Cap3_SP_344 | Capybara microvirus Cap3_SP_344 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5891 | 41.911 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496768 | Capybara microvirus Cap3_SP_333 | Capybara microvirus Cap3_SP_333 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5957 | 42.437 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496767 | Capybara microvirus Cap3_SP_332 | Capybara microvirus Cap3_SP_332 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5963 | 35.536 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496766 | Capybara microvirus Cap3_SP_330 | Capybara microvirus Cap3_SP_330 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6002 | 39.354 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496765 | Capybara microvirus Cap3_SP_320 | Capybara microvirus Cap3_SP_320 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6097 | 39.036 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496764 | Capybara microvirus Cap3_SP_319 | Capybara microvirus Cap3_SP_319 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6107 | 35.32 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496763 | Capybara microvirus Cap3_SP_316 | Capybara microvirus Cap3_SP_316 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6140 | 29.951 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496762 | Capybara microvirus Cap3_SP_315 | Capybara microvirus Cap3_SP_315 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6146 | 30.996 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496761 | Capybara microvirus Cap3_SP_297 | Capybara microvirus Cap3_SP_297 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6394 | 35.127 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496760 | Capybara microvirus Cap3_SP_293 | Capybara microvirus Cap3_SP_293 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6415 | 35.261 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496759 | Capybara microvirus Cap3_SP_264 | Capybara microvirus Cap3_SP_264 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6887 | 36.141 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496758 | Capybara microvirus Cap3_SP_188 | Capybara microvirus Cap3_SP_188 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4709 | 36.526 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496757 | Capybara microvirus Cap1_SP_263 | Capybara microvirus Cap1_SP_263 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4161 | 31.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496756 | Capybara microvirus Cap1_SP_259 | Capybara microvirus Cap1_SP_259 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4203 | 38.829 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496755 | Capybara microvirus Cap1_SP_244 | Capybara microvirus Cap1_SP_244 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4404 | 29.882 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496754 | Capybara microvirus Cap1_SP_240 | Capybara microvirus Cap1_SP_240 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4482 | 39.201 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496753 | Capybara microvirus Cap1_SP_230 | Capybara microvirus Cap1_SP_230 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4530 | 44.79 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496752 | Capybara microvirus Cap1_SP_228 | Capybara microvirus Cap1_SP_228 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4558 | 50.351 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496751 | Capybara microvirus Cap1_SP_223 | Capybara microvirus Cap1_SP_223 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4590 | 37.429 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496750 | Capybara microvirus Cap1_SP_222 | Capybara microvirus Cap1_SP_222 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4592 | 35.475 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496749 | Capybara microvirus Cap1_SP_217 | Capybara microvirus Cap1_SP_217 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4610 | 47.093 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496748 | Capybara microvirus Cap1_SP_216 | Capybara microvirus Cap1_SP_216 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4611 | 34.266 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496747 | Capybara microvirus Cap1_SP_209 | Capybara microvirus Cap1_SP_209 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4635 | 51.241 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496746 | Capybara microvirus Cap1_SP_206 | Capybara microvirus Cap1_SP_206 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4672 | 41.567 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496745 | Capybara microvirus Cap1_SP_200 | Capybara microvirus Cap1_SP_200 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4718 | 35.481 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496744 | Capybara microvirus Cap1_SP_192 | Capybara microvirus Cap1_SP_192 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4757 | 41.097 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496743 | Capybara microvirus Cap1_SP_188 | Capybara microvirus Cap1_SP_188 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4785 | 36.155 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496742 | Capybara microvirus Cap1_SP_178 | Capybara microvirus Cap1_SP_178 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4852 | 45.899 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496741 | Capybara microvirus Cap1_SP_175 | Capybara microvirus Cap1_SP_175 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4866 | 37.217 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496740 | Capybara microvirus Cap1_SP_168 | Capybara microvirus Cap1_SP_168 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4905 | 41.651 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496739 | Capybara microvirus Cap1_SP_167 | Capybara microvirus Cap1_SP_167 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4915 | 46.307 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496738 | Capybara microvirus Cap1_SP_166 | Capybara microvirus Cap1_SP_166 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4936 | 40.154 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496737 | Capybara microvirus Cap1_SP_164 | Capybara microvirus Cap1_SP_164 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4939 | 40.454 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496736 | Capybara microvirus Cap1_SP_163 | Capybara microvirus Cap1_SP_163 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4955 | 43.35 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496735 | Capybara microvirus Cap1_SP_162 | Capybara microvirus Cap1_SP_162 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4956 | 38.64 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496734 | Capybara microvirus Cap1_SP_161 | Capybara microvirus Cap1_SP_161 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4979 | 31.914 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496733 | Capybara microvirus Cap1_SP_160 | Capybara microvirus Cap1_SP_160 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4989 | 47.444 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496732 | Capybara microvirus Cap1_SP_159 | Capybara microvirus Cap1_SP_159 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4996 | 45.176 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496731 | Capybara microvirus Cap1_SP_158 | Capybara microvirus Cap1_SP_158 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5001 | 39.832 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496730 | Capybara microvirus Cap1_SP_151 | Capybara microvirus Cap1_SP_151 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5059 | 32.338 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496729 | Capybara microvirus Cap1_SP_148 | Capybara microvirus Cap1_SP_148 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5067 | 41.78 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496728 | Capybara microvirus Cap1_SP_147 | Capybara microvirus Cap1_SP_147 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5074 | 33.859 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496727 | Capybara microvirus Cap1_SP_145 | Capybara microvirus Cap1_SP_145 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5085 | 35.398 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496726 | Capybara microvirus Cap1_SP_143 | Capybara microvirus Cap1_SP_143 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5123 | 37.244 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496725 | Capybara microvirus Cap1_SP_142 | Capybara microvirus Cap1_SP_142 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5124 | 43.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496724 | Capybara microvirus Cap1_SP_141 | Capybara microvirus Cap1_SP_141 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5132 | 35.405 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496723 | Capybara microvirus Cap1_SP_140 | Capybara microvirus Cap1_SP_140 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5134 | 37.768 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496722 | Capybara microvirus Cap1_SP_137 | Capybara microvirus Cap1_SP_137 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5188 | 50.328 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496721 | Capybara microvirus Cap1_SP_135 | Capybara microvirus Cap1_SP_135 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5207 | 37.277 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496720 | Capybara microvirus Cap1_SP_133 | Capybara microvirus Cap1_SP_133 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5221 | 47.175 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496719 | Capybara microvirus Cap1_SP_131 | Capybara microvirus Cap1_SP_131 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5243 | 40.454 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496718 | Capybara microvirus Cap1_SP_128 | Capybara microvirus Cap1_SP_128 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4514 | 35.755 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496717 | Capybara microvirus Cap1_SP_127 | Capybara microvirus Cap1_SP_127 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5267 | 36.434 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496716 | Capybara microvirus Cap1_SP_124 | Capybara microvirus Cap1_SP_124 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5278 | 36.984 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496715 | Capybara microvirus Cap1_SP_123 | Capybara microvirus Cap1_SP_123 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5291 | 36.874 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496714 | Capybara microvirus Cap1_SP_121 | Capybara microvirus Cap1_SP_121 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5314 | 44.242 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496713 | Capybara microvirus Cap1_SP_119 | Capybara microvirus Cap1_SP_119 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5316 | 30.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496712 | Capybara microvirus Cap1_SP_118 | Capybara microvirus Cap1_SP_118 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5318 | 38.247 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496711 | Capybara microvirus Cap1_SP_116 | Capybara microvirus Cap1_SP_116 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5364 | 43.568 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496710 | Capybara microvirus Cap1_SP_115 | Capybara microvirus Cap1_SP_115 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5375 | 35.907 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496709 | Capybara microvirus Cap1_SP_109 | Capybara microvirus Cap1_SP_109 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5489 | 31.08 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496708 | Capybara microvirus Cap1_SP_108 | Capybara microvirus Cap1_SP_108 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5490 | 38.288 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496707 | Capybara microvirus Cap1_SP_107 | Capybara microvirus Cap1_SP_107 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5508 | 36.71 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496706 | Capybara microvirus Cap1_SP_106 | Capybara microvirus Cap1_SP_106 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5508 | 35.712 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496705 | Capybara microvirus Cap1_SP_104 | Capybara microvirus Cap1_SP_104 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5562 | 40.399 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496704 | Capybara microvirus Cap1_SP_99 | Capybara microvirus Cap1_SP_99 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5610 | 37.023 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496703 | Capybara microvirus Cap1_SP_95 | Capybara microvirus Cap1_SP_95 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5647 | 38.675 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496702 | Capybara microvirus Cap1_SP_94 | Capybara microvirus Cap1_SP_94 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5663 | 37.118 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496701 | Capybara microvirus Cap1_SP_93 | Capybara microvirus Cap1_SP_93 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5675 | 34.89 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496700 | Capybara microvirus Cap1_SP_92 | Capybara microvirus Cap1_SP_92 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5680 | 30.775 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496699 | Capybara microvirus Cap1_SP_90 | Capybara microvirus Cap1_SP_90 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5707 | 34.116 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496698 | Capybara microvirus Cap1_SP_87 | Capybara microvirus Cap1_SP_87 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5735 | 33.88 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496697 | Capybara microvirus Cap1_SP_84 | Capybara microvirus Cap1_SP_84 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5767 | 44.044 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496696 | Capybara microvirus Cap1_SP_83 | Capybara microvirus Cap1_SP_83 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5779 | 39.644 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496695 | Capybara microvirus Cap1_SP_82 | Capybara microvirus Cap1_SP_82 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5785 | 32.031 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496694 | Capybara microvirus Cap1_SP_81 | Capybara microvirus Cap1_SP_81 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5791 | 40.442 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496693 | Capybara microvirus Cap1_SP_77 | Capybara microvirus Cap1_SP_77 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5865 | 39.267 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496692 | Capybara microvirus Cap1_SP_76 | Capybara microvirus Cap1_SP_76 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5869 | 46.584 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496691 | Capybara microvirus Cap1_SP_70 | Capybara microvirus Cap1_SP_70 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5986 | 33.512 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496690 | Capybara microvirus Cap1_SP_66 | Capybara microvirus Cap1_SP_66 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6036 | 39.761 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496689 | Capybara microvirus Cap1_SP_64 | Capybara microvirus Cap1_SP_64 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6096 | 30.184 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496688 | Capybara microvirus Cap1_SP_61 | Capybara microvirus Cap1_SP_61 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6173 | 40.094 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496687 | Capybara microvirus Cap1_SP_60 | Capybara microvirus Cap1_SP_60 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6242 | 37.536 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496686 | Capybara microvirus Cap1_SP_59 | Capybara microvirus Cap1_SP_59 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6209 | 40.039 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496685 | Capybara microvirus Cap1_SP_55 | Capybara microvirus Cap1_SP_55 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6272 | 36.878 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496684 | Capybara microvirus Cap1_SP_52 | Capybara microvirus Cap1_SP_52 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6356 | 35.746 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496683 | Capybara microvirus Cap1_SP_51 | Capybara microvirus Cap1_SP_51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6391 | 34.768 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496682 | Capybara microvirus Cap1_SP_50 | Capybara microvirus Cap1_SP_50 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6391 | 38.945 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496681 | Capybara microvirus Cap1_SP_48 | Capybara microvirus Cap1_SP_48 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6415 | 35.635 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496680 | Capybara microvirus Cap1_SP_45 | Capybara microvirus Cap1_SP_45 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5585 | 33.321 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK496679 | Capybara microvirus Cap1_SP_22 | Capybara microvirus Cap1_SP_22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5881 | 35.436 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_011399 | Ralstonia phage RSM3 | Ralstonia phage RSM3 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8929 | 59.648 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| NC_007461 | Chlamydia phage 4 | Chlamydia phage 4 unclassified Chlamydiamicrovirus Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4530 | 41.17 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| JQ340774 | Enterobacter phage Enc34 | Enterobacter phage Enc34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60496 | 51.084 | Unclassified | Unclassified | Siphoviridae | Enterobacter |
| MH444512 | Bifidobacterium phage PMBT6 | Bifidobacterium phage PMBT6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36607 | 61.644 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| MG944226 | Microbacterium phage Casey | Microbacterium phage Casey unclassified Pikminvirus Pikminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39307 | 61.279 | Pikminvirus | Unclassified | Siphoviridae | Microbacterium |
| MG944218 | Microbacterium phage Pikmin | Microbacterium phage Pikmin Microbacterium virus Pikmin Pikminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39307 | 61.269 | Pikminvirus | Unclassified | Siphoviridae | Microbacterium |
| MG944216 | Microbacterium phage Pajaza | Microbacterium phage Pajaza unclassified Pikminvirus Pikminvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39307 | 61.244 | Pikminvirus | Unclassified | Siphoviridae | Microbacterium |
| LR535921 | Salmonella phage SPFM2 | Salmonella phage SPFM2 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240111 | 48.615 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535920 | Salmonella phage SPFM3 | Salmonella phage SPFM3 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240198 | 48.619 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535919 | Salmonella phage SPFM21 | Salmonella phage SPFM21 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240196 | 48.621 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535918 | Salmonella phage SPFM22 | Salmonella phage SPFM22 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240196 | 48.623 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535916 | Salmonella phage SPFM19 | Salmonella phage SPFM19 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.621 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535915 | Salmonella phage SPFM16 | Salmonella phage SPFM16 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 233195 | 48.629 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535914 | Salmonella phage SPFM17 | Salmonella phage SPFM17 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239842 | 48.64 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535910 | Salmonella phage SPFM13 | Salmonella phage SPFM13 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241405 | 48.573 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535909 | Salmonella phage SPFM11 | Salmonella phage SPFM11 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.62 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| LR535902 | Salmonella phage SPFM4 | Salmonella phage SPFM4 unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240197 | 48.618 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| MK765669 | Tortoise microvirus 111 | Tortoise microvirus 111 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4383 | 47.798 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765668 | Tortoise microvirus 8 | Tortoise microvirus 8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.708 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765667 | Tortoise microvirus 110 | Tortoise microvirus 110 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5554 | 43.86 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765666 | Tortoise microvirus 109 | Tortoise microvirus 109 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5823 | 41.456 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765665 | Tortoise microvirus 108 | Tortoise microvirus 108 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5837 | 42.436 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765664 | Tortoise microvirus 107 | Tortoise microvirus 107 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5998 | 42.181 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765663 | Tortoise microvirus 8 | Tortoise microvirus 8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.981 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765662 | Tortoise microvirus 106 | Tortoise microvirus 106 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4845 | 40.888 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765661 | Tortoise microvirus 105 | Tortoise microvirus 105 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5554 | 44.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765660 | Tortoise microvirus 104 | Tortoise microvirus 104 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5663 | 40.102 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765659 | Tortoise microvirus 103 | Tortoise microvirus 103 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5671 | 37.524 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765658 | Tortoise microvirus 3 | Tortoise microvirus 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5823 | 41.594 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765657 | Tortoise microvirus 2 | Tortoise microvirus 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5975 | 42.226 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765656 | Tortoise microvirus 8 | Tortoise microvirus 8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.708 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765655 | Tortoise microvirus 102 | Tortoise microvirus 102 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5554 | 43.86 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765654 | Tortoise microvirus 101 | Tortoise microvirus 101 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.685 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765653 | Tortoise microvirus 100 | Tortoise microvirus 100 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4597 | 35.784 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765652 | Tortoise microvirus 99 | Tortoise microvirus 99 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5788 | 41.137 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765651 | Tortoise microvirus 98 | Tortoise microvirus 98 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5816 | 41.506 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765650 | Tortoise microvirus 97 | Tortoise microvirus 97 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6243 | 37.434 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765649 | Tortoise microvirus 96 | Tortoise microvirus 96 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5122 | 44.299 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765648 | Tortoise microvirus 95 | Tortoise microvirus 95 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5781 | 41.757 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765647 | Tortoise microvirus 94 | Tortoise microvirus 94 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5820 | 42.629 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765646 | Tortoise microvirus 93 | Tortoise microvirus 93 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4242 | 50.094 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765645 | Tortoise microvirus 92 | Tortoise microvirus 92 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4294 | 49.697 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765644 | Tortoise microvirus 91 | Tortoise microvirus 91 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4298 | 49.581 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765643 | Tortoise microvirus 90 | Tortoise microvirus 90 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4337 | 48.697 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765642 | Tortoise microvirus 89 | Tortoise microvirus 89 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4345 | 41.864 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765641 | Tortoise microvirus 88 | Tortoise microvirus 88 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4370 | 56.339 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765640 | Tortoise microvirus 87 | Tortoise microvirus 87 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4392 | 37.682 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765639 | Tortoise microvirus 86 | Tortoise microvirus 86 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4483 | 49.61 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765638 | Tortoise microvirus 85 | Tortoise microvirus 85 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5537 | 38.776 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765637 | Tortoise microvirus 84 | Tortoise microvirus 84 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5775 | 41.905 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765636 | Tortoise microvirus 83 | Tortoise microvirus 83 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 44.048 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765635 | Tortoise microvirus 82 | Tortoise microvirus 82 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5853 | 59.047 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765634 | Tortoise microvirus 81 | Tortoise microvirus 81 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5915 | 56.991 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765633 | Tortoise microvirus 80 | Tortoise microvirus 80 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5973 | 42.893 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765632 | Tortoise microvirus 79 | Tortoise microvirus 79 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5977 | 55.044 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765631 | Tortoise microvirus 78 | Tortoise microvirus 78 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6078 | 63.508 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765630 | Tortoise microvirus 77 | Tortoise microvirus 77 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6096 | 63.091 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765629 | Tortoise microvirus 76 | Tortoise microvirus 76 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6290 | 62.21 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765628 | Tortoise microvirus 75 | Tortoise microvirus 75 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6343 | 59.53 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765627 | Tortoise microvirus 74 | Tortoise microvirus 74 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6430 | 60.871 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765626 | Tortoise microvirus 73 | Tortoise microvirus 73 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6526 | 60.788 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765625 | Tortoise microvirus 72 | Tortoise microvirus 72 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6530 | 59.219 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765624 | Tortoise microvirus 71 | Tortoise microvirus 71 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6456 | 61.261 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765623 | Tortoise microvirus 70 | Tortoise microvirus 70 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6549 | 56.222 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765622 | Tortoise microvirus 69 | Tortoise microvirus 69 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4392 | 37.773 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765621 | Tortoise microvirus 53 | Tortoise microvirus 53 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4395 | 47.713 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765620 | Tortoise microvirus 68 | Tortoise microvirus 68 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4951 | 41.911 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765619 | Tortoise microvirus 67 | Tortoise microvirus 67 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5507 | 37.48 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765618 | Tortoise microvirus 66 | Tortoise microvirus 66 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5650 | 40.372 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765617 | Tortoise microvirus 65 | Tortoise microvirus 65 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5676 | 37.615 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765616 | Tortoise microvirus 64 | Tortoise microvirus 64 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5735 | 41.622 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765615 | Tortoise microvirus 63 | Tortoise microvirus 63 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5807 | 41.433 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765614 | Tortoise microvirus 62 | Tortoise microvirus 62 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5810 | 41.583 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765613 | Tortoise microvirus 61 | Tortoise microvirus 61 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5835 | 41.731 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765612 | Tortoise microvirus 60 | Tortoise microvirus 60 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5944 | 40.949 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765611 | Tortoise microvirus 59 | Tortoise microvirus 59 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5983 | 42.303 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765610 | Tortoise microvirus 58 | Tortoise microvirus 58 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5985 | 42.155 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765609 | Tortoise microvirus 57 | Tortoise microvirus 57 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5985 | 38.312 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765608 | Tortoise microvirus 56 | Tortoise microvirus 56 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6126 | 35.896 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765607 | Tortoise microvirus 55 | Tortoise microvirus 55 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5895 | 41.425 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765606 | Tortoise microvirus 54 | Tortoise microvirus 54 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5891 | 41.453 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765605 | Tortoise microvirus 53 | Tortoise microvirus 53 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4372 | 56.725 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765604 | Tortoise microvirus 52 | Tortoise microvirus 52 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.753 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765603 | Tortoise microvirus 51 | Tortoise microvirus 51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4395 | 47.713 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765602 | Tortoise microvirus 50 | Tortoise microvirus 50 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4829 | 38.621 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765601 | Tortoise microvirus 49 | Tortoise microvirus 49 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5403 | 37.664 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765600 | Tortoise microvirus 48 | Tortoise microvirus 48 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5705 | 37.704 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765599 | Tortoise microvirus 47 | Tortoise microvirus 47 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5729 | 57.095 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765598 | Tortoise microvirus 46 | Tortoise microvirus 46 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5751 | 58.164 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765597 | Tortoise microvirus 45 | Tortoise microvirus 45 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5764 | 39.4 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765596 | Tortoise microvirus 44 | Tortoise microvirus 44 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5895 | 41.408 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765595 | Tortoise microvirus 43 | Tortoise microvirus 43 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5908 | 61.171 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765594 | Tortoise microvirus 13 | Tortoise microvirus 13 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5912 | 36.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765593 | Tortoise microvirus 42 | Tortoise microvirus 42 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5948 | 41.812 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765592 | Tortoise microvirus 41 | Tortoise microvirus 41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5982 | 42.377 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765591 | Tortoise microvirus 10 | Tortoise microvirus 10 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5986 | 38.172 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765590 | Tortoise microvirus 40 | Tortoise microvirus 40 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6154 | 36.009 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765589 | Tortoise microvirus 39 | Tortoise microvirus 39 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6204 | 59.139 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765588 | Tortoise microvirus 38 | Tortoise microvirus 38 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6338 | 64.263 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765587 | Tortoise microvirus 37 | Tortoise microvirus 37 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6418 | 58.242 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765586 | Tortoise microvirus 36 | Tortoise microvirus 36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5735 | 41.133 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765585 | Tortoise microvirus 35 | Tortoise microvirus 35 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4217 | 36.78 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765584 | Tortoise microvirus 34 | Tortoise microvirus 34 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4250 | 49.718 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765583 | Tortoise microvirus 33 | Tortoise microvirus 33 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4346 | 41.118 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765582 | Tortoise microvirus 32 | Tortoise microvirus 32 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4374 | 46.182 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765581 | Tortoise microvirus 31 | Tortoise microvirus 31 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.64 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765580 | Tortoise microvirus 30 | Tortoise microvirus 30 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4444 | 38.771 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765579 | Tortoise microvirus 29 | Tortoise microvirus 29 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4513 | 42.544 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765578 | Tortoise microvirus 28 | Tortoise microvirus 28 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4531 | 47.252 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765577 | Tortoise microvirus 27 | Tortoise microvirus 27 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4545 | 43.212 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765576 | Tortoise microvirus 26 | Tortoise microvirus 26 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4600 | 37.761 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765575 | Tortoise microvirus 25 | Tortoise microvirus 25 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4607 | 46.343 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765574 | Tortoise microvirus 24 | Tortoise microvirus 24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4732 | 45.562 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765573 | Tortoise microvirus 23 | Tortoise microvirus 23 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4841 | 41.293 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765572 | Tortoise microvirus 22 | Tortoise microvirus 22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4851 | 39.27 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765571 | Tortoise microvirus 21 | Tortoise microvirus 21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4587 | 41.465 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765570 | Tortoise microvirus 20 | Tortoise microvirus 20 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5355 | 40.299 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765569 | Tortoise microvirus 19 | Tortoise microvirus 19 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5409 | 41.893 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765568 | Tortoise microvirus 18 | Tortoise microvirus 18 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5438 | 35.417 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765567 | Tortoise microvirus 17 | Tortoise microvirus 17 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5684 | 37.614 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765566 | Tortoise microvirus 16 | Tortoise microvirus 16 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5764 | 39.33 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765565 | Tortoise microvirus 15 | Tortoise microvirus 15 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5837 | 41.323 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765564 | Tortoise microvirus 14 | Tortoise microvirus 14 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5850 | 36.051 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765563 | Tortoise microvirus 13 | Tortoise microvirus 13 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5912 | 36.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765562 | Tortoise microvirus 12 | Tortoise microvirus 12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5961 | 42.275 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765561 | Tortoise microvirus 11 | Tortoise microvirus 11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5983 | 38.743 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765560 | Tortoise microvirus 10 | Tortoise microvirus 10 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5986 | 38.172 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765559 | Tortoise microvirus 9 | Tortoise microvirus 9 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4383 | 47.821 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765558 | Tortoise microvirus 8 | Tortoise microvirus 8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4389 | 37.981 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765557 | Tortoise microvirus 7 | Tortoise microvirus 7 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5554 | 44.094 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765556 | Tortoise microvirus 6 | Tortoise microvirus 6 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5665 | 40.088 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765555 | Tortoise microvirus 5 | Tortoise microvirus 5 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5672 | 37.518 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765554 | Tortoise microvirus 4 | Tortoise microvirus 4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5788 | 41.154 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765553 | Tortoise microvirus 3 | Tortoise microvirus 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5823 | 41.594 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765552 | Tortoise microvirus 2 | Tortoise microvirus 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5975 | 42.226 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK765551 | Tortoise microvirus 1 | Tortoise microvirus 1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6243 | 37.338 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK249234 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4232 | 47.164 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249233 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4170 | 55.06 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249232 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4237 | 42.743 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249231 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4238 | 59.462 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249230 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4302 | 49.558 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249229 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4320 | 54.653 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249228 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4205 | 38.906 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249227 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4318 | 47.244 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249226 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4276 | 50.094 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249225 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4351 | 51.919 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249224 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4293 | 46.401 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249223 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4368 | 51.832 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249222 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4391 | 50.808 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249221 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4422 | 51.289 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249220 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4400 | 50.432 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249219 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4445 | 45.287 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249218 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4387 | 56.394 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249217 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4401 | 46.308 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249216 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4459 | 44.539 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249215 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4410 | 47.959 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249214 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4487 | 53.688 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249213 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4502 | 50.444 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249212 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4484 | 53.078 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249211 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4350 | 55.494 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249210 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4483 | 57.484 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249209 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4438 | 55.273 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249208 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4443 | 44.564 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249207 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4460 | 44.619 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249206 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4460 | 55.404 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249205 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4549 | 50.077 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249204 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4521 | 53.24 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249203 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 54.67 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249202 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4471 | 53.5 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249201 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4538 | 54.495 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249200 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4547 | 51.946 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249199 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4335 | 42.999 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249198 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4551 | 52.362 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249197 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4488 | 52.161 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249196 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4558 | 51.514 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249195 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4498 | 49.6 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249194 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4570 | 54.004 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249193 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4503 | 54.053 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249192 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4581 | 49.88 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249191 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4507 | 38.34 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249190 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4510 | 53.902 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249189 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4499 | 50.633 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249188 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4591 | 51.165 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249187 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4617 | 56.27 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249186 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4621 | 54.663 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249185 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4590 | 54.619 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249184 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4596 | 50.914 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249183 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4608 | 47.439 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249182 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4618 | 53.53 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249181 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4647 | 56.961 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249180 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4706 | 46.876 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249179 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4680 | 57.436 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249178 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4692 | 50.298 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249177 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4721 | 49.587 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249176 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4706 | 46.77 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249175 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4700 | 51.021 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249174 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4646 | 48.192 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249173 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4721 | 56.704 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249172 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4733 | 58.568 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249171 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4630 | 54.6 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249170 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4615 | 48.732 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249169 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4768 | 50.315 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249168 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4779 | 52.249 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249167 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4825 | 55.15 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249166 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4799 | 48.593 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249165 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4858 | 48.209 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249164 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4711 | 52.473 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249163 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4774 | 46.23 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249162 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4789 | 49.697 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249161 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4765 | 50.073 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249160 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4870 | 55.298 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249159 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4901 | 56.948 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249158 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4633 | 47.874 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249157 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4901 | 42.42 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249156 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4980 | 55.663 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249155 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4637 | 53.871 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249154 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4667 | 48.168 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249153 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4581 | 50.207 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249152 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4731 | 51.067 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249151 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4521 | 52.201 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249150 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4633 | 50.14 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249149 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4565 | 50.011 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249148 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6565 | 39.513 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249147 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4461 | 55.257 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249146 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4498 | 35.705 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249145 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4727 | 48.36 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249144 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4841 | 54.699 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249143 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4474 | 55.811 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249142 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4689 | 52.868 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249141 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4557 | 54.883 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249140 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4694 | 52.748 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249139 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4582 | 51.44 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249138 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4231 | 43.961 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249137 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4565 | 51.172 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249136 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4656 | 51.224 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249135 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4324 | 54.14 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249134 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4483 | 57.417 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249133 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4379 | 43.549 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249132 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4675 | 50.439 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249131 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4594 | 54.049 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249130 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4660 | 50.451 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249129 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4625 | 53.708 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249128 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4544 | 48.438 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK249127 | Blackfly microvirus SF02 | Blackfly microvirus SF02 Microvirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4873 | 48.369 | Microvirus | Unclassified | Microviridae | Unspecified |
| MK308637 | Microbacterium phage Schubert | Microbacterium phage Schubert Microbacterium virus Schubert Schubertvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38820 | 61.419 | Schubertvirus | Unclassified | Siphoviridae | Microbacterium |
| MH825712 | Microbacterium phage ValentiniPuff | Microbacterium phage ValentiniPuff Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62517 | 67.148 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH697589 | Microbacterium phage Neferthena | Microbacterium phage Neferthena Microbacterium virus Neferthena Neferthenavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41706 | 64.386 | Neferthenavirus | Unclassified | Siphoviridae | Microbacterium |
| MH590604 | Microbacterium phage ColaCorta | Microbacterium phage ColaCorta unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40494 | 62.031 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MH590590 | Microbacterium phage Schnapsidee | Microbacterium phage Schnapsidee unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41872 | 63.362 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH513982 | Microbacterium phage Sansa | Microbacterium phage Sansa unclassified Elerivirus Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40306 | 61.792 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MH513981 | Microbacterium phage Papafritta | Microbacterium phage Papafritta unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41573 | 63.399 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH479922 | Microbacterium phage Redfield | Microbacterium phage Redfield unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41930 | 63.341 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153811 | Microbacterium phage Teagan | Microbacterium phage Teagan Microbacterium virus Teagan Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41793 | 63.451 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153806 | Microbacterium phage Oats | Microbacterium phage Oats unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.393 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153805 | Microbacterium phage Martin | Microbacterium phage Martin unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41812 | 63.549 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH153802 | Microbacterium phage Gargoyle | Microbacterium phage Gargoyle unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41803 | 63.512 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH047631 | Microbacterium phage Triscuit | Microbacterium phage Triscuit Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67539 | 58.283 | Unclassified | Unclassified | Siphoviridae | Microbacterium |
| MH045565 | Microbacterium phage Antoinette | Microbacterium phage Antoinette unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41858 | 63.436 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045564 | Microbacterium phage TeddyBear | Microbacterium phage TeddyBear unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.388 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045563 | Microbacterium phage Robinson | Microbacterium phage Robinson unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41874 | 63.474 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045562 | Microbacterium phage Raptor | Microbacterium phage Raptor unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41801 | 63.405 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045560 | Microbacterium phage Nagem | Microbacterium phage Nagem unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41846 | 63.48 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045559 | Microbacterium phage Etna | Microbacterium phage Etna unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41908 | 63.413 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045558 | Microbacterium phage Dave | Microbacterium phage Dave unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41858 | 63.417 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045555 | Microbacterium phage BeeBee8 | Microbacterium phage BeeBee8 unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42027 | 63.35 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MH045554 | Microbacterium phage Bandik | Microbacterium phage Bandik unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41804 | 63.547 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG962367 | Microbacterium phage Gelo | Microbacterium phage Gelo unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41562 | 63.354 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG962360 | Microbacterium phage AlexAdler | Microbacterium phage AlexAdler unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41834 | 63.408 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG944222 | Microbacterium phage StingRay | Microbacterium phage StingRay unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41597 | 63.488 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG944219 | Microbacterium phage PuppyEggo | Microbacterium phage PuppyEggo unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41803 | 63.304 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG925352 | Microbacterium phage Nattles | Microbacterium phage Nattles unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41544 | 63.364 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG925347 | Microbacterium phage Lucky3 | Microbacterium phage Lucky3 unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39640 | 64.087 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MG925345 | Microbacterium phage Koji | Microbacterium phage Koji Microbacterium virus Koji Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39403 | 64.173 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MG925343 | Microbacterium phage Golden | Microbacterium phage Golden Microbacterium virus Golden Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39640 | 64.084 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MG920061 | Microbacterium phage Bonino | Microbacterium phage Bonino unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41534 | 63.355 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839030 | Microbacterium phage Balsa | Microbacterium phage Balsa unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41862 | 63.377 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839029 | Microbacterium phage Ilzat | Microbacterium phage Ilzat Microbacterium virus Ilzat Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41525 | 63.485 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839028 | Microbacterium phage Tenda | Microbacterium phage Tenda unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41553 | 63.43 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839027 | Microbacterium phage Eleri | Microbacterium phage Eleri Microbacterium virus Eleri Elerivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40366 | 61.975 | Elerivirus | Unclassified | Siphoviridae | Microbacterium |
| MG839026 | Microbacterium phage Superfresh | Microbacterium phage Superfresh unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41860 | 63.399 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839025 | Microbacterium phage Raccoon | Microbacterium phage Raccoon unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41894 | 63.389 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839024 | Microbacterium phage Peppino | Microbacterium phage Peppino unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41932 | 63.343 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839023 | Microbacterium phage Peep | Microbacterium phage Peep unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41856 | 63.432 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839022 | Microbacterium phage Ludgate | Microbacterium phage Ludgate unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41799 | 63.454 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839021 | Microbacterium phage Knox | Microbacterium phage Knox unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41797 | 63.344 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839020 | Microbacterium phage Kale | Microbacterium phage Kale unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41558 | 63.432 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839019 | Microbacterium phage Hamlet | Microbacterium phage Hamlet Microbacterium virus Hamlet Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41934 | 63.402 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839018 | Microbacterium phage Espinosa | Microbacterium phage Espinosa unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41553 | 63.411 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839017 | Microbacterium phage Baines | Microbacterium phage Baines unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.352 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839016 | Microbacterium phage AxiPup | Microbacterium phage AxiPup unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41770 | 63.33 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MG839015 | Microbacterium phage Aubergine | Microbacterium phage Aubergine unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41555 | 63.434 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| KT624200 | Bacillus phage SP-15 | Bacillus phage SP-15 Bacillus virus SP15 Thornevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221908 | 38.608 | Thornevirus | Unclassified | Myoviridae | Bacillus |
| GQ357915 | Delftia phage PhiW-14 | Delftia phage PhiW-14 Delftia virus PhiW14 Ionavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157486 | 56.277 | Ionavirus | Unclassified | Myoviridae | Delftia |
| MK719754 | Rheinheimera phage vB_RspM_Barba35A | Rheinheimera phage vB_RspM_Barba35A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80206 | 38.178 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719753 | Rheinheimera phage vB_RspM_Barba34A | Rheinheimera phage vB_RspM_Barba34A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719752 | Rheinheimera phage vB_RspM_Barba33A | Rheinheimera phage vB_RspM_Barba33A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80336 | 38.218 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719751 | Rheinheimera phage vB_RspM_Barba32A | Rheinheimera phage vB_RspM_Barba32A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.226 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719750 | Rheinheimera phage vB_RspM_Barba31A | Rheinheimera phage vB_RspM_Barba31A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84615 | 38.271 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719749 | Rheinheimera phage vB_RspM_Barba30A | Rheinheimera phage vB_RspM_Barba30A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80237 | 38.173 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719748 | Rheinheimera phage vB_RspM_Barba29S | Rheinheimera phage vB_RspM_Barba29S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81390 | 38.193 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719747 | Rheinheimera phage vB_RspM_Barba29A | Rheinheimera phage vB_RspM_Barba29A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719746 | Rheinheimera phage vB_RspM_Barba28S | Rheinheimera phage vB_RspM_Barba28S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81595 | 38.406 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719745 | Rheinheimera phage vB_RspM_Barba28A | Rheinheimera phage vB_RspM_Barba28A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80270 | 38.204 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719744 | Rheinheimera phage vB_RspM_Barba27S | Rheinheimera phage vB_RspM_Barba27S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719743 | Rheinheimera phage vB_RspM_Barba26S | Rheinheimera phage vB_RspM_Barba26S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719742 | Rheinheimera phage vB_RspM_Barba26A | Rheinheimera phage vB_RspM_Barba26A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82164 | 38.387 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719741 | Rheinheimera phage vB_RspM_Barba25S | Rheinheimera phage vB_RspM_Barba25S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80355 | 38.224 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719740 | Rheinheimera phage vB_RspM_Barba25A | Rheinheimera phage vB_RspM_Barba25A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80284 | 38.203 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719739 | Rheinheimera phage vB_RspM_Barba24S | Rheinheimera phage vB_RspM_Barba24S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80340 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719738 | Rheinheimera phage vB_RspM_Barba23S | Rheinheimera phage vB_RspM_Barba23S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.228 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719737 | Rheinheimera phage vB_RspM_Barba23A | Rheinheimera phage vB_RspM_Barba23A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83415 | 38.404 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719736 | Rheinheimera phage vB_RspM_Barba22S | Rheinheimera phage vB_RspM_Barba22S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.222 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719735 | Rheinheimera phage vB_RspM_Barba22A | Rheinheimera phage vB_RspM_Barba22A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.224 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719734 | Rheinheimera phage vB_RspM_Barba21S | Rheinheimera phage vB_RspM_Barba21S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80785 | 38.22 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719733 | Rheinheimera phage Barba21A | Rheinheimera phage Barba21A Rheinheimera virus Barba21A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80361 | 38.28 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719732 | Rheinheimera phage vB_RspM_Barba20S | Rheinheimera phage vB_RspM_Barba20S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81477 | 38.246 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719731 | Rheinheimera phage vB_RspM_Barba20A | Rheinheimera phage vB_RspM_Barba20A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719730 | Rheinheimera phage vB_RspM_Barba19A | Rheinheimera phage vB_RspM_Barba19A Rheinheimera virus Barba19A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84608 | 38.247 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719729 | Rheinheimera phage vB_RspM_Barba18A | Rheinheimera phage vB_RspM_Barba18A Rheinheimera virus Barba18A Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719728 | Rheinheimera phage vB_RspM_Barba17S | Rheinheimera phage vB_RspM_Barba17S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81537 | 38.27 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719727 | Rheinheimera phage vB_RspM_Barba17A | Rheinheimera phage vB_RspM_Barba17A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.229 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719726 | Rheinheimera phage vB_RspM_Barba16S | Rheinheimera phage vB_RspM_Barba16S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80226 | 38.238 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719725 | Rheinheimera phage vB_RspM_Barba15S | Rheinheimera phage vB_RspM_Barba15S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80303 | 38.207 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719724 | Rheinheimera phage vB_RspM_Barba15A | Rheinheimera phage vB_RspM_Barba15A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82059 | 38.166 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719723 | Rheinheimera phage vB_RspM_Barba14S | Rheinheimera phage vB_RspM_Barba14S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80312 | 38.216 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719722 | Rheinheimera phage vB_RspM_Barba14A | Rheinheimera phage vB_RspM_Barba14A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80332 | 38.231 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719721 | Rheinheimera phage vB_RspM_Barba13A | Rheinheimera phage vB_RspM_Barba13A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719720 | Rheinheimera phage vB_RspM_Barba12S | Rheinheimera phage vB_RspM_Barba12S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80342 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719719 | Rheinheimera phage vB_RspM_Barba12A | Rheinheimera phage vB_RspM_Barba12A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719718 | Rheinheimera phage vB_RspM_Barba11S | Rheinheimera phage vB_RspM_Barba11S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80353 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719717 | Rheinheimera phage vB_RspM_Barba10S | Rheinheimera phage vB_RspM_Barba10S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80469 | 38.199 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719716 | Rheinheimera phage vB_RspM_Barba10A | Rheinheimera phage vB_RspM_Barba10A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.225 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719715 | Rheinheimera phage vB_RspM_Barba9A | Rheinheimera phage vB_RspM_Barba9A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80203 | 38.151 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719714 | Rheinheimera phage Barba8S | Rheinheimera phage Barba8S Rheinheimera virus Barba8S Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81920 | 38.315 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719713 | Rheinheimera phage vB_RspM_Barba8A | Rheinheimera phage vB_RspM_Barba8A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80236 | 38.185 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719712 | Rheinheimera phage vB_RspM_Barba7A | Rheinheimera phage vB_RspM_Barba7A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719711 | Rheinheimera phage vB_RspM_Barba6A | Rheinheimera phage vB_RspM_Barba6A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80332 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719710 | Rheinheimera phage Barba5S | Rheinheimera phage Barba5S Rheinheimera virus Barba5S Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82744 | 38.218 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719709 | Rheinheimera phage vB_RspM_Barba5A | Rheinheimera phage vB_RspM_Barba5A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80236 | 38.189 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719708 | Rheinheimera phage vB_RspM_Barba4S | Rheinheimera phage vB_RspM_Barba4S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80400 | 38.246 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719707 | Rheinheimera phage vB_RspM_Barba4A | Rheinheimera phage vB_RspM_Barba4A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80327 | 38.214 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719706 | Rheinheimera phage vB_RspM_Barba3S | Rheinheimera phage vB_RspM_Barba3S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719705 | Rheinheimera phage vB_RspM_Barba3A | Rheinheimera phage vB_RspM_Barba3A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81004 | 38.23 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719704 | Rheinheimera phage vB_RspM_Barba2S | Rheinheimera phage vB_RspM_Barba2S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719703 | Rheinheimera phage vB_RspM_Barba2A | Rheinheimera phage vB_RspM_Barba2A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80352 | 38.223 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719702 | Rheinheimera phage vB_RspM_Barba1S | Rheinheimera phage vB_RspM_Barba1S unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80341 | 38.227 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK719701 | Rheinheimera phage vB_RspM_Barba1A | Rheinheimera phage vB_RspM_Barba1A unclassified Barbavirus Barbavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80240 | 38.202 | Barbavirus | Unclassified | Myoviridae | Rheinheimera |
| MK759918 | Bacillus phage vB_BtS_B83 | Bacillus phage vB_BtS_B83 Bacillus virus B83 Pushchinovirus Skryabinvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49952 | 35.79 | Pushchinovirus | Skryabinvirinae | Siphoviridae | Bacillus |
| MK796244 | Vibrio phage Achelous | Vibrio phage Achelous unclassified Cetovirus Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121089 | 40.036 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MK759854 | Shigella phage DS8 | Shigella phage DS8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44605 | 46.517 | Unclassified | Unclassified | Siphoviridae | Shigella |
| MH518298 | Pseudomonas phage SCYZ1 | Pseudomonas phage SCYZ1 unclassified Krylovvirus Krylovvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47475 | 52.731 | Krylovvirus | Unclassified | Podoviridae | Pseudomonas |
| MK494128 | Mycobacterium phage Heliosoles | Mycobacterium phage Heliosoles unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50577 | 64.013 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494127 | Mycobacterium phage Noella | Mycobacterium phage Noella unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50079 | 64.045 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494110 | Mycobacterium phage SwagPigglett | Mycobacterium phage SwagPigglett unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58930 | 61.259 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK494096 | Mycobacterium phage Mainiac | Mycobacterium phage Mainiac unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50828 | 64.02 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524506 | Mycobacterium phage Zeuska | Mycobacterium phage Zeuska unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53598 | 63.736 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524499 | Mycobacterium phage Tripl3t | Mycobacterium phage Tripl3t unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53565 | 63.689 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524496 | Mycobacterium phage Veracruz | Mycobacterium phage Veracruz unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50062 | 64.049 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK524493 | Mycobacterium phage Darionha | Mycobacterium phage Darionha unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41901 | 66.566 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK380016 | Klebsiella phage Kund-ULIP54 | Klebsiella phage Kund-ULIP54 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41109 | 53.022 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK359343 | Mycobacterium phage Pollywog | Mycobacterium phage Pollywog unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58397 | 61.251 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359341 | Mycobacterium phage GingkoMaracino | Mycobacterium phage GingkoMaracino unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50013 | 64.033 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359315 | Mycobacterium phage MisterCuddles | Mycobacterium phage MisterCuddles unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57754 | 61.355 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359307 | Mycobacterium phage Ochi17 | Mycobacterium phage Ochi17 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58069 | 61.014 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK310140 | Mycobacterium phage Gemma | Mycobacterium phage Gemma unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50574 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK380015 | Klebsiella phage Kund-ULIP47 | Klebsiella phage Kund-ULIP47 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41397 | 52.975 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK380014 | Klebsiella phage K1-ULIP33 | Klebsiella phage K1-ULIP33 unclassified Molineuxvirinae Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44122 | 50.403 | Unclassified | Molineuxvirinae | Autographiviridae | Klebsiella |
| MK284519 | Mycobacterium phage Norbert | Mycobacterium phage Norbert unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50078 | 64.05 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK284518 | Mycobacterium phage ACFishhook | Mycobacterium phage ACFishhook unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47343 | 63.99 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK112543 | Mycobacterium Phage Kalb97 | Mycobacterium Phage Kalb97 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49943 | 64.115 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK016496 | Mycobacterium phage IrishSherpFalk | Mycobacterium phage IrishSherpFalk unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58575 | 61.641 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727554 | Mycobacterium phage MuchMore | Mycobacterium phage MuchMore unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48545 | 64.171 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH727548 | Mycobacterium phage Galactic | Mycobacterium phage Galactic unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57861 | 61.447 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669014 | Mycobacterium phage Spikelee | Mycobacterium phage Spikelee unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58604 | 61.038 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669012 | Mycobacterium phage QuickMath | Mycobacterium phage QuickMath unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57526 | 61.706 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH669001 | Mycobacterium phage EleanorGeorge | Mycobacterium phage EleanorGeorge unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59482 | 61.731 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH590596 | Mycobacterium phage Mantra | Mycobacterium phage Mantra unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57001 | 61.545 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH513975 | Mycobacterium phage Misomonster | Mycobacterium phage Misomonster unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48504 | 64.325 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH536825 | Mycobacterium phage Ollie | Mycobacterium phage Ollie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50721 | 63.972 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH509442 | Mycobacterium phage Aminay | Mycobacterium phage Aminay unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60430 | 67.834 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450128 | Mycobacterium phage Reba | Mycobacterium phage Reba unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49412 | 63.958 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450124 | Mycobacterium phage Marius | Mycobacterium phage Marius unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50570 | 64.018 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450119 | Mycobacterium phage Kalnoky | Mycobacterium phage Kalnoky unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50571 | 64.031 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371121 | Mycobacterium phage Phranny | Mycobacterium phage Phranny unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48024 | 64.024 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH155871 | Mycobacterium phage Mattes | Mycobacterium phage Mattes unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58074 | 61.31 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH155865 | Mycobacterium phage BobaPhett | Mycobacterium phage BobaPhett unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59815 | 61.326 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077585 | Mycobacterium phage TChen | Mycobacterium phage TChen Mycobacterium virus TChen Thetabobvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57742 | 62.338 | Thetabobvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH077583 | Mycobacterium phage PGHhamlin | Mycobacterium phage PGHhamlin unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50840 | 64.024 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020243 | Mycobacterium phage TootsiePop | Mycobacterium phage TootsiePop unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57455 | 62.947 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020242 | Mycobacterium phage Misha28 | Mycobacterium phage Misha28 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57455 | 62.945 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020235 | Mycobacterium phage Batiatus | Mycobacterium phage Batiatus unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57737 | 61.836 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MH001448 | Mycobacterium phage UncleRicky | Mycobacterium phage UncleRicky unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57256 | 61.337 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MG757161 | Mycobacterium phage LugYA | Mycobacterium phage LugYA unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50878 | 64.004 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF373840 | Mycobacterium phage Fred313 | Mycobacterium phage Fred313 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50053 | 63.974 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185732 | Mycobacterium phage Stagni | Mycobacterium phage Stagni unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50856 | 64.032 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185729 | Mycobacterium phage KADY | Mycobacterium phage KADY unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50898 | 64.207 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY965063 | Mycobacterium phage PurpleHaze | Mycobacterium phage PurpleHaze unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48596 | 63.999 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY464936 | Mycobacterium phage Idleandcovert | Mycobacterium phage Idleandcovert unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49156 | 63.793 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KY348865 | Mycobacterium phage Bubbles123 | Mycobacterium phage Bubbles123 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57647 | 61.375 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KX808129 | Mycobacterium phage Sabinator | Mycobacterium phage Sabinator unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50883 | 64.006 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX670788 | Mycobacterium phage Louie6 | Mycobacterium phage Louie6 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50899 | 64.211 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX664448 | Mycobacterium phage Watson | Mycobacterium phage Watson unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50841 | 64.045 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX640831 | Mycobacterium phage Margo | Mycobacterium phage Margo unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50087 | 63.983 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX670828 | Mycobacterium phage Todacoro | Mycobacterium phage Todacoro unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50066 | 64.045 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX507362 | Mycobacterium phage Aglet | Mycobacterium phage Aglet unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50866 | 64.055 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU985091 | Mycobacterium phage Hercules11 | Mycobacterium phage Hercules11 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50892 | 63.992 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU984914 | Mycobacterium phage Malinsilva | Mycobacterium phage Malinsilva unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50839 | 64.026 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU578077 | Mycobacterium phage Marie | Mycobacterium phage Marie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT381277 | Mycobacterium phage Wooldri | Mycobacterium phage Wooldri unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50797 | 64.004 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT326767 | Mycobacterium phage Texage | Mycobacterium phage Texage unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50081 | 64.044 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KT359365 | Mycobacterium phage DaHudson | Mycobacterium phage DaHudson unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50841 | 64.053 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KR997968 | Mycobacterium phage Quico | Mycobacterium phage Quico unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58671 | 61.688 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KR997931 | Mycobacterium phage Girafales | Mycobacterium phage Girafales unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58456 | 61.687 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KR935214 | Mycobacterium phage Kimberlium | Mycobacterium phage Kimberlium unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56826 | 61.44 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KP027202 | Mycobacterium phage Taurus | Mycobacterium phage Taurus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 64.001 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP017310 | Mycobacterium phage Phoxy | Mycobacterium phage Phoxy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49267 | 64.209 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KP017311 | Mycobacterium phage Spike509 | Mycobacterium phage Spike509 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50989 | 64.145 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM592966 | Mycobacterium phage QuinnKiro | Mycobacterium phage QuinnKiro unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50066 | 64.049 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM347889 | Mycobacterium phage BuzzLyseyear | Mycobacterium phage BuzzLyseyear unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59419 | 61.132 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101124 | Mycobacterium phage Squirty | Mycobacterium phage Squirty Mycobacterium virus Squirty Squirtyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60285 | 62.405 | Squirtyvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM101119 | Mycobacterium phage Tiffany | Mycobacterium phage Tiffany unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50768 | 63.991 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661272 | Mycobacterium phage Methuselah | Mycobacterium phage Methuselah Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50891 | 64.198 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN698994 | Mycobacterium phage DS6A | Mycobacterium phage DS6A Mycobacterium virus DS6A Hnatkovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60588 | 68.438 | Hnatkovirus | Unclassified | Siphoviridae | Mycobacterium |
| JN542517 | Mycobacterium virus Drago | Mycobacterium virus Drago Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54411 | 61.17 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704114 | Mycobacterium phage Vix | Mycobacterium phage Vix unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50963 | 64.046 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JF704111 | Mycobacterium phage Rockstar | Mycobacterium phage Rockstar Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47780 | 64.28 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK656892 | Staphylococcus phage vBSP-A2 | Staphylococcus phage vBSP-A2 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136528 | 30.447 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY926699 | Lactococcus phage PLgY-30 | Lactococcus phage PLgY-30 unclassified Uwajimavirus Uwajimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29284 | 37.41 | Uwajimavirus | Unclassified | Siphoviridae | Lactococcus |
| KY926698 | Lactococcus phage PLgY-16 | Lactococcus phage PLgY-16 unclassified Uwajimavirus Uwajimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29342 | 37.373 | Uwajimavirus | Unclassified | Siphoviridae | Lactococcus |
| KY888143 | Lactococcus phage PLgW-1 | Lactococcus phage PLgW-1 Lactococcus virus PLgW1 Uwajimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30189 | 37.322 | Uwajimavirus | Unclassified | Siphoviridae | Lactococcus |
| MK521906 | Klebsiella phage vB_KpnM_FZ14 | Klebsiella phage vB_KpnM_FZ14 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49370 | 48.584 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MK224592 | Mycobacterium phage Henu3 PeY-2017 | Mycobacterium phage Henu3 PeY-2017 unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61276 | 61.172 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK814762 | Microbacterium phage Bustleton | Microbacterium phage Bustleton unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39184 | 64.432 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MK757447 | Microbacterium phage Greys | Microbacterium phage Greys unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41797 | 63.406 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK521907 | Klebsiella phage vB_KpnS_FZ41 | Klebsiella phage vB_KpnS_FZ41 unclassified Sugarlandvirus Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106104 | 45.221 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| MK521905 | Klebsiella phage vB_KpnP_FZ12 | Klebsiella phage vB_KpnP_FZ12 unclassified Przondovirus Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39519 | 53.063 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK521904 | Klebsiella phage vB_KpnS_FZ10 | Klebsiella phage vB_KpnS_FZ10 Klebsiella virus FZ10 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50381 | 50.66 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH160767 | Escherichia phage vB_EcoM-Ro157c2YLVW | Escherichia phage vB_EcoM-Ro157c2YLVW unclassified Punavirus Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80732 | 47.764 | Punavirus | Unclassified | Myoviridae | Escherichia |
| MF041990 | Lactobacillus phage phiEF-1.1 | Lactobacillus phage phiEF-1.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39130 | 39.438 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MH115576 | Acinetobacter phage AM106 | Acinetobacter phage AM106 unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52031 | 40.172 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| MK685667 | Klebsiella phage vB_KpnM_IME346 | Klebsiella phage vB_KpnM_IME346 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49482 | 49.052 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MH370375 | Salmonella phage S124 | Salmonella phage S124 Salmonella virus S124 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112564 | 40.119 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370387 | Salmonella phage S149 | Salmonella phage S149 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41391 | 47.435 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MH370386 | Salmonella phage S147 | Salmonella phage S147 Salmonella virus S147 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111447 | 39.985 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370385 | Salmonella phage S142 | Salmonella phage S142 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43119 | 49.616 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370384 | Salmonella phage S138 | Salmonella phage S138 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43119 | 49.614 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370383 | Salmonella phage S137 | Salmonella phage S137 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50593 | 48.362 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH370382 | Salmonella phage S135 | Salmonella phage S135 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50591 | 48.364 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH370381 | Salmonella phage S134 | Salmonella phage S134 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43118 | 49.652 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370380 | Salmonella phage S133 | Salmonella phage S133 Salmonella virus S133 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110926 | 40.061 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370379 | Salmonella phage S132 | Salmonella phage S132 Salmonella virus S132 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116832 | 39.944 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370378 | Salmonella phage S131 | Salmonella phage S131 Salmonella virus S131 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110091 | 39.216 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370377 | Salmonella phage S130 | Salmonella phage S130 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110091 | 39.054 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370376 | Salmonella phage S126 | Salmonella phage S126 Salmonella virus S126 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111999 | 40.1 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370374 | Salmonella phage S123 | Salmonella phage S123 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43467 | 49.564 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370373 | Salmonella phage S120 | Salmonella phage S120 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43467 | 49.562 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370372 | Salmonella phage S119 | Salmonella phage S119 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43876 | 49.521 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370371 | Salmonella phage S118 | Salmonella phage S118 Salmonella virus S118 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157013 | 44.801 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH370370 | Salmonella phage S117 | Salmonella phage S117 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158110 | 44.668 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH370369 | Salmonella phage S116 | Salmonella phage S116 Salmonella virus S116 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110664 | 40.085 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370368 | Salmonella phage S115 | Salmonella phage S115 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157946 | 44.568 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH370367 | Salmonella phage S114 | Salmonella phage S114 Salmonella virus S114 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110926 | 40.062 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370366 | Salmonella phage S113 | Salmonella phage S113 Salmonella virus S113 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112582 | 39.96 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MH370365 | Salmonella phage S111 | Salmonella phage S111 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43421 | 49.672 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370364 | Salmonella phage S107 | Salmonella phage S107 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49818 | 48.583 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH370363 | Salmonella phage S106 | Salmonella phage S106 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42976 | 49.702 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370362 | Salmonella phage S104 | Salmonella phage S104 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43118 | 49.615 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370361 | Salmonella phage S103 | Salmonella phage S103 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42441 | 49.655 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370360 | Salmonella phage S102 | Salmonella phage S102 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42439 | 49.655 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370359 | Salmonella phage S101 | Salmonella phage S101 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42621 | 49.668 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH370358 | Salmonella phage S100 | Salmonella phage S100 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43468 | 49.565 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| GQ919031 | Streptomyces phage ZL12 | Streptomyces phage ZL12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90435 | 69.472 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MK689364 | Edwardsiella virus pEtSU | Edwardsiella virus pEtSU Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 276734 | 43.173 | Petsuvirus | Unclassified | Myoviridae | Edwardsiella |
| MG944236 | Xanthomonas phage XPV3 | Xanthomonas phage XPV3 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47046 | 61.151 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944234 | Xanthomonas phage XPV1 | Xanthomonas phage XPV1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46503 | 61.121 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944233 | Xanthomonas phage XPP9 | Xanthomonas phage XPP9 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48669 | 61.012 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944231 | Xanthomonas phage XPP6 | Xanthomonas phage XPP6 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46281 | 60.995 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944229 | Xanthomonas phage XPP3 | Xanthomonas phage XPP3 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49612 | 60.991 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG944228 | Xanthomonas phage XPP2 | Xanthomonas phage XPP2 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46480 | 60.871 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| LC465543 | Shigella phage SfPhi01 | Shigella phage SfPhi01 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168000 | 35.292 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| JQ287645 | Sulfolobus virus STSV2 | Sulfolobus virus STSV2 unclassified Bicaudaviridae Bicaudaviridae Viruses | 76107 | 35.098 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| KT945994 | Enterococcus phage vB_EfaS_IME197 | Enterococcus phage vB_EfaS_IME197 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41098 | 34.058 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KM378617 | Vibrio phage VpKK5 | Vibrio phage VpKK5 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56637 | 51.318 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| KJ920400 | Bacillus phage Waukesha92 | Bacillus phage Waukesha92 Bacillus virus Waukesha92 Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45648 | 35.452 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| KF926093 | Lactococcus phage phiL47 | Lactococcus phage phiL47 Lactococcus virus phiL47 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128546 | 32.532 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KC171647 | Lactobacillus phage PL-1 | Lactobacillus phage PL-1 Lactobacillus virus PL1 Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38880 | 44.882 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| KC171646 | Lactobacillus phage J-1 | Lactobacillus phage J-1 Lactobacillus virus J1 Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40931 | 44.768 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| KC460990 | Serratia phage Eta | Serratia phage Eta Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42724 | 49.913 | Unclassified | Unclassified | Siphoviridae | Serratia |
| GU967410 | Lactobacillus phage LBR48 | Lactobacillus phage LBR48 Lactobacillus virus LBR48 Anamdongvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48211 | 45.877 | Anamdongvirus | Unclassified | Myoviridae | Lactobacillus |
| EU030939 | Sulfolobus spindle-shaped virus 5 | Sulfolobus spindle-shaped virus 5 Alphafusellovirus Fuselloviridae Viruses | 15330 | 38.48 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| EU030938 | Sulfolobus spindle-shaped virus 4 | Sulfolobus spindle-shaped virus 4 Alphafusellovirus Fuselloviridae Viruses | 15135 | 38.474 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK721200 | Enterococcus phage vB_EfaS_Ef5.3 | Enterococcus phage vB_EfaS_Ef5.3 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39115 | 34.838 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK721192 | Enterococcus phage vB_EfaM_Ef2.3 | Enterococcus phage vB_EfaM_Ef2.3 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147289 | 35.865 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MK721186 | Enterococcus phage vB_EfaS_Ef5.2 | Enterococcus phage vB_EfaS_Ef5.2 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41418 | 34.461 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK721199 | Enterococcus phage vB_EfaS_Ef5.1 | Enterococcus phage vB_EfaS_Ef5.1 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41141 | 34.513 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK721191 | Enterococcus phage vB_EfaS_Ef5.4 | Enterococcus phage vB_EfaS_Ef5.4 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40685 | 34.671 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK693030 | Enterococcus phage vB_EfaM_Ef2.1 | Enterococcus phage vB_EfaM_Ef2.1 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140938 | 35.872 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MK620905 | Microbacterium phage TinSulphur | Microbacterium phage TinSulphur unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41807 | 63.535 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK620904 | Microbacterium phage Pherbot | Microbacterium phage Pherbot unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39171 | 64.415 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MK620903 | Microbacterium phage MillyPhilly | Microbacterium phage MillyPhilly unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41800 | 63.505 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK620902 | Microbacterium phage Phriends | Microbacterium phage Phriends unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41606 | 63.558 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK620901 | Microbacterium phage PrincePhergus | Microbacterium phage PrincePhergus unclassified Kojivirus Kojivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39191 | 64.395 | Kojivirus | Unclassified | Siphoviridae | Microbacterium |
| MK620895 | Microbacterium phage Riyhil | Microbacterium phage Riyhil unclassified Ilzatvirus Ilzatvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41812 | 63.546 | Ilzatvirus | Unclassified | Siphoviridae | Microbacterium |
| MK639187 | Shigella phage SSE1 | Shigella phage SSE1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169744 | 37.546 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| MK673511 | Salmonella phage ZCSE2 | Salmonella phage ZCSE2 Salmonella virus ZCSE2 Loughboroughvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53965 | 45.843 | Loughboroughvirus | Unclassified | Myoviridae | Salmonella |
| MK584893 | Staphylococcus phage vBSM-A1 | Staphylococcus phage vBSM-A1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140654 | 30.325 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK721198 | Enterococcus phage vB_EfaP_Ef7.4 | Enterococcus phage vB_EfaP_Ef7.4 Enterococcus virus Ef74 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18415 | 33.098 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721197 | Enterococcus phage vB_EfaP_Efmus2 | Enterococcus phage vB_EfaP_Efmus2 unclassified Copernicusvirus Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18366 | 33.246 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721196 | Enterococcus phage vB_EfaP_Ef6.3 | Enterococcus phage vB_EfaP_Ef6.3 Enterococcus virus Ef63 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18136 | 32.637 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721195 | Enterococcus phage vB_EfaP_Efmus1 | Enterococcus phage vB_EfaP_Efmus1 Enterococcus virus Efmus1 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17927 | 33.542 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721194 | Enterococcus phage vB_EfaS_Ef7.1 | Enterococcus phage vB_EfaS_Ef7.1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58018 | 40.032 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MK721193 | Enterococcus phage vB_EfaP_Efmus4 | Enterococcus phage vB_EfaP_Efmus4 Enterococcus virus Efmus4 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18186 | 33.427 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721190 | Enterococcus phage vB_EfaS_Ef6.4 | Enterococcus phage vB_EfaS_Ef6.4 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41133 | 34.542 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK721189 | Enterococcus phage vB_EfaS_Ef2.2 | Enterococcus phage vB_EfaS_Ef2.2 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58400 | 39.92 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MK721188 | Enterococcus phage vB_EfaP_Ef6.2 | Enterococcus phage vB_EfaP_Ef6.2 Enterococcus virus Ef62 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17966 | 32.823 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721187 | Enterococcus phage vB_EfaS_Ef6.1 | Enterococcus phage vB_EfaS_Ef6.1 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40429 | 34.958 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK721185 | Enterococcus phage vB_EfaP_Efmus3 | Enterococcus phage vB_EfaP_Efmus3 Enterococcus virus Efmus3 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18286 | 33.337 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721184 | Enterococcus phage vB_EfaP_Ef7.3 | Enterococcus phage vB_EfaP_Ef7.3 Enterococcus virus Ef73 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18818 | 32.952 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK721183 | Enterococcus phage vB_EfaP_Ef7.2 | Enterococcus phage vB_EfaP_Ef7.2 Enterococcus virus Ef72 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18737 | 33.159 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK685668 | Shigella phage VB_Ship_A7 | Shigella phage VB_Ship_A7 Shigella virus A7 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40058 | 48.405 | Berlinvirus | Studiervirinae | Autographiviridae | Shigella |
| MK560763 | Virgibacillus phage Mimir87 | Virgibacillus phage Mimir87 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48016 | 36.859 | Unclassified | Unclassified | Siphoviridae | Virgibacillus |
| MG649967 | Vibrio phage Thalassa | Vibrio phage Thalassa Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128602 | 40.24 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MG649966 | Vibrio phage Ceto | Vibrio phage Ceto Cetovirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128241 | 39.927 | Cetovirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| MF153391 | Escherichia phage ST20 | Escherichia phage ST20 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44517 | 50.808 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MH051333 | Escherichia phage vB_EcoM-Ro121c4YLVW | Escherichia phage vB_EcoM-Ro121c4YLVW unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151671 | 39.099 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MK075006 | Staphylococcus phage phiSP119-3 | Staphylococcus phage phiSP119-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44311 | 35.625 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| MK075005 | Staphylococcus phage phiSP119-2 | Staphylococcus phage phiSP119-2 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40011 | 34.091 | Unclassified | Trabyvirinae | Siphoviridae | Staphylococcus |
| MK075004 | Staphylococcus phage phiSP119-1 | Staphylococcus phage phiSP119-1 unclassified Fibralongavirus Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49795 | 36.421 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| MK075003 | Staphylococcus phage phiSP44-1 | Staphylococcus phage phiSP44-1 Staphylococcus virus phiSP44-1 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42565 | 35.9 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MK075001 | Staphylococcus phage phiSP15-1 | Staphylococcus phage phiSP15-1 unclassified Fibralongavirus Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45164 | 35.469 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| AB797215 | Klebsiella virus 0507KN21 | Klebsiella virus 0507KN21 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159991 | 46.705 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| KY000079 | Acinetobacter phage AM24 | Acinetobacter phage AM24 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97177 | 37.258 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MK310183 | Phage NC-B | Phage NC-B unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76641 | 42.077 | Kuravirus | Unclassified | Podoviridae | Unspecified |
| MK570225 | Enterococcus phage vB_EfaS_PHB08 | Enterococcus phage vB_EfaS_PHB08 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55244 | 40.055 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MK577502 | Aeromonas phage Asfd_1 | Aeromonas phage Asfd_1 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168962 | 42.343 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| MK599315 | Pseudomonas phage PA1C | Pseudomonas phage PA1C unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 304671 | 43.626 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MK449012 | Streptococcus phage Javan94 | Streptococcus phage Javan94 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34293 | 38.757 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449011 | Streptococcus phage Javan92 | Streptococcus phage Javan92 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37554 | 43.543 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449010 | Streptococcus phage Javan90 | Streptococcus phage Javan90 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37029 | 38.572 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449009 | Streptococcus phage Javan88 | Streptococcus phage Javan88 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40047 | 38.293 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449008 | Streptococcus phage Javan84 | Streptococcus phage Javan84 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43572 | 41.21 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449007 | Streptococcus phage Javan8 | Streptococcus phage Javan8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34464 | 43.782 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449006 | Streptococcus phage Javan78 | Streptococcus phage Javan78 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36493 | 37.818 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449005 | Streptococcus phage Javan74 | Streptococcus phage Javan74 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40231 | 40.026 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449004 | Streptococcus phage Javan68 | Streptococcus phage Javan68 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36400 | 40.739 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449003 | Streptococcus phage Javan648 | Streptococcus phage Javan648 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40810 | 33.842 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449002 | Streptococcus phage Javan642 | Streptococcus phage Javan642 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39641 | 33.945 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449001 | Streptococcus phage Javan640 | Streptococcus phage Javan640 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36232 | 35.927 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK449000 | Streptococcus phage Javan64 | Streptococcus phage Javan64 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40738 | 38.146 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448999 | Streptococcus phage Javan638 | Streptococcus phage Javan638 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35105 | 43.35 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448998 | Streptococcus phage Javan636 | Streptococcus phage Javan636 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42341 | 36.667 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448997 | Streptococcus phage Javan630 | Streptococcus phage Javan630 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48058 | 40.563 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448996 | Streptococcus phage Javan628 | Streptococcus phage Javan628 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37871 | 37.406 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448995 | Streptococcus phage Javan626 | Streptococcus phage Javan626 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41682 | 39.991 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448994 | Streptococcus phage Javan618 | Streptococcus phage Javan618 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39286 | 40.796 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448993 | Streptococcus phage Javan616 | Streptococcus phage Javan616 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40339 | 37.512 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448992 | Streptococcus phage Javan6 | Streptococcus phage Javan6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33800 | 40.269 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448991 | Streptococcus phage Javan590 | Streptococcus phage Javan590 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33345 | 41.083 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448990 | Streptococcus phage Javan584 | Streptococcus phage Javan584 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42506 | 39.999 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448989 | Streptococcus phage Javan580 | Streptococcus phage Javan580 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20214 | 43.485 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448988 | Streptococcus phage Javan578 | Streptococcus phage Javan578 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36699 | 41.214 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448987 | Streptococcus phage Javan576 | Streptococcus phage Javan576 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44767 | 40.898 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448986 | Streptococcus phage Javan570 | Streptococcus phage Javan570 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37934 | 41.224 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448985 | Streptococcus phage Javan566 | Streptococcus phage Javan566 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36044 | 40.756 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448984 | Streptococcus phage Javan562 | Streptococcus phage Javan562 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35422 | 40.54 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448983 | Streptococcus phage Javan558 | Streptococcus phage Javan558 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39898 | 41.142 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448982 | Streptococcus phage Javan554 | Streptococcus phage Javan554 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37127 | 40.782 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448981 | Streptococcus phage Javan548 | Streptococcus phage Javan548 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32773 | 41.842 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448980 | Streptococcus phage Javan536 | Streptococcus phage Javan536 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40798 | 41.509 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK448979 | Streptococcus phage Javan534 | Streptococcus phage Javan534 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39888 | 42.429 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448978 | Streptococcus phage Javan532 | Streptococcus phage Javan532 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39587 | 37.707 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448977 | Streptococcus phage Javan530 | Streptococcus phage Javan530 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41512 | 38.627 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448976 | Streptococcus phage Javan528 | Streptococcus phage Javan528 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37381 | 39.442 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448975 | Streptococcus phage Javan526 | Streptococcus phage Javan526 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41107 | 38.227 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448974 | Streptococcus phage Javan524 | Streptococcus phage Javan524 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40618 | 39.785 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448973 | Streptococcus phage Javan522 | Streptococcus phage Javan522 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40673 | 38.293 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448972 | Streptococcus phage Javan520 | Streptococcus phage Javan520 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35057 | 37.75 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448971 | Streptococcus phage Javan52 | Streptococcus phage Javan52 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44039 | 36.168 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448970 | Streptococcus phage Javan516 | Streptococcus phage Javan516 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45960 | 37.221 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448969 | Streptococcus phage Javan514 | Streptococcus phage Javan514 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33123 | 37.726 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448968 | Streptococcus phage Javan512 | Streptococcus phage Javan512 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39734 | 39.769 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448967 | Streptococcus phage Javan510 | Streptococcus phage Javan510 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37969 | 38.142 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448966 | Streptococcus phage Javan508 | Streptococcus phage Javan508 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40757 | 38.295 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448965 | Streptococcus phage Javan506 | Streptococcus phage Javan506 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40925 | 39.238 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448964 | Streptococcus phage Javan502 | Streptococcus phage Javan502 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35302 | 38.094 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448963 | Streptococcus phage Javan498 | Streptococcus phage Javan498 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31625 | 38.489 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448962 | Streptococcus phage Javan496 | Streptococcus phage Javan496 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32917 | 38.032 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448961 | Streptococcus phage Javan494 | Streptococcus phage Javan494 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39352 | 39.319 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448960 | Streptococcus phage Javan492 | Streptococcus phage Javan492 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41080 | 38.668 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448959 | Streptococcus phage Javan488 | Streptococcus phage Javan488 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40527 | 39.398 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448958 | Streptococcus phage Javan486 | Streptococcus phage Javan486 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41286 | 38.989 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448957 | Streptococcus phage Javan484 | Streptococcus phage Javan484 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46761 | 37.358 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448956 | Streptococcus phage Javan48 | Streptococcus phage Javan48 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41305 | 36.243 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448955 | Streptococcus phage Javan478 | Streptococcus phage Javan478 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41634 | 37.698 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448954 | Streptococcus phage Javan476 | Streptococcus phage Javan476 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40953 | 38.598 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448953 | Streptococcus phage Javan474 | Streptococcus phage Javan474 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43679 | 37.544 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448952 | Streptococcus phage Javan472 | Streptococcus phage Javan472 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40157 | 39.251 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448951 | Streptococcus phage Javan470 | Streptococcus phage Javan470 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38785 | 39.68 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448950 | Streptococcus phage Javan464 | Streptococcus phage Javan464 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42236 | 37.755 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448949 | Streptococcus phage Javan460 | Streptococcus phage Javan460 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41323 | 39.3 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448948 | Streptococcus phage Javan46 | Streptococcus phage Javan46 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38824 | 36.204 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448947 | Streptococcus phage Javan458 | Streptococcus phage Javan458 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40618 | 39.783 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448946 | Streptococcus phage Javan456 | Streptococcus phage Javan456 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44730 | 38.938 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448945 | Streptococcus phage Javan454 | Streptococcus phage Javan454 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39144 | 37.978 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448944 | Streptococcus phage Javan452 | Streptococcus phage Javan452 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43424 | 38.071 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448943 | Streptococcus phage Javan450 | Streptococcus phage Javan450 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34125 | 38.084 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448942 | Streptococcus phage Javan448 | Streptococcus phage Javan448 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25927 | 37.744 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448941 | Streptococcus phage Javan446 | Streptococcus phage Javan446 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45151 | 37.842 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448940 | Streptococcus phage Javan444 | Streptococcus phage Javan444 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36704 | 41.603 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448939 | Streptococcus phage Javan440 | Streptococcus phage Javan440 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41549 | 40.039 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448938 | Streptococcus phage Javan44 | Streptococcus phage Javan44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37765 | 36.801 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448937 | Streptococcus phage Javan436 | Streptococcus phage Javan436 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13056 | 38.725 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448936 | Streptococcus phage Javan426 | Streptococcus phage Javan426 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37936 | 42.603 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448935 | Streptococcus phage Javan424 | Streptococcus phage Javan424 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37709 | 42.632 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448934 | Streptococcus phage Javan422 | Streptococcus phage Javan422 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40394 | 35.327 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448933 | Streptococcus phage Javan420 | Streptococcus phage Javan420 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45151 | 37.503 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448932 | Streptococcus phage Javan42 | Streptococcus phage Javan42 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36054 | 39.824 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448931 | Streptococcus phage Javan414 | Streptococcus phage Javan414 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35546 | 38.001 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448930 | Streptococcus phage Javan406 | Streptococcus phage Javan406 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37950 | 36.377 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448929 | Streptococcus phage Javan400 | Streptococcus phage Javan400 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36561 | 34.944 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448928 | Streptococcus phage Javan40 | Streptococcus phage Javan40 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34653 | 43.396 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448927 | Streptococcus phage Javan394 | Streptococcus phage Javan394 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36484 | 33.905 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448926 | Streptococcus phage Javan392 | Streptococcus phage Javan392 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39795 | 36.097 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448925 | Streptococcus phage Javan386 | Streptococcus phage Javan386 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37026 | 34.762 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448924 | Streptococcus phage Javan384 | Streptococcus phage Javan384 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37542 | 34.99 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448923 | Streptococcus phage Javan380 | Streptococcus phage Javan380 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39487 | 44.645 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448922 | Streptococcus phage Javan38 | Streptococcus phage Javan38 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35085 | 44.167 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448921 | Streptococcus phage Javan374 | Streptococcus phage Javan374 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39974 | 39.013 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448920 | Streptococcus phage Javan372 | Streptococcus phage Javan372 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32918 | 38.681 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448919 | Streptococcus phage Javan370 | Streptococcus phage Javan370 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38574 | 39.35 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448918 | Streptococcus phage Javan366 | Streptococcus phage Javan366 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30862 | 39.625 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448917 | Streptococcus phage Javan362 | Streptococcus phage Javan362 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41015 | 40.093 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448916 | Streptococcus phage Javan36 | Streptococcus phage Javan36 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37764 | 36.8 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448915 | Streptococcus phage Javan350 | Streptococcus phage Javan350 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39079 | 39.594 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448914 | Streptococcus phage Javan348 | Streptococcus phage Javan348 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31633 | 39.576 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448913 | Streptococcus phage Javan346 | Streptococcus phage Javan346 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33554 | 36.303 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448912 | Streptococcus phage Javan340 | Streptococcus phage Javan340 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31480 | 39.581 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448911 | Streptococcus phage Javan34 | Streptococcus phage Javan34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38065 | 36.971 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448910 | Streptococcus phage Javan336 | Streptococcus phage Javan336 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42688 | 40.002 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| MK448909 | Streptococcus phage Javan330 | Streptococcus phage Javan330 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34560 | 40.344 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448908 | Streptococcus phage Javan326 | Streptococcus phage Javan326 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41238 | 40.455 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448907 | Streptococcus phage Javan320 | Streptococcus phage Javan320 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33228 | 36.514 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448906 | Streptococcus phage Javan32 | Streptococcus phage Javan32 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35085 | 44.17 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448905 | Streptococcus phage Javan318 | Streptococcus phage Javan318 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39412 | 38.176 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448904 | Streptococcus phage Javan316 | Streptococcus phage Javan316 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36441 | 40.257 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448903 | Streptococcus phage Javan310 | Streptococcus phage Javan310 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38675 | 39.387 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448902 | Streptococcus phage Javan308 | Streptococcus phage Javan308 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43790 | 39.842 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| MK448901 | Streptococcus phage Javan306 | Streptococcus phage Javan306 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32976 | 40.011 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448900 | Streptococcus phage Javan290 | Streptococcus phage Javan290 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36384 | 42.774 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448899 | Streptococcus phage Javan288 | Streptococcus phage Javan288 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38767 | 38.476 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK448898 | Streptococcus phage Javan284 | Streptococcus phage Javan284 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37997 | 39.024 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448897 | Streptococcus phage Javan28 | Streptococcus phage Javan28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37965 | 35.372 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448896 | Streptococcus phage Javan278 | Streptococcus phage Javan278 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33366 | 37.967 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448895 | Streptococcus phage Javan274 | Streptococcus phage Javan274 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36635 | 35.832 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448894 | Streptococcus phage Javan272 | Streptococcus phage Javan272 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31070 | 37.705 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448893 | Streptococcus phage Javan270 | Streptococcus phage Javan270 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35527 | 38.075 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448892 | Streptococcus phage Javan268 | Streptococcus phage Javan268 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35214 | 42.006 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448891 | Streptococcus phage Javan266 | Streptococcus phage Javan266 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35613 | 40.676 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448890 | Streptococcus phage Javan264 | Streptococcus phage Javan264 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31478 | 38.995 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448889 | Streptococcus phage Javan262 | Streptococcus phage Javan262 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22717 | 38.082 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448888 | Streptococcus phage Javan26 | Streptococcus phage Javan26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33765 | 40.172 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448887 | Streptococcus phage Javan258 | Streptococcus phage Javan258 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39028 | 37.596 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448886 | Streptococcus phage Javan254 | Streptococcus phage Javan254 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38556 | 39.296 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448885 | Streptococcus phage Javan252 | Streptococcus phage Javan252 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39051 | 39.364 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448884 | Streptococcus phage Javan246 | Streptococcus phage Javan246 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38065 | 40.883 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448883 | Streptococcus phage Javan244 | Streptococcus phage Javan244 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35462 | 41.461 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448882 | Streptococcus phage Javan242 | Streptococcus phage Javan242 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33252 | 38.422 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448881 | Streptococcus phage Javan240 | Streptococcus phage Javan240 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33013 | 38.67 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448880 | Streptococcus phage Javan238 | Streptococcus phage Javan238 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27408 | 39.7 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK448879 | Streptococcus phage Javan226 | Streptococcus phage Javan226 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37369 | 37.467 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448878 | Streptococcus phage Javan224 | Streptococcus phage Javan224 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38781 | 38.011 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448877 | Streptococcus phage Javan220 | Streptococcus phage Javan220 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41432 | 39.431 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448876 | Streptococcus phage Javan214 | Streptococcus phage Javan214 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44595 | 36.652 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448875 | Streptococcus phage Javan210 | Streptococcus phage Javan210 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38639 | 39.139 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448874 | Streptococcus phage Javan206 | Streptococcus phage Javan206 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37284 | 38.649 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448873 | Streptococcus phage Javan202 | Streptococcus phage Javan202 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 40.613 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448872 | Streptococcus phage Javan20 | Streptococcus phage Javan20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34653 | 43.393 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448871 | Streptococcus phage Javan196 | Streptococcus phage Javan196 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.281 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448870 | Streptococcus phage Javan192 | Streptococcus phage Javan192 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35580 | 38.449 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448869 | Streptococcus phage Javan190 | Streptococcus phage Javan190 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35580 | 38.449 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448868 | Streptococcus phage Javan188 | Streptococcus phage Javan188 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40415 | 39.275 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448867 | Streptococcus phage Javan186 | Streptococcus phage Javan186 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.276 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448866 | Streptococcus phage Javan184 | Streptococcus phage Javan184 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.281 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448865 | Streptococcus phage Javan182 | Streptococcus phage Javan182 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33810 | 38.74 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448864 | Streptococcus phage Javan180 | Streptococcus phage Javan180 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30782 | 39.036 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448863 | Streptococcus phage Javan18 | Streptococcus phage Javan18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32012 | 40.288 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448862 | Streptococcus phage Javan178 | Streptococcus phage Javan178 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39669 | 40.261 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448861 | Streptococcus phage Javan174 | Streptococcus phage Javan174 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38137 | 41.828 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448860 | Streptococcus phage Javan172 | Streptococcus phage Javan172 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39869 | 36.923 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448859 | Streptococcus phage Javan170 | Streptococcus phage Javan170 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42762 | 38.82 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448858 | Streptococcus phage Javan166 | Streptococcus phage Javan166 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45241 | 38.598 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448857 | Streptococcus phage Javan164 | Streptococcus phage Javan164 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37145 | 37.494 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448856 | Streptococcus phage Javan158 | Streptococcus phage Javan158 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19446 | 38.501 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448855 | Streptococcus phage Javan150 | Streptococcus phage Javan150 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33705 | 39.703 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448854 | Streptococcus phage Javan146 | Streptococcus phage Javan146 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45121 | 37.57 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448853 | Streptococcus phage Javan144 | Streptococcus phage Javan144 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36243 | 38.079 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448852 | Streptococcus phage Javan140 | Streptococcus phage Javan140 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41182 | 39.748 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448851 | Streptococcus phage Javan14 | Streptococcus phage Javan14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43470 | 36.034 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448850 | Streptococcus phage Javan132 | Streptococcus phage Javan132 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43082 | 38.394 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448849 | Streptococcus phage Javan128 | Streptococcus phage Javan128 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40001 | 37.687 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448848 | Streptococcus phage Javan126 | Streptococcus phage Javan126 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37145 | 37.494 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448847 | Streptococcus phage Javan124 | Streptococcus phage Javan124 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38473 | 40.834 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448846 | Streptococcus phage Javan122 | Streptococcus phage Javan122 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39241 | 40.792 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448845 | Streptococcus phage Javan118 | Streptococcus phage Javan118 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37401 | 38.272 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448844 | Streptococcus phage Javan116 | Streptococcus phage Javan116 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28958 | 40.372 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448843 | Streptococcus phage Javan112 | Streptococcus phage Javan112 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54687 | 35.508 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448842 | Streptococcus phage Javan110 | Streptococcus phage Javan110 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31374 | 37.461 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448841 | Streptococcus phage Javan108 | Streptococcus phage Javan108 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54689 | 35.51 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448840 | Streptococcus phage Javan104 | Streptococcus phage Javan104 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37952 | 44.216 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448839 | Streptococcus phage Javan102 | Streptococcus phage Javan102 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54689 | 35.508 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448838 | Streptococcus phage Javan100 | Streptococcus phage Javan100 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33082 | 43.057 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448837 | Streptococcus phage Javan10 | Streptococcus phage Javan10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37275 | 36.99 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448836 | Streptococcus phage Javan95 | Streptococcus phage Javan95 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38797 | 43.403 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448835 | Streptococcus phage Javan93 | Streptococcus phage Javan93 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37554 | 43.543 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448834 | Streptococcus phage Javan91 | Streptococcus phage Javan91 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38343 | 38.429 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448833 | Streptococcus phage Javan87 | Streptococcus phage Javan87 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38593 | 38.766 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448832 | Streptococcus phage Javan85 | Streptococcus phage Javan85 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43572 | 41.21 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448831 | Streptococcus phage Javan83 | Streptococcus phage Javan83 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36493 | 37.818 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448830 | Streptococcus phage Javan73 | Streptococcus phage Javan73 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32459 | 37.9 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448829 | Streptococcus phage Javan7 | Streptococcus phage Javan7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37719 | 36.968 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448828 | Streptococcus phage Javan69 | Streptococcus phage Javan69 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33053 | 41.2 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448827 | Streptococcus phage Javan65 | Streptococcus phage Javan65 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35085 | 44.17 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448826 | Streptococcus phage Javan645 | Streptococcus phage Javan645 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36232 | 35.927 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448825 | Streptococcus phage Javan639 | Streptococcus phage Javan639 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38797 | 43.403 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448824 | Streptococcus phage Javan637 | Streptococcus phage Javan637 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39686 | 33.944 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448823 | Streptococcus phage Javan633 | Streptococcus phage Javan633 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37871 | 37.406 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448822 | Streptococcus phage Javan629 | Streptococcus phage Javan629 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39712 | 36.344 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448821 | Streptococcus phage Javan623 | Streptococcus phage Javan623 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38640 | 35.057 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448820 | Streptococcus phage Javan617 | Streptococcus phage Javan617 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33966 | 35.062 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448819 | Streptococcus phage Javan599 | Streptococcus phage Javan599 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39405 | 41.32 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448818 | Streptococcus phage Javan597 | Streptococcus phage Javan597 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40915 | 40.626 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448817 | Streptococcus phage Javan59 | Streptococcus phage Javan59 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42741 | 38.937 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448816 | Streptococcus phage Javan589 | Streptococcus phage Javan589 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34523 | 40.64 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448815 | Streptococcus phage Javan585 | Streptococcus phage Javan585 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31965 | 41.411 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448814 | Streptococcus phage Javan583 | Streptococcus phage Javan583 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33393 | 41.724 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448813 | Streptococcus phage Javan579 | Streptococcus phage Javan579 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19142 | 39.949 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448812 | Streptococcus phage Javan577 | Streptococcus phage Javan577 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40914 | 40.216 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448811 | Streptococcus phage Javan575 | Streptococcus phage Javan575 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41571 | 40.968 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448810 | Streptococcus phage Javan573 | Streptococcus phage Javan573 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38506 | 41.76 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448809 | Streptococcus phage Javan57 | Streptococcus phage Javan57 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34653 | 43.393 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448808 | Streptococcus phage Javan569 | Streptococcus phage Javan569 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33242 | 41.066 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448807 | Streptococcus phage Javan567 | Streptococcus phage Javan567 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33345 | 41.074 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448806 | Streptococcus phage Javan565 | Streptococcus phage Javan565 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41469 | 40.365 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448805 | Streptococcus phage Javan563 | Streptococcus phage Javan563 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33344 | 41.075 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448804 | Streptococcus phage Javan553 | Streptococcus phage Javan553 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30807 | 41.575 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448803 | Streptococcus phage Javan551 | Streptococcus phage Javan551 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32773 | 41.833 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448802 | Streptococcus phage Javan55 | Streptococcus phage Javan55 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34653 | 43.399 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448801 | Streptococcus phage Javan549 | Streptococcus phage Javan549 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37540 | 41.137 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448800 | Streptococcus phage Javan539 | Streptococcus phage Javan539 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39889 | 42.428 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448799 | Streptococcus phage Javan535 | Streptococcus phage Javan535 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39888 | 42.429 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448798 | Streptococcus phage Javan533 | Streptococcus phage Javan533 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37968 | 38.14 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448797 | Streptococcus phage Javan531 | Streptococcus phage Javan531 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39734 | 39.744 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448796 | Streptococcus phage Javan53 | Streptococcus phage Javan53 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47880 | 36.652 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448795 | Streptococcus phage Javan527 | Streptococcus phage Javan527 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47290 | 38.042 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448794 | Streptococcus phage Javan525 | Streptococcus phage Javan525 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39016 | 38.205 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448793 | Streptococcus phage Javan523 | Streptococcus phage Javan523 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41530 | 38.507 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448792 | Streptococcus phage Javan521 | Streptococcus phage Javan521 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38833 | 37.695 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448791 | Streptococcus phage Javan517 | Streptococcus phage Javan517 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42031 | 38.405 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448790 | Streptococcus phage Javan515 | Streptococcus phage Javan515 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37744 | 38.07 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448789 | Streptococcus phage Javan513 | Streptococcus phage Javan513 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40617 | 39.779 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448788 | Streptococcus phage Javan511 | Streptococcus phage Javan511 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41512 | 38.625 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448787 | Streptococcus phage Javan51 | Streptococcus phage Javan51 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34465 | 43.778 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448786 | Streptococcus phage Javan509 | Streptococcus phage Javan509 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34125 | 38.086 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448785 | Streptococcus phage Javan507 | Streptococcus phage Javan507 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39345 | 37.824 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448784 | Streptococcus phage Javan505 | Streptococcus phage Javan505 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40618 | 39.78 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448783 | Streptococcus phage Javan503 | Streptococcus phage Javan503 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37606 | 38.292 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448782 | Streptococcus phage Javan501 | Streptococcus phage Javan501 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43457 | 38.074 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448781 | Streptococcus phage Javan5 | Streptococcus phage Javan5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40382 | 36.811 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448780 | Streptococcus phage Javan499 | Streptococcus phage Javan499 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36676 | 41.171 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448779 | Streptococcus phage Javan497 | Streptococcus phage Javan497 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39626 | 39.772 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448778 | Streptococcus phage Javan493 | Streptococcus phage Javan493 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40316 | 38.288 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448777 | Streptococcus phage Javan491 | Streptococcus phage Javan491 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35709 | 37.999 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448776 | Streptococcus phage Javan49 | Streptococcus phage Javan49 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34479 | 43.806 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448775 | Streptococcus phage Javan489 | Streptococcus phage Javan489 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41958 | 38.088 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448774 | Streptococcus phage Javan487 | Streptococcus phage Javan487 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33236 | 38.103 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448773 | Streptococcus phage Javan485 | Streptococcus phage Javan485 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36008 | 37.805 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448772 | Streptococcus phage Javan483 | Streptococcus phage Javan483 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41000 | 38.107 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448771 | Streptococcus phage Javan481 | Streptococcus phage Javan481 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35002 | 38.186 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448770 | Streptococcus phage Javan477 | Streptococcus phage Javan477 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38834 | 37.714 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448769 | Streptococcus phage Javan471 | Streptococcus phage Javan471 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38056 | 36.042 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448768 | Streptococcus phage Javan47 | Streptococcus phage Javan47 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41191 | 36.401 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448767 | Streptococcus phage Javan467 | Streptococcus phage Javan467 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34125 | 38.084 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448766 | Streptococcus phage Javan465 | Streptococcus phage Javan465 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38523 | 39.748 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448765 | Streptococcus phage Javan459 | Streptococcus phage Javan459 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39014 | 38.209 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448764 | Streptococcus phage Javan457 | Streptococcus phage Javan457 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30892 | 39.127 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448763 | Streptococcus phage Javan451 | Streptococcus phage Javan451 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33798 | 38.641 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448762 | Streptococcus phage Javan447 | Streptococcus phage Javan447 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41213 | 38.687 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448761 | Streptococcus phage Javan445 | Streptococcus phage Javan445 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40136 | 36.64 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448760 | Streptococcus phage Javan443 | Streptococcus phage Javan443 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40136 | 36.638 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448759 | Streptococcus phage Javan439 | Streptococcus phage Javan439 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41549 | 40.039 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448758 | Streptococcus phage Javan433 | Streptococcus phage Javan433 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41549 | 40.039 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448757 | Streptococcus phage Javan427 | Streptococcus phage Javan427 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42333 | 39.905 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448756 | Streptococcus phage Javan425 | Streptococcus phage Javan425 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37660 | 36.455 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448755 | Streptococcus phage Javan423 | Streptococcus phage Javan423 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37660 | 36.455 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448754 | Streptococcus phage Javan411 | Streptococcus phage Javan411 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36561 | 34.944 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448753 | Streptococcus phage Javan407 | Streptococcus phage Javan407 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38109 | 35.335 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448752 | Streptococcus phage Javan405 | Streptococcus phage Javan405 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38222 | 42.271 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448751 | Streptococcus phage Javan399 | Streptococcus phage Javan399 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38443 | 35.185 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK448750 | Streptococcus phage Javan393 | Streptococcus phage Javan393 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42146 | 35.6 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448749 | Streptococcus phage Javan39 | Streptococcus phage Javan39 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43433 | 36.746 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448748 | Streptococcus phage Javan389 | Streptococcus phage Javan389 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39475 | 36.147 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448747 | Streptococcus phage Javan385 | Streptococcus phage Javan385 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42560 | 35.738 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448746 | Streptococcus phage Javan383 | Streptococcus phage Javan383 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37490 | 35.639 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448745 | Streptococcus phage Javan381 | Streptococcus phage Javan381 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39487 | 44.645 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448744 | Streptococcus phage Javan377 | Streptococcus phage Javan377 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39487 | 44.645 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448743 | Streptococcus phage Javan371 | Streptococcus phage Javan371 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39366 | 39.25 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448742 | Streptococcus phage Javan37 | Streptococcus phage Javan37 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37128 | 36.851 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448741 | Streptococcus phage Javan369 | Streptococcus phage Javan369 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37447 | 41.154 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448740 | Streptococcus phage Javan367 | Streptococcus phage Javan367 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40055 | 40.3 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448739 | Streptococcus phage Javan363 | Streptococcus phage Javan363 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29384 | 39.681 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448738 | Streptococcus phage Javan355 | Streptococcus phage Javan355 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39310 | 41.051 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448737 | Streptococcus phage Javan351 | Streptococcus phage Javan351 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37175 | 39.871 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448736 | Streptococcus phage Javan35 | Streptococcus phage Javan35 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37764 | 36.794 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448735 | Streptococcus phage Javan349 | Streptococcus phage Javan349 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37433 | 40.112 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448734 | Streptococcus phage Javan347 | Streptococcus phage Javan347 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30906 | 38.912 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448733 | Streptococcus phage Javan345 | Streptococcus phage Javan345 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37444 | 41.665 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448732 | Streptococcus phage Javan331 | Streptococcus phage Javan331 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35176 | 39.982 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448731 | Streptococcus phage Javan33 | Streptococcus phage Javan33 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35793 | 40.078 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448730 | Streptococcus phage Javan311 | Streptococcus phage Javan311 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40399 | 45.142 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448729 | Streptococcus phage Javan31 | Streptococcus phage Javan31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39484 | 36.134 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448728 | Streptococcus phage Javan3 | Streptococcus phage Javan3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34657 | 43.397 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448727 | Streptococcus phage Javan291 | Streptococcus phage Javan291 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39605 | 43.176 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448726 | Streptococcus phage Javan29 | Streptococcus phage Javan29 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34545 | 43.769 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448725 | Streptococcus phage Javan275 | Streptococcus phage Javan275 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30398 | 37.756 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448724 | Streptococcus phage Javan273 | Streptococcus phage Javan273 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37618 | 37.065 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448723 | Streptococcus phage Javan271 | Streptococcus phage Javan271 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40704 | 35.478 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448722 | Streptococcus phage Javan269 | Streptococcus phage Javan269 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36580 | 36.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448721 | Streptococcus phage Javan263 | Streptococcus phage Javan263 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34637 | 37.723 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448720 | Streptococcus phage Javan261 | Streptococcus phage Javan261 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25306 | 38.635 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448719 | Streptococcus phage Javan255 | Streptococcus phage Javan255 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39881 | 42.266 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448718 | Streptococcus phage Javan253 | Streptococcus phage Javan253 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 40.129 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448717 | Streptococcus phage Javan25 | Streptococcus phage Javan25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35085 | 44.173 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448716 | Streptococcus phage Javan249 | Streptococcus phage Javan249 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41477 | 38.115 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448715 | Streptococcus phage Javan247 | Streptococcus phage Javan247 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35684 | 41.374 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448714 | Streptococcus phage Javan241 | Streptococcus phage Javan241 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36502 | 38.255 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448713 | Streptococcus phage Javan239 | Streptococcus phage Javan239 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36677 | 41.167 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448712 | Streptococcus phage Javan237 | Streptococcus phage Javan237 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27408 | 39.7 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK448711 | Streptococcus phage Javan235 | Streptococcus phage Javan235 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45029 | 37.667 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448710 | Streptococcus phage Javan233 | Streptococcus phage Javan233 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39808 | 37.402 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448709 | Streptococcus phage Javan231 | Streptococcus phage Javan231 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39808 | 37.402 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448708 | Streptococcus phage Javan23 | Streptococcus phage Javan23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37964 | 35.376 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448707 | Streptococcus phage Javan227 | Streptococcus phage Javan227 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42048 | 37.055 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448706 | Streptococcus phage Javan221 | Streptococcus phage Javan221 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35845 | 39.35 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448705 | Streptococcus phage Javan215 | Streptococcus phage Javan215 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40164 | 40.955 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448704 | Streptococcus phage Javan213 | Streptococcus phage Javan213 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40311 | 40.485 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448703 | Streptococcus phage Javan207 | Streptococcus phage Javan207 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45067 | 36.854 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448702 | Streptococcus phage Javan199 | Streptococcus phage Javan199 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41831 | 38.471 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448701 | Streptococcus phage Javan197 | Streptococcus phage Javan197 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34770 | 38.605 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448700 | Streptococcus phage Javan191 | Streptococcus phage Javan191 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44570 | 38.236 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448699 | Streptococcus phage Javan189 | Streptococcus phage Javan189 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.278 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448698 | Streptococcus phage Javan187 | Streptococcus phage Javan187 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28059 | 35.251 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448697 | Streptococcus phage Javan185 | Streptococcus phage Javan185 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.274 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448696 | Streptococcus phage Javan183 | Streptococcus phage Javan183 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.278 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448695 | Streptococcus phage Javan181 | Streptococcus phage Javan181 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33542 | 42.833 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448694 | Streptococcus phage Javan179 | Streptococcus phage Javan179 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41483 | 39.312 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448693 | Streptococcus phage Javan177 | Streptococcus phage Javan177 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 39.276 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448692 | Streptococcus phage Javan173 | Streptococcus phage Javan173 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44662 | 44.176 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448691 | Streptococcus phage Javan17 | Streptococcus phage Javan17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39484 | 36.141 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448690 | Streptococcus phage Javan169 | Streptococcus phage Javan169 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44153 | 38.5 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448689 | Streptococcus phage Javan163 | Streptococcus phage Javan163 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40438 | 39.789 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448688 | Streptococcus phage Javan161 | Streptococcus phage Javan161 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42720 | 38.811 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448687 | Streptococcus phage Javan159 | Streptococcus phage Javan159 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36052 | 41.179 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448686 | Streptococcus phage Javan157 | Streptococcus phage Javan157 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40260 | 39.709 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448685 | Streptococcus phage Javan155 | Streptococcus phage Javan155 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36243 | 38.079 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448684 | Streptococcus phage Javan151 | Streptococcus phage Javan151 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35995 | 36.236 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448683 | Streptococcus phage Javan149 | Streptococcus phage Javan149 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41754 | 37.594 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448682 | Streptococcus phage Javan143 | Streptococcus phage Javan143 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37721 | 38.209 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448681 | Streptococcus phage Javan141 | Streptococcus phage Javan141 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42448 | 38.002 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448680 | Streptococcus phage Javan139 | Streptococcus phage Javan139 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35722 | 36.84 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448679 | Streptococcus phage Javan137 | Streptococcus phage Javan137 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37335 | 37.557 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448678 | Streptococcus phage Javan135 | Streptococcus phage Javan135 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39497 | 39.889 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448677 | Streptococcus phage Javan133 | Streptococcus phage Javan133 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38145 | 37.373 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| MK448676 | Streptococcus phage Javan131 | Streptococcus phage Javan131 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44876 | 38.649 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448675 | Streptococcus phage Javan129 | Streptococcus phage Javan129 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42310 | 38.814 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448674 | Streptococcus phage Javan123 | Streptococcus phage Javan123 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37428 | 41.589 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448673 | Streptococcus phage Javan119 | Streptococcus phage Javan119 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48294 | 38.353 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448672 | Streptococcus phage Javan117 | Streptococcus phage Javan117 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40234 | 37.744 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448671 | Streptococcus phage Javan115 | Streptococcus phage Javan115 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38959 | 39.801 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448670 | Streptococcus phage Javan113 | Streptococcus phage Javan113 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37901 | 44.191 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448669 | Streptococcus phage Javan11 | Streptococcus phage Javan11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36754 | 39.745 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448668 | Streptococcus phage Javan107 | Streptococcus phage Javan107 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37952 | 44.216 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448667 | Streptococcus phage Javan105 | Streptococcus phage Javan105 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54689 | 35.51 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448666 | Streptococcus phage Javan101 | Streptococcus phage Javan101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37952 | 44.216 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK448665 | Streptococcus phage Javan1 | Streptococcus phage Javan1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38191 | 40.098 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK552140 | Burkholderia phage BcepSaruman | Burkholderia phage BcepSaruman Burkholderia virus BcepSaruman Sarumanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 263735 | 58.139 | Sarumanvirus | Unclassified | Myoviridae | Burkholderia |
| KR347354 | Propionibacterium phage Moyashi | Propionibacterium phage Moyashi unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29254 | 54.587 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR347353 | Propionibacterium phage Enoki | Propionibacterium phage Enoki unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29347 | 54.588 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR347352 | Propionibacterium phage BruceLethal | Propionibacterium phage BruceLethal unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29249 | 54.039 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337654 | Propionibacterium phage Ouroboros | Propionibacterium phage Ouroboros Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29506 | 53.928 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337652 | Propionibacterium phage Stormborn | Propionibacterium phage Stormborn Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29330 | 53.802 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337651 | Propionibacterium phage Attacne | Propionibacterium phage Attacne Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28876 | 54.706 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337650 | Propionibacterium phage Lauchelly | Propionibacterium phage Lauchelly Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29517 | 53.86 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337649 | Propionibacterium phage Keiki | Propionibacterium phage Keiki Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29339 | 54.027 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337648 | Propionibacterium phage SKKY | Propionibacterium phage SKKY Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29594 | 54.599 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337647 | Propionibacterium phage Solid | Propionibacterium phage Solid Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29440 | 53.916 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337646 | Propionibacterium phage Wizzo | Propionibacterium phage Wizzo Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29463 | 54.441 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337645 | Propionibacterium phage Kubed | Propionibacterium phage Kubed Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29461 | 54.526 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337644 | Propionibacterium phage Procrass1 | Propionibacterium phage Procrass1 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29347 | 54.04 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR337643 | Propionibacterium phage MrAK | Propionibacterium phage MrAK Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29726 | 54.417 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MK483107 | Vibrio phage vB_VpP_BA6 | Vibrio phage vB_VpP_BA6 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50520 | 41.766 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MH051335 | Escherichia phage vB_EcoM-Ro157lw | Escherichia phage vB_EcoM-Ro157lw unclassified Wifcevirus Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72179 | 46.013 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MH571750 | Escherichia phage vB_EcoM-Ro111lw | Escherichia phage vB_EcoM-Ro111lw unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86950 | 38.837 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MH160766 | Escherichia phage vB_EcoM-Ro121lw | Escherichia phage vB_EcoM-Ro121lw unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149803 | 39.08 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MH051334 | Escherichia phage vB_EcoS-Ro145c2YLVW | Escherichia phage vB_EcoS-Ro145c2YLVW unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42145 | 50.566 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK310226 | Halobacterium virus ChaoS9 | Halobacterium virus ChaoS9 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55145 | 65.273 | Myohalovirus | Unclassified | Myoviridae | Halobacterium |
| AP019562 | Staphylococcus phage SP276 | Staphylococcus phage SP276 Staphylococcus virus SP276 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40147 | 35.353 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| AP019561 | Staphylococcus phage SP197 | Staphylococcus phage SP197 Staphylococcus virus SP197 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 35.673 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| AP019560 | Staphylococcus phage SP120 | Staphylococcus phage SP120 Staphylococcus virus SP120 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40530 | 34.991 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| AP019559 | Escherichia phage SP15 | Escherichia phage SP15 Escherichia virus SP15 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110964 | 39.134 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK511066 | Pseudomonas phage vB_Pae_BR319a | Pseudomonas phage vB_Pae_BR319a unclassified Citexvirus Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35343 | 62.42 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| MK511065 | Pseudomonas phage vB_Pae_BR141c | Pseudomonas phage vB_Pae_BR141c unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39730 | 63.501 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511064 | Pseudomonas phage vB_Pae_BR58a | Pseudomonas phage vB_Pae_BR58a unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29041 | 63.328 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511063 | Pseudomonas phage vB_Pae_CF177a | Pseudomonas phage vB_Pae_CF177a unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37399 | 63.649 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511061 | Pseudomonas phage vB_Pae_CF125a | Pseudomonas phage vB_Pae_CF125a unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39659 | 63.506 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511060 | Pseudomonas phage vB_Pae_CF118c | Pseudomonas phage vB_Pae_CF118c unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32181 | 63.407 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511059 | Pseudomonas phage vB_Pae_CF78a | Pseudomonas phage vB_Pae_CF78a unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39705 | 63.523 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511058 | Pseudomonas phage vB_Pae_CF53b | Pseudomonas phage vB_Pae_CF53b unclassified Beetrevirus Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28078 | 63.334 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511057 | Pseudomonas phage vB_Pae_CF5a | Pseudomonas phage vB_Pae_CF5a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36361 | 62.905 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511051 | Pseudomonas phage vB_Pae_BR181a | Pseudomonas phage vB_Pae_BR181a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511050 | Pseudomonas phage vB_Pae_BR150a | Pseudomonas phage vB_Pae_BR150a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511049 | Pseudomonas phage vB_Pae_BR326a | Pseudomonas phage vB_Pae_BR326a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511048 | Pseudomonas phage vB_Pae_BR320a | Pseudomonas phage vB_Pae_BR320a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511047 | Pseudomonas phage vB_Pae_BR319b | Pseudomonas phage vB_Pae_BR319b unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61853 | 63.67 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511046 | Pseudomonas phage vB_Pae_BR313a | Pseudomonas phage vB_Pae_BR313a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.662 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511045 | Pseudomonas phage vB_Pae_BR208a | Pseudomonas phage vB_Pae_BR208a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511044 | Pseudomonas phage vB_Pae_BR197a | Pseudomonas phage vB_Pae_BR197a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511043 | Pseudomonas phage vB_Pae_BR228a | Pseudomonas phage vB_Pae_BR228a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61684 | 63.662 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511042 | Pseudomonas phage vB_Pae_CF126a | Pseudomonas phage vB_Pae_CF126a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511041 | Pseudomonas phage vB_Pae_CF69a | Pseudomonas phage vB_Pae_CF69a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511040 | Pseudomonas phage vB_Pae_CF63a | Pseudomonas phage vB_Pae_CF63a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62695 | 63.683 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511039 | Pseudomonas phage vB_Pae_CF24a | Pseudomonas phage vB_Pae_CF24a unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61772 | 63.665 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MK511038 | Pseudomonas phage vB_Pae_BR58b | Pseudomonas phage vB_Pae_BR58b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38471 | 64.425 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511037 | Pseudomonas phage vB_Pae_BR141d | Pseudomonas phage vB_Pae_BR141d unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27510 | 64.42 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511036 | Pseudomonas phage vB_Pae_BR327a | Pseudomonas phage vB_Pae_BR327a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37474 | 64.156 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511035 | Pseudomonas phage vB_Pae_BR313b | Pseudomonas phage vB_Pae_BR313b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37758 | 64.349 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511034 | Pseudomonas phage vB_Pae_BR243b | Pseudomonas phage vB_Pae_BR243b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38215 | 64.192 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511033 | Pseudomonas phage vB_Pae_BR205a | Pseudomonas phage vB_Pae_BR205a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37715 | 64.322 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511032 | Pseudomonas phage vB_Pae_BR161b | Pseudomonas phage vB_Pae_BR161b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39613 | 64.072 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511031 | Pseudomonas phage vB_Pae_BR52b | Pseudomonas phage vB_Pae_BR52b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31014 | 64.477 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511030 | Pseudomonas phage vB_Pae_CF213b | Pseudomonas phage vB_Pae_CF213b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37615 | 64.309 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511029 | Pseudomonas phage vB_Pae_CF183b | Pseudomonas phage vB_Pae_CF183b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38416 | 64.169 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511028 | Pseudomonas phage vB_Pae_CF177b | Pseudomonas phage vB_Pae_CF177b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37615 | 64.336 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511027 | Pseudomonas phage vB_Pae_CF136a | Pseudomonas phage vB_Pae_CF136a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37654 | 64.344 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511026 | Pseudomonas phage vB_Pae_CF121a | Pseudomonas phage vB_Pae_CF121a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37617 | 64.386 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511025 | Pseudomonas phage vB_Pae_CF118b | Pseudomonas phage vB_Pae_CF118b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37734 | 64.385 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511024 | Pseudomonas phage vB_Pae_CF81b | Pseudomonas phage vB_Pae_CF81b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37636 | 64.313 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511023 | Pseudomonas phage vB_Pae_CF77b | Pseudomonas phage vB_Pae_CF77b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37617 | 64.351 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511022 | Pseudomonas phage vB_Pae_CF74a | Pseudomonas phage vB_Pae_CF74a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44244 | 64.092 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511021 | Pseudomonas phage vB_Pae_CF65b | Pseudomonas phage vB_Pae_CF65b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37629 | 64.331 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511020 | Pseudomonas phage vB_Pae_CF57b | Pseudomonas phage vB_Pae_CF57b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37702 | 64.333 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511019 | Pseudomonas phage vB_Pae_CF53c | Pseudomonas phage vB_Pae_CF53c unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26171 | 64.51 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511018 | Pseudomonas phage vB_Pae_CF28a | Pseudomonas phage vB_Pae_CF28a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37395 | 64.091 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511017 | Pseudomonas phage vB_Pae_CF23b | Pseudomonas phage vB_Pae_CF23b unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37682 | 64.307 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK511016 | Pseudomonas phage vB_Pae_BR141b | Pseudomonas phage vB_Pae_BR141b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45372 | 58.69 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511015 | Pseudomonas phage vB_Pae_BR313c | Pseudomonas phage vB_Pae_BR313c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33113 | 58.992 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511014 | Pseudomonas phage vB_Pae_BR243a | Pseudomonas phage vB_Pae_BR243a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46492 | 59.283 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511013 | Pseudomonas phage vB_Pae_BR161a | Pseudomonas phage vB_Pae_BR161a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42421 | 59.162 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511012 | Pseudomonas phage vB_Pae_BR153a | Pseudomonas phage vB_Pae_BR153a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50013 | 61.806 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511011 | Pseudomonas phage vB_Pae_CF213a | Pseudomonas phage vB_Pae_CF213a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52658 | 58.908 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511010 | Pseudomonas phage vB_Pae_CF177c | Pseudomonas phage vB_Pae_CF177c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52755 | 58.912 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511009 | Pseudomonas phage vB_Pae_CF136b | Pseudomonas phage vB_Pae_CF136b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37242 | 59.24 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511008 | Pseudomonas phage vB_Pae_CF124b | Pseudomonas phage vB_Pae_CF124b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29118 | 58.795 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511007 | Pseudomonas phage vB_Pae_CF121b | Pseudomonas phage vB_Pae_CF121b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42495 | 58.868 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511006 | Pseudomonas phage vB_Pae_CF118a | Pseudomonas phage vB_Pae_CF118a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52758 | 58.916 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511005 | Pseudomonas phage vB_Pae_CF81a | Pseudomonas phage vB_Pae_CF81a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52823 | 58.935 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511004 | Pseudomonas phage vB_Pae_CF74b | Pseudomonas phage vB_Pae_CF74b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34669 | 59.064 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511003 | Pseudomonas phage vB_Pae_CF65a | Pseudomonas phage vB_Pae_CF65a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52662 | 58.917 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511002 | Pseudomonas phage vB_Pae_CF57a | Pseudomonas phage vB_Pae_CF57a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52664 | 58.911 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK511001 | Pseudomonas phage vB_Pae_CF23a | Pseudomonas phage vB_Pae_CF23a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52800 | 58.936 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510993 | Pseudomonas phage vB_Pae_BR201a | Pseudomonas phage vB_Pae_BR201a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38378 | 60.764 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510992 | Pseudomonas phage vB_Pae_BR144a | Pseudomonas phage vB_Pae_BR144a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39861 | 61.381 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510991 | Pseudomonas phage vB_Pae_BR141a | Pseudomonas phage vB_Pae_BR141a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38304 | 61.367 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510990 | Pseudomonas phage vB_Pae_BR133a | Pseudomonas phage vB_Pae_BR133a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41217 | 61.494 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510989 | Pseudomonas phage vB_Pae_BR322a | Pseudomonas phage vB_Pae_BR322a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38344 | 61.347 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510988 | Pseudomonas phage vB_Pae_BR299a | Pseudomonas phage vB_Pae_BR299a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40745 | 61.556 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510987 | Pseudomonas phage vB_Pae_BR233a | Pseudomonas phage vB_Pae_BR233a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40425 | 61.549 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510986 | Pseudomonas phage vB_Pae_BR213a | Pseudomonas phage vB_Pae_BR213a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30609 | 61.6 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510985 | Pseudomonas phage vB_Pae_BR204a | Pseudomonas phage vB_Pae_BR204a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40441 | 61.559 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510984 | Pseudomonas phage vB_Pae_BR200a | Pseudomonas phage vB_Pae_BR200a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38217 | 61.269 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510983 | Pseudomonas phage vB_Pae_BR178a | Pseudomonas phage vB_Pae_BR178a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38345 | 61.348 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510982 | Pseudomonas phage vB_Pae_BR161c | Pseudomonas phage vB_Pae_BR161c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38285 | 61.382 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510981 | Pseudomonas phage vB_Pae_BR143a | Pseudomonas phage vB_Pae_BR143a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38370 | 61.353 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510980 | Pseudomonas phage vB_Pae_BR123a | Pseudomonas phage vB_Pae_BR123a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38344 | 61.347 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510979 | Pseudomonas phage vB_Pae_BR52a | Pseudomonas phage vB_Pae_BR52a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38353 | 61.351 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510978 | Pseudomonas phage vB_Pae_CF208a | Pseudomonas phage vB_Pae_CF208a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40412 | 61.563 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510977 | Pseudomonas phage vB_Pae_CF183a | Pseudomonas phage vB_Pae_CF183a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37635 | 61.36 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510976 | Pseudomonas phage vB_Pae_CF165a | Pseudomonas phage vB_Pae_CF165a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40778 | 61.545 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510975 | Pseudomonas phage vB_Pae_CF145a | Pseudomonas phage vB_Pae_CF145a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40711 | 61.58 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510974 | Pseudomonas phage vB_Pae_CF140a | Pseudomonas phage vB_Pae_CF140a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40393 | 61.565 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510973 | Pseudomonas phage vB_Pae_CF127a | Pseudomonas phage vB_Pae_CF127a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38239 | 61.39 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510972 | Pseudomonas phage vB_Pae_CF121c | Pseudomonas phage vB_Pae_CF121c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38379 | 61.304 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510971 | Pseudomonas phage vB_Pae_CF79a | Pseudomonas phage vB_Pae_CF79a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41155 | 61.599 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510970 | Pseudomonas phage vB_Pae_CF67a | Pseudomonas phage vB_Pae_CF67a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35420 | 61.35 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510969 | Pseudomonas phage vB_Pae_CF60a | Pseudomonas phage vB_Pae_CF60a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35225 | 61.593 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510968 | Pseudomonas phage vB_Pae_CF55b | Pseudomonas phage vB_Pae_CF55b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42511 | 61.492 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510967 | Pseudomonas phage vB_Pae_CF54a | Pseudomonas phage vB_Pae_CF54a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41186 | 61.164 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510966 | Pseudomonas phage vB_Pae_CF53a | Pseudomonas phage vB_Pae_CF53a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38375 | 61.342 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510965 | Pseudomonas phage vB_Pae_CF34a | Pseudomonas phage vB_Pae_CF34a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41462 | 61.524 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510964 | Pseudomonas phage vB_Pae_CF28b | Pseudomonas phage vB_Pae_CF28b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40439 | 61.572 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510963 | Pseudomonas phage vB_Pae_CF24b | Pseudomonas phage vB_Pae_CF24b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40414 | 61.597 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK510962 | Pseudomonas phage vB_Pae_CF3a | Pseudomonas phage vB_Pae_CF3a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33681 | 60.948 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| LC473039 | Escherichia phage L27 | Escherichia phage L27 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86019 | 38.936 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MH427377 | Escherichia phage vB_EcoM_Sa157lw | Escherichia phage vB_EcoM_Sa157lw unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155887 | 44.944 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| MK301443 | Lactococcus phage CW09 | Lactococcus phage CW09 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21677 | 35.941 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MK301442 | Lactococcus phage 20R03M | Lactococcus phage 20R03M Lactococcus virus 20R03M Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21750 | 35.959 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MK301441 | Lactococcus phage MV16 | Lactococcus phage MV16 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32855 | 35.039 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MK301440 | Lactococcus phage MV10L | Lactococcus phage MV10L Lactococcus virus MV10L Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31374 | 34.876 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MK301439 | Lactococcus phage AV09 | Lactococcus phage AV09 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28732 | 35.299 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MK301438 | Lactococcus phage 8V08 | Lactococcus phage 8V08 Lactococcus virus 8V08 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32312 | 35.071 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MK301437 | Lactococcus phage 1W08F | Lactococcus phage 1W08F unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30633 | 34.887 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MK373799 | Escherichia phage vB_EcoS_EASG3 | Escherichia phage vB_EcoS_EASG3 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120715 | 39.037 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK373798 | Escherichia phage vB_EcoS_G29-2 | Escherichia phage vB_EcoS_G29-2 Escherichia virus G29-2 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51739 | 44.017 | Hanrivervirus | Tempevirinae | Drexlerviridae | Escherichia |
| MK373797 | Escherichia phage vB_EcoS_HASG4 | Escherichia phage vB_EcoS_HASG4 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120603 | 39.024 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK373796 | Escherichia phage vB_EcoS_HdH2 | Escherichia phage vB_EcoS_HdH2 Escherichia virus HdH2 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120120 | 39.321 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK373795 | Escherichia phage vB_EcoS_HdSG1 | Escherichia phage vB_EcoS_HdSG1 unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63595 | 43.865 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MK373794 | Escherichia phage vB_EcoS_HdK1 | Escherichia phage vB_EcoS_HdK1 unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64059 | 44.07 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MK373793 | Escherichia phage vB_EcoS_MM01 | Escherichia phage vB_EcoS_MM01 unclassified Rogunavirinae Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43157 | 43.766 | Unclassified | Rogunavirinae | Drexlerviridae | Escherichia |
| MK373792 | Escherichia phage vB_EcoS_VAH1 | Escherichia phage vB_EcoS_VAH1 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124537 | 38.605 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MK373791 | Escherichia phage vB_EcoS_WFI | Escherichia phage vB_EcoS_WFI unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45536 | 54.572 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MK373790 | Escherichia phage vB_EcoS_WF5505 | Escherichia phage vB_EcoS_WF5505 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47082 | 54.301 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MK373789 | Escherichia phage vB_EcoS_PTXU06 | Escherichia phage vB_EcoS_PTXU06 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45904 | 54.442 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MK373788 | Escherichia phage vB_EcoM_Schickermooser | Escherichia phage vB_EcoM_Schickermooser Escherichia virus Schickermooser Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151194 | 39.008 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| MK373787 | Escherichia phage vB_EcoM_G8 | Escherichia phage vB_EcoM_G8 Escherichia virus G8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169650 | 35.326 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373786 | Escherichia phage vB_EcoM_R5505 | Escherichia phage vB_EcoM_R5505 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167317 | 35.362 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373785 | Escherichia phage vB_EcoM_OE5505 | Escherichia phage vB_EcoM_OE5505 Escherichia virus OE5505 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168756 | 35.221 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373784 | Escherichia phage vB_EcoM_MM02 | Escherichia phage vB_EcoM_MM02 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169201 | 37.603 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373783 | Escherichia phage vB_EcoM_KWBSE43-6 | Escherichia phage vB_EcoM_KWBSE43-6 Escherichia virus KWBSE43-6 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158607 | 46.091 | Taipeivirus | Unclassified | Ackermannviridae | Escherichia |
| MK373782 | Escherichia phage vB_EcoM_KAW3E185 | Escherichia phage vB_EcoM_KAW3E185 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170187 | 37.55 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373781 | Escherichia phage vB_EcoM_KAW1E185 | Escherichia phage vB_EcoM_KAW1E185 Escherichia virus KAW1E185 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164987 | 35.368 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373780 | Escherichia phage vB_EcoM_HdK5 | Escherichia phage vB_EcoM_HdK5 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139328 | 43.627 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MK373779 | Escherichia phage vB_EcoM_G9062 | Escherichia phage vB_EcoM_G9062 Escherichia virus G9062 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168670 | 35.296 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373778 | Escherichia phage vB_EcoM_WFbE185 | Escherichia phage vB_EcoM_WFbE185 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170429 | 37.605 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373777 | Escherichia phage vB_EcoM_WFC | Escherichia phage vB_EcoM_WFC Escherichia virus WFC Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72472 | 46.011 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MK373776 | Escherichia phage vB_EcoM_WFH | Escherichia phage vB_EcoM_WFH Escherichia virus WFH Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71283 | 46.118 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MK373775 | Escherichia phage vB_EcoM_WFK | Escherichia phage vB_EcoM_WFK unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164590 | 37.576 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373774 | Escherichia phage vB_EcoM_WFL6982 | Escherichia phage vB_EcoM_WFL6982 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164279 | 37.559 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK373773 | Escherichia phage vB_EcoP_KAW1A4500 | Escherichia phage vB_EcoP_KAW1A4500 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44241 | 45.184 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MK373772 | Escherichia phage PTXU04 | Escherichia phage PTXU04 Escherichia virus PTXU04 Xuquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61600 | 52.844 | Xuquatrovirus | Unclassified | Podoviridae | Escherichia |
| MK373771 | Escherichia phage vB_EcoP_R4596 | Escherichia phage vB_EcoP_R4596 unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45130 | 44.839 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MK373770 | Escherichia phage vB_EcoP_WFI101126 | Escherichia phage vB_EcoP_WFI101126 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77307 | 42.066 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MK482688 | Escherichia phage vB_EcoM_LMP25 | Escherichia phage vB_EcoM_LMP25 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88409 | 38.906 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MK482689 | Escherichia phage vB_EcoM_LMP33 | Escherichia phage vB_EcoM_LMP33 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88409 | 38.911 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MK482690 | Escherichia phage vB_EcoM_LMP34 | Escherichia phage vB_EcoM_LMP34 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84898 | 39.012 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MK493325 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170397 | 40.646 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| MK493324 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169811 | 40.646 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| MK493323 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171770 | 40.58 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| MK493322 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169803 | 40.631 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| MK493321 | Cyanophage S-RIM4 | Cyanophage S-RIM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175462 | 41.196 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MK552141 | Burkholderia phage BcepSauron | Burkholderia phage BcepSauron Burkholderia virus BcepSauron Sarumanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262653 | 58.097 | Sarumanvirus | Unclassified | Myoviridae | Burkholderia |
| MK573636 | Escherichia phage vB_EcoM_PHB13 | Escherichia phage vB_EcoM_PHB13 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165641 | 40.408 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK578530 | Salmonella phage vB_SenS_SB3 | Salmonella phage vB_SenS_SB3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41147 | 50.009 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MK605246 | Nodularia phage vB_NspS-kac68v162 | Nodularia phage vB_NspS-kac68v162 unclassified Ravarandavirus Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144444 | 40.189 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| MK605245 | Nodularia phage vB_NspS-kac68v161 | Nodularia phage vB_NspS-kac68v161 Nodularia virus kac68v161 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145400 | 40.235 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| MK605244 | Nodularia phage vB_NspS-kac65v162 | Nodularia phage vB_NspS-kac65v162 unclassified Ravarandavirus Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147033 | 40.292 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| MK605243 | Nodularia phage vB_NspS-kac65v161 | Nodularia phage vB_NspS-kac65v161 unclassified Ravarandavirus Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146735 | 40.311 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| MK605242 | Nodularia phage vB_NspS-kac65v151 | Nodularia phage vB_NspS-kac65v151 Nodularia virus kac65v151 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146618 | 40.31 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| MK433578 | Klebsiella phage ST15-OXA48phi14.1 | Klebsiella phage ST15-OXA48phi14.1 Klebsiella virus ST15OXA48phi14-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33839 | 52.747 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK422450 | Klebsiella phage ST13-OXA48phi12.4 | Klebsiella phage ST13-OXA48phi12.4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59049 | 52.119 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK422448 | Klebsiella phage ST512-KPC3phi13.3 | Klebsiella phage ST512-KPC3phi13.3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63032 | 54.33 | Unclassified | Unclassified | Podoviridae | Klebsiella |
| MK416010 | Klebsiella phage ST11-OXA245phi3.2 | Klebsiella phage ST11-OXA245phi3.2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60118 | 55.586 | Unclassified | Unclassified | Podoviridae | Klebsiella |
| MK416012 | Klebsiella phage ST437-OXA245phi4.2 | Klebsiella phage ST437-OXA245phi4.2 Klebsiella virus ST437OXA245phi4-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18281 | 50.501 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK675901 | Shewanella phage S0112 | Shewanella phage S0112 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62286 | 44.7 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| MK562505 | Shigella phage Silverhawkium | Shigella phage Silverhawkium unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88821 | 39.071 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MK562504 | Enterobacteria phage CHB7 | Enterobacteria phage CHB7 unclassified Suspvirus Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87998 | 40.049 | Suspvirus | Ounavirinae | Myoviridae | Enterobacteria |
| MK562502 | Shigella phage KPS64 | Shigella phage KPS64 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90227 | 39.045 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MG550112 | Haloferax tailed virus 1 | Haloferax tailed virus 1 unclassified Haloviruses Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 38059 | 54.077 | Unclassified | Unclassified | Unclassified | Haloferax |
| MG252615 | Escherichia phage vB_EcoS_HSE2 | Escherichia phage vB_EcoS_HSE2 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41317 | 50.873 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK514283 | Pseudomonas phage vB_PaeP_TF17 | Pseudomonas phage vB_PaeP_TF17 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43580 | 45.207 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MK310184 | Phage NC-G | Phage NC-G unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170164 | 39.522 | Dhakavirus | Tevenvirinae | Myoviridae | Unspecified |
| CP038007 | Klebsiella phage 020009 | Klebsiella phage 020009 unclassified bacterial viruses Viruses | 55032 | 52.686 | Unclassified | Unclassified | Unclassified | Klebsiella |
| MK448236 | Klebsiella phage ST899-OXA48phi17.1 | Klebsiella phage ST899-OXA48phi17.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35476 | 51.643 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK448228 | Klebsiella phage ST15-VIM1phi2.1 | Klebsiella phage ST15-VIM1phi2.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46342 | 53.586 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK433582 | Klebsiella phage ST258-KPC3phi16.2 | Klebsiella phage ST258-KPC3phi16.2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33326 | 51.455 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK433581 | Klebsiella phage ST258-KPC3phi16.1 | Klebsiella phage ST258-KPC3phi16.1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39643 | 52.758 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK433580 | Klebsiella phage ST11-OXA48phi15.3 | Klebsiella phage ST11-OXA48phi15.3 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33326 | 51.47 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK433577 | Klebsiella phage ST512-KPC3phi13.6 | Klebsiella phage ST512-KPC3phi13.6 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39643 | 52.758 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK422454 | Klebsiella phage ST340-VIM1phi10.2 | Klebsiella phage ST340-VIM1phi10.2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33326 | 51.461 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK422453 | Klebsiella phage ST13-OXA48phi12.1 | Klebsiella phage ST13-OXA48phi12.1 Klebsiella virus ST13OXA48phi12-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39086 | 53.526 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK422452 | Klebsiella phage ST13-OXA48phi12.2 | Klebsiella phage ST13-OXA48phi12.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34141 | 50.485 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK422449 | Klebsiella phage ST512-KPC3phi13.2 | Klebsiella phage ST512-KPC3phi13.2 Klebsiella virus ST512KPC3phi13-2 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32302 | 51.873 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416008 | Klebsiella phage ST405-OXA48phi1.3 | Klebsiella phage ST405-OXA48phi1.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32010 | 50.887 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK416009 | Klebsiella phage ST11-OXA245phi3.1 | Klebsiella phage ST11-OXA245phi3.1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33326 | 51.5 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416011 | Klebsiella phage ST437-OXA245phi4.1 | Klebsiella phage ST437-OXA245phi4.1 Klebsiella virus ST437OXA245phi4-1 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39642 | 52.76 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416016 | Klebsiella phage ST101-KPC2phi6.2 | Klebsiella phage ST101-KPC2phi6.2 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11454 | 58.958 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416020 | Klebsiella phage ST11-VIM1phi8.4 | Klebsiella phage ST11-VIM1phi8.4 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33016 | 51.59 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| MK416021 | Klebsiella phage ST846-OXA48phi9.1 | Klebsiella phage ST846-OXA48phi9.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38370 | 53.331 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MK514282 | Erwinia phage Derbicus | Erwinia phage Derbicus Erwinia virus Derbicus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223950 | 50.564 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| MK514281 | Erwinia phage Rebecca | Erwinia phage Rebecca unclassified Agricanvirus Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273731 | 49.811 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MK580972 | Stenotrophomonas phage YB07 | Stenotrophomonas phage YB07 unclassified Menderavirus Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159862 | 54.128 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| MK016664 | Synechococcus phage S-B68 | Synechococcus phage S-B68 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163982 | 51.745 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MK016662 | Synechococcus phage S-B28 | Synechococcus phage S-B28 Synechococcus virus SB28 Qadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45424 | 42.059 | Qadamvirus | Unclassified | Autographiviridae | Synechococcus |
| MK524178 | Escherichia phage PHB12 | Escherichia phage PHB12 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168791 | 37.591 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MG250486 | Raoultella phage Ro1 | Raoultella phage Ro1 Raoultella virus Ro1 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145759 | 44.548 | Mydovirus | Vequintavirinae | Myoviridae | Raoultella |
| MG250484 | Citrobacter phage CF1 ERZ-2017 | Citrobacter phage CF1 ERZ-2017 Citrobacter virus CF1 Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171911 | 38.741 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| MG250483 | Aeromonas phage Ah1 | Aeromonas phage Ah1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221116 | 42.05 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| MF370225 | Salmonella phage ST11 | Salmonella phage ST11 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82101 | 39.02 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KY683736 | Escherichia phage V18 | Escherichia phage V18 Escherichia virus V18 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129454 | 43.626 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KY683735 | Escherichia phage ECD7 | Escherichia phage ECD7 Escherichia virus ECD7 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164706 | 40.542 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| KY626163 | Salmonella phage SE40 | Salmonella phage SE40 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28914 | 50.173 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KY626162 | Salmonella phage Si3 | Salmonella phage Si3 Salmonella virus Si3 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84419 | 39.013 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MK327947 | Escherichia phage vB_EcoM_G5211 | Escherichia phage vB_EcoM_G5211 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164278 | 40.44 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327946 | Escherichia phage vB_EcoM_G4507 | Escherichia phage vB_EcoM_G4507 Escherichia virus G4507 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168828 | 35.367 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327945 | Escherichia phage vB_EcoM_G4500 | Escherichia phage vB_EcoM_G4500 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168363 | 35.32 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327944 | Escherichia phage vB_EcoM_G2540-3 | Escherichia phage vB_EcoM_G2540-3 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168654 | 35.333 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327943 | Escherichia phage vB_EcoM_G53 | Escherichia phage vB_EcoM_G53 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167834 | 37.752 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327942 | Escherichia phage vB_EcoM_G50 | Escherichia phage vB_EcoM_G50 Escherichia virus G50 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167728 | 35.515 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327941 | Escherichia phage vB_EcoM_G37-3 | Escherichia phage vB_EcoM_G37-3 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167832 | 40.331 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327940 | Escherichia phage vB_EcoM_G29 | Escherichia phage vB_EcoM_G29 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168241 | 35.328 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327939 | Escherichia phage vB_EcoM_G4498 | Escherichia phage vB_EcoM_G4498 Escherichia virus G4498 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167826 | 35.46 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327938 | Escherichia phage vB_EcoM_Goslar | Escherichia phage vB_EcoM_Goslar Escherichia virus Goslar Goslarvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 237307 | 46.53 | Goslarvirus | Unclassified | Myoviridae | Escherichia |
| MK327937 | Escherichia phage vB_EcoM_G10400 | Escherichia phage vB_EcoM_G10400 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170182 | 35.286 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327936 | Escherichia phage vB_EcoM_G2540 | Escherichia phage vB_EcoM_G2540 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168886 | 35.356 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327935 | Escherichia phage vB_EcoM_G2494 | Escherichia phage vB_EcoM_G2494 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168327 | 40.324 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327934 | Escherichia phage vB_EcoM_G2469 | Escherichia phage vB_EcoM_G2469 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170452 | 37.568 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327933 | Escherichia phage vB_EcoM_G2285 | Escherichia phage vB_EcoM_G2285 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166675 | 37.544 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327932 | Escherichia phage vB_EcoM_G2248 | Escherichia phage vB_EcoM_G2248 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170678 | 40.524 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327931 | Escherichia phage vB_EcoM_G17 | Escherichia phage vB_EcoM_G17 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 370817 | 34.335 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MK327930 | Escherichia phage EdH4 | Escherichia phage EdH4 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136031 | 43.653 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MK327929 | Escherichia phage D5505 | Escherichia phage D5505 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168049 | 35.388 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK327928 | Escherichia phage vB_EcoM_G2133 | Escherichia phage vB_EcoM_G2133 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168959 | 35.332 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KC847113 | Bacillus phage PBP180 | Bacillus phage PBP180 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28205 | 40.801 | Unclassified | Unclassified | Myoviridae | Bacillus |
| AP019535 | Pseudomonas phage PA01 | Pseudomonas phage PA01 Pseudomonas virus PA01 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66220 | 55.402 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| AP019527 | Lactococcus phage phiQ1 | Lactococcus phage phiQ1 Lactococcus virus phiQ1 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59833 | 34.412 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| AP019522 | Staphylococcus phage MR003 | Staphylococcus phage MR003 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132152 | 29.96 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH844558 | Exiguobacterium phage vB_EauM-23 | Exiguobacterium phage vB_EauM-23 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37660 | 52.079 | Unclassified | Unclassified | Myoviridae | Exiguobacterium |
| MH709121 | Salmonella phage SenALZ1 | Salmonella phage SenALZ1 Salmonella virus SenALZ1 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154811 | 44.56 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH709120 | Salmonella phage SenASZ3 | Salmonella phage SenASZ3 Salmonella virus SenASZ3 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157630 | 44.74 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MK359316 | Mycobacterium phage Cueylyss | Mycobacterium phage Cueylyss unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48484 | 64.064 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK359300 | Mycobacterium phage AFIS | Mycobacterium phage AFIS unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51737 | 63.687 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279913 | Mycobacterium phage PotatoSplit | Mycobacterium phage PotatoSplit unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50886 | 63.99 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279903 | Mycobacterium phage Zetzy | Mycobacterium phage Zetzy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48229 | 64.028 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH976517 | Mycobacterium phage Popcicle | Mycobacterium phage Popcicle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50075 | 64.054 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH651179 | Mycobacterium phage MadMarie | Mycobacterium phage MadMarie unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50849 | 64.013 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH576959 | Mycobacterium phage Pita2 | Mycobacterium phage Pita2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51486 | 63.4 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH450126 | Mycobacterium phage PhishRPhriends | Mycobacterium phage PhishRPhriends unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49756 | 63.924 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH399776 | Mycobacterium phage Grum1 | Mycobacterium phage Grum1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50883 | 64.065 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH371110 | Mycobacterium phage Magnar | Mycobacterium phage Magnar unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51428 | 63.726 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH338240 | Mycobacterium phage Sabia | Mycobacterium phage Sabia unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50855 | 64.021 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH155864 | Mycobacterium phage Beauxregard13 | Mycobacterium phage Beauxregard13 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50884 | 63.995 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH051259 | Mycobacterium phage TNguyen7 | Mycobacterium phage TNguyen7 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50413 | 63.805 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH047633 | Mycobacterium phage Pistachio | Mycobacterium phage Pistachio unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50006 | 63.782 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH047632 | Mycobacterium phage Puppy | Mycobacterium phage Puppy unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50084 | 63.781 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH020238 | Mycobacterium phage Lilith | Mycobacterium phage Lilith unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50841 | 64.053 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925355 | Mycobacterium phage OlanP | Mycobacterium phage OlanP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50755 | 63.966 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG925341 | Mycobacterium phage EpicPhail | Mycobacterium phage EpicPhail unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50884 | 63.993 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF140410 | Mycobacterium phage Et2Brutus | Mycobacterium phage Et2Brutus unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52497 | 63.798 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF185733 | Mycobacterium phage StepMih | Mycobacterium phage StepMih unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50841 | 64.013 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MF141539 | Mycobacterium phage MyraDee | Mycobacterium phage MyraDee unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50514 | 62.694 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX576639 | Mycobacterium phage DudeLittle | Mycobacterium phage DudeLittle unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52917 | 63.479 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX458237 | Mycobacterium phage Penny1 | Mycobacterium phage Penny1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50884 | 63.995 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KU255188 | Mycobacterium phage JenCasNa | Mycobacterium phage JenCasNa unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50877 | 63.999 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM677210 | Mycobacterium phage Larenn | Mycobacterium phage Larenn unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52967 | 63.54 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KM197169 | Mycobacterium phage Piro94 | Mycobacterium phage Piro94 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52647 | 63.419 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ510415 | Mycobacterium phage Phantastic | Mycobacterium phage Phantastic unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50101 | 63.805 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF981601 | Mycobacterium phage Echild | Mycobacterium phage Echild unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53159 | 63.69 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC661281 | Mycobacterium phage Jobu08 | Mycobacterium phage Jobu08 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50679 | 64.032 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK474470 | Vibrio phage vB_VpaS_OWB | Vibrio phage vB_VpaS_OWB Vibrio virus OWB Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43264 | 48.74 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MK554696 | Fusobacterium phage Fnu1 | Fusobacterium phage Fnu1 Fusobacterium virus FNU1 Latrobevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130914 | 24.95 | Latrobevirus | Unclassified | Siphoviridae | Fusobacterium |
| MK542015 | Escherichia phage ZCEC5 | Escherichia phage ZCEC5 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43372 | 50.689 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MK552327 | Pseudomonas phage Psa21 | Pseudomonas phage Psa21 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 305260 | 43.147 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MK450543 | Natrialba phage PhiCh1 | Natrialba phage PhiCh1 Natrialba virus PhiCh1 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58487 | 61.898 | Myohalovirus | Unclassified | Myoviridae | Natrialba |
| MK575466 | Vibrio phage Rostov 7 | Vibrio phage Rostov 7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45903 | 45.69 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MK568062 | Salmonella phage LSE7621 | Salmonella phage LSE7621 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50936 | 46.185 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MK424903 | Pseudoalteromonas phage GXT1010 | Pseudoalteromonas phage GXT1010 unclassified Corticovirus Corticovirus Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 12527 | 43.195 | Corticovirus | Unclassified | Corticoviridae | Pseudoalteromonas |
| MK524177 | Escherichia phage PHB10 | Escherichia phage PHB10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57228 | 50.252 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MK524176 | Escherichia phage PHB11 | Escherichia phage PHB11 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87553 | 38.902 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| MK542821 | Escherichia phage HZP2 | Escherichia phage HZP2 Escherichia virus HZP2 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32448 | 48.305 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK059749 | Mycobacterium phage CRB2 | Mycobacterium phage CRB2 unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72217 | 69.655 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT932700 | Enterococcus phage vB_EfaP_IME195 | Enterococcus phage vB_EfaP_IME195 Enterococcus virus IME195 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18655 | 33.203 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MH102285 | Salmonella phage vB_SenS_PHB06 | Salmonella phage vB_SenS_PHB06 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84406 | 39.187 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MK310182 | Escherichia phage NC-A | Escherichia phage NC-A Escherichia virus NCA Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39536 | 48.816 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK411820 | Acinetobacter phage vB_AbaS_D0 | Acinetobacter phage vB_AbaS_D0 unclassified Lokivirus Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43051 | 45.481 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| MH844531 | Klebsiella phage GH-K3 | Klebsiella phage GH-K3 Klebsiella virus GHK3 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49427 | 50.169 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KY697807 | Microcystis phage MACPNOA1 | Microcystis phage MACPNOA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42473 | 68.265 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MH491167 | Cronobacter phage GW1 | Cronobacter phage GW1 Cronobacter virus GW1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39695 | 53.183 | Kayfunavirus | Studiervirinae | Autographiviridae | Cronobacter |
| MK301608 | Vibrio virus vB_VspP_SBP1 | Vibrio virus vB_VspP_SBP1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78071 | 44.645 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MK069556 | Microcystis phage Me-ZS1 | Microcystis phage Me-ZS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49665 | 58.278 | Unclassified | Unclassified | Siphoviridae | Microcystis |
| MF431617 | Lentibacter virus ICBM1 | Lentibacter virus ICBM1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40163 | 47.023 | Siovirus | Cobavirinae | Zobellviridae | Lentibacter |
| MF431616 | Lentibacter virus vB_LenP_ICBM2 | Lentibacter virus vB_LenP_ICBM2 unclassified Siovirus Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40907 | 47.816 | Siovirus | Cobavirinae | Zobellviridae | Lentibacter |
| MF431615 | Lentibacter virus vB_LenP_ICBM3 | Lentibacter virus vB_LenP_ICBM3 unclassified Siovirus Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40497 | 47.25 | Siovirus | Cobavirinae | Zobellviridae | Lentibacter |
| MH107118 | Staphylococcus phage CSA13 | Staphylococcus phage CSA13 Staphylococcus virus CSA13 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17034 | 28.966 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| AP019418 | Pseudomonas phage PA02 | Pseudomonas phage PA02 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279095 | 36.835 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| LR215724 | Yersinia phage fPS-65 | Yersinia phage fPS-65 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167058 | 35.327 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| LR215723 | Yersinia phage fPS-90 | Yersinia phage fPS-90 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167132 | 35.384 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| LR215722 | Yersinia phage fPS-2 | Yersinia phage fPS-2 Yersinia virus fPS-2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169060 | 35.347 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| LR215721 | Staphylococcus phage Stab22 | Staphylococcus phage Stab22 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154499 | 30.879 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| LR215720 | Staphylococcus phage Stab23 | Staphylococcus phage Stab23 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155962 | 30.61 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| LR215719 | Staphylococcus phage Stab21 | Staphylococcus phage Stab21 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153797 | 30.324 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| LR215718 | Staphylococcus phage Stab20 | Staphylococcus phage Stab20 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153338 | 30.206 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK388859 | Klebsiella phage ST405-OXA48phi1.1 | Klebsiella phage ST405-OXA48phi1.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35232 | 53.755 | Unclassified | Unclassified | Myoviridae | Klebsiella |
| MK340761 | Pseudomonas phage vB_PaeM_SCUT-S2 | Pseudomonas phage vB_PaeM_SCUT-S2 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94434 | 49.345 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK340760 | Pseudomonas phage vB_PaeM_SCUT-S1 | Pseudomonas phage vB_PaeM_SCUT-S1 Pseudomonas virus S1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66086 | 55.432 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MH992131 | Thermus phage TSP4 | Thermus phage TSP4 unclassified Oshimavirus Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83142 | 56.004 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| MH939157 | Meiothermus phage MMP17 | Meiothermus phage MMP17 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33172 | 63.858 | Unclassified | Unclassified | Myoviridae | Meiothermus |
| MK387309 | Pseudoalteromonas phage C7 | Pseudoalteromonas phage C7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42261 | 40.631 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MK348510 | Staphylococcus phage vB_SauS_fPfSau02 | Staphylococcus phage vB_SauS_fPfSau02 Staphylococcus virus fPfSau02 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45108 | 33.708 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MK331931 | Staphylococcus phage Sa30 | Staphylococcus phage Sa30 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146737 | 30.352 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK331930 | Staphylococcus phage CH1 | Staphylococcus phage CH1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138057 | 30.426 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH142220 | Acidithiobacillus phage AcaML1 | Acidithiobacillus phage AcaML1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59362 | 65.471 | Unclassified | Unclassified | Myoviridae | Acidithiobacillus |
| MH142219 | Acidithiobacillus phage AcaML1 | Acidithiobacillus phage AcaML1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59361 | 65.471 | Unclassified | Unclassified | Myoviridae | Acidithiobacillus |
| MH925094 | Vibrio phage VspSw_1 | Vibrio phage VspSw_1 Vibrio virus VspSw1 Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113760 | 43.836 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MH925093 | Vibrio phage VspDsh_1 | Vibrio phage VspDsh_1 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46692 | 46.715 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MH925092 | Vibrio phage VpaJT_1 | Vibrio phage VpaJT_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60177 | 49.246 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MH925091 | Vibrio phage ValSw4_1 | Vibrio phage ValSw4_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79545 | 45.681 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MH925090 | Vibrio phage ValLY_3 | Vibrio phage ValLY_3 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76310 | 48.78 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MH791416 | Pseudomonas phage PaSzW-1 | Pseudomonas phage PaSzW-1 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94315 | 49.364 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MH791415 | Enterococcus phage EfsWh-1 | Enterococcus phage EfsWh-1 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58036 | 39.849 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| MH791414 | Aeromonas phage Aswh_1 | Aeromonas phage Aswh_1 unclassified Ishigurovirus Ishigurovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229730 | 38.37 | Ishigurovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MH791413 | Escherichia phage EcSzw_1 | Escherichia phage EcSzw_1 unclassified Suspvirus Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86813 | 39.97 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| MH791412 | Salmonella phage SeSz-2 | Salmonella phage SeSz-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45049 | 45.972 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH791411 | Escherichia phage Ecwhy_1 | Escherichia phage Ecwhy_1 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 354537 | 35.571 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MH791410 | Salmonella phage Seszw_1 | Salmonella phage Seszw_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45881 | 45.938 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH791409 | Escherichia phage EcWhh-1 | Escherichia phage EcWhh-1 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170524 | 39.442 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MH791407 | Aeromonas phage AsFcp_4 | Aeromonas phage AsFcp_4 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 220450 | 41.919 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| MH791406 | Pseudomonas phage PaSz-4 | Pseudomonas phage PaSz-4 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61942 | 64.438 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MH791405 | Salmonella phage SeSz-1 | Salmonella phage SeSz-1 Salmonella virus SeSz1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155297 | 44.768 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MH791404 | Aeromonas phage Assk | Aeromonas phage Assk unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231839 | 37.288 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MH791403 | Pseudomonas phage PaTs-2 | Pseudomonas phage PaTs-2 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61820 | 64.583 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MH791402 | Salmonella phage Segz_1 | Salmonella phage Segz_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48285 | 46.416 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH791401 | Aeromonas phage AsFcp_2 | Aeromonas phage AsFcp_2 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219171 | 42.022 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| MH791400 | Escherichia phage EcSzw-2 | Escherichia phage EcSzw-2 unclassified Suspvirus Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86752 | 39.984 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| MH791398 | Aeromonas phage Asswx_1 | Aeromonas phage Asswx_1 unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229103 | 38.812 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MH791397 | Enterococcus phage EfsSzw-1 | Enterococcus phage EfsSzw-1 unclassified Schiekvirus Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150272 | 36.834 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MH791396 | Salmonella phage SeZq-1 | Salmonella phage SeZq-1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60188 | 56.588 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MH791395 | Salmonella phage SeWh-1 | Salmonella phage SeWh-1 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59878 | 56.583 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MK360025 | Enterococcus phage vB_EfaP_Zip | Enterococcus phage vB_EfaP_Zip Enterococcus virus Zip Minhovirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18742 | 35.044 | Minhovirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MK360024 | Enterococcus phage vB_EfaS_Max | Enterococcus phage vB_EfaS_Max unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 34.711 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MK330684 | Aeromonas phage ZPAH7B | Aeromonas phage ZPAH7B Aeromonas virus ZPAH7 Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30791 | 56.78 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MK305884 | Mycobacterium phage BQuat | Mycobacterium phage BQuat unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41893 | 66.591 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MK388689 | Pantoea phage vB_PagM_LIET2 | Pantoea phage vB_PagM_LIET2 Pantoea virus LIET2 Lietduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74710 | 53.989 | Lietduovirus | Unclassified | Myoviridae | Pantoea |
| MK335533 | Shigella phage vB_SsoS_008 | Shigella phage vB_SsoS_008 Shigella virus 008 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50414 | 45.656 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MK327140 | Klebsiella phage YX3973 | Klebsiella phage YX3973 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46907 | 46.899 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MG867727 | Escherichia phage vB_EcoM-G28 | Escherichia phage vB_EcoM-G28 Escherichia virus G28 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170121 | 35.346 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MK417516 | Staphylococcus virus J-Sa36 | Staphylococcus virus J-Sa36 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148312 | 30.387 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK417515 | Staphylococcus virus Sa87 | Staphylococcus virus Sa87 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149108 | 30.332 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK417514 | Staphylococcus virus Sa83 | Staphylococcus virus Sa83 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146004 | 30.314 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK224498 | Pseudomonas phage Henu5 | Pseudomonas phage Henu5 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92558 | 49.364 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK224497 | Mycobacterium phage Henu3 | Mycobacterium phage Henu3 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58899 | 67.612 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MK408758 | Campylobacter phage CP20 | Campylobacter phage CP20 unclassified Firehammervirus Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174820 | 27.351 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MK372342 | Escherichia phage vB_EcoS_IME542 | Escherichia phage vB_EcoS_IME542 Escherichia virus IME542 Christensenvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46553 | 43.568 | Christensenvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MK340941 | Acinetobacter phage AbTJ | Acinetobacter phage AbTJ Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42670 | 39.323 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MK240351 | Acinetobacter phage Henu6 | Acinetobacter phage Henu6 unclassified Zedzedvirus Zedzedvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166890 | 34.333 | Zedzedvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MK234886 | Escherichia phage AnYang | Escherichia phage AnYang unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169215 | 39.494 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MK211557 | Staphylococcus phage Henu2 | Staphylococcus phage Henu2 Staphylococcus virus Henu2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43513 | 35.04 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK284528 | Bacillus phage DK3 | Bacillus phage DK3 Bacillus virus DK3 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26865 | 31.055 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| MK284527 | Bacillus phage DK2 | Bacillus phage DK2 Bacillus virus DK2 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26357 | 30.914 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| MK284526 | Bacillus phage DK1 | Bacillus phage DK1 Bacillus virus DK1 Hemphillvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27180 | 30.868 | Hemphillvirus | Northropvirinae | Salasmaviridae | Bacillus |
| MF683623 | Aeromonas phage CF7 | Aeromonas phage CF7 Aeromonas virus CF7 Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42439 | 58.95 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MG707188 | Pseudomonas phage TC7 | Pseudomonas phage TC7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43748 | 62.531 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MK317959 | Pseudomonas phage EPa61 | Pseudomonas phage EPa61 Pseudomonas virus EPa61 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65905 | 55.134 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK295205 | Shigella phage vB_SflM_004 | Shigella phage vB_SflM_004 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85887 | 38.619 | Unclassified | Ounavirinae | Myoviridae | Shigella |
| MK295204 | Shigella phage vB_SdyM_006 | Shigella phage vB_SdyM_006 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166138 | 31.498 | Unclassified | Tevenvirinae | Myoviridae | Shigella |
| MK295203 | Escherichia phage vB_EcoM_005 | Escherichia phage vB_EcoM_005 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155101 | 39.327 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| MK291445 | Paracoccus phage vB_PyeM_Pyei1 | Paracoccus phage vB_PyeM_Pyei1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50161 | 65.493 | Unclassified | Unclassified | Myoviridae | Paracoccus |
| MK291444 | Paracoccus phage vB_PthS_Pthi1 | Paracoccus phage vB_PthS_Pthi1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39547 | 63.843 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| MK291443 | Paracoccus phage vB_PsuS_Psul1 | Paracoccus phage vB_PsuS_Psul1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37901 | 60.901 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| MK291442 | Paracoccus phage vB_PkoS_Pkon1 | Paracoccus phage vB_PkoS_Pkon1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49723 | 60.626 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| MK291441 | Paracoccus phage vB_PbeS_Pben1 | Paracoccus phage vB_PbeS_Pben1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39879 | 64.262 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| MK286578 | Salmonella phage Season12 | Salmonella phage Season12 unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59047 | 56.467 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MK290764 | Staphylococcus virus B122 | Staphylococcus virus B122 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43108 | 34.987 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MK288022 | Bacillus phage pW4 | Bacillus phage pW4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80919 | 34.024 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MK288021 | Bacillus phage pW2 | Bacillus phage pW2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160627 | 32.536 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MK278861 | Klebsiella phage KpKT21phi1 | Klebsiella phage KpKT21phi1 Klebsiella virus KpKT21phi1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49106 | 50.89 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK278860 | Acinetobacter phage AbTZA1 | Acinetobacter phage AbTZA1 Acinetobacter virus AbTZA1 Hadassahvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168223 | 36.345 | Hadassahvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MK278859 | Acinetobacter phage AbKT21phiIII | Acinetobacter phage AbKT21phiIII Acinetobacter virus AbKT21III Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40898 | 39.359 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MK249872 | Vibrio phage LP.1 | Vibrio phage LP.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46791 | 46.359 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK249871 | Vibrio phage LP.2 | Vibrio phage LP.2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37128 | 43.075 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MK257744 | Thermus phage YS40_Isch | Thermus phage YS40_Isch Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162184 | 33.275 | Unclassified | Unclassified | Myoviridae | Thermus |
| MK250904 | Staphylococcus phage VB-SavM-JYL02 | Staphylococcus phage VB-SavM-JYL02 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141384 | 30.18 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MK240575 | Thermobifida phage P318 | Thermobifida phage P318 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48045 | 54.57 | Unclassified | Unclassified | Siphoviridae | Thermobifida |
| MF001363 | Salmonella phage SH9 | Salmonella phage SH9 Salmonella virus SH9 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111607 | 40.145 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KY694971 | Citrobacter phage CF1 DK-2017 | Citrobacter phage CF1 DK-2017 Citrobacter virus DK2017 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50339 | 42.647 | Tlsvirus | Tempevirinae | Drexlerviridae | Citrobacter |
| MH884512 | Bacillus phage vB_BpsS-140 | Bacillus phage vB_BpsS-140 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55091 | 39.841 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MK213796 | Escherichia phage T1 | Escherichia phage T1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48836 | 45.552 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MK279909 | Mycobacterium phage DrLupo | Mycobacterium phage DrLupo Mycobacterium virus DrLupo Barnyardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70030 | 57.501 | Barnyardvirus | Unclassified | Siphoviridae | Mycobacterium |
| MK279899 | Arthrobacter phage Maja | Arthrobacter phage Maja Arthrobacter virus Maja Majavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36776 | 65.053 | Majavirus | Unclassified | Siphoviridae | Arthrobacter |
| MK203841 | Klebsiella phage Henu1 | Klebsiella phage Henu1 Klebsiella virus Henu1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40352 | 53.142 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK095606 | Pantoea phage vB_PagS_AAS23 | Pantoea phage vB_PagS_AAS23 Pantoea virus AAS23 Sauletekiovirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51170 | 47.645 | Sauletekiovirus | Unclassified | Drexlerviridae | Pantoea |
| MK095605 | Pantoea phage vB_PagS_MED16 | Pantoea phage vB_PagS_MED16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46103 | 55.111 | Unclassified | Unclassified | Siphoviridae | Pantoea |
| MK213795 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3526 | 52.184 | Levivirus | Unclassified | Leviviridae | Escherichia |
| MK301532 | Escherichia phage Ro45lw | Escherichia phage Ro45lw Escherichia virus Ro45lw Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39793 | 52.19 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MK033137 | Pseudomonas phage UNO-SLW1 | Pseudomonas phage UNO-SLW1 Pseudomonas virus UNOSLW1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39636 | 57.952 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MK033136 | Enterobacter phage EcpYZU01 | Enterobacter phage EcpYZU01 Enterobacter virus EcpYZU01 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39767 | 51.751 | Kayfunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| MH673674 | Thermus phage phiMa | Thermus phage phiMa Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51843 | 67.301 | Unclassified | Unclassified | Myoviridae | Thermus |
| MH673673 | Thermus phage phiLo | Thermus phage phiLo Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178531 | 50.717 | Unclassified | Unclassified | Myoviridae | Thermus |
| MG557979 | Lactobacillus phage Lpa804 | Lactobacillus phage Lpa804 Lactobacillus virus Lpa804 Harbinvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142567 | 36.623 | Harbinvirus | Unclassified | Herelleviridae | Lactobacillus |
| MH992513 | Aeromonas phage ZPAH7 | Aeromonas phage ZPAH7 Aeromonas virus ZPAH7 Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30791 | 56.78 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MG197810 | Klebsiella phage 1611E-K2-1 | Klebsiella phage 1611E-K2-1 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47797 | 48.915 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MF564201 | Escherichia phage vB_EcoS-95 | Escherichia phage vB_EcoS-95 Escherichia virus 95 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50910 | 44.83 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| MH230177 | Aeromonas phage SD04 | Aeromonas phage SD04 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59646 | 44.689 | Seuratvirus | Queuovirinae | Siphoviridae | Aeromonas |
| MH356730 | Staphylococcus phage VB-SauS-SA2 | Staphylococcus phage VB-SauS-SA2 unclassified Sextaecvirus Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89055 | 31.917 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| MH940212 | Salmonella phage PS5 | Salmonella phage PS5 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158400 | 44.8 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MG602477 | Bacillus phage BCP01 | Bacillus phage BCP01 unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157958 | 39.871 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MG602476 | Vibrio phage VVP001 | Vibrio phage VVP001 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76423 | 48.835 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MH992223 | Apis mellifera associated microvirus 47 | Apis mellifera associated microvirus 47 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4117 | 60.068 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992222 | Apis mellifera associated microvirus 54 | Apis mellifera associated microvirus 54 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4211 | 41.677 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992221 | Apis mellifera associated microvirus 36 | Apis mellifera associated microvirus 36 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4231 | 46.561 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992220 | Apis mellifera associated microvirus 41 | Apis mellifera associated microvirus 41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4259 | 57.83 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992219 | Apis mellifera associated microvirus 46 | Apis mellifera associated microvirus 46 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4269 | 52.097 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992218 | Apis mellifera associated microvirus 22 | Apis mellifera associated microvirus 22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4278 | 44.483 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992217 | Apis mellifera associated microvirus 42 | Apis mellifera associated microvirus 42 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4279 | 55.083 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992216 | Apis mellifera associated microvirus 43 | Apis mellifera associated microvirus 43 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4286 | 49.697 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992215 | Apis mellifera associated microvirus 52 | Apis mellifera associated microvirus 52 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4331 | 50.958 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992214 | Apis mellifera associated microvirus 53 | Apis mellifera associated microvirus 53 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4332 | 45.476 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992213 | Apis mellifera associated microvirus 15 | Apis mellifera associated microvirus 15 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4346 | 58.997 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992212 | Apis mellifera associated microvirus 10 | Apis mellifera associated microvirus 10 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4348 | 41.835 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992211 | Apis mellifera associated microvirus 19 | Apis mellifera associated microvirus 19 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4355 | 54.627 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992210 | Apis mellifera associated microvirus 18 | Apis mellifera associated microvirus 18 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4392 | 58.675 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992209 | Apis mellifera associated microvirus 17 | Apis mellifera associated microvirus 17 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4394 | 54.939 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992208 | Apis mellifera associated microvirus 51 | Apis mellifera associated microvirus 51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4412 | 47.892 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992207 | Apis mellifera associated microvirus 25 | Apis mellifera associated microvirus 25 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4431 | 40.713 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992206 | Apis mellifera associated microvirus 40 | Apis mellifera associated microvirus 40 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4436 | 60.099 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992205 | Apis mellifera associated microvirus 61 | Apis mellifera associated microvirus 61 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4448 | 48.022 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992204 | Apis mellifera associated microvirus 35 | Apis mellifera associated microvirus 35 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4469 | 42.403 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992203 | Apis mellifera associated microvirus 29 | Apis mellifera associated microvirus 29 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4470 | 42.483 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992202 | Apis mellifera associated microvirus 50 | Apis mellifera associated microvirus 50 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4485 | 54.693 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992201 | Apis mellifera associated microvirus 45 | Apis mellifera associated microvirus 45 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4502 | 53.332 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992200 | Apis mellifera associated microvirus 12 | Apis mellifera associated microvirus 12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4507 | 61.105 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992199 | Apis mellifera associated microvirus 39 | Apis mellifera associated microvirus 39 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4519 | 62.58 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992198 | Apis mellifera associated microvirus 4 | Apis mellifera associated microvirus 4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 50.099 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992197 | Apis mellifera associated microvirus 21 | Apis mellifera associated microvirus 21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4548 | 43.624 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992196 | Apis mellifera associated microvirus 60 | Apis mellifera associated microvirus 60 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4557 | 43.954 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992195 | Apis mellifera associated microvirus 28 | Apis mellifera associated microvirus 28 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4559 | 47.379 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992194 | Apis mellifera associated microvirus 26 | Apis mellifera associated microvirus 26 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4566 | 50.964 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992193 | Apis mellifera associated microvirus 32 | Apis mellifera associated microvirus 32 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4569 | 46.553 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992192 | Apis mellifera associated microvirus 49 | Apis mellifera associated microvirus 49 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4576 | 55.878 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992191 | Apis mellifera associated microvirus 23 | Apis mellifera associated microvirus 23 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4578 | 46.789 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992190 | Apis mellifera associated microvirus 30 | Apis mellifera associated microvirus 30 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4582 | 43.016 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992189 | Apis mellifera associated microvirus 24 | Apis mellifera associated microvirus 24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4600 | 45.717 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992188 | Apis mellifera associated microvirus 9 | Apis mellifera associated microvirus 9 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4608 | 42.622 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992187 | Apis mellifera associated microvirus 5 | Apis mellifera associated microvirus 5 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4633 | 45.241 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992186 | Apis mellifera associated microvirus 58 | Apis mellifera associated microvirus 58 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4708 | 42.396 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992185 | Apis mellifera associated microvirus 14 | Apis mellifera associated microvirus 14 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4719 | 57.406 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992184 | Apis mellifera associated microvirus 13 | Apis mellifera associated microvirus 13 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4737 | 51.488 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992183 | Apis mellifera associated microvirus 33 | Apis mellifera associated microvirus 33 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4756 | 44.365 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992182 | Apis mellifera associated microvirus 16 | Apis mellifera associated microvirus 16 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4882 | 52.315 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992181 | Apis mellifera associated microvirus 8 | Apis mellifera associated microvirus 8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4893 | 43.286 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992180 | Apis mellifera associated microvirus 59 | Apis mellifera associated microvirus 59 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4506 | 45.939 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992179 | Apis mellifera associated microvirus 4 | Apis mellifera associated microvirus 4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 50.099 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992178 | Apis mellifera associated microvirus 27 | Apis mellifera associated microvirus 27 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4587 | 42.359 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992177 | Apis mellifera associated microvirus 24 | Apis mellifera associated microvirus 24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4600 | 45.696 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992176 | Apis mellifera associated microvirus 5 | Apis mellifera associated microvirus 5 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4633 | 45.241 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992175 | Apis mellifera associated microvirus 11 | Apis mellifera associated microvirus 11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4661 | 64.664 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992174 | Apis mellifera associated microvirus 3 | Apis mellifera associated microvirus 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4770 | 51.593 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992173 | Apis mellifera associated microvirus 56 | Apis mellifera associated microvirus 56 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4859 | 44.927 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992172 | Apis mellifera associated microvirus 4 | Apis mellifera associated microvirus 4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 50.055 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992171 | Apis mellifera associated microvirus 48 | Apis mellifera associated microvirus 48 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4135 | 56.638 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992170 | Apis mellifera associated microvirus 31 | Apis mellifera associated microvirus 31 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4345 | 52.589 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992169 | Apis mellifera associated microvirus 6 | Apis mellifera associated microvirus 6 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4410 | 40.567 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992168 | Apis mellifera associated microvirus 57 | Apis mellifera associated microvirus 57 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4702 | 31.88 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992167 | Apis mellifera associated microvirus 55 | Apis mellifera associated microvirus 55 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6391 | 38.945 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992166 | Apis mellifera associated microvirus 44 | Apis mellifera associated microvirus 44 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4538 | 58.308 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992165 | Apis mellifera associated microvirus 7 | Apis mellifera associated microvirus 7 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4703 | 49.585 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992164 | Apis mellifera associated microvirus 38 | Apis mellifera associated microvirus 38 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4195 | 55.781 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992163 | Apis mellifera associated microvirus 37 | Apis mellifera associated microvirus 37 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4387 | 48.416 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992162 | Apis mellifera associated microvirus 4 | Apis mellifera associated microvirus 4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 50.099 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992161 | Apis mellifera associated microvirus 2 | Apis mellifera associated microvirus 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4584 | 48.909 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992160 | Apis mellifera associated microvirus 24 | Apis mellifera associated microvirus 24 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4600 | 45.696 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992159 | Apis mellifera associated microvirus 11 | Apis mellifera associated microvirus 11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4661 | 64.643 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992158 | Apis mellifera associated microvirus 20 | Apis mellifera associated microvirus 20 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4691 | 44.98 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992157 | Apis mellifera associated microvirus 3 | Apis mellifera associated microvirus 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4770 | 51.614 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992156 | Apis mellifera associated microvirus 1 | Apis mellifera associated microvirus 1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4469 | 52.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992155 | Apis mellifera associated microvirus 60 | Apis mellifera associated microvirus 60 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4557 | 43.954 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992154 | Apis mellifera associated microvirus 34 | Apis mellifera associated microvirus 34 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4728 | 44.247 | Unclassified | Unclassified | Microviridae | Unspecified |
| MK165658 | Pseudomonas phage vB_PaeS_SCUT-S4 | Pseudomonas phage vB_PaeS_SCUT-S4 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42932 | 53.834 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK165657 | Pseudomonas phage vB_PaeS_SCUT-S3 | Pseudomonas phage vB_PaeS_SCUT-S3 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42622 | 53.489 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| MK002701 | Halobacterium phage phiH | Halobacterium phage phiH Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58072 | 63.678 | Myohalovirus | Unclassified | Myoviridae | Halobacterium |
| MK097141 | Vibrio phage P23 | Vibrio phage P23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40063 | 42.54 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MH688453 | Klebsiella phage TSK1 | Klebsiella phage TSK1 Klebsiella virus TSK1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49861 | 50.452 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MK064565 | Sulfolobales Beppu rod-shaped virus 1 | Sulfolobales Beppu rod-shaped virus 1 Japarudivirus SBRV1 Japarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 34025 | 26.783 | Japarudivirus | Unclassified | Rudiviridae | Unspecified |
| MK064564 | Sulfolobales Beppu filamentous virus 3 | Sulfolobales Beppu filamentous virus 3 Deltalipothrixvirus SBFV3 Deltalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 31324 | 37.926 | Deltalipothrixvirus | Unclassified | Lipothrixviridae | Unspecified |
| MK064563 | Sulfolobales Beppu filamentous virus 2 | Sulfolobales Beppu filamentous virus 2 Alphalipothrixvirus SBFV2 Alphalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 38091 | 39.458 | Alphalipothrixvirus | Unclassified | Lipothrixviridae | Unspecified |
| MK064562 | Sulfolobales Beppu filamentous virus 1 | Sulfolobales Beppu filamentous virus 1 unclassified Lipothrixviridae Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 37919 | 35.555 | Unclassified | Unclassified | Lipothrixviridae | Unspecified |
| MH892384 | Streptococcus phage SW32 | Streptococcus phage SW32 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36587 | 39.238 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892370 | Streptococcus phage SW23 | Streptococcus phage SW23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31422 | 37.031 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892365 | Streptococcus phage SW17 | Streptococcus phage SW17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32005 | 36.844 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892350 | Streptococcus phage SW16 | Streptococcus phage SW16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32106 | 36.953 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892385 | Streptococcus phage SW33 | Streptococcus phage SW33 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36587 | 39.238 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892383 | Streptococcus phage SW31 | Streptococcus phage SW31 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36587 | 39.238 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892382 | Streptococcus phage SWK6 | Streptococcus phage SWK6 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34999 | 39.007 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892381 | Streptococcus phage SWK5 | Streptococcus phage SWK5 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35849 | 38.899 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892380 | Streptococcus phage SWK4 | Streptococcus phage SWK4 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35341 | 38.694 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892379 | Streptococcus phage SWK3 | Streptococcus phage SWK3 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35443 | 38.716 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892378 | Streptococcus phage SWK2 | Streptococcus phage SWK2 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36594 | 39.296 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892377 | Streptococcus phage SWK1 | Streptococcus phage SWK1 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36089 | 39.167 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892376 | Streptococcus phage SW1151 | Streptococcus phage SW1151 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35056 | 39.599 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892375 | Streptococcus phage SW30 | Streptococcus phage SW30 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34158 | 38.46 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892374 | Streptococcus phage SW28 | Streptococcus phage SW28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32093 | 36.88 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892373 | Streptococcus phage SW27 | Streptococcus phage SW27 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33953 | 38.25 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892372 | Streptococcus phage SW26 | Streptococcus phage SW26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31283 | 36.985 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892371 | Streptococcus phage SW25 | Streptococcus phage SW25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32013 | 37.197 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892369 | Streptococcus phage SW22 | Streptococcus phage SW22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31738 | 37.35 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892368 | Streptococcus phage SW20 | Streptococcus phage SW20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32567 | 36.81 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892367 | Streptococcus phage SW19 | Streptococcus phage SW19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35153 | 38.096 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892366 | Streptococcus phage SW18 | Streptococcus phage SW18 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34825 | 39.434 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892364 | Streptococcus phage SW15 | Streptococcus phage SW15 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34289 | 39.386 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892363 | Streptococcus phage SW14 | Streptococcus phage SW14 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35913 | 39.866 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892362 | Streptococcus phage SW13 | Streptococcus phage SW13 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36587 | 39.238 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892361 | Streptococcus phage SW12 | Streptococcus phage SW12 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35382 | 38.754 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892360 | Streptococcus phage SW11 | Streptococcus phage SW11 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36250 | 38.234 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892359 | Streptococcus phage SW10 | Streptococcus phage SW10 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34336 | 38.758 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892358 | Streptococcus phage SW9 | Streptococcus phage SW9 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33999 | 38.713 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892357 | Streptococcus phage SW8 | Streptococcus phage SW8 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33930 | 39.107 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892356 | Streptococcus phage SW21 | Streptococcus phage SW21 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35351 | 38.777 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892355 | Streptococcus phage SW4 | Streptococcus phage SW4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34556 | 38.153 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH892354 | Streptococcus phage SW3 | Streptococcus phage SW3 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37181 | 38.641 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892353 | Streptococcus phage SW2 | Streptococcus phage SW2 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37222 | 38.805 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892352 | Streptococcus phage SW1 | Streptococcus phage SW1 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34821 | 38.85 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH892351 | Streptococcus phage SW6 | Streptococcus phage SW6 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34276 | 38.642 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH973663 | Streptococcus phage SW24 | Streptococcus phage SW24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32092 | 38.309 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH973662 | Streptococcus phage SW7 | Streptococcus phage SW7 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35917 | 39.124 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH973661 | Streptococcus phage SW5 | Streptococcus phage SW5 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36836 | 38.777 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MG450654 | Synechococcus phage S-CBWM1 | Synechococcus phage S-CBWM1 Synechococcus virus SCBWM1 Aokuangvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139069 | 51.623 | Aokuangvirus | Unclassified | Myoviridae | Synechococcus |
| KP162168 | Paracoccus phage vB_PmaS-R3 | Paracoccus phage vB_PmaS-R3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42093 | 56.368 | Unclassified | Unclassified | Siphoviridae | Paracoccus |
| MK054237 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 17757 | 39.511 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054236 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 18548 | 38.953 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054235 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15789 | 39.141 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054234 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15911 | 39.325 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054233 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15548 | 38.835 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054232 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15575 | 38.902 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054231 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15412 | 38.846 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054230 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15881 | 38.883 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054229 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16213 | 38.778 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054228 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16451 | 38.812 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054227 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 14609 | 37.545 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054226 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16026 | 38.712 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054225 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15815 | 38.868 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054224 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16044 | 38.631 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054223 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15766 | 38.862 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054222 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15750 | 38.597 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054221 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15029 | 38.259 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054220 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16319 | 38.716 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054219 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 14972 | 37.877 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054218 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11658 | 39.792 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054217 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11593 | 40.024 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054216 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11887 | 39.775 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054215 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 17148 | 39.41 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054214 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11661 | 39.799 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054213 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15176 | 38.528 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054212 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11433 | 39.876 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054211 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11323 | 40.148 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054210 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 14967 | 38.886 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054209 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 11682 | 39.908 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054208 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15529 | 37.601 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054207 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15088 | 37.473 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054206 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 18208 | 39.268 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054205 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16184 | 37.809 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK054204 | Sulfolobus spindle-shaped virus | Sulfolobus spindle-shaped virus unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 17195 | 39.035 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| MK095211 | Pectobacterium phage Zenivior_B1 | Pectobacterium phage Zenivior_B1 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43724 | 49.001 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095210 | Pectobacterium phage Zenivior | Pectobacterium phage Zenivior Pectobacterium virus Zenivior Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43724 | 48.998 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095209 | Pectobacterium phage Phoria | Pectobacterium phage Phoria Pectobacterium virus Phoria Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43825 | 48.634 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095208 | Pectobacterium phage Nobby_B4 | Pectobacterium phage Nobby_B4 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42830 | 48.998 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095207 | Pectobacterium phage Nobby_B3 | Pectobacterium phage Nobby_B3 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42818 | 48.993 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095206 | Pectobacterium phage Nobby_B2 | Pectobacterium phage Nobby_B2 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42837 | 49.016 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095205 | Pectobacterium phage Nobby_B1 | Pectobacterium phage Nobby_B1 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42836 | 49.003 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095204 | Pectobacterium phage Koot_B1 | Pectobacterium phage Koot_B1 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43402 | 49.018 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095203 | Pectobacterium phage Koot | Pectobacterium phage Koot Pectobacterium virus Koot Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43402 | 49.023 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095202 | Pectobacterium phage Khlen | Pectobacterium phage Khlen Pectobacterium virus Khlen Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44013 | 48.915 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095201 | Pectobacterium phage Ekidair | Pectobacterium phage Ekidair unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43495 | 48.884 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095200 | Pectobacterium phage Clickz_B8 | Pectobacterium phage Clickz_B8 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43310 | 49.194 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095199 | Pectobacterium phage Clickz_B7 | Pectobacterium phage Clickz_B7 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43310 | 49.19 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095198 | Pectobacterium phage Clickz_B6 | Pectobacterium phage Clickz_B6 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43314 | 49.192 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095197 | Pectobacterium phage Clickz_B5 | Pectobacterium phage Clickz_B5 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43312 | 49.192 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095196 | Pectobacterium phage Clickz_B4 | Pectobacterium phage Clickz_B4 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43312 | 49.19 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095195 | Pectobacterium phage Clickz_B3 | Pectobacterium phage Clickz_B3 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43310 | 49.19 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095194 | Pectobacterium phage Clickz_B2 | Pectobacterium phage Clickz_B2 unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43312 | 49.192 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095193 | Pectobacterium phage Clickz | Pectobacterium phage Clickz Pectobacterium virus Clickz Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43310 | 49.192 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK095192 | Pectobacterium phage Astalicious | Pectobacterium phage Astalicious unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42818 | 49.012 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH937491 | Streptococcus phage CHPC1037 | Streptococcus phage CHPC1037 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36551 | 38.683 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937489 | Streptococcus phage CHPC1034 | Streptococcus phage CHPC1034 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33826 | 39.147 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937482 | Streptococcus phage CHPC982 | Streptococcus phage CHPC982 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35246 | 38.861 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937476 | Streptococcus phage CHPC933 | Streptococcus phage CHPC933 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32182 | 40.128 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937507 | Streptococcus phage CHPC1152 | Streptococcus phage CHPC1152 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35353 | 39.561 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937505 | Streptococcus phage CHPC1109 | Streptococcus phage CHPC1109 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33791 | 39.297 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937495 | Streptococcus phage CHPC1045 | Streptococcus phage CHPC1045 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34096 | 38.667 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937488 | Streptococcus phage CHPC1033 | Streptococcus phage CHPC1033 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36827 | 38.548 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937484 | Streptococcus phage CHPC1008 | Streptococcus phage CHPC1008 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34844 | 40.125 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937481 | Streptococcus phage CHPC979 | Streptococcus phage CHPC979 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34277 | 38.685 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937479 | Streptococcus phage CHPC952 | Streptococcus phage CHPC952 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36775 | 39.372 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937474 | Streptococcus phage CHPC930 | Streptococcus phage CHPC930 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36350 | 38.586 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937468 | Streptococcus phage CHPC879 | Streptococcus phage CHPC879 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36011 | 38.224 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937459 | Streptococcus phage CHPC595 | Streptococcus phage CHPC595 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35697 | 39.071 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937457 | Streptococcus phage CHPC1248 | Streptococcus phage CHPC1248 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38383 | 39.51 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937511 | Streptococcus phage CHPC1247 | Streptococcus phage CHPC1247 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35543 | 39.76 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937510 | Streptococcus phage CHPC1246 | Streptococcus phage CHPC1246 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36876 | 39.229 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937509 | Streptococcus phage CHPC1230 | Streptococcus phage CHPC1230 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39384 | 39.473 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937508 | Streptococcus phage CHPC1156 | Streptococcus phage CHPC1156 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34912 | 38.998 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937506 | Streptococcus phage CHPC1148 | Streptococcus phage CHPC1148 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39069 | 38.025 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937504 | Streptococcus phage CHPC1091 | Streptococcus phage CHPC1091 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37028 | 38.441 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937503 | Streptococcus phage CHPC1084 | Streptococcus phage CHPC1084 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36776 | 39.371 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937502 | Streptococcus phage CHPC1083 | Streptococcus phage CHPC1083 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36471 | 39.316 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937501 | Streptococcus phage CHPC1073 | Streptococcus phage CHPC1073 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34017 | 38.907 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937500 | Streptococcus phage CHPC1067 | Streptococcus phage CHPC1067 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34355 | 39.028 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937499 | Streptococcus phage CHPC1062 | Streptococcus phage CHPC1062 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40037 | 38.467 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937498 | Streptococcus phage CHPC1057 | Streptococcus phage CHPC1057 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34845 | 40.143 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937497 | Streptococcus phage CHPC1048 | Streptococcus phage CHPC1048 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34812 | 38.728 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937496 | Streptococcus phage CHPC1046 | Streptococcus phage CHPC1046 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34790 | 38.735 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937494 | Streptococcus phage CHPC1042 | Streptococcus phage CHPC1042 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42019 | 39.256 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937493 | Streptococcus phage CHPC1041 | Streptococcus phage CHPC1041 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38840 | 38.136 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937492 | Streptococcus phage CHPC1040 | Streptococcus phage CHPC1040 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35851 | 38.911 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937490 | Streptococcus phage CHPC1036 | Streptococcus phage CHPC1036 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36333 | 38.706 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937487 | Streptococcus phage CHPC1029 | Streptococcus phage CHPC1029 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35920 | 39.115 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937486 | Streptococcus phage CHPC1027 | Streptococcus phage CHPC1027 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35928 | 38.897 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937485 | Streptococcus phage CHPC1014 | Streptococcus phage CHPC1014 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35260 | 38.976 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937483 | Streptococcus phage CHPC1005 | Streptococcus phage CHPC1005 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37598 | 38.494 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937480 | Streptococcus phage CHPC954 | Streptococcus phage CHPC954 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37464 | 38.728 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937478 | Streptococcus phage CHPC951 | Streptococcus phage CHPC951 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36471 | 39.313 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937477 | Streptococcus phage CHPC950 | Streptococcus phage CHPC950 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35299 | 39.015 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937475 | Streptococcus phage CHPC931 | Streptococcus phage CHPC931 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36490 | 39.2 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937473 | Streptococcus phage CHPC929 | Streptococcus phage CHPC929 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40874 | 38.487 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937472 | Streptococcus phage CHPC928 | Streptococcus phage CHPC928 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34022 | 38.801 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937471 | Streptococcus phage CHPC927 | Streptococcus phage CHPC927 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37303 | 38.975 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937470 | Streptococcus phage CHPC925 | Streptococcus phage CHPC925 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34759 | 38.479 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937469 | Streptococcus phage CHPC919 | Streptococcus phage CHPC919 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37281 | 38.269 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937467 | Streptococcus phage CHPC877 | Streptococcus phage CHPC877 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39965 | 38.364 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937466 | Streptococcus phage CHPC875 | Streptococcus phage CHPC875 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36576 | 39.02 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937465 | Streptococcus phage CHPC873 | Streptococcus phage CHPC873 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38258 | 38.084 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937464 | Streptococcus phage CHPC869 | Streptococcus phage CHPC869 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37080 | 39.064 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937463 | Streptococcus phage CHPC676 | Streptococcus phage CHPC676 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40402 | 38.758 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937462 | Streptococcus phage CHPC663 | Streptococcus phage CHPC663 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38531 | 38.93 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937460 | Streptococcus phage CHPC640 | Streptococcus phage CHPC640 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40404 | 38.768 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| MH937458 | Streptococcus phage CHPC572 | Streptococcus phage CHPC572 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37005 | 38.511 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK138527 | Escherichia phage vB_EcoS_PNS1 | Escherichia phage vB_EcoS_PNS1 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43945 | 54.361 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MK138526 | Pseudomonas phage vB_PaeM_LCK69 | Pseudomonas phage vB_PaeM_LCK69 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92319 | 49.395 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MK044829 | Streptococcus phage 109751 | Streptococcus phage 109751 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39520 | 39.375 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MK044828 | Streptococcus phage 33888 | Streptococcus phage 33888 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34256 | 39.932 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH937461 | Streptococcus phage CHPC642 | Streptococcus phage CHPC642 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35715 | 38.776 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MK165087 | Escherichia phage BRET | Escherichia phage BRET unclassified Nonagvirus Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59550 | 43.365 | Nonagvirus | Queuovirinae | Siphoviridae | Escherichia |
| MK144667 | Pseudomonas phage Spike | Pseudomonas phage Spike unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37729 | 64.34 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MK134560 | Klebsiella phage kpssk3 | Klebsiella phage kpssk3 Klebsiella virus kpssk3 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40539 | 52.803 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MK085976 | Bacillus phage AP631 | Bacillus phage AP631 unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39549 | 35.007 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| MK071653 | Meiothermus phage MMP7 | Meiothermus phage MMP7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32864 | 63.982 | Unclassified | Unclassified | Myoviridae | Meiothermus |
| MK034952 | Pseudomonas phage Dobby | Pseudomonas phage Dobby Pseudomonas virus Dobby Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37152 | 62.215 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| MH809534 | Yersinia phage PYPS50 | Yersinia phage PYPS50 Yersinia virus PYPS50 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39648 | 48.736 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| MH972263 | Staphylococcus phage JBug18 | Staphylococcus phage JBug18 unclassified Andhravirus Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18547 | 29.59 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MH972262 | Staphylococcus phage Pontiff | Staphylococcus phage Pontiff unclassified Andhravirus Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18364 | 29.955 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MH972261 | Staphylococcus phage Pike | Staphylococcus phage Pike unclassified Andhravirus Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18376 | 29.909 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MH972260 | Staphylococcus phage Pabna | Staphylococcus phage Pabna Staphylococcus virus Pabna Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17700 | 29.362 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MK105819 | Synechococcus phage S-H1 | Synechococcus phage S-H1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39158 | 53.718 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| MH844529 | Staphylococcus phage 812h1 | Staphylococcus phage 812h1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150582 | 30.389 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MH844528 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150390 | 30.367 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG779474 | Lactococcus phage MP1 | Lactococcus phage MP1 Lactococcus virus MP1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32575 | 35.008 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346250 | Lactococcus phage MW18S | Lactococcus phage MW18S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31877 | 34.774 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346249 | Lactococcus phage 16W23 | Lactococcus phage 16W23 Lactococcus virus 16W23 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32477 | 34.76 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346248 | Lactococcus phage 17W12M | Lactococcus phage 17W12M unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33017 | 34.888 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346247 | Lactococcus phage 17W11 | Lactococcus phage 17W11 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30984 | 34.937 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346246 | Lactococcus phage 11W16L | Lactococcus phage 11W16L unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29964 | 34.752 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346245 | Lactococcus phage 10W24 | Lactococcus phage 10W24 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31866 | 34.912 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346244 | Lactococcus phage 6W18L | Lactococcus phage 6W18L unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30918 | 34.834 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346243 | Lactococcus phage 6W06 | Lactococcus phage 6W06 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32793 | 34.87 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346242 | Lactococcus phage 8R06S | Lactococcus phage 8R06S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31034 | 34.9 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346240 | Lactococcus phage 2R06A | Lactococcus phage 2R06A unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31627 | 35.207 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346239 | Lactococcus phage 2R15S2 | Lactococcus phage 2R15S2 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31468 | 34.966 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346238 | Lactococcus phage 2R15S | Lactococcus phage 2R15S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32505 | 34.795 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346237 | Lactococcus phage 2R15M | Lactococcus phage 2R15M unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31632 | 34.914 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346236 | Lactococcus phage 2R14S | Lactococcus phage 2R14S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30165 | 34.782 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX346241 | Lactococcus phage 3R07S | Lactococcus phage 3R07S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31192 | 34.942 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG557619 | Staphylococcus phage HSA84 | Staphylococcus phage HSA84 Staphylococcus virus HSA84 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42650 | 34.567 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| MG557618 | Staphylococcus phage HSA30 | Staphylococcus phage HSA30 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140358 | 30.18 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG471392 | Salmonella phage BSP161 | Salmonella phage BSP161 Salmonella virus BSP161 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39688 | 48.72 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| MG432151 | Pseudomonas phage Delta | Pseudomonas phage Delta unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45970 | 52.258 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KY210139 | Xanthomonas phage KPhi1 | Xanthomonas phage KPhi1 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46077 | 62.817 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| MG584725 | Bacillus phage BVE2 | Bacillus phage BVE2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20021 | 33.395 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MG280946 | Salmonella phage KFS-SE1 | Salmonella phage KFS-SE1 Salmonella virus KFSSE1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59715 | 56.711 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| MH179478 | Pseudomonas phage 67PfluR64PP | Pseudomonas phage 67PfluR64PP unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40748 | 55.907 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MH179475 | Pseudomonas phage 71PfluR64PP | Pseudomonas phage 71PfluR64PP unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40582 | 55.889 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MH179473 | Aeromonas phage 25AhydR2PP | Aeromonas phage 25AhydR2PP Aeromonas virus 25AhydR2PP Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 54.897 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| MH179472 | Pseudomonas phage 22PfluR64PP | Pseudomonas phage 22PfluR64PP Pseudomonas virus 22PfluR64PP Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40822 | 55.806 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MH730557 | Vibrio phage fNo16 | Vibrio phage fNo16 unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10594 | 47.357 | Unclassified | Unclassified | Corticoviridae | Vibrio |
| MG945909 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6925 | 39.668 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945870 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5757 | 48.237 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945869 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5392 | 49.258 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945868 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5305 | 50.179 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945867 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5567 | 48.536 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945866 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5338 | 31.285 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945865 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5511 | 30.503 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945864 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5015 | 46.082 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945863 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4700 | 51.34 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945862 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4859 | 46.429 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945861 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4627 | 52.107 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945860 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4316 | 49.583 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945859 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5390 | 31.558 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945858 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5259 | 47.385 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945857 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5425 | 30.396 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945856 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4650 | 39.892 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945855 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5500 | 48.055 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945854 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4759 | 48.939 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945853 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4900 | 46.673 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945852 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 44.271 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945851 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5010 | 40 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945850 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5827 | 41.651 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945849 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5296 | 45.374 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945848 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 44.357 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945847 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5557 | 42.739 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945846 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4664 | 42.324 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945845 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5745 | 48.982 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945844 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5291 | 33.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945843 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4696 | 57.751 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945842 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3963 | 52.94 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945841 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4701 | 57.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945840 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4552 | 56.854 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945839 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4383 | 59.457 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945838 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4521 | 45.455 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945837 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4734 | 49.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945836 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4360 | 52.89 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945835 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4666 | 51.329 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945834 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4615 | 46.717 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945833 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4027 | 50.658 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945832 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5065 | 39.585 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945831 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6589 | 37.563 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945830 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5614 | 41.486 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945829 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6868 | 39.531 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945828 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6662 | 41.429 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945827 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4700 | 47.149 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945826 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4477 | 39.245 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945825 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5725 | 40.541 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945824 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6238 | 40.83 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945823 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6028 | 38.852 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945822 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5051 | 35.32 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945821 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4813 | 40.785 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945820 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5063 | 49.299 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945819 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5302 | 47.661 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945818 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5492 | 41.715 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945817 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4737 | 42.537 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945816 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4902 | 45.43 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945815 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5974 | 43.321 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945814 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4884 | 36.364 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945813 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5922 | 45.829 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945812 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5732 | 40.579 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945811 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5012 | 41.54 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945810 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4686 | 46.009 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945809 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5496 | 42.977 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945808 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5304 | 45.965 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945807 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4361 | 41.756 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945806 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4410 | 39.161 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945805 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5052 | 38.242 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945804 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6406 | 39.51 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945803 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7259 | 38.587 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945802 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5224 | 43.07 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945801 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5190 | 47.168 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945800 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6286 | 42.682 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945799 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6253 | 43.211 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945798 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4764 | 39.274 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945797 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5562 | 48.723 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945796 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4989 | 46.603 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945795 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5799 | 35.127 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945794 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5733 | 38.025 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945793 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4803 | 44.743 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945792 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5805 | 46.994 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945791 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6904 | 35.269 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945790 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5914 | 45.079 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945789 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6772 | 42.661 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945788 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5882 | 35.447 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945787 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5008 | 44.509 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945786 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5164 | 45.372 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945785 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4683 | 46.744 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945784 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5870 | 45.468 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945783 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5102 | 46.08 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945782 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4470 | 40.761 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945781 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5572 | 44.311 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945780 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5468 | 47.11 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945779 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4716 | 39.546 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945778 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5401 | 40.789 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945777 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4978 | 37.646 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945776 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6480 | 37.407 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945775 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4923 | 37.843 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945774 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7173 | 39.774 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945773 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5186 | 39.896 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945772 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6363 | 34.669 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945771 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5302 | 47.548 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945770 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5060 | 44.921 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945769 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6569 | 39.9 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945768 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4916 | 44.996 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945767 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5656 | 34.972 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945766 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6310 | 44.01 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945765 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5073 | 47.467 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945764 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4855 | 39.011 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945763 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5855 | 37.541 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945762 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6895 | 45.25 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945761 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4710 | 45.966 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945760 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5017 | 41.24 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945759 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5048 | 41.244 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945758 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4715 | 44.942 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945757 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6426 | 39.169 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945756 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5044 | 41.534 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945755 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6683 | 41.104 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945754 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4945 | 39.373 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945753 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5886 | 44.818 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945752 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6584 | 39.642 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945751 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5364 | 38.945 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945750 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6629 | 37.728 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945749 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4827 | 40.066 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945748 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6288 | 43.305 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945747 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4590 | 41.634 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945746 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5289 | 45.926 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945745 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6993 | 39.382 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945744 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4998 | 45.698 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945743 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5882 | 36.569 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945742 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5624 | 40.025 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945741 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5136 | 45.814 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945740 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4997 | 42.506 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945739 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4563 | 43.984 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945738 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4731 | 39.949 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945737 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5044 | 53.251 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945736 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4980 | 45.663 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945735 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6577 | 39.79 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945734 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5887 | 41.821 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945733 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4427 | 38.197 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945732 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6116 | 39.83 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945731 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4925 | 44 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945730 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5868 | 43.763 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945729 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4705 | 39.469 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945728 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4707 | 41.173 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945727 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4647 | 46.008 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945726 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4897 | 39.902 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945725 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6289 | 40.038 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945724 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5100 | 41.255 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945723 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6791 | 36.902 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945722 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6137 | 33.388 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945721 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5105 | 48.697 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945720 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5235 | 40.267 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945719 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5019 | 39.809 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945718 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4948 | 40.724 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945717 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4545 | 44.224 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945716 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 44.933 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945715 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5427 | 36.908 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945714 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5102 | 49.863 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945713 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5882 | 45.104 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945712 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4466 | 38.356 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945711 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4902 | 49.245 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945710 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4846 | 47.338 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945709 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5934 | 43.091 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945708 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5170 | 46.499 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945707 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5283 | 40.734 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945706 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4742 | 42.535 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945705 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4946 | 43.975 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945704 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4774 | 45.161 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945703 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5120 | 44.023 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945702 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4934 | 47.426 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945701 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4883 | 44.583 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945700 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5006 | 41.47 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945699 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4909 | 45.773 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945698 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6095 | 41.526 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945697 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5895 | 41.561 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945696 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6170 | 45.122 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945695 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5712 | 49.072 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945694 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5967 | 44.16 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945693 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5399 | 42.749 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945692 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5887 | 41.821 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945691 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4905 | 45.138 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945690 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5900 | 41.153 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945689 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5382 | 51.765 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945688 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6473 | 33.91 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945687 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4768 | 40.667 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945686 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5045 | 41.407 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945685 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7099 | 39.245 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945684 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4972 | 41.834 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945683 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5665 | 43.619 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945682 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4280 | 55.164 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945681 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5008 | 41.174 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945680 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 44.933 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945679 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4860 | 47.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945678 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5192 | 32.3 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945677 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5633 | 37.103 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945676 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4777 | 39.858 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945675 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5022 | 44.066 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945674 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4946 | 43.934 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945673 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4906 | 45.536 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945672 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5379 | 35.917 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945671 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4922 | 43.966 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945670 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4404 | 55.813 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945669 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4342 | 37.54 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945668 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4726 | 40.14 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945667 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4952 | 39.54 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945666 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6887 | 38.885 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945665 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4592 | 38.284 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945664 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6039 | 45.024 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945663 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5021 | 49.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945662 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6121 | 46.61 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945647 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5381 | 35.588 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945646 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4919 | 42.427 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945645 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4998 | 44.198 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945644 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4855 | 42.451 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945643 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6194 | 44.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945642 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5758 | 44.251 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945641 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5687 | 35.713 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945640 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4802 | 36.651 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945639 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5385 | 40.223 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945638 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6519 | 38.395 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945637 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6296 | 43.091 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945636 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6002 | 35.755 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945635 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5144 | 34.176 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945634 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6244 | 44.971 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945633 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5695 | 45.549 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945632 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5884 | 43.916 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945631 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5611 | 42.524 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945630 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5564 | 38.048 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945629 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5721 | 44.992 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945628 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4777 | 51.246 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945627 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5367 | 38.401 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945626 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5170 | 46.112 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945625 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5743 | 41.494 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945624 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5769 | 38.447 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945623 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6248 | 39.677 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945622 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5419 | 41.318 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945621 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4766 | 42.048 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945620 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5079 | 45.895 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945619 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5346 | 36.626 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945618 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5948 | 40.249 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945617 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4966 | 46.899 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945616 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6027 | 40.966 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945615 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5456 | 35.887 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945614 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5996 | 44.947 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945613 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4548 | 38.171 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945612 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4388 | 37.215 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945611 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5849 | 40.793 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945610 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4336 | 43.173 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945609 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4710 | 42.951 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945608 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4940 | 47.713 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945607 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5317 | 39.496 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945606 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6082 | 43.686 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945605 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5689 | 40.288 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945604 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5069 | 39.633 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945603 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4988 | 41.079 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945602 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5038 | 44.442 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945601 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4937 | 38.242 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945600 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5325 | 36.939 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945599 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5104 | 39.773 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945598 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4979 | 42.74 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945597 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5256 | 45.624 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945596 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4845 | 44.83 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945595 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5760 | 36.528 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945594 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4916 | 54.292 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945593 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6069 | 34.404 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945592 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4883 | 41.081 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945591 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5089 | 53.96 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945590 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5415 | 43.73 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945589 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4960 | 47.48 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945588 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6200 | 36.79 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945587 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4860 | 43.807 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945586 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4928 | 43.202 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945585 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6205 | 38.276 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945584 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4459 | 39.673 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945583 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4774 | 36.971 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945582 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5740 | 36.08 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945581 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5922 | 40.831 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945580 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6537 | 47.147 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945579 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4724 | 35.605 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945578 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5840 | 41.182 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945577 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4648 | 37.715 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945576 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5684 | 45.936 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945575 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6589 | 43.148 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945574 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5514 | 44.487 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945573 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4963 | 40.701 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945572 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6133 | 40.698 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945571 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4464 | 39.315 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945570 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4864 | 43.956 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945569 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5621 | 49.013 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945568 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4714 | 40.348 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945567 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5793 | 43.363 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945566 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5133 | 44.243 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945565 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5058 | 46.085 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945564 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5703 | 49.167 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945563 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6094 | 40.154 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945562 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4572 | 40.486 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945561 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5053 | 44.033 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945560 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4933 | 44.679 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945559 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5076 | 46.414 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945558 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6149 | 40.511 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945557 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6032 | 41.048 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945556 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4458 | 37.865 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945555 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5394 | 38.561 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945554 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6618 | 37.791 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945553 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4976 | 41.6 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945552 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4477 | 50.972 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945551 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4935 | 39.98 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945550 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5490 | 42.386 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945549 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4480 | 41.094 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945548 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4986 | 46.27 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945547 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4400 | 44.318 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945546 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4296 | 44.181 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945545 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5092 | 39.336 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945544 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6902 | 37.018 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945543 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5036 | 53.197 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945542 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5005 | 43.117 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945541 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4916 | 42.331 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945540 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4852 | 37.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945539 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5616 | 46.688 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945538 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5241 | 45.507 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945537 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5318 | 43.174 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945536 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4699 | 39.2 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945535 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5612 | 41.269 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945534 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4767 | 46.151 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945533 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6293 | 39.282 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945532 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4851 | 52.298 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945531 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5439 | 42.103 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945530 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5193 | 43.019 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945529 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4606 | 41.424 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945528 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4801 | 41.97 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945527 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6394 | 36.347 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945526 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5006 | 35.318 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945525 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5080 | 38.583 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945524 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4455 | 38.025 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945523 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5614 | 41.112 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945522 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5707 | 36.779 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945521 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6330 | 40.664 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945520 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4991 | 40.834 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945519 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5590 | 42.755 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945518 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6904 | 41.367 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945517 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4985 | 44.955 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945516 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5469 | 29.914 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945515 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5948 | 47.394 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945514 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6042 | 43.595 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945513 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6181 | 38.699 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945512 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4783 | 40.623 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945511 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5973 | 46.626 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945510 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5022 | 37.495 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945509 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5803 | 44.098 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945508 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5030 | 36.978 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945507 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4793 | 39.808 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945506 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4930 | 39.067 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945505 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5591 | 41.764 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945504 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4703 | 45.652 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945503 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5535 | 42.349 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945502 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5562 | 36.228 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945501 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4451 | 39.25 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945500 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4579 | 35.532 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945499 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5128 | 43.916 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945498 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5663 | 44.994 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945497 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5052 | 43.29 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945496 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5427 | 40.778 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945495 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5923 | 37.852 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945494 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6101 | 38.781 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945493 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6228 | 41.522 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945492 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5634 | 40.149 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945491 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4861 | 39.477 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945490 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4506 | 38.26 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945489 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4760 | 36.828 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945488 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5986 | 40.01 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945487 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5308 | 41.541 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945486 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4911 | 46.447 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945485 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5773 | 34.817 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945484 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4613 | 43.594 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945483 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5962 | 38.796 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945482 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6161 | 41.6 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945481 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4254 | 40.103 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945480 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5651 | 44.222 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945479 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4800 | 36.25 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945478 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5985 | 45.564 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945477 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5061 | 48.805 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945476 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5052 | 31.888 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945475 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 47.399 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945474 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 54.891 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945473 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4370 | 48.581 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945472 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4518 | 45.529 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945471 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7885 | 35.32 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945470 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6709 | 42.749 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945469 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6615 | 36.901 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945468 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4476 | 38.472 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945467 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5083 | 50.246 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945466 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6112 | 38.285 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945465 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5119 | 46.376 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945464 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5948 | 39.509 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945463 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4934 | 38.974 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945462 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5924 | 37.829 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945461 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5241 | 43.236 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945460 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5417 | 37.419 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945459 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4926 | 43.179 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945458 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6258 | 40.988 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945457 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6140 | 38.974 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945456 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5858 | 45.015 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945455 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5859 | 44.188 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945454 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5274 | 40.14 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945453 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5837 | 45.451 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945452 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5728 | 44.955 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945451 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 8312 | 37.68 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945450 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5219 | 42.939 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945449 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6056 | 48.085 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945448 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6471 | 43.131 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945447 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5900 | 46.068 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945446 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6260 | 37.476 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945445 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5824 | 41.157 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945444 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5028 | 44.73 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945443 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6208 | 38.611 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945442 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6285 | 38.632 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945441 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5245 | 42.078 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945440 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5259 | 37.65 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945439 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4783 | 51.077 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945438 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4842 | 46.097 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945437 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5507 | 38.006 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945436 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6331 | 38.114 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945435 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4800 | 42.458 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945434 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6456 | 40.273 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945433 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6079 | 42.145 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945432 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6267 | 41.36 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945431 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4675 | 41.07 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945430 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5668 | 44.866 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945429 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5104 | 42.927 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945428 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4780 | 42.343 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945427 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5785 | 44.494 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945426 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5637 | 45.769 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945425 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6799 | 42.742 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945424 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5021 | 42.92 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945423 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5313 | 47.977 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945422 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6917 | 39.005 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945421 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5975 | 37.556 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945420 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6405 | 42.81 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945419 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6118 | 38.542 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945418 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4685 | 43.074 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945417 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6109 | 38.844 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945416 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5385 | 46.388 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945415 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5022 | 45.639 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945414 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5836 | 41.57 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945413 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4859 | 39.967 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945412 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6827 | 41.277 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945411 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6527 | 37.628 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945410 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5385 | 46.388 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945409 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5775 | 31.515 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945408 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6522 | 39.482 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945407 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4932 | 43.917 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945406 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6632 | 43.139 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945405 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4658 | 46.179 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945404 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6479 | 37.382 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945403 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4756 | 41.884 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945402 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4988 | 44.166 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945401 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7021 | 39.496 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945400 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6196 | 42.673 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945399 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6330 | 40.269 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945398 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5025 | 42.527 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945397 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5730 | 39.25 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945396 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6230 | 44.767 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945395 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5791 | 39.492 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945394 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5345 | 45.987 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945393 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4964 | 42.143 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945392 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5634 | 41.072 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945391 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5367 | 47.475 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945390 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6762 | 35.507 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945389 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5202 | 46.175 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945388 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5664 | 45.392 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945387 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4832 | 46.316 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945386 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5635 | 37.657 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945385 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5208 | 46.409 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945384 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5947 | 41.786 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945383 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5841 | 43.571 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945382 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5785 | 45.687 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945381 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5503 | 47.683 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945365 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4847 | 38.147 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945364 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5017 | 39.227 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945363 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4815 | 43.074 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945362 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5585 | 35.237 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945361 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5057 | 50.168 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945360 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5990 | 45.91 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945359 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4687 | 40.431 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945358 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6243 | 41.855 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945357 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4915 | 39.471 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945356 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6477 | 39.802 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945355 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4947 | 42.046 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945354 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6352 | 34.95 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945353 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6177 | 42.432 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945352 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6694 | 39.647 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945351 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5466 | 38.767 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945350 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6602 | 40.563 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945349 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5877 | 37.417 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945348 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5065 | 48.095 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945347 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5373 | 45.84 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945346 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4888 | 44.558 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945345 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5345 | 40.206 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945344 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4447 | 39.6 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945343 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4440 | 39.527 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945342 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4954 | 52.422 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945341 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4942 | 40.429 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945340 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4904 | 43.699 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945339 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5099 | 43.185 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945338 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5790 | 35.699 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945337 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6494 | 39.498 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945336 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5831 | 34.985 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945335 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5399 | 39.563 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945334 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5633 | 43.724 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945333 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4884 | 45.188 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945332 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7203 | 39.345 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945331 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4732 | 45.816 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945330 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4774 | 44.533 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945329 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4774 | 41.328 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945328 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5499 | 36.407 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945327 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6499 | 39.298 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945326 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6060 | 30.71 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945325 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4865 | 47.605 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945324 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4577 | 43.325 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945323 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7391 | 39.386 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945322 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4314 | 42.721 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945321 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6378 | 38.962 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945320 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5064 | 49.19 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945319 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5213 | 46.71 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945318 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4885 | 43.071 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945317 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5278 | 37.798 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945316 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4433 | 38.665 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945315 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5130 | 41.267 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945314 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6245 | 37.182 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945313 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4314 | 42.675 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945312 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5318 | 40.391 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945311 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5356 | 37.509 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945310 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5250 | 41.619 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945309 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6176 | 43.264 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945308 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5221 | 42.214 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945307 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5488 | 43.331 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945306 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5331 | 37.254 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945286 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5332 | 34.734 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945285 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5537 | 38.956 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945284 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6216 | 36.116 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945283 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5297 | 55.579 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945282 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6013 | 40.063 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945281 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5913 | 43.819 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945280 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5076 | 34.89 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945279 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6651 | 37.919 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945278 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5430 | 39.006 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945277 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5100 | 43.039 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945276 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6827 | 42.581 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945275 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6914 | 43.824 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945266 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6115 | 35.061 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945265 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4797 | 48.968 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945264 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6084 | 51.545 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945263 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4781 | 37.691 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945262 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6049 | 38.601 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945261 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6769 | 38.174 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945260 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5655 | 36.145 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945259 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6435 | 42.859 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945258 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7454 | 41.079 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945257 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6235 | 46.945 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945256 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6576 | 41.271 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945255 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7027 | 40.729 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945254 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6079 | 45.09 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945253 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6329 | 36.625 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945252 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4462 | 38.906 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945241 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6384 | 41.886 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945240 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5475 | 44.621 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945239 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6483 | 40.753 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945238 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5060 | 49.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945237 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4765 | 46.065 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945236 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5200 | 41.038 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945235 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5636 | 41.43 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945234 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4772 | 46.794 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945233 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6127 | 46.189 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945232 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5101 | 45.834 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945231 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4994 | 48.338 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945230 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5631 | 33.653 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945229 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7064 | 39.609 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945228 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4907 | 39.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945227 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5153 | 45.779 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945216 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6327 | 44.144 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945215 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4955 | 42.341 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945214 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5746 | 41.055 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945213 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4873 | 44.511 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945212 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6126 | 32.272 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945211 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4986 | 44.585 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945210 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6386 | 39.226 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945209 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6679 | 37.76 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945208 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4830 | 38.344 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945207 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5057 | 46.886 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945206 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4557 | 49.133 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945205 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4902 | 45.553 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945204 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4677 | 46.269 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945203 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5108 | 46.378 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945202 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4865 | 49.394 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945191 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6277 | 37.661 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945190 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4946 | 46.098 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945189 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5961 | 41.537 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945188 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7323 | 39.533 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945187 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6796 | 42.128 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945186 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5631 | 48.766 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945185 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6042 | 40.5 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945184 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5588 | 35.916 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945183 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7622 | 40.554 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945182 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6609 | 39.113 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945181 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5000 | 44.58 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945180 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6775 | 31.985 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945179 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4786 | 39.323 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945178 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4228 | 39.38 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945177 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6003 | 38.331 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945166 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4619 | 41.394 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945165 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4884 | 38.677 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945164 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4422 | 63.455 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945163 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4938 | 43.034 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945162 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5707 | 45.698 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945161 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4797 | 59.641 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945160 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6086 | 40.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945159 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7189 | 39.157 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945158 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5009 | 42.703 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945157 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5257 | 47.422 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945156 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4910 | 41.731 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945155 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6213 | 47.159 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945154 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4682 | 44.041 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945153 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4913 | 42.866 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945142 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5391 | 47.616 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945141 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5568 | 44.235 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945140 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6131 | 46.974 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945139 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5185 | 47.367 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945138 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5505 | 47.593 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945137 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6002 | 42.136 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945136 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5433 | 47.837 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945135 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5474 | 49.068 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945134 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5105 | 42.625 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945133 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6038 | 40.245 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945132 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4595 | 49.162 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945131 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3143 | 39.294 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945130 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6208 | 40.512 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945129 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4511 | 50.964 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG945128 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7099 | 39.287 | Unclassified | Unclassified | Microviridae | Unspecified |
| LR027391 | Escherichia virus vB_Eco_mar005P1 | Escherichia virus vB_Eco_mar005P1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167658 | 37.717 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| LR027390 | Escherichia virus vB_Eco_mar005P1 | Escherichia virus vB_Eco_mar005P1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167773 | 37.724 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| LR027389 | Escherichia phage vB_Eco_mar003J3 | Escherichia phage vB_Eco_mar003J3 Escherichia virus mar003J3 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115471 | 39.775 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| LR027388 | Escherichia phage vB_Eco_mar001J1 | Escherichia phage vB_Eco_mar001J1 Escherichia virus mar001J1 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50343 | 44.395 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LR027387 | Escherichia virus vB_Eco_mar005P1 | Escherichia virus vB_Eco_mar005P1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167773 | 37.725 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| LR027386 | Escherichia virus vB_Eco_mar005P1 | Escherichia virus vB_Eco_mar005P1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167773 | 37.719 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| LR027385 | Escherichia phage vB_Eco_mar001J1 | Escherichia phage vB_Eco_mar001J1 Escherichia virus mar001J1 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50343 | 44.382 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LR027384 | Escherichia phage vB_Eco_mar004NP2 | Escherichia phage vB_Eco_mar004NP2 Escherichia virus mar004NP2 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107601 | 38.944 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| LR027383 | Escherichia virus vB_Eco_mar005P1 | Escherichia virus vB_Eco_mar005P1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167651 | 37.714 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MF093736 | Shigella phage vB_SsoS-ISF002 | Shigella phage vB_SsoS-ISF002 Shigella virus ISF002 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50564 | 45.532 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| AP018932 | Escherichia phage KIT03 | Escherichia phage KIT03 Escherichia virus KIT03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166848 | 35.342 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LC375533 | Pectobacterium phage PPWS2 | Pectobacterium phage PPWS2 Pectobacterium virus PPWS2 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44716 | 50.968 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| LR031359 | Enterococcus phage 156 | Enterococcus phage 156 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141133 | 35.789 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MH817022 | Bacillus phage vB_BthP-Goe4 | Bacillus phage vB_BthP-Goe4 Bacillus virus Goe4 Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25722 | 30.433 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KJ936628 | Vibrio phage VPp1 | Vibrio phage VPp1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50431 | 41.346 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MH992123 | Escherichia phage OLB145 | Escherichia phage OLB145 unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70173 | 41.237 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MH992122 | Escherichia phage OLB35 | Escherichia phage OLB35 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169140 | 37.539 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MH992121 | Escherichia phage MLF4 | Escherichia phage MLF4 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167379 | 35.387 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH717097 | Escherichia phage C1 | Escherichia phage C1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46667 | 46.596 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MH717096 | Escherichia phage N7 | Escherichia phage N7 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38829 | 46.635 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| MH920640 | Synechococcus phage S-E7 | Synechococcus phage S-E7 unclassified Leucotheavirus Leucotheavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177622 | 39.894 | Leucotheavirus | Unclassified | Myoviridae | Synechococcus |
| MH920639 | Synechococcus phage S-P4 | Synechococcus phage S-P4 Synechococcus virus SP4 Leucotheavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158485 | 39.946 | Leucotheavirus | Unclassified | Myoviridae | Synechococcus |
| MH920638 | Synechococcus phage S-B05 | Synechococcus phage S-B05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40500 | 38.259 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| MH926057 | Corynebacterium phage Kimchi1738 | Corynebacterium phage Kimchi1738 unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66915 | 52.257 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MH926056 | Corynebacterium phage Cruella | Corynebacterium phage Cruella unclassified Ceetrepovirus Ceetrepovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66714 | 52.34 | Ceetrepovirus | Unclassified | Siphoviridae | Corynebacterium |
| MH983004 | Lactobacillus phage BH1 | Lactobacillus phage BH1 Lactobacillus virus BH1 Behunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40721 | 44.834 | Behunavirus | Unclassified | Siphoviridae | Lactobacillus |
| MH931004 | Termite gut associated microvirus 2 | Termite gut associated microvirus 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4637 | 45.913 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH931003 | Termite gut associated microvirus 1 | Termite gut associated microvirus 1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4898 | 46.978 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883726 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4694 | 48.999 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883725 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4923 | 40.524 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883724 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5770 | 44.731 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883723 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4870 | 53.368 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883722 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5003 | 37.777 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883721 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6257 | 42.64 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883720 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5903 | 41.928 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883719 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4969 | 44.254 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883718 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4837 | 40.418 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883717 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5022 | 45.639 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883715 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4942 | 40.429 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883714 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5028 | 39.757 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883713 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 7029 | 38.683 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883712 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5754 | 35.401 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883711 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5311 | 42.892 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883710 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5910 | 37.039 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG883709 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5731 | 31.391 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH992510 | Escherichia phage vB_EcoM_DalCa | Escherichia phage vB_EcoM_DalCa Escherichia virus DalCa Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166040 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH992509 | Salmonella phage vB_SenS_phi135 | Salmonella phage vB_SenS_phi135 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43142 | 49.877 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH917278 | Shigella phage phi2457T | Shigella phage phi2457T unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50219 | 45.288 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MH898687 | Edwardsiella phage Edno5 | Edwardsiella phage Edno5 unclassified Gofduovirus Gofduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40844 | 50.803 | Gofduovirus | Unclassified | Myoviridae | Edwardsiella |
| MH884514 | Bacillus phage vB_BpsM-61 | Bacillus phage vB_BpsM-61 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48160 | 43.142 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| MH884513 | Bacillus phage vB_BpsS-36 | Bacillus phage vB_BpsS-36 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50485 | 41.143 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MH884511 | Exiguobacterium phage vB_EalM-132 | Exiguobacterium phage vB_EalM-132 unclassified Spounavirinae Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145804 | 40.62 | Unclassified | Spounavirinae | Herelleviridae | Exiguobacterium |
| MH884510 | Exiguobacterium phage vB_EalM-137 | Exiguobacterium phage vB_EalM-137 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41601 | 50.431 | Unclassified | Unclassified | Myoviridae | Exiguobacterium |
| MH884509 | Bacillus phage vB_BboS-125 | Bacillus phage vB_BboS-125 Bacillus virus 125 Elmenteitavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58528 | 48.568 | Elmenteitavirus | Unclassified | Myoviridae | Bacillus |
| MH884508 | Bacillus phage vB_BcoS-136 | Bacillus phage vB_BcoS-136 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160590 | 32.163 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MH160392 | Erwinia phage phiEaP8 | Erwinia phage phiEaP8 Erwinia virus phiEaP8 Yonginvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75929 | 46.841 | Yonginvirus | Erskinevirinae | Schitoviridae | Erwinia |
| MK088078 | Marinobacter phage AS1 | Marinobacter phage AS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36994 | 57.044 | Unclassified | Unclassified | Siphoviridae | Marinobacter |
| MH929319 | Serratia phage vB_SmaA_3M | Serratia phage vB_SmaA_3M Serratia virus 3M Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159398 | 51.414 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| KX278418 | Pectobacterium phage PP16 | Pectobacterium phage PP16 Pectobacterium virus PP16 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44268 | 51.936 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MK069403 | CrAssphage sp. O-152 | CrAssphage sp. O-152 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96283 | 29.21 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MH159197 | Staphylococcus phage VB_SavM_JYL01 | Staphylococcus phage VB_SavM_JYL01 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141384 | 30.172 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY744240 | Weissella phage PWc | Weissella phage PWc Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38783 | 37.617 | Unclassified | Unclassified | Siphoviridae | Weissella |
| MH837543 | Lactobacillus phage LR2 | Lactobacillus phage LR2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43695 | 38.13 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MH837542 | Lactobacillus phage LR1 | Lactobacillus phage LR1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43290 | 39.848 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MK047718 | Escherichia phage p000y | Escherichia phage p000y unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169872 | 37.654 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK047717 | Escherichia phage p000v | Escherichia phage p000v unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167803 | 37.62 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MK005300 | Salmonella phage STG2 | Salmonella phage STG2 Salmonella virus STG2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114275 | 40.118 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| LC063634 | Pectobacterium phage PPWS1 | Pectobacterium phage PPWS1 Pectobacterium virus PPWS1 Kotilavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44539 | 51.063 | Kotilavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| LR025198 | Escherichia phage vB_Eco_SLUR26 | Escherichia phage vB_Eco_SLUR26 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60165 | 44.473 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LR025197 | Escherichia phage vB_Eco_SLUR75 | Escherichia phage vB_Eco_SLUR75 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59880 | 44.738 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LR025196 | Escherichia phage vB_Eco_SLUR63 | Escherichia phage vB_Eco_SLUR63 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58998 | 44.793 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LR025195 | Escherichia phage vB_Eco_SLUR76 | Escherichia phage vB_Eco_SLUR76 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60165 | 44.468 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LT630001 | Enterococcus phage Idefix | Enterococcus phage Idefix Enterococcus virus Idefix Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18168 | 33.201 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| MH853358 | Streptococcus phage LF4 | Streptococcus phage LF4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37421 | 37.022 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH853357 | Streptococcus phage LF3 | Streptococcus phage LF3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32205 | 40.23 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH853355 | Streptococcus phage LF1 | Streptococcus phage LF1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37421 | 36.904 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| MH809535 | Yersinia phage PYPS2T | Yersinia phage PYPS2T Yersinia virus PYPS2T Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169604 | 35.404 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| MH809533 | Burkholderia phage phiE058 | Burkholderia phage phiE058 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44121 | 64.124 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MH809532 | Burkholderia phage phiE131 | Burkholderia phage phiE131 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44121 | 64.124 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| MH880817 | Enterococcus phage EfV12-phi1 | Enterococcus phage EfV12-phi1 Enterococcus virus EfV12 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152770 | 37.02 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MH845412 | Cronobacter phage CS01 | Cronobacter phage CS01 Cronobacter virus CS01 Gyeonggidovirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48195 | 50.109 | Gyeonggidovirus | Unclassified | Drexlerviridae | Cronobacter |
| MH713599 | Acinetobacter phage KARL-1 | Acinetobacter phage KARL-1 Acinetobacter virus KARL1 Lazarusvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166560 | 36.786 | Lazarusvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| MH458951 | Bacillus phage vB_BthS_BMBphi | Bacillus phage vB_BthS_BMBphi Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49277 | 39.708 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MG948468 | Pantoea phage vB_PagS_Vid5 | Pantoea phage vB_PagS_Vid5 Pantoea virus Vid5 Vidquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61437 | 48.821 | Vidquintavirus | Unclassified | Siphoviridae | Pantoea |
| MH445381 | Escherichia phage P1 | Escherichia phage P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66750 | 47.634 | Punavirus | Unclassified | Myoviridae | Escherichia |
| MH445380 | Escherichia phage P1 | Escherichia phage P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126505 | 49.902 | Punavirus | Unclassified | Myoviridae | Escherichia |
| MH422554 | Escherichia virus P1 | Escherichia virus P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97213 | 47.915 | Punavirus | Unclassified | Myoviridae | Escherichia |
| MH884648 | Caulobacter phage Kronos | Caulobacter phage Kronos Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42424 | 66.29 | Unclassified | Unclassified | Siphoviridae | Caulobacter |
| MH825706 | Streptomyces phage Microdon | Streptomyces phage Microdon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53356 | 68.562 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH800200 | Acinetobacter phage vB_AbaP_46-62_Aci07 | Acinetobacter phage vB_AbaP_46-62_Aci07 Acinetobacter virus Aci07 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42330 | 39.135 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MH800199 | Acinetobacter phage vB_AbaM_B09_Aci02-2 | Acinetobacter phage vB_AbaM_B09_Aci02-2 Acinetobacter virus Aci02-2 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104354 | 37.182 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MH800198 | Acinetobacter phage vB_AbaM_B09_Aci01-1 | Acinetobacter phage vB_AbaM_B09_Aci01-1 Acinetobacter virus Aci01-1 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103628 | 37.186 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MH816966 | Escherichia phage FEC19 | Escherichia phage FEC19 Escherichia virus FEC19 Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68472 | 46.117 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| MH837626 | Escherichia phage vB_vPM_PD112 | Escherichia phage vB_vPM_PD112 Escherichia virus PD112 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168084 | 35.364 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH816848 | Escherichia phage vB_vPM_PD06 | Escherichia phage vB_vPM_PD06 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149506 | 37.652 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MH779515 | Mycobacterium phage Sweets | Mycobacterium phage Sweets unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41896 | 66.593 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779514 | Mycobacterium phage Paito | Mycobacterium phage Paito unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42311 | 66.023 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779513 | Mycobacterium phage Olga | Mycobacterium phage Olga unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41902 | 66.591 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779509 | Mycobacterium phage Kasen3 | Mycobacterium phage Kasen3 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41890 | 66.579 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779505 | Mycobacterium phage Grizzly | Mycobacterium phage Grizzly unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41752 | 66.758 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| MH779619 | Enterobacteria phage vB_EcoP_IME390 | Enterobacteria phage vB_EcoP_IME390 Enterobacteria virus IME390 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38396 | 49.065 | Teseptimavirus | Studiervirinae | Autographiviridae | Enterobacteria |
| MH763831 | Acinetobacter phage vB_AbaP_B09_Aci08 | Acinetobacter phage vB_AbaP_B09_Aci08 Acinetobacter virus Aci08 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42067 | 39.285 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MH751506 | Escherichia phage T2 | Escherichia phage T2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163832 | 35.317 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH746814 | Acinetobacter phage vB_AbaM_B09_Aci05 | Acinetobacter phage vB_AbaM_B09_Aci05 Acinetobacter virus Aci05 Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102789 | 37.207 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| MH752385 | Bacillus phage Ray17 | Bacillus phage Ray17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43733 | 44.552 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MH729874 | Klebsiella phage KP179 | Klebsiella phage KP179 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162630 | 39.346 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MH717099 | Escherichia phage C5 | Escherichia phage C5 Escherichia virus C5 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39551 | 48.815 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MH717098 | Escherichia phage N30 | Escherichia phage N30 Escherichia virus N30 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39266 | 48.615 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MH389777 | Dickeya phage vB_DsoM_JA11 | Dickeya phage vB_DsoM_JA11 Dickeya virus JA11 Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 255356 | 44.47 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| MG589387 | Enterobacter phage phiT5282H | Enterobacter phage phiT5282H Enterobacter virus phiT5282H Novemvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31978 | 52.458 | Novemvirus | Peduovirinae | Myoviridae | Enterobacter |
| MG589386 | Pseudomonas phage phiPA01_EW | Pseudomonas phage phiPA01_EW unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46403 | 52.456 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MG589385 | Pseudomonas phage phiPA01_302 | Pseudomonas phage phiPA01_302 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46093 | 52.524 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| MG589384 | Enterobacter phage phi63_307 | Enterobacter phage phi63_307 unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85263 | 43.777 | Kolesnikvirus | Ounavirinae | Myoviridae | Enterobacter |
| MG589383 | Shigella phage phi25-307 | Shigella phage phi25-307 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167544 | 37.55 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| KT184316 | Enterobacteria phage XTG1 | Enterobacteria phage XTG1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89635 | 38.9 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| KT184315 | Enterobacteria phage KhF3 | Enterobacteria phage KhF3 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88016 | 38.862 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| KT184314 | Enterobacteria phage KhF2 | Enterobacteria phage KhF2 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88309 | 38.83 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| KT184313 | Enterobacteria phage KhF1 | Enterobacteria phage KhF1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88356 | 38.773 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| KT184312 | Enterobacteria phage Kha5h | Enterobacteria phage Kha5h Enterobacteria virus Kha5h Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167318 | 35.548 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KT184311 | Enterobacteria phage GiZh | Enterobacteria phage GiZh Enterobacteria virus GiZh Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166514 | 35.451 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KT184310 | Enterobacteria phage ATK48 | Enterobacteria phage ATK48 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169729 | 37.644 | Mosigvirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KT184309 | Enterobacteria phage ATK47 | Enterobacteria phage ATK47 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170020 | 37.649 | Mosigvirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KT184308 | Enterobacteria phage Aplg8 | Enterobacteria phage Aplg8 Enterobacteria virus Aplg8 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168496 | 35.533 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MG000860 | Bacillus phage BSP9 | Bacillus phage BSP9 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155958 | 42.008 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF630922 | Escherichia phage IMM-001 | Escherichia phage IMM-001 unclassified Guelphvirus Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32486 | 44.318 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MF630921 | Escherichia phage IMM-002 | Escherichia phage IMM-002 Escherichia virus IMM002 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40060 | 53.19 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| AP017903 | Azobacteroides phage ProJPt-Bp1 | Azobacteroides phage ProJPt-Bp1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99517 | 42.309 | Unclassified | Unclassified | Podoviridae | Azobacteroides |
| MH363708 | Escherichia phage C130_2 | Escherichia phage C130_2 Escherichia virus C1302 Giessenvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41775 | 55.43 | Giessenvirus | Unclassified | Podoviridae | Escherichia |
| MH725810 | Pseudomonas phage PaYy-2 | Pseudomonas phage PaYy-2 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92348 | 49.333 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MH719082 | Escherichia phage vB_EcoM_IME392 | Escherichia phage vB_EcoM_IME392 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116460 | 45.43 | Unclassified | Unclassified | Myoviridae | Escherichia |
| MH384260 | Staphylococcus phage SA780ruMSSAST101 | Staphylococcus phage SA780ruMSSAST101 Staphylococcus virus SA780ruMSSAST101 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43184 | 33.554 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MH717709 | Escherichia phage LL2 | Escherichia phage LL2 Escherichia virus LL2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39382 | 50.34 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| MH727593 | Acidovorax phage ACPWH | Acidovorax phage ACPWH Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42499 | 64.437 | Unclassified | Unclassified | Siphoviridae | Acidovorax |
| MH733496 | Escherichia phage PGN829.1 | Escherichia phage PGN829.1 unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74437 | 42.879 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MH310935 | Vibrio phage ICP1_2011_B | Vibrio phage ICP1_2011_B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125128 | 37.19 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MH310933 | Vibrio phage ICP1_2011_A | Vibrio phage ICP1_2011_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126861 | 37.137 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MH807819 | Pectobacterium phage Slant | Pectobacterium phage Slant unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44149 | 48.957 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH807818 | Pectobacterium phage Nobby | Pectobacterium phage Nobby Pectobacterium virus Nobby Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42818 | 48.996 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH807816 | Pectobacterium phage Momine | Pectobacterium phage Momine unclassified Phimunavirus Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43402 | 49.039 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH807814 | Pectobacterium phage Lelidair | Pectobacterium phage Lelidair Pectobacterium virus Lelidair Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43686 | 48.659 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH807811 | Pectobacterium phage Gaspode | Pectobacterium phage Gaspode Pectobacterium virus Gaspode Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44364 | 48.808 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MH675927 | Escherichia phage vB_vPM_PD114 | Escherichia phage vB_vPM_PD114 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150354 | 37.543 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MH669274 | Escherichia phage PD38 | Escherichia phage PD38 unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72213 | 42.884 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MH487649 | Bacillus phage BC01 | Bacillus phage BC01 unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158385 | 39.895 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH752840 | Escherichia phage vB_EcoM-Pr121LW | Escherichia phage vB_EcoM-Pr121LW unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134575 | 43.582 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MG746602 | Klebsiella phage vB_Kpn_F48 | Klebsiella phage vB_Kpn_F48 Klebsiella virus F48 Marfavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170764 | 40.783 | Marfavirus | Unclassified | Myoviridae | Klebsiella |
| MH674343 | Methanobacterium virus Drs3 | Methanobacterium virus Drs3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37129 | 41.183 | Unclassified | Unclassified | Siphoviridae | Methanobacterium |
| KY981272 | Curvibacter phage P26059B | Curvibacter phage P26059B Curvibacter virus P26059B Kalppathivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41344 | 54.252 | Kalppathivirus | Unclassified | Autographiviridae | Curvibacter |
| KY981271 | Curvibacter phage P26059A | Curvibacter phage P26059A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84008 | 43.573 | Unclassified | Unclassified | Siphoviridae | Curvibacter |
| MG967618 | Bacillus phage v_B-Bak10 | Bacillus phage v_B-Bak10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82931 | 33.81 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MG967617 | Bacillus phage v_B-Bak6 | Bacillus phage v_B-Bak6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80764 | 33.927 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MG967616 | Bacillus phage v_B-Bak1 | Bacillus phage v_B-Bak1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80764 | 33.928 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MH460462 | Dickeya phage vB_DsoM_JA33 | Dickeya phage vB_DsoM_JA33 unclassified Salmondvirus Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 255356 | 44.465 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| MH460463 | Dickeya phage vB_DsoM_AD1 | Dickeya phage vB_DsoM_AD1 Dickeya virus AD1 Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261658 | 49.366 | Alexandravirus | Unclassified | Myoviridae | Dickeya |
| MH460461 | Dickeya phage vB_DsoM_JA29 | Dickeya phage vB_DsoM_JA29 Dickeya virus JA29 Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 253323 | 43.843 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| MH460460 | Dickeya phage vB_DsoM_JA13 | Dickeya phage vB_DsoM_JA13 unclassified Salmondvirus Salmondvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 254061 | 44.535 | Salmondvirus | Unclassified | Myoviridae | Dickeya |
| MH460459 | Dickeya phage vB_DsoP_JA10 | Dickeya phage vB_DsoP_JA10 Dickeya virus JA10 Ningirsuvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40131 | 51.451 | Ningirsuvirus | Studiervirinae | Autographiviridae | Dickeya |
| MH327491 | Marine virus AG-345-A17 | Marine virus AG-345-A17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11424 | 34.506 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MH327490 | Marine virus AG-345-A17 | Marine virus AG-345-A17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16365 | 31.738 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| MH327487 | Marine virus AG-341-P01 | Marine virus AG-341-P01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13641 | 38.362 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH327486 | Marine virus AG-341-P01 | Marine virus AG-341-P01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14735 | 36.736 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH327485 | Marine virus AG-341-P01 | Marine virus AG-341-P01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18878 | 36.307 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319765 | Marine virus AG-345-P14 | Marine virus AG-345-P14 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67213 | 34.766 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319764 | Marine virus AG-345-P14 | Marine virus AG-345-P14 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79455 | 36.389 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319760 | Marine virus AG-345-O18 | Marine virus AG-345-O18 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11481 | 33.96 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319759 | Marine virus AG-345-O18 | Marine virus AG-345-O18 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36824 | 34.755 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319758 | Marine virus AG-345-O18 | Marine virus AG-345-O18 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48117 | 36.463 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319755 | Marine virus AG-345-L05 | Marine virus AG-345-L05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32905 | 35.171 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319754 | Marine virus AG-345-L05 | Marine virus AG-345-L05 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149201 | 34.832 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319753 | Marine virus AG-345-F15 | Marine virus AG-345-F15 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13583 | 32.423 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319752 | Marine virus AG-345-F15 | Marine virus AG-345-F15 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19248 | 35.754 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319751 | Marine virus AG-345-F15 | Marine virus AG-345-F15 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24865 | 36.787 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319750 | Marine virus AG-345-F15 | Marine virus AG-345-F15 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35407 | 35.329 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319749 | Marine virus AG-345-F15 | Marine virus AG-345-F15 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139752 | 34.855 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319746 | Marine virus AG-345-E15 | Marine virus AG-345-E15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14292 | 36.811 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319745 | Marine virus AG-345-E15 | Marine virus AG-345-E15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15781 | 33.002 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319743 | Marine virus AG-345-E15 | Marine virus AG-345-E15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16760 | 34.881 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319742 | Marine virus AG-345-E15 | Marine virus AG-345-E15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35937 | 36.539 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319741 | Marine virus AG-345-E15 | Marine virus AG-345-E15 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66256 | 36.28 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319738 | Marine virus AG-345-E08 | Marine virus AG-345-E08 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22821 | 37.803 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319737 | Marine virus AG-345-E08 | Marine virus AG-345-E08 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35082 | 37.224 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319734 | Marine virus AG-345-E02 | Marine virus AG-345-E02 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13970 | 35.225 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319733 | Marine virus AG-345-E02 | Marine virus AG-345-E02 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22073 | 33.502 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319732 | Marine virus AG-345-E02 | Marine virus AG-345-E02 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69654 | 36.309 | Salacisavirus | Unclassified | Myoviridae | Unspecified |
| MH319725 | Marine virus AG-345-D19 | Marine virus AG-345-D19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10654 | 38.971 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319724 | Marine virus AG-345-D19 | Marine virus AG-345-D19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11603 | 37.085 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319723 | Marine virus AG-345-D19 | Marine virus AG-345-D19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13629 | 38.646 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319722 | Marine virus AG-345-D19 | Marine virus AG-345-D19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15952 | 35.832 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MH319720 | Marine virus AG-345-D19 | Marine virus AG-345-D19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66178 | 37.754 | Unclassified | Unclassified | Myoviridae | Unspecified |
| KR049063 | Enterococcus phage EFLK1 | Enterococcus phage EFLK1 Enterococcus virus EFLK1 Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130952 | 35.888 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| KP339049 | Enterococcus phage EFDG1 | Enterococcus phage EFDG1 Enterococcus virus EFDG1 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147589 | 37.203 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MH136584 | Staphylococcus phage HYZ21 | Staphylococcus phage HYZ21 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139675 | 30.423 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MF623055 | Pseudomonas phage BrSP1 | Pseudomonas phage BrSP1 Pseudomonas virus BrSP1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66189 | 55.68 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MF468274 | Shigella phage Sfin-1 | Shigella phage Sfin-1 Shigella virus Sfin1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50403 | 45.202 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MH310936 | Vibrio phage ICP1_2012_A | Vibrio phage ICP1_2012_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121418 | 37.174 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MH310934 | Vibrio phage ICP1_2006_E | Vibrio phage ICP1_2006_E Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128298 | 37.156 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MH412654 | Synechococcus phage S-T4 | Synechococcus phage S-T4 Synechococcus virus ST4 Tamkungvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181082 | 38.927 | Tamkungvirus | Unclassified | Myoviridae | Synechococcus |
| MH424446 | Salmonella virus VSt472 | Salmonella virus VSt472 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46905 | 45.643 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MH424445 | Salmonella virus VSt10 | Salmonella virus VSt10 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41581 | 50.032 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH424444 | Salmonella virus VSiP | Salmonella virus VSiP unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43110 | 51.211 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH424443 | Salmonella virus VSe103 | Salmonella virus VSe103 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42262 | 50.002 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH588546 | Caulobacter phage CcrBL9 | Caulobacter phage CcrBL9 Caulobacter virus CcrBL9 Bertelyvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 322272 | 63.679 | Bertelyvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| MH588545 | Caulobacter phage CcrPW | Caulobacter phage CcrPW Caulobacter virus CcrPW Colossusvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 308141 | 62.242 | Colossusvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| MH588544 | Caulobacter phage CcrBL10 | Caulobacter phage CcrBL10 Caulobacter virus CcrBL10 Poindextervirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 220934 | 65.687 | Poindextervirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| MH633487 | Klebsiella phage NJR15 | Klebsiella phage NJR15 Klebsiella virus NJR15 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49468 | 51.043 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH633486 | Klebsiella phage NJS3 | Klebsiella phage NJS3 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49387 | 51.084 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH633485 | Klebsiella phage NJS2 | Klebsiella phage NJS2 Klebsiella virus NJS2 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50132 | 50.786 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH633484 | Klebsiella phage TAH8 | Klebsiella phage TAH8 Klebsiella virus TAH8 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49344 | 51.092 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH638312 | Bacillus phage Kioshi | Bacillus phage Kioshi unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165676 | 39.481 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH638311 | Bacillus phage OmnioDeoPrimus | Bacillus phage OmnioDeoPrimus unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161833 | 38.765 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH638310 | Bacillus phage Kamfam | Bacillus phage Kamfam unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161830 | 37.886 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH638309 | Bacillus phage Hobo | Bacillus phage Hobo unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165213 | 39.605 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH645904 | Vibrio phage vB_VpS_PG07 | Vibrio phage vB_VpS_PG07 Vibrio virus PG07 Pogseptimavirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112106 | 43.65 | Pogseptimavirus | Unclassified | Demerecviridae | Vibrio |
| MH675552 | Bacteroides phage crAss001 | Bacteroides phage crAss001 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 102679 | 34.743 | Unclassified | Unclassified | Podoviridae | Bacteroides |
| MH375644 | Vibrio phage YC | Vibrio phage YC Vibrio virus YC Campanilevirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147890 | 47.33 | Campanilevirus | Unclassified | Ackermannviridae | Vibrio |
| MH595538 | Vibrio phage BONAISHI | Vibrio phage BONAISHI Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 288967 | 42.23 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MF754116 | Vibrio phage vB_VpaS_KF6 | Vibrio phage vB_VpaS_KF6 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75904 | 48.904 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MF754115 | Vibrio phage vB_VpaS_KF5 | Vibrio phage vB_VpaS_KF5 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76402 | 48.831 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MF754114 | Vibrio phage vB_VpaS_KF4 | Vibrio phage vB_VpaS_KF4 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75501 | 49.05 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MF754113 | Vibrio phage vB_VpaS_KF3 | Vibrio phage vB_VpaS_KF3 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75955 | 49.029 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MF754112 | Vibrio phage vB_VpaP_KF2 | Vibrio phage vB_VpaP_KF2 Vibrio virus KF2 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43571 | 49.482 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MF754111 | Vibrio phage vB_VpaP_KF1 | Vibrio phage vB_VpaP_KF1 Vibrio virus KF1 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43237 | 49.49 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| MG878892 | Salmonella phage vB_SpuP_Spp16 | Salmonella phage vB_SpuP_Spp16 Salmonella virus Spp16 Panjvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41832 | 39.458 | Panjvirus | Melnykvirinae | Autographiviridae | Salmonella |
| KX912252 | Pseudoalteromonas phage PHS3 | Pseudoalteromonas phage PHS3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35626 | 40.849 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MH447526 | Sulfolobus filamentous virus 1 | Sulfolobus filamentous virus 1 Alphalipothrixvirus SFV1 Alphalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 37311 | 35.802 | Alphalipothrixvirus | Unclassified | Lipothrixviridae | Sulfolobus |
| MG253653 | Lactococcus phage vB_Llc_bIBBF13 | Lactococcus phage vB_Llc_bIBBF13 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29374 | 34.806 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253652 | Lactococcus phage vB_Llc_bIBBF12 | Lactococcus phage vB_Llc_bIBBF12 Lactococcus virus bIBBF12 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29454 | 34.824 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253651 | Lactococcus phage vB_Llc_bIBBEg1 | Lactococcus phage vB_Llc_bIBBEg1 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29659 | 34.684 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253650 | Lactococcus phage vB_Llc_bIBB77s | Lactococcus phage vB_Llc_bIBB77s unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29423 | 34.864 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253649 | Lactococcus phage vB_Llc_bIBB24tp1 | Lactococcus phage vB_Llc_bIBB24tp1 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29736 | 34.712 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253648 | Lactococcus phage vB_Llc_bIBB14s | Lactococcus phage vB_Llc_bIBB14s unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30249 | 34.679 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG253647 | Lactococcus phage vB_Llc_bIBB5g1 | Lactococcus phage vB_Llc_bIBB5g1 Lactococcus virus bIBB5g1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30132 | 34.667 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF346933 | Lactococcus phage vB_Llc_bIBBF14 | Lactococcus phage vB_Llc_bIBBF14 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29837 | 34.739 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MH059636 | Erwinia phage Cronus | Erwinia phage Cronus unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175774 | 38.378 | Unclassified | Tevenvirinae | Myoviridae | Erwinia |
| MF477236 | Nocardia phage NTR1 | Nocardia phage NTR1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65275 | 68.167 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| MH700630 | Sinorhizobium phage phiM6 | Sinorhizobium phage phiM6 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68176 | 42.949 | Unclassified | Unclassified | Podoviridae | Sinorhizobium |
| NC_030476 | Chimpanzee faeces associated microphage 2 | Chimpanzee faeces associated microphage 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5184 | 38.31 | Unclassified | Unclassified | Microviridae | Unspecified |
| KT989433 | Streptomyces phage phiSAJS1 | Streptomyces phage phiSAJS1 unclassified Woodruffvirus Woodruffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56451 | 68.33 | Woodruffvirus | Unclassified | Siphoviridae | Streptomyces |
| KY888885 | Mastigocladus phage CHP58 | Mastigocladus phage CHP58 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40740 | 43.883 | Unclassified | Unclassified | Podoviridae | Mastigocladus |
| MH629685 | Synechococcus virus S-PRM1 | Synechococcus virus S-PRM1 unclassified Kanaloavirus Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144311 | 40.743 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| MH550421 | Enterobacteria phage T6 | Enterobacteria phage T6 Enterobacteria virus T6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168706 | 35.216 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH547045 | Enterobacteria phage O276 | Enterobacteria phage O276 Enterobacteria virus O276 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41969 | 50.683 | Lambdavirus | Unclassified | Siphoviridae | Enterobacteria |
| MH533020 | Acinetobacter phage ABPH49 | Acinetobacter phage ABPH49 unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149960 | 50.8 | Certrevirus | Vequintavirinae | Myoviridae | Acinetobacter |
| MH426725 | Erwinia phage SunLIRen | Erwinia phage SunLIRen unclassified Kolesnikvirus Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84559 | 43.753 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| MH706966 | Enterobacteria phage mEp021 | Enterobacteria phage mEp021 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54655 | 45.767 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| MH684921 | Sphingomonas phage Scott | Sphingomonas phage Scott Sphingomonas virus Scott Scottvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43077 | 57.43 | Scottvirus | Unclassified | Autographiviridae | Sphingomonas |
| MH643778 | Pseudomonas phage YMC12/01/R24 | Pseudomonas phage YMC12/01/R24 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46148 | 64.839 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MH643777 | Pseudomonas phage YMC12/01/R960 | Pseudomonas phage YMC12/01/R960 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37011 | 64.546 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| MH638294 | Ralstonia phage GP4 | Ralstonia phage GP4 Ralstonia virus GP4 Gervaisevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61129 | 64.014 | Gervaisevirus | Unclassified | Podoviridae | Ralstonia |
| MF974397 | Leptospira phage LE4 | Leptospira phage LE4 Leptospira virus LE4 Nylescharonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47866 | 37.285 | Nylescharonvirus | Unclassified | Myoviridae | Leptospira |
| MF974396 | Leptospira phage LE3 | Leptospira phage LE3 Leptospira virus LE3 Nylescharonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48288 | 37.312 | Nylescharonvirus | Unclassified | Myoviridae | Leptospira |
| JX421753 | Enterobacteria phage T7M | Enterobacteria phage T7M Enterobacteria virus T7M Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38202 | 49.906 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacteria |
| KY953156 | Pectobacterium phage vB_PatP_CB5 | Pectobacterium phage vB_PatP_CB5 Pectobacterium virus CB5 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44549 | 48.984 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| NC_004821 | Bacillus phage phBC6A52 | Bacillus phage phBC6A52 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38472 | 34.724 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_004820 | Bacillus phage phBC6A51 | Bacillus phage phBC6A51 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61395 | 37.689 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| NC_004586 | Streptococcus phage 315.3 | Streptococcus phage 315.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34419 | 38.026 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| NC_004170 | Pseudomonas phage phi13 | Pseudomonas phage phi13 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 2981 | 56.189 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| MH370477 | Escherichia phage SRT7 | Escherichia phage SRT7 Escherichia virus SRT7 Foetvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39883 | 50.535 | Foetvirus | Studiervirinae | Autographiviridae | Escherichia |
| MH572526 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5502 | 36.169 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572525 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5508 | 33.442 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572524 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4594 | 37.941 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572523 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4215 | 35.018 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572522 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4171 | 34.86 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572521 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4152 | 38.512 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572520 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5708 | 54.573 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572519 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 56.998 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572518 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5230 | 51.721 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572517 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5321 | 51.851 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572516 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4615 | 47.649 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572515 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4857 | 49.949 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572514 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4172 | 40.34 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572513 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4221 | 40.109 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572512 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4426 | 53.412 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572511 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4189 | 52.9 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572510 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4126 | 52.496 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572509 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4368 | 41.941 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572508 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4479 | 40.366 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572507 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4353 | 42.821 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572506 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4115 | 33.803 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572505 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4041 | 34.298 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572504 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4093 | 44.491 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572503 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4449 | 31.715 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572502 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4415 | 39.366 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572501 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4215 | 36.94 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572500 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4361 | 37.767 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572499 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4348 | 37.052 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572498 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4367 | 36.593 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572497 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4367 | 36.616 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572496 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4346 | 36.631 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572495 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4316 | 47.127 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572494 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4497 | 38.937 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572493 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4230 | 38.582 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572492 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4141 | 51.026 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572491 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5874 | 53.83 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572490 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5668 | 55.452 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572489 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5339 | 53.137 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572488 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4403 | 55.916 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572487 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4521 | 49.502 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572486 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4425 | 43.412 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572485 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4568 | 36.405 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572484 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4207 | 53.245 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572483 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4187 | 53.141 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572482 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4272 | 50.445 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572481 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4459 | 51.76 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572480 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4338 | 47.787 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572479 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4210 | 53.373 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572478 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4247 | 53.968 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572477 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4301 | 54.499 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572476 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4353 | 54.353 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572475 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4309 | 53.818 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572474 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4317 | 57.378 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572473 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4344 | 60.198 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572472 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4199 | 42.105 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572471 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4076 | 34.151 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572470 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4028 | 34.384 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572469 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4398 | 40.45 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572468 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4313 | 35.66 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572467 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4227 | 35.628 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572466 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4177 | 47.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572465 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4084 | 36.337 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572464 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4278 | 36.536 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572463 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3979 | 33.752 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572462 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4370 | 38.924 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572461 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4305 | 39.791 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572460 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4425 | 40.407 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572459 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4518 | 45.883 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572458 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4299 | 52.594 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572457 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4245 | 53.922 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572456 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4281 | 50.572 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572455 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4627 | 50.421 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572454 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4096 | 43.604 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572453 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4249 | 39.303 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572452 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4451 | 44.844 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572451 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4650 | 49.011 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572450 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4443 | 46.973 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572449 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4240 | 50.896 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572448 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4226 | 55.632 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572447 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4351 | 60.607 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572446 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4393 | 49.715 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572445 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4701 | 48.33 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572444 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4399 | 48.784 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572443 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4437 | 45.368 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572442 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4248 | 48.988 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572441 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4152 | 52.842 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572440 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4106 | 45.884 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572439 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4385 | 49.624 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572438 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4169 | 41.569 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572437 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4542 | 42.074 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572436 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4415 | 43.239 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572435 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4572 | 40.114 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572434 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4319 | 40.519 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572433 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4457 | 40.139 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572432 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4286 | 39.454 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572431 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4337 | 41.734 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572430 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4432 | 41.674 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572428 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4242 | 42.928 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572427 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4214 | 42.264 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572426 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4452 | 34.996 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572425 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4409 | 39.828 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572424 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4125 | 48.388 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572423 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4123 | 49.454 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572422 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4642 | 39.961 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572421 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4440 | 35.023 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572420 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4260 | 51.714 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572419 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4268 | 47.071 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572418 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4661 | 47.479 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572417 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4270 | 43.77 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572416 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4212 | 48.457 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572415 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4147 | 33.663 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572414 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4137 | 35.098 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572413 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4544 | 43.552 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572412 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4421 | 43.022 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572411 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4188 | 52.698 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572410 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4237 | 47.77 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572409 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4124 | 53.661 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572408 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4681 | 51.592 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572407 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4755 | 55.605 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572406 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 50.417 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572405 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4781 | 53.19 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572404 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4352 | 47.036 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572403 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4250 | 42.141 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572402 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4090 | 40.244 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572401 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4143 | 39.681 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572400 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4264 | 40.525 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572399 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4105 | 39.44 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572398 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4355 | 48.014 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572397 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4428 | 55.352 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572396 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4188 | 51.528 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572395 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4163 | 52.822 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572394 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4107 | 52.277 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572393 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4361 | 41.734 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572392 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4421 | 40.602 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572391 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4069 | 36.717 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572390 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4048 | 36.512 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572389 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4053 | 34.394 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572388 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4252 | 34.384 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572387 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4345 | 41.703 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572386 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4438 | 41.911 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572385 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4432 | 41.99 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572384 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4466 | 41.738 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572383 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4299 | 40.8 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572382 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4183 | 44.896 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572381 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4381 | 43.255 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572380 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4417 | 42.993 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572379 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3997 | 38.504 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572378 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4174 | 37.494 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572377 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4035 | 37.745 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572376 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4098 | 35.896 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572375 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4348 | 36.362 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572374 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4478 | 42.229 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572373 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4082 | 42.185 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572372 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4305 | 39.117 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572371 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3984 | 39.935 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572370 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4098 | 40.117 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572369 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4518 | 34.108 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572368 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4380 | 42.26 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572367 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4566 | 41.962 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572366 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4184 | 52.653 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572365 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4419 | 51.188 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572364 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4087 | 50.599 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572363 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4239 | 50.955 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572362 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4215 | 51.151 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572361 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4244 | 52.521 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572360 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4272 | 52.528 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572359 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4383 | 49.213 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572358 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4388 | 54.353 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572357 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4521 | 55.541 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572356 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4504 | 54.84 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572355 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4537 | 52.7 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572354 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4568 | 53.897 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572353 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4745 | 55.933 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572352 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5090 | 53.674 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572351 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4227 | 49.278 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572350 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4466 | 53.851 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572349 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4246 | 49.647 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572348 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4255 | 49.001 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572347 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4417 | 44.963 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572346 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4273 | 48.654 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572345 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4324 | 47.248 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572344 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4533 | 49.151 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572343 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4136 | 39.072 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572342 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4211 | 41.724 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572341 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4246 | 41.05 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572340 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4153 | 43.559 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572339 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4497 | 43.54 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572338 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4400 | 46.614 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572337 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4846 | 46.265 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572336 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4309 | 50.592 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572335 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4148 | 57.112 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572334 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4248 | 56.497 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572333 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4467 | 56.638 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572332 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4589 | 49.989 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572331 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4181 | 47.548 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572330 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4265 | 49.004 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572329 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4292 | 48.392 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572328 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4053 | 48.828 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572327 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4194 | 50.501 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572326 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4345 | 51.07 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572325 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4018 | 50.473 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572324 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4320 | 43.218 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572323 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4084 | 49.682 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572322 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4195 | 53.754 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572321 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4340 | 52.074 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572320 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4326 | 52.913 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572319 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4285 | 49.522 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572318 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4447 | 54.599 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572317 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4178 | 52.106 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572316 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4266 | 51.547 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572315 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4342 | 49.332 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572314 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4207 | 53.744 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572313 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4412 | 42.724 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572312 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4247 | 41.676 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572311 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4465 | 41.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572310 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4486 | 39.924 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572309 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4511 | 39.348 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572308 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4413 | 43.598 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572307 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4325 | 40.023 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572306 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4544 | 40.515 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572305 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4571 | 41.588 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572304 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4518 | 41.943 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572303 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4331 | 40.984 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572302 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4416 | 41.259 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572301 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4470 | 38.881 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572300 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4526 | 40.102 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572299 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4221 | 39.327 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572298 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4176 | 38.985 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572297 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4686 | 32.458 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572296 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4116 | 34.645 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572295 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4122 | 34.546 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572294 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4499 | 41.698 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572293 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4150 | 44.819 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572292 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4241 | 42.183 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572291 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4248 | 41.902 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572290 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4368 | 38.37 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572289 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4202 | 39.648 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572288 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4308 | 36.142 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572287 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4365 | 36.678 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572286 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4106 | 46.59 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572285 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4148 | 45.13 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572284 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4228 | 45.932 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572283 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4316 | 41.196 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572282 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4200 | 38.31 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572281 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 3988 | 36.384 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572280 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4334 | 39.248 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572279 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4438 | 38.373 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572278 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4750 | 52.674 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572277 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4429 | 44.999 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572276 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4767 | 45.186 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572275 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4618 | 42.962 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572274 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4780 | 45.251 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572273 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4189 | 48.317 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572272 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4237 | 51.546 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572271 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4141 | 49.312 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572270 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4449 | 55.968 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH572269 | Microviridae sp. | Microviridae sp. unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4446 | 51.844 | Unclassified | Unclassified | Microviridae | Unspecified |
| MH636380 | Cylindrospermopsis raciborskii virus RM-2018a | Cylindrospermopsis raciborskii virus RM-2018a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104363 | 39.016 | Unclassified | Unclassified | Siphoviridae | Cylindrospermopsis |
| MG770228 | Escherichia phage PMBT57 | Escherichia phage PMBT57 unclassified Enquatrovirus Enquatrovirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70857 | 41.371 | Enquatrovirus | Enquatrovirinae | Schitoviridae | Escherichia |
| NC_030472 | Chimpanzee faeces associated microphage 1 | Chimpanzee faeces associated microphage 1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4851 | 45.455 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_030458 | Chimpanzee faeces associated microphage 3 | Chimpanzee faeces associated microphage 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5016 | 45.355 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_029014 | Parabacteroides phage YZ-2015b | Parabacteroides phage YZ-2015b unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5001 | 40.152 | Unclassified | Unclassified | Microviridae | Parabacteroides |
| NC_029012 | Parabacteroides phage YZ-2015a | Parabacteroides phage YZ-2015a unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4623 | 42.808 | Unclassified | Unclassified | Microviridae | Parabacteroides |
| NC_028994 | Gokushovirinae GNX3R | Gokushovirinae GNX3R unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4273 | 40.019 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_028993 | Gokushovirinae GAIR4 | Gokushovirinae GAIR4 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4468 | 42.01 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_028902 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4184 | 51.243 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| NC_027648 | Microviridae Fen7940_21 | Microviridae Fen7940_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4453 | 52.773 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027647 | Microviridae Fen7918_21 | Microviridae Fen7918_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4199 | 44.534 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027646 | Microviridae Fen7895_21 | Microviridae Fen7895_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4424 | 43.716 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027645 | Gokushovirinae Fen7875_21 | Gokushovirinae Fen7875_21 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4631 | 53.444 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_027644 | Microviridae Fen7786_21 | Microviridae Fen7786_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4686 | 43.064 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027643 | Microviridae Fen685_11 | Microviridae Fen685_11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4654 | 45.982 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027642 | Gokushovirinae Fen672_31 | Gokushovirinae Fen672_31 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4624 | 49.632 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_027641 | Microviridae Fen4707_41 | Microviridae Fen4707_41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4552 | 44.464 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027640 | Microviridae Fen418_41 | Microviridae Fen418_41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4949 | 42.797 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027639 | Microviridae Fen2266_11 | Microviridae Fen2266_11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4366 | 40.609 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027638 | Microviridae Bog9017_22 | Microviridae Bog9017_22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 41.94 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027637 | Gokushovirinae Bog8989_22 | Gokushovirinae Bog8989_22 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4656 | 39.798 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_027636 | Gokushovirinae Bog5712_52 | Gokushovirinae Bog5712_52 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4501 | 51.744 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_027635 | Microviridae Bog5275_51 | Microviridae Bog5275_51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4815 | 47.622 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027634 | Microviridae Bog1249_12 | Microviridae Bog1249_12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4451 | 42.373 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_027633 | Gokushovirinae Bog1183_53 | Gokushovirinae Bog1183_53 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4609 | 36.537 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_026665 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6083 | 30.659 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_026013 | Microviridae IME-16 | Microviridae IME-16 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5755 | 37.063 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_025454 | Ralstonia phage RS603 | Ralstonia phage RS603 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7679 | 59.37 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| NC_023586 | Ralstonia phage 1 NP-2014 | Ralstonia phage 1 NP-2014 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8460 | 58.487 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| NC_008574 | Ralstonia phage RSM1 | Ralstonia phage RSM1 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 9004 | 59.951 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| NC_022790 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4129 | 39.574 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| NC_021805 | Cellulophaga phage phi12a:1 | Cellulophaga phage phi12a:1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6478 | 34.023 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| NC_021797 | Cellulophaga phage phi12:2 | Cellulophaga phage phi12:2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6453 | 34.806 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| NC_021866 | Ralstonia phage RSS20 | Ralstonia phage RSS20 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7219 | 40.546 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| NC_021862 | Ralstonia phage RSS30 | Ralstonia phage RSS30 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8576 | 61.719 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| NC_021569 | Stenotrophomonas phage SMA7 | Stenotrophomonas phage SMA7 Subteminivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7069 | 62.343 | Subteminivirus | Unclassified | Inoviridae | Stenotrophomonas |
| NC_021562 | Vibrio phage VFJ | Vibrio phage VFJ Saetivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8555 | 44.29 | Saetivirus | Unclassified | Inoviridae | Vibrio |
| NC_019548 | Ralstonia phage RSS0 | Ralstonia phage RSS0 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7288 | 62.047 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| NC_015785 | Microviridae phi-CA82 | Microviridae phi-CA82 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5514 | 39.717 | Unclassified | Unclassified | Microviridae | Unspecified |
| NC_015586 | Stenotrophomonas phage phiSHP2 | Stenotrophomonas phage phiSHP2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5819 | 61.505 | Unclassified | Unclassified | Inoviridae | Stenotrophomonas |
| NC_016162 | Vibrio phage VCY-phi | Vibrio phage VCY-phi Vicialiavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7103 | 41.405 | Vicialiavirus | Unclassified | Inoviridae | Vibrio |
| NC_015297 | Ralstonia phage PE226 | Ralstonia phage PE226 Parhipatevirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5475 | 61.662 | Parhipatevirus | Unclassified | Inoviridae | Ralstonia |
| NC_012868 | Escherichia phage St-1 | Escherichia phage St-1 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6094 | 45.208 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_012757 | Vibrio phage VEJ | Vibrio phage VEJ Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6842 | 42.955 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| NC_010392 | Phage Gifsy-1 | Phage Gifsy-1 unclassified bacterial viruses Viruses | 48491 | 51.098 | Unclassified | Unclassified | Unclassified | Unspecified |
| NC_010391 | Salmonella phage Fels-1 | Salmonella phage Fels-1 unclassified bacterial viruses Viruses | 42723 | 51.555 | Unclassified | Unclassified | Unclassified | Salmonella |
| NC_009987 | Spiroplasma virus SkV1CR23x | Spiroplasma virus SkV1CR23x Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7870 | 22.211 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| NC_005131 | Ralstonia phage p12J | Ralstonia phage p12J unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7118 | 56.954 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| NC_008575 | Ralstonia phage RSS1 | Ralstonia phage RSS1 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6662 | 62.609 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| NC_007821 | Enterobacteria phage WA13 sensu lato | Enterobacteria phage WA13 sensu lato unclassified Alphatrevirus Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 44.825 | Alphatrevirus | Bullavirinae | Microviridae | Enterobacteria |
| NC_007817 | Escherichia phage ID2 Moscow/ID/2001 | Escherichia phage ID2 Moscow/ID/2001 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.753 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| NC_007856 | Enterobacteria phage ID18 sensu lato | Enterobacteria phage ID18 sensu lato Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.224 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| NC_007189 | Stenotrophomonas phage SMA9 | Stenotrophomonas phage SMA9 Staminivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6907 | 62.386 | Staminivirus | Unclassified | Inoviridae | Stenotrophomonas |
| NC_003287 | Escherichia virus M13 | Escherichia virus M13 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.737 | Inovirus | Unclassified | Inoviridae | Escherichia |
| NC_006294 | Vibrio phage KSF1 | Vibrio phage KSF1 Capistrivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7107 | 44.379 | Capistrivirus | Unclassified | Inoviridae | Vibrio |
| NC_005948 | Vibrio virus Vf33 | Vibrio virus Vf33 Villovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7965 | 45.662 | Villovirus | Unclassified | Inoviridae | Vibrio |
| NC_001270 | Spiroplasma phage SVTS2 | Spiroplasma phage SVTS2 Suturavirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6825 | 22.652 | Suturavirus | Unclassified | Plectroviridae | Spiroplasma |
| NC_004736 | Vibrio phage VGJ | Vibrio phage VGJ Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7542 | 43.37 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| NC_003327 | Vibrio phage VSK | Vibrio phage VSK Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6882 | 43.723 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| NC_004306 | Vibrio virus fs1 | Vibrio virus fs1 Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6340 | 43.423 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| NC_004175 | Pseudomonas phage phi12 | Pseudomonas phage phi12 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 4100 | 56.024 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_004174 | Pseudomonas phage phi12 | Pseudomonas phage phi12 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 2322 | 56.977 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_004173 | Pseudomonas phage phi12 | Pseudomonas phage phi12 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 6751 | 54.111 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_004172 | Pseudomonas phage phi13 | Pseudomonas phage phi13 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 6458 | 58.362 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_004171 | Pseudomonas phage phi13 | Pseudomonas phage phi13 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 4213 | 57.869 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003793 | Spiroplasma phage C74 | Spiroplasma phage C74 Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7768 | 23.159 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| NC_003716 | Pseudomonas virus phi6 | Pseudomonas virus phi6 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 4063 | 56.756 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003715 | Pseudomonas virus phi6 | Pseudomonas virus phi6 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 6374 | 55.522 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003714 | Pseudomonas virus phi6 | Pseudomonas virus phi6 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 2948 | 55.156 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003460 | Propionibacterium phage B5 | Propionibacterium phage B5 Bifilivirus Paulinoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5804 | 64.283 | Bifilivirus | Unclassified | Paulinoviridae | Propionibacterium |
| NC_003438 | Spiroplasma virus SpV4 | Spiroplasma virus SpV4 Spiromicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4421 | 32.052 | Spiromicrovirus | Gokushovirinae | Microviridae | Spiroplasma |
| NC_003301 | Pseudomonas phage phi8 | Pseudomonas phage phi8 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 3192 | 54.449 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003300 | Pseudomonas phage phi8 | Pseudomonas phage phi8 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 4741 | 55.263 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_003299 | Pseudomonas phage phi8 | Pseudomonas phage phi8 Cystovirus Cystoviridae Mindivirales Vidaverviricetes Duplornaviricota Orthornavirae Riboviria Viruses | 7051 | 54.021 | Cystovirus | Unclassified | Cystoviridae | Pseudomonas |
| NC_002643 | Bdellovibrio phage phiMH2K | Bdellovibrio phage phiMH2K Bdellomicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4594 | 46.104 | Bdellomicrovirus | Gokushovirinae | Microviridae | Bdellovibrio |
| NC_002180 | Chlamydia virus CPAR39 | Chlamydia virus CPAR39 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4532 | 40.622 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| NC_002363 | Vibrio phage VfO4K68 | Vibrio phage VfO4K68 Versovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6891 | 47.236 | Versovirus | Unclassified | Inoviridae | Vibrio |
| NC_002362 | Vibrio phage VfO3K6 | Vibrio phage VfO3K6 Versovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8784 | 45.162 | Versovirus | Unclassified | Inoviridae | Vibrio |
| NC_002194 | Chlamydia phage 2 | Chlamydia phage 2 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4563 | 40.938 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| NC_001998 | Chlamydia phage CPG1 | Chlamydia phage CPG1 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 40.98 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| NC_001956 | Vibrio phage fs2 | Vibrio phage fs2 Saetivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8651 | 44.527 | Saetivirus | Unclassified | Inoviridae | Vibrio |
| NC_001954 | Escherichia phage If1 | Escherichia phage If1 Infulavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8454 | 43.707 | Infulavirus | Unclassified | Inoviridae | Escherichia |
| NC_001890 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4215 | 48.541 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| NC_001741 | Chlamydia virus Chp1 | Chlamydia virus Chp1 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4877 | 36.498 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| NC_001730 | Escherichia phage phiK | Escherichia phage phiK Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6089 | 44.917 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| NC_001426 | Escherichia virus BZ13 | Escherichia virus BZ13 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3466 | 47.865 | Levivirus | Unclassified | Leviviridae | Escherichia |
| NC_001418 | Pseudomonas phage Pf3 | Pseudomonas phage Pf3 Tertilicivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5833 | 45.363 | Tertilicivirus | Unclassified | Inoviridae | Pseudomonas |
| NC_002014 | Salmonella phage IKe | Salmonella phage IKe Lineavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6883 | 40.549 | Lineavirus | Unclassified | Inoviridae | Salmonella |
| NC_001396 | Xanthomonas virus Cf1c | Xanthomonas virus Cf1c Coriovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7308 | 58.073 | Coriovirus | Unclassified | Inoviridae | Xanthomonas |
| NC_001365 | Spiroplasma phage R8A2B | Spiroplasma phage R8A2B Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8273 | 22.894 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| NC_001341 | Acholeplasma phage MV-L51 | Acholeplasma phage MV-L51 Plectrovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4491 | 33.289 | Plectrovirus | Unclassified | Plectroviridae | Acholeplasma |
| NC_001332 | Escherichia virus I22 | Escherichia virus I22 Lineavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6744 | 42.719 | Lineavirus | Unclassified | Inoviridae | Escherichia |
| NC_001331 | Pseudomonas virus Pf1 | Pseudomonas virus Pf1 Primolicivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7349 | 61.491 | Primolicivirus | Unclassified | Inoviridae | Pseudomonas |
| NC_001330 | Escherichia phage alpha3 | Escherichia phage alpha3 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6087 | 45.178 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| KT804908 | Acinetobacter phage IME-200 | Acinetobacter phage IME-200 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41243 | 39.313 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KM411960 | Pseudomonas phage phi176 | Pseudomonas phage phi176 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73048 | 54.853 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KM411959 | Pseudomonas phage Pa2 | Pseudomonas phage Pa2 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73008 | 54.857 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KM411958 | Pseudomonas phage RWG | Pseudomonas phage RWG Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72646 | 54.869 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MF448571 | Lactococcus phage 96403 | Lactococcus phage 96403 Lactococcus virus 96403 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26874 | 34.476 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448570 | Lactococcus phage 96401 | Lactococcus phage 96401 Lactococcus virus 96401 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29306 | 35.16 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448569 | Lactococcus phage 88605 | Lactococcus phage 88605 Lactococcus virus 88605 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31508 | 34.467 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448568 | Lactococcus phage 79201 | Lactococcus phage 79201 Lactococcus virus 79201 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31123 | 34.932 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448567 | Lactococcus phage 66901 | Lactococcus phage 66901 Lactococcus virus 66901 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28917 | 34.71 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448566 | Lactococcus phage 63302 | Lactococcus phage 63302 Lactococcus virus 63302 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31883 | 34.774 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448565 | Lactococcus phage 62605 | Lactococcus phage 62605 Lactococcus virus 62605 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31014 | 34.562 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448564 | Lactococcus phage 62601 | Lactococcus phage 62601 Lactococcus virus 62601 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30852 | 34.62 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448563 | Lactococcus phage 57001 | Lactococcus phage 57001 Lactococcus virus 57001 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31819 | 34.3 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448562 | Lactococcus phage 56301 | Lactococcus phage 56301 Lactococcus virus 56301 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32555 | 33.958 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448561 | Lactococcus phage 56003 | Lactococcus phage 56003 Lactococcus virus 56003 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30926 | 34.524 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448559 | Lactococcus phage 38507 | Lactococcus phage 38507 Lactococcus virus 38507 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32458 | 33.56 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448558 | Lactococcus phage 38503 | Lactococcus phage 38503 Lactococcus virus 38503 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27948 | 34.414 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448557 | Lactococcus phage 37201 | Lactococcus phage 37201 Lactococcus virus 37201 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30989 | 34.79 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448556 | Lactococcus phage 30804 | Lactococcus phage 30804 Lactococcus virus 30804 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30401 | 34.193 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448555 | Lactococcus phage 16802 | Lactococcus phage 16802 Lactococcus virus 16802 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28721 | 35.239 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448554 | Lactococcus phage 05601 | Lactococcus phage 05601 Lactococcus virus 05601 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32895 | 34.084 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448572 | Lactococcus phage 96603 | Lactococcus phage 96603 Lactococcus virus 96603 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30690 | 34.904 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF448560 | Lactococcus phage 51701 | Lactococcus phage 51701 Lactococcus virus 51701 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32529 | 34.357 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG893203 | Vibrio phage VP06 | Vibrio phage VP06 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75893 | 48.989 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MH445453 | Klebsiella phage NJS1 | Klebsiella phage NJS1 Klebsiella virus NJS1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49292 | 50.665 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MH626557 | Eggerthella phage PMBT5 | Eggerthella phage PMBT5 Eggerthella virus PMBT5 Lentavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30930 | 51.3 | Lentavirus | Unclassified | Siphoviridae | Eggerthella |
| MH606185 | Bacillus phage BSP38 | Bacillus phage BSP38 Bacillus virus BSP38 Jeonjuvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153268 | 41.747 | Jeonjuvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH599087 | Vibrio phage R01 | Vibrio phage R01 unclassified Mardecavirus Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75514 | 49.424 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| MH536815 | Streptomyces phage Crosby | Streptomyces phage Crosby Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54036 | 68.317 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH509445 | Mycobacterium phage Cborch11 | Mycobacterium phage Cborch11 unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68508 | 57.582 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| MH431933 | Paenibacillus phage DevRi | Paenibacillus phage DevRi unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38520 | 41.493 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431932 | Paenibacillus phage Arcticfreeze | Paenibacillus phage Arcticfreeze unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38518 | 41.485 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH450123 | Arthrobacter phage MargaretKali | Arthrobacter phage MargaretKali Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39448 | 61.06 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| MG581355 | Escherichia phage B2 | Escherichia phage B2 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44283 | 54.671 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| MH248069 | Clostridium phage CPS2 | Clostridium phage CPS2 Clostridium virus CPS2 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17961 | 33.3 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| MH560358 | Escherichia virus KFS-EC | Escherichia virus KFS-EC unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164715 | 40.47 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| MH426726 | Erwinia phage Pavtok | Erwinia phage Pavtok Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61401 | 61.983 | Unclassified | Unclassified | Podoviridae | Erwinia |
| MH426724 | Erwinia phage Wellington | Erwinia phage Wellington Erwinia virus Wellington Wellingtonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244950 | 52.115 | Wellingtonvirus | Unclassified | Myoviridae | Erwinia |
| MF431726 | Vibrio phage vB_ValP_IME271 | Vibrio phage vB_ValP_IME271 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50345 | 41.444 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MH460824 | Paenibacillus phage Unity | Paenibacillus phage Unity Paenibacillus virus Unity Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50316 | 49.11 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH553563 | Enterobacteria phage RB18 | Enterobacteria phage RB18 Enterobacteria virus RB18 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166677 | 35.371 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH431934 | Paenibacillus phage Gryphonian | Paenibacillus phage Gryphonian unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38541 | 41.457 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH538296 | Bacillus phage Maceta | Bacillus phage Maceta unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45023 | 39.991 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH015257 | Ruegeria phage vB_RpoP-V14 | Ruegeria phage vB_RpoP-V14 unclassified Aorunvirus Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74792 | 47.912 | Aorunvirus | Rhodovirinae | Schitoviridae | Ruegeria |
| MH540083 | Synechococcus T7-like phage S-TIP37 | Synechococcus T7-like phage S-TIP37 Synechococcus virus STIP37 Igirivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45477 | 50.483 | Igirivirus | Unclassified | Autographiviridae | Synechococcus |
| MH538193 | Bacillus phage Saddex | Bacillus phage Saddex unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142353 | 38.985 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MH536736 | Pseudomonas phage E79 | Pseudomonas phage E79 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66061 | 55.029 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MH517022 | Acinetobacter phage SH-Ab 15599 | Acinetobacter phage SH-Ab 15599 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143204 | 38.459 | Unclassified | Unclassified | Ackermannviridae | Acinetobacter |
| MH512890 | Cyanophage S-TIM4 | Cyanophage S-TIM4 Synechococcus virus STIM4 Thaumasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176495 | 37.657 | Thaumasvirus | Unclassified | Myoviridae | Synechococcus |
| MH460827 | Paenibacillus phage Halcyone | Paenibacillus phage Halcyone Paenibacillus virus Halcyone Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55560 | 48.56 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH460826 | Paenibacillus phage Heath | Paenibacillus phage Heath unclassified Halcyonevirus Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55560 | 48.571 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH460825 | Paenibacillus phage Scottie | Paenibacillus phage Scottie Paenibacillus virus Scottie Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55990 | 48.519 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH464253 | Shigella phage SFPH2 | Shigella phage SFPH2 Shigella virus SFPH2 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40387 | 52.41 | Kayfunavirus | Studiervirinae | Autographiviridae | Shigella |
| MH454084 | Paenibacillus phage Toothless | Paenibacillus phage Toothless unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38832 | 41.978 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454083 | Paenibacillus phage Saudage | Paenibacillus phage Saudage unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37962 | 41.873 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454082 | Paenibacillus phage Genki | Paenibacillus phage Genki unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38540 | 41.458 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454081 | Paenibacillus phage LincolnB | Paenibacillus phage LincolnB unclassified Wanderervirus Wanderervirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40437 | 42.333 | Wanderervirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| MH454080 | Paenibacillus phage Ley | Paenibacillus phage Ley unclassified Halcyonevirus Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56465 | 47.975 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454079 | Paenibacillus phage Jacopo | Paenibacillus phage Jacopo Paenibacillus virus Jacopo Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38526 | 41.559 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454078 | Paenibacillus phage Eltigre | Paenibacillus phage Eltigre Paenibacillus virus Eltigre Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38675 | 41.417 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454077 | Paenibacillus phage Bloom | Paenibacillus phage Bloom unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38519 | 41.502 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH454076 | Paenibacillus phage Ash | Paenibacillus phage Ash unclassified Halcyonevirus Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56468 | 47.997 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431938 | Paenibacillus phage C7Cdelta | Paenibacillus phage C7Cdelta Paenibacillus virus C7Cdelta Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55774 | 48.008 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431937 | Paenibacillus phage Lucielle | Paenibacillus phage Lucielle Paenibacillus virus Lucielle Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37947 | 41.821 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431936 | Paenibacillus phage Kawika | Paenibacillus phage Kawika Paenibacillus virus Kawika Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40769 | 41.782 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431935 | Paenibacillus phage Honeybear | Paenibacillus phage Honeybear unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40054 | 41.941 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431931 | Paenibacillus phage Yyerffej | Paenibacillus phage Yyerffej Paenibacillus virus Yyerffej Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43126 | 40.551 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MH431930 | Paenibacillus phage Wanderer | Paenibacillus phage Wanderer Paenibacillus virus Wanderer Wanderervirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40448 | 42.353 | Wanderervirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| MH401416 | Staphylococcus phage SA345ruMSSAST8 | Staphylococcus phage SA345ruMSSAST8 Staphylococcus virus SA345ruMSSAST8 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42735 | 33.062 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MH401415 | Staphylococcus phage SA537ruMSSAST97 | Staphylococcus phage SA537ruMSSAST97 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43458 | 33.349 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MH401414 | Staphylococcus phage SA45ruMSSAST97 | Staphylococcus phage SA45ruMSSAST97 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43454 | 33.329 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MH384261 | Staphylococcus phage SA137ruMSSAST121PVL | Staphylococcus phage SA137ruMSSAST121PVL Staphylococcus virus SA137ruMSSAST121PVL Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45999 | 33.305 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MH384259 | Staphylococcus phage SA1014ruMSSAST7 | Staphylococcus phage SA1014ruMSSAST7 Staphylococcus virus SA1014ruMSSAST7 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43504 | 33.08 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| MH252365 | Ralstonia phage phiRSP | Ralstonia phage phiRSP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44526 | 62.256 | Unclassified | Unclassified | Myoviridae | Ralstonia |
| CP013974 | Erwinia phage LS-2018a | Erwinia phage LS-2018a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59759 | 51.017 | Unclassified | Unclassified | Myoviridae | Erwinia |
| KX397364 | Erwinia phage vB_EamM_Asesino | Erwinia phage vB_EamM_Asesino Erskinevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246290 | 51.211 | Erskinevirus | Unclassified | Myoviridae | Erwinia |
| MH221129 | Salmonella phage SPAsTU | Salmonella phage SPAsTU unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 250739 | 49.541 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| MH587638 | Klebsiella phage vB_KpnP_IME321 | Klebsiella phage vB_KpnP_IME321 Klebsiella virus IME321 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39906 | 52.767 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MH378443 | Escherichia virus phiX174 | Escherichia virus phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| MH378442 | Escherichia virus phiX174 | Escherichia virus phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.783 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| MH378441 | Escherichia virus phiX174 | Escherichia virus phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.82 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| MH494197 | Escherichia phage CMSTMSU | Escherichia phage CMSTMSU unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 386442 | 35.597 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MH383160 | Escherichia phage UB | Escherichia phage UB unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 353081 | 34.009 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| MH382836 | Pseudomonas phage phCDa | Pseudomonas phage phCDa Pseudomonas virus phCDa Shizishanvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72821 | 53.268 | Shizishanvirus | Unclassified | Schitoviridae | Pseudomonas |
| MH370478 | Pseudomonas phage SRT6 | Pseudomonas phage SRT6 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91364 | 49.301 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MH355584 | Escherichia phage vB_EcoM_JB75 | Escherichia phage vB_EcoM_JB75 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167208 | 35.27 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH355583 | Enterococcus phage vB_EfaS_LM99 | Enterococcus phage vB_EfaS_LM99 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40203 | 35.005 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MH178096 | Aeromonas phage AsXd-1 | Aeromonas phage AsXd-1 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39014 | 51.233 | Unclassified | Hendrixvirinae | Siphoviridae | Aeromonas |
| MF158038 | Shigella phage Sf11 SMD-2017 | Shigella phage Sf11 SMD-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46454 | 45.985 | Unclassified | Unclassified | Siphoviridae | Shigella |
| KX711710 | Pseudomonas phage vB_Pae-TbilisiM32 | Pseudomonas phage vB_Pae-TbilisiM32 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42965 | 62.265 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KY658673 | Vibrio phage H2 PGK-2017 | Vibrio phage H2 PGK-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46149 | 43.136 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KM612262 | Vibrio phage H2 SGB-2014 | Vibrio phage H2 SGB-2014 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39530 | 50.549 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| HE611333 | Pseudomonas phage tf | Pseudomonas phage tf Pseudomonas virus tf Krylovvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46271 | 53.202 | Krylovvirus | Unclassified | Podoviridae | Pseudomonas |
| JF974339 | Enterobacteria phage IME10 | Enterobacteria phage IME10 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39646 | 47.495 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| MG641188 | Lepus americanus faeces associated microvirus SHP1 6472 | Lepus americanus faeces associated microvirus SHP1 6472 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6357 | 37.124 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641187 | Alces alces faeces associated microvirus MP21 4718 | Alces alces faeces associated microvirus MP21 4718 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4662 | 48.263 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641186 | Alces alces faeces associated microvirus MP18 4940 | Alces alces faeces associated microvirus MP18 4940 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4892 | 36.958 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641185 | Alces alces faeces associated microvirus MP15 5067 | Alces alces faeces associated microvirus MP15 5067 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5004 | 36.771 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641184 | Alces alces faeces associated microvirus MP12 5423 | Alces alces faeces associated microvirus MP12 5423 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5360 | 37.91 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641183 | Alces alces faeces associated microvirus MP11 5517 | Alces alces faeces associated microvirus MP11 5517 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5398 | 32.29 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641182 | Alces alces faeces associated microvirus MP10 5560 | Alces alces faeces associated microvirus MP10 5560 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5497 | 38.039 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641181 | Alces alces faeces associated microvirus MP3 6497 | Alces alces faeces associated microvirus MP3 6497 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6434 | 36.509 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641180 | Lynx canadensis associated microvirus CLP 9366 | Lynx canadensis associated microvirus CLP 9366 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5265 | 49.383 | Unclassified | Unclassified | Microviridae | Unspecified |
| MG641179 | Lynx canadensis associated microvirus CLP 9413 | Lynx canadensis associated microvirus CLP 9413 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5312 | 49.812 | Unclassified | Unclassified | Microviridae | Unspecified |
| KY707339 | Pseudomonas phage JBD68 | Pseudomonas phage JBD68 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39841 | 61.08 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MG641885 | Salmonella phage PMBT28 | Salmonella phage PMBT28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48113 | 58.614 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| MF443126 | Lactococcus phage 05802 | Lactococcus phage 05802 Lactococcus virus 05802 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22434 | 35.727 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443125 | Lactococcus phage 50102 | Lactococcus phage 50102 Lactococcus virus 50102 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22242 | 36.022 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443124 | Lactococcus phage 50504 | Lactococcus phage 50504 Lactococcus virus 50504 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21562 | 35.92 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443122 | Lactococcus phage 74001 | Lactococcus phage 74001 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21879 | 35.916 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443121 | Lactococcus phage 37203 | Lactococcus phage 37203 Lactococcus virus 37203 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22029 | 35.921 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443120 | Lactococcus phage 62403 | Lactococcus phage 62403 Lactococcus virus 62403 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21704 | 35.851 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443119 | Lactococcus phage 62402 | Lactococcus phage 62402 Lactococcus virus 62402 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21980 | 35.983 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MF443123 | Lactococcus phage 62606 | Lactococcus phage 62606 Lactococcus virus 62606 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22185 | 36.407 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| MH268168 | Pseudomonas phage PFP1 | Pseudomonas phage PFP1 Pseudomonas virus PFP1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40914 | 55.815 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MH206184 | Xanthomonas phage phiLF | Xanthomonas phage phiLF unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6062 | 59.865 | Unclassified | Unclassified | Inoviridae | Xanthomonas |
| MH206183 | Xanthomonas phage phiXv2 | Xanthomonas phage phiXv2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6564 | 59.552 | Unclassified | Unclassified | Inoviridae | Xanthomonas |
| MH218848 | Xanthomonas phage phiLf2 | Xanthomonas phage phiLf2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6372 | 60.217 | Unclassified | Unclassified | Inoviridae | Xanthomonas |
| MH015259 | Ruegeria phage vB_RpoP-V17 | Ruegeria phage vB_RpoP-V17 unclassified Aorunvirus Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74665 | 47.933 | Aorunvirus | Rhodovirinae | Schitoviridae | Ruegeria |
| MH015258 | Ruegeria phage vB_RpoS-V16 | Ruegeria phage vB_RpoS-V16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61382 | 63.641 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MH015256 | Ruegeria phage vB_RpoP-V13 | Ruegeria phage vB_RpoP-V13 Ruegeria virus V13 Pomeroyivirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74830 | 50.771 | Pomeroyivirus | Rhodovirinae | Schitoviridae | Ruegeria |
| MH015255 | Ruegeria phage vB_RpoS-V10 | Ruegeria phage vB_RpoS-V10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147480 | 56.363 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MH015254 | Ruegeria phage vB_RpoS-V11 | Ruegeria phage vB_RpoS-V11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59549 | 64.025 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MH015253 | Ruegeria phage vB_RpoP-V21 | Ruegeria phage vB_RpoP-V21 unclassified Aorunvirus Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74665 | 47.907 | Aorunvirus | Rhodovirinae | Schitoviridae | Ruegeria |
| MH015252 | Ruegeria phage vB_RpoS-V18 | Ruegeria phage vB_RpoS-V18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59111 | 64.029 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MH015251 | Ruegeria phage vB_RpoMi-V15 | Ruegeria phage vB_RpoMi-V15 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4248 | 57.698 | Unclassified | Unclassified | Microviridae | Ruegeria |
| MH015250 | Ruegeria phage vB_RpoP-V12 | Ruegeria phage vB_RpoP-V12 Ruegeria virus V12 Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74675 | 47.936 | Aorunvirus | Rhodovirinae | Schitoviridae | Ruegeria |
| MH015249 | Ruegeria phage vB_RpoS-V7 | Ruegeria phage vB_RpoS-V7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59573 | 64.123 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| MH285980 | Escherichia phage Eco_BIFF | Escherichia phage Eco_BIFF Escherichia virus BIFF Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49372 | 45.258 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MH248138 | Erwinia phage vB_EamM_Alexandra | Erwinia phage vB_EamM_Alexandra Erwinia virus Alexandra Alexandravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266532 | 50.126 | Alexandravirus | Unclassified | Myoviridae | Erwinia |
| MH252123 | Klebsiella phage ZCKP1 | Klebsiella phage ZCKP1 Klebsiella virus ZCKP1 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150925 | 39.102 | Phapecoctavirus | Unclassified | Myoviridae | Klebsiella |
| MH359124 | Escherichia phage SF | Escherichia phage SF unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168695 | 37.614 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MH229865 | Streptomyces phage LazerLemon | Streptomyces phage LazerLemon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54798 | 68.092 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH229864 | Streptomyces phage UNTPL | Streptomyces phage UNTPL Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54495 | 68.335 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH229863 | Streptomyces phage JackieB | Streptomyces phage JackieB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54912 | 68.231 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH229862 | Streptomyces phage Henoccus | Streptomyces phage Henoccus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55137 | 68.219 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH271296 | Gordonia phage Emperor | Gordonia phage Emperor Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16604 | 70.116 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MH031343 | Escherichia phage vB_EcoP_S523 | Escherichia phage vB_EcoP_S523 Escherichia virus S523 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39825 | 48.806 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| MH358458 | Escherichia phage phi G17 | Escherichia phage phi G17 unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68270 | 43.529 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MH238467 | Pasteurella phage Pm86 | Pasteurella phage Pm86 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56048 | 38.526 | Unclassified | Unclassified | Siphoviridae | Pasteurella |
| MH238466 | Pasteurella phage AFS-2018a | Pasteurella phage AFS-2018a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33296 | 39.521 | Unclassified | Unclassified | Myoviridae | Pasteurella |
| MH375074 | Enterococcus phage LY0323 | Enterococcus phage LY0323 unclassified Efquatrovirus Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40876 | 34.673 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MH358359 | Salmonella phage vB_SpuS_Sp4 | Salmonella phage vB_SpuS_Sp4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43614 | 49.491 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MH445500 | Alteromonas phage JH01 | Alteromonas phage JH01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46500 | 44.394 | Unclassified | Unclassified | Siphoviridae | Alteromonas |
| KY381600 | Lactobacillus phage P2 | Lactobacillus phage P2 unclassified Maenadvirus Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77937 | 39.2 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MG781191 | Escherichia phage vB_EcoM-fHoEco02 | Escherichia phage vB_EcoM-fHoEco02 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167064 | 35.335 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MG781190 | Escherichia phage vB_EcoM-fFiEco06 | Escherichia phage vB_EcoM-fFiEco06 Escherichia virus fFiEco06 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167076 | 35.34 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MF893341 | Ralstonia phage RPSC1 | Ralstonia phage RPSC1 Ralstonia virus RPSC1 Stompelvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39628 | 61.555 | Stompelvirus | Unclassified | Autographiviridae | Ralstonia |
| MG748548 | Escherichia phage EP335 | Escherichia phage EP335 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76622 | 42.531 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MG748547 | Escherichia phage EP75 | Escherichia phage EP75 Escherichia virus EP75 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158143 | 44.644 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| MF418016 | Bacillus phage vB_BceM-HSE3 | Bacillus phage vB_BceM-HSE3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124002 | 36.206 | Unclassified | Unclassified | Myoviridae | Bacillus |
| MG022440 | Phage 206 | Phage 206 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138073 | 43.702 | Vequintavirus | Vequintavirinae | Myoviridae | Unspecified |
| MG022439 | Phage 203 | Phage 203 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138073 | 43.702 | Vequintavirus | Vequintavirinae | Myoviridae | Unspecified |
| MG020111 | Lactobacillus phage Lb | Lactobacillus phage Lb Lactobacillus virus Lb Heilongjiangvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43416 | 41.738 | Heilongjiangvirus | Unclassified | Myoviridae | Lactobacillus |
| AP014888 | Vibrio phage CKB-S2 | Vibrio phage CKB-S2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34713 | 46.006 | Unclassified | Unclassified | Podoviridae | Vibrio |
| AP014887 | Vibrio phage CKB-S1 | Vibrio phage CKB-S1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48157 | 61.378 | Unclassified | Unclassified | Myoviridae | Vibrio |
| AP014858 | Vibrio phage RYC | Vibrio phage RYC Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158066 | 39.042 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MH133207 | Acinetobacter phage vB_AbaM_B9 | Acinetobacter phage vB_AbaM_B9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93641 | 32.57 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| MF740800 | Salmonella phage vB_SenM_PA13076 | Salmonella phage vB_SenM_PA13076 unclassified Rosemountvirus Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52474 | 46.112 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| KR296684 | Salmonella phage 19 | Salmonella phage 19 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94766 | 45.642 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KR296685 | Salmonella phage 21 | Salmonella phage 21 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51896 | 46.936 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KR296693 | Salmonella phage 39 | Salmonella phage 39 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21437 | 45.193 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KR296695 | Salmonella phage 41 | Salmonella phage 41 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91721 | 45.594 | Unclassified | Unclassified | Myoviridae | Salmonella |
| MF443784 | Metallosphaera turreted icosahedral virus | Metallosphaera turreted icosahedral virus unclassified archaeal viruses Viruses | 9895 | 53.38 | Unclassified | Unclassified | Unclassified | Metallosphaera |
| MF443783 | Metallosphaera turreted icosahedral virus | Metallosphaera turreted icosahedral virus unclassified archaeal viruses Viruses | 9780 | 53.998 | Unclassified | Unclassified | Unclassified | Metallosphaera |
| MH220877 | Oenococcus phage phiOE33PA | Oenococcus phage phiOE33PA Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39866 | 36.287 | Unclassified | Unclassified | Siphoviridae | Oenococcus |
| LT999987 | Pseudomonas phage vB_PaeS_PAO1_HW12 | Pseudomonas phage vB_PaeS_PAO1_HW12 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37496 | 64.268 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| LT996082 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 28069 | 35.185 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996081 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 34470 | 36.115 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996080 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 12974 | 36.396 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996079 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 11055 | 34.826 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996078 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 26652 | 37.273 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996077 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 7439 | 37.666 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996076 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 36866 | 35.664 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996075 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 14429 | 36.641 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996074 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 15005 | 37.148 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996073 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 5111 | 37.155 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT996072 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 23491 | 35.695 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT993247 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 6633 | 38.339 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT993229 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 36216 | 35.026 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT993227 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 15727 | 35.207 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT993225 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 16305 | 36.983 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT993224 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 21235 | 35.814 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985593 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 9951 | 36.559 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985592 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 5025 | 38.109 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985591 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 6905 | 34.584 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985590 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 17506 | 36.896 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985589 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 7398 | 37.524 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985588 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 8385 | 37.102 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985587 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 5758 | 38.746 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985586 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 14542 | 34.968 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985585 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 23587 | 35.685 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985584 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 6204 | 39.023 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985583 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 36021 | 35.032 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985582 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 24374 | 35.714 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985581 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 6622 | 38.765 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985580 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 10239 | 36.107 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985579 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 5030 | 35.686 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985578 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 11224 | 35.7 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985577 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 12763 | 36.857 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985576 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 34407 | 36.094 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985575 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 34460 | 36.106 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985574 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 36213 | 35.059 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985573 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 18019 | 36.018 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985572 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 15730 | 35.168 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985571 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 24589 | 36.11 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985570 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 35985 | 35.809 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985569 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 24166 | 35.111 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT985568 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 37620 | 35.439 | Unclassified | Unclassified | Unclassified | Unspecified |
| LT992259 | Yersinia phage fEV-1 | Yersinia phage fEV-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38622 | 52.092 | Unclassified | Unclassified | Myoviridae | Yersinia |
| LT986678 | Poophage MBI-2016a | Poophage MBI-2016a unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5752 | 40.421 | Unclassified | Unclassified | Microviridae | Unspecified |
| LT961732 | Escherichia phage SECphi27 | Escherichia phage SECphi27 Escherichia virus SECphi27 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51811 | 44.691 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| LT960610 | Bacillus phage SBSphiC | Bacillus phage SBSphiC unclassified Spounavirinae Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144615 | 40.451 | Unclassified | Spounavirinae | Herelleviridae | Bacillus |
| LT960609 | Escherichia phage SECphi18 | Escherichia phage SECphi18 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44798 | 54.804 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LT960608 | Bacillus phage SBSphiJ | Bacillus phage SBSphiJ unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156875 | 42.132 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| LT960607 | Escherichia phage SECphi17 | Escherichia phage SECphi17 unclassified Bullavirinae Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5538 | 46.009 | Unclassified | Bullavirinae | Microviridae | Escherichia |
| LT907986 | Escherichia phage vB_Eco_SLUR25 | Escherichia phage vB_Eco_SLUR25 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58998 | 44.778 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LT962907 | Yersinia phage fPS-10 | Yersinia phage fPS-10 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39179 | 45.57 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT962906 | Yersinia phage fPS16 | Yersinia phage fPS16 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39227 | 45.555 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT962475 | Yersinia phage fPS-54-ocr | Yersinia phage fPS-54-ocr Yersinia virus fPS54ocr Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40075 | 45.535 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT962380 | Yersinia phage fPS-85 | Yersinia phage fPS-85 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40429 | 45.475 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT962379 | Yersinia phage fPS-53 | Yersinia phage fPS-53 Yersinia virus fPS53 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40694 | 45.461 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961846 | Yersinia phage fPS-64 | Yersinia phage fPS-64 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39326 | 45.611 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961845 | Yersinia phage fPS-59 | Yersinia phage fPS-59 Yersinia virus fPS59 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38391 | 45.714 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961844 | Yersinia phage fPS-21 | Yersinia phage fPS-21 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39180 | 45.564 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961843 | Yersinia phage fPS-50 | Yersinia phage fPS-50 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39764 | 45.554 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961842 | Yersinia phage fPS-86 | Yersinia phage fPS-86 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39024 | 45.633 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961841 | Yersinia phage fPS-89 | Yersinia phage fPS-89 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40405 | 45.465 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961840 | Yersinia phage fPS-7 | Yersinia phage fPS-7 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38967 | 45.631 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961838 | Yersinia phage fPS-19 | Yersinia phage fPS-19 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38938 | 45.662 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961837 | Yersinia phage fPS-52 | Yersinia phage fPS-52 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39888 | 45.487 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT961836 | Yersinia phage fPS-26 | Yersinia phage fPS-26 unclassified Helsettvirus Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38792 | 45.698 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT960606 | Yersinia phage fPS-9 | Yersinia phage fPS-9 Yersinia virus fPS9 Helsettvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39034 | 45.604 | Helsettvirus | Studiervirinae | Autographiviridae | Yersinia |
| LT960552 | Yersinia phage fHe-Yen9-03 | Yersinia phage fHe-Yen9-03 unclassified Eneladusvirus Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 352596 | 31.681 | Eneladusvirus | Unclassified | Myoviridae | Yersinia |
| LT960551 | Yersinia phage fHe-Yen9-04 | Yersinia phage fHe-Yen9-04 Yersinia virus Yen9-04 Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 354378 | 31.646 | Eneladusvirus | Unclassified | Myoviridae | Yersinia |
| LT898437 | Escherichia virus MS2 | Escherichia virus MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3534 | 52.094 | Levivirus | Unclassified | Leviviridae | Escherichia |
| LT607758 | Klebsiella phage PMBT1 | Klebsiella phage PMBT1 Klebsiella virus PMBT1 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175206 | 41.882 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| LT600745 | Clostridium phage HM2 | Clostridium phage HM2 Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17470 | 29.399 | Unclassified | Unclassified | Salasmaviridae | Clostridium |
| LT598654 | Phage NCTB | Phage NCTB Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 257877 | 42.009 | Unclassified | Unclassified | Myoviridae | Unspecified |
| LT594300 | Escherichia phage LM33_P1 | Escherichia phage LM33_P1 Escherichia virus LM33P1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38979 | 50.186 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| LN890663 | Acinetobacter phage Loki | Acinetobacter phage Loki Acinetobacter virus Loki Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41308 | 44.347 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| LN885237 | Lactobacillus phage EV3 | Lactobacillus phage EV3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34834 | 36.447 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| KJ174318 | Salmonella phage vB-SalM-SJ3 | Salmonella phage vB-SalM-SJ3 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162910 | 44.377 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KF806589 | Erwinia phage Ea35-70 | Erwinia phage Ea35-70 Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271084 | 49.881 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| AP018715 | Enterococcus phage phiM1EF22 | Enterococcus phage phiM1EF22 unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143046 | 35.953 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| AP018714 | Enterococcus phage phiEF17H | Enterococcus phage phiEF17H unclassified Kochikohdavirus Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143638 | 35.964 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| MH023293 | Escherichia phage vB_EcoS Sa179lw | Escherichia phage vB_EcoS Sa179lw Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46833 | 45.974 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| MG973030 | Erwinia phage vB_EamM-Bue1 | Erwinia phage vB_EamM-Bue1 Erwinia virus Bue1 Nezavisimistyvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164037 | 50.204 | Nezavisimistyvirus | Unclassified | Ackermannviridae | Erwinia |
| KY971611 | Pseudomonas phage PspYZU08 | Pseudomonas phage PspYZU08 Pseudomonas virus PspYZU08 Pijolavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40184 | 55.261 | Pijolavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KY971610 | Pseudomonas phage PspYZU05 | Pseudomonas phage PspYZU05 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166442 | 35.479 | Unclassified | Tevenvirinae | Myoviridae | Pseudomonas |
| KY971609 | Pseudomonas phage PspYZU01 | Pseudomonas phage PspYZU01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58279 | 63.122 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MH221128 | Salmonella phage STsAS | Salmonella phage STsAS unclassified Seoulvirus Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197215 | 49.338 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| MH243438 | Escherichia phage vB_EcoM_NBG1 | Escherichia phage vB_EcoM_NBG1 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168869 | 37.652 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| MG251392 | Salmonella virus VSe102 | Salmonella virus VSe102 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86365 | 38.989 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG251391 | Salmonella virus VSe11 | Salmonella virus VSe11 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86360 | 39 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KY575286 | Klebsiella phage SH-Kp 160016 | Klebsiella phage SH-Kp 160016 unclassified Webervirus Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49170 | 50.504 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KY450753 | Klebsiella phage SH-Kp 152234 | Klebsiella phage SH-Kp 152234 Klebsiella virus SHKp152234 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40578 | 52.851 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX278419 | Pectobacterium phage PP90 | Pectobacterium phage PP90 Pectobacterium virus PP90 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44570 | 48.889 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| MF044458 | Escherichia phage ST32 | Escherichia phage ST32 Escherichia virus ST32 Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53092 | 44.137 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| MF101922 | Ruegeria phage DSS3-P22 | Ruegeria phage DSS3-P22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4248 | 57.698 | Unclassified | Unclassified | Microviridae | Ruegeria |
| MF098558 | Pseudoalteromonas phage KB12-38 | Pseudoalteromonas phage KB12-38 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78271 | 42.223 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MH203051 | Escherichia phage Gostya9 | Escherichia phage Gostya9 Escherichia virus Gostya9 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101665 | 39.371 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MH191398 | Erwinia phage Faunus | Erwinia phage Faunus Erwinia virus Faunus Faunusvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54065 | 43.649 | Faunusvirus | Cleopatravirinae | Chaseviridae | Erwinia |
| MH341453 | Listeria phage PSU-VKH-LP041 | Listeria phage PSU-VKH-LP041 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48286 | 35.82 | Unclassified | Unclassified | Myoviridae | Listeria |
| MH341452 | Listeria phage PSU-VKH-LP040 | Listeria phage PSU-VKH-LP040 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39585 | 37.092 | Unclassified | Unclassified | Siphoviridae | Listeria |
| MH341451 | Listeria phage PSU-VKH-LP019 | Listeria phage PSU-VKH-LP019 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38601 | 35.678 | Unclassified | Unclassified | Siphoviridae | Listeria |
| MH155870 | Streptomyces phage Ibantik | Streptomyces phage Ibantik Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56362 | 57.441 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| MH000604 | Streptococcus phage D1811 | Streptococcus phage D1811 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34083 | 39.025 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH000603 | Streptococcus phage D1024 | Streptococcus phage D1024 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37757 | 38.144 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MH000602 | Streptococcus phage D5842 | Streptococcus phage D5842 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36501 | 39.125 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY984068 | Erwinia phage vB_EamM_Y3 | Erwinia phage vB_EamM_Y3 Erwinia virus Y3 Sasquatchvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 261365 | 47.178 | Sasquatchvirus | Unclassified | Myoviridae | Erwinia |
| MH243439 | Escherichia phage vB_EcoM_NBG2 | Escherichia phage vB_EcoM_NBG2 Escherichia virus NBG2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166083 | 35.448 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MG596800 | Pseudomonas phage PMBT14 | Pseudomonas phage PMBT14 Pseudomonas virus PMBT14 Knuthellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47820 | 54.548 | Knuthellervirus | Unclassified | Siphoviridae | Pseudomonas |
| KY514265 | Pectobacterium phage vB_PatP_CB3 | Pectobacterium phage vB_PatP_CB3 unclassified Cbunavirus Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76169 | 48.669 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| KY514264 | Pectobacterium phage vB_PatP_CB1 | Pectobacterium phage vB_PatP_CB1 Pectobacterium virus CB1 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76041 | 48.691 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| KY549659 | Pectobacterium phage vB_PatP_CB4 | Pectobacterium phage vB_PatP_CB4 Pectobacterium virus CB4 Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76621 | 48.643 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MG966531 | Escherichia phage Ebrios | Escherichia phage Ebrios Escherichia virus Ebrios Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39752 | 52.853 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MG986485 | Escherichia phage SH2026Stx1 | Escherichia phage SH2026Stx1 Escherichia virus SH2026Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61564 | 49.391 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| MF034659 | Pasteurella phage vB_PmuP_PHB02 | Pasteurella phage vB_PmuP_PHB02 Pasteurella virus PHB02 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38670 | 40.846 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| KX587949 | Klebsiella phage phiKpS2 | Klebsiella phage phiKpS2 Klebsiella virus KpS2 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44024 | 53.998 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KY926791 | Salmonella phage SE1 (in:Nonagvirus) | Salmonella phage SE1 (in:Nonagvirus) Salmonella virus SE1Kor Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60030 | 43.385 | Nonagvirus | Queuovirinae | Siphoviridae | Salmonella |
| DQ003260 | Salmonella phage SE1 (in:P22virus) | Salmonella phage SE1 (in:P22virus) Salmonella virus SE1Spa Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 46.985 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MH179477 | Aeromonas phage 60AhydR15PP | Aeromonas phage 60AhydR15PP unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165795 | 41.233 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| MH179480 | Pseudomonas phage 98PfluR60PP | Pseudomonas phage 98PfluR60PP unclassified Littlefixvirus Littlefixvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74361 | 47.272 | Littlefixvirus | Unclassified | Schitoviridae | Pseudomonas |
| MH179471 | Aeromonas phage 14AhydR10PP | Aeromonas phage 14AhydR10PP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48335 | 55.858 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MH203384 | Enterococcus phage vB_EfaS_AL2 | Enterococcus phage vB_EfaS_AL2 Enterococcus virus AL2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40836 | 34.514 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MH179479 | Aeromonas phage 85AhydR10PP | Aeromonas phage 85AhydR10PP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47194 | 56.092 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MH179476 | Aeromonas phage 50AhydR13PP | Aeromonas phage 50AhydR13PP unclassified Tulanevirus Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164983 | 41.074 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| MH179474 | Aeromonas phage 62AhydR11PP | Aeromonas phage 62AhydR11PP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43755 | 57.196 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MH179470 | Aeromonas phage 13AhydR10PP | Aeromonas phage 13AhydR10PP Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47828 | 55.804 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| LC348380 | Escherichia phage T2 | Escherichia phage T2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163825 | 35.318 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LC348379 | Escherichia phage PP01 | Escherichia phage PP01 Escherichia virus PP01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167812 | 35.459 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MH105080 | Marinomonas phage YY | Marinomonas phage YY unclassified Corticoviridae Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 8828 | 57.046 | Unclassified | Unclassified | Corticoviridae | Marinomonas |
| MF155889 | Vibrio virus CTXphi | Vibrio virus CTXphi Affertcholeramvirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6698 | 43.595 | Affertcholeramvirus | Unclassified | Inoviridae | Vibrio |
| MH059638 | Pectobacterium phage Nepra | Pectobacterium phage Nepra Pectobacterium virus Nepra Cbunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74509 | 48.701 | Cbunavirus | Unclassified | Schitoviridae | Pectobacterium |
| MH059633 | Xanthomonas phage Carpasina | Xanthomonas phage Carpasina Xanthomonas virus Carpasina Carpasinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61939 | 52.356 | Carpasinavirus | Unclassified | Myoviridae | Xanthomonas |
| MH193369 | Enterococcus phage LY0322 | Enterococcus phage LY0322 Enterococcus virus LY0322 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40934 | 34.841 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MH051918 | Escherichia phage vB_EcoS_IME347 | Escherichia phage vB_EcoS_IME347 Escherichia virus IME347 Badaguanvirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50048 | 49.706 | Badaguanvirus | Tunavirinae | Drexlerviridae | Escherichia |
| MH051917 | Enterobacteria phage vB_EcoM_IME341 | Enterobacteria phage vB_EcoM_IME341 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172379 | 39.515 | Dhakavirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH051916 | Enterobacteria phage vB_EcoM_IME340 | Enterobacteria phage vB_EcoM_IME340 Enterobacteria virus IME340 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165549 | 35.549 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH051915 | Enterobacteria phage vB_EcoM_IME339 | Enterobacteria phage vB_EcoM_IME339 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164366 | 35.64 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH051914 | Enterobacteria phage vB_EcoM_IME338 | Enterobacteria phage vB_EcoM_IME338 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85675 | 38.78 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| MH051913 | Enterobacteria phage vB_EcoM_IME281 | Enterobacteria phage vB_EcoM_IME281 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170531 | 39.442 | Dhakavirus | Tevenvirinae | Myoviridae | Enterobacteria |
| MH051912 | Enterobacteria phage vB_EcoS_IME167 | Enterobacteria phage vB_EcoS_IME167 unclassified Tunavirus Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49794 | 45.548 | Tunavirus | Tunavirinae | Drexlerviridae | Enterobacteria |
| MH051911 | Enterobacteria phage vB_EcoS_IME18 | Enterobacteria phage vB_EcoS_IME18 Escherichia virus IME18 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50354 | 45.615 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MH128984 | Burkholderia phage vB_BmuP_KL4 | Burkholderia phage vB_BmuP_KL4 Burkholderia virus KL4 Kelquatrovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42250 | 63.183 | Kelquatrovirus | Unclassified | Podoviridae | Burkholderia |
| MH105773 | Vibrio phage Rostov 6 | Vibrio phage Rostov 6 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39934 | 50.739 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| MH113815 | Pseudomonas phage Alpheus | Pseudomonas phage Alpheus Pseudomonas virus Alpheus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45756 | 57.824 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MH113814 | Pseudomonas phage Achelous | Pseudomonas phage Achelous Pseudomonas virus Achelous Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46585 | 57.345 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MH113813 | Pseudomonas phage Nerthus | Pseudomonas phage Nerthus Pseudomonas virus Nerthus Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45779 | 58.505 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MH113812 | Pseudomonas phage Njord | Pseudomonas phage Njord Pseudomonas virus Njord Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46644 | 56.62 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MH059635 | Dickeya phage Amaethon | Dickeya phage Amaethon unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41436 | 39.791 | Kafunavirus | Unclassified | Podoviridae | Dickeya |
| MH059634 | Dickeya phage Sucellus | Dickeya phage Sucellus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39826 | 41.902 | Unclassified | Unclassified | Siphoviridae | Dickeya |
| MH107246 | Synechococcus phage S-B64 | Synechococcus phage S-B64 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151867 | 41.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KY379511 | Pseudoalteromonas phage PHS21 | Pseudoalteromonas phage PHS21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35802 | 40.626 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KX987999 | Bacillus phage BCP12 | Bacillus phage BCP12 unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155985 | 39.846 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX257490 | Pseudoalteromonas virus vB_PspP-H6/1 | Pseudoalteromonas virus vB_PspP-H6/1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36753 | 45.033 | Unclassified | Unclassified | Podoviridae | Pseudoalteromonas |
| MG710484 | Bacillus phage BtiUFT6.51-F | Bacillus phage BtiUFT6.51-F unclassified Camtrevirus Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42076 | 35.334 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| MG656408 | Staphylococcus phage B1 | Staphylococcus phage B1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148884 | 30.254 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MF405094 | Staphylococcus phage JA1 | Staphylococcus phage JA1 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147135 | 30.252 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY448244 | Erwinia phage vB_EamM_Yoloswag | Erwinia phage vB_EamM_Yoloswag Erwinia virus Yoloswag Yoloswagvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 259700 | 46.915 | Yoloswagvirus | Unclassified | Myoviridae | Erwinia |
| LT986460 | Pseudomonas phage GP100 | Pseudomonas phage GP100 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50547 | 50.949 | Unclassified | Unclassified | Zobellviridae | Pseudomonas |
| MG793454 | Mycobacterium phage SWU2 | Mycobacterium phage SWU2 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50013 | 62.666 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MH107770 | Pseudomonas phage vB_PaeP_130_113 | Pseudomonas phage vB_PaeP_130_113 Pseudomonas virus 130-113 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44205 | 62.357 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MH107769 | Staphylococcus phage vB_SauM_0414_108 | Staphylococcus phage vB_SauM_0414_108 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151627 | 30.388 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG966533 | Pseudoalteromonas phage Cr39582 | Pseudoalteromonas phage Cr39582 Pseudoalteromonas virus Cr39582 Corticovirus Corticoviridae Vinavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10584 | 41.903 | Corticovirus | Unclassified | Corticoviridae | Pseudoalteromonas |
| MG652450 | Ralstonia phage RsoP1IDN | Ralstonia phage RsoP1IDN Ralstonia virus RsoP1IDN Higashivirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41135 | 62.747 | Higashivirus | Okabevirinae | Autographiviridae | Ralstonia |
| HE956708 | Yersinia phage phiR2-01 | Yersinia phage phiR2-01 Yersinia virus phiR201 Haartmanvirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122696 | 40.512 | Haartmanvirus | Markadamsvirinae | Demerecviridae | Yersinia |
| KY914478 | Escherichia phage ArgO145 | Escherichia phage ArgO145 Escherichia virus ArgO145 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62020 | 50.002 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| MG775042 | Citrobacter phage vB_CroP_CrRp3 | Citrobacter phage vB_CroP_CrRp3 Citrobacter CrRp3 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44349 | 45.155 | Vectrevirus | Molineuxvirinae | Autographiviridae | Citrobacter |
| MG775043 | Citrobacter phage vB_CroM_CrRp10 | Citrobacter phage vB_CroM_CrRp10 Citrobacter virus CrRp10 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168403 | 35.508 | Tequatrovirus | Tevenvirinae | Myoviridae | Citrobacter |
| MF547663 | Clostridioides phage LIBA2945 | Clostridioides phage LIBA2945 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132508 | 26.272 | Unclassified | Unclassified | Siphoviridae | Clostridioides |
| MF547662 | Clostridioides phage LIBA6276 | Clostridioides phage LIBA6276 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134243 | 26.141 | Unclassified | Unclassified | Siphoviridae | Clostridioides |
| MF278336 | Alteromonas virus vB_AspP-H4/4 | Alteromonas virus vB_AspP-H4/4 Alteromonas virus H4-4 Foturvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47631 | 40.761 | Foturvirus | Unclassified | Autographiviridae | Alteromonas |
| KU747973 | Pseudoalteromonas phage vB_PspS-H40/1 | Pseudoalteromonas phage vB_PspS-H40/1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45306 | 40.147 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MG018929 | Pseudomonas phage uligo | Pseudomonas phage uligo Pseudomonas virus uligo Uliginvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47868 | 56.301 | Uliginvirus | Colwellvirinae | Autographiviridae | Pseudomonas |
| MG727702 | Paenibacillus phage Likha | Paenibacillus phage Likha Paenibacillus virus Likha Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39778 | 41.302 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727700 | Paenibacillus phage Tadhana | Paenibacillus phage Tadhana Paenibacillus virus Tadhana Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37880 | 42.112 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727699 | Paenibacillus phage Pagassa | Paenibacillus phage Pagassa Paenibacillus virus Pagassa Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40035 | 42.001 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727696 | Paenibacillus phage Kiel007 | Paenibacillus phage Kiel007 unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37985 | 41.811 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727695 | Paenibacillus phage BN12 | Paenibacillus phage BN12 unclassified Sitaravirus Sitaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39485 | 42.56 | Sitaravirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727698 | Paenibacillus phage PBL1c | Paenibacillus phage PBL1c Paenibacillus virus PBL1c Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40611 | 41.247 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| MG727697 | Paenibacillus phage Dragolir | Paenibacillus phage Dragolir Paenibacillus virus Dragolir Dragolirvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41131 | 43.981 | Dragolirvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| MG775261 | Pseudomonas phage PollyC | Pseudomonas phage PollyC Pseudomonas virus PollyC Pollyceevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41651 | 57.595 | Pollyceevirus | Unclassified | Autographiviridae | Pseudomonas |
| MG775260 | Pseudomonas phage Littlefix | Pseudomonas phage Littlefix Pseudomonas virus Littlefix Littlefixvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74895 | 48.734 | Littlefixvirus | Unclassified | Schitoviridae | Pseudomonas |
| MG775259 | Pseudomonas phage Bjorn | Pseudomonas phage Bjorn Pseudomonas virus Bjorn Bjornvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45936 | 53.076 | Bjornvirus | Unclassified | Podoviridae | Pseudomonas |
| MG775258 | Pseudomonas phage Henninger | Pseudomonas phage Henninger Pseudomonas virus Henninger Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40923 | 57.586 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| MG099933 | Escherichia phage VB_EcoS-Golestan | Escherichia phage VB_EcoS-Golestan Escherichia virus Golestan Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44829 | 50.601 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| MG676225 | Aeromonas phage AhSzw-1 | Aeromonas phage AhSzw-1 Aeromonas virus AhSzw1 Shenzhenvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115739 | 43.82 | Shenzhenvirus | Unclassified | Demerecviridae | Aeromonas |
| MG676224 | Aeromonas phage AhSzq-1 | Aeromonas phage AhSzq-1 Aeromonas virus AhSzq1 Shenzhenvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112558 | 43.861 | Shenzhenvirus | Unclassified | Demerecviridae | Aeromonas |
| MH107028 | Campylobacter phage CP39 | Campylobacter phage CP39 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130715 | 26.044 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| MH028956 | Staphylococcus phage phiSA_BS2 | Staphylococcus phage phiSA_BS2 Staphylococcus virus BS2 Baoshanvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149229 | 29.608 | Baoshanvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG999954 | Enterobacter phage myPSH1140 | Enterobacter phage myPSH1140 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172614 | 39.883 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| MG711516 | Ralstonia phage RsoP1EGY | Ralstonia phage RsoP1EGY Ralstonia virus RsoP1EGY Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41297 | 58.97 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MG983840 | Escherichia phage myPSH1131 | Escherichia phage myPSH1131 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76163 | 42.253 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MH102284 | Salmonella phage vB_SenS_PHB07 | Salmonella phage vB_SenS_PHB07 Salmonella virus PHB07 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51818 | 43.359 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| MG266157 | Dickeya phage PP35 | Dickeya phage PP35 unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152048 | 49.304 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| MF893340 | Pseudomonas phage PPSC2 | Pseudomonas phage PPSC2 unclassified Otagovirus Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97330 | 47.51 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| KY744566 | Pectobacterium phage POP72 | Pectobacterium phage POP72 unclassified Axomammavirus Axomammavirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44760 | 49.707 | Axomammavirus | Molineuxvirinae | Autographiviridae | Pectobacterium |
| KY618819 | Pseudomonas phage PAXYB1 | Pseudomonas phage PAXYB1 Pseudomonas virus PAXYB1 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43337 | 62.289 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MG976803 | Escherichia phage myPSH2311 | Escherichia phage myPSH2311 unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68712 | 42.367 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MH042230 | Acinetobacter phage vB_AbaP_D2 | Acinetobacter phage vB_AbaP_D2 Acinetobacter virus D2 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39964 | 39.233 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MH078572 | Staphylococcus phage phiSA_BS1 | Staphylococcus phage phiSA_BS1 Staphylococcus virus BS1 Baoshanvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142978 | 29.799 | Baoshanvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG972768 | Klebsiella phage myPSH1235 | Klebsiella phage myPSH1235 Klebsiella virus myPSH1235 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45135 | 53.663 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| AP018361 | Lactobacillus phage T25 | Lactobacillus phage T25 Lactobacillus virus T25 Sukhumvitvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38421 | 44.541 | Sukhumvitvirus | Unclassified | Siphoviridae | Lactobacillus |
| MG969427 | Anoxybacillus phage A403 | Anoxybacillus phage A403 Anoxybacillus virus A403 Tandoganvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40847 | 36.769 | Tandoganvirus | Unclassified | Siphoviridae | Anoxybacillus |
| MG969415 | Salmonella phage GE_vB_TR | Salmonella phage GE_vB_TR Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42436 | 47.658 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MG969414 | Salmonella phage GE_vB_NS7 | Salmonella phage GE_vB_NS7 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55248 | 38.742 | Unclassified | Ounavirinae | Myoviridae | Salmonella |
| MG969413 | Salmonella phage GE_vB_N8 | Salmonella phage GE_vB_N8 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51134 | 38.831 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MG969412 | Salmonella phage GE_vB_N5 | Salmonella phage GE_vB_N5 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148669 | 39.488 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MG969411 | Salmonella phage GE_vB_MG | Salmonella phage GE_vB_MG unclassified Seunavirus Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172009 | 46.049 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| MG969410 | Salmonella phage GE_vB_M5 | Salmonella phage GE_vB_M5 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44432 | 49.572 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MG969409 | Salmonella phage GE_vB_M4 | Salmonella phage GE_vB_M4 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44895 | 49.787 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MG969408 | Salmonella phage GE_vB_HIL | Salmonella phage GE_vB_HIL unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45334 | 49.724 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MG969407 | Salmonella phage GE_vB_BS | Salmonella phage GE_vB_BS unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90041 | 39.566 | Unclassified | Ounavirinae | Myoviridae | Salmonella |
| MG969406 | Salmonella phage GE_vB_B3 | Salmonella phage GE_vB_B3 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109375 | 39.196 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG969405 | Salmonella phage GE_vB_B1 | Salmonella phage GE_vB_B1 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59703 | 38.911 | Unclassified | Ounavirinae | Myoviridae | Salmonella |
| MG969404 | Salmonella phage GE_vB_7A | Salmonella phage GE_vB_7A unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85780 | 38.957 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG788324 | Lactobacillus phage PM411 | Lactobacillus phage PM411 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69649 | 42.311 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| MG732930 | Enterobacter phage Ec_L1 | Enterobacter phage Ec_L1 Enterobacter virus EcL1 Eclunavirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51894 | 48.241 | Eclunavirus | Unclassified | Drexlerviridae | Enterobacter |
| MF580775 | Streptococcus phage V2 | Streptococcus phage V2 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34698 | 38.714 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580774 | Streptococcus phage STP2 | Streptococcus phage STP2 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35286 | 38.452 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580773 | Streptococcus phage STP1 | Streptococcus phage STP1 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35283 | 38.492 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580772 | Streptococcus phage R1 | Streptococcus phage R1 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34652 | 38.737 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580771 | Streptococcus phage MM25 | Streptococcus phage MM25 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34059 | 38.976 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580770 | Streptococcus phage M19 | Streptococcus phage M19 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34328 | 38.866 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580769 | Streptococcus phage L5A1 | Streptococcus phage L5A1 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36884 | 38.86 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580768 | Streptococcus phage C0 | Streptococcus phage C0 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34718 | 38.957 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580767 | Streptococcus phage B5 | Streptococcus phage B5 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35636 | 38.554 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580766 | Streptococcus phage B0 | Streptococcus phage B0 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36773 | 38.452 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580764 | Streptococcus phage 31B4 | Streptococcus phage 31B4 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36438 | 38.946 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580763 | Streptococcus phage 16B8 | Streptococcus phage 16B8 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34379 | 38.617 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580762 | Streptococcus phage 9B4 | Streptococcus phage 9B4 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34636 | 38.659 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580761 | Streptococcus phage 9A | Streptococcus phage 9A unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35604 | 38.962 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580759 | Streptococcus phage 7A5 | Streptococcus phage 7A5 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35936 | 38.304 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580765 | Streptococcus phage A0 | Streptococcus phage A0 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35481 | 38.849 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MF580760 | Streptococcus phage 7T | Streptococcus phage 7T unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35141 | 38.727 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| CP027117 | Bacillus phage EZ-2018a | Bacillus phage EZ-2018a unclassified Jenstvirus Jenstvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118485 | 39.9 | Jenstvirus | Unclassified | Siphoviridae | Bacillus |
| LN866626 | Klebsiella phage vB_KpnP_KpV289 | Klebsiella phage vB_KpnP_KpV289 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41054 | 52.565 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| AJ748296 | Sulfolobus islandicus rudivirus 1 variant XX | Sulfolobus islandicus rudivirus 1 variant XX Icerudivirus SIRV1 Icerudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 33641 | 25.282 | Icerudivirus | Unclassified | Rudiviridae | Sulfolobus |
| MF774689 | Salmonella phage SP1a | Salmonella phage SP1a unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14427 | 39.142 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MF774688 | Salmonella phage SP1a | Salmonella phage SP1a unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47894 | 38.713 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MF774687 | Salmonella phage SP1a | Salmonella phage SP1a unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44399 | 38.638 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MG963916 | Escherichia phage PDX | Escherichia phage PDX unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138427 | 43.548 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| MG941013 | Actinomyces phage xhp1 | Actinomyces phage xhp1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35031 | 65.291 | Unclassified | Unclassified | Siphoviridae | Actinomyces |
| MG897799 | Pseudomonas phage vB_PaeM_LS1 | Pseudomonas phage vB_PaeM_LS1 Pseudomonas virus LS1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66095 | 55.693 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MG812496 | Gordonia phage SallySpecial | Gordonia phage SallySpecial Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15896 | 70.131 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| MG873442 | Salmonella phage SE131 | Salmonella phage SE131 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89910 | 44.218 | Unclassified | Unclassified | Podoviridae | Salmonella |
| MG897800 | Pseudomonas phage vB_PaeS_C1 | Pseudomonas phage vB_PaeS_C1 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43133 | 53.576 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| KY939598 | Alteromonadaceae phage B23 | Alteromonadaceae phage B23 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35132 | 59.507 | Unclassified | Unclassified | Myoviridae | Unspecified |
| MG894052 | Klebsiella phage JY917 | Klebsiella phage JY917 Klebsiella virus JY917 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37655 | 50.432 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MF288922 | Bacillus phage Janet | Bacillus phage Janet unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160705 | 37.964 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF288921 | Bacillus phage OTooleKemple52 | Bacillus phage OTooleKemple52 unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161807 | 37.884 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF288919 | Bacillus phage AaronPhadgers | Bacillus phage AaronPhadgers unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161772 | 38.654 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF288918 | Bacillus phage Bubs | Bacillus phage Bubs unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162449 | 38.78 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF288917 | Bacillus phage PPIsBest | Bacillus phage PPIsBest unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162281 | 38.637 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| AP018375 | Staphylococcus phage phiSA039 | Staphylococcus phage phiSA039 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141038 | 30.378 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY787213 | Salmonella phage BSP101 | Salmonella phage BSP101 Salmonella virus BSP101 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157665 | 44.452 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KY787212 | Salmonella phage BSP22A | Salmonella phage BSP22A unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110741 | 40.079 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MG214783 | Sinorhizobium phage HMSP1-Susan | Sinorhizobium phage HMSP1-Susan Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51963 | 52.512 | Unclassified | Unclassified | Siphoviridae | Sinorhizobium |
| MG845683 | Pseudomonas phage phiNV3 | Pseudomonas phage phiNV3 Pseudomonas virus NV3 Kirikabuvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43184 | 58.29 | Kirikabuvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| MG765274 | Lactobacillus phage Maenad | Lactobacillus phage Maenad Lactobacillus virus Maenad Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78646 | 39.074 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MG833025 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38553 | 48.391 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MG957431 | Vibrio phage Rostov-1 | Vibrio phage Rostov-1 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37247 | 42.774 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY684082 | Shigella phage SFN6B | Shigella phage SFN6B Shigella virus SFN6B Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43036 | 50.943 | Drulisvirus | Slopekvirinae | Autographiviridae | Shigella |
| KY620117 | Salmonella phage BSPM4 | Salmonella phage BSPM4 Salmonella virus BSPM4 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59097 | 56.541 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KY963424 | Shigella phage SSP1 | Shigella phage SSP1 Shigella virus SSP1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113299 | 39.076 | Tequintavirus | Markadamsvirinae | Demerecviridae | Shigella |
| MF775705 | Lactococcus phage LP1502c | Lactococcus phage LP1502c unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30247 | 35.108 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775704 | Lactococcus phage LP1502b | Lactococcus phage LP1502b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30227 | 35.114 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775703 | Lactococcus phage LP1502a | Lactococcus phage LP1502a Lactococcus virus LP1502a Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30247 | 35.121 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775702 | Lactococcus phage LP1407 | Lactococcus phage LP1407 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30508 | 34.909 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775701 | Lactococcus phage LP1110 | Lactococcus phage LP1110 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30528 | 34.889 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775700 | Lactococcus phage LP1011 | Lactococcus phage LP1011 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30542 | 34.87 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775699 | Lactococcus phage LP1005 | Lactococcus phage LP1005 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30541 | 34.891 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775698 | Lactococcus phage LP0903 | Lactococcus phage LP0903 Lactococcus virus LP0903 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31049 | 34.9 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775697 | Lactococcus phage LP0604 | Lactococcus phage LP0604 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30547 | 34.933 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775696 | Lactococcus phage LP0509 | Lactococcus phage LP0509 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30547 | 34.936 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775695 | Lactococcus phage LP0304 | Lactococcus phage LP0304 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30344 | 34.883 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775694 | Lactococcus phage LP0212 | Lactococcus phage LP0212 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29916 | 34.767 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775693 | Lactococcus phage LP0209 | Lactococcus phage LP0209 Lactococcus virus LP0209 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30339 | 34.761 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775692 | Lactococcus phage LP0202 | Lactococcus phage LP0202 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30236 | 34.932 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775691 | Lactococcus phage LP0109 | Lactococcus phage LP0109 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30537 | 34.83 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775690 | Lactococcus phage LP0004d | Lactococcus phage LP0004d unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30104 | 34.816 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775689 | Lactococcus phage LP0004c | Lactococcus phage LP0004c unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30044 | 34.799 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775688 | Lactococcus phage LP0004b | Lactococcus phage LP0004b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30953 | 34.73 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775686 | Lactococcus phage LP9908 | Lactococcus phage LP9908 Lactococcus virus LP9908 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29668 | 34.829 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775685 | Lactococcus phage LP9903 | Lactococcus phage LP9903 Lactococcus virus LP9903 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29557 | 34.96 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775684 | Lactococcus phage LP9801 | Lactococcus phage LP9801 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30204 | 34.86 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775683 | Lactococcus phage LP9701 | Lactococcus phage LP9701 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30545 | 34.919 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775682 | Lactococcus phage LP9609 | Lactococcus phage LP9609 Lactococcus virus LP9609 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30945 | 34.765 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775681 | Lactococcus phage LP9406 | Lactococcus phage LP9406 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30237 | 34.838 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775680 | Lactococcus phage LP9405b | Lactococcus phage LP9405b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29294 | 34.929 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775679 | Lactococcus phage LP9405a | Lactococcus phage LP9405a unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30083 | 34.973 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775678 | Lactococcus phage LP9404 | Lactococcus phage LP9404 Lactococcus virus LP9404 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30072 | 34.946 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775677 | Lactococcus phage LP9210 | Lactococcus phage LP9210 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29525 | 35.143 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775676 | Lactococcus phage LP9207 | Lactococcus phage LP9207 Lactococcus virus LP9207 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29203 | 35.01 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775675 | Lactococcus phage LP9206c | Lactococcus phage LP9206c unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29291 | 34.956 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775674 | Lactococcus phage LP9206b | Lactococcus phage LP9206b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30309 | 34.854 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775673 | Lactococcus phage LP9206a | Lactococcus phage LP9206a unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29097 | 34.949 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775672 | Lactococcus phage LP9205b | Lactococcus phage LP9205b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30296 | 34.816 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775671 | Lactococcus phage LP9205a | Lactococcus phage LP9205a unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29098 | 34.944 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775670 | Lactococcus phage LP9104 | Lactococcus phage LP9104 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29157 | 35.155 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775669 | Lactococcus phage LP8511 | Lactococcus phage LP8511 Lactococcus virus LP8511 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29953 | 35.112 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF775687 | Lactococcus phage LP0004a | Lactococcus phage LP0004a unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30093 | 34.819 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MF431741 | Bacteriophage T5-like saus176N | Bacteriophage T5-like saus176N unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.745 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431740 | Bacteriophage T5-like cott162 | Bacteriophage T5-like cott162 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.744 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431739 | Bacteriophage T5-like chee158 | Bacteriophage T5-like chee158 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.746 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431738 | Bacteriophage T5-like poul149 | Bacteriophage T5-like poul149 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.746 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431737 | Escherichia phage saus132 | Escherichia phage saus132 Escherichia virus saus132 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.745 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MF431736 | Bacteriophage T5-like chee130_1 | Bacteriophage T5-like chee130_1 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121986 | 39.745 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431735 | Bacteriophage T5-like poul124 | Bacteriophage T5-like poul124 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120629 | 39.264 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431734 | Bacteriophage T5-like saus111K | Bacteriophage T5-like saus111K unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120620 | 39.263 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431733 | Bacteriophage T5-like saus47N | Bacteriophage T5-like saus47N unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120622 | 39.262 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431732 | Bacteriophage T5-like pork29 | Bacteriophage T5-like pork29 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120622 | 39.263 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431731 | Bacteriophage T5-like pork27 | Bacteriophage T5-like pork27 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120618 | 39.264 | Tequintavirus | Markadamsvirinae | Demerecviridae | Unspecified |
| MF431730 | Escherichia phage chee24 | Escherichia phage chee24 Escherichia virus chee24 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120622 | 39.263 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MG766219 | Staphylococcus phage vB_SauP_phiAGO1.9 | Staphylococcus phage vB_SauP_phiAGO1.9 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17637 | 28.962 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MG710528 | Escherichia phage GER2 | Escherichia phage GER2 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88114 | 49.243 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| MG366115 | Acinetobacter phage PBAB25 | Acinetobacter phage PBAB25 unclassified Lokivirus Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40260 | 42.837 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| MG366114 | Acinetobacter phage PBAB08 | Acinetobacter phage PBAB08 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42312 | 38.573 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| KU737350 | Bacillus phage SageFayge | Bacillus phage SageFayge unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162359 | 38.698 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| LC371242 | Escherichia phage EcS1 | Escherichia phage EcS1 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175437 | 37.829 | Winklervirus | Unclassified | Myoviridae | Escherichia |
| AY129339 | Mycobacterium phage Barnyard | Mycobacterium phage Barnyard Barnyardvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70797 | 57.313 | Barnyardvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG025518 | Gokushovirus MK-2017 | Gokushovirus MK-2017 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4907 | 41.329 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| MG820644 | Propionibacterium phage pa33 | Propionibacterium phage pa33 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29135 | 54.412 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820643 | Propionibacterium phage pa35 | Propionibacterium phage pa35 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.271 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820642 | Propionibacterium phage pa28 | Propionibacterium phage pa28 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29733 | 54.138 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820641 | Propionibacterium phage pa615 | Propionibacterium phage pa615 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29307 | 54.414 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820640 | Propionibacterium phage pa15 | Propionibacterium phage pa15 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29309 | 54.29 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820639 | Propionibacterium phage pa9-6919-4 | Propionibacterium phage pa9-6919-4 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29784 | 54.079 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820638 | Propionibacterium phage pa6919-4 | Propionibacterium phage pa6919-4 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29784 | 54.079 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820637 | Propionibacterium phage pa63 | Propionibacterium phage pa63 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29446 | 53.912 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820636 | Propionibacterium phage pa29399-1-D_2 | Propionibacterium phage pa29399-1-D_2 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28892 | 54.378 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820635 | Propionibacterium phage pa29399-1-D_1 | Propionibacterium phage pa29399-1-D_1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29370 | 54.072 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820634 | Propionibacterium phage pa27 | Propionibacterium phage pa27 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29593 | 53.915 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820633 | Propionibacterium phage pa59 | Propionibacterium phage pa59 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29507 | 53.858 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820632 | Propionibacterium phage pa310 | Propionibacterium phage pa310 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29508 | 53.853 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MG820645 | Propionibacterium phage pa3-SS3 | Propionibacterium phage pa3-SS3 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29137 | 54.412 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| MF490241 | Pseudomonas phage vB_PaeM_E215 | Pseudomonas phage vB_PaeM_E215 Pseudomonas virus E215 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66789 | 55.629 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MF490240 | Pseudomonas phage vB_PaeM_E217 | Pseudomonas phage vB_PaeM_E217 Pseudomonas virus E217 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66291 | 55.647 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MF490239 | Pseudomonas phage vB_PaeS_S218 | Pseudomonas phage vB_PaeS_S218 unclassified Yuavirus Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61680 | 64.45 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| MF490238 | Pseudomonas phage vB_PaeP_DEV | Pseudomonas phage vB_PaeP_DEV unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72697 | 54.862 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| MF490237 | Pseudomonas phage vB_PaeP_E220 | Pseudomonas phage vB_PaeP_E220 unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62874 | 63.832 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MF490236 | Pseudomonas phage vB_PaeP_PYO2 | Pseudomonas phage vB_PaeP_PYO2 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72697 | 54.843 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| HG792103 | Escherichia phage P8983 | Escherichia phage P8983 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61343 | 50.231 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| MG845684 | Pseudomonas phage NV1 | Pseudomonas phage NV1 Pseudomonas virus NV1 Vicosavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45058 | 52.115 | Vicosavirus | Unclassified | Podoviridae | Pseudomonas |
| MG845865 | Escherichia phage YZ1 | Escherichia phage YZ1 Escherichia virus YZ1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38823 | 50.375 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MF403009 | Agrobacterium phage Atu_ph08 | Agrobacterium phage Atu_ph08 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58966 | 59.672 | Unclassified | Unclassified | Podoviridae | Agrobacterium |
| MF403008 | Agrobacterium phage Atu_ph07 | Agrobacterium phage Atu_ph07 Agrobacterium virus Atuph07 Polybotosvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 490380 | 37.06 | Polybotosvirus | Unclassified | Myoviridae | Agrobacterium |
| MF403007 | Agrobacterium phage Atu_ph04 | Agrobacterium phage Atu_ph04 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143349 | 49.44 | Unclassified | Unclassified | Ackermannviridae | Agrobacterium |
| MF403006 | Agrobacterium phage Atu_ph03 | Agrobacterium phage Atu_ph03 Agrobacterium virus Atuph03 Atuphduovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45175 | 54.734 | Atuphduovirus | Unclassified | Autographiviridae | Agrobacterium |
| MF403005 | Agrobacterium phage Atu_ph02 | Agrobacterium phage Atu_ph02 Agrobacterium virus Atuph02 Atuphduovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45423 | 54.829 | Atuphduovirus | Unclassified | Autographiviridae | Agrobacterium |
| CP018841 | Staphylococcus phage HOB 14.1.R1 | Staphylococcus phage HOB 14.1.R1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18659 | 32.885 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| AP018476 | Mycobacterium phage C3 | Mycobacterium phage C3 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51141 | 63.122 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018471 | Mycobacterium phage Y10 | Mycobacterium phage Y10 Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57990 | 67.888 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018469 | Mycobacterium phage Y10 | Mycobacterium phage Y10 Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57990 | 67.882 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF327006 | Shigella phage Sf20 | Shigella phage Sf20 unclassified Krischvirus Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163982 | 40.564 | Krischvirus | Tevenvirinae | Myoviridae | Shigella |
| MF327007 | Shigella phage Sf21 | Shigella phage Sf21 Shigella virus Sf21 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166002 | 35.451 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MF327009 | Shigella phage Sf25 | Shigella phage Sf25 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168573 | 35.346 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MF327008 | Shigella phage Sf24 | Shigella phage Sf24 Shigella virus Sf24 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168112 | 35.321 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MF327004 | Shigella phage Sf17 | Shigella phage Sf17 Shigella virus Sf17 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90092 | 39.032 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MF327003 | Shigella phage Sf14 | Shigella phage Sf14 Shigella virus Sf14 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87575 | 39.065 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MG835858 | Klebsiella phage KPN N98 | Klebsiella phage KPN N98 unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59214 | 56.289 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MG832642 | Escherichia phage HZ2R8 | Escherichia phage HZ2R8 Escherichia virus HZ2R8 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39482 | 48.615 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MG711460 | Faecalibacterium phage FP_Mushu | Faecalibacterium phage FP_Mushu Faecalibacterium virus Mushu Mushuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36636 | 60.31 | Mushuvirus | Unclassified | Myoviridae | Faecalibacterium |
| MG835450 | Pontimonas phage phiPsal1 | Pontimonas phage phiPsal1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31069 | 53.278 | Unclassified | Unclassified | Siphoviridae | Pontimonas |
| MG765276 | Lactobacillus phage Nyseid | Lactobacillus phage Nyseid Lactobacillus virus Nyseid Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79221 | 37.067 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MG770897 | Staphylococcus phage SH-St 15644 | Staphylococcus phage SH-St 15644 Staphylococcus virus 15644 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45111 | 33.349 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| MG770379 | Klebsiella phage vB_KpnM_KpS110 | Klebsiella phage vB_KpnM_KpS110 Klebsiella virus KpS110 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156801 | 46.143 | Taipeivirus | Unclassified | Ackermannviridae | Klebsiella |
| MG763894 | Bacillus phage HonestAbe | Bacillus phage HonestAbe unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161129 | 38.652 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MG751100 | Klebsiella phage KP1 | Klebsiella phage KP1 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167989 | 39.601 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| MG744354 | Lactobacillus phage Satyr | Lactobacillus phage Satyr Lactobacillus virus Satyr Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81076 | 39.417 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MG736918 | Erwinia phage vB_EamP-S2 | Erwinia phage vB_EamP-S2 Erwinia virus S2 Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45495 | 49.792 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| MG721208 | Staphylococcus phage vB_SauM_LM12 | Staphylococcus phage vB_SauM_LM12 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143652 | 30.247 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MG711467 | Faecalibacterium phage FP_Taranis | Faecalibacterium phage FP_Taranis Faecalibacterium virus Taranis Taranisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54286 | 55.799 | Taranisvirus | Unclassified | Myoviridae | Faecalibacterium |
| MG711466 | Faecalibacterium phage FP_Toutatis | Faecalibacterium phage FP_Toutatis Faecalibacterium virus Toutatis Toutatisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54725 | 53.906 | Toutatisvirus | Unclassified | Myoviridae | Faecalibacterium |
| MG711465 | Faecalibacterium phage FP_Brigit | Faecalibacterium phage FP_Brigit Faecalibacterium virus Brigit Brigitvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61716 | 55.796 | Brigitvirus | Unclassified | Myoviridae | Faecalibacterium |
| MG711464 | Faecalibacterium phage FP_Lugh | Faecalibacterium phage FP_Lugh Faecalibacterium virus Lugh Lughvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34075 | 59.894 | Lughvirus | Unclassified | Siphoviridae | Faecalibacterium |
| MG711463 | Faecalibacterium phage FP_oengus | Faecalibacterium phage FP_oengus Faecalibacterium virus Oengus Oengusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58391 | 49.794 | Oengusvirus | Unclassified | Siphoviridae | Faecalibacterium |
| MG711462 | Faecalibacterium phage FP_Epona | Faecalibacterium phage FP_Epona Faecalibacterium virus Epona Eponavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49573 | 57.092 | Eponavirus | Unclassified | Myoviridae | Faecalibacterium |
| MG711461 | Faecalibacterium phage FP_Lagaffe | Faecalibacterium phage FP_Lagaffe Faecalibacterium virus Lagaffe Lagaffevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48211 | 55.906 | Lagaffevirus | Unclassified | Myoviridae | Faecalibacterium |
| MF327005 | Shigella phage Sf19 | Shigella phage Sf19 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90375 | 39.024 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MG708276 | Enterococcus phage PMBT2 | Enterococcus phage PMBT2 Enterococcus virus PMBT2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41489 | 34.691 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MG189906 | Stenotrophomonas phage vB_SmaS_DLP_5 | Stenotrophomonas phage vB_SmaS_DLP_5 Stenotrophomonas virus DLP5 Delepquintavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 96542 | 58.369 | Delepquintavirus | Unclassified | Siphoviridae | Stenotrophomonas |
| KU245542 | Pseudomonas phage SM1 | Pseudomonas phage SM1 Pseudomonas virus SM1 Samunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93191 | 55.24 | Samunavirus | Unclassified | Siphoviridae | Pseudomonas |
| CP017837 | Proteobacteria phage MWH-Nonnen-W8red 2 | Proteobacteria phage MWH-Nonnen-W8red 2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41775 | 30.533 | Unclassified | Unclassified | Myoviridae | Unspecified |
| KX130863 | Shigella phage SHSML-45 | Shigella phage SHSML-45 Shigella virus SHSML45 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108050 | 38.738 | Tequintavirus | Markadamsvirinae | Demerecviridae | Shigella |
| CP017838 | Proteobacteria phage MWH-Nonnen-W8red 1 | Proteobacteria phage MWH-Nonnen-W8red 1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43334 | 33.092 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MF276773 | Erwinia phage EtG | Erwinia phage EtG Erwinia virus EtG Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30413 | 54.138 | Eganvirus | Peduovirinae | Myoviridae | Erwinia |
| MF768470 | Pseudomonas phage SL1 | Pseudomonas phage SL1 Pseudomonas virus SL1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65847 | 55.585 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| MF768469 | Pseudomonas phage SL4 | Pseudomonas phage SL4 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44194 | 52.288 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| AP018487 | Mycobacterium phage HC | Mycobacterium phage HC unclassified Brujitavirus Brujitavirus Chebruvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46305 | 67.172 | Brujitavirus | Chebruvirinae | Siphoviridae | Mycobacterium |
| AP018486 | Mycobacterium phage PP | Mycobacterium phage PP unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51509 | 64.486 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018485 | Mycobacterium phage B1 | Mycobacterium phage B1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45961 | 64.059 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG766218 | Staphylococcus phage vB_SauP_phiAGO1.3 | Staphylococcus phage vB_SauP_phiAGO1.3 Staphylococcus virus phiAGO13 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17603 | 28.955 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MG592672 | Vibrio phage 3.058.O._10N.286.46.B8 | Vibrio phage 3.058.O._10N.286.46.B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101642 | 41.068 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592671 | Vibrio phage 2.275.O._10N.286.54.E11 | Vibrio phage 2.275.O._10N.286.54.E11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348911 | 38.093 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592670 | Vibrio phage 2.159.B._10N.261.46.F12 | Vibrio phage 2.159.B._10N.261.46.F12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592669 | Vibrio phage 2.159.A._10N.261.46.F12 | Vibrio phage 2.159.A._10N.261.46.F12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592668 | Vibrio phage 2.130.O._10N.222.46.C2 | Vibrio phage 2.130.O._10N.222.46.C2 unclassified Nahantvirus Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75797 | 43.11 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592667 | Vibrio phage 2.117.O._10N.261.45.E9 | Vibrio phage 2.117.O._10N.261.45.E9 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55795 | 49.47 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MG592666 | Vibrio phage 2.096.O._10N.286.48.B5 | Vibrio phage 2.096.O._10N.286.48.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44683 | 38.784 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592665 | Vibrio phage 2.095.B._10N.286.46.E10 | Vibrio phage 2.095.B._10N.286.46.E10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44649 | 42.359 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592664 | Vibrio phage 2.095.A._10N.286.46.E10 | Vibrio phage 2.095.A._10N.286.46.E10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44649 | 42.364 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592662 | Vibrio phage 2.058.O._10N.286.46.B8 | Vibrio phage 2.058.O._10N.286.46.B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101637 | 41.065 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592661 | Vibrio phage 2.044.O._10N.261.51.B8 | Vibrio phage 2.044.O._10N.261.51.B8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45020 | 44.816 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592660 | Vibrio phage 1.293.O._10N.261.52.E1 | Vibrio phage 1.293.O._10N.261.52.E1 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48275 | 44.141 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592659 | Vibrio phage 1.291.O._10N.286.55.F6 | Vibrio phage 1.291.O._10N.286.55.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43662 | 42.969 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592658 | Vibrio phage 1.289.A._10N.286.55.E8 | Vibrio phage 1.289.A._10N.286.55.E8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44529 | 43.392 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592657 | Vibrio phage 1.287.O._10N.286.55.C7 | Vibrio phage 1.287.O._10N.286.55.C7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45562 | 43.334 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592656 | Vibrio phage 1.286.O._10N.286.55.C4 | Vibrio phage 1.286.O._10N.286.55.C4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49131 | 42.993 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592655 | Vibrio phage 1.285.O._10N.286.55.C12 | Vibrio phage 1.285.O._10N.286.55.C12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59437 | 48.552 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592654 | Vibrio phage 1.284.A._10N.286.55.A5 | Vibrio phage 1.284.A._10N.286.55.A5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45648 | 43.454 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592653 | Vibrio phage 1.283.C._10N.286.55.A1 | Vibrio phage 1.283.C._10N.286.55.A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59530 | 48.619 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592652 | Vibrio phage 1.283.B._10N.286.55.A1 | Vibrio phage 1.283.B._10N.286.55.A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59530 | 48.621 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592651 | Vibrio phage 1.283.A._10N.286.55.A1 | Vibrio phage 1.283.A._10N.286.55.A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59530 | 48.621 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592650 | Vibrio phage 1.282.A._10N.286.54.F8 | Vibrio phage 1.282.A._10N.286.54.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59359 | 48.577 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592649 | Vibrio phage 1.281.O._10N.286.54.F7 | Vibrio phage 1.281.O._10N.286.54.F7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.591 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592648 | Vibrio phage 1.280.O._10N.286.54.F6 | Vibrio phage 1.280.O._10N.286.54.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.591 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592647 | Vibrio phage 1.278.O._10N.286.54.E8 | Vibrio phage 1.278.O._10N.286.54.E8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49282 | 42.669 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592646 | Vibrio phage 1.277.B._10N.286.54.E7 | Vibrio phage 1.277.B._10N.286.54.E7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59419 | 48.602 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592645 | Vibrio phage 1.277.A._10N.286.54.E7 | Vibrio phage 1.277.A._10N.286.54.E7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.588 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592644 | Vibrio phage 1.276.O._10N.286.54.E4 | Vibrio phage 1.276.O._10N.286.54.E4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44334 | 42.184 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592643 | Vibrio phage 1.275.O._10N.286.54.E11 | Vibrio phage 1.275.O._10N.286.54.E11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37915 | 44.304 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592642 | Vibrio phage 1.274.O._10N.286.54.E1 | Vibrio phage 1.274.O._10N.286.54.E1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59409 | 48.572 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592641 | Vibrio phage 1.273.O._10N.286.54.C7 | Vibrio phage 1.273.O._10N.286.54.C7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48145 | 42.86 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592640 | Vibrio phage 1.272.O._10N.286.54.C4 | Vibrio phage 1.272.O._10N.286.54.C4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.591 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592639 | Vibrio phage 1.271.B._10N.286.54.B4 | Vibrio phage 1.271.B._10N.286.54.B4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.589 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592638 | Vibrio phage 1.271.A._10N.286.54.B4 | Vibrio phage 1.271.A._10N.286.54.B4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.586 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592637 | Vibrio phage 1.270.B._10N.286.54.A8 | Vibrio phage 1.270.B._10N.286.54.A8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59294 | 48.59 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592636 | Vibrio phage 1.270.A._10N.286.54.A8 | Vibrio phage 1.270.A._10N.286.54.A8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59294 | 48.592 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592635 | Vibrio phage 1.269.O._10N.286.54.A6 | Vibrio phage 1.269.O._10N.286.54.A6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59738 | 48.652 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592634 | Vibrio phage 1.268.B._10N.286.54.A11 | Vibrio phage 1.268.B._10N.286.54.A11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.588 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592633 | Vibrio phage 1.268.A._10N.286.54.A11 | Vibrio phage 1.268.A._10N.286.54.A11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.588 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592632 | Vibrio phage 1.267.O._10N.286.54.A1 | Vibrio phage 1.267.O._10N.286.54.A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59414 | 48.579 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592631 | Vibrio phage 1.266.O._10N.286.52.F9 | Vibrio phage 1.266.O._10N.286.52.F9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34788 | 43.262 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592630 | Vibrio phage 1.265.O._10N.286.52.F6 | Vibrio phage 1.265.O._10N.286.52.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43630 | 42.18 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592629 | Vibrio phage 1.264.O._10N.286.51.F2 | Vibrio phage 1.264.O._10N.286.51.F2 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47739 | 42.77 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592628 | Vibrio phage 1.263.B._10N.286.51.B1 | Vibrio phage 1.263.B._10N.286.51.B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49640 | 43.042 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592627 | Vibrio phage 1.263.A._10N.286.51.B1 | Vibrio phage 1.263.A._10N.286.51.B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49640 | 43.038 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592626 | Vibrio phage 1.262.O._10N.286.51.A9 | Vibrio phage 1.262.O._10N.286.51.A9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47635 | 39.072 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592625 | Vibrio phage 1.261.O._10N.286.51.A7 | Vibrio phage 1.261.O._10N.286.51.A7 Vibrio virus 51A7 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71992 | 43.166 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592623 | Vibrio phage 1.257.O._10N.286.46.A4 | Vibrio phage 1.257.O._10N.286.46.A4 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32371 | 45.776 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592622 | Vibrio phage 1.256.O._10N.286.45.F8 | Vibrio phage 1.256.O._10N.286.45.F8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48207 | 42.967 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592621 | Vibrio phage 1.255.O._10N.286.45.F1 | Vibrio phage 1.255.O._10N.286.45.F1 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159885 | 44.335 | Unclassified | Unclassified | Ackermannviridae | Vibrio |
| MG592620 | Vibrio phage 1.254.O._10N.286.45.C8 | Vibrio phage 1.254.O._10N.286.45.C8 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32699 | 45.466 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592619 | Vibrio phage 1.253.O._10N.286.45.B12 | Vibrio phage 1.253.O._10N.286.45.B12 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47008 | 43.018 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592618 | Vibrio phage 1.251.O._10N.261.55.E5 | Vibrio phage 1.251.O._10N.261.55.E5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59649 | 48.566 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592617 | Vibrio phage 1.250.O._10N.261.55.E11 | Vibrio phage 1.250.O._10N.261.55.E11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59981 | 48.93 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592614 | Vibrio phage 1.248.O._10N.261.54.F1 | Vibrio phage 1.248.O._10N.261.54.F1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48301 | 42.881 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592613 | Vibrio phage 1.247.B._10N.261.54.E12 | Vibrio phage 1.247.B._10N.261.54.E12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43896 | 44.054 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592612 | Vibrio phage 1.247.A._10N.261.54.E12 | Vibrio phage 1.247.A._10N.261.54.E12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43896 | 44.056 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592611 | Vibrio phage 1.246.O._10N.261.54.E10 | Vibrio phage 1.246.O._10N.261.54.E10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47133 | 44.761 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592610 | Vibrio phage 1.245.O._10N.261.54.C7 | Vibrio phage 1.245.O._10N.261.54.C7 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71702 | 43.12 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MG592609 | Vibrio phage 1.244.A._10N.261.54.C3 | Vibrio phage 1.244.A._10N.261.54.C3 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159885 | 44.335 | Unclassified | Unclassified | Ackermannviridae | Vibrio |
| MG592608 | Vibrio phage 1.243.O._10N.261.54.B5 | Vibrio phage 1.243.O._10N.261.54.B5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48414 | 42.94 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592607 | Vibrio phage 1.242.O._10N.261.54.B2 | Vibrio phage 1.242.O._10N.261.54.B2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36910 | 42.753 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592606 | Vibrio phage 1.240.O._10N.261.52.F8 | Vibrio phage 1.240.O._10N.261.52.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36710 | 42.531 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592605 | Vibrio phage 1.239.O._10N.261.52.F6 | Vibrio phage 1.239.O._10N.261.52.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38618 | 43.021 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592604 | Vibrio phage 1.238.B._10N.261.52.F10 | Vibrio phage 1.238.B._10N.261.52.F10 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70467 | 43.189 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MG592603 | Vibrio phage 1.238.A._10N.261.52.F10 | Vibrio phage 1.238.A._10N.261.52.F10 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70494 | 43.188 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| MG592602 | Vibrio phage 1.237.B._10N.261.52.C5 | Vibrio phage 1.237.B._10N.261.52.C5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60160 | 48.763 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592601 | Vibrio phage 1.237.A._10N.261.52.C5 | Vibrio phage 1.237.A._10N.261.52.C5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60097 | 48.763 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592600 | Vibrio phage 1.236.O._10N.261.52.C4 | Vibrio phage 1.236.O._10N.261.52.C4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42646 | 42.325 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592599 | Vibrio phage 1.235.O._10N.261.52.B2 | Vibrio phage 1.235.O._10N.261.52.B2 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47017 | 43.025 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592598 | Vibrio phage 1.233.B._10N.261.51.E6 | Vibrio phage 1.233.B._10N.261.51.E6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36823 | 43.114 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592597 | Vibrio phage 1.233.A._10N.261.51.E6 | Vibrio phage 1.233.A._10N.261.51.E6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36823 | 43.114 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592596 | Vibrio phage 1.232.O._10N.261.51.E11 | Vibrio phage 1.232.O._10N.261.51.E11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56736 | 43.896 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592595 | Vibrio phage 1.231.O._10N.261.49.F8 | Vibrio phage 1.231.O._10N.261.49.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44527 | 42.143 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592594 | Vibrio phage 1.228.O._10N.261.49.C1 | Vibrio phage 1.228.O._10N.261.49.C1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36667 | 42.897 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592593 | Vibrio phage 1.226.O._10N.261.48.E5 | Vibrio phage 1.226.O._10N.261.48.E5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35655 | 42.979 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592592 | Vibrio phage 1.225.O._10N.261.48.B7 | Vibrio phage 1.225.O._10N.261.48.B7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52615 | 44.242 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592591 | Vibrio phage 1.224.A._10N.261.48.B1 | Vibrio phage 1.224.A._10N.261.48.B1 Vibrio virus 48B1 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71915 | 43.063 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592590 | Vibrio phage 1.223.O._10N.261.48.A9 | Vibrio phage 1.223.O._10N.261.48.A9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49535 | 44.443 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592589 | Vibrio phage 1.219.O._10N.261.45.E2 | Vibrio phage 1.219.O._10N.261.45.E2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592588 | Vibrio phage 1.217.O._10N.261.45.A1 | Vibrio phage 1.217.O._10N.261.45.A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592587 | Vibrio phage 1.216.O._10N.222.55.C12 | Vibrio phage 1.216.O._10N.222.55.C12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41359 | 42.603 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592586 | Vibrio phage 1.215.B._10N.222.54.F7 | Vibrio phage 1.215.B._10N.222.54.F7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80834 | 45.548 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592585 | Vibrio phage 1.215.A._10N.222.54.F7 | Vibrio phage 1.215.A._10N.222.54.F7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80834 | 45.548 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592584 | Vibrio phage 1.214.O._10N.222.54.F11 | Vibrio phage 1.214.O._10N.222.54.F11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44205 | 43.629 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592583 | Vibrio phage 1.213.O._10N.222.54.F10 | Vibrio phage 1.213.O._10N.222.54.F10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42443 | 42.568 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592582 | Vibrio phage 1.211.B._10N.222.52.F11 | Vibrio phage 1.211.B._10N.222.52.F11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37169 | 43.749 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592581 | Vibrio phage 1.211.A._10N.222.52.F11 | Vibrio phage 1.211.A._10N.222.52.F11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37169 | 43.749 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592580 | Vibrio phage 1.210.O._10N.222.52.C2 | Vibrio phage 1.210.O._10N.222.52.C2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48224 | 42.039 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592579 | Vibrio phage 1.209.O._10N.222.52.B2 | Vibrio phage 1.209.O._10N.222.52.B2 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43712 | 48.001 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592578 | Vibrio phage 1.208.B._10N.222.52.A7 | Vibrio phage 1.208.B._10N.222.52.A7 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48927 | 44.464 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592577 | Vibrio phage 1.207.B._10N.222.51.C2 | Vibrio phage 1.207.B._10N.222.51.C2 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55793 | 49.476 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MG592576 | Vibrio phage 1.206.O._10N.222.51.B10 | Vibrio phage 1.206.O._10N.222.51.B10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37551 | 42.926 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592575 | Vibrio phage 1.205.O._10N.222.51.A7 | Vibrio phage 1.205.O._10N.222.51.A7 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57861 | 44.925 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592574 | Vibrio phage 1.204.O._10N.222.46.F12 | Vibrio phage 1.204.O._10N.222.46.F12 Vibrio virus Cyclit Cyclitvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44168 | 43.516 | Cyclitvirus | Unclassified | Autographiviridae | Vibrio |
| MG592573 | Vibrio phage 1.202.O._10N.222.45.E8 | Vibrio phage 1.202.O._10N.222.45.E8 Vibrio virus Canoe Canoevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32014 | 44.559 | Canoevirus | Peduovirinae | Myoviridae | Vibrio |
| MG592572 | Vibrio phage 1.201.B._10N.286.55.F1 | Vibrio phage 1.201.B._10N.286.55.F1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50506 | 44.292 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592571 | Vibrio phage 1.200.O._10N.286.55.E1 | Vibrio phage 1.200.O._10N.286.55.E1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.593 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592570 | Vibrio phage 1.199.B._10N.286.55.C10 | Vibrio phage 1.199.B._10N.286.55.C10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48312 | 43.043 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592569 | Vibrio phage 1.199.A._10N.286.55.C10 | Vibrio phage 1.199.A._10N.286.55.C10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48312 | 43.045 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592568 | Vibrio phage 1.198.B._10N.286.54.F4 | Vibrio phage 1.198.B._10N.286.54.F4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44343 | 41.858 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592567 | Vibrio phage 1.198.A._10N.286.54.F4 | Vibrio phage 1.198.A._10N.286.54.F4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44472 | 41.905 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592566 | Vibrio phage 1.197.A._10N.286.54.F2 | Vibrio phage 1.197.A._10N.286.54.F2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44587 | 43.618 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592565 | Vibrio phage 1.196.O._10N.286.54.E12 | Vibrio phage 1.196.O._10N.286.54.E12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59414 | 48.579 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592564 | Vibrio phage 1.195.O._10N.286.54.C8 | Vibrio phage 1.195.O._10N.286.54.C8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59738 | 48.652 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592563 | Vibrio phage 1.194.O._10N.286.54.B1 | Vibrio phage 1.194.O._10N.286.54.B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59294 | 48.58 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592562 | Vibrio phage 1.193.O._10N.286.52.C6 | Vibrio phage 1.193.O._10N.286.52.C6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132560 | 38.085 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592561 | Vibrio phage 1.191.O._10N.286.52.B4 | Vibrio phage 1.191.O._10N.286.52.B4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48354 | 44.197 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592560 | Vibrio phage 1.190.O._10N.286.51.F12 | Vibrio phage 1.190.O._10N.286.51.F12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21785 | 45.935 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592559 | Vibrio phage 1.189.O._10N.286.51.B5 | Vibrio phage 1.189.O._10N.286.51.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36855 | 42.797 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592558 | Vibrio phage 1.189.C._10N.286.51.B5 | Vibrio phage 1.189.C._10N.286.51.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36855 | 42.8 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592557 | Vibrio phage 1.189.B._10N.286.51.B5 | Vibrio phage 1.189.B._10N.286.51.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36855 | 42.797 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592556 | Vibrio phage 1.188.C._10N.286.51.A6 | Vibrio phage 1.188.C._10N.286.51.A6 unclassified Mukerjeevirus Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72305 | 42.983 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592555 | Vibrio phage 1.188.B._10N.286.51.A6 | Vibrio phage 1.188.B._10N.286.51.A6 unclassified Mukerjeevirus Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72305 | 42.983 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592554 | Vibrio phage 1.188.A._10N.286.51.A6 | Vibrio phage 1.188.A._10N.286.51.A6 Vibrio virus 51A6 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72305 | 42.983 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592553 | Vibrio phage 1.187.O._10N.286.49.F1 | Vibrio phage 1.187.O._10N.286.49.F1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133254 | 37.927 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592552 | Vibrio phage 1.186.O._10N.286.49.E3 | Vibrio phage 1.186.O._10N.286.49.E3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48643 | 42.783 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592551 | Vibrio phage 1.185.O._10N.286.49.C2 | Vibrio phage 1.185.O._10N.286.49.C2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43397 | 46.045 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592550 | Vibrio phage 1.184.A._10N.286.49.A5 | Vibrio phage 1.184.A._10N.286.49.A5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33272 | 43.794 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592549 | Vibrio phage 1.183.O._10N.286.48.B7 | Vibrio phage 1.183.O._10N.286.48.B7 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37411 | 44.016 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592548 | Vibrio phage 1.182.O._10N.286.46.E1 | Vibrio phage 1.182.O._10N.286.46.E1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36910 | 42.728 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592547 | Vibrio phage 1.181.O._10N.286.46.C9 | Vibrio phage 1.181.O._10N.286.46.C9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50228 | 43.378 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592546 | Vibrio phage 1.179.O._10N.286.45.F12 | Vibrio phage 1.179.O._10N.286.45.F12 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31498 | 45.682 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592545 | Vibrio phage 1.178.O._10N.286.45.E12 | Vibrio phage 1.178.O._10N.286.45.E12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39934 | 41.849 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592544 | Vibrio phage 1.177.O._10N.286.45.E10 | Vibrio phage 1.177.O._10N.286.45.E10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45439 | 42.116 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592543 | Vibrio phage 1.176.O._10N.261.55.F5 | Vibrio phage 1.176.O._10N.261.55.F5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36463 | 42.934 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592542 | Vibrio phage 1.175.O._10N.261.55.B3 | Vibrio phage 1.175.O._10N.261.55.B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49066 | 44.514 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592541 | Vibrio phage 1.174.O._10N.261.55.A8 | Vibrio phage 1.174.O._10N.261.55.A8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47063 | 43.95 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592540 | Vibrio phage 1.173.O._10N.261.55.A11 | Vibrio phage 1.173.O._10N.261.55.A11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48063 | 42.758 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592539 | Vibrio phage 1.172.O._10N.261.52.F5 | Vibrio phage 1.172.O._10N.261.52.F5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36314 | 42.683 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592538 | Vibrio phage 1.171.O._10N.261.52.F12 | Vibrio phage 1.171.O._10N.261.52.F12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60216 | 48.725 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592537 | Vibrio phage 1.170.O._10N.261.52.C3 | Vibrio phage 1.170.O._10N.261.52.C3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133692 | 38.696 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592536 | Vibrio phage 1.169.O._10N.261.52.B1 | Vibrio phage 1.169.O._10N.261.52.B1 Vibrio virus 52B1 Mukerjeevirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72290 | 42.941 | Mukerjeevirus | Unclassified | Schitoviridae | Vibrio |
| MG592535 | Vibrio phage 1.168.O._10N.261.52.A10 | Vibrio phage 1.168.O._10N.261.52.A10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37551 | 42.555 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592534 | Vibrio phage 1.167.O._10N.261.51.F2 | Vibrio phage 1.167.O._10N.261.51.F2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35811 | 43.048 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592533 | Vibrio phage 1.166.O._10N.261.51.C7 | Vibrio phage 1.166.O._10N.261.51.C7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21800 | 45.968 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592532 | Vibrio phage 1.165.O._10N.261.51.B7 | Vibrio phage 1.165.O._10N.261.51.B7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37046 | 42.99 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592531 | Vibrio phage 1.164.O._10N.261.51.A7 | Vibrio phage 1.164.O._10N.261.51.A7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48235 | 44.028 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592530 | Vibrio phage 1.162.O._10N.261.48.E3 | Vibrio phage 1.162.O._10N.261.48.E3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37360 | 42.81 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592529 | Vibrio phage 1.161.O._10N.261.48.C5 | Vibrio phage 1.161.O._10N.261.48.C5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140668 | 37.68 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592528 | Vibrio phage 1.160.O._10N.261.48.B11 | Vibrio phage 1.160.O._10N.261.48.B11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60734 | 48.655 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592527 | Vibrio phage 1.159.O._10N.261.46.F12 | Vibrio phage 1.159.O._10N.261.46.F12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.947 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592526 | Vibrio phage 1.158.O._10N.261.45.E12 | Vibrio phage 1.158.O._10N.261.45.E12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46507 | 44.619 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592525 | Vibrio phage 1.157.O._10N.261.45.B7 | Vibrio phage 1.157.O._10N.261.45.B7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35855 | 42.92 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592524 | Vibrio phage 1.156.O._10N.261.45.A6 | Vibrio phage 1.156.O._10N.261.45.A6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592523 | Vibrio phage 1.155.O._10N.222.55.B3 | Vibrio phage 1.155.O._10N.222.55.B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29029 | 44.001 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592522 | Vibrio phage 1.154.O._10N.222.52.B12 | Vibrio phage 1.154.O._10N.222.52.B12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37136 | 42.207 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592521 | Vibrio phage 1.152.O._10N.222.46.E1 | Vibrio phage 1.152.O._10N.222.46.E1 unclassified Nahantvirus Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75798 | 43.104 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592520 | Vibrio phage 1.151.O._10N.222.46.B1 | Vibrio phage 1.151.O._10N.222.46.B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44307 | 44.374 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592519 | Vibrio phage 1.150.O._10N.222.46.A6 | Vibrio phage 1.150.O._10N.222.46.A6 unclassified Nahantvirus Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75796 | 43.106 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592518 | Vibrio phage 1.149.O._10N.286.55.A12 | Vibrio phage 1.149.O._10N.286.55.A12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48481 | 42.747 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592517 | Vibrio phage 1.148.O._10N.286.54.A10 | Vibrio phage 1.148.O._10N.286.54.A10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36789 | 42.447 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592516 | Vibrio phage 1.147.O._10N.286.49.E9 | Vibrio phage 1.147.O._10N.286.49.E9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37360 | 42.813 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592515 | Vibrio phage 1.144.O._10N.286.45.B3 | Vibrio phage 1.144.O._10N.286.45.B3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44418 | 42.634 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592514 | Vibrio phage 1.143.O._10N.261.55.C8 | Vibrio phage 1.143.O._10N.261.55.C8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44527 | 42.147 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592513 | Vibrio phage 1.142.O._10N.261.49.E11 | Vibrio phage 1.142.O._10N.261.49.E11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592511 | Vibrio phage 1.139.B._10N.261.48.C6 | Vibrio phage 1.139.B._10N.261.48.C6 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44094 | 48.022 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592510 | Vibrio phage 1.139.A._10N.261.48.C6 | Vibrio phage 1.139.A._10N.261.48.C6 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43893 | 47.955 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592509 | Vibrio phage 1.138.O._10N.261.48.A1 | Vibrio phage 1.138.O._10N.261.48.A1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32510 | 45.798 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592508 | Vibrio phage 1.137.O._10N.261.46.B5 | Vibrio phage 1.137.O._10N.261.46.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37958 | 42.41 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592507 | Vibrio phage 1.136.O._10N.261.45.E11 | Vibrio phage 1.136.O._10N.261.45.E11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592506 | Vibrio phage 1.135.O._10N.222.54.B6 | Vibrio phage 1.135.O._10N.222.54.B6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41918 | 42.545 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592505 | Vibrio phage 1.134.O._10N.222.52.B8 | Vibrio phage 1.134.O._10N.222.52.B8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36379 | 42.956 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592504 | Vibrio phage 1.133.O._10N.222.51.E4 | Vibrio phage 1.133.O._10N.222.51.E4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37087 | 42.899 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592503 | Vibrio phage 1.132.O._10N.222.49.F8 | Vibrio phage 1.132.O._10N.222.49.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36926 | 43.205 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592502 | Vibrio phage 1.131.O._10N.222.49.A8 | Vibrio phage 1.131.O._10N.222.49.A8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37933 | 42.839 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592501 | Vibrio phage 1.127.O._10N.286.52.E12 | Vibrio phage 1.127.O._10N.286.52.E12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44915 | 42.298 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592499 | Vibrio phage 1.124.O._10N.286.49.B1 | Vibrio phage 1.124.O._10N.286.49.B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47604 | 43.522 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592498 | Vibrio phage 1.123.O._10N.286.48.F3 | Vibrio phage 1.123.O._10N.286.48.F3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46071 | 48.297 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592497 | Vibrio phage 1.122.B._10N.286.46.F8 | Vibrio phage 1.122.B._10N.286.46.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44523 | 43.629 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592496 | Vibrio phage 1.122.A._10N.286.46.F8 | Vibrio phage 1.122.A._10N.286.46.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44523 | 43.625 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592495 | Vibrio phage 1.121.O._10N.286.46.C4 | Vibrio phage 1.121.O._10N.286.46.C4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145590 | 39.686 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592494 | Vibrio phage 1.119.O._10N.261.51.A9 | Vibrio phage 1.119.O._10N.261.51.A9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44527 | 42.15 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592493 | Vibrio phage 1.118.B._10N.261.49.F6 | Vibrio phage 1.118.B._10N.261.49.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60458 | 48.796 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592492 | Vibrio phage 1.118.A._10N.261.49.F6 | Vibrio phage 1.118.A._10N.261.49.F6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60458 | 48.796 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592491 | Vibrio phage 1.117.O._10N.261.45.E9 | Vibrio phage 1.117.O._10N.261.45.E9 Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55794 | 49.471 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| MG592490 | Vibrio phage 1.116.O._10N.222.52.C10 | Vibrio phage 1.116.O._10N.222.52.C10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36314 | 42.683 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592489 | Vibrio phage 1.115.B._10N.222.49.B11 | Vibrio phage 1.115.B._10N.222.49.B11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37416 | 43.222 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592488 | Vibrio phage 1.115.A._10N.222.49.B11 | Vibrio phage 1.115.A._10N.222.49.B11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37416 | 43.225 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592487 | Vibrio phage 1.113.A._10N.286.51.E7 | Vibrio phage 1.113.A._10N.286.51.E7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44150 | 43.343 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592486 | Vibrio phage 1.112.O._10N.286.46.B11 | Vibrio phage 1.112.O._10N.286.46.B11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48149 | 43.083 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592485 | Vibrio phage 1.111.B._10N.286.45.E6 | Vibrio phage 1.111.B._10N.286.45.E6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40209 | 43.575 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592484 | Vibrio phage 1.111.A._10N.286.45.E6 | Vibrio phage 1.111.A._10N.286.45.E6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40209 | 43.575 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592483 | Vibrio phage 1.110.O._10N.261.52.C1 | Vibrio phage 1.110.O._10N.261.52.C1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37556 | 42.923 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592482 | Vibrio phage 1.108.O._10N.222.51.A4 | Vibrio phage 1.108.O._10N.222.51.A4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36584 | 42.792 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592478 | Vibrio phage 1.106.O._10N.286.51.F7 | Vibrio phage 1.106.O._10N.286.51.F7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47070 | 42.8 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592477 | Vibrio phage 1.105.O._10N.286.49.B4 | Vibrio phage 1.105.O._10N.286.49.B4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49123 | 42.809 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592476 | Vibrio phage 1.104.O._10N.286.49.A12 | Vibrio phage 1.104.O._10N.286.49.A12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60990 | 48.751 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592475 | Vibrio phage 1.103.O._10N.261.52.F2 | Vibrio phage 1.103.O._10N.261.52.F2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61748 | 48.635 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592473 | Vibrio phage 1.101.O._10N.261.45.C6 | Vibrio phage 1.101.O._10N.261.45.C6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130250 | 37.359 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592472 | Vibrio phage 1.100.O._10N.261.45.C3 | Vibrio phage 1.100.O._10N.261.45.C3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51268 | 44.406 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592471 | Vibrio phage 1.098.O._10N.286.51.B9 | Vibrio phage 1.098.O._10N.286.51.B9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59851 | 48.557 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592470 | Vibrio phage 1.097.O._10N.286.49.B3 | Vibrio phage 1.097.O._10N.286.49.B3 Vibrio virus 49B3 Dorisvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76918 | 42.596 | Dorisvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592468 | Vibrio phage 1.094.O._10N.286.55.E12 | Vibrio phage 1.094.O._10N.286.55.E12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 44.285 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592467 | Vibrio phage 1.093.O._10N.286.55.E10 | Vibrio phage 1.093.O._10N.286.55.E10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 44.287 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592466 | Vibrio phage 1.091.O._10N.286.52.B12 | Vibrio phage 1.091.O._10N.286.52.B12 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43134 | 42.767 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592465 | Vibrio phage 1.090.B._10N.286.48.F1 | Vibrio phage 1.090.B._10N.286.48.F1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41868 | 42.519 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592464 | Vibrio phage 1.089.O._10N.261.51.F9 | Vibrio phage 1.089.O._10N.261.51.F9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59851 | 48.561 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592463 | Vibrio phage 1.088.O._10N.261.46.A1 | Vibrio phage 1.088.O._10N.261.46.A1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60385 | 48.782 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592462 | Vibrio phage 1.087.A._10N.261.45.F9 | Vibrio phage 1.087.A._10N.261.45.F9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45674 | 44.119 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592461 | Vibrio phage 1.086.O._10N.222.51.F8 | Vibrio phage 1.086.O._10N.222.51.F8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50835 | 44.192 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592460 | Vibrio phage 1.085.O._10N.222.51.E3 | Vibrio phage 1.085.O._10N.222.51.E3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37820 | 42.739 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592459 | Vibrio phage 1.084.O._10N.261.49.F5 | Vibrio phage 1.084.O._10N.261.49.F5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141906 | 37.103 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592458 | Vibrio phage 1.083.O._10N.286.52.B9 | Vibrio phage 1.083.O._10N.286.52.B9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45120 | 38.708 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592457 | Vibrio phage 1.082.O._10N.261.49.E4 | Vibrio phage 1.082.O._10N.261.49.E4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35810 | 42.996 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592456 | Vibrio phage 1.081.O._10N.286.52.C2 | Vibrio phage 1.081.O._10N.286.52.C2 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239318 | 42.413 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| MG592454 | Vibrio phage 1.079.O._10N.286.45.E9 | Vibrio phage 1.079.O._10N.286.45.E9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42375 | 42.782 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592453 | Vibrio phage 1.077.O._10N.261.45.A10 | Vibrio phage 1.077.O._10N.261.45.A10 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44047 | 49.379 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592452 | Vibrio phage 1.076.O._10N.286.51.B7 | Vibrio phage 1.076.O._10N.286.51.B7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47914 | 42.843 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592451 | Vibrio phage 1.075.O._10N.286.55.B10 | Vibrio phage 1.075.O._10N.286.55.B10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 44.289 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592450 | Vibrio phage 1.074.O._10N.222.49.B7 | Vibrio phage 1.074.O._10N.222.49.B7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36922 | 42.609 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592449 | Vibrio phage 1.072.O._10N.286.48.A12 | Vibrio phage 1.072.O._10N.286.48.A12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43788 | 44.773 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592448 | Vibrio phage 1.071.A._10N.286.46.A12 | Vibrio phage 1.071.A._10N.286.46.A12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48114 | 43.31 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592447 | Vibrio phage 1.070.O._10N.261.45.B2 | Vibrio phage 1.070.O._10N.261.45.B2 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44087 | 49.396 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592445 | Vibrio phage 1.068.O._10N.261.51.F8 | Vibrio phage 1.068.O._10N.261.51.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47038 | 44.76 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592444 | Vibrio phage 1.067.O._10N.261.52.C9 | Vibrio phage 1.067.O._10N.261.52.C9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40955 | 42.542 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592443 | Vibrio phage 1.066.O._10N.286.46.E8 | Vibrio phage 1.066.O._10N.286.46.E8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47053 | 44.75 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592442 | Vibrio phage 1.064.O._10N.261.52.E2 | Vibrio phage 1.064.O._10N.261.52.E2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36584 | 42.789 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592441 | Vibrio phage 1.063.O._10N.261.45.C7 | Vibrio phage 1.063.O._10N.261.45.C7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128641 | 37.336 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592439 | Vibrio phage 1.061.O._10N.286.55.C2 | Vibrio phage 1.061.O._10N.286.55.C2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41667 | 43.075 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592438 | Vibrio phage 1.060.A._10N.261.48.B5 | Vibrio phage 1.060.A._10N.261.48.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41105 | 43.255 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592436 | Vibrio phage 1.056.O._10N.261.48.C11 | Vibrio phage 1.056.O._10N.261.48.C11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47054 | 44.757 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592435 | Vibrio phage 1.055.O._10N.286.55.E9 | Vibrio phage 1.055.O._10N.286.55.E9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47199 | 40.929 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592434 | Vibrio phage 1.054.O._10N.261.52.A1 | Vibrio phage 1.054.O._10N.261.52.A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41774 | 42.318 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592433 | Vibrio phage 1.052.A._10N.286.46.C3 | Vibrio phage 1.052.A._10N.286.46.C3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42889 | 46.772 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592432 | Vibrio phage 1.050.O._10N.286.48.A6 | Vibrio phage 1.050.O._10N.286.48.A6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45285 | 41.641 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592431 | Vibrio phage 1.049.O._10N.286.54.B5 | Vibrio phage 1.049.O._10N.286.54.B5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45021 | 41.598 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592429 | Vibrio phage 1.047.O._10N.286.55.F2 | Vibrio phage 1.047.O._10N.286.55.F2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46106 | 43.3 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592428 | Vibrio phage 1.046.O._10N.286.52.E3 | Vibrio phage 1.046.O._10N.286.52.E3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62503 | 48.322 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592425 | Vibrio phage 1.042.O._10N.286.45.B8 | Vibrio phage 1.042.O._10N.286.45.B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49730 | 43.175 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592423 | Vibrio phage 1.039.O._10N.286.55.A2 | Vibrio phage 1.039.O._10N.286.55.A2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40972 | 42.524 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592422 | Vibrio phage 1.038.O._10N.286.51.C2 | Vibrio phage 1.038.O._10N.286.51.C2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49567 | 43.168 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592421 | Vibrio phage 1.037.O._10N.261.52.F7 | Vibrio phage 1.037.O._10N.261.52.F7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43111 | 44.968 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592420 | Vibrio phage 1.036.O._10N.286.45.C3 | Vibrio phage 1.036.O._10N.286.45.C3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40172 | 42.798 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592419 | Vibrio phage 1.034.X._10N.261.46.B7 | Vibrio phage 1.034.X._10N.261.46.B7 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44383 | 48.021 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592418 | Vibrio phage 1.034.O._10N.261.46.B7 | Vibrio phage 1.034.O._10N.261.46.B7 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44398 | 48.022 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592417 | Vibrio phage 1.033.O._10N.222.49.B8 | Vibrio phage 1.033.O._10N.222.49.B8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61206 | 48.353 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592416 | Vibrio phage 1.032.O._10N.261.54.F5 | Vibrio phage 1.032.O._10N.261.54.F5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60399 | 48.445 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592415 | Vibrio phage 1.031.O._10N.261.46.F8 | Vibrio phage 1.031.O._10N.261.46.F8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152942 | 43.159 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592414 | Vibrio phage 1.030.O._10N.222.55.F9 | Vibrio phage 1.030.O._10N.222.55.F9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45827 | 43.893 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592413 | Vibrio phage 1.029.O._10N.261.55.A7 | Vibrio phage 1.029.O._10N.261.55.A7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46513 | 42.554 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592412 | Vibrio phage 1.028.O._10N.286.45.B6 | Vibrio phage 1.028.O._10N.286.45.B6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31617 | 45.944 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592411 | Vibrio phage 1.027.O._10N.286.54.B8 | Vibrio phage 1.027.O._10N.286.54.B8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59297 | 48.593 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592410 | Vibrio phage 1.026.O._10N.222.49.C7 | Vibrio phage 1.026.O._10N.222.49.C7 Vibrio virus 49C7 Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75880 | 43.102 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592409 | Vibrio phage 1.025.O._10N.222.46.B6 | Vibrio phage 1.025.O._10N.222.46.B6 unclassified Nahantvirus Nahantvirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75797 | 43.105 | Nahantvirus | Pontosvirinae | Schitoviridae | Vibrio |
| MG592408 | Vibrio phage 1.024.O._10N.261.45.F8 | Vibrio phage 1.024.O._10N.261.45.F8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58892 | 49.061 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592407 | Vibrio phage 1.023.O._10N.222.51.B4 | Vibrio phage 1.023.O._10N.222.51.B4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59872 | 48.625 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592406 | Vibrio phage 1.022.O._10N.286.45.A10 | Vibrio phage 1.022.O._10N.286.45.A10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62440 | 48.555 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592405 | Vibrio phage 1.021.C._10N.222.51.F9 | Vibrio phage 1.021.C._10N.222.51.F9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43743 | 46.515 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592404 | Vibrio phage 1.021.B._10N.222.51.F9 | Vibrio phage 1.021.B._10N.222.51.F9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43743 | 46.517 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592403 | Vibrio phage 1.021.A._10N.222.51.F9 | Vibrio phage 1.021.A._10N.222.51.F9 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43743 | 46.517 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG592401 | Vibrio phage 1.017.O._10N.286.55.C11 | Vibrio phage 1.017.O._10N.286.55.C11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51290 | 44.287 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592400 | Vibrio phage 1.016.O._10N.286.46.A11 | Vibrio phage 1.016.O._10N.286.46.A11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48078 | 43.288 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592399 | Vibrio phage 1.015.O._10N.222.51.E5 | Vibrio phage 1.015.O._10N.222.51.E5 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42586 | 47.612 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| MG592398 | Vibrio phage 1.013.O._10N.286.54.F9 | Vibrio phage 1.013.O._10N.286.54.F9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44457 | 43.863 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592397 | Vibrio phage 1.012.O._10N.261.48.C12 | Vibrio phage 1.012.O._10N.261.48.C12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59979 | 48.607 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592395 | Vibrio phage 1.009.O._10N.261.51.C9 | Vibrio phage 1.009.O._10N.261.51.C9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44443 | 57.692 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592393 | Vibrio phage 1.007.O._10N.261.55.F9 | Vibrio phage 1.007.O._10N.261.55.F9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49244 | 42.846 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592392 | Vibrio phage 1.005.O._10N.286.48.F2 | Vibrio phage 1.005.O._10N.286.48.F2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50301 | 43.961 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MG592391 | Vibrio phage 1.004.O._10N.261.54.A2 | Vibrio phage 1.004.O._10N.261.54.A2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42511 | 44.984 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MG592390 | Vibrio phage 1.003.O._10N.286.48.A2 | Vibrio phage 1.003.O._10N.286.48.A2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41891 | 42.842 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG159787 | Gluconobacter phage GC1 | Gluconobacter phage GC1 Gluconobacter virus GC1 Gammatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 16523 | 50.475 | Gammatectivirus | Unclassified | Tectiviridae | Gluconobacter |
| MF422185 | Bacillus phage BSP10 | Bacillus phage BSP10 unclassified Nitunavirus Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153767 | 42.101 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF580962 | Salinibacter virus M31CR41-3 | Salinibacter virus M31CR41-3 unclassified Kairosalinivirus Kairosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50128 | 59.743 | Kairosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580960 | Salinibacter virus M31CC-1 | Salinibacter virus M31CC-1 unclassified Kryptosalinivirus Kryptosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53808 | 52.994 | Kryptosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MF580958 | Salinibacter virus M8CR30-4 | Salinibacter virus M8CR30-4 unclassified Holosalinivirus Holosalinivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35434 | 64.562 | Holosalinivirus | Unclassified | Siphoviridae | Salinibacter |
| MG640035 | Vibrio phage Athena1 | Vibrio phage Athena1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39826 | 42.389 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MF407276 | Bacillus phage vB_BveP-Goe6 | Bacillus phage vB_BveP-Goe6 Bacillus virus Goe6 Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19105 | 39.78 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| AP018480 | Mycobacterium virus D29 | Mycobacterium virus D29 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49136 | 63.532 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018479 | Mycobacterium phage GS4E | Mycobacterium phage GS4E unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49823 | 63.378 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018478 | Mycobacterium phage A6 | Mycobacterium phage A6 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49728 | 63.761 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018477 | Mycobacterium phage BK1 | Mycobacterium phage BK1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49728 | 63.761 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AP018470 | Mycobacterium phage Y2 | Mycobacterium phage Y2 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57990 | 67.882 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| MF172979 | Erysipelothrix phage phi1605 | Erysipelothrix phage phi1605 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90000 | 38.038 | Unclassified | Unclassified | Siphoviridae | Erysipelothrix |
| KY084243 | Pectobacterium phage PP74 | Pectobacterium phage PP74 Pectobacterium virus PP74 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39790 | 48.635 | Berlinvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| LC208019 | Aeropyrum globular virus 1 | Aeropyrum globular virus 1 unclassified archaeal viruses Viruses | 18222 | 55.801 | Unclassified | Unclassified | Unclassified | Aeropyrum |
| MG708275 | Streptococcus phage vB_SthS_VA460 | Streptococcus phage vB_SthS_VA460 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41169 | 38.755 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MG708274 | Streptococcus phage vB_SthS_VA214 | Streptococcus phage vB_SthS_VA214 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 38.23 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| MG708273 | Streptococcus phage vB_SthS_VA698 | Streptococcus phage vB_SthS_VA698 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33290 | 38.402 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| CP025713 | Escherichia phage YDC107_2 | Escherichia phage YDC107_2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18617 | 47.274 | Unclassified | Unclassified | Podoviridae | Escherichia |
| CP025712 | Escherichia phage YDC107_1 | Escherichia phage YDC107_1 unclassified Ravinvirus Ravinvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46701 | 49.888 | Ravinvirus | Unclassified | Siphoviridae | Escherichia |
| KY435490 | Escherichia virus K1E | Escherichia virus K1E Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44246 | 44.903 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| AP018463 | Mycobacterium phage PR | Mycobacterium phage PR Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61267 | 67.593 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| AP018462 | Mycobacterium phage D12 | Mycobacterium phage D12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61290 | 67.605 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| MG646672 | Salmonella phage BPS17L1 | Salmonella phage BPS17L1 Salmonella virus BPS17L1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84916 | 38.861 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG646671 | Salmonella phage BPS17S6 | Salmonella phage BPS17S6 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87628 | 38.798 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG646670 | Salmonella phage BPS15S6 | Salmonella phage BPS15S6 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87609 | 38.802 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG646669 | Salmonella phage BPS17W1 | Salmonella phage BPS17W1 Salmonella virus BPS17W1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87609 | 38.802 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| MG646668 | Salmonella phage BPS11T2 | Salmonella phage BPS11T2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43797 | 49.718 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MG696115 | Plesiomonas phage phiP4-7 | Plesiomonas phage phiP4-7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39825 | 48.665 | Unclassified | Unclassified | Siphoviridae | Plesiomonas |
| MG696114 | Proteus phage phiP4-3 | Proteus phage phiP4-3 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167849 | 31.507 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| MG673519 | Escherichia phage LAMP | Escherichia phage LAMP unclassified Kuravirus Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68521 | 42.235 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| MG564297 | Staphylococcus phage UPMK_2 | Staphylococcus phage UPMK_2 Staphylococcus virus UPMK2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40955 | 35.393 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KX011169 | Bacillus phage SalinJah | Bacillus phage SalinJah unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161140 | 38.72 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF974178 | Pseudomonas phage YS35 | Pseudomonas phage YS35 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93296 | 49.359 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| MG271909 | Synechococcus phage S-LBS1 | Synechococcus phage S-LBS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34641 | 60.218 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| MG065692 | Campylobacter phage C13 | Campylobacter phage C13 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99911 | 38.278 | Tequintavirus | Markadamsvirinae | Demerecviridae | Campylobacter |
| MG065691 | Campylobacter phage A11a | Campylobacter phage A11a unclassified bacterial viruses Viruses | 174114 | 50.187 | Unclassified | Unclassified | Unclassified | Campylobacter |
| MG065690 | Campylobacter phage A135 | Campylobacter phage A135 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143281 | 43.561 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065689 | Campylobacter phage A110a | Campylobacter phage A110a unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146366 | 37.408 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065688 | Campylobacter phage A110b | Campylobacter phage A110b unclassified Braunvirinae Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44238 | 44.123 | Unclassified | Braunvirinae | Drexlerviridae | Campylobacter |
| MG065687 | Campylobacter phage A136 | Campylobacter phage A136 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48078 | 43.981 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065685 | Campylobacter phage A112a | Campylobacter phage A112a unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25343 | 43.539 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065684 | Campylobacter phage A138 | Campylobacter phage A138 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35310 | 42.458 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065682 | Campylobacter phage A112b | Campylobacter phage A112b unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134946 | 43.527 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065681 | Campylobacter phage A139 | Campylobacter phage A139 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49946 | 43.771 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065680 | Campylobacter phage A113 | Campylobacter phage A113 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49946 | 43.771 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065679 | Campylobacter phage A13a | Campylobacter phage A13a unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47249 | 43.929 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065677 | Campylobacter phage A13b | Campylobacter phage A13b unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140869 | 43.506 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065676 | Campylobacter phage A116 | Campylobacter phage A116 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59527 | 43.699 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065675 | Campylobacter phage A118 | Campylobacter phage A118 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50132 | 43.796 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065674 | Campylobacter phage A140 | Campylobacter phage A140 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140874 | 43.506 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065673 | Campylobacter phage A119 | Campylobacter phage A119 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28832 | 44.246 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065672 | Campylobacter phage A141 | Campylobacter phage A141 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140869 | 43.506 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065671 | Campylobacter phage A120 | Campylobacter phage A120 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47516 | 43.59 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065670 | Campylobacter phage A121 | Campylobacter phage A121 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47658 | 43.598 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065669 | Campylobacter phage A123 | Campylobacter phage A123 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50156 | 43.827 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065668 | Campylobacter phage A126 | Campylobacter phage A126 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43082 | 39.049 | Unclassified | Ounavirinae | Myoviridae | Campylobacter |
| MG065667 | Campylobacter phage A127 | Campylobacter phage A127 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47287 | 43.847 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065666 | Campylobacter phage A12a | Campylobacter phage A12a unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135398 | 43.62 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065665 | Campylobacter phage A131 | Campylobacter phage A131 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47181 | 43.85 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065664 | Campylobacter phage A132 | Campylobacter phage A132 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70739 | 39.213 | Unclassified | Ounavirinae | Myoviridae | Campylobacter |
| MG065663 | Campylobacter phage A134 | Campylobacter phage A134 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47181 | 43.85 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065662 | Campylobacter phage A142 | Campylobacter phage A142 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140869 | 43.506 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065661 | Campylobacter phage C9 | Campylobacter phage C9 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128230 | 43.48 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065659 | Campylobacter phage C5 | Campylobacter phage C5 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75622 | 43.132 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065658 | Campylobacter phage C7 | Campylobacter phage C7 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134946 | 43.527 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065657 | Campylobacter phage C4 | Campylobacter phage C4 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49946 | 43.771 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065656 | Campylobacter phage C3 | Campylobacter phage C3 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146366 | 37.408 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065655 | Campylobacter phage C2 | Campylobacter phage C2 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140158 | 43.649 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065654 | Campylobacter phage C15 | Campylobacter phage C15 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135398 | 43.62 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065653 | Campylobacter phage C14 | Campylobacter phage C14 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87512 | 43.223 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065652 | Campylobacter phage C11 | Campylobacter phage C11 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58772 | 43.635 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065650 | Campylobacter phage B15 | Campylobacter phage B15 unclassified Phapecoctavirus Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150742 | 39.085 | Phapecoctavirus | Unclassified | Myoviridae | Campylobacter |
| MG065649 | Campylobacter phage A143 | Campylobacter phage A143 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49946 | 43.771 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065648 | Campylobacter phage A144 | Campylobacter phage A144 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35197 | 54.525 | Dhillonvirus | Unclassified | Siphoviridae | Campylobacter |
| MG065647 | Campylobacter phage D# | Campylobacter phage D# unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137061 | 43.619 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065646 | Campylobacter phage D1 | Campylobacter phage D1 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137167 | 43.621 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065645 | Campylobacter phage A145 | Campylobacter phage A145 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146364 | 37.408 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065644 | Campylobacter phage A147 | Campylobacter phage A147 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143296 | 43.567 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| MG065643 | Campylobacter phage E6 | Campylobacter phage E6 unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49946 | 43.771 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065642 | Campylobacter phage A148 | Campylobacter phage A148 unclassified Epseptimavirus Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113314 | 40.307 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Campylobacter |
| MG065641 | Campylobacter phage A14a | Campylobacter phage A14a unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47249 | 43.929 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065640 | Campylobacter phage A14b | Campylobacter phage A14b unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47385 | 43.915 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065639 | Campylobacter phage A150 | Campylobacter phage A150 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146366 | 37.408 | Unclassified | Vequintavirinae | Myoviridae | Campylobacter |
| MG065638 | Campylobacter phage A15a | Campylobacter phage A15a unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22680 | 43.558 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065637 | Campylobacter phage A15b | Campylobacter phage A15b unclassified Tunavirinae Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22375 | 43.812 | Unclassified | Tunavirinae | Drexlerviridae | Campylobacter |
| MG065636 | Campylobacter phage A16a | Campylobacter phage A16a unclassified Hanrivervirus Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50168 | 43.956 | Hanrivervirus | Tempevirinae | Drexlerviridae | Campylobacter |
| MG065635 | Campylobacter phage C12 | Campylobacter phage C12 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137603 | 43.591 | Vequintavirus | Vequintavirinae | Myoviridae | Campylobacter |
| KX379666 | Lactococcus phage 3R16S | Lactococcus phage 3R16S Lactococcus virus 3R16S Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30446 | 34.98 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379664 | Lactococcus phage 10W18 | Lactococcus phage 10W18 Lactococcus virus 10W18 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32393 | 34.949 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379665 | Lactococcus phage i0139 | Lactococcus phage i0139 Lactococcus virus i0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29513 | 35.1 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379667 | Lactococcus phage 4R15L | Lactococcus phage 4R15L unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30782 | 34.432 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379668 | Lactococcus phage 4R16L2 | Lactococcus phage 4R16L2 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30913 | 34.574 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379669 | Lactococcus phage 10W22S | Lactococcus phage 10W22S unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30934 | 34.939 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379670 | Lactococcus phage 19W07F | Lactococcus phage 19W07F unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30890 | 34.681 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379671 | Lactococcus phage 13W11L | Lactococcus phage 13W11L Lactococcus virus 13W11L Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29952 | 34.709 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379672 | Lactococcus phage 16W12L | Lactococcus phage 16W12L unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32782 | 34.766 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX379673 | Lactococcus phage MW18L | Lactococcus phage MW18L unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33013 | 34.814 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KU935715 | Acinetobacter phage vB_AbaM_ME3 | Acinetobacter phage vB_AbaM_ME3 Acinetobacter virus ME3 Metrivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 234900 | 30.764 | Metrivirus | Unclassified | Myoviridae | Acinetobacter |
| KX815270 | Methylophilaceae phage P19250A | Methylophilaceae phage P19250A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38749 | 35.454 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| KU574722 | Pectinobacterium phage vB_PcaM_CBB | Pectinobacterium phage vB_PcaM_CBB Pectinobacterium virus CBB Mimasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 378379 | 35.919 | Mimasvirus | Unclassified | Myoviridae | Pectobacterium |
| MG676466 | Pseudomonas phage TC6 | Pseudomonas phage TC6 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49796 | 45.062 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| MG603697 | Vibrio phage Vp_R1 | Vibrio phage Vp_R1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112127 | 40.324 | Unclassified | Unclassified | Podoviridae | Vibrio |
| MG563770 | Escherichia phage EMCL318 | Escherichia phage EMCL318 unclassified Gequatrovirus Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5489 | 45.145 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| MG545917 | Vibrio phage VEN | Vibrio phage VEN Vibrio virus VEN Trungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44603 | 43.531 | Trungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| MG655270 | Erwinia phage vB_EamM_Mortimer | Erwinia phage vB_EamM_Mortimer unclassified Agricanvirus Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273914 | 49.815 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MG655269 | Erwinia phage vB_EamM_MadMel | Erwinia phage vB_EamM_MadMel unclassified Agricanvirus Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 275000 | 49.738 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MG655268 | Erwinia phage vB_EamM_Desertfox | Erwinia phage vB_EamM_Desertfox Erwinia virus Desertfox Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 272458 | 49.877 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MG655267 | Erwinia phage vB_EamM_Bosolaphorus | Erwinia phage vB_EamM_Bosolaphorus unclassified Agricanvirus Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 272228 | 49.838 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MG599035 | Acinetobacter phage SWH-Ab-3 | Acinetobacter phage SWH-Ab-3 Acinetobacter virus SWHAb3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41730 | 39.382 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MG674163 | Acinetobacter phage SH-Ab 15497 | Acinetobacter phage SH-Ab 15497 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43420 | 47.87 | Unclassified | Unclassified | Siphoviridae | Acinetobacter |
| MG407615 | Salmonella phage Bp96115 | Salmonella phage Bp96115 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41264 | 47.378 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KX898400 | Pseudomonas phage JG054 | Pseudomonas phage JG054 unclassified Nipunavirus Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57839 | 57.641 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| KX898399 | Pseudomonas phage JG012 | Pseudomonas phage JG012 unclassified Nipunavirus Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58359 | 58.442 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| KX455876 | Aeromonas phage Ahp2 | Aeromonas phage Ahp2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47331 | 57.049 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| MG552615 | Klebsiella phage GML-KpCol1 | Klebsiella phage GML-KpCol1 Klebsiella virus KpCol1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50249 | 50.97 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MG241338 | Salmonella phage YSP2 | Salmonella phage YSP2 Salmonella virus YSP2 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50316 | 42.873 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| KU886222 | Erwinia phage vB_EamM_Special G | Erwinia phage vB_EamM_Special G Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273224 | 49.791 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| JN412589 | Mycobacterium phage Patience | Mycobacterium phage Patience Patiencevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70506 | 50.304 | Patiencevirus | Unclassified | Siphoviridae | Mycobacterium |
| KY652725 | Klebsiella phage vB_KpnP_BIS33 | Klebsiella phage vB_KpnP_BIS33 Klebsiella virus BIS33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41697 | 52.704 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KY652724 | Klebsiella phage vB_KpnP_IL33 | Klebsiella phage vB_KpnP_IL33 Klebsiella virus IL33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41335 | 52.495 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KY652723 | Klebsiella phage vB_KpnP_PRA33 | Klebsiella phage vB_KpnP_PRA33 Klebsiella virus PRA33 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40605 | 52.516 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX231829 | Enterobacter phage Tyrion | Enterobacter phage Tyrion unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41760 | 50.587 | Uetakevirus | Unclassified | Podoviridae | Enterobacter |
| KU886225 | Erwinia phage vB_EamM_Deimos-Minion | Erwinia phage vB_EamM_Deimos-Minion Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 273501 | 49.861 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| KU886224 | Erwinia phage vB_EamM_RAY | Erwinia phage vB_EamM_RAY Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271182 | 49.872 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| KU886223 | Erwinia phage vB_EamM_Simmy50 | Erwinia phage vB_EamM_Simmy50 Agricanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 271088 | 49.855 | Agricanvirus | Unclassified | Myoviridae | Erwinia |
| MG543995 | Staphylococcus phage UPMK_1 | Staphylococcus phage UPMK_1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152788 | 31.917 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KX669227 | Escherichia phage ZG49 | Escherichia phage ZG49 Escherichia virus ZG49 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40291 | 49.815 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| KX405003 | Salmonella phage BPS15Q2 | Salmonella phage BPS15Q2 Salmonella virus BPS15Q2 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89817 | 38.857 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KX405002 | Salmonella phage BPS11Q3 | Salmonella phage BPS11Q3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43788 | 49.705 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KX231828 | Enterobacter phage Arya | Enterobacter phage Arya unclassified Jilinvirus Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41918 | 54.108 | Jilinvirus | Unclassified | Myoviridae | Enterobacter |
| KT206225 | Mycobacterium phage Phlei | Mycobacterium phage Phlei unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50418 | 60.062 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ410134 | Mycobacterium phage CRB1 | Mycobacterium phage CRB1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52963 | 63.227 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ410132 | Mycobacterium phage 20ES | Mycobacterium phage 20ES unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53124 | 63.427 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ192196 | Mycobacterium phage 40AC | Mycobacterium phage 40AC unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53396 | 63.338 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| JX899358 | Mycobacterium phage First | Mycobacterium phage First unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53028 | 63.425 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| MG208881 | Escherichia phage St11Ph5 | Escherichia phage St11Ph5 unclassified Gamaleyavirus Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72444 | 42.643 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| MF459647 | Erwinia phage vB_EamM_Joad | Erwinia phage vB_EamM_Joad unclassified Risingsunvirus Risingsunvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235374 | 48.294 | Risingsunvirus | Unclassified | Myoviridae | Erwinia |
| MF459646 | Erwinia phage vB_EamM_RisingSun | Erwinia phage vB_EamM_RisingSun Erwinia virus Risingsun Risingsunvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 235108 | 48.318 | Risingsunvirus | Unclassified | Myoviridae | Erwinia |
| KX776161 | Salmonella phage UPF_BP1 | Salmonella phage UPF_BP1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39902 | 47.559 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MF716957 | Ralstonia virus RSIBR1 | Ralstonia virus RSIBR1 unclassified Habenivirus Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6945 | 61.253 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| MG309718 | Lactococcus phage CB14b | Lactococcus phage CB14b unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29339 | 34.773 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG309717 | Lactococcus phage CB14a | Lactococcus phage CB14a unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29339 | 34.783 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| MG488277 | Escherichia phage EG1 | Escherichia phage EG1 Escherichia virus EG1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39919 | 48.461 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| MG459218 | Acinetobacter phage SWH-Ab-1 | Acinetobacter phage SWH-Ab-1 Acinetobacter virus SWHAb1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41567 | 39.416 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MG387042 | Salmonella phage SP3 | Salmonella phage SP3 Salmonella virus SP3 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109306 | 39.022 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MG324354 | Corynebacterium phage phi674 | Corynebacterium phage phi674 Corynebacterium virus phi674 Ikedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43193 | 50.749 | Ikedavirus | Unclassified | Siphoviridae | Corynebacterium |
| MG324353 | Corynebacterium phage phi673 | Corynebacterium phage phi673 Corynebacterium virus phi673 Ikedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44530 | 51.087 | Ikedavirus | Unclassified | Siphoviridae | Corynebacterium |
| MG251388 | Pseudomonas phage Psp6 | Pseudomonas phage Psp6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42789 | 57.103 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MG214764 | Citromicrobium phage vB_Cib_ssDNA_P1 | Citromicrobium phage vB_Cib_ssDNA_P1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4360 | 59.312 | Unclassified | Unclassified | Microviridae | Citromicrobium |
| MG252693 | Lactobacillus phage Lenus | Lactobacillus phage Lenus Lactobacillus virus Lenus Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78891 | 37.204 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| MG269973 | TM7 phage DolZOral124_53_65 | TM7 phage DolZOral124_53_65 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38840 | 52.49 | Unclassified | Unclassified | Podoviridae | Unspecified |
| MG050172 | Escherichia phage DTL | Escherichia phage DTL Escherichia virus DTL Loudonvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45814 | 44.251 | Loudonvirus | Braunvirinae | Drexlerviridae | Escherichia |
| MG018930 | Pseudomonas phage ventosus | Pseudomonas phage ventosus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 97425 | 48.987 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| MG018928 | Pseudomonas phage inbricus | Pseudomonas phage inbricus Pseudomonas virus inbricus Inbricusvirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70211 | 55.856 | Inbricusvirus | Rothmandenesvirinae | Schitoviridae | Pseudomonas |
| MG018927 | Pseudomonas phage nickie | Pseudomonas phage nickie Pseudomonas virus nickie Nickievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112225 | 57.36 | Nickievirus | Unclassified | Siphoviridae | Pseudomonas |
| MG018926 | Pseudomonas phage tabernarius | Pseudomonas phage tabernarius Pseudomonas virus tabernarius Tabernariusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67498 | 52.296 | Tabernariusvirus | Unclassified | Myoviridae | Pseudomonas |
| MG209611 | Aphanizomenon phage vB_AphaS-CL131 | Aphanizomenon phage vB_AphaS-CL131 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 112793 | 39.682 | Unclassified | Unclassified | Siphoviridae | Aphanizomenon |
| LN681537 | Clostridium phage phiCD211 | Clostridium phage phiCD211 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131704 | 26.415 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| KP010413 | Salmonella phage vB_SPuM_SP116 | Salmonella phage vB_SPuM_SP116 Salmonella virus SP116 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87510 | 38.839 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| KY963371 | Bacillus phage Carmel_SA | Bacillus phage Carmel_SA unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40165 | 34.856 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| KY963370 | Bacillus phage Negev_SA | Bacillus phage Negev_SA unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40375 | 35.158 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| KY963369 | Bacillus phage Tavor_SA | Bacillus phage Tavor_SA unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40397 | 35.168 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| MF979564 | Pectobacterium phage DU_PP_V | Pectobacterium phage DU_PP_V Pectobacterium virus DUPPV Hongcheonvirus Mccorquodalevirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106185 | 39.879 | Hongcheonvirus | Mccorquodalevirinae | Demerecviridae | Pectobacterium |
| MF979563 | Pectobacterium phage DU_PP_IV | Pectobacterium phage DU_PP_IV unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145233 | 50.315 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| MF979562 | Pectobacterium phage DU_PP_III | Pectobacterium phage DU_PP_III unclassified Astrithrvirus Astrithrvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11504 | 37.396 | Astrithrvirus | Unclassified | Podoviridae | Pectobacterium |
| MF979561 | Pectobacterium phage DU_PP_II | Pectobacterium phage DU_PP_II Pectobacterium virus DUPPII Unyawovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40142 | 47.917 | Unyawovirus | Studiervirinae | Autographiviridae | Pectobacterium |
| MF979560 | Pectobacterium phage DU_PP_I | Pectobacterium phage DU_PP_I unclassified Certrevirus Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144959 | 50.299 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| MF979559 | Ralstonia phage DU_RP_I | Ralstonia phage DU_RP_I Ralstonia virus DURPI Gyeongsanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40657 | 58.789 | Gyeongsanvirus | Unclassified | Autographiviridae | Ralstonia |
| MG201401 | Escherichia phage PGT2 | Escherichia phage PGT2 Escherichia virus PGT2 Ermolevavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42686 | 51.649 | Ermolevavirus | Unclassified | Autographiviridae | Escherichia |
| MG018224 | Stenotrophomonas phage DLP4 | Stenotrophomonas phage DLP4 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63945 | 65.059 | Pamexvirus | Unclassified | Siphoviridae | Stenotrophomonas |
| KX778611 | Serratia phage SM9-3Y | Serratia phage SM9-3Y Serratia virus SM9-3Y Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39631 | 50.741 | Teetrevirus | Studiervirinae | Autographiviridae | Serratia |
| KY883658 | Vibrio phage JSF27 | Vibrio phage JSF27 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48690 | 42.456 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| KY883657 | Vibrio phage JSF23 | Vibrio phage JSF23 Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50325 | 42.482 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| KY883656 | Vibrio phage JSF9 | Vibrio phage JSF9 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40007 | 50.629 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KY883655 | Vibrio phage JSF12 | Vibrio phage JSF12 Vibrio virus JSF12 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111671 | 42.713 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| KY883654 | Vibrio phage JSF10 | Vibrio phage JSF10 Vibrio virus JSF10 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111671 | 42.705 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| KY883653 | Vibrio phage JSF34 | Vibrio phage JSF34 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39378 | 42.724 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883652 | Vibrio phage JSF24 | Vibrio phage JSF24 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39378 | 42.724 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883651 | Vibrio phage JSF20 | Vibrio phage JSF20 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39378 | 42.724 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883650 | Vibrio phage JSF18 | Vibrio phage JSF18 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38570 | 42.733 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883649 | Vibrio phage JSF36 | Vibrio phage JSF36 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38581 | 42.746 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883648 | Vibrio phage JSF35 | Vibrio phage JSF35 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38688 | 42.708 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883647 | Vibrio phage JSF33 | Vibrio phage JSF33 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39661 | 50.642 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KY883646 | Vibrio phage JSF32 | Vibrio phage JSF32 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38581 | 42.767 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883645 | Vibrio phage JSF31 | Vibrio phage JSF31 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38581 | 42.754 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883644 | Vibrio phage JSF30 | Vibrio phage JSF30 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38056 | 42.795 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883643 | Vibrio phage JSF28 | Vibrio phage JSF28 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38489 | 42.695 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883642 | Vibrio phage JSF15 | Vibrio phage JSF15 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39643 | 50.713 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KY883641 | Vibrio phage JSF11 | Vibrio phage JSF11 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39341 | 42.854 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| KY883640 | Vibrio phage JSF17 | Vibrio phage JSF17 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125174 | 37.187 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883639 | Vibrio phage JSF14 | Vibrio phage JSF14 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125096 | 37.19 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883638 | Vibrio phage JSF13 | Vibrio phage JSF13 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128814 | 37.158 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883637 | Vibrio phage JSF2 | Vibrio phage JSF2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126082 | 37.083 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883636 | Vibrio phage JSF1 | Vibrio phage JSF1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126082 | 37.073 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883635 | Vibrio phage JSF6 | Vibrio phage JSF6 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133685 | 37.122 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY883634 | Vibrio phage JSF5 | Vibrio phage JSF5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132142 | 37.152 | Unclassified | Unclassified | Myoviridae | Vibrio |
| MF574151 | Vibrio phage JSF25 | Vibrio phage JSF25 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38593 | 42.728 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| MF805716 | Pseudomonas phage SL2 | Pseudomonas phage SL2 Pseudomonas virus SL2 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279696 | 36.9 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| MF893271 | Lysinibacillus phage vB_LspM-01 | Lysinibacillus phage vB_LspM-01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50753 | 37.265 | Unclassified | Unclassified | Myoviridae | Lysinibacillus |
| KJ190157 | Escherichia phage vB_EcoS_FFH_1 | Escherichia phage vB_EcoS_FFH_1 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 108483 | 39.24 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| MF807953 | Escherichia phage Ayreon | Escherichia phage Ayreon Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44708 | 50.067 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| KT232076 | Escherichia virus Lambda | Escherichia virus Lambda Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49205 | 49.747 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| MF497422 | Vibrio phage vB_VspS_VS-ABTNL-3 | Vibrio phage vB_VspS_VS-ABTNL-3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43573 | 43.949 | Unclassified | Unclassified | Podoviridae | Vibrio |
| KY996523 | Clostridium phage CPS1 | Clostridium phage CPS1 Clostridium virus CPS1 Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19089 | 28.257 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| KC262634 | Pseudomonas virus H66 | Pseudomonas virus H66 Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65270 | 62.628 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| MG049919 | Shigella phage vB_SflS-ISF001 | Shigella phage vB_SflS-ISF001 Shigella virus ISF001 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.585 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| MF999224 | Lactobacillus phage LJ | Lactobacillus phage LJ Lactobacillus virus LJ Junavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44260 | 44.781 | Junavirus | Unclassified | Siphoviridae | Lactobacillus |
| MF996376 | Escherichia phage SRT8 | Escherichia phage SRT8 Escherichia virus SRT8 Sertoctavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49579 | 48.034 | Sertoctavirus | Tunavirinae | Drexlerviridae | Escherichia |
| MF975721 | Pseudomonas phage VW-6B | Pseudomonas phage VW-6B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35306 | 56.755 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MF975720 | Pseudomonas phage VW-6S | Pseudomonas phage VW-6S Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37917 | 56.901 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| MF988720 | Pseudoalteromonas phage J2-1 | Pseudoalteromonas phage J2-1 Pseudoalteromonas virus J2-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142204 | 37.92 | Unclassified | Unclassified | Myoviridae | Pseudoalteromonas |
| MF959999 | Marinobacter phage PS3 | Marinobacter phage PS3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42168 | 55.689 | Unclassified | Unclassified | Siphoviridae | Marinobacter |
| MF959998 | Marinobacter phage PS6 | Marinobacter phage PS6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58226 | 51.903 | Unclassified | Unclassified | Siphoviridae | Marinobacter |
| MG197996 | Escherichia phage APC_JM3.2 | Escherichia phage APC_JM3.2 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39761 | 47.338 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| KY565347 | Geobacillus phage TP-84 | Geobacillus phage TP-84 Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47703 | 45.542 | Saundersvirus | Unclassified | Siphoviridae | Geobacillus |
| MF398190 | Staphylococcus phage vB_SauM-fRuSau02 | Staphylococcus phage vB_SauM-fRuSau02 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148464 | 30.223 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY652726 | Klebsiella phage vB_KpnM_BIS47 | Klebsiella phage vB_KpnM_BIS47 Klebsiella virus BIS47 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147443 | 44.622 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| AB276040 | Ralstonia virus RSA1 | Ralstonia virus RSA1 Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38760 | 65.348 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| MF498773 | Aeromonas phage AS-szw | Aeromonas phage AS-szw unclassified Ceceduovirus Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229957 | 38.725 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| KY114934 | Salmonella phage SP01 | Salmonella phage SP01 Salmonella virus SP01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 117842 | 38.982 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KX011028 | Staphylococcus phage pSco-10 | Staphylococcus phage pSco-10 unclassified Twortvirinae Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101986 | 31.113 | Unclassified | Twortvirinae | Herelleviridae | Staphylococcus |
| MF612071 | Klebsiella phage AltoGao | Klebsiella phage AltoGao Klebsiella virus AltoGao Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43012 | 53.292 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| MF140401 | Arthrobacter phage Caterpillar | Arthrobacter phage Caterpillar unclassified Gordonvirus Gordonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56622 | 51.06 | Gordonvirus | Unclassified | Siphoviridae | Arthrobacter |
| MF498775 | Aeromonas phage AS-sw | Aeromonas phage AS-sw Aeromonas virus ASsw Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 230024 | 38.657 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MF158045 | Shigella phage Sf22 | Shigella phage Sf22 Shigella virus Sf22 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166283 | 35.509 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MF681663 | Salmonella phage LVR16A | Salmonella phage LVR16A Salmonella virus LVR16A Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111601 | 40.078 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| MF663761 | Klebsiella phage vB_KpnM_KpV79 | Klebsiella phage vB_KpnM_KpV79 Klebsiella virus KpV52 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47760 | 48.955 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| MF614100 | Klebsiella phage vB_KpnS_IME279 | Klebsiella phage vB_KpnS_IME279 Klebsiella virus IME279 Sortsnevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42518 | 59.307 | Sortsnevirus | Unclassified | Podoviridae | Klebsiella |
| MF621978 | Caulobacter phage Lullwater | Caulobacter phage Lullwater Caulobacter virus Lullwater Lullwatervirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44638 | 54.653 | Lullwatervirus | Unclassified | Autographiviridae | Caulobacter |
| MF805809 | Escherichia phage vB_EcoM_PHB05 | Escherichia phage vB_EcoM_PHB05 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147659 | 37.536 | Unclassified | Vequintavirinae | Myoviridae | Escherichia |
| MF663786 | Bordetella phage vB_BbrM_PHB04 | Bordetella phage vB_BbrM_PHB04 Bordetella virus PHB04 Phabquatrovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94005 | 65.464 | Phabquatrovirus | Unclassified | Myoviridae | Bordetella |
| AP018399 | Xanthomonas phage XacN1 | Xanthomonas phage XacN1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 384670 | 50.439 | Unclassified | Unclassified | Myoviridae | Xanthomonas |
| MF787246 | Lactobacillus phage LpeD | Lactobacillus phage LpeD Lactobacillus virus LpeD Elpedvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145162 | 34.112 | Elpedvirus | Unclassified | Herelleviridae | Lactobacillus |
| MF788075 | Pseudomonas phage IME180 | Pseudomonas phage IME180 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49494 | 44.616 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| MF158046 | Shigella phage Sf23 | Shigella phage Sf23 Shigella virus Sf23 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167678 | 35.382 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| MF158044 | Shigella phage Sf18 | Shigella phage Sf18 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90270 | 39.003 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MF158043 | Shigella phage Sf16 | Shigella phage Sf16 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88580 | 39.017 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MF158042 | Shigella phage Sd1 | Shigella phage Sd1 Shigella virus Sd1 Wilsonroadvirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48262 | 44.499 | Wilsonroadvirus | Rogunavirinae | Drexlerviridae | Shigella |
| MF158041 | Shigella phage Sf15 | Shigella phage Sf15 unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88474 | 38.998 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MF158040 | Shigella phage Sf13 | Shigella phage Sf13 Shigella virus Sf13 Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87570 | 38.911 | Mooglevirus | Ounavirinae | Myoviridae | Shigella |
| MF158039 | Shigella phage Sf12 | Shigella phage Sf12 Shigella virus Sf12 Eastlansingvirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47647 | 44.328 | Eastlansingvirus | Rogunavirinae | Drexlerviridae | Shigella |
| MF158037 | Salmonella phage St162 | Salmonella phage St162 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42701 | 51.217 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MF158036 | Salmonella phage St161 | Salmonella phage St161 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29178 | 50.494 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MF614628 | Sinorhizobium phage phi3LM21 | Sinorhizobium phage phi3LM21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41448 | 61.236 | Unclassified | Unclassified | Siphoviridae | Sinorhizobium |
| MF614627 | Sinorhizobium phage phi2LM21 | Sinorhizobium phage phi2LM21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46600 | 60.648 | Unclassified | Unclassified | Siphoviridae | Sinorhizobium |
| KY549443 | Enterococcus phage EFP01 | Enterococcus phage EFP01 Enterococcus virus EFP01 Schiekvirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155053 | 37.018 | Schiekvirus | Brockvirinae | Herelleviridae | Enterococcus |
| MF695815 | Klebsiella phage KPP5665-2 | Klebsiella phage KPP5665-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39241 | 51.576 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MF186604 | Methanosarcina spherical virus | Methanosarcina spherical virus unclassified Tectiviridae Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 10567 | 38.478 | Unclassified | Unclassified | Tectiviridae | Methanosarcina |
| MF595878 | Caldibacillus phage CBP1 | Caldibacillus phage CBP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37316 | 41.867 | Unclassified | Unclassified | Siphoviridae | Caldibacillus |
| KY612839 | Vibrio phage pVco-5 | Vibrio phage pVco-5 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74325 | 38.716 | Unclassified | Unclassified | Schitoviridae | Vibrio |
| KY295894 | Escherichia phage vB_EcoP_F | Escherichia phage vB_EcoP_F Escherichia virus F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39300 | 49.954 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| KY295899 | Escherichia phage vB_EcoP_R | Escherichia phage vB_EcoP_R unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44941 | 44.959 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| KY295897 | Escherichia phage vB_EcoP_K | Escherichia phage vB_EcoP_K Escherichia virus K Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37775 | 45.117 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| KY295893 | Escherichia phage vB_EcoP_D | Escherichia phage vB_EcoP_D unclassified Vectrevirus Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44931 | 44.973 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| KY295892 | Escherichia phage vB_EcoP_C | Escherichia phage vB_EcoP_C Escherichia virus C Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44970 | 44.966 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| KY295891 | Escherichia phage vB_EcoP_B | Escherichia phage vB_EcoP_B Escherichia virus B Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44018 | 45.036 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| MF285618 | Serratia phage vB_SmaM_ 2050HW | Serratia phage vB_SmaM_ 2050HW Serratia virus 2050HW Moabitevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 276025 | 46.793 | Moabitevirus | Unclassified | Myoviridae | Serratia |
| MF285617 | Leclercia phage 10164RH | Leclercia phage 10164RH unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39300 | 50.774 | Teetrevirus | Studiervirinae | Autographiviridae | Leclercia |
| MF285616 | Leclercia phage 10164-302 | Leclercia phage 10164-302 Leclercia virus 10164-302 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39064 | 50.819 | Teetrevirus | Studiervirinae | Autographiviridae | Leclercia |
| MF285620 | Serratia phage 2050H2 | Serratia phage 2050H2 Serratia virus 2050H2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39216 | 50.418 | Teetrevirus | Studiervirinae | Autographiviridae | Serratia |
| MF285619 | Serratia phage 2050H1 | Serratia phage 2050H1 unclassified Miltonvirus Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159631 | 51.957 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| MF285615 | Klebsiella phage 2044-307w | Klebsiella phage 2044-307w Klebsiella virus 2044-307w Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40048 | 52.892 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| MF001367 | Staphylococcus phage SA4 | Staphylococcus phage SA4 unclassified Rakietenvirinae Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10440 | 29.262 | Unclassified | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| MF001366 | Salmonella phage ST1 | Salmonella phage ST1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42285 | 49.926 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MF001365 | Staphylococcus phage SA3 | Staphylococcus phage SA3 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142827 | 30.215 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MF001362 | Salmonella phage SP1 | Salmonella phage SP1 Salmonella virus SP1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156585 | 44.674 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| MF001361 | Enterococcus phage EF5 | Enterococcus phage EF5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141996 | 31.939 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MF001360 | Escherichia phage ECO4 | Escherichia phage ECO4 Escherichia virus ECO4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168638 | 35.466 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MF001359 | Escherichia phage EC121 | Escherichia phage EC121 Escherichia virus EC121 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168805 | 35.529 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| MF001358 | Enterococcus phage EF1 | Enterococcus phage EF1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141996 | 31.939 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| MF001356 | Salmonella phage SG2 | Salmonella phage SG2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33010 | 50.136 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| MF001355 | Proteus phage PM2 | Proteus phage PM2 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163469 | 31.157 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| MF001354 | Salmonella phage SG1 | Salmonella phage SG1 Salmonella virus SG1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169805 | 35.276 | Tequatrovirus | Tevenvirinae | Myoviridae | Salmonella |
| MF765814 | Bacillus phage Taffo16 | Bacillus phage Taffo16 unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164241 | 37.786 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF678790 | Enterococcus phage phiSHEF5 | Enterococcus phage phiSHEF5 Enterococcus virus phiSHEF5 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41598 | 34.747 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MF678789 | Enterococcus phage phiSHEF4 | Enterococcus phage phiSHEF4 Enterococcus virus phiSHEF4 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41081 | 34.724 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MF678788 | Enterococcus phage phiSHEF2 | Enterococcus phage phiSHEF2 Enterococcus virus phiSHEF2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41712 | 34.554 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| MF668275 | Brevibacterium phage LuckyBarnes | Brevibacterium phage LuckyBarnes Brevibacterium virus LuckyBarnes Luckybarnesvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50774 | 61.933 | Luckybarnesvirus | Unclassified | Siphoviridae | Brevibacterium |
| MF375456 | Xanthomonas phage phi Xc10 | Xanthomonas phage phi Xc10 Xanthomonas virus Xc10 Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44597 | 62.435 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| KY705290 | Streptococcus phage P9903 | Streptococcus phage P9903 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35256 | 38.725 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705289 | Streptococcus phage P9902 | Streptococcus phage P9902 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35535 | 38.866 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705288 | Streptococcus phage P9901 | Streptococcus phage P9901 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35211 | 38.866 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705287 | Streptococcus phage P9854 | Streptococcus phage P9854 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36979 | 38.938 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705286 | Streptococcus phage P9853 | Streptococcus phage P9853 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34469 | 39.726 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705285 | Streptococcus phage P9852 | Streptococcus phage P9852 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34638 | 40.04 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705284 | Streptococcus phage P9851 | Streptococcus phage P9851 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35835 | 39.074 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705283 | Streptococcus phage P8922 | Streptococcus phage P8922 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34509 | 38.819 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705282 | Streptococcus phage P8921 | Streptococcus phage P8921 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34516 | 39.002 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705281 | Streptococcus phage P7955 | Streptococcus phage P7955 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36931 | 38.935 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705280 | Streptococcus phage P7954 | Streptococcus phage P7954 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35688 | 38.862 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705279 | Streptococcus phage P7953 | Streptococcus phage P7953 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35786 | 39.071 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705278 | Streptococcus phage P7952 | Streptococcus phage P7952 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37594 | 39.022 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705277 | Streptococcus phage P7951 | Streptococcus phage P7951 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34123 | 39.771 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705276 | Streptococcus phage P7633 | Streptococcus phage P7633 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35921 | 38.974 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705275 | Streptococcus phage P7632 | Streptococcus phage P7632 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35988 | 39.096 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705274 | Streptococcus phage P7631 | Streptococcus phage P7631 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36164 | 38.878 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705273 | Streptococcus phage P7602 | Streptococcus phage P7602 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37202 | 38.654 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705272 | Streptococcus phage P7601 | Streptococcus phage P7601 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35395 | 38.5 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705271 | Streptococcus phage P7574 | Streptococcus phage P7574 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36298 | 38.749 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705270 | Streptococcus phage P7573 | Streptococcus phage P7573 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37715 | 38.685 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705269 | Streptococcus phage P7572 | Streptococcus phage P7572 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33410 | 39.578 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705268 | Streptococcus phage P7571 | Streptococcus phage P7571 unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34907 | 39.388 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705267 | Streptococcus phage P7154 | Streptococcus phage P7154 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34507 | 38.792 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705266 | Streptococcus phage P7152 | Streptococcus phage P7152 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35663 | 38.903 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705265 | Streptococcus phage P7151 | Streptococcus phage P7151 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35025 | 38.789 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705264 | Streptococcus phage P7134 | Streptococcus phage P7134 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34019 | 38.943 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705263 | Streptococcus phage P7133 | Streptococcus phage P7133 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33284 | 38.841 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705262 | Streptococcus phage P7132 | Streptococcus phage P7132 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35843 | 39.04 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705261 | Streptococcus phage P5652 | Streptococcus phage P5652 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33448 | 39.288 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705260 | Streptococcus phage P5651 | Streptococcus phage P5651 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34292 | 39.129 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705259 | Streptococcus phage P5641 | Streptococcus phage P5641 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37235 | 38.848 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705258 | Streptococcus phage P4761 | Streptococcus phage P4761 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38139 | 38.957 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705257 | Streptococcus phage P3684 | Streptococcus phage P3684 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35444 | 38.875 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705256 | Streptococcus phage P3681 | Streptococcus phage P3681 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36168 | 38.412 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY705255 | Streptococcus phage P0095 | Streptococcus phage P0095 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35966 | 38.114 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705254 | Streptococcus phage P0094 | Streptococcus phage P0094 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33335 | 38.548 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705253 | Streptococcus phage P0093 | Streptococcus phage P0093 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34932 | 38.575 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705252 | Streptococcus phage P0092 | Streptococcus phage P0092 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34580 | 38.438 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY705251 | Streptococcus phage P0091 | Streptococcus phage P0091 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35653 | 39.203 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| JX412914 | Escherichia virus M13 | Escherichia virus M13 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.768 | Inovirus | Unclassified | Inoviridae | Escherichia |
| KX826077 | Salmonella phage UPF_BP2 | Salmonella phage UPF_BP2 Salmonella virus UPFBP2 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54894 | 46.009 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| MF476925 | Klebsiella phage YMC16/01/N133_KPN_BP | Klebsiella phage YMC16/01/N133_KPN_BP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58387 | 58.88 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| MF476924 | Klebsiella phage YMC15/11/N53_KPN_BP | Klebsiella phage YMC15/11/N53_KPN_BP unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59100 | 56.301 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MF415413 | Klebsiella phage KPN N54 | Klebsiella phage KPN N54 unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59100 | 56.301 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MF415412 | Klebsiella phage KPN N141 | Klebsiella phage KPN N141 Klebsiella virus KPN N141 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49090 | 50.959 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MF415411 | Klebsiella phage KPN U2874 | Klebsiella phage KPN U2874 unclassified Yonseivirus Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59087 | 56.302 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| MF415410 | Klebsiella phage KPN N137 | Klebsiella phage KPN N137 Klebsiella virus N137 Yonseivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59100 | 56.301 | Yonseivirus | Unclassified | Siphoviridae | Klebsiella |
| KY942057 | Dickeya phage XF4 | Dickeya phage XF4 unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151519 | 49.382 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| KY942056 | Dickeya phage JA15 | Dickeya phage JA15 unclassified Limestonevirus Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153757 | 49.2 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| KY629563 | Sphingobium phage Lacusarx | Sphingobium phage Lacusarx Sphingobium virus Lacusarx Lacusarxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130138 | 60.175 | Lacusarxvirus | Unclassified | Siphoviridae | Sphingobium |
| KX911187 | Rathayibacter phage NCPPB3778 | Rathayibacter phage NCPPB3778 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44520 | 52.934 | Unclassified | Unclassified | Siphoviridae | Rathayibacter |
| KY630163 | Salmonella phage S8 | Salmonella phage S8 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158432 | 44.527 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KJ206563 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138070 | 30.389 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ206562 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138134 | 30.387 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ206561 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139542 | 30.422 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ206560 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139656 | 30.417 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ206559 | Staphylococcus phage 812 | Staphylococcus phage 812 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142096 | 30.396 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| MF612073 | Klebsiella phage SopranoGao | Klebsiella phage SopranoGao Klebsiella virus SopranoGao Lastavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61644 | 57.266 | Lastavirus | Unclassified | Podoviridae | Klebsiella |
| MF612072 | Klebsiella phage MezzoGao | Klebsiella phage MezzoGao Klebsiella virus MezzoGao Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49807 | 50.977 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| MF360958 | Salicola phage SCTP-2 | Salicola phage SCTP-2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 440001 | 29.987 | Unclassified | Unclassified | Myoviridae | Unspecified |
| LC216347 | Pectobacterium phage PPWS4 | Pectobacterium phage PPWS4 Pectobacterium virus PPWS4 Pektosvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39906 | 48.905 | Pektosvirus | Studiervirinae | Autographiviridae | Pectobacterium |
| KX889068 | Vibrio phage vB_VspP_pVa5 | Vibrio phage vB_VspP_pVa5 Vibrio virus PVA5 Galateavirus Pontosvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78145 | 43.158 | Galateavirus | Pontosvirinae | Schitoviridae | Vibrio |
| CP011970 | Peptoclostridium phage phiCDIF1296T | Peptoclostridium phage phiCDIF1296T Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131326 | 26.427 | Unclassified | Unclassified | Siphoviridae | Peptoclostridium |
| MF156580 | Bacillus phage RadRaab | Bacillus phage RadRaab unclassified Claudivirus Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23946 | 30.623 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MF156578 | Bacillus phage KonjoTrouble | Bacillus phage KonjoTrouble Bacillus virus KonjoTrouble Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26061 | 30.125 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MF156577 | Bacillus phage Juan | Bacillus phage Juan Bacillus virus Juan Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25032 | 30.565 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| MF370965 | Pseudoalteromonas phage SL25 | Pseudoalteromonas phage SL25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33906 | 40.668 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| MF370964 | Pseudoalteromonas phage J2-1 | Pseudoalteromonas phage J2-1 Pseudoalteromonas virus J2-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120295 | 35.844 | Unclassified | Unclassified | Myoviridae | Pseudoalteromonas |
| MF428481 | Staphylococcus phage SN8 | Staphylococcus phage SN8 Staphylococcus virus SN8 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39803 | 35.791 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MF428480 | Staphylococcus phage SN10 | Staphylococcus phage SN10 unclassified Coventryvirus Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39779 | 35.805 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MF428479 | Staphylococcus phage SN11 | Staphylococcus phage SN11 unclassified Coventryvirus Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40108 | 35.813 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MF428478 | Staphylococcus phage SN13 | Staphylococcus phage SN13 unclassified Coventryvirus Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40462 | 35.7 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| MF498901 | Bacillus phage Anthony | Bacillus phage Anthony unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159983 | 38.048 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF479730 | Aeromonas phage AS-gz | Aeromonas phage AS-gz Aeromonas virus Asgz Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162422 | 41.127 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| MF448340 | Aeromonas phage AS-zj | Aeromonas phage AS-zj Aeromonas virus ASzj Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229929 | 38.717 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| MF431252 | Salmonella phage vB_SenS_PVP-SE2 | Salmonella phage vB_SenS_PVP-SE2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42425 | 49.982 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KM359505 | Prochlorococcus phage P-TIM68 | Prochlorococcus phage P-TIM68 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197361 | 34.321 | Unclassified | Unclassified | Myoviridae | Prochlorococcus |
| KY829116 | Acinetobacter phage WCHABP1 | Acinetobacter phage WCHABP1 Acinetobacter virus WCHABP1 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45888 | 37.626 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KY888680 | Acinetobacter phage WCHABP5 | Acinetobacter phage WCHABP5 Acinetobacter virus WCHABP5 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40409 | 39.395 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| MF288920 | Bacillus phage Zainny | Bacillus phage Zainny unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162692 | 38.747 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| MF356679 | Escherichia phage D6 | Escherichia phage D6 unclassified Punavirus Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91159 | 47.01 | Punavirus | Unclassified | Myoviridae | Escherichia |
| MF351863 | Synechococcus phage Bellamy | Synechococcus phage Bellamy Synechococcus virus Bellamy Bellamyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 204930 | 41.105 | Bellamyvirus | Unclassified | Myoviridae | Synechococcus |
| MF161328 | Streptococcus phage D4276 | Streptococcus phage D4276 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39707 | 38.036 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KY670595 | Acinetobacter phage WCHABP12 | Acinetobacter phage WCHABP12 Acinetobacter virus WCHABP12 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45415 | 37.578 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KY290958 | Aeromonas phage SW69-9 | Aeromonas phage SW69-9 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173097 | 43.893 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| KY290957 | Aeromonas phage Riv-10 | Aeromonas phage Riv-10 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174311 | 43.831 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| KY290956 | Aeromonas phage L9-6 | Aeromonas phage L9-6 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173578 | 43.864 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| KY290955 | Aeromonas phage 65.2 | Aeromonas phage 65.2 unclassified Ishigurovirus Ishigurovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 236567 | 37.154 | Ishigurovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| KY290954 | Aeromonas phage 56 | Aeromonas phage 56 Aeromonas virus 56 Popoffvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43551 | 55.422 | Popoffvirus | Unclassified | Myoviridae | Aeromonas |
| KY290953 | Aeromonas phage 51 | Aeromonas phage 51 unclassified Popoffvirus Popoffvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43551 | 55.427 | Popoffvirus | Unclassified | Myoviridae | Aeromonas |
| KY290952 | Aeromonas phage 32 | Aeromonas phage 32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48252 | 56.619 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| KY290951 | Aeromonas phage 31.2 | Aeromonas phage 31.2 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172957 | 43.916 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| KY290950 | Aeromonas phage 59.1 | Aeromonas phage 59.1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46057 | 53.972 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| KY290949 | Aeromonas phage Asp37 | Aeromonas phage Asp37 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47977 | 56.802 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| KY290948 | Aeromonas phage 44RR2.8t.2 | Aeromonas phage 44RR2.8t.2 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173590 | 43.88 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| KY290947 | Aeromonas phage 3 | Aeromonas phage 3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46349 | 57.02 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| KY979132 | Acidovorax phage ACP17 | Acidovorax phage ACP17 Acidovorax virus ACP17 Busanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156281 | 58.738 | Busanvirus | Unclassified | Myoviridae | Acidovorax |
| MF481197 | Marinomonas phage CB5A | Marinomonas phage CB5A Marinomonas virus CB5A Murciavirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45385 | 43.217 | Murciavirus | Colwellvirinae | Autographiviridae | Marinomonas |
| KY794643 | Staphylococcus phage vB_Sau_S24 | Staphylococcus phage vB_Sau_S24 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139997 | 30.863 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY794642 | Staphylococcus phage vB_Sau_Clo6 | Staphylococcus phage vB_Sau_Clo6 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143734 | 30.863 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY794641 | Staphylococcus phage vB_Sau_CG | Staphylococcus phage vB_Sau_CG unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142934 | 30.512 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KX495186 | Bacillus phage PK16 | Bacillus phage PK16 unclassified Caeruleovirus Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158127 | 39.858 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX258185 | Klebsiella phage vB_KpnM_KpV477 | Klebsiella phage vB_KpnM_KpV477 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168272 | 39.321 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| KT630649 | Salmonella phage SEN34 | Salmonella phage SEN34 Salmonella virus SEN34 Brunovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40740 | 49.912 | Brunovirus | Unclassified | Myoviridae | Salmonella |
| KT240186 | Dickeya phage BF25/12 | Dickeya phage BF25/12 Dickeya virus BF25-12 Stompvirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43872 | 52.186 | Stompvirus | Corkvirinae | Autographiviridae | Dickeya |
| MF150911 | Ralstonia phage DU_RP_II | Ralstonia phage DU_RP_II Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42091 | 62.165 | Unclassified | Unclassified | Podoviridae | Ralstonia |
| MF402939 | Escherichia phage OSYSP | Escherichia phage OSYSP Escherichia virus OSYSP Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 110901 | 39.162 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KU356691 | Pseudomonas phage ZC08 | Pseudomonas phage ZC08 Pseudomonas virus ZC08 Zicotriavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70774 | 43.074 | Zicotriavirus | Unclassified | Schitoviridae | Pseudomonas |
| KU356689 | Pseudomonas phage ZC01 | Pseudomonas phage ZC01 unclassified Abidjanvirus Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57061 | 63.381 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KU356690 | Pseudomonas phage ZC03 | Pseudomonas phage ZC03 Pseudomonas virus ZC03 Zicotriavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69844 | 42.601 | Zicotriavirus | Unclassified | Schitoviridae | Pseudomonas |
| KY860935 | Salmonella phage LPST10 | Salmonella phage LPST10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47657 | 45.532 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| KY065149 | Vibrio phage JSF7 | Vibrio phage JSF7 Vibrio virus JSF7 Tawavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46318 | 48.307 | Tawavirus | Unclassified | Autographiviridae | Vibrio |
| KY065148 | Vibrio phage JSF3 | Vibrio phage JSF3 unclassified Pacinivirus Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69396 | 34.573 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| KY065147 | Vibrio phage JSF4 | Vibrio phage JSF4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124261 | 37.084 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY821088 | Bacillus phage Harambe | Bacillus phage Harambe Bacillus virus Harambe Harambevirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21684 | 35.293 | Harambevirus | Unclassified | Salasmaviridae | Bacillus |
| KY417925 | Ochrobactrum phage POI1126 | Ochrobactrum phage POI1126 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59782 | 56.243 | Unclassified | Unclassified | Podoviridae | Ochrobactrum |
| KX669658 | Ochrobactrum phage POA1180 | Ochrobactrum phage POA1180 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41655 | 56.615 | Unclassified | Unclassified | Myoviridae | Ochrobactrum |
| KT224359 | Bacillus phage TsarBomba | Bacillus phage TsarBomba Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162486 | 40.074 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT588075 | Acinetobacter phage Ab105-2phi | Acinetobacter phage Ab105-2phi unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61304 | 39.353 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| KY661963 | Liberibacter phage P-JXGC-3 | Liberibacter phage P-JXGC-3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31449 | 39.966 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| KY389316 | Klebsiella phage K5-4 | Klebsiella phage K5-4 Klebsiella virus K5-4 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40163 | 53.136 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KY389315 | Klebsiella phage K5-2 | Klebsiella phage K5-2 Klebsiella virus K5-2 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41116 | 53.342 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KY082667 | Acinetobacter phage SH-Ab 15519 | Acinetobacter phage SH-Ab 15519 Acinetobacter virus SH-Ab 15519 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40493 | 39.464 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KY926793 | Propionibacterium phage PacnesP2 | Propionibacterium phage PacnesP2 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30016 | 54.404 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KY926792 | Propionibacterium phage PacnesP1 | Propionibacterium phage PacnesP1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29533 | 54.011 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KY979108 | Escherichia phage ECP1 | Escherichia phage ECP1 Escherichia virus ECP1 Wongtaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40469 | 51.009 | Wongtaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| MF188997 | Salmonella phage vB_SalP_PM43 | Salmonella phage vB_SalP_PM43 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40574 | 47.461 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| MF346584 | Acinetobacter phage AbP2 | Acinetobacter phage AbP2 Acinetobacter virus AbP2 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45373 | 37.837 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KX578044 | Leuconostoc phage CHA | Leuconostoc phage CHA unclassified Unaquatrovirus Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28142 | 35.992 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KX578043 | Leuconostoc phage CHB | Leuconostoc phage CHB unclassified Unaquatrovirus Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28265 | 35.946 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KX578042 | Leuconostoc phage Ln-7 | Leuconostoc phage Ln-7 Leuconostoc virus Ln7 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28093 | 36.301 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KX555527 | Leuconostoc phage LDG | Leuconostoc phage LDG Leuconostoc virus LDG Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26559 | 36.274 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KY994101 | Pseudomonas phage vB_PaeM_G1 | Pseudomonas phage vB_PaeM_G1 Pseudomonas virus G1 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87646 | 54.682 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KY296501 | Xenohaliotis phage pCXc-HR2015 | Xenohaliotis phage pCXc-HR2015 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35728 | 38.866 | Unclassified | Unclassified | Siphoviridae | Xenohaliotis |
| KY296500 | Xenohaliotis phage pCXc-HC2016 | Xenohaliotis phage pCXc-HC2016 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35736 | 38.866 | Unclassified | Unclassified | Siphoviridae | Xenohaliotis |
| MF166859 | Pasteurella phage PHB01 | Pasteurella phage PHB01 Pasteurella virus PHB01 Wuhanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37287 | 40.725 | Wuhanvirus | Unclassified | Autographiviridae | Pasteurella |
| KY268296 | Acinetobacter phage vB_AbaP_AS11 | Acinetobacter phage vB_AbaP_AS11 Acinetobacter virus AS11 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41642 | 39.292 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KY268295 | Acinetobacter phage vB_AbaP_AS12 | Acinetobacter phage vB_AbaP_AS12 Acinetobacter virus AS12 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41402 | 39.315 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KT187252 | Bacillus phage PBC6 | Bacillus phage PBC6 unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157147 | 39.91 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT070868 | Bacillus phage PBC5 | Bacillus phage PBC5 unclassified Saundersvirus Saundersvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56332 | 40.256 | Saundersvirus | Unclassified | Siphoviridae | Bacillus |
| KT070867 | Bacillus phage PBC2 | Bacillus phage PBC2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168689 | 34.419 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| MF044457 | Escherichia phage ST0 | Escherichia phage ST0 Escherichia virus ST0 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170496 | 37.665 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KY979247 | Flavobacterium phage VK58 | Flavobacterium phage VK58 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46448 | 29.988 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979246 | Flavobacterium phage VK52 | Flavobacterium phage VK52 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46455 | 29.837 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979245 | Flavobacterium phage VK48 | Flavobacterium phage VK48 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46570 | 30.013 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979244 | Flavobacterium phage VK42 | Flavobacterium phage VK42 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46455 | 29.837 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979243 | Flavobacterium phage VK20 | Flavobacterium phage VK20 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46496 | 29.826 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979242 | Flavobacterium phage V182 | Flavobacterium phage V182 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49099 | 29.752 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979241 | Flavobacterium phage V165 | Flavobacterium phage V165 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49013 | 29.727 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979240 | Flavobacterium phage V157 | Flavobacterium phage V157 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48565 | 29.894 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979239 | Flavobacterium phage V156 | Flavobacterium phage V156 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48564 | 29.897 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979238 | Flavobacterium phage FCV-20 | Flavobacterium phage FCV-20 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46496 | 29.826 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979237 | Flavobacterium phage FCV-16 | Flavobacterium phage FCV-16 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46481 | 29.823 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979236 | Flavobacterium phage FCV-10 | Flavobacterium phage FCV-10 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46482 | 29.824 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY979235 | Flavobacterium phage FCV-1 | Flavobacterium phage FCV-1 Flavobacterium virus FCV1 Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46450 | 29.823 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY992520 | Flavobacterium phage V181 | Flavobacterium phage V181 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49014 | 29.728 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY992519 | Flavobacterium phage V175 | Flavobacterium phage V175 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49016 | 29.743 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY951964 | Flavobacterium phage FCV-11 | Flavobacterium phage FCV-11 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46481 | 29.825 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY951963 | Flavobacterium phage FCV-3 | Flavobacterium phage FCV-3 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46496 | 29.826 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY368640 | Bacillus phage vB_BsuM-Goe3 | Bacillus phage vB_BsuM-Goe3 Bacillus virus vBBsuMGoe3 Grisebachstrassevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156430 | 41.927 | Grisebachstrassevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KY368639 | Bacillus phage vB_BsuM-Goe2 | Bacillus phage vB_BsuM-Goe2 unclassified Okubovirus Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146141 | 40.265 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| KX098390 | Erwinia phage vB_EamP_Rexella | Erwinia phage vB_EamP_Rexella Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75448 | 46.905 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| KX098389 | Erwinia phage vB_EamP_Frozen | Erwinia phage vB_EamP_Frozen Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75147 | 46.92 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| KX098391 | Erwinia phage vB_EamP_Gutmeister | Erwinia phage vB_EamP_Gutmeister Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71173 | 46.928 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| KY888882 | Bacillus phage Flapjack | Bacillus phage Flapjack unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166137 | 37.599 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT588073 | Acinetobacter phage Ab105-3phi | Acinetobacter phage Ab105-3phi unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63785 | 39.415 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| KY658680 | Vibrio phage pVa-8 | Vibrio phage pVa-8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53227 | 43.02 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658679 | Vibrio phage pVa-4 | Vibrio phage pVa-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54295 | 43.072 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658678 | Vibrio phage pVa-3 | Vibrio phage pVa-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54344 | 43.063 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658677 | Vibrio phage P3 | Vibrio phage P3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53242 | 43.013 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658676 | Vibrio phage P2 | Vibrio phage P2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46149 | 43.136 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658675 | Vibrio phage H20 | Vibrio phage H20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53224 | 43.022 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KY658674 | Vibrio phage H8 | Vibrio phage H8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46157 | 43.135 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581100 | Vibrio phage pVa-7 | Vibrio phage pVa-7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54268 | 43.064 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581099 | Vibrio phage Strym | Vibrio phage Strym Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53226 | 43.022 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581098 | Vibrio phage VaK | Vibrio phage VaK Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53216 | 43.089 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581097 | Vibrio phage pVa-6 | Vibrio phage pVa-6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53227 | 43.021 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581095 | Vibrio phage pVa-1 | Vibrio phage pVa-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53227 | 43.021 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581094 | Vibrio phage pVa-2 | Vibrio phage pVa-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53286 | 43.019 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581093 | Vibrio phage Pel | Vibrio phage Pel Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53227 | 43.021 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581092 | Vibrio phage Len | Vibrio phage Len Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53226 | 43.022 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581091 | Vibrio phage Cla | Vibrio phage Cla Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53226 | 43.022 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX581090 | Vibrio phage Her | Vibrio phage Her Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53226 | 43.022 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| MF176161 | Bacillus phage Deep-Purple | Bacillus phage Deep-Purple Bacillus virus DeepPurple Deurplevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36278 | 38.357 | Deurplevirus | Unclassified | Siphoviridae | Bacillus |
| MF036692 | Serratia phage X20 | Serratia phage X20 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172450 | 38.655 | Winklervirus | Unclassified | Myoviridae | Serratia |
| MF036691 | Serratia phage CBH8 | Serratia phage CBH8 unclassified Winklervirus Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171175 | 38.736 | Winklervirus | Unclassified | Myoviridae | Serratia |
| MF036690 | Serratia phage CHI14 | Serratia phage CHI14 Serratia virus CHI14 Winklervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171175 | 38.735 | Winklervirus | Unclassified | Myoviridae | Serratia |
| KY985004 | Escherichia phage vB_EcoS_SH2 | Escherichia phage vB_EcoS_SH2 Escherichia virus SH2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49088 | 45.475 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| KY945241 | Synechococcus phage S-H35 | Synechococcus phage S-H35 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174231 | 41.188 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KY962008 | Escherichia phage ST31 | Escherichia phage ST31 Escherichia virus ST31 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39693 | 49.989 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| MF074189 | Sinorhizobium phage phiM5 | Sinorhizobium phage phiM5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44005 | 60.975 | Unclassified | Unclassified | Podoviridae | Sinorhizobium |
| MF063068 | Pseudomonas phage Noxifer | Pseudomonas phage Noxifer Pseudomonas virus Noxifer Noxifervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 278136 | 53.526 | Noxifervirus | Unclassified | Myoviridae | Pseudomonas |
| MF042361 | Pseudomonas phage Skulduggery | Pseudomonas phage Skulduggery Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62978 | 60.709 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| MF042360 | Pseudomonas phage Phabio | Pseudomonas phage Phabio unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 309157 | 42.998 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| KM576124 | Klebsiella phage phiBO1E | Klebsiella phage phiBO1E Klebsiella virus BO1E Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43865 | 53.852 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KT820175 | Ruegeria phage 45A6 | Ruegeria phage 45A6 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53160 | 61.544 | Unclassified | Unclassified | Podoviridae | Ruegeria |
| KX241618 | Xanthomonas phage Xoo-sp2 | Xanthomonas phage Xoo-sp2 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60497 | 66.532 | Pamexvirus | Unclassified | Siphoviridae | Xanthomonas |
| KX077896 | Streptococcus phage phiJH1301-2 | Streptococcus phage phiJH1301-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67195 | 38.634 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX077895 | Streptococcus phage phiZJ20091101-4 | Streptococcus phage phiZJ20091101-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41057 | 41.601 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX077894 | Streptococcus phage phiZJ20091101-3 | Streptococcus phage phiZJ20091101-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22861 | 43.069 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX077893 | Streptococcus phage phiZJ20091101-2 | Streptococcus phage phiZJ20091101-2 unclassified bacterial viruses Viruses | 11295 | 37.707 | Unclassified | Unclassified | Unclassified | Streptococcus |
| KX077892 | Streptococcus phage phiJH1301-3 | Streptococcus phage phiJH1301-3 unclassified bacterial viruses Viruses | 14113 | 41.033 | Unclassified | Unclassified | Unclassified | Streptococcus |
| KX077891 | Streptococcus phage phiJH1301-1 | Streptococcus phage phiJH1301-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45451 | 41.585 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX077890 | Streptococcus phage phiLP081102 | Streptococcus phage phiLP081102 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36636 | 41.396 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX077889 | Streptococcus phage phiZJ20091101-1 | Streptococcus phage phiZJ20091101-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33253 | 40.875 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY499642 | Vibrio phage pVa-21 | Vibrio phage pVa-21 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231998 | 44.576 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KY056620 | Staphylococcus phage P240 | Staphylococcus phage P240 Staphylococcus virus P240 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45985 | 33.111 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| KT878766 | Staphylococcus phage P1105 | Staphylococcus phage P1105 Staphylococcus virus P1105 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42282 | 33.463 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KT809369 | Staphylococcus phage P630 | Staphylococcus phage P630 Staphylococcus virus P630 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40448 | 33.747 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KT809368 | Staphylococcus phage P282 | Staphylococcus phage P282 Staphylococcus virus P282 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41960 | 32.96 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY779849 | Staphylococcus phage qdsa002 | Staphylococcus phage qdsa002 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142499 | 30.259 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY779848 | Staphylococcus phage qdsa001 | Staphylococcus phage qdsa001 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135563 | 29.858 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY554776 | Lactococcus phage AM12 | Lactococcus phage AM12 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125842 | 32.556 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY947509 | Bacillus phage SerPounce | Bacillus phage SerPounce Bacillus virus SerPounce Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27206 | 30.431 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KY945355 | Mycobacterium phage Shandong1 | Mycobacterium phage Shandong1 unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60618 | 67.46 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KY940711 | Bradyrhizobium phage BDU-MI-1 | Bradyrhizobium phage BDU-MI-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121342 | 61.645 | Unclassified | Unclassified | Myoviridae | Bradyrhizobium |
| KY921761 | Bacillus phage BeachBum | Bacillus phage BeachBum Bacillus virus BeachBum Harambevirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21054 | 35.409 | Harambevirus | Unclassified | Salasmaviridae | Bacillus |
| KY271396 | Klebsiella phage 2 LV-2017 | Klebsiella phage 2 LV-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44400 | 53.725 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KY744235 | Sulfolobus islandicus rod-shaped virus 6 | Sulfolobus islandicus rod-shaped virus 6 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35439 | 25.54 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744234 | Sulfolobus islandicus rod-shaped virus 11 | Sulfolobus islandicus rod-shaped virus 11 Usarudivirus SIRV11 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 33356 | 25.8 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744233 | Sulfolobus islandicus rod-shaped virus 5 | Sulfolobus islandicus rod-shaped virus 5 Usarudivirus SIRV5 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36306 | 25.434 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744232 | Sulfolobus islandicus rod-shaped virus 7 | Sulfolobus islandicus rod-shaped virus 7 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 34190 | 25.566 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744231 | Sulfolobus islandicus rod-shaped virus 4 | Sulfolobus islandicus rod-shaped virus 4 Usarudivirus SIRV4 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35035 | 25.623 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744230 | Sulfolobus islandicus rod-shaped virus 10 | Sulfolobus islandicus rod-shaped virus 10 Usarudivirus SIRV10 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32735 | 25.383 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744229 | Sulfolobus islandicus rod-shaped virus 8 | Sulfolobus islandicus rod-shaped virus 8 Usarudivirus SIRV8 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36493 | 25.079 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY744228 | Sulfolobus islandicus rod-shaped virus 9 | Sulfolobus islandicus rod-shaped virus 9 Usarudivirus SIRV9 Usarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36391 | 24.932 | Usarudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KY853667 | Xanthomonas phage Xf409 | Xanthomonas phage Xf409 Xanthomonas virus Xf109 Xylivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8280 | 59.746 | Xylivirus | Unclassified | Inoviridae | Xanthomonas |
| KX513871 | Human gut microviridae SH-CHD8 | Human gut microviridae SH-CHD8 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6312 | 37.231 | Unclassified | Unclassified | Microviridae | Unspecified |
| KX513875 | Human gut microviridae SH-CHD12 | Human gut microviridae SH-CHD12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6363 | 39.132 | Unclassified | Unclassified | Microviridae | Unspecified |
| KX513877 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5726 | 45.651 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513876 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5637 | 47.295 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513874 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4997 | 40.845 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513873 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5631 | 50.346 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513872 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5024 | 40.267 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513870 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 43.759 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513869 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4939 | 44.564 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513868 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6042 | 43.595 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513867 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5018 | 41.052 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513866 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4664 | 44.404 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513865 | Human gut gokushovirus | Human gut gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5135 | 43.934 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KX513864 | Human gokushovirus | Human gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5508 | 49.401 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KT945995 | Enterococcus phage vB_EfaP_IME199 | Enterococcus phage vB_EfaP_IME199 Enterococcus virus IME199 Minhovirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18838 | 34.627 | Minhovirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| KX171213 | Staphylococcus phage vB_SscM-2 | Staphylococcus phage vB_SscM-2 unclassified Sciuriunavirus Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139682 | 31.824 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KX171212 | Staphylococcus phage vB_SscM-1 | Staphylococcus phage vB_SscM-1 Staphylococcus virus SscM1 Sciuriunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139681 | 31.827 | Sciuriunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KX171211 | Salmonella phage vB_SenM-2 | Salmonella phage vB_SenM-2 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158986 | 44.706 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KX171210 | Pseudomonas phage vB_Pae1396P-5 | Pseudomonas phage vB_Pae1396P-5 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72508 | 54.724 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KX171209 | Pseudomonas phage vB_Pae575P-3 | Pseudomonas phage vB_Pae575P-3 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72728 | 54.724 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KX171208 | Pseudomonas phage vB_Pae436M-8 | Pseudomonas phage vB_Pae436M-8 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66922 | 55.757 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KX833905 | Lactococcus phage PLg-TB25 | Lactococcus phage PLg-TB25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38122 | 34.542 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KY629621 | Streptococcus virus MS1 | Streptococcus virus MS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56075 | 42.261 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY554777 | Lactococcus phage LW81 | Lactococcus phage LW81 Lactococcus virus LW81 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128179 | 32.57 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554775 | Lactococcus phage AM11 | Lactococcus phage AM11 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126161 | 32.546 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554774 | Lactococcus phage AM9 | Lactococcus phage AM9 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125294 | 32.536 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554773 | Lactococcus phage AM8 | Lactococcus phage AM8 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126177 | 32.546 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554772 | Lactococcus phage AM5 | Lactococcus phage AM5 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128178 | 32.552 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554771 | Lactococcus phage AM4 | Lactococcus phage AM4 Lactococcus virus AM4 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132949 | 32.508 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554770 | Lactococcus phage AM3 | Lactococcus phage AM3 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126032 | 32.499 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554769 | Lactococcus phage AM2 | Lactococcus phage AM2 unclassified Audreyjarvisvirus Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125656 | 32.532 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554768 | Lactococcus phage AM1 | Lactococcus phage AM1 Lactococcus virus AM1 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125658 | 32.551 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KY554767 | Lactococcus phage AM7 | Lactococcus phage AM7 unclassified Teubervirus Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62252 | 34.237 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554766 | Lactococcus phage AM6 | Lactococcus phage AM6 Lactococcus virus AM6 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62054 | 34.246 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554765 | Lactococcus phage LW4 | Lactococcus phage LW4 unclassified Teubervirus Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60217 | 34.334 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554764 | Lactococcus phage LW33 | Lactococcus phage LW33 unclassified Teubervirus Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59899 | 34.376 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554763 | Lactococcus phage LW32 | Lactococcus phage LW32 unclassified Teubervirus Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60161 | 34.341 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554762 | Lactococcus phage LW31 | Lactococcus phage LW31 Lactococcus virus LW31 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60551 | 34.292 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| KY554761 | Lactococcus phage R31 | Lactococcus phage R31 Lactococcus virus R31 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27203 | 35.378 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KY554760 | Lactococcus phage R3.4 | Lactococcus phage R3.4 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27704 | 35.381 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KX443552 | Cronobacter phage vB_CsaM_leE | Cronobacter phage vB_CsaM_leE Cronobacter virus leE Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181570 | 44.68 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| KX431560 | Cronobacter phage vB_CsaM_leN | Cronobacter phage vB_CsaM_leN unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179516 | 44.897 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| KX431559 | Cronobacter phage vB_CsaM_leB | Cronobacter phage vB_CsaM_leB Cronobacter virus leB Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177907 | 44.871 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| LC168164 | Tenacibaculum phage pT24 | Tenacibaculum phage pT24 Tenacibaculum virus pT24 Kungbxnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 234670 | 28.717 | Kungbxnavirus | Unclassified | Myoviridae | Tenacibaculum |
| CP017906 | Vibrio phage K05K4_VK05K4_2 | Vibrio phage K05K4_VK05K4_2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 13327 | 46.35 | Unclassified | Unclassified | Inoviridae | Vibrio |
| CP017905 | Vibrio phage K05K4_VK05K4_1 | Vibrio phage K05K4_VK05K4_1 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 21012 | 46.354 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KY271401 | Klebsiella phage 1 LV-2017 | Klebsiella phage 1 LV-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29880 | 50.422 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KY271400 | Klebsiella phage 6 LV-2017 | Klebsiella phage 6 LV-2017 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19260 | 53.172 | Unclassified | Unclassified | Podoviridae | Klebsiella |
| KY271399 | Klebsiella phage 5 LV-2017 | Klebsiella phage 5 LV-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47014 | 52.884 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KY271398 | Klebsiella phage 4LV2017 | Klebsiella phage 4LV2017 Klebsiella virus 4LV2017 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33540 | 50.403 | Felsduovirus | Peduovirinae | Myoviridae | Klebsiella |
| KY271397 | Klebsiella phage 3LV2017 | Klebsiella phage 3LV2017 Klebsiella virus 3LV2017 Reginaelenavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35100 | 54.43 | Reginaelenavirus | Peduovirinae | Myoviridae | Klebsiella |
| KY271395 | Klebsiella phage 2b LV-2017 | Klebsiella phage 2b LV-2017 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44279 | 53.007 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KY271402 | Klebsiella phage 48ST307 | Klebsiella phage 48ST307 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52338 | 52.375 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KY389064 | Staphylococcus phage phi879 | Staphylococcus phage phi879 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41448 | 32.014 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY389063 | Staphylococcus phage phi575 | Staphylococcus phage phi575 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42160 | 31.848 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY703222 | Escherichia phage phiC120 | Escherichia phage phiC120 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186570 | 37.626 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KY709296 | Shewanella phage SppYZU05 | Shewanella phage SppYZU05 Shewanella virus SppYZU05 Yushanvirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54319 | 50.64 | Yushanvirus | Nefertitivirinae | Chaseviridae | Shewanella |
| KY705409 | Escherichia phage vB_EcoM_ECOO78 | Escherichia phage vB_EcoM_ECOO78 Escherichia virus ECOO78 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41289 | 53.075 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| KY780482 | Klebsiella phage KOX1 | Klebsiella phage KOX1 Klebsiella virus KOX1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50526 | 51.237 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KY624616 | Corynebacterium phage LGCM-V9 | Corynebacterium phage LGCM-V9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.049 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624615 | Corynebacterium phage LGCM-V8 | Corynebacterium phage LGCM-V8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.049 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624614 | Corynebacterium phage LGCM-V7 | Corynebacterium phage LGCM-V7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.049 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624613 | Corynebacterium phage LGCM-V5 | Corynebacterium phage LGCM-V5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.049 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624612 | Corynebacterium phage LGCM-V6 | Corynebacterium phage LGCM-V6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.049 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624611 | Corynebacterium phage LGCM-V4 | Corynebacterium phage LGCM-V4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36674 | 51.042 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY624610 | Corynebacterium phage LGCM-V3 | Corynebacterium phage LGCM-V3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.046 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY613597 | Corynebacterium phage LGCM-V2 | Corynebacterium phage LGCM-V2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.046 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY653129 | Staphylococcus phage IME1365_01 | Staphylococcus phage IME1365_01 unclassified Rockefellervirus Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44875 | 35.349 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| KY653127 | Corynebacterium phage IME1320_01 | Corynebacterium phage IME1320_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40068 | 58.004 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY653126 | Staphylococcus phage IME1354_01 | Staphylococcus phage IME1354_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42706 | 34.166 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KY653125 | Staphylococcus phage IME1346_01 | Staphylococcus phage IME1346_01 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42689 | 33.428 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY653124 | Staphylococcus phage IME1367_01 | Staphylococcus phage IME1367_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43135 | 33.657 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KY653123 | Staphylococcus phage IME1361_01 | Staphylococcus phage IME1361_01 Staphylococcus virus IME1361-01 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43516 | 32.85 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY653120 | Staphylococcus phage IME1348_01 | Staphylococcus phage IME1348_01 Staphylococcus virus IME1348-01 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42371 | 34.686 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| KY653119 | Morganella phage IME1369_02 | Morganella phage IME1369_02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39387 | 50.022 | Unclassified | Unclassified | Siphoviridae | Morganella |
| KY653118 | Morganella phage IME1369_01 | Morganella phage IME1369_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47739 | 45.82 | Unclassified | Unclassified | Siphoviridae | Morganella |
| KY653116 | Staphylococcus phage IME1318_01 | Staphylococcus phage IME1318_01 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45236 | 34.488 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| KX179905 | Ralstonia phage Rs551 | Ralstonia phage Rs551 Ralstonia virus RS551 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7929 | 60.827 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| KY000003 | Salmonella phage STP07 | Salmonella phage STP07 unclassified Kuttervirus Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160342 | 44.463 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KY914485 | Aeromonas phage phiA8-29 | Aeromonas phage phiA8-29 Aeromonas virus A8-29 Tedavirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 144974 | 48.18 | Tedavirus | Unclassified | Ackermannviridae | Aeromonas |
| KY817360 | Rhodococcus phage Toil | Rhodococcus phage Toil Rhodococcus virus Toil Epsilontectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 17253 | 54.472 | Epsilontectivirus | Unclassified | Tectiviridae | Rhodococcus |
| KU925172 | Escherichia phage PE37 | Escherichia phage PE37 Escherichia virus PE37 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166423 | 35.353 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KX822733 | Pseudoalteromonas phage PH357 | Pseudoalteromonas phage PH357 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136203 | 34.576 | Unclassified | Unclassified | Myoviridae | Pseudoalteromonas |
| KX756572 | Pectobacterium phage PP2 | Pectobacterium phage PP2 Pectobacterium virus PP2 Wanjuvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41841 | 51.622 | Wanjuvirus | Melnykvirinae | Autographiviridae | Pectobacterium |
| KX689784 | Escherichia phage JSS1 | Escherichia phage JSS1 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39024 | 49.923 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| LT841304 | Escherichia phage vB_EcoS_swan01 | Escherichia phage vB_EcoS_swan01 Escherichia virus swan01 Warwickvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50865 | 44.74 | Warwickvirus | Tempevirinae | Drexlerviridae | Escherichia |
| KY626177 | Marinomonas phage CPG1g | Marinomonas phage CPG1g unclassified Murciavirus Murciavirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44244 | 43.36 | Murciavirus | Colwellvirinae | Autographiviridae | Marinomonas |
| KY626176 | Marinomonas phage CPP1m | Marinomonas phage CPP1m Marinomonas virus CPP1m Murciavirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44245 | 43.363 | Murciavirus | Colwellvirinae | Autographiviridae | Marinomonas |
| KY421186 | Flavobacterium phage FL-1 | Flavobacterium phage FL-1 unclassified Ficleduovirus Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53088 | 32.422 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KY056619 | Brucella phage Iz | Brucella phage Iz unclassified Perisivirus Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41446 | 48.12 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KY581279 | Staphylococcus phage pSa-3 | Staphylococcus phage pSa-3 unclassified Kayvirus Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137836 | 30.405 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY318515 | Yersinia phage fHe-Yen3-01 | Yersinia phage fHe-Yen3-01 Yersinia virus fHeYen301 Pokrovskaiavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42771 | 47.672 | Pokrovskaiavirus | Melnykvirinae | Autographiviridae | Yersinia |
| KU977420 | Stx1 converting phage AU6Stx1 | Stx1 converting phage AU6Stx1 Escherichia virus AU6Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57123 | 48.732 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| KU977419 | Stx1 converting phage AU5Stx1 | Stx1 converting phage AU5Stx1 Escherichia virus AU5Stx1 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58371 | 49.775 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| KY555147 | Caulobacter phage Ccr34 | Caulobacter phage Ccr34 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216240 | 66.152 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY555146 | Caulobacter phage Ccr32 | Caulobacter phage Ccr32 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215799 | 66.16 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY555145 | Caulobacter phage Ccr29 | Caulobacter phage Ccr29 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229319 | 66.161 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY555144 | Caulobacter phage Ccr5 | Caulobacter phage Ccr5 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218729 | 66.225 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY555143 | Caulobacter phage Ccr2 | Caulobacter phage Ccr2 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 220299 | 66.129 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY555142 | Caulobacter phage Ccr10 | Caulobacter phage Ccr10 unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219348 | 66.124 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| KY695241 | Wolbachia phage sr1WOdamA | Wolbachia phage sr1WOdamA Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11696 | 34.986 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| KY695239 | Wolbachia phage sr1WOdamA | Wolbachia phage sr1WOdamA Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11689 | 34.118 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| KY709687 | Salmonella phage 29485 | Salmonella phage 29485 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51738 | 48.415 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| KY695153 | Staphylococcus phage SA7 | Staphylococcus phage SA7 Staphylococcus virus SA7 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34730 | 34.129 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KY593455 | Yersinia phage fHe-Yen9-01 | Yersinia phage fHe-Yen9-01 unclassified Tegunavirus Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167773 | 34.791 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| KX905163 | Clostridioides phage phiSemix9P1 | Clostridioides phage phiSemix9P1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56606 | 26.889 | Unclassified | Unclassified | Myoviridae | Clostridioides |
| KY677846 | Escherichia phage phiLLS | Escherichia phage phiLLS Escherichia virus phiLLS Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107263 | 38.989 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KY657202 | Salmonella phage GJL01 | Salmonella phage GJL01 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50407 | 42.843 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| KY622015 | Shewanella phage SppYZU01 | Shewanella phage SppYZU01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43567 | 55.723 | Unclassified | Unclassified | Myoviridae | Shewanella |
| KY606587 | Dinoroseobacter phage vB_DshS-R5C | Dinoroseobacter phage vB_DshS-R5C Dinoroseobacter virus D5C Nanhaivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77874 | 61.546 | Nanhaivirus | Unclassified | Siphoviridae | Dinoroseobacter |
| KY249644 | Synechococcus virus S-ESS1 | Synechococcus virus S-ESS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60362 | 60.904 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| KY176369 | Salmonella phage STP03 | Salmonella phage STP03 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43428 | 49.576 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KY363465 | Providencia phage vB_PreS_PR1 | Providencia phage vB_PreS_PR1 Providencia virus PR1 Priunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118537 | 39.518 | Priunavirus | Unclassified | Siphoviridae | Providencia |
| KX147096 | Serratia phage vB-Sru-IME250 | Serratia phage vB-Sru-IME250 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154938 | 47.351 | Taipeivirus | Unclassified | Ackermannviridae | Serratia |
| LC210520 | Thermus phage phiOH16 | Thermus phage phiOH16 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6533 | 58.442 | Unclassified | Unclassified | Inoviridae | Thermus |
| LC035386 | Thermus phage phiOH3 | Thermus phage phiOH3 Thermus virus OH3 Thomixvirus Paulinoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5688 | 58.034 | Thomixvirus | Unclassified | Paulinoviridae | Thermus |
| KT630647 | Salmonella phage SEN8 | Salmonella phage SEN8 Salmonella virus SEN8 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35203 | 51.936 | Felsduovirus | Peduovirinae | Myoviridae | Salmonella |
| KY653237 | Salmonella phage alphaalpha | Salmonella phage alphaalpha unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.663 | Sinsheimervirus | Bullavirinae | Microviridae | Salmonella |
| KY619305 | Escherichia phage vB_EcoS_ESCO41 | Escherichia phage vB_EcoS_ESCO41 Escherichia virus ESCO41 Nouzillyvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50800 | 46.085 | Nouzillyvirus | Unclassified | Drexlerviridae | Escherichia |
| KY566218 | Corynebacterium phage LGCM-VI | Corynebacterium phage LGCM-VI Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36671 | 51.046 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KM879221 | Delftia phage RG-2014 | Delftia phage RG-2014 Delftia virus RG2014 Dendoorenvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73882 | 59.889 | Dendoorenvirus | Unclassified | Schitoviridae | Delftia |
| CP011103 | Listeria phage LWP01 | Listeria phage LWP01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41913 | 45.506 | Unclassified | Unclassified | Siphoviridae | Listeria |
| KY442063 | Staphylococcus phage Andhra | Staphylococcus phage Andhra Staphylococcus virus Andhra Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18546 | 29.823 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KT717085 | Streptococcus phage 128 | Streptococcus phage 128 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34593 | 38.999 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KT717084 | Streptococcus phage 53 | Streptococcus phage 53 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34239 | 38.9 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KT717083 | Streptococcus phage 73 | Streptococcus phage 73 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36733 | 38.826 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KU873925 | Pseudomonas phage pf16 | Pseudomonas phage pf16 Pseudomonas virus pf16 Chakrabartyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158136 | 52.653 | Chakrabartyvirus | Unclassified | Myoviridae | Pseudomonas |
| KY630187 | Serratia phage BF | Serratia phage BF Serratia virus BF Eneladusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 357154 | 34.423 | Eneladusvirus | Unclassified | Myoviridae | Serratia |
| KY464836 | Ralstonia phage RS-PI-1 | Ralstonia phage RS-PI-1 Ralstonia virus RSPI1 Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43211 | 61.482 | Ampunavirus | Okabevirinae | Autographiviridae | Ralstonia |
| KY472224 | Enterococcus phage EF-P10 | Enterococcus phage EF-P10 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57408 | 39.815 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| KU862660 | Pseudomonas phage phiPMW | Pseudomonas phage phiPMW Pseudomonas virus PMW Plaisancevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103218 | 45.149 | Plaisancevirus | Unclassified | Myoviridae | Pseudomonas |
| KY471386 | Staphylococcus phage St 134 | Staphylococcus phage St 134 Staphylococcus virus St134 Andhravirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18275 | 30.096 | Andhravirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KU836751 | Bacillus phage Leo2 | Bacillus phage Leo2 unclassified Andromedavirus Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48589 | 42.086 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| KT949345 | Aeromonas phage Ahp1 | Aeromonas phage Ahp1 Aeromonas virus Ahp1 Ahphunavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42167 | 58.864 | Ahphunavirus | Melnykvirinae | Autographiviridae | Aeromonas |
| KP966108 | Burkholderia phage AP3 | Burkholderia phage AP3 Burkholderia virus AP3 Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36499 | 64.407 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| KX223815 | Lactobacillus phage P1 | Lactobacillus phage P1 Lactobacillus virus P1 Maenadvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73787 | 38.962 | Maenadvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| KY065477 | Streptococcus phage IPP37 | Streptococcus phage IPP37 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39221 | 40.139 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065505 | Streptococcus phage IPP69 | Streptococcus phage IPP69 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40787 | 40.476 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065504 | Streptococcus phage IPP68 | Streptococcus phage IPP68 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36203 | 40.955 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065503 | Streptococcus phage IPP67 | Streptococcus phage IPP67 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33607 | 40.798 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065502 | Streptococcus phage IPP66 | Streptococcus phage IPP66 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37969 | 37.844 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065501 | Streptococcus phage IPP65 | Streptococcus phage IPP65 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39046 | 38.514 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065500 | Streptococcus phage IPP64 | Streptococcus phage IPP64 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39689 | 41.1 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065499 | Streptococcus phage IPP63 | Streptococcus phage IPP63 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34280 | 40.088 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065498 | Streptococcus phage IPP62 | Streptococcus phage IPP62 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39369 | 40.844 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065497 | Streptococcus phage IPP61 | Streptococcus phage IPP61 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59870 | 39.851 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065496 | Streptococcus phage IPP60 | Streptococcus phage IPP60 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41977 | 40.11 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065495 | Streptococcus phage IPP58 | Streptococcus phage IPP58 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32027 | 40.076 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065494 | Streptococcus phage IPP57 | Streptococcus phage IPP57 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33084 | 40.186 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065493 | Streptococcus phage IPP55 | Streptococcus phage IPP55 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37847 | 38.706 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065492 | Streptococcus phage IPP54 | Streptococcus phage IPP54 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38900 | 38.375 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065491 | Streptococcus phage IPP53 | Streptococcus phage IPP53 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37235 | 38.273 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065490 | Streptococcus phage IPP52 | Streptococcus phage IPP52 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36805 | 38.15 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065489 | Streptococcus phage IPP51 | Streptococcus phage IPP51 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32793 | 39.039 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065488 | Streptococcus phage IPP50 | Streptococcus phage IPP50 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40284 | 39.805 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065487 | Streptococcus phage IPP48 | Streptococcus phage IPP48 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37848 | 38.528 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065486 | Streptococcus phage IPP46 | Streptococcus phage IPP46 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40232 | 41.241 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065485 | Streptococcus phage IPP45 | Streptococcus phage IPP45 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37387 | 40.439 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065484 | Streptococcus phage IPP44 | Streptococcus phage IPP44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30572 | 38.853 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065483 | Streptococcus phage IPP43 | Streptococcus phage IPP43 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33032 | 39.132 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065482 | Streptococcus phage IPP42 | Streptococcus phage IPP42 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35061 | 39.331 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065481 | Streptococcus phage IPP41 | Streptococcus phage IPP41 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34915 | 39.562 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065480 | Streptococcus phage IPP40 | Streptococcus phage IPP40 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40449 | 40.041 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065479 | Streptococcus phage IPP39 | Streptococcus phage IPP39 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37761 | 38.362 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065478 | Streptococcus phage IPP38 | Streptococcus phage IPP38 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40403 | 39.972 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065476 | Streptococcus phage IPP36 | Streptococcus phage IPP36 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40781 | 40.19 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065475 | Streptococcus phage IPP35 | Streptococcus phage IPP35 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40164 | 39.877 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065474 | Streptococcus phage IPP34 | Streptococcus phage IPP34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38867 | 39.972 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065473 | Streptococcus phage IPP32 | Streptococcus phage IPP32 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41421 | 40.265 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065472 | Streptococcus phage IPP31 | Streptococcus phage IPP31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39828 | 40.03 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065471 | Streptococcus phage IPP30 | Streptococcus phage IPP30 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36844 | 40.118 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065470 | Streptococcus phage IPP29 | Streptococcus phage IPP29 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33953 | 39.914 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065469 | Streptococcus phage IPP28 | Streptococcus phage IPP28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33045 | 39.74 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065468 | Streptococcus phage IPP27 | Streptococcus phage IPP27 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32649 | 38.154 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065467 | Streptococcus phage IPP26 | Streptococcus phage IPP26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34001 | 37.693 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065466 | Streptococcus phage IPP25 | Streptococcus phage IPP25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33895 | 37.917 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065465 | Streptococcus phage IPP24 | Streptococcus phage IPP24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34638 | 39.63 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065464 | Streptococcus phage IPP23 | Streptococcus phage IPP23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33903 | 37.917 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065463 | Streptococcus phage IPP22 | Streptococcus phage IPP22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36344 | 40.161 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065462 | Streptococcus phage IPP21 | Streptococcus phage IPP21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37192 | 40.208 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065461 | Streptococcus phage IPP20 | Streptococcus phage IPP20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37441 | 40.14 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065460 | Streptococcus phage IPP19 | Streptococcus phage IPP19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33895 | 40.451 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065459 | Streptococcus phage IPP18 | Streptococcus phage IPP18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34235 | 40.231 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065458 | Streptococcus phage IPP17 | Streptococcus phage IPP17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34864 | 40.245 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065457 | Streptococcus phage IPP16 | Streptococcus phage IPP16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34329 | 37.516 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065456 | Streptococcus phage IPP15 | Streptococcus phage IPP15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38083 | 40.257 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065455 | Streptococcus phage IPP14 | Streptococcus phage IPP14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37252 | 38.441 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065454 | Streptococcus phage IPP12 | Streptococcus phage IPP12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34428 | 40.252 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065453 | Streptococcus phage IPP11 | Streptococcus phage IPP11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37518 | 40.591 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065452 | Streptococcus phage IPP10 | Streptococcus phage IPP10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40648 | 40.041 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065451 | Streptococcus phage IPP9 | Streptococcus phage IPP9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41185 | 39.99 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065450 | Streptococcus phage IPP8 | Streptococcus phage IPP8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40080 | 40.127 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY065449 | Streptococcus phage IPP5 | Streptococcus phage IPP5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33495 | 38.895 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY579375 | Sulfolobus spindle-shaped virus 3 | Sulfolobus spindle-shaped virus 3 unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 15230 | 38.234 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| KY563228 | Sulfolobus spindle-shaped virus Lassen | Sulfolobus spindle-shaped virus Lassen unclassified Alphafusellovirus Alphafusellovirus Fuselloviridae Viruses | 16271 | 37.078 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| AP017972 | Vibrio phage pTD1 | Vibrio phage pTD1 Vibrio virus pTD1 Tidunavirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239276 | 44.546 | Tidunavirus | Gorgonvirinae | Myoviridae | Vibrio |
| KX119183 | Helicobacter phage SwC388G | Helicobacter phage SwC388G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12309 | 36.177 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119182 | Helicobacter phage Is3180G | Helicobacter phage Is3180G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19736 | 37.135 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119181 | Helicobacter phage Pt259G | Helicobacter phage Pt259G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11604 | 36.177 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119178 | Helicobacter phage Pt5303G | Helicobacter phage Pt5303G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16009 | 35.73 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119206 | Helicobacter phage UKEN32U | Helicobacter phage UKEN32U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29882 | 36.691 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119205 | Helicobacter phage DeM53M | Helicobacter phage DeM53M unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28068 | 36.223 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119204 | Helicobacter phage Sw577G | Helicobacter phage Sw577G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26906 | 36.297 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119203 | Helicobacter phage PtB89G | Helicobacter phage PtB89G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27363 | 37.405 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119202 | Helicobacter phage Pt1293U | Helicobacter phage Pt1293U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30071 | 36.933 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119201 | Helicobacter phage FrANT170U | Helicobacter phage FrANT170U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31200 | 37.234 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119200 | Helicobacter phage FrMEG235U | Helicobacter phage FrMEG235U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31236 | 37.255 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119199 | Helicobacter phage Pt5771G | Helicobacter phage Pt5771G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29801 | 37.039 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119198 | Helicobacter phage Pt5322G | Helicobacter phage Pt5322G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28341 | 36.908 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119197 | Helicobacter phage PtB92G | Helicobacter phage PtB92G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30548 | 36.893 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119196 | Helicobacter phage Pt4481G | Helicobacter phage Pt4481G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25388 | 36.817 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119195 | Helicobacter phage FrGC43A | Helicobacter phage FrGC43A unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32975 | 36.264 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119194 | Helicobacter phage FrG12G | Helicobacter phage FrG12G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28565 | 36.422 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119193 | Helicobacter phage FrB58M | Helicobacter phage FrB58M unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22559 | 35.986 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119192 | Helicobacter phage Pt1918U | Helicobacter phage Pt1918U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28670 | 36.216 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119191 | Helicobacter phage Pt4497U | Helicobacter phage Pt4497U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29393 | 36.199 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119190 | Helicobacter phage Pt4472G | Helicobacter phage Pt4472G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27572 | 36.559 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119189 | Helicobacter phage Pt21299RU | Helicobacter phage Pt21299RU unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23008 | 37.118 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119188 | Helicobacter phage FrB41M | Helicobacter phage FrB41M unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29388 | 35.583 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119177 | Helicobacter phage SwA626G | Helicobacter phage SwA626G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30977 | 36.55 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119176 | Helicobacter phage Pt1846U | Helicobacter phage Pt1846U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27960 | 36.996 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119175 | Helicobacter phage Pt22899G | Helicobacter phage Pt22899G unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30078 | 37.22 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KX119174 | Helicobacter phage UKEN31U | Helicobacter phage UKEN31U unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30456 | 36.715 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KY398841 | Escherichia phage vB_EcoS_CEB_EC3a | Escherichia phage vB_EcoS_CEB_EC3a Escherichia virus EC3a Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44234 | 44.217 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| KY295898 | Escherichia phage P AB-2017 | Escherichia phage P AB-2017 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41184 | 51.284 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| KY295896 | Escherichia phage L AB-2017 | Escherichia phage L AB-2017 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41039 | 51.127 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| KY295895 | Escherichia phage G AB-2017 | Escherichia phage G AB-2017 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41519 | 50.782 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| KX160220 | Lactococcus phage Dub35A | Lactococcus phage Dub35A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35188 | 35.381 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160219 | Lactococcus phage C41431 | Lactococcus phage C41431 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31446 | 35.769 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160218 | Lactococcus phage 98204 | Lactococcus phage 98204 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38334 | 36.576 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160217 | Lactococcus phage 98203 | Lactococcus phage 98203 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35867 | 36.315 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160216 | Lactococcus phage 98202 | Lactococcus phage 98202 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35636 | 36.143 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160215 | Lactococcus phage 98104 | Lactococcus phage 98104 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34989 | 36.117 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160214 | Lactococcus phage 98103 | Lactococcus phage 98103 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35904 | 35.754 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160213 | Lactococcus phage 98102 | Lactococcus phage 98102 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34421 | 35.807 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160212 | Lactococcus phage 98101 | Lactococcus phage 98101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34635 | 35.868 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160211 | Lactococcus phage 62503 | Lactococcus phage 62503 Lactococcus virus 62503 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30556 | 36.32 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| KX160210 | Lactococcus phage 62502 | Lactococcus phage 62502 unclassified Vedamuthuvirus Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30616 | 36.089 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| KX160209 | Lactococcus phage 58502 | Lactococcus phage 58502 unclassified Vedamuthuvirus Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30238 | 36.203 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| KX160208 | Lactococcus phage 53802 | Lactococcus phage 53802 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36445 | 35.967 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160207 | Lactococcus phage 53801 | Lactococcus phage 53801 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36378 | 36.264 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160206 | Lactococcus phage 50902 | Lactococcus phage 50902 Lactococcus virus 50902 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30745 | 36.045 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| KX160205 | Lactococcus phage 49801 | Lactococcus phage 49801 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35800 | 35.869 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX160204 | Lactococcus phage 38502 | Lactococcus phage 38502 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34067 | 35.392 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| JX290549 | Pectobacterium phage PhiM1 | Pectobacterium phage PhiM1 Pectobacterium virus fM1 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43827 | 49.182 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| KF156340 | Synechococcus phage S-MbCM100 | Synechococcus phage S-MbCM100 Synechococcus virus SMbCM100 Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170438 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| JQ067084 | Pseudomonas phage PaMx25 | Pseudomonas phage PaMx25 Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57899 | 58.595 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| KX552041 | Escherichia phage ESCO13 | Escherichia phage ESCO13 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149813 | 39.125 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| KX664695 | Escherichia phage ESCO5 | Escherichia phage ESCO5 Escherichia virus ESCO5 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149312 | 38.961 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| KY250482 | Corynebacterium phage phi16 | Corynebacterium phage phi16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58200 | 52.241 | Unclassified | Unclassified | Siphoviridae | Corynebacterium |
| KY385423 | Klebsiella phage vB_KpnP_KpV74 | Klebsiella phage vB_KpnP_KpV74 Klebsiella virus KpV74 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44094 | 54.141 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KY363359 | Streptococcus phage Str03 | Streptococcus phage Str03 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32296 | 40.253 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY349816 | Streptococcus phage Str01 | Streptococcus phage Str01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37030 | 37.934 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KY006853 | Erythrobacter phage vB_EliS_R6L | Erythrobacter phage vB_EliS_R6L Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65675 | 66.52 | Unclassified | Unclassified | Siphoviridae | Erythrobacter |
| LT714109 | Salmonella virus BTP1 | Salmonella virus BTP1 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40525 | 47.45 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KY030782 | Bacillus phage phi3T | Bacillus phage phi3T unclassified Spbetavirus Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128375 | 34.959 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| AP012540 | Stx2-converting phage Stx2a_WGPS8 | Stx2-converting phage Stx2a_WGPS8 Escherichia virus WGPS8 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55023 | 51.03 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| AP012539 | Stx2-converting phage Stx2a_WGPS6 | Stx2-converting phage Stx2a_WGPS6 Escherichia virus WGPS6 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57849 | 50.678 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| AP012538 | Stx2-converting phage Stx2a_WGPS4 | Stx2-converting phage Stx2a_WGPS4 unclassified Pankowvirus Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58538 | 50.922 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| AP012537 | Stx2-converting phage Stx2a_WGPS2 | Stx2-converting phage Stx2a_WGPS2 Escherichia virus WGPS2 Sawaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58326 | 51.38 | Sawaravirus | Unclassified | Siphoviridae | Unspecified |
| AP012536 | Stx2-converting phage Stx2a_1447 | Stx2-converting phage Stx2a_1447 unclassified Sawaravirus Sawaravirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57034 | 51.333 | Sawaravirus | Unclassified | Siphoviridae | Unspecified |
| AP012535 | Stx2-converting phage Stx2a_WGPS9 | Stx2-converting phage Stx2a_WGPS9 Escherichia virus WGPS9 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56224 | 49.945 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AP012534 | Stx2-converting phage Stx2a_F765 | Stx2-converting phage Stx2a_F765 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63552 | 50.176 | Oslovirus | Sepvirinae | Podoviridae | Unspecified |
| AP012533 | Stx2-converting phage Stx2a_F723 | Stx2-converting phage Stx2a_F723 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62274 | 49.319 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AP012532 | Stx2-converting phage Stx2a_F451 | Stx2-converting phage Stx2a_F451 Escherichia virus F451 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64900 | 49.767 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AP012531 | Stx2-converting phage Stx2a_F422 | Stx2-converting phage Stx2a_F422 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63253 | 49.451 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AP012530 | Stx2-converting phage Stx2a_F349 | Stx2-converting phage Stx2a_F349 unclassified Pankowvirus Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59872 | 50.997 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| AP012529 | Stx2-converting phage Stx2a_F403 | Stx2-converting phage Stx2a_F403 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63369 | 49.477 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AB008550 | Pseudomonas virus phiCTX | Pseudomonas virus phiCTX Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35580 | 62.622 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| KX232515 | Staphylococcus phage 3 AJ-2017 | Staphylococcus phage 3 AJ-2017 Staphylococcus virus 3AJ2017 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43922 | 33.277 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KX532239 | Staphylococcus phage StAP1 | Staphylococcus phage StAP1 unclassified Silviavirus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135502 | 30.001 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KY014601 | Escherichia phage Utah | Escherichia phage Utah unclassified Chivirus Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59024 | 56.418 | Chivirus | Unclassified | Siphoviridae | Escherichia |
| KX660669 | Aquamicrobium phage P14 | Aquamicrobium phage P14 Aquamicrobium virus P14 Aqualcavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40551 | 57.833 | Aqualcavirus | Unclassified | Autographiviridae | Aquamicrobium |
| KT454806 | Escherichia phage SUSP2 | Escherichia phage SUSP2 Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88698 | 40.158 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| KT454805 | Escherichia phage phiSUSP1 | Escherichia phage phiSUSP1 Suspvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90743 | 39.761 | Suspvirus | Ounavirinae | Myoviridae | Escherichia |
| KX130960 | Escherichia phage vB_EcoS-IME253 | Escherichia phage vB_EcoS-IME253 Escherichia virus IME253 Rtpvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46717 | 44.215 | Rtpvirus | Braunvirinae | Drexlerviridae | Escherichia |
| KY420199 | Proteus phage vB_PvuS_Pm34 | Proteus phage vB_PvuS_Pm34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39558 | 41.008 | Unclassified | Unclassified | Siphoviridae | Proteus |
| KX853514 | Virus Rctr16k | Virus Rctr16k Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16739 | 67.842 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| KX853513 | Virus Rctr41k | Virus Rctr41k Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41271 | 57.244 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| EU078592 | Escherichia phage DE3 | Escherichia phage DE3 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42925 | 51.884 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| KX431888 | Pseudomonas phage UNO-SLW1 | Pseudomonas phage UNO-SLW1 Pseudomonas virus UNOSLW1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39215 | 57.899 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KX449361 | Pseudomonas phage UNO-SLW2 | Pseudomonas phage UNO-SLW2 unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39167 | 57.891 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KX449362 | Pseudomonas phage UNO-SLW3 | Pseudomonas phage UNO-SLW3 unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39092 | 57.874 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KX449363 | Pseudomonas phage UNO-SLW4 | Pseudomonas phage UNO-SLW4 unclassified Pifdecavirus Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39136 | 57.89 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KY117485 | Ralstonia phage phiAp1 | Ralstonia phage phiAp1 Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44793 | 60.967 | Unclassified | Krylovirinae | Autographiviridae | Ralstonia |
| KY379853 | Salmonella phage LPSE1 | Salmonella phage LPSE1 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41854 | 49.828 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KY316062 | Ralstonia phage RS-PII-1 | Ralstonia phage RS-PII-1 Ralstonia virus RSPII1 Sukuvirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42042 | 63.177 | Sukuvirus | Okabevirinae | Autographiviridae | Ralstonia |
| KY303907 | Enterococcus phage EF-P29 | Enterococcus phage EF-P29 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58984 | 39.768 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| KY290756 | Vibrio phage Vp670 | Vibrio phage Vp670 Vibrio virus Vp670 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43121 | 43.401 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| KX845404 | Klebsiella phage vB_Kpn_IME260 | Klebsiella phage vB_Kpn_IME260 Klebsiella virus IME260 Sugarlandvirus Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123490 | 45.372 | Sugarlandvirus | Unclassified | Demerecviridae | Klebsiella |
| KX987127 | Thermus phage G20c | Thermus phage G20c unclassified Oshimavirus Oshimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81291 | 57.634 | Oshimavirus | Unclassified | Siphoviridae | Thermus |
| HF569092 | Brucella phage Wb | Brucella phage Wb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.166 | Perisivirus | Unclassified | Podoviridae | Brucella |
| HF569091 | Brucella phage Tb | Brucella phage Tb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.173 | Perisivirus | Unclassified | Podoviridae | Brucella |
| HF569090 | Brucella phage R/C | Brucella phage R/C Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38208 | 48.163 | Perisivirus | Unclassified | Podoviridae | Brucella |
| HF569089 | Brucella phage Fi | Brucella phage Fi unclassified Perisivirus Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.182 | Perisivirus | Unclassified | Podoviridae | Brucella |
| HF569088 | Brucella phage Bk2 | Brucella phage Bk2 unclassified Perisivirus Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.177 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KU510289 | Acinetobacter phage LZ35 | Acinetobacter phage LZ35 Acinetobacter virus LZ35 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44885 | 37.948 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KX879643 | Streptococcus phage CHPC1151 | Streptococcus phage CHPC1151 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33507 | 38.473 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX879642 | Streptococcus phage CHPC926 | Streptococcus phage CHPC926 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30077 | 37.025 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX879641 | Streptococcus phage CHPC577 | Streptococcus phage CHPC577 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35129 | 36.827 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KJ802832 | Salmonella phage 9NA | Salmonella phage 9NA Nonanavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52869 | 42.908 | Nonanavirus | Unclassified | Siphoviridae | Salmonella |
| KY203335 | Pasteurella phage PMP-GADVASU-IND | Pasteurella phage PMP-GADVASU-IND Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46335 | 53.392 | Unclassified | Unclassified | Siphoviridae | Pasteurella |
| KY206887 | Clostridium phage Clo-PEP-1 | Clostridium phage Clo-PEP-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50401 | 34.114 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KY245950 | Acinetobacter phage AJO1 | Acinetobacter phage AJO1 Daemvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10823 | 41.347 | Daemvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KY245949 | Acinetobacter phage AJO1 | Acinetobacter phage AJO1 Daemvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16127 | 40.95 | Daemvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KX879602 | Liberibacter phage HHCA1-2 | Liberibacter phage HHCA1-2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38989 | 39.314 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| KX879601 | Liberibacter phage SGCA5-1 | Liberibacter phage SGCA5-1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36022 | 42.144 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| KY038360 | Staphylococcus phage MRSA107 | Staphylococcus phage MRSA107 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24435 | 58.142 | Unclassified | Unclassified | Myoviridae | Staphylococcus |
| KY045851 | Pseudoalteromonas phage C5a | Pseudoalteromonas phage C5a Pseudoalteromonas virus C5a Catalunyavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35209 | 42.231 | Catalunyavirus | Peduovirinae | Myoviridae | Pseudoalteromonas |
| KT264835 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4492 | 36.955 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264834 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4975 | 31.739 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264833 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4826 | 43.349 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264832 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5682 | 32.207 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264831 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4880 | 36.004 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264830 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5007 | 37.827 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264829 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4883 | 45.034 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264828 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5060 | 42.332 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264827 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5111 | 44.884 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264826 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5100 | 44.098 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264825 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5003 | 39.196 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264824 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4879 | 39.598 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264823 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5008 | 39.277 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264822 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4811 | 42.756 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264821 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5583 | 40.695 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264820 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5596 | 38.992 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264819 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4913 | 44.209 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264818 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5015 | 41.735 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264817 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5077 | 48.986 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264816 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5077 | 49.064 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264815 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4631 | 53.444 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264814 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4832 | 47.351 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264813 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4783 | 53.627 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264812 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4672 | 47.389 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264811 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4383 | 52.909 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264810 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4862 | 49.239 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264809 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4876 | 47.17 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264808 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4812 | 46.405 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264807 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4957 | 37.381 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264806 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5053 | 44.053 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264805 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5657 | 43.362 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264804 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4785 | 41.463 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264803 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5121 | 47.295 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264802 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5065 | 45.923 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264801 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5564 | 50.485 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264800 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5562 | 49.964 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264799 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5179 | 48.639 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264798 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5477 | 47.435 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264797 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5326 | 48.442 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264796 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5417 | 46.649 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264795 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5338 | 46.403 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264794 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5267 | 48.016 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264793 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5830 | 45.249 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264792 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5730 | 45.864 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264791 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6028 | 43.215 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264790 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5895 | 46.243 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264789 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5548 | 49.099 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264788 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5672 | 48.519 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264787 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5209 | 33.73 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264786 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5804 | 41.833 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264785 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5767 | 48.916 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264784 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5560 | 46.277 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264783 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4623 | 32.49 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264782 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4710 | 32.675 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264781 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4268 | 40.558 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264780 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4651 | 48.248 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264779 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4850 | 37.897 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264778 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5109 | 49.344 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264777 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5109 | 49.344 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264776 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5330 | 45.947 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264775 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4710 | 39.002 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264774 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5051 | 31.835 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264773 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5379 | 37.163 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264772 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5147 | 38.294 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264771 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5149 | 38.202 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264770 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5482 | 39.949 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264769 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4821 | 41.734 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264768 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5140 | 46.44 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264767 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4833 | 44.093 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264766 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4847 | 42.604 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264765 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5013 | 40.116 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264764 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5515 | 48.685 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264763 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5506 | 48.075 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264762 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5205 | 45.053 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264761 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5301 | 48.198 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264760 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5215 | 45.618 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264759 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5293 | 47.478 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264758 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4824 | 42.62 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264757 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4936 | 47.468 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264756 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5050 | 45.842 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264755 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5389 | 49.935 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264754 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5208 | 48.214 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264753 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5133 | 44.243 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264752 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5111 | 41.655 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264751 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5131 | 46.677 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264750 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5065 | 46.12 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| KT264749 | Gokushovirus WZ-2015a | Gokushovirus WZ-2015a unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5242 | 45.727 | Gokushovirus | Gokushovirinae | Microviridae | Unspecified |
| LT608331 | Pseudomonas phage vB_PaeS_PcyII-40_PfII40a | Pseudomonas phage vB_PaeS_PcyII-40_PfII40a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37033 | 64.343 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KT919973 | Vibrio phage phi-ST2 | Vibrio phage phi-ST2 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 250485 | 41.231 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| KT919972 | Vibrio phage phi-Grn1 | Vibrio phage phi-Grn1 unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 248605 | 41.259 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| KU230356 | Nodularia phage vB_NpeS-2AV2 | Nodularia phage vB_NpeS-2AV2 Nodularia virus 2AV2 Ravarandavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139100 | 40.278 | Ravarandavirus | Unclassified | Siphoviridae | Nodularia |
| AP017925 | Ralstonia phage RP31 | Ralstonia phage RP31 unclassified Ripduovirus Ripduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 276958 | 53.346 | Ripduovirus | Unclassified | Myoviridae | Ralstonia |
| AP017924 | Ralstonia phage RP12 | Ralstonia phage RP12 Ralstonia virus RP12 Ripduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279845 | 53.404 | Ripduovirus | Unclassified | Myoviridae | Ralstonia |
| KU517658 | Clostridium phage HM T | Clostridium phage HM T Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38039 | 29.025 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| HE956710 | Yersinia phage phi80-18 | Yersinia phage phi80-18 Yersinia virus Phi80-18 Pokrovskaiavirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42406 | 47.637 | Pokrovskaiavirus | Melnykvirinae | Autographiviridae | Yersinia |
| KU057941 | Clostridium phage CDSH1 | Clostridium phage CDSH1 unclassified Leicestervirus Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41619 | 30.815 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| CP013282 | Bacillus phage pGIL02 | Bacillus phage pGIL02 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14961 | 39.703 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| KX268652 | Acinetobacter phage vB_AbaS_TRS1 | Acinetobacter phage vB_AbaS_TRS1 unclassified Vieuvirus Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40749 | 40.256 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| KU197014 | Xanthomonas phage XAJ2 | Xanthomonas phage XAJ2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49241 | 47.434 | Unclassified | Unclassified | Siphoviridae | Xanthomonas |
| KU197013 | Xanthomonas phage XAJ24 | Xanthomonas phage XAJ24 Xanthomonas virus XAJ24 Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44862 | 58.205 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| KX228400 | Clostridium phage CDKM15 | Clostridium phage CDKM15 Clostridioides virus CDKM15 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50605 | 28.683 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| KX228399 | Clostridium phage CDKM9 | Clostridium phage CDKM9 Clostridioides virus CDKM9 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49822 | 28.975 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| KU238070 | Stx converting phage vB_EcoS_ST2-8624 | Stx converting phage vB_EcoS_ST2-8624 Enterobacteria virus ST2-8624 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62822 | 49.607 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| KU238069 | Stx converting phage vB_EcoS_P22 | Stx converting phage vB_EcoS_P22 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62331 | 49.292 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| KU238068 | Stx converting phage vB_EcoS_P32 | Stx converting phage vB_EcoS_P32 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62535 | 49.296 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| KU238067 | Stx converting phage vB_EcoS_P27 | Stx converting phage vB_EcoS_P27 Escherichia virus P27 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61580 | 49.169 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| HE956707 | Yersinia phage phiR8-01 | Yersinia phage phiR8-01 Yersinia virus R8-01 Pienvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42085 | 51.845 | Pienvirus | Melnykvirinae | Autographiviridae | Yersinia |
| KY006474 | Mycobacterium phage Prann | Mycobacterium phage Prann unclassified Pegunavirus Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68398 | 66.638 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KY000085 | Staphylococcus phage SCH111 | Staphylococcus phage SCH111 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18018 | 29.348 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KY000084 | Staphylococcus phage SCH1 | Staphylococcus phage SCH1 Staphylococcus virus SCH1 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18023 | 29.34 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KY000083 | Pseudomonas phage PA10 | Pseudomonas phage PA10 Pseudomonas virus PA10 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91212 | 49.305 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KY000082 | Pseudomonas phage PA5 | Pseudomonas phage PA5 Pseudomonas virus PA5 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66182 | 55.489 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KY000081 | Klebsiella phage KPV811 | Klebsiella phage KPV811 Klebsiella virus KPV811 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42641 | 51.347 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KY000080 | Klebsiella phage KPV15 | Klebsiella phage KPV15 unclassified Jiaodavirus Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167034 | 39.522 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| KY073123 | Serratia phage vB-Sru-IME250 | Serratia phage vB-Sru-IME250 Taipeivirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154938 | 47.351 | Taipeivirus | Unclassified | Ackermannviridae | Serratia |
| KX827371 | Staphylococcus phage SpT99F3 | Staphylococcus phage SpT99F3 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40751 | 35.587 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KX827370 | Staphylococcus phage SpT252 | Staphylococcus phage SpT252 unclassified Coventryvirus Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40093 | 35.695 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| KX827369 | Staphylococcus phage SpT152 | Staphylococcus phage SpT152 Staphylococcus virus SpT152 Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41087 | 35.537 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| KX827368 | Staphylococcus phage SpT5 | Staphylococcus phage SpT5 unclassified Coventryvirus Coventryvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39804 | 35.798 | Coventryvirus | Unclassified | Siphoviridae | Staphylococcus |
| KY073228 | Pseudomonas phage Zigelbrucke | Pseudomonas phage Zigelbrucke Pseudomonas virus Zigelbrucke Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92338 | 49.318 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KY022433 | Vibrio phage J14 | Vibrio phage J14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88973 | 45.401 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KU595434 | Xanthomonas phage f29-Xaj | Xanthomonas phage f29-Xaj unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41865 | 49.798 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Xanthomonas |
| KU595433 | Xanthomonas phage f30-Xaj | Xanthomonas phage f30-Xaj Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44262 | 59.882 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| KU595432 | Xanthomonas phage f20-Xaj | Xanthomonas phage f20-Xaj Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43851 | 59.83 | Pradovirus | Unclassified | Autographiviridae | Xanthomonas |
| KJ019165 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179208 | 40.277 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019164 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179177 | 40.264 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019163 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171179 | 39.375 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019162 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179254 | 40.277 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019161 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172684 | 39.143 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019160 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179210 | 40.282 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019159 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225436 | 41.583 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019158 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171292 | 39.367 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019157 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171292 | 39.367 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019156 | Synechococcus phage ACG-2014e | Synechococcus phage ACG-2014e Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189418 | 38.938 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019155 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221967 | 41.575 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019154 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172681 | 39.132 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019153 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171166 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019152 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221396 | 41.535 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019151 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224313 | 41.544 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019150 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 226536 | 41.612 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019149 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222893 | 41.597 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019148 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215097 | 41.503 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019147 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222322 | 41.582 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019146 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225418 | 41.602 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019145 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225175 | 41.616 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019144 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 214751 | 41.535 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019143 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224502 | 41.595 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019142 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222619 | 41.574 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019141 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222089 | 41.583 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019140 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179115 | 40.295 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019139 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179208 | 40.273 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019138 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171295 | 39.364 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019137 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171289 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019136 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179110 | 40.265 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019135 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171274 | 39.356 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019134 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.135 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019133 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172874 | 39.112 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019132 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172261 | 39.111 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019131 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179111 | 40.286 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019130 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179204 | 40.28 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019129 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179142 | 40.276 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019128 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176140 | 39.104 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019127 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179192 | 40.278 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019126 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179203 | 40.282 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019125 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179206 | 40.28 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019124 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179260 | 40.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019123 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219936 | 41.582 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019122 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171285 | 39.364 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019121 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179109 | 40.273 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019120 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179130 | 40.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019119 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179165 | 40.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019118 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179203 | 40.27 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019117 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179155 | 40.271 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019116 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170403 | 39.391 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019115 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179043 | 40.279 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019114 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171192 | 39.375 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019113 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179254 | 40.266 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019112 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179206 | 40.271 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019111 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221929 | 41.571 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019110 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.131 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019109 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.128 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019108 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.125 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019107 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 217673 | 41.544 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019106 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223389 | 41.597 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019105 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215432 | 41.565 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019104 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.129 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019103 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 217937 | 41.557 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019102 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224791 | 41.612 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019101 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216994 | 41.503 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019100 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216273 | 41.527 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019099 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218152 | 41.559 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019098 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222767 | 41.622 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019097 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221746 | 41.508 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019096 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219775 | 41.504 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019095 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222709 | 41.557 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019094 | Synechococcus phage ACG-2014e | Synechococcus phage ACG-2014e Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189421 | 38.943 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019093 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222009 | 41.568 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019092 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224378 | 41.621 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019090 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 224948 | 41.593 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019089 | Synechococcus phage ACG-2014j | Synechococcus phage ACG-2014j Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192117 | 38.657 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019088 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171276 | 39.376 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019087 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171238 | 39.381 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019086 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 226849 | 41.547 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019085 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222326 | 41.543 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019084 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171183 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019083 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179110 | 40.281 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019082 | Synechococcus phage ACG-2014i | Synechococcus phage ACG-2014i Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 190768 | 39.008 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019081 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171192 | 39.376 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019080 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178897 | 40.265 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019079 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179111 | 40.256 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019078 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179107 | 40.271 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019077 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179209 | 40.279 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019076 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172372 | 39.402 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019075 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179101 | 40.268 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019074 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179204 | 40.28 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019073 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178993 | 40.28 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019072 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179116 | 40.278 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019071 | Synechococcus phage ACG-2014g | Synechococcus phage ACG-2014g unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174885 | 39.258 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019070 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179254 | 40.269 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019069 | Synechococcus phage ACG-2014j | Synechococcus phage ACG-2014j Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192108 | 38.566 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019068 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171231 | 39.371 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019067 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171192 | 39.376 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019066 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215767 | 41.568 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019065 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171179 | 39.376 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019064 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175640 | 39.08 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019063 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173725 | 39.111 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019062 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179323 | 40.292 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019061 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172691 | 39.135 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019060 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.133 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019059 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 228143 | 41.619 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019057 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179110 | 40.279 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019056 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179142 | 40.248 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019055 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171294 | 39.358 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019054 | Synechococcus phage ACG-2014e | Synechococcus phage ACG-2014e Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189418 | 38.931 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019053 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225391 | 41.636 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019051 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.125 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019050 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179207 | 40.271 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019049 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.128 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019048 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178873 | 40.27 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019047 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179095 | 40.275 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019046 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179208 | 40.279 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019045 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 226158 | 41.635 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019044 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172681 | 39.131 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019043 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172688 | 39.131 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019042 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172686 | 39.12 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019041 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172802 | 39.146 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019040 | Synechococcus phage ACG-2014b | Synechococcus phage ACG-2014b Synechococcus virus ACG2014bSyn7803C61 Nereusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154990 | 39.226 | Nereusvirus | Unclassified | Myoviridae | Synechococcus |
| KJ019039 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171296 | 39.369 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019038 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171073 | 39.358 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019037 | Synechococcus phage ACG-2014f | Synechococcus phage ACG-2014f Synechococcus virus AC2014fSyn7803C8 Atlauavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225059 | 41.595 | Atlauavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019036 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179107 | 40.285 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019035 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158300 | 40.254 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019034 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179206 | 40.269 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019033 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171035 | 39.373 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019032 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179233 | 40.268 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019031 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179110 | 40.276 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019030 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171279 | 39.366 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019029 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179200 | 40.271 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019028 | Synechococcus phage ACG-2014d | Synechococcus phage ACG-2014d Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179116 | 40.281 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KJ019027 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176034 | 39.109 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ019026 | Synechococcus phage ACG-2014a | Synechococcus phage ACG-2014a unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171282 | 39.36 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KJ210330 | Staphylococcus phage GRCS | Staphylococcus phage GRCS Staphylococcus virus GRCS Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17869 | 28.899 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KX982260 | Alteromonas phage PB15 | Alteromonas phage PB15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37333 | 45.523 | Unclassified | Unclassified | Siphoviridae | Alteromonas |
| KX879752 | Halomonas phage QHHSV-1 | Halomonas phage QHHSV-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37270 | 66.802 | Unclassified | Unclassified | Siphoviridae | Halomonas |
| KX828711 | Shigella phage SH7 | Shigella phage SH7 Shigella virus SH7 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164870 | 35.446 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KX828710 | Shigella phage SH6 | Shigella phage SH6 Shigella virus SH6 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.828 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| KX815338 | Streptomyces phage Joe | Streptomyces phage Joe unclassified Camvirus Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48941 | 65.477 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| KT898134 | Aeromonas phage phiARM81mr | Aeromonas phage phiARM81mr Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60012 | 60.878 | Unclassified | Unclassified | Podoviridae | Aeromonas |
| KT898133 | Aeromonas phage phiARM81ld | Aeromonas phage phiARM81ld Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47457 | 58.603 | Unclassified | Unclassified | Myoviridae | Aeromonas |
| KT630644 | Salmonella phage SEN1 | Salmonella phage SEN1 Salmonella virus SEN1 Eganvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29733 | 53.012 | Eganvirus | Peduovirinae | Myoviridae | Salmonella |
| KT630645 | Salmonella phage SEN4 | Salmonella phage SEN4 Salmonella virus SEN4 Senquatrovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33509 | 53.365 | Senquatrovirus | Peduovirinae | Myoviridae | Salmonella |
| KT630646 | Salmonella phage SEN5 | Salmonella phage SEN5 unclassified Senquatrovirus Senquatrovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33509 | 53.365 | Senquatrovirus | Peduovirinae | Myoviridae | Salmonella |
| KT630648 | Salmonella phage SEN22 | Salmonella phage SEN22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41338 | 47.828 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| LN881738 | Clostridium phage slur17 | Clostridium phage slur17 unclassified Leicestervirus Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41318 | 30.822 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| KJ190158 | Escherichia phage FFH2 | Escherichia phage FFH2 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139020 | 43.609 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KX712071 | Klebsiella phage vB_KpnP_KpV766 | Klebsiella phage vB_KpnP_KpV766 Klebsiella virus KpV766 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41283 | 52.583 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX712070 | Klebsiella phage vB_KpnP_KpV767 | Klebsiella phage vB_KpnP_KpV767 Klebsiella virus KpV767 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40395 | 52.281 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX237516 | Klebsiella phage vB_KpnM_KpV52 | Klebsiella phage vB_KpnM_KpV52 Klebsiella virus KpV80 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47405 | 48.955 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| KX237515 | Klebsiella phage vB_KpnS_KpV522 | Klebsiella phage vB_KpnS_KpV522 Klebsiella virus KpV522 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51099 | 50.848 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KX237514 | Klebsiella phage vB_KpnP_KpV48 | Klebsiella phage vB_KpnP_KpV48 Klebsiella virus KpV48 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44623 | 54.129 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KX961632 | Bacillus phage SBP8a | Bacillus phage SBP8a unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161643 | 38.667 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX961631 | Bacillus phage QCM11 | Bacillus phage QCM11 Bacillus virus QCM11 Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26054 | 30.441 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KX961630 | Bacillus phage QCM8 | Bacillus phage QCM8 unclassified Tsarbombavirus Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164911 | 39.898 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX961629 | Bacillus phage BJ4 | Bacillus phage BJ4 unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160732 | 38.742 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU052488 | Lactobacillus phage SA-C12 | Lactobacillus phage SA-C12 Lactobacillus virus SAC12 Lenusvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79099 | 37.491 | Lenusvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| KX156762 | Staphylococcus phage vB_SauS_IMEP5 | Staphylococcus phage vB_SauS_IMEP5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44677 | 34.261 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KX834009 | Mycobacterium phage Goldilocks | Mycobacterium phage Goldilocks unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75728 | 62.933 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KX774374 | Salinivibrio phage SMHB1 | Salinivibrio phage SMHB1 Salinivibrio virus SMHB1 Playavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32826 | 50.795 | Playavirus | Peduovirinae | Myoviridae | Salinivibrio |
| KX181651 | Xanthomonas phage Xf109 | Xanthomonas phage Xf109 Xanthomonas virus Xf109 Xylivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7190 | 59.597 | Xylivirus | Unclassified | Inoviridae | Xanthomonas |
| LT546029 | Enterococcus phage VFW | Enterococcus phage VFW Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85865 | 30.346 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KJ024807 | Bacillus phage vB_BtS_BMBtp3 | Bacillus phage vB_BtS_BMBtp3 Bacillus virus BMBtp3 Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51366 | 35.446 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| KX349327 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175117 | 39.562 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349326 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175341 | 39.562 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349325 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174137 | 39.511 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349324 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176285 | 39.556 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349322 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174142 | 39.507 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349321 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174353 | 39.505 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349320 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174352 | 39.558 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349319 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173817 | 39.583 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349318 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174785 | 39.63 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349317 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174193 | 39.563 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349316 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174353 | 39.558 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349315 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174046 | 39.569 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349314 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174351 | 39.539 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349312 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174790 | 39.573 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349311 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176183 | 39.548 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349309 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174204 | 39.571 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349308 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175190 | 39.561 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349307 | Cyanophage S-RIM12 | Cyanophage S-RIM12 unclassified Brizovirus Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173821 | 39.578 | Brizovirus | Unclassified | Myoviridae | Synechococcus |
| KX349306 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180111 | 41.103 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349305 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179559 | 41.112 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349304 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179560 | 41.092 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349303 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180780 | 41.087 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349302 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179558 | 41.113 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349301 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179558 | 41.098 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349300 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179565 | 41.107 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349299 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179558 | 41.094 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349298 | Cyanophage S-RIM14 | Cyanophage S-RIM14 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179558 | 41.098 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| KX349297 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193281 | 40.636 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349296 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197625 | 40.323 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349295 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192972 | 40.617 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349294 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197450 | 40.367 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349293 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197868 | 40.343 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349292 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 193001 | 40.637 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349291 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197629 | 40.345 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KX349290 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170397 | 40.646 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349289 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171216 | 40.63 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349288 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169803 | 40.631 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349287 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170397 | 40.645 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349286 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169483 | 40.621 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349285 | Synechococcus phage S-RIM8 | Synechococcus phage S-RIM8 Synechococcus virus SRIM8 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171770 | 40.579 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KX349284 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175334 | 42.163 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349283 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175247 | 42.173 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349282 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175448 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349281 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175440 | 42.164 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349280 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175307 | 42.163 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349279 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175436 | 42.16 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349278 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175312 | 42.171 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349277 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175309 | 42.164 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349276 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175432 | 42.152 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349275 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175261 | 42.169 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349274 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175648 | 42.169 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349273 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175311 | 42.17 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349272 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175431 | 42.156 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349271 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175439 | 42.168 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349270 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175438 | 42.162 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349269 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175435 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349268 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175437 | 42.15 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349267 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175435 | 42.171 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349266 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174272 | 42.182 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349265 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175425 | 42.163 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349264 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175434 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349263 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175444 | 42.15 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349262 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175145 | 42.136 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349261 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174895 | 42.172 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349260 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175436 | 42.154 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349259 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174273 | 42.196 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349258 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175429 | 42.157 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349257 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175432 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349256 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175435 | 42.169 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349255 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175381 | 42.16 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349254 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175441 | 42.148 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349253 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175438 | 42.153 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349252 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175424 | 42.151 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349251 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175433 | 42.177 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349250 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175438 | 42.148 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349249 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175436 | 42.147 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349248 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174268 | 42.197 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349247 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175437 | 42.176 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349246 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175455 | 42.171 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349245 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175423 | 42.162 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349244 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175441 | 42.153 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349243 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175442 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349242 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175440 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349241 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175439 | 42.163 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349240 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174265 | 42.192 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349239 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175434 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349238 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175433 | 42.141 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349237 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175340 | 42.158 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349236 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174281 | 42.19 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349235 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175440 | 42.149 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349234 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175083 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349233 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175447 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349232 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175432 | 42.155 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349231 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175439 | 42.156 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349230 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175777 | 42.164 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349229 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175083 | 42.157 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349228 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175431 | 42.159 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349227 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175432 | 42.158 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KX349226 | Synechococcus phage S-RIM2 | Synechococcus phage S-RIM2 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175410 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| KU686213 | Synechococcus phage S-CAM7 | Synechococcus phage S-CAM7 Synechococcus virus SCAM7 Mazuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 214214 | 41.263 | Mazuvirus | Unclassified | Myoviridae | Synechococcus |
| KU686212 | Synechococcus phage S-CAM7 | Synechococcus phage S-CAM7 Synechococcus virus SCAM7 Mazuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 216121 | 41.18 | Mazuvirus | Unclassified | Myoviridae | Synechococcus |
| KU686211 | Synechococcus phage S-WAM2 | Synechococcus phage S-WAM2 Synechococcus virus SWAM2 Cymopoleiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186386 | 41.321 | Cymopoleiavirus | Unclassified | Myoviridae | Synechococcus |
| KU686210 | Synechococcus phage S-WAM1 | Synechococcus phage S-WAM1 unclassified Libanvirus Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 185102 | 44.659 | Libanvirus | Unclassified | Myoviridae | Synechococcus |
| KU686209 | Synechococcus phage S-CAM22 | Synechococcus phage S-CAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172345 | 39.913 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686208 | Synechococcus phage S-CAM22 | Synechococcus phage S-CAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172395 | 39.902 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686207 | Synechococcus phage S-CAM22 | Synechococcus phage S-CAM22 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172230 | 39.919 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686206 | Synechococcus phage S-CAM9 | Synechococcus phage S-CAM9 Synechococcus virus SCAM9 Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174830 | 39.046 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| KU686205 | Synechococcus phage S-CAM9 | Synechococcus phage S-CAM9 Synechococcus virus SCAM9 Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174659 | 39.014 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| KU686204 | Synechococcus phage S-CAM9 | Synechococcus phage S-CAM9 Synechococcus virus SCAM9 Kanaloavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174812 | 39.045 | Kanaloavirus | Unclassified | Myoviridae | Synechococcus |
| KU686203 | Synechococcus phage S-CAM8 | Synechococcus phage S-CAM8 unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172342 | 39.248 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KU686202 | Synechococcus phage S-CAM4 | Synechococcus phage S-CAM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191615 | 38.586 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686201 | Synechococcus phage S-CAM4 | Synechococcus phage S-CAM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191983 | 38.583 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686200 | Synechococcus phage S-CAM4 | Synechococcus phage S-CAM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191937 | 38.578 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU686199 | Synechococcus phage S-CAM3 | Synechococcus phage S-CAM3 Synechococcus virus SCAM3 Charybdisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 198190 | 41.558 | Charybdisvirus | Unclassified | Myoviridae | Synechococcus |
| KU686198 | Synechococcus phage S-CAM3 | Synechococcus phage S-CAM3 Synechococcus virus SCAM3 Charybdisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197800 | 41.552 | Charybdisvirus | Unclassified | Myoviridae | Synechococcus |
| KU686197 | Synechococcus phage S-CAM3 | Synechococcus phage S-CAM3 Synechococcus virus SCAM3 Charybdisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197843 | 41.565 | Charybdisvirus | Unclassified | Myoviridae | Synechococcus |
| KU686196 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197533 | 43.045 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| KU686195 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197534 | 43.036 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| KU686194 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197263 | 43.033 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| KU686193 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197533 | 43.039 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| KU686192 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 197539 | 43.036 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| KT070866 | Bacillus phage PBC4 | Bacillus phage PBC4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80647 | 33.983 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KF279412 | Mycobacterium phage Bane1 | Mycobacterium phage Bane1 Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69309 | 68.867 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KX591654 | Klebsiella phage vB_KpnP_KpV763 | Klebsiella phage vB_KpnP_KpV763 Klebsiella virus KpV763 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40765 | 53.222 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KX656785 | Leptospira phage Lin_34 | Leptospira phage Lin_34 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87308 | 39.376 | Unclassified | Unclassified | Myoviridae | Leptospira |
| KX879627 | Campylobacter phage vB_CjeM_Los1 | Campylobacter phage vB_CjeM_Los1 Campylobacter virus Los1 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134073 | 26.196 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| KX856662 | Klebsiella phage KP-Rio/2015 | Klebsiella phage KP-Rio/2015 Klebsiella virus KPRio2015 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43557 | 54.134 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KX190835 | Bacillus phage vB_BtS_BMBtp15 | Bacillus phage vB_BtS_BMBtp15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34986 | 35.417 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KX190833 | Bacillus phage vB_BtS_BMBtp14 | Bacillus phage vB_BtS_BMBtp14 Bacillus virus BMBtp14 Bembunaquatrovirus Skryabinvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50740 | 36.762 | Bembunaquatrovirus | Skryabinvirinae | Siphoviridae | Bacillus |
| KX190832 | Bacillus phage vB_BtS_BMBtp13 | Bacillus phage vB_BtS_BMBtp13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26471 | 35.001 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KT372714 | Bacillus phage vB_BtS_BMBtp16 | Bacillus phage vB_BtS_BMBtp16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34979 | 35.055 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KF279416 | Mycobacterium phage SargentShorty9 | Mycobacterium phage SargentShorty9 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53693 | 63.684 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KF279413 | Mycobacterium phage Bane2 | Mycobacterium phage Bane2 Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69306 | 68.868 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KF279411 | Mycobacterium phage Adawi | Mycobacterium phage Adawi Coopervirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70236 | 68.792 | Coopervirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KC862300 | Pseudomonas phage PAK_P4 | Pseudomonas phage PAK_P4 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93147 | 49.267 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| JQ698665 | Mycobacterium virus Nepal | Mycobacterium virus Nepal Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53972 | 63.548 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KX534337 | Escherichia phage Greed | Escherichia phage Greed unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60042 | 44.64 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| KX467643 | Sulfolobus islandicus filamentous virus 2 | Sulfolobus islandicus filamentous virus 2 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 39399 | 33.536 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Sulfolobus |
| KU927497 | Salmonella phage 100268_sal2 | Salmonella phage 100268_sal2 Salmonella virus 100268sal2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125114 | 40.218 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KT852578 | Bacillus phage BMBtp1 | Bacillus phage BMBtp1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35838 | 34.896 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KJ002054 | Shewanella phage Spp001 | Shewanella phage Spp001 Shewanella virus Spp001 Yushanvirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54791 | 49.417 | Yushanvirus | Nefertitivirinae | Chaseviridae | Shewanella |
| HQ918180 | Klebsiella phage KP27 | Klebsiella phage KP27 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174413 | 41.817 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| GU295964 | Klebsiella phage KP15 | Klebsiella phage KP15 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174436 | 41.831 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| KX550065 | Streptococcus phage SpGS-1 | Streptococcus phage SpGS-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37631 | 38.28 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KX507046 | Vibrio phage S4-7 | Vibrio phage S4-7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129756 | 36.823 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KX507045 | Vibrio phage 2E1 | Vibrio phage 2E1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40823 | 41.67 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| LT614807 | Cronobacter phage Pet-CM3-4 | Cronobacter phage Pet-CM3-4 unclassified Karamvirus Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171975 | 39.806 | Karamvirus | Tevenvirinae | Myoviridae | Cronobacter |
| KU927493 | Salmonella phage 118970_sal3 | Salmonella phage 118970_sal3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77375 | 50.701 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KX607102 | Acidianus two-tailed virus variant 1 | Acidianus two-tailed virus variant 1 unclassified Bicaudavirus Bicaudavirus Bicaudaviridae Viruses | 61783 | 41.34 | Bicaudavirus | Unclassified | Bicaudaviridae | Acidianus |
| KX607101 | Acidianus two-tailed virus 2 | Acidianus two-tailed virus 2 unclassified Bicaudavirus Bicaudavirus Bicaudaviridae Viruses | 57900 | 40.684 | Bicaudavirus | Unclassified | Bicaudaviridae | Acidianus |
| KX236333 | Campylobacter phage PC14 | Campylobacter phage PC14 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134927 | 26.16 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| KX456213 | Lactococcus phage 98201 | Lactococcus phage 98201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36202 | 35.951 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456212 | Lactococcus phage 86501 | Lactococcus phage 86501 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33035 | 35.499 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456211 | Lactococcus phage 63301 | Lactococcus phage 63301 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32409 | 35.132 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456210 | Lactococcus phage 62501 | Lactococcus phage 62501 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34084 | 35.82 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456209 | Lactococcus phage 56701 | Lactococcus phage 56701 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34117 | 35.903 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456208 | Lactococcus phage 50901 | Lactococcus phage 50901 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33440 | 35.592 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456207 | Lactococcus phage 50101 | Lactococcus phage 50101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31987 | 35.618 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KX456206 | Lactococcus phage 28201 | Lactococcus phage 28201 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34396 | 35.821 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KU884565 | Pseudomonas phage PaMx46 | Pseudomonas phage PaMx46 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43266 | 45.243 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KU884564 | Pseudomonas phage PaMx43 | Pseudomonas phage PaMx43 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43223 | 45.335 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KU884563 | Pseudomonas phage PaMx41 | Pseudomonas phage PaMx41 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43490 | 45.215 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KU884562 | Pseudomonas phage PaMx35 | Pseudomonas phage PaMx35 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43733 | 45.044 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KU884561 | Pseudomonas phage PaMx33 | Pseudomonas phage PaMx33 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43265 | 45.247 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KX229736 | Campylobacter phage PC5 | Campylobacter phage PC5 unclassified Fletchervirus Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131095 | 26.139 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| KX523699 | Salmonella phage IME207 | Salmonella phage IME207 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47564 | 46.415 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| KX712143 | Sulfolobus islandicus rudivirus 3 | Sulfolobus islandicus rudivirus 3 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32995 | 25.519 | Rudivirus | Unclassified | Rudiviridae | Sulfolobus |
| KX284704 | Enterococcus phage SANTOR1 | Enterococcus phage SANTOR1 Enterococcus virus SANTOR1 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37933 | 34.54 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| KR131750 | Enterococcus phage Ec-ZZ2 | Enterococcus phage Ec-ZZ2 Enterococcus virus EcZZ2 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41170 | 34.596 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| KT990215 | Escherichia phage GA2A | Escherichia phage GA2A Escherichia virus GA2A Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40470 | 51.097 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| KX009778 | Escherichia phage UFV-AREG1 | Escherichia phage UFV-AREG1 Escherichia virus AREG1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170788 | 35.279 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KX397373 | Erwinia phage vB_EamM_Stratton | Erwinia phage vB_EamM_Stratton unclassified Erskinevirus Erskinevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 243953 | 51.332 | Erskinevirus | Unclassified | Myoviridae | Erwinia |
| KX397372 | Erwinia phage vB_EamM_Phobos | Erwinia phage vB_EamM_Phobos unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 229501 | 49.137 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| KX397371 | Erwinia phage vB_EamM_Parshik | Erwinia phage vB_EamM_Parshik unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241050 | 51.038 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| KX397370 | Erwinia phage Machina | Erwinia phage Machina Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241654 | 51.013 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| KX397369 | Erwinia phage vB_EamM_Kwan | Erwinia phage vB_EamM_Kwan Wellingtonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246390 | 52.102 | Wellingtonvirus | Unclassified | Myoviridae | Erwinia |
| KX397367 | Erwinia phage vB_EamM_EarlPhillipIV | Erwinia phage vB_EamM_EarlPhillipIV unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223935 | 50.589 | Derbicusvirus | Unclassified | Myoviridae | Erwinia |
| KX397366 | Erwinia phage vB_EamM_ChrisDB | Erwinia phage vB_EamM_ChrisDB unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244840 | 49.414 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| KX397365 | Erwinia phage vB_EamM_Caitlin | Erwinia phage vB_EamM_Caitlin unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 241147 | 52.17 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| KU927500 | Salmonella phage 118970_sal1 | Salmonella phage 118970_sal1 Salmonella virus 118970sal1 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59518 | 56.492 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KJ748011 | Escherichia phage PE3-1 | Escherichia phage PE3-1 Escherichia virus PE3-1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39093 | 49.927 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| KT809302 | Haloarcula californiae icosahedral virus 1 | Haloarcula californiae icosahedral virus 1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 31314 | 68.126 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| LT603033 | Escherichia phage vB_Eco_slurp01 | Escherichia phage vB_Eco_slurp01 unclassified Asteriusvirus Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348043 | 34.052 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| LN997803 | Escherichia phage phi467 | Escherichia phage phi467 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38594 | 47.067 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| KX522565 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65653 | 34.492 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| KX397368 | Erwinia phage vB_EamM_Huxley | Erwinia phage vB_EamM_Huxley unclassified Machinavirus Machinavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240761 | 51.069 | Machinavirus | Unclassified | Myoviridae | Erwinia |
| AY150271 | Salmonella phage epsilon15 | Salmonella phage epsilon15 Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39672 | 50.834 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| KT381865 | Pelagibaca phage vB_PeaS-P1 | Pelagibaca phage vB_PeaS-P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38868 | 63.518 | Unclassified | Unclassified | Siphoviridae | Pelagibaca |
| KT381864 | Thiobacimonas phage vB_ThpS-P1 | Thiobacimonas phage vB_ThpS-P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39591 | 66.735 | Unclassified | Unclassified | Siphoviridae | Thiobacimonas |
| KX534341 | Escherichia phage Pride | Escherichia phage Pride unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44899 | 54.467 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KX534340 | Phage Wrath | Phage Wrath Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29238 | 34.766 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| KX534339 | Escherichia phage Sloth | Escherichia phage Sloth unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45206 | 54.429 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KX534338 | Escherichia phage Lust | Escherichia phage Lust unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41942 | 54.542 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KX534336 | Escherichia phage Gluttony | Escherichia phage Gluttony unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44513 | 54.456 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KX534335 | Escherichia phage Envy | Escherichia phage Envy unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45206 | 54.457 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KX365748 | Pseudoalteromonas phage BS5 | Pseudoalteromonas phage BS5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39949 | 40.597 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KX365747 | Pseudoalteromonas phage PH1 | Pseudoalteromonas phage PH1 unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42685 | 42.237 | Kafunavirus | Unclassified | Podoviridae | Pseudoalteromonas |
| KX077179 | Rhodovulum phage vB_RhkS_P1 | Rhodovulum phage vB_RhkS_P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38773 | 67.454 | Unclassified | Unclassified | Siphoviridae | Rhodovulum |
| KU847400 | Bacillus phage Crookii | Bacillus phage Crookii unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154012 | 37.165 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| LT546030 | Enterococcus phage VPE25 | Enterococcus phage VPE25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86524 | 30.342 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KU599889 | Flavobacterium phage 1H | Flavobacterium phage 1H Flavobacterium virus 1H Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39290 | 31.441 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| KU599888 | Flavobacterium phage 23T | Flavobacterium phage 23T Flavobacterium virus 23T Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43226 | 31.389 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| KU599887 | Flavobacterium phage 2A | Flavobacterium phage 2A Flavobacterium virus 2A Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46575 | 31.744 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| KU599886 | Flavobacterium phage Fpv20 | Flavobacterium phage Fpv20 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89937 | 26.944 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KU599885 | Flavobacterium phage Fpv11 | Flavobacterium phage Fpv11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.01 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599884 | Flavobacterium phage Fpv10 | Flavobacterium phage Fpv10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42702 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599883 | Flavobacterium phage Fpv8 | Flavobacterium phage Fpv8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599882 | Flavobacterium phage Fpv7 | Flavobacterium phage Fpv7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42894 | 32.037 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599881 | Flavobacterium phage Fpv6 | Flavobacterium phage Fpv6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42820 | 31.985 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599880 | Flavobacterium phage Fpv5 | Flavobacterium phage Fpv5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KU599879 | Flavobacterium phage Fpv3 | Flavobacterium phage Fpv3 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88421 | 27.222 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KU599878 | Flavobacterium phage Fpv2 | Flavobacterium phage Fpv2 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89010 | 26.995 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KU599877 | Flavobacterium phage Fpv1 | Flavobacterium phage Fpv1 Flavobacterium virus Fpv1 Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88994 | 26.992 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KT881477 | Salmonella phage fSE4S | Salmonella phage fSE4S Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41768 | 49.782 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KT336321 | Streptococcus phage phiSC070807 | Streptococcus phage phiSC070807 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57638 | 39.606 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT336320 | Streptococcus phage phiNJ3 | Streptococcus phage phiNJ3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53802 | 39.748 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| EU982300 | Pseudomonas phage DVM-2008 | Pseudomonas phage DVM-2008 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22689 | 59.425 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| AY616033 | Burkholderia phage BcepB1A | Burkholderia phage BcepB1A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47399 | 54.453 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| AY569307 | Sulfolobus turreted icosahedral virus 1 | Sulfolobus turreted icosahedral virus 1 Alphaturrivirus Turriviridae Belfryvirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 17663 | 36.007 | Alphaturrivirus | Unclassified | Turriviridae | Sulfolobus |
| AY328853 | Vibrio phage VP16C | Vibrio phage VP16C Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47537 | 59.377 | Unclassified | Unclassified | Myoviridae | Vibrio |
| AY328852 | Vibrio phage VP16T | Vibrio phage VP16T Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49575 | 58.792 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KX349902 | Bacillus phage Kida | Bacillus phage Kida unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162151 | 38.666 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX458241 | Pseudomonas phage Andromeda | Pseudomonas phage Andromeda Pseudomonas virus Andromeda Bifseptvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40008 | 58.233 | Bifseptvirus | Unclassified | Autographiviridae | Pseudomonas |
| KU981050 | Weissella phage WCP30 | Weissella phage WCP30 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33697 | 41.229 | Unclassified | Unclassified | Siphoviridae | Weissella |
| JN600960 | Enterobacteria phage IME10 | Enterobacteria phage IME10 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39646 | 47.495 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| JF719731 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.365 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| JF719730 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.204 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| JF719729 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.186 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| JF773396 | Liberibacter phage FP2 | Liberibacter phage FP2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38552 | 39.274 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| HQ728265 | Erwinia phage vB_EamP-L1 | Erwinia phage vB_EamP-L1 Erwinia virus L1 Elunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39282 | 51.937 | Elunavirus | Studiervirinae | Autographiviridae | Erwinia |
| JF412302 | EBPR siphovirus 1 | EBPR siphovirus 1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54466 | 55.034 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| JF412301 | EBPR siphovirus 6 | EBPR siphovirus 6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11444 | 66.76 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| JF412300 | EBPR siphovirus 5 | EBPR siphovirus 5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13119 | 63.015 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| JF412299 | EBPR siphovirus 4 | EBPR siphovirus 4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10050 | 65.423 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| JF412297 | EBPR siphovirus 2 | EBPR siphovirus 2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48954 | 62.939 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| HQ641342 | Vibrio phage ICP3_2009_A | Vibrio phage ICP3_2009_A unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38983 | 42.839 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| HQ268735 | Streptococcus phage Dp-1 | Streptococcus phage Dp-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56506 | 40.337 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| HM596271 | Leuconostoc phage phiMH1 | Leuconostoc phage phiMH1 Leuconostoc virus MH1 Seongbukvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38709 | 38.678 | Seongbukvirus | Unclassified | Siphoviridae | Leuconostoc |
| GU071100 | Cyanophage 9515-10a | Cyanophage 9515-10a Prochlorococcus virus 951510a Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47055 | 39.197 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| GQ422451 | Enterobacteria phage Phi75 | Enterobacteria phage Phi75 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41136 | 47.469 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| GU942563 | Vibrio virus CTXphi | Vibrio virus CTXphi Affertcholeramvirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 17070 | 46.257 | Affertcholeramvirus | Unclassified | Inoviridae | Vibrio |
| GU071092 | Prochlorococcus phage P-SSM2 | Prochlorococcus phage P-SSM2 Prochlorococcus virus PSSM2 Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252407 | 35.503 | Salacisavirus | Unclassified | Myoviridae | Prochlorococcus |
| GQ153916 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 37.521 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| KX377933 | Escherichia phage vB_EcoM_Alf5 | Escherichia phage vB_EcoM_Alf5 Escherichia virus Alf5 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87662 | 39.018 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KX349903 | Bacillus phage DirtyBetty | Bacillus phage DirtyBetty unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162415 | 38.724 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KX349901 | Bacillus phage Stitch | Bacillus phage Stitch Bacillus virus Stitch Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24320 | 30.362 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KX349900 | Bacillus phage Claudi | Bacillus phage Claudi Bacillus virus Claudi Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26504 | 30.26 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KX349899 | Bacillus phage Aurora | Bacillus phage Aurora Bacillus virus Aurora Claudivirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25908 | 30.662 | Claudivirus | Northropvirinae | Salasmaviridae | Bacillus |
| KX227760 | Bacillus phage PfEFR-5 | Bacillus phage PfEFR-5 unclassified Hubeivirus Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43773 | 35.392 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| KX227759 | Bacillus phage PfIS075 | Bacillus phage PfIS075 unclassified Cecivirus Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48709 | 36.484 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| KX227758 | Bacillus phage PfNC7401 | Bacillus phage PfNC7401 unclassified Cecivirus Cecivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48055 | 36.552 | Cecivirus | Unclassified | Siphoviridae | Bacillus |
| KX227757 | Bacillus phage PfEFR-4 | Bacillus phage PfEFR-4 Bacillus virus PfEFR4 Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43223 | 35.437 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| KX190834 | Bacillus phage BMBtpLA3 | Bacillus phage BMBtpLA3 unclassified Lwoffvirus Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37385 | 37.796 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| KU848187 | Lactobacillus phage PLE2 | Lactobacillus phage PLE2 Lactobacillus virus PLE2 Pleeduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35068 | 45.326 | Pleeduovirus | Unclassified | Siphoviridae | Lactobacillus |
| KU848186 | Lactobacillus phage PLE3 | Lactobacillus phage PLE3 Lactobacillus virus PLE3 Pleetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41005 | 44.931 | Pleetrevirus | Unclassified | Siphoviridae | Lactobacillus |
| KX245890 | Citrobacter phage vB_CfrM_CfP1 | Citrobacter phage vB_CfrM_CfP1 unclassified Pseudotevenvirus Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180219 | 43.119 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| KU708005 | Pseudomonas phage phiAH14b | Pseudomonas phage phiAH14b unclassified bacterial viruses Viruses | 16812 | 58.904 | Unclassified | Unclassified | Unclassified | Pseudomonas |
| KU708004 | Pseudomonas phage phiAH14a | Pseudomonas phage phiAH14a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55060 | 58.135 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KT870145 | Roseobacter phage DSS3P8 | Roseobacter phage DSS3P8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146135 | 56.348 | Unclassified | Unclassified | Siphoviridae | Roseobacter |
| KU867307 | Salmonella phage vB_SnwM_CGG4-1 | Salmonella phage vB_SnwM_CGG4-1 unclassified Gelderlandvirus Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159878 | 37.148 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| KX266637 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266636 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266635 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266634 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266633 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266632 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266631 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266630 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266629 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266628 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266627 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266626 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266625 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266624 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266623 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266622 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266621 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266620 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266619 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266618 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266617 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266616 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266615 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266614 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266613 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266612 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266611 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266610 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266609 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266608 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266607 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266606 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266605 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266604 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266603 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266602 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266601 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266600 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266599 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266598 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266597 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266596 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266595 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266594 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266593 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266592 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266591 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266590 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266589 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266588 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266587 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266586 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266585 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266584 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266583 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266582 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266581 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266580 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266579 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266578 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266577 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266576 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266575 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266574 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266573 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266572 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266571 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266570 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266569 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266568 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266567 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266566 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266565 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266564 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266563 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266562 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266561 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266560 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266559 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266558 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266557 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266556 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266555 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266554 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266553 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266552 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266551 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266550 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266549 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266548 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266547 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266546 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266545 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266544 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266543 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266542 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266541 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266540 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266539 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266538 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266537 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266536 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266535 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266534 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266533 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266532 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266531 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266530 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266529 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266528 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266527 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266526 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266525 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266524 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266523 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266522 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266521 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266520 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266519 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266518 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266517 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266516 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266515 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266514 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266513 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266512 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266511 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266510 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266509 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266508 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266507 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266506 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266505 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266504 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266503 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266502 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266501 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266500 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266499 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266498 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266497 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266496 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266495 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266494 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.01 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266493 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266492 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266491 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266490 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266489 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266488 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266487 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266486 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266485 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266484 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266483 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266482 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266481 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266480 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266479 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266478 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266477 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266476 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266475 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266474 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266473 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266472 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266471 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266470 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266469 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266468 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266467 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266466 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266465 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266464 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266463 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266462 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266461 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266460 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266459 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266458 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266457 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266456 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266455 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266454 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266453 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266452 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266451 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266450 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266449 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266448 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266447 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266446 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266445 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266444 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266443 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266442 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266441 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266440 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266439 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266438 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266437 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.01 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266436 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266435 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266434 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266433 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266432 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266431 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266430 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266429 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266428 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266427 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266426 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266425 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266424 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266423 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266422 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266421 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266420 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.867 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266419 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266418 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266417 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266416 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266415 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266414 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266413 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266412 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266411 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266410 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266409 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266408 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266407 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266406 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266405 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266404 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266403 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266402 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266401 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266400 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266399 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266398 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266397 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266396 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.01 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266395 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266394 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266393 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266392 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266391 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266390 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266389 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266388 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266387 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266386 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266385 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266384 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266383 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266382 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266381 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266380 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266379 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266378 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266377 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266376 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266375 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266374 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266373 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266372 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266371 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266370 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266369 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266368 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266367 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266366 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266365 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266364 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266363 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266362 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266361 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266360 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266359 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266358 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266357 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266356 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266355 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266354 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266353 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266352 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266351 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266350 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266349 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266348 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266347 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266346 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266345 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266344 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266343 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.01 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266342 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266341 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266340 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266339 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266338 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266337 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266336 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266335 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266334 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266333 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266332 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266331 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266330 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266329 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266328 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266327 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266326 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266325 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266324 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266323 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266322 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266321 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266320 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.885 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266319 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266318 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266317 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266316 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266315 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.903 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266314 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266313 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266312 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266311 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266310 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.975 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266309 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266308 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266307 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266306 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266305 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266304 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266303 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.992 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266302 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266301 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.921 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KX266300 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KU687351 | Citrobacter phage SH5 | Citrobacter phage SH5 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39832 | 52.516 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| KU687350 | Citrobacter phage SH4 | Citrobacter phage SH4 Citrobacter virus SH4 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39274 | 52.587 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| KU687349 | Citrobacter phage SH3 | Citrobacter phage SH3 Citrobacter virus SH3 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39444 | 50.641 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| KU687348 | Citrobacter phage SH2 | Citrobacter phage SH2 Citrobacter virus SH2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39158 | 50.723 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| KU687347 | Citrobacter phage SH1 | Citrobacter phage SH1 Citrobacter virus SH1 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39434 | 50.953 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| KT932701 | Enterococcus phage vB_EfaS_IME196 | Enterococcus phage vB_EfaS_IME196 Enterococcus virus IME196 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38886 | 34.974 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| KT932699 | Enterococcus phage vB_EfaS_IME198 | Enterococcus phage vB_EfaS_IME198 unclassified Saphexavirus Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58000 | 40.019 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| KM879463 | Arthrobacter phage vB_ArtM-ArV1 | Arthrobacter phage vB_ArtM-ArV1 Klausavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71200 | 61.621 | Klausavirus | Unclassified | Myoviridae | Arthrobacter |
| KX196154 | Shewanella phage SFCi1 | Shewanella phage SFCi1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42279 | 59.067 | Unclassified | Unclassified | Myoviridae | Shewanella |
| KX211991 | Klebsiella phage KpV475 | Klebsiella phage KpV475 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42201 | 54.198 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KX198615 | Vibrio phage vB_VhaS-a | Vibrio phage vB_VhaS-a Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82196 | 47.294 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KX198614 | Vibrio phage vB_VhaS-tm | Vibrio phage vB_VhaS-tm Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59742 | 46.62 | Unclassified | Queuovirinae | Siphoviridae | Vibrio |
| KM366100 | Staphylococcus phage BP39 | Staphylococcus phage BP39 Staphylococcus virus BP39 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17641 | 29.018 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KX130865 | Shigella phage SHSML-52-1 | Shigella phage SHSML-52-1 Shigella virus SHSML521 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169621 | 37.576 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| KX130864 | Shigella phage SHBML-50-1 | Shigella phage SHBML-50-1 Shigella virus SHBML501 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166634 | 35.372 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KX130862 | Shigella phage SHFML-26 | Shigella phage SHFML-26 Shigella virus SHFML26 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168993 | 35.382 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KX130861 | Shigella phage SHFML-11 | Shigella phage SHFML-11 Shigella virus SHFML11 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170650 | 35.238 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KX130727 | Escherichia phage phT4A | Escherichia phage phT4A unclassified Slopekvirus Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171598 | 41.451 | Slopekvirus | Tevenvirinae | Myoviridae | Escherichia |
| KX130726 | Escherichia phage ECA2 | Escherichia phage ECA2 Escherichia virus ECA2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38890 | 50.463 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| KX245013 | Salmonella phage MA12 | Salmonella phage MA12 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41224 | 49.876 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KX245012 | Salmonella phage GG32 | Salmonella phage GG32 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157855 | 44.545 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KR704915 | Chimpanzee faeces associated microphage 3 | Chimpanzee faeces associated microphage 3 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5016 | 45.355 | Unclassified | Unclassified | Microviridae | Unspecified |
| KR704914 | Chimpanzee faeces associated microphage 2 | Chimpanzee faeces associated microphage 2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5184 | 38.31 | Unclassified | Unclassified | Microviridae | Unspecified |
| KR704913 | Chimpanzee faeces associated microphage 1 | Chimpanzee faeces associated microphage 1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4851 | 45.455 | Unclassified | Unclassified | Microviridae | Unspecified |
| KX373692 | Lactococcus phage M6654 | Lactococcus phage M6654 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22350 | 35.866 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373691 | Lactococcus phage M6653 | Lactococcus phage M6653 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22157 | 35.943 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373690 | Lactococcus phage M6202 | Lactococcus phage M6202 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22251 | 35.949 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373689 | Lactococcus phage M6165 | Lactococcus phage M6165 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22179 | 35.732 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373688 | Lactococcus phage M6162 | Lactococcus phage M6162 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21697 | 35.793 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373687 | Lactococcus phage M5938 | Lactococcus phage M5938 Lactococcus virus M5938 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22240 | 35.863 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373686 | Lactococcus phage D4412 | Lactococcus phage D4412 Lactococcus virus D4412 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22884 | 35.431 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KX373685 | Lactococcus phage D4410 | Lactococcus phage D4410 Lactococcus virus D4410 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22825 | 35.706 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| KP994390 | Pseudomonas phage YH30 | Pseudomonas phage YH30 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72192 | 54.916 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KU682439 | Stenotrophomonas phage vB_SmaS-DLP_6 | Stenotrophomonas phage vB_SmaS-DLP_6 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168489 | 55.754 | Unclassified | Unclassified | Ackermannviridae | Stenotrophomonas |
| LN898172 | Pseudomonas phage vB_PaeS_PM105 | Pseudomonas phage vB_PaeS_PM105 Pseudomonas virus PM105 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39593 | 63.1 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| LN610582 | Pseudomonas phage vB_PaeM_PAO1_Ab06 | Pseudomonas phage vB_PaeM_PAO1_Ab06 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84759 | 54.598 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| LN610581 | Pseudomonas phage vB_PaeM_PAO1_Ab04 | Pseudomonas phage vB_PaeM_PAO1_Ab04 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86668 | 54.559 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| LN610576 | Pseudomonas phage vB_PaeM_PAO1_Ab17 | Pseudomonas phage vB_PaeM_PAO1_Ab17 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83598 | 54.591 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| LN610573 | Pseudomonas phage vB_PaeM_PAO1_Ab03 | Pseudomonas phage vB_PaeM_PAO1_Ab03 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86246 | 54.677 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| HG962376 | Pseudomonas phage vB_PaeS_SCH_Ab26 | Pseudomonas phage vB_PaeS_SCH_Ab26 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43056 | 53.426 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| HG962375 | Pseudomonas phage vB_PaeP_C2-10_Ab09 | Pseudomonas phage vB_PaeP_C2-10_Ab09 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72028 | 54.897 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KU204984 | Pseudomonas phage AAT-1 | Pseudomonas phage AAT-1 unclassified Pamexvirus Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57599 | 65.894 | Pamexvirus | Unclassified | Siphoviridae | Pseudomonas |
| KT287080 | Enterobacter phage phiEap-2 | Enterobacter phage phiEap-2 unclassified Cornellvirus Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40491 | 51.972 | Cornellvirus | Guernseyvirinae | Siphoviridae | Enterobacter |
| KR905068 | Lactobacillus phage iA2 | Lactobacillus phage iA2 Lactobacillus virus iA2 Iaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34155 | 45.639 | Iaduovirus | Unclassified | Siphoviridae | Lactobacillus |
| KR905070 | Lactobacillus phage iLp1308 | Lactobacillus phage iLp1308 Lactobacillus virus iLp1308 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34176 | 45.049 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| KR905069 | Lactobacillus phage iLp84 | Lactobacillus phage iLp84 Lactobacillus virus iLp84 Pleetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39399 | 45.085 | Pleetrevirus | Unclassified | Siphoviridae | Lactobacillus |
| KR905067 | Lactobacillus phage CL2 | Lactobacillus phage CL2 Lactobacillus virus CL2 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38751 | 44.879 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| KR905066 | Lactobacillus phage CL1 | Lactobacillus phage CL1 Lactobacillus virus CL1 Colunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39474 | 44.774 | Colunavirus | Unclassified | Siphoviridae | Lactobacillus |
| JX094500 | Cronobacter phage CR5 | Cronobacter phage CR5 unclassified Derbicusvirus Derbicusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223989 | 50.057 | Derbicusvirus | Unclassified | Myoviridae | Cronobacter |
| KX078569 | Morganella phage vB_MmoM_MP1 | Morganella phage vB_MmoM_MP1 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163201 | 34.747 | Unclassified | Tevenvirinae | Myoviridae | Morganella |
| KX078568 | Morganella phage vB_MmoP_MP2 | Morganella phage vB_MmoP_MP2 Morganella virus MP2 Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39616 | 46.913 | Minipunavirus | Studiervirinae | Autographiviridae | Morganella |
| KX130668 | Escherichia phage vB_EcoS_NBD2 | Escherichia phage vB_EcoS_NBD2 Escherichia virus NBD2 Vilniusvirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51802 | 49.817 | Vilniusvirus | Unclassified | Drexlerviridae | Escherichia |
| KU927499 | Escherichia phage 64795_ec1 | Escherichia phage 64795_ec1 Escherichia virus 64795ec1 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39235 | 48.658 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| KU927498 | Salmonella phage 64795_sal4 | Salmonella phage 64795_sal4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42134 | 46.926 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU927496 | Salmonella phage 101962B_sal5 | Salmonella phage 101962B_sal5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42126 | 46.914 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU927495 | Salmonella phage 103203_sal4 | Salmonella phage 103203_sal4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42428 | 46.816 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU927494 | Salmonella phage 103203_sal5 | Salmonella phage 103203_sal5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40443 | 46.525 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU927492 | Salmonella phage 146851_sal4 | Salmonella phage 146851_sal4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42154 | 46.923 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU927491 | Salmonella phage 146851_sal5 | Salmonella phage 146851_sal5 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42850 | 46.751 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU521356 | Pseudomonas phage KTN4 | Pseudomonas phage KTN4 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279593 | 36.893 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| KU234533 | Nostoc phage A1 | Nostoc phage A1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68304 | 36.5 | Unclassified | Unclassified | Myoviridae | Nostoc |
| KU234532 | Nostoc phage N1 | Nostoc phage N1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64960 | 35.445 | Unclassified | Unclassified | Myoviridae | Nostoc |
| KU678392 | Streptococcus virus 9874 | Streptococcus virus 9874 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32649 | 36.617 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KU678391 | Streptococcus virus 9873 | Streptococcus virus 9873 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32813 | 36.9 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KU678390 | Streptococcus virus 9872 | Streptococcus virus 9872 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33105 | 36.84 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KU678389 | Streptococcus virus 9871 | Streptococcus virus 9871 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32729 | 37.059 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KU577463 | Bacillus phage Deep Blue | Bacillus phage Deep Blue Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157501 | 39.947 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT716399 | Pseudomonas phage POR1 | Pseudomonas phage POR1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55349 | 55.506 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| KT734862 | Pseudomonas phage PAE1 | Pseudomonas phage PAE1 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62181 | 64.233 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| KR997933 | Mycobacterium phage Madruga | Mycobacterium phage Madruga Patiencevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69377 | 50.406 | Patiencevirus | Unclassified | Siphoviridae | Mycobacterium |
| KU948710 | Pseudomonas phage PEV2 | Pseudomonas phage PEV2 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72697 | 54.863 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KU885990 | Roseobacter phage RD-1410Ws-07 | Roseobacter phage RD-1410Ws-07 unclassified Sanyabayvirus Sanyabayvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76298 | 50.008 | Sanyabayvirus | Rhodovirinae | Schitoviridae | Roseobacter |
| KU885989 | Roseobacter phage RD-1410W1-01 | Roseobacter phage RD-1410W1-01 Roseobacter virus RD1410W1-01 Aoqinvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72674 | 49.527 | Aoqinvirus | Rhodovirinae | Schitoviridae | Roseobacter |
| KU885988 | Dinoroseobacter phage DS-1410Ws-06 | Dinoroseobacter phage DS-1410Ws-06 Dinoroseobacter virus DS1410Ws06 Sanyabayvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76466 | 50.005 | Sanyabayvirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| KP090454 | Escherichia phage P694 | Escherichia phage P694 Escherichia virus P694 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40477 | 48.42 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| KP090453 | Escherichia phage P483 | Escherichia phage P483 Escherichia virus P483 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40829 | 48.444 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| KR698074 | Escherichia phage APECc02 | Escherichia phage APECc02 Escherichia virus APECc02 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135400 | 43.592 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KR422353 | Escherichia phage phiAPCEc03 | Escherichia phage phiAPCEc03 Escherichia virus phiAPCEc03 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103737 | 38.933 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KR422352 | Escherichia phage APCEc01 | Escherichia phage APCEc01 Escherichia virus APCEc01 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168771 | 37.707 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KT253150 | Rhodobacter phage RcSaxon | Rhodobacter phage RcSaxon Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36081 | 65.442 | Cronusvirus | Unclassified | Siphoviridae | Rhodobacter |
| KR935217 | Rhodobacter phage RcCronus | Rhodobacter phage RcCronus Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35985 | 65.427 | Cronusvirus | Unclassified | Siphoviridae | Rhodobacter |
| KR935216 | Rhodobacter phage RcRhea | Rhodobacter phage RcRhea Cronusvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36065 | 65.446 | Cronusvirus | Unclassified | Siphoviridae | Rhodobacter |
| KR935215 | Rhodobacter phage RcSpartan | Rhodobacter phage RcSpartan Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44194 | 54.885 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| KR935213 | Rhodobacter phage RcTitan | Rhodobacter phage RcTitan Titanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44496 | 55.14 | Titanvirus | Unclassified | Siphoviridae | Rhodobacter |
| KF626667 | Xylella phage Prado | Xylella phage Prado Pradovirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43940 | 63.022 | Pradovirus | Unclassified | Autographiviridae | Xylella |
| KF626666 | Xylella phage Paz | Xylella phage Paz unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43869 | 60.168 | Unclassified | Okabevirinae | Autographiviridae | Xylella |
| AP014629 | Edwardsiella phage GF-2 | Edwardsiella phage GF-2 Edwardsiella virus GF2 Gofduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43129 | 51.269 | Gofduovirus | Unclassified | Myoviridae | Edwardsiella |
| KC688701 | Croceibacter phage P2559Y | Croceibacter phage P2559Y Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43153 | 38.922 | Unclassified | Unclassified | Siphoviridae | Croceibacter |
| KX066068 | Pseudomonas phage VSW-3 | Pseudomonas phage VSW-3 Pseudomonas virus VSW3 Napahaivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40556 | 57.54 | Napahaivirus | Unclassified | Autographiviridae | Pseudomonas |
| KX017521 | Salmonella phage 118970_sal2 | Salmonella phage 118970_sal2 Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114180 | 40.278 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KX017520 | Salmonella phage 64795_sal3 | Salmonella phage 64795_sal3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45342 | 45.596 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| KU992911 | Staphylococcus phage SLPW | Staphylococcus phage SLPW Staphylococcus virus SLPW Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17861 | 29.354 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KU984980 | Propionibacterium phage PFR2 | Propionibacterium phage PFR2 unclassified Pulverervirus Pulverervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39640 | 64.955 | Pulverervirus | Unclassified | Siphoviridae | Propionibacterium |
| KU984979 | Propionibacterium phage PFR1 | Propionibacterium phage PFR1 Pulverervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38071 | 64.842 | Pulverervirus | Unclassified | Siphoviridae | Propionibacterium |
| KU504502 | Vibrio phage 919TP | Vibrio phage 919TP unclassified Longwoodvirus Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33133 | 48.921 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| KT895374 | Bacillus phage vB_BpuM-BpSp | Bacillus phage vB_BpuM-BpSp unclassified Takahashivirus Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 255569 | 25.885 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| KU878088 | Bacillus phage AR9 | Bacillus phage AR9 unclassified Takahashivirus Takahashivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 251042 | 27.752 | Takahashivirus | Unclassified | Myoviridae | Bacillus |
| KR060090 | Pseudoalteromonas phage Pq0 | Pseudoalteromonas phage Pq0 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33399 | 40.286 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KX058879 | Vibrio phage VcP032 | Vibrio phage VcP032 Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33108 | 48.858 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| KR296686 | Salmonella phage 22 | Salmonella phage 22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41733 | 47.116 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KR296687 | Salmonella phage 25 | Salmonella phage 25 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41803 | 47.064 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KR296688 | Salmonella phage 34 | Salmonella phage 34 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41783 | 47.079 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KR296689 | Salmonella phage 35 | Salmonella phage 35 Salmonella virus 35 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55391 | 56.845 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KR296690 | Salmonella phage 36 | Salmonella phage 36 Salmonella virus 36 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41085 | 43.522 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| KR296691 | Salmonella phage 37 | Salmonella phage 37 Salmonella virus 37 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60216 | 56.527 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KR296692 | Salmonella phage 38 | Salmonella phage 38 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156833 | 44.616 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KR296694 | Salmonella phage 40 | Salmonella phage 40 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63263 | 44.939 | Unclassified | Vequintavirinae | Myoviridae | Salmonella |
| KR091942 | Salmonella phage 18-India | Salmonella phage 18-India Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33133 | 44.373 | Unclassified | Unclassified | Myoviridae | Salmonella |
| KP881332 | Staphylococcus phage Stau2 | Staphylococcus phage Stau2 Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133798 | 29.974 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ746502 | Rhizobium phage vB_RleM_PPF1 | Rhizobium phage vB_RleM_PPF1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54506 | 61.9 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| KU892558 | Lactococcus phage PLgT-1 | Lactococcus phage PLgT-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40273 | 35.401 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KR052143 | Pseudomonas phage vB_PaeM_MAG1 | Pseudomonas phage vB_PaeM_MAG1 Pseudomonas virus MAG1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94555 | 49.248 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KR052142 | Pseudomonas phage vB_PaeP_MAG4 | Pseudomonas phage vB_PaeP_MAG4 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72979 | 54.82 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KP085586 | Shigella phage pSf-2 | Shigella phage pSf-2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50109 | 45.443 | Tunavirus | Tunavirinae | Drexlerviridae | Shigella |
| JN225449 | Enterobacteriaphage UAB_Phi87 | Enterobacteriaphage UAB_Phi87 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87603 | 38.887 | Felixounavirus | Ounavirinae | Myoviridae | Enterobacteria |
| KU878969 | Escherichia phage MX01 | Escherichia phage MX01 Escherichia virus MX01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168929 | 39.564 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| KU878968 | Escherichia phage WG01 | Escherichia phage WG01 Escherichia virus WG01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169936 | 39.623 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| KX129925 | Pseudomonas phage NP1 | Pseudomonas phage NP1 Nipunavirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58566 | 58.399 | Nipunavirus | Queuovirinae | Siphoviridae | Pseudomonas |
| LC102730 | Pseudomonas phage S12-1 | Pseudomonas phage S12-1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66257 | 55.581 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC102729 | Pseudomonas phage phiR18 | Pseudomonas phage phiR18 Kochitakasuvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63560 | 60.349 | Kochitakasuvirus | Unclassified | Podoviridae | Pseudomonas |
| GQ422450 | Enterobacteria phage UAB_Phi20 | Enterobacteria phage UAB_Phi20 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41809 | 47.241 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| AB775549 | Pseudomonas phage PPpW-4 | Pseudomonas phage PPpW-4 Pseudomonas virus PPpW4 Phutvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41386 | 56.775 | Phutvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| AB775548 | Pseudomonas phage PPpW-3 | Pseudomonas phage PPpW-3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43564 | 61.11 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| KT780304 | Bacillus phage VMY22 | Bacillus phage VMY22 Bacillus virus VMY22 Mingyongvirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18609 | 36.364 | Mingyongvirus | Unclassified | Salasmaviridae | Bacillus |
| KM242061 | Escherichia phage CICC 80001 | Escherichia phage CICC 80001 Escherichia virus CICC80001 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38810 | 48.82 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| KU194206 | Escherichia phage JMPW1 | Escherichia phage JMPW1 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49628 | 45.559 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| KU194205 | Escherichia phage JMPW2 | Escherichia phage JMPW2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50298 | 45.376 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| KT876726 | Flavobacterium phage FpV21 | Flavobacterium phage FpV21 unclassified Fipvunavirus Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89748 | 26.948 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KT876725 | Flavobacterium phage FpV9 | Flavobacterium phage FpV9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42696 | 32.008 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KT876724 | Flavobacterium phage FpV4 | Flavobacterium phage FpV4 Flavobacterium virus Fpv4 Fipvunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88308 | 27.218 | Fipvunavirus | Unclassified | Podoviridae | Flavobacterium |
| KT339321 | Acinetobacter phage phiAB6 | Acinetobacter phage phiAB6 Acinetobacter virus phiAB6 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40570 | 39.468 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KP671755 | Vibrio phage ValKK3 | Vibrio phage ValKK3 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 248088 | 41.235 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| KC862299 | Pseudomonas phage PAK_P3 | Pseudomonas phage PAK_P3 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88097 | 54.827 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KU253712 | Bacillus phage Eldridge | Bacillus phage Eldridge Bacillus virus Eldridge Eldridgevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165868 | 40.407 | Eldridgevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT725776 | Bacillus phage vB_BceS-MY192 | Bacillus phage vB_BceS-MY192 Bacillus virus MY192 Hubeivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44696 | 34.979 | Hubeivirus | Unclassified | Siphoviridae | Bacillus |
| KU998233 | Gordonia phage BritBrat | Gordonia phage BritBrat Gordonia virus Britbrat Britbratvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55524 | 64.999 | Britbratvirus | Unclassified | Siphoviridae | Gordonia |
| KR053196 | Tsukamurella phage TPA4 | Tsukamurella phage TPA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56212 | 70.542 | Unclassified | Unclassified | Siphoviridae | Tsukamurella |
| KR053195 | Gordonia phage GMA1 | Gordonia phage GMA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41207 | 65.712 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR053194 | Gordonia phage GAL1 | Gordonia phage GAL1 Gordonia virus GAL1 Galunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49979 | 63.547 | Galunavirus | Unclassified | Siphoviridae | Gordonia |
| KR233165 | Escherichia phage PEC04 | Escherichia phage PEC04 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167552 | 35.472 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KR233164 | Salmonella phage NR01 | Salmonella phage NR01 Salmonella virus NR01 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111325 | 38.838 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KR269720 | Klebsiella phage PKO111 | Klebsiella phage PKO111 Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168758 | 39.394 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| KR269719 | Klebsiella phage PKP126 | Klebsiella phage PKP126 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50934 | 50.73 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KR269718 | Escherichia phage HY03 | Escherichia phage HY03 Escherichia virus HY03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170770 | 35.302 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN889993 | Bacteriophage 4A | Bacteriophage 4A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48214 | 65.081 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LN889992 | Pseudomonas phage 15b | Pseudomonas phage 15b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48452 | 49.628 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| LN889991 | Bacteriophage 15G | Bacteriophage 15G Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48452 | 49.602 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| LN889756 | Pseudomonas phage shl2 | Pseudomonas phage shl2 Pseudomonas virus shl2 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40466 | 56.606 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KU867876 | Escherichia phage vB_EcoM-UFV13 | Escherichia phage vB_EcoM-UFV13 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165772 | 35.537 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KU878967 | Salmonella phage 118970_sal4 | Salmonella phage 118970_sal4 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42418 | 46.813 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KU951147 | Salmonella phage f3SE | Salmonella phage f3SE Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41867 | 49.798 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KU951146 | Salmonella phage f2SE | Salmonella phage f2SE Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41865 | 49.796 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KX015771 | Salmonella phage phSE-5 | Salmonella phage phSE-5 unclassified Tlsvirus Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49178 | 42.881 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| KX015770 | Salmonella phage phSE-2 | Salmonella phage phSE-2 Salmonella virus phSE2 Tlsvirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49167 | 42.889 | Tlsvirus | Tempevirinae | Drexlerviridae | Salmonella |
| KR029087 | Mycobacterium phage PDRPxv | Mycobacterium phage PDRPxv Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69171 | 66.347 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KR029086 | Mycobacterium phage PDRPv | Mycobacterium phage PDRPv Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69110 | 66.351 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KT160311 | Vibrio phage H188 | Vibrio phage H188 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50364 | 43.634 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KF188409 | Lactobacillus phage phiJB | Lactobacillus phage phiJB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36969 | 47.702 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| KT321315 | Enterobacter phage phiEap-3 | Enterobacter phage phiEap-3 Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175814 | 42.005 | Slopekvirus | Tevenvirinae | Myoviridae | Enterobacter |
| KU737352 | Bacillus phage Nemo | Bacillus phage Nemo unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162375 | 38.787 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737351 | Bacillus phage NotTheCreek | Bacillus phage NotTheCreek unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161929 | 38.711 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737349 | Bacillus phage DIGNKC | Bacillus phage DIGNKC unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161552 | 38.67 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737348 | Bacillus phage Zuko | Bacillus phage Zuko unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163345 | 38.63 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737347 | Bacillus phage Phrodo | Bacillus phage Phrodo unclassified Bequatrovirus Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164443 | 37.802 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737346 | Bacillus phage Vinny | Bacillus phage Vinny unclassified Bastillevirus Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162865 | 38 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737345 | Bacillus phage Juglone | Bacillus phage Juglone Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164227 | 37.786 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU737344 | Bacillus phage Nigalana | Bacillus phage Nigalana unclassified Wphvirus Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163041 | 38.75 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| AP014927 | Ralstonia phage RSF1 | Ralstonia phage RSF1 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222888 | 52.259 | Chiangmaivirus | Unclassified | Myoviridae | Ralstonia |
| AP014693 | Ralstonia phage RSL2 | Ralstonia phage RSL2 Chiangmaivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223932 | 52.057 | Chiangmaivirus | Unclassified | Myoviridae | Ralstonia |
| KU598975 | Staphylococcus phage CNPx | Staphylococcus phage CNPx Staphylococcus virus CNPx Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43293 | 34.661 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| KR902987 | Propionibacterium phage PAC10 | Propionibacterium phage PAC10 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29704 | 54.036 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902986 | Propionibacterium phage PAC9 | Propionibacterium phage PAC9 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29786 | 54.039 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902985 | Propionibacterium phage PAC8 | Propionibacterium phage PAC8 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29488 | 54.032 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902984 | Propionibacterium phage PAC7 | Propionibacterium phage PAC7 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29551 | 54.066 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902983 | Propionibacterium phage PAC6 | Propionibacterium phage PAC6 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29609 | 54.058 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902982 | Propionibacterium phage PAC5 | Propionibacterium phage PAC5 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29428 | 54.169 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902981 | Propionibacterium phage PAC4 | Propionibacterium phage PAC4 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29581 | 54.041 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902980 | Propionibacterium phage PAC3 | Propionibacterium phage PAC3 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29526 | 54.027 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902979 | Propionibacterium phage PAC2 | Propionibacterium phage PAC2 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29602 | 54.023 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KR902978 | Propionibacterium phage PAC1 | Propionibacterium phage PAC1 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29605 | 54.042 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KP064094 | Escherichia phage CAjan | Escherichia phage CAjan Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59670 | 44.708 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| KP836356 | Marinitoga camini virus 2 | Marinitoga camini virus 2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50307 | 28.732 | Unclassified | Unclassified | Siphoviridae | Marinitoga |
| JQ015307 | Pectobacterium phage phiTE | Pectobacterium phage phiTE Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142349 | 50.137 | Certrevirus | Vequintavirinae | Myoviridae | Pectobacterium |
| JF712866 | Vibrio phage phiVC8 | Vibrio phage phiVC8 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39422 | 50.807 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KU726251 | Enterobacteria phage SEGD1 | Enterobacteria phage SEGD1 Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 239461 | 48.512 | Seoulvirus | Unclassified | Myoviridae | Enterobacteria |
| KU831549 | Bacillus phage vB_BsuP-Goe1 | Bacillus phage vB_BsuP-Goe1 Bacillus virus Goe1 Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18379 | 37.494 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| KU130131 | Pseudomonas phage vB_PsyM_KIL3b | Pseudomonas phage vB_PsyM_KIL3b Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92095 | 44.721 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU130130 | Pseudomonas phage vB_PsyM_KIL5 | Pseudomonas phage vB_PsyM_KIL5 Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93385 | 44.971 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU130129 | Pseudomonas phage vB_PsyM_KIL4 | Pseudomonas phage vB_PsyM_KIL4 Pseudomonas virus KIL2 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92816 | 44.897 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU130128 | Pseudomonas phage vB_PsyM_KIL3 | Pseudomonas phage vB_PsyM_KIL3 Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92068 | 44.727 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU130127 | Pseudomonas phage vB_PsyM_KIL2 | Pseudomonas phage vB_PsyM_KIL2 Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92466 | 44.786 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU130126 | Pseudomonas phage vB_PsyM_KIL1 | Pseudomonas phage vB_PsyM_KIL1 Pseudomonas virus KIL4 Flaumdravirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90552 | 44.811 | Flaumdravirus | Unclassified | Myoviridae | Pseudomonas |
| KU198331 | Pseudomonas phage NP3 | Pseudomonas phage NP3 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66063 | 55.412 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KM209320 | Dickeya phage phiDP23.1 | Dickeya phage phiDP23.1 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 150933 | 49.329 | Unclassified | Aglimvirinae | Ackermannviridae | Dickeya |
| KM209294 | Dickeya phage phiDP23.1 | Dickeya phage phiDP23.1 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 12099 | 50.459 | Unclassified | Aglimvirinae | Ackermannviridae | Dickeya |
| KM209271 | Dickeya phage phiDP10.3 | Dickeya phage phiDP10.3 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27033 | 48.53 | Unclassified | Aglimvirinae | Ackermannviridae | Dickeya |
| KM209270 | Dickeya phage phiDP10.3 | Dickeya phage phiDP10.3 unclassified Aglimvirinae Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129650 | 49.766 | Unclassified | Aglimvirinae | Ackermannviridae | Dickeya |
| LN881737 | Escherichia phage slur13 | Escherichia phage slur13 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167299 | 35.514 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881726 | Escherichia phage slur02 | Escherichia phage slur02 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167298 | 35.516 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881725 | Escherichia phage slur01 | Escherichia phage slur01 unclassified Seuratvirus Seuratvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61235 | 44.429 | Seuratvirus | Queuovirinae | Siphoviridae | Escherichia |
| LN881736 | Escherichia phage slur14 | Escherichia phage slur14 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167467 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881735 | Escherichia phage slur12 | Escherichia phage slur12 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136898 | 43.617 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| LN881734 | Escherichia phage slur11 | Escherichia phage slur11 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167298 | 35.509 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881733 | Escherichia phage slur08 | Escherichia phage slur08 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167467 | 35.405 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881732 | Escherichia phage slur07 | Escherichia phage slur07 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167124 | 35.429 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881731 | Escherichia phage slur06 | Escherichia phage slur06 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44515 | 54.534 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LN881730 | Escherichia phage slur05 | Escherichia phage slur05 unclassified Dhillonvirus Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43900 | 54.544 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LN881729 | Escherichia phage slur04 | Escherichia phage slur04 Escherichia virus slur04 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167298 | 35.505 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881728 | Escherichia phage slur03 | Escherichia phage slur03 Escherichia virus slur03 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167467 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| LN881727 | Escherichia phage slur16 | Escherichia phage slur16 Escherichia virus slur16 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136896 | 43.613 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KP836355 | Marinitoga camini virus 1 | Marinitoga camini virus 1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53412 | 28.065 | Unclassified | Unclassified | Siphoviridae | Marinitoga |
| KC357596 | Vibrio phage VFJ | Vibrio phage VFJ Saetivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8555 | 44.29 | Saetivirus | Unclassified | Inoviridae | Vibrio |
| KU665491 | Bacillus phage Mgbh1 | Bacillus phage Mgbh1 Bacillus virus Mgbh1 Magadivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58951 | 45.414 | Magadivirus | Unclassified | Siphoviridae | Bacillus |
| KU640380 | Bacillus phage Shbh1 | Bacillus phage Shbh1 Bacillus virus Shbh1 Shalavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138081 | 36.428 | Shalavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KU666550 | Klebsiella phage KpV71 | Klebsiella phage KpV71 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43267 | 53.979 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KU761955 | Pseudomonas phage phiMK | Pseudomonas phage phiMK Pseudomonas virus phiMK Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93129 | 49.288 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KU743887 | Pseudomonas phage phiNFS | Pseudomonas phage phiNFS Pseudomonas virus NFS Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42351 | 62.263 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KT755656 | Paenibacillus phage Tripp | Paenibacillus phage Tripp Paenibacillus virus Tripp Halcyonevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54439 | 48.315 | Halcyonevirus | Unclassified | Siphoviridae | Paenibacillus |
| KM501444 | Shigella phage pSs-1 | Shigella phage pSs-1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164999 | 35.544 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| KM366099 | Salmonella phage BP63 | Salmonella phage BP63 Salmonella virus BP63 Rosemountvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52437 | 46.021 | Rosemountvirus | Unclassified | Myoviridae | Salmonella |
| KM366098 | Salmonella phage BP12C | Salmonella phage BP12C Salmonella virus BP12C Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60606 | 56.395 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KM366097 | Salmonella phage BP12B | Salmonella phage BP12B Salmonella virus BP12B Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43602 | 47.557 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| KM366096 | Salmonella phage BP12A | Salmonella phage BP12A Salmonella virus BP12A Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39696 | 48.952 | Berlinvirus | Studiervirinae | Autographiviridae | Salmonella |
| KR054033 | Pseudomonas phage DL68 | Pseudomonas phage DL68 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66111 | 55.724 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KR054032 | Pseudomonas phage DL64 | Pseudomonas phage DL64 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72378 | 54.953 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KR054031 | Pseudomonas phage DL62 | Pseudomonas phage DL62 Pseudomonas virus DL62 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42508 | 62.195 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KR054030 | Pseudomonas phage DL60 | Pseudomonas phage DL60 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66103 | 54.917 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KR054029 | Pseudomonas phage DL54 | Pseudomonas phage DL54 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45673 | 52.394 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KR054028 | Pseudomonas phage DL52 | Pseudomonas phage DL52 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65867 | 54.903 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT008108 | Yersinia phage phiYe-F10 | Yersinia phage phiYe-F10 Yersinia virus YeF10 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39210 | 50.699 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| KT736033 | Pseudomonas phage K8 | Pseudomonas phage K8 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93879 | 49.355 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT337359 | Streptococcus phage phiARI0468b-3 | Streptococcus phage phiARI0468b-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31936 | 39.764 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337362 | Streptococcus phage phiARI0598b | Streptococcus phage phiARI0598b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35788 | 40.097 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337364 | Streptococcus phage phiARI0639b | Streptococcus phage phiARI0639b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32821 | 37.832 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337339 | Streptococcus phage phiARI0004 | Streptococcus phage phiARI0004 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41105 | 40.659 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337340 | Streptococcus phage phiARI0031 | Streptococcus phage phiARI0031 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41887 | 41.208 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337341 | Streptococcus phage phiARI0131-1 | Streptococcus phage phiARI0131-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42927 | 39.597 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337342 | Streptococcus phage phiARI0131-2 | Streptococcus phage phiARI0131-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34563 | 40.187 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337352 | Streptococcus phage phiARI0460-1 | Streptococcus phage phiARI0460-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41807 | 39.69 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337354 | Streptococcus phage phiARI0462 | Streptococcus phage phiARI0462 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40498 | 40.543 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337355 | Streptococcus phage phiARI0468-1 | Streptococcus phage phiARI0468-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41063 | 40.806 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337356 | Streptococcus phage phiARI0468-2 | Streptococcus phage phiARI0468-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42186 | 40.362 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337358 | Streptococcus phage phiARI0468-4 | Streptococcus phage phiARI0468-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37395 | 38.131 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337365 | Streptococcus phage phiARI0746 | Streptococcus phage phiARI0746 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33181 | 39.586 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT337370 | Streptococcus phage phiARI0923 | Streptococcus phage phiARI0923 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33477 | 38.952 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KU310944 | Pseudomonas phage YMC11/06/C171_PPU_BP | Pseudomonas phage YMC11/06/C171_PPU_BP Pseudomonas virus C171 Kantovirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45417 | 60.354 | Kantovirus | Corkvirinae | Autographiviridae | Pseudomonas |
| KU310943 | Pseudomonas phage YMC11/07/P54_PAE_BP | Pseudomonas phage YMC11/07/P54_PAE_BP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50183 | 62.18 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KU535860 | Pseudomonas phage O4 | Pseudomonas phage O4 unclassified Paundecimvirus Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50509 | 44.568 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| KU594607 | Cyanophage S-RIM44 | Cyanophage S-RIM44 Synechococcus virus SRIM44 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192999 | 40.625 | Vellamovirus | Unclassified | Myoviridae | Synechococcus |
| KU594606 | Cyanophage S-RIM32 | Cyanophage S-RIM32 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194437 | 39.868 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KU594605 | Cyanophage S-RIM50 | Cyanophage S-RIM50 Synechococcus virus SRIM50 Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174307 | 40.281 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| KT361657 | Paenibacillus phage Diane | Paenibacillus phage Diane Vegasvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45653 | 43.649 | Vegasvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361656 | Paenibacillus phage Vadim | Paenibacillus phage Vadim Vegasvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45653 | 43.653 | Vegasvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361655 | Paenibacillus phage Hayley | Paenibacillus phage Hayley Vegasvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44256 | 43.459 | Vegasvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361654 | Paenibacillus phage Vegas | Paenibacillus phage Vegas Vegasvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45653 | 43.642 | Vegasvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361653 | Paenibacillus phage Paisley | Paenibacillus phage Paisley Harrisonvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44172 | 40.043 | Harrisonvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361652 | Paenibacillus phage Xenia | Paenibacillus phage Xenia unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 41.525 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KT361651 | Paenibacillus phage Harrison | Paenibacillus phage Harrison Harrisonvirus Gochnauervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44249 | 40.157 | Harrisonvirus | Gochnauervirinae | Siphoviridae | Paenibacillus |
| KT361650 | Paenibacillus phage Willow | Paenibacillus phage Willow unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37994 | 41.857 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KT361649 | Paenibacillus phage Fern | Paenibacillus phage Fern Paenibacillus virus Fern Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37995 | 41.858 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KP313532 | Achromobacter phage phiAxp-1 | Achromobacter phage phiAxp-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45045 | 56.017 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| KU522583 | Enterobacteria phage ECGD1 | Enterobacteria phage ECGD1 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146647 | 37.473 | Unclassified | Vequintavirinae | Myoviridae | Enterobacteria |
| KP793119 | Lactococcus phage 936 group phage Phi10.5 | Lactococcus phage 936 group phage Phi10.5 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31805 | 34.84 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793111 | Lactococcus phage 936 group phage Phi19.2 | Lactococcus phage 936 group phage Phi19.2 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29984 | 34.735 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793135 | Lactococcus phage 936 group phage Phi.16 | Lactococcus phage 936 group phage Phi.16 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31561 | 34.834 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793134 | Lactococcus phage 936 group phage PhiS0139 | Lactococcus phage 936 group phage PhiS0139 Lactococcus virus PhiS0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30116 | 35.197 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793133 | Lactococcus phage 936 group phage PhiLj | Lactococcus phage 936 group phage PhiLj Lactococcus virus PhiLj Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28983 | 35.138 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793132 | Lactococcus phage 936 group phage PhiM1127 | Lactococcus phage 936 group phage PhiM1127 Lactococcus virus PhiM1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32682 | 34.79 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793131 | Lactococcus phage 936 group phage PhiE1127 | Lactococcus phage 936 group phage PhiE1127 Lactococcus virus PhiE1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32270 | 34.62 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793130 | Lactococcus phage 936 group phage Phi155 | Lactococcus phage 936 group phage Phi155 Lactococcus virus Phi155 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32150 | 34.936 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793129 | Lactococcus phage 936 group phage PhiJF1 | Lactococcus phage 936 group phage PhiJF1 Lactococcus virus PhiJF1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31901 | 34.579 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793128 | Lactococcus phage 936 group phage PhiM.16 | Lactococcus phage 936 group phage PhiM.16 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31869 | 34.956 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793127 | Lactococcus phage 936 group phage Phi40 | Lactococcus phage 936 group phage Phi40 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31631 | 34.874 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793126 | Lactococcus phage 936 group phage PhiM.5 | Lactococcus phage 936 group phage PhiM.5 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31498 | 34.824 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793125 | Lactococcus phage 936 group phage Phi91127 | Lactococcus phage 936 group phage Phi91127 Lactococcus virus Phi91127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31495 | 34.821 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793124 | Lactococcus phage 936 group phage Phi44 | Lactococcus phage 936 group phage Phi44 Lactococcus virus Phi44 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31379 | 34.622 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793123 | Lactococcus phage 936 group phage Phi4.2 | Lactococcus phage 936 group phage Phi4.2 Lactococcus virus Phi42 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31375 | 34.489 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793122 | Lactococcus phage 936 group phage PhiL.6 | Lactococcus phage 936 group phage PhiL.6 Lactococcus virus PhiL6 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31181 | 34.55 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793121 | Lactococcus phage 936 group phage Phi109 | Lactococcus phage 936 group phage Phi109 Lactococcus virus Phi109 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31033 | 34.921 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793120 | Lactococcus phage 936 group phage PhiL.18 | Lactococcus phage 936 group phage PhiL.18 Lactococcus virus PhiL18 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30882 | 35.007 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793118 | Lactococcus phage 936 group phage PhiF0139 | Lactococcus phage 936 group phage PhiF0139 Lactococcus virus PhiF0139 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30811 | 34.669 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793117 | Lactococcus phage 936 group phage PhiG | Lactococcus phage 936 group phage PhiG unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30594 | 34.618 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793116 | Lactococcus phage 936 group phage Phi13.16 | Lactococcus phage 936 group phage Phi13.16 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30577 | 34.67 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793115 | Lactococcus phage 936 group phage Phi114 | Lactococcus phage 936 group phage Phi114 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30532 | 34.682 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793114 | Lactococcus phage 936 group phage Phi17 | Lactococcus phage 936 group phage Phi17 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30463 | 34.8 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793113 | Lactococcus phage 936 group phage PhiF.17 | Lactococcus phage 936 group phage PhiF.17 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30389 | 34.713 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793112 | Lactococcus phage 936 group phage Phi129 | Lactococcus phage 936 group phage Phi129 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30178 | 35.032 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793110 | Lactococcus phage 936 group phage Phi43 | Lactococcus phage 936 group phage Phi43 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29416 | 35.073 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793109 | Lactococcus phage 936 group phage PhiC0139 | Lactococcus phage 936 group phage PhiC0139 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29032 | 35.364 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793108 | Lactococcus phage 936 group phage Phi5.12 | Lactococcus phage 936 group phage Phi5.12 Lactococcus virus Phi512 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29026 | 34.745 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793107 | Lactococcus phage 936 group phage PhiD.18 | Lactococcus phage 936 group phage PhiD.18 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28833 | 35.31 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793106 | Lactococcus phage 936 group phage PhiA1127 | Lactococcus phage 936 group phage PhiA1127 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28873 | 35.171 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793105 | Lactococcus phage 936 group phage Phi19.3 | Lactococcus phage 936 group phage Phi19.3 Lactococcus virus Phi193 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28811 | 34.792 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793104 | Lactococcus phage 936 group phage PhiB1127 | Lactococcus phage 936 group phage PhiB1127 Lactococcus virus PhiB1127 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28785 | 35.459 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793103 | Lactococcus phage 936 group phage Phi19 | Lactococcus phage 936 group phage Phi19 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28437 | 35.25 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793102 | Lactococcus phage 936 group phage PhiA.16 | Lactococcus phage 936 group phage PhiA.16 Lactococcus virus PhiA16 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28349 | 35.246 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KP793101 | Lactococcus phage 936 group phage Phi4 | Lactococcus phage 936 group phage Phi4 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28764 | 35.141 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KT337360 | Streptococcus phage phiARI0578 | Streptococcus phage phiARI0578 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28706 | 39.81 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KJ628499 | Acinetobacter phage vB_AbaM_phiAbaA1 | Acinetobacter phage vB_AbaM_phiAbaA1 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104906 | 37.774 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| KC595513 | Bacillus phage Shanette | Bacillus phage Shanette Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138877 | 40.77 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| KC595512 | Bacillus phage JL | Bacillus phage JL Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137918 | 40.802 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| KC595511 | Bacillus phage Basilisk | Bacillus phage Basilisk Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82008 | 33.906 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KT321317 | Achromobacter phage phiAxp-3 | Achromobacter phage phiAxp-3 Jwalphavirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72825 | 55.202 | Jwalphavirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| KT724718 | Brucella phage BiPBO1 | Brucella phage BiPBO1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46877 | 53.32 | Unclassified | Unclassified | Siphoviridae | Brucella |
| KU645528 | Acidianus tailed spindle virus | Acidianus tailed spindle virus unclassified Bicaudaviridae Bicaudaviridae Viruses | 70812 | 36.41 | Unclassified | Unclassified | Bicaudaviridae | Acidianus |
| KU298437 | Escherichia phage phiON-2011 | Escherichia phage phiON-2011 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60897 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| KU497559 | Pseudomonas phage K5 | Pseudomonas phage K5 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93754 | 49.353 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KU199710 | Pseudomonas phage JBD44 | Pseudomonas phage JBD44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49033 | 58.493 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KU199709 | Pseudomonas phage JBD93 | Pseudomonas phage JBD93 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36629 | 64.293 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KU199708 | Pseudomonas phage JBD69 | Pseudomonas phage JBD69 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36938 | 63.931 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KU199707 | Pseudomonas phage JBD16C | Pseudomonas phage JBD16C unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36436 | 63.811 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| HG813241 | Cronobacter phage Dev2 | Cronobacter phage Dev2 Cronobacter virus Dev2 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38966 | 52.559 | Kayfunavirus | Studiervirinae | Autographiviridae | Cronobacter |
| KT344895 | Staphylococcus phage phiJB | Staphylococcus phage phiJB Staphylococcus virus phiJB Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43012 | 34.73 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| FN436268 | Acaryochloris phage A-HIS1 | Acaryochloris phage A-HIS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55653 | 47.047 | Unclassified | Unclassified | Siphoviridae | Acaryochloris |
| LC121084 | Ralstonia phage RSP15 | Ralstonia phage RSP15 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167619 | 44.702 | Unclassified | Unclassified | Ackermannviridae | Ralstonia |
| KT345706 | Vibrio phage vB_VorS-PVo5 | Vibrio phage vB_VorS-PVo5 Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80578 | 40.577 | Unclassified | Ermolyevavirinae | Demerecviridae | Vibrio |
| KP340288 | Pseudomonas phage phiKTN6 | Pseudomonas phage phiKTN6 Pseudomonas virus KTN6 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65994 | 55.513 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KP340287 | Pseudomonas phage phiKT28 | Pseudomonas phage phiKT28 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66381 | 55.644 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KP343639 | Ralstonia phage phiITL-1 | Ralstonia phage phiITL-1 Ralstonia virus ITL1 Serkorvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38731 | 62.751 | Serkorvirus | Unclassified | Autographiviridae | Ralstonia |
| KT887559 | Pseudomonas phage phi3 | Pseudomonas phage phi3 Pseudomonas virus phi3 Phitrevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32637 | 63.854 | Phitrevirus | Peduovirinae | Myoviridae | Pseudomonas |
| KT887558 | Pseudomonas phage phi2 | Pseudomonas phage phi2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41871 | 62.115 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KT887557 | Pseudomonas phage phi1 | Pseudomonas phage phi1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57218 | 57.491 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KU160496 | Bacillus phage vB_BhaS-171 | Bacillus phage vB_BhaS-171 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38975 | 40.816 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KU160495 | Exiguobacterium phage vB_EauS-123 | Exiguobacterium phage vB_EauS-123 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30924 | 47.73 | Unclassified | Unclassified | Siphoviridae | Exiguobacterium |
| KU160494 | Vibrio phage vB_VmeM-32 | Vibrio phage vB_VmeM-32 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 199912 | 35.611 | Unclassified | Tevenvirinae | Myoviridae | Vibrio |
| KU255730 | Escherichia phage phiE142 | Escherichia phage phiE142 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121442 | 37.355 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| LN913017 | Pseudomonas phage HC15b2 | Pseudomonas phage HC15b2 unclassified Ghunavirus Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40346 | 56.6 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| LN907801 | Bacteriophage sp. | Bacteriophage sp. unclassified bacterial viruses Viruses | 37087 | 64.319 | Unclassified | Unclassified | Unclassified | Unspecified |
| KU052038 | Escherichia phage SerU-LTIIb | Escherichia phage SerU-LTIIb unclassified bacterial viruses Viruses | 10126 | 46.06 | Unclassified | Unclassified | Unclassified | Escherichia |
| KU052037 | Escherichia phage Rac-SA53 | Escherichia phage Rac-SA53 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52291 | 47.278 | Unclassified | Unclassified | Myoviridae | Escherichia |
| KR072689 | Paracoccus phage Shpa | Paracoccus phage Shpa Paracoccus virus Shpa Vhulanivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38261 | 64.674 | Vhulanivirus | Unclassified | Siphoviridae | Paracoccus |
| KC900378 | Pseudomonas phage LKA5 | Pseudomonas phage LKA5 unclassified Hollowayvirus Hollowayvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64746 | 63.284 | Hollowayvirus | Unclassified | Podoviridae | Pseudomonas |
| KC821634 | Cellulophaga phage phi47:1 | Cellulophaga phage phi47:1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54016 | 33.472 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| KC821633 | Cellulophaga phage phi13:2 | Cellulophaga phage phi13:2 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72369 | 32.873 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821632 | Cellulophaga phage phi4:1 | Cellulophaga phage phi4:1 Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145865 | 32.672 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| KC821631 | Cellulophaga phage phi48:1 | Cellulophaga phage phi48:1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6478 | 34.208 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| KC821630 | Cellulophaga phage phi3:1 | Cellulophaga phage phi3:1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54427 | 33.448 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| KC821629 | Cellulophaga phage phi38:2 | Cellulophaga phage phi38:2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54012 | 33.476 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| KC821628 | Cellulophaga phage phi18:4 | Cellulophaga phage phi18:4 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6478 | 34.27 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| KC821627 | Cellulophaga phage phi18:2 | Cellulophaga phage phi18:2 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38476 | 36.581 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821626 | Cellulophaga phage phi39:1 | Cellulophaga phage phi39:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28760 | 31.332 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| KC821625 | Cellulophaga phage phi13:1 | Cellulophaga phage phi13:1 Cbastvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76666 | 30.223 | Cbastvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821624 | Cellulophaga phage phi14:2 | Cellulophaga phage phi14:2 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100418 | 29.605 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821623 | Cellulophaga phage phi12a:1 | Cellulophaga phage phi12a:1 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6478 | 34.023 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| KC821622 | Cellulophaga phage phi46:3 | Cellulophaga phage phi46:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72961 | 32.696 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821621 | Cellulophaga phage phi19:2 | Cellulophaga phage phi19:2 Cbastvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78275 | 30.182 | Cbastvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821620 | Cellulophaga phage phi18:3 | Cellulophaga phage phi18:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71443 | 32.884 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821619 | Cellulophaga phage phi18:1 | Cellulophaga phage phi18:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39189 | 36.515 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821618 | Cellulophaga phage phi10:1 | Cellulophaga phage phi10:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53664 | 31.535 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| KC821617 | Cellulophaga phage phi17:1 | Cellulophaga phage phi17:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38776 | 36.476 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821616 | Cellulophaga phage phiSM | Cellulophaga phage phiSM Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54566 | 33.464 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| KC821615 | Cellulophaga phage phi12:3 | Cellulophaga phage phi12:3 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39151 | 36.584 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821614 | Cellulophaga phage phi38:1 | Cellulophaga phage phi38:1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72534 | 38.055 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821613 | Cellulophaga phage phi12:1 | Cellulophaga phage phi12:1 Helsingorvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39148 | 36.587 | Helsingorvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC821612 | Cellulophaga phage phi40:1 | Cellulophaga phage phi40:1 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72529 | 38.055 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821611 | Cellulophaga phage phi46:1 | Cellulophaga phage phi46:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34844 | 38.256 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| KC821610 | Cellulophaga phage phi3ST:2 | Cellulophaga phage phi3ST:2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54015 | 33.47 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| KC821609 | Cellulophaga phage phi17:2 | Cellulophaga phage phi17:2 Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145343 | 32.651 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| KC821608 | Cellulophaga phage phi19:3 | Cellulophaga phage phi19:3 crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75994 | 32.547 | Unclassified | Unclassified | Podoviridae | Cellulophaga |
| KC821607 | Cellulophaga phage phi19:1 | Cellulophaga phage phi19:1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57447 | 31.239 | Unclassified | Unclassified | Siphoviridae | Cellulophaga |
| KC821606 | Cellulophaga phage phi12:2 | Cellulophaga phage phi12:2 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6453 | 34.806 | Unclassified | Unclassified | Microviridae | Cellulophaga |
| KC821604 | Cellulophaga phage phiST | Cellulophaga phage phiST Cbastvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78269 | 30.183 | Cbastvirus | Unclassified | Siphoviridae | Cellulophaga |
| KC758116 | Pseudomonas phage LKO4 | Pseudomonas phage LKO4 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61818 | 64.426 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| KU183006 | Klebsiella phage vB_KpnP_IME205 | Klebsiella phage vB_KpnP_IME205 Klebsiella virus IME205 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41310 | 52.213 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KU297675 | Pseudomonas phage PaoP5 | Pseudomonas phage PaoP5 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93464 | 49.508 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC105990 | Pseudomonas phage KPP22M3 | Pseudomonas phage KPP22M3 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64415 | 55.642 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC105989 | Pseudomonas phage KPP22M2 | Pseudomonas phage KPP22M2 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64415 | 55.642 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC105988 | Pseudomonas phage KPP22M1 | Pseudomonas phage KPP22M1 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64415 | 55.642 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LC105987 | Pseudomonas phage KPP22 | Pseudomonas phage KPP22 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64415 | 55.644 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT381879 | Caulobacter phage Percy | Caulobacter phage Percy Caulobacter virus Percy Percyvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44974 | 60.902 | Percyvirus | Unclassified | Autographiviridae | Caulobacter |
| KF188410 | Lactobacillus phage phiLdb | Lactobacillus phage phiLdb Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33996 | 41.952 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| KP861230 | Acinetobacter phage YMC11/11/R3177 | Acinetobacter phage YMC11/11/R3177 Acinetobacter virus R3177 Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47575 | 39.832 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| KP792623 | Rhodoferax phage P26218 | Rhodoferax phage P26218 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36315 | 56.696 | Unclassified | Unclassified | Podoviridae | Rhodoferax |
| KC310806 | Synechococcus phage S-CBP2 | Synechococcus phage S-CBP2 Synechococcus virus SCBP2 Kembevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46237 | 55.034 | Kembevirus | Unclassified | Autographiviridae | Synechococcus |
| KC310805 | Synechococcus phage S-CBP42 | Synechococcus phage S-CBP42 Synechococcus virus SCBP42 Aegirvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45218 | 54.551 | Aegirvirus | Unclassified | Autographiviridae | Synechococcus |
| KC310804 | Synechococcus phage S-CBP4 | Synechococcus phage S-CBP4 Synechococcus virus SCBP4 Poseidonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44147 | 44.381 | Poseidonvirus | Unclassified | Autographiviridae | Synechococcus |
| KC310803 | Synechococcus phage S-CBP3 | Synechococcus phage S-CBP3 Synechococcus virus SCBP3 Lirvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45871 | 46.949 | Lirvirus | Unclassified | Autographiviridae | Synechococcus |
| KC310802 | Synechococcus phage S-CBP1 | Synechococcus phage S-CBP1 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46547 | 47.578 | Unclassified | Unclassified | Autographiviridae | Synechococcus |
| KM896878 | Escherichia phage YD-2008.s | Escherichia phage YD-2008.s Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44613 | 54.614 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| LN889995 | Pseudomonas phage 17A | Pseudomonas phage 17A Pseudomonas virus 17A Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40242 | 57.711 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KT968832 | Pseudomonas phage YMC11/11/R1836 | Pseudomonas phage YMC11/11/R1836 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37714 | 64.249 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KT968831 | Pseudomonas phage YMC11/02/R656 | Pseudomonas phage YMC11/02/R656 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60919 | 58.739 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KT964103 | Klebsiella phage KpV41 | Klebsiella phage KpV41 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44203 | 53.967 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KT962247 | Cellulophaga phage phi17:2_18 | Cellulophaga phage phi17:2_18 unclassified Lightbulbvirus Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145343 | 32.651 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| KT962246 | Cellulophaga phage phi4:1_18 | Cellulophaga phage phi4:1_18 Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145865 | 32.672 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| KT962245 | Cellulophaga phage phi4:1_13 | Cellulophaga phage phi4:1_13 unclassified Lightbulbvirus Lightbulbvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145865 | 32.672 | Lightbulbvirus | Unclassified | Podoviridae | Cellulophaga |
| KU064779 | Pseudomonas phage PPPL-1 | Pseudomonas phage PPPL-1 Pseudomonas virus PPPL1 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41149 | 57.003 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KT970646 | Bacillus phage phiS58 | Bacillus phage phiS58 unclassified Waukeshavirus Waukeshavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46635 | 35.435 | Waukeshavirus | Unclassified | Siphoviridae | Bacillus |
| KT970645 | Bacillus phage phi4J1 | Bacillus phage phi4J1 Bacillus virus phi4J1 Rockvillevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41486 | 35.889 | Rockvillevirus | Unclassified | Siphoviridae | Bacillus |
| KJ716334 | Staphylococcus phage SA97 | Staphylococcus phage SA97 Staphylococcus virus SA97 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40592 | 34.246 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KC969441 | Pseudomonas phage MBL | Pseudomonas phage MBL unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42519 | 62.137 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KF766125 | Shigella phage 75/02 Stx | Shigella phage 75/02 Stx Diegovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60875 | 49.117 | Diegovirus | Sepvirinae | Podoviridae | Shigella |
| KT962832 | Salmonella phage fSE1C | Salmonella phage fSE1C Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41720 | 49.732 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KT967075 | Bacillus phage phi4I1 | Bacillus phage phi4I1 unclassified Camtrevirus Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41999 | 35.282 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| KR709303 | Staphylococcus phage 55-3 | Staphylococcus phage 55-3 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42309 | 35.645 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KR709302 | Staphylococcus phage 55-2 | Staphylococcus phage 55-2 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41898 | 35.742 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KR093656 | Moraxella phage Mcat32 | Moraxella phage Mcat32 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30958 | 42.677 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093655 | Moraxella phage Mcat31 | Moraxella phage Mcat31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30974 | 42.678 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093654 | Moraxella phage Mcat30 | Moraxella phage Mcat30 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30974 | 42.678 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093653 | Moraxella phage Mcat29 | Moraxella phage Mcat29 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34549 | 42.23 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093652 | Moraxella phage Mcat28 | Moraxella phage Mcat28 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54840 | 42.237 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093651 | Moraxella phage Mcat27 | Moraxella phage Mcat27 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41487 | 42.57 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093650 | Moraxella phage Mcat26 | Moraxella phage Mcat26 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37967 | 42.218 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093649 | Moraxella phage Mcat25 | Moraxella phage Mcat25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41268 | 42.607 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093648 | Moraxella phage Mcat24 | Moraxella phage Mcat24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39290 | 42.754 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093647 | Moraxella phage Mcat23 | Moraxella phage Mcat23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34758 | 42.649 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093646 | Moraxella phage Mcat22 | Moraxella phage Mcat22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40408 | 42.484 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093645 | Moraxella phage Mcat21 | Moraxella phage Mcat21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23410 | 42.26 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093644 | Moraxella phage Mcat20 | Moraxella phage Mcat20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23629 | 42.714 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093643 | Moraxella phage Mcat19 | Moraxella phage Mcat19 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36471 | 42.376 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093642 | Moraxella phage Mcat18 | Moraxella phage Mcat18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51889 | 42.213 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093641 | Moraxella phage Mcat17 | Moraxella phage Mcat17 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60303 | 42.456 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093640 | Moraxella phage Mcat16 | Moraxella phage Mcat16 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46782 | 42.672 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093639 | Moraxella phage Mcat15 | Moraxella phage Mcat15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46083 | 42.706 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093638 | Moraxella phage Mcat14 | Moraxella phage Mcat14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17422 | 43.393 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093637 | Moraxella phage Mcat13 | Moraxella phage Mcat13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34664 | 42.773 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093636 | Moraxella phage Mcat12 | Moraxella phage Mcat12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26845 | 43.565 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093635 | Moraxella phage Mcat11 | Moraxella phage Mcat11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25085 | 43.919 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093634 | Moraxella phage Mcat10 | Moraxella phage Mcat10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24841 | 44.145 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093633 | Moraxella phage Mcat9 | Moraxella phage Mcat9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36406 | 43.941 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093632 | Moraxella phage Mcat8 | Moraxella phage Mcat8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40710 | 41.39 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093631 | Moraxella phage Mcat7 | Moraxella phage Mcat7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42027 | 43.624 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093630 | Moraxella phage Mcat6 | Moraxella phage Mcat6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54619 | 43.078 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093629 | Moraxella phage Mcat5 | Moraxella phage Mcat5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47875 | 43.302 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093628 | Moraxella phage Mcat4 | Moraxella phage Mcat4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42819 | 43.214 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093627 | Moraxella phage Mcat3 | Moraxella phage Mcat3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31332 | 43.792 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093626 | Moraxella phage Mcat2 | Moraxella phage Mcat2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40578 | 43.639 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KR093625 | Moraxella phage Mcat1 | Moraxella phage Mcat1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40551 | 43.619 | Unclassified | Unclassified | Siphoviridae | Moraxella |
| KJ749827 | Escherichia phage ECBP5 | Escherichia phage ECBP5 Escherichia virus ECBP5 Gajwadongvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45403 | 45.891 | Gajwadongvirus | Unclassified | Autographiviridae | Escherichia |
| LN887844 | Pseudomonas phage VCM | Pseudomonas phage VCM Pseudomonas virus VCM Otagovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98765 | 48.508 | Otagovirus | Unclassified | Myoviridae | Pseudomonas |
| HG803185 | Escherichia phage P13357 | Escherichia phage P13357 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60896 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| HG803184 | Escherichia phage P13353 | Escherichia phage P13353 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60896 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| HG803183 | Escherichia phage P13368 | Escherichia phage P13368 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60896 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| HG803182 | Escherichia phage P13363 | Escherichia phage P13363 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60896 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| KP881232 | Sinorhizobium phage phiM9 | Sinorhizobium phage phiM9 unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149218 | 49.763 | Unclassified | Unclassified | Ackermannviridae | Sinorhizobium |
| KT372698 | Pseudomonas phage Triton | Pseudomonas phage Triton Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.927 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372697 | Pseudomonas phage Smee | Pseudomonas phage Smee Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66278 | 54.993 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372696 | Pseudomonas phage Poseidon | Pseudomonas phage Poseidon Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.921 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372695 | Pseudomonas phage Nessie | Pseudomonas phage Nessie Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.927 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372694 | Pseudomonas phage Nemo | Pseudomonas phage Nemo Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66283 | 54.999 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372693 | Pseudomonas phage Kula | Pseudomonas phage Kula Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.927 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372692 | Pseudomonas phage Kraken | Pseudomonas phage Kraken Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.924 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372691 | Pseudomonas phage Jollyroger | Pseudomonas phage Jollyroger Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65795 | 54.937 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT372690 | Pseudomonas phage Gallinipper | Pseudomonas phage Gallinipper Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65917 | 54.937 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT932418 | Vibrio phage qdvp001 | Vibrio phage qdvp001 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134742 | 35.349 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KT728931 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5712 | 47.146 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KT728930 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5215 | 46.654 | Unclassified | Unclassified | Inoviridae | Vibrio |
| HQ832595 | Pseudomonas phage PaP1 | Pseudomonas phage PaP1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 91715 | 49.362 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT934943 | Klebsiella phage vB_KpnM_KB57 | Klebsiella phage vB_KpnM_KB57 Klebsiella virus KB57 Mydovirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142987 | 44.616 | Mydovirus | Vequintavirinae | Myoviridae | Klebsiella |
| KT934381 | Propionibacterium phage PA1-14 | Propionibacterium phage PA1-14 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29407 | 53.851 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KT852574 | Yersinia phage vB_YenP_AP10 | Yersinia phage vB_YenP_AP10 Yersinia virus AP10 Apdecimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39235 | 51.757 | Apdecimavirus | Studiervirinae | Autographiviridae | Yersinia |
| KT151958 | Brevibacillus phage Powder | Brevibacillus phage Powder Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52992 | 38.142 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| KT151959 | Brevibacillus phage Sundance | Brevibacillus phage Sundance Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134270 | 35.504 | Unclassified | Unclassified | Siphoviridae | Brevibacillus |
| KT207918 | Bacillus phage Eyuki | Bacillus phage Eyuki Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162252 | 38.773 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT307976 | Bacillus phage AvesoBmore | Bacillus phage AvesoBmore Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167431 | 37.69 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KT151957 | Brevibacillus phage SecTim467 | Brevibacillus phage SecTim467 Jenstvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130482 | 42.706 | Jenstvirus | Unclassified | Siphoviridae | Brevibacillus |
| KT151956 | Brevibacillus phage Osiris | Brevibacillus phage Osiris Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52955 | 38.102 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| KT151955 | Brevibacillus phage Jenst | Brevibacillus phage Jenst Jenstvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126341 | 42.886 | Jenstvirus | Unclassified | Siphoviridae | Brevibacillus |
| KT388103 | Acinetobacter phage vB_AbaP_PD-AB9 | Acinetobacter phage vB_AbaP_PD-AB9 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40938 | 39.338 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KT388102 | Acinetobacter phage vB_AbaP_PD-6A3 | Acinetobacter phage vB_AbaP_PD-6A3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41563 | 39.477 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KP211958 | Cyanophage P-TIM40 | Cyanophage P-TIM40 Prochlorococcus virus PTIM40 Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 188632 | 40.717 | Libanvirus | Unclassified | Myoviridae | Prochlorococcus |
| KJ676859 | Bacillus phage JBP901 | Bacillus phage JBP901 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159492 | 39.658 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KP063902 | Bacillus phage BalMu-1 | Bacillus phage BalMu-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39873 | 42.761 | Unclassified | Unclassified | Myoviridae | Bacillus |
| KP063903 | Bacillus phage BalMu-1 | Bacillus phage BalMu-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39861 | 42.726 | Unclassified | Unclassified | Myoviridae | Bacillus |
| KT254133 | Pbunalikevirus phiFenriz | Pbunalikevirus phiFenriz Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65760 | 54.938 | Pbunavirus | Unclassified | Myoviridae | Unspecified |
| KT254132 | Pbunalikevirus phiHabibi | Pbunalikevirus phiHabibi Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65763 | 54.918 | Pbunavirus | Unclassified | Myoviridae | Unspecified |
| KT254131 | Pbunalikevirus phiMoody | Pbunalikevirus phiMoody Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65908 | 54.902 | Pbunavirus | Unclassified | Myoviridae | Unspecified |
| KT254130 | Pbunalikevirus phiVader | Pbunalikevirus phiVader Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65762 | 54.925 | Pbunavirus | Unclassified | Myoviridae | Unspecified |
| KT588074 | Acinetobacter phage Ab105-1phi | Acinetobacter phage Ab105-1phi Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41496 | 39.486 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| KT825490 | Escherichia phage C119 | Escherichia phage C119 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47319 | 44.238 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KT367887 | Klebsiella phage vB_Kp3 | Klebsiella phage vB_Kp3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48493 | 57.076 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| KT367886 | Klebsiella phage Kp2 | Klebsiella phage Kp2 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43963 | 53.784 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KT367885 | Klebsiella phage vB_Kp1 | Klebsiella phage vB_Kp1 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40114 | 53.283 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KT804923 | Pseudomonas phage C11 | Pseudomonas phage C11 unclassified Pakpunavirus Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94109 | 49.391 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KP282680 | Sulfolobales Virus YNP2 | Sulfolobales Virus YNP2 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 34783 | 42.653 | Unclassified | Unclassified | Unclassified | Unspecified |
| KP282679 | Sulfolobales virus YNP1 | Sulfolobales virus YNP1 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 40860 | 38.338 | Unclassified | Unclassified | Unclassified | Unspecified |
| KR011061 | Skermania phage SPI1 | Skermania phage SPI1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55748 | 67.785 | Unclassified | Unclassified | Siphoviridae | Skermania |
| KJ081346 | Bacillus phage BCP8-2 | Bacillus phage BCP8-2 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159071 | 39.452 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| HG792105 | Escherichia phage P14437 | Escherichia phage P14437 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61465 | 50.227 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| HG792104 | Escherichia phage P13771 | Escherichia phage P13771 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61566 | 50.153 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| HG792102 | Escherichia phage P13803 | Escherichia phage P13803 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60941 | 50.242 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| KR136260 | Polaribacter phage P12002S | Polaribacter phage P12002S Incheonvrus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49847 | 28.933 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| KR136259 | Polaribacter phage P12002L | Polaribacter phage P12002L Incheonvrus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48689 | 28.935 | Unclassified | Unclassified | Siphoviridae | Polaribacter |
| JX194239 | Staphylococcus virus SA11 | Staphylococcus virus SA11 Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136326 | 30.038 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KR270151 | Salmonella phage f18SE | Salmonella phage f18SE Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41868 | 49.799 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KT001919 | Klebsiella phage Miro | Klebsiella phage Miro Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176055 | 41.779 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| KT001918 | Klebsiella phage Matisse | Klebsiella phage Matisse Slopekvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176081 | 41.774 | Slopekvirus | Tevenvirinae | Myoviridae | Klebsiella |
| KT001917 | Escherichia phage Murica | Escherichia phage Murica Escherichia virus Murica Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135391 | 43.614 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KT001916 | Citrobacter phage Michonne | Citrobacter phage Michonne unclassified Mooglevirus Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 90000 | 38.849 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| KT001915 | Citrobacter phage Merlin | Citrobacter phage Merlin Moonvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172733 | 38.763 | Moonvirus | Tevenvirinae | Myoviridae | Citrobacter |
| KT001914 | Caulobacter phage Seuss | Caulobacter phage Seuss Caulobacter virus Seuss Seussvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85138 | 61.403 | Seussvirus | Unclassified | Siphoviridae | Caulobacter |
| KT001913 | Caulobacter phage Sansa | Caulobacter phage Sansa Caulobacter virus Sansa Sansavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80297 | 65.751 | Sansavirus | Unclassified | Siphoviridae | Caulobacter |
| KT001912 | Bacillus phage Silence | Bacillus phage Silence Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40001 | 38.289 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| LN887948 | Escherichia phage slur09 | Escherichia phage slur09 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111751 | 39.015 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| KT429161 | Staphylococcus phage StB20-like | Staphylococcus phage StB20-like Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40670 | 33.29 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KT429160 | Staphylococcus phage SPbeta-like | Staphylococcus phage SPbeta-like Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127726 | 30.534 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| LN828717 | Synechococcus phage S-PM2 | Synechococcus phage S-PM2 Synechococcus virus SPM2 Nodensvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 186736 | 37.797 | Nodensvirus | Unclassified | Myoviridae | Synechococcus |
| KP282678 | Sulfolobus monocaudavirus SMV4 | Sulfolobus monocaudavirus SMV4 unclassified Bicaudaviridae Bicaudaviridae Viruses | 51711 | 38.615 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| KP282677 | Sulfolobus monocaudavirus SMV3 | Sulfolobus monocaudavirus SMV3 unclassified Bicaudaviridae Bicaudaviridae Viruses | 64323 | 38.363 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| KP282676 | Hydrogenobaculum phage 1 | Hydrogenobaculum phage 1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 19351 | 38.184 | Unclassified | Unclassified | Unclassified | Hydrogenobaculum |
| KP282675 | Acidianus rod-shaped virus 2 | Acidianus rod-shaped virus 2 Hoswirudivirus ARV2 Hoswirudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 29763 | 32.349 | Hoswirudivirus | Unclassified | Rudiviridae | Acidianus |
| KP282674 | Acidianus bottle-shaped virus 3 strain ABV3 | Acidianus bottle-shaped virus 3 strain ABV3 Bottigliavirus ABV3 Bottigliavirus Ampullaviridae Viruses | 28489 | 31.654 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| KP282673 | Acidianus bottle-shaped virus 2 strain ABV2 | Acidianus bottle-shaped virus 2 strain ABV2 Bottigliavirus ABV2 Bottigliavirus Ampullaviridae Viruses | 22613 | 33.33 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| KT021004 | Thermobifida phage P1312 | Thermobifida phage P1312 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60284 | 65.906 | Unclassified | Unclassified | Siphoviridae | Thermobifida |
| KP238129 | Sulfolobus monocaudavirus SMV2 | Sulfolobus monocaudavirus SMV2 unclassified Bicaudaviridae Bicaudaviridae Viruses | 50918 | 43.552 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| KM209228 | Dickeya phage phiD3 | Dickeya phage phiD3 Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152308 | 49.357 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| JQ067087 | Pseudomonas phage PaMx11 | Pseudomonas phage PaMx11 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59878 | 64.453 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JQ067089 | Pseudomonas phage PaMx28 | Pseudomonas phage PaMx28 Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55108 | 66.406 | Pamexvirus | Unclassified | Siphoviridae | Pseudomonas |
| JQ067092 | Pseudomonas phage PaMx42 | Pseudomonas phage PaMx42 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43225 | 54.635 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| JQ067093 | Pseudomonas phage PaMx74 | Pseudomonas phage PaMx74 Pamexvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58637 | 68.382 | Pamexvirus | Unclassified | Siphoviridae | Pseudomonas |
| KR063268 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5087 | 46.963 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KR063267 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5714 | 47.252 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KR869157 | Pseudomonas phage vB_PaeM_CEB_DP1 | Pseudomonas phage vB_PaeM_CEB_DP1 Pseudomonas virus DP1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66158 | 55.55 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KT339177 | Lactococcus phage GE1 | Lactococcus phage GE1 Lactococcus virus GE1 Chertseyvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24847 | 37.872 | Chertseyvirus | Unclassified | Siphoviridae | Lactococcus |
| KP682391 | Escherichia phage PA51 | Escherichia phage PA51 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61388 | 49.818 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682376 | Escherichia phage PA12 | Escherichia phage PA12 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61388 | 49.818 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682392 | Escherichia phage PA52 | Escherichia phage PA52 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65334 | 50.067 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682390 | Escherichia phage PA50 | Escherichia phage PA50 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682389 | Escherichia phage PA45 | Escherichia phage PA45 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65334 | 50.064 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682388 | Escherichia phage PA44 | Escherichia phage PA44 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682387 | Escherichia phage PA42 | Escherichia phage PA42 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.904 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682386 | Escherichia phage PA36 | Escherichia phage PA36 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64021 | 49.98 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682385 | Escherichia phage PA33 | Escherichia phage PA33 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682384 | Escherichia phage PA32 | Escherichia phage PA32 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.904 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682383 | Escherichia phage PA30 | Escherichia phage PA30 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682382 | Escherichia phage PA29 | Escherichia phage PA29 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682381 | Escherichia phage PA28 | Escherichia phage PA28 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61292 | 49.741 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682380 | Escherichia phage PA27 | Escherichia phage PA27 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64021 | 49.984 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682379 | Escherichia phage PA21 | Escherichia phage PA21 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.904 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682378 | Escherichia phage PA18 | Escherichia phage PA18 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682377 | Escherichia phage PA16 | Escherichia phage PA16 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682375 | Escherichia phage PA11 | Escherichia phage PA11 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682374 | Escherichia phage PA8 | Escherichia phage PA8 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63584 | 50.167 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| KP682373 | Escherichia phage PA5 | Escherichia phage PA5 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682372 | Escherichia phage PA4 | Escherichia phage PA4 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62708 | 49.903 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KP682371 | Escherichia phage PA2 | Escherichia phage PA2 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63569 | 50.169 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| KJ959591 | Pseudomonas phage PAN70 | Pseudomonas phage PAN70 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39673 | 62.12 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| AB063393 | Thermus virus IN93 | Thermus virus IN93 Gammasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 19604 | 65.936 | Gammasphaerolipovirus | Unclassified | Sphaerolipoviridae | Thermus |
| KR153145 | Streptococcus phage APCM01 | Streptococcus phage APCM01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31075 | 39.028 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AY048850 | Natrinema virus SNJ1 | Natrinema virus SNJ1 Betasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 16341 | 61.104 | Betasphaerolipovirus | Unclassified | Sphaerolipoviridae | Natrinema |
| KF279417 | Mycobacterium phage Quink | Mycobacterium phage Quink unclassified Kostyavirus Kostyavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76586 | 62.993 | Kostyavirus | Unclassified | Siphoviridae | Mycobacterium |
| KT388093 | Streptococcus phage A25 | Streptococcus phage A25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33900 | 38.448 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KT626446 | Bacillus phage phi4B1 | Bacillus phage phi4B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38663 | 35.892 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| KT588072 | Enterococcus phage IME-EFm5 | Enterococcus phage IME-EFm5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42265 | 35.509 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KT319626 | Listeria phage WIL-3 | Listeria phage WIL-3 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10610 | 36.277 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KT319620 | Listeria phage WIL-3 | Listeria phage WIL-3 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10387 | 37.335 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KT319607 | Listeria phage WIL-3 | Listeria phage WIL-3 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15899 | 37.493 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KT319602 | Listeria phage WIL-3 | Listeria phage WIL-3 unclassified Pecentumvirus Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14414 | 36.284 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KT381880 | Citrobacter phage Margaery | Citrobacter phage Margaery Citrobacter virus Margaery Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178182 | 44.868 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| KT221034 | Streptomyces phage SF3 | Streptomyces phage SF3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60934 | 69.112 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| KP790011 | Gordonia phage Gsput1 | Gordonia phage Gsput1 Gordonia virus Gsput1 Gesputvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43505 | 62.832 | Gesputvirus | Unclassified | Siphoviridae | Gordonia |
| KP790010 | Gordonia phage GordDuk1 | Gordonia phage GordDuk1 unclassified Gordtnkvirus Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76276 | 50.653 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| KP790009 | Gordonia phage Gmala1 | Gordonia phage Gmala1 unclassified Gordtnkvirus Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75167 | 50.798 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| KP790008 | Gordonia phage GordTnk2 | Gordonia phage GordTnk2 Gordtnkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75987 | 50.694 | Gordtnkvirus | Unclassified | Siphoviridae | Gordonia |
| AB916497 | Ralstonia phage RS138 | Ralstonia phage RS138 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 65.103 | Unclassified | Unclassified | Siphoviridae | Ralstonia |
| KT321316 | Achromobacter phage phiAxp-2 | Achromobacter phage phiAxp-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62220 | 60.111 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| LN878149 | Cronobacter phage Dev-CD-23823 | Cronobacter phage Dev-CD-23823 Cronobacter virus DevCD23823 Cronosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41620 | 53.84 | Cronosvirus | Melnykvirinae | Autographiviridae | Cronobacter |
| KR537872 | Stenotrophomonas phage vB_SmaS-DLP_1 | Stenotrophomonas phage vB_SmaS-DLP_1 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42887 | 53.711 | Septimatrevirus | Unclassified | Siphoviridae | Stenotrophomonas |
| KR537871 | Stenotrophomonas phage vB_SmaS-DLP_2 | Stenotrophomonas phage vB_SmaS-DLP_2 unclassified Septimatrevirus Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42593 | 53.715 | Septimatrevirus | Unclassified | Siphoviridae | Stenotrophomonas |
| KM873719 | Flavobacterium phage FCL-2 | Flavobacterium phage FCL-2 Flavobacterium virus FCL2 Ficleduovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47142 | 30.198 | Ficleduovirus | Unclassified | Myoviridae | Flavobacterium |
| KP137439 | Mannheimia phage vB_MhS_3927AP1 | Mannheimia phage vB_MhS_3927AP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52049 | 41.371 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| KP137437 | Mannheimia phage vB_MhS_1152AP2 | Mannheimia phage vB_MhS_1152AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52139 | 41.094 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| KP137433 | Mannheimia phage vB_MhS_535AP2 | Mannheimia phage vB_MhS_535AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50078 | 41.31 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| KP137440 | Mannheimia phage vB_MhM_3927AP2 | Mannheimia phage vB_MhM_3927AP2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33755 | 43.137 | Unclassified | Unclassified | Myoviridae | Mannheimia |
| KP137438 | Mannheimia phage vB_MhM_2256AP1 | Mannheimia phage vB_MhM_2256AP1 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34926 | 41.519 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| KP137436 | Mannheimia phage vB_MhM_1127AP1 | Mannheimia phage vB_MhM_1127AP1 Mannheimia virus 1127AP1 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36745 | 41.984 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| KP137435 | Mannheimia phage vB_MhS_587AP2 | Mannheimia phage vB_MhS_587AP2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48594 | 41.285 | Unclassified | Unclassified | Siphoviridae | Mannheimia |
| KP137434 | Mannheimia phage vB_MhM_587AP1 | Mannheimia phage vB_MhM_587AP1 unclassified Baylorvirus Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35764 | 42.062 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| KP137432 | Mannheimia phage vB_MhM_535AP1 | Mannheimia phage vB_MhM_535AP1 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34565 | 41.574 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| KP054477 | Lactobacillus phage LfeInf | Lactobacillus phage LfeInf Lactobacillus virus LfeInf Hopescreekvirus Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 106071 | 38.162 | Hopescreekvirus | Unclassified | Herelleviridae | Lactobacillus |
| KP027015 | Lactobacillus phage LfeSau | Lactobacillus phage LfeSau Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31703 | 48.024 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| KR011064 | Tsukamurella phage TIN4 | Tsukamurella phage TIN4 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76268 | 59.29 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| KR011063 | Tsukamurella phage TIN3 | Tsukamurella phage TIN3 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76269 | 59.291 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| KR011062 | Tsukamurella phage TIN2 | Tsukamurella phage TIN2 Tinduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76964 | 58.92 | Tinduovirus | Unclassified | Siphoviridae | Tsukamurella |
| JQ309827 | Enterococcus phage vB_Efae230P-4 | Enterococcus phage vB_Efae230P-4 Enterococcus virus Efae230P4 Copernicusvirus Sarlesvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17972 | 32.757 | Copernicusvirus | Sarlesvirinae | Rountreeviridae | Enterococcus |
| KP696448 | Bacillus phage Stills | Bacillus phage Stills Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80798 | 35.481 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| KP696447 | Bacillus phage Stahl | Bacillus phage Stahl Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80148 | 35.257 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| KR604693 | Pectobacterium phage Peat1 | Pectobacterium phage Peat1 Pectobacterium virus Peat1 Phimunavirus Corkvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45633 | 48.862 | Phimunavirus | Corkvirinae | Autographiviridae | Pectobacterium |
| KT321314 | Enterobacter phage phiEap-1 | Enterobacter phage phiEap-1 Enterobacter virus Eap1 Eapunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39133 | 51.688 | Eapunavirus | Studiervirinae | Autographiviridae | Enterobacter |
| KP994596 | Pseudoalteromonas phage H103 | Pseudoalteromonas phage H103 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43190 | 41.174 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KM434186 | Pseudomonas phage vB_PaeM_PS24 | Pseudomonas phage vB_PaeM_PS24 Pseudomonas virus PS24 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84583 | 54.7 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KM434185 | Pseudomonas phage vB_Pae_PS9N | Pseudomonas phage vB_Pae_PS9N Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43047 | 53.764 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| KM434184 | Pseudomonas phage vB_Pae_PS44 | Pseudomonas phage vB_Pae_PS44 Pseudomonas virus PS44 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68871 | 55.242 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| HG798901 | Clostridium phage phiCDHM11 | Clostridium phage phiCDHM11 Clostridioides virus phiCDHM11 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32000 | 29.256 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| KM360178 | Escherichia phage vB_EcoM-ep3 | Escherichia phage vB_EcoM-ep3 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42351 | 53.347 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| KT353109 | Cronobacter phage PBES 02 | Cronobacter phage PBES 02 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149732 | 50.725 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| KT221033 | Streptomyces phage SF1 | Streptomyces phage SF1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43150 | 69.089 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| KT239446 | Klebsiella phage JD18 | Klebsiella phage JD18 Jiaodavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166313 | 39.599 | Jiaodavirus | Tevenvirinae | Myoviridae | Klebsiella |
| KP202972 | Paenibacillus phage HB10c2 | Paenibacillus phage HB10c2 Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35644 | 41.844 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KF148616 | Campylobacter phage CP8 | Campylobacter phage CP8 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132667 | 26.045 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| JX569801 | Campylobacter phage CP30A | Campylobacter phage CP30A Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133572 | 26.127 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| KF582788 | Escherichia phage vB_EcoM_JS09 | Escherichia phage vB_EcoM_JS09 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169148 | 37.649 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KR063280 | Gordonia phage GMA6 | Gordonia phage GMA6 Gordonia virus GMA6 Bendigovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83324 | 58.179 | Bendigovirus | Unclassified | Myoviridae | Gordonia |
| KR063279 | Gordonia phage GMA3 | Gordonia phage GMA3 Gordonia virus GMA3 Gamtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77779 | 51.263 | Gamtrevirus | Unclassified | Siphoviridae | Gordonia |
| KR063281 | Gordonia phage GMA2 | Gordonia phage GMA2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103424 | 53.372 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR063278 | Gordonia phage GMA7 | Gordonia phage GMA7 Gordonia virus GMA7 Getseptimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73419 | 56.611 | Getseptimavirus | Unclassified | Siphoviridae | Gordonia |
| KR053201 | Gordonia phage GTE8 | Gordonia phage GTE8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67617 | 65.957 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR053199 | Gordonia phage GMA4 | Gordonia phage GMA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45537 | 66.368 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR053197 | Gordonia phage GRU3 | Gordonia phage GRU3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17727 | 66.509 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR053200 | Gordonia phage GTE6 | Gordonia phage GTE6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56982 | 67.797 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR053198 | Gordonia phage GMA5 | Gordonia phage GMA5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17562 | 66.353 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| KR052482 | Sinorhizobium phage phiN3 | Sinorhizobium phage phiN3 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 206713 | 49.075 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| KR052481 | Sinorhizobium phage phiM19 | Sinorhizobium phage phiM19 unclassified Emdodecavirus Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 188047 | 49.027 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| KR052480 | Sinorhizobium phage phiM7 | Sinorhizobium phage phiM7 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 188427 | 49.037 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| AB716666 | Klebsiella phage NTUH-K2044-K1-1 | Klebsiella phage NTUH-K2044-K1-1 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43871 | 54.179 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| LC066596 | Ralstonia phage RSS-TH1 | Ralstonia phage RSS-TH1 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7273 | 62.12 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| KT186243 | Staphylococcus phage vB_SauS_phi2 | Staphylococcus phage vB_SauS_phi2 Staphylococcus virus phi2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44222 | 33.716 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| KT176190 | Escherichia phage QL01 | Escherichia phage QL01 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170527 | 39.647 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| KM236239 | Citrobacter phage Moogle | Citrobacter phage Moogle Mooglevirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87999 | 38.982 | Mooglevirus | Ounavirinae | Myoviridae | Citrobacter |
| HG934470 | Salmonella phage vB_SenS-Ent3 | Salmonella phage vB_SenS-Ent3 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42764 | 49.792 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| HG934469 | Salmonella phage vB_SenS-Ent2 | Salmonella phage vB_SenS-Ent2 unclassified Jerseyvirus Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42093 | 49.916 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| HE775250 | Salmonella phage Ent1 | Salmonella phage Ent1 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42391 | 49.789 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KP774835 | Citrobacter phage CVT22 | Citrobacter phage CVT22 Citrobacter virus CVT22 Citrovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47636 | 41.632 | Citrovirus | Unclassified | Zobellviridae | Citrobacter |
| KP893290 | Staphylococcus phage B236 | Staphylococcus phage B236 Staphylococcus virus B236 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43228 | 35.553 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KP893289 | Staphylococcus phage B166 | Staphylococcus phage B166 Staphylococcus virus B166 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42881 | 34.829 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KP719134 | Enterobacteria phage JenK1 | Enterobacteria phage JenK1 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60747 | 43.862 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| KP719133 | Enterobacteria phage JenP2 | Enterobacteria phage JenP2 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59802 | 43.214 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| KP719132 | Enterobacteria phage JenP1 | Enterobacteria phage JenP1 Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60754 | 43.232 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| HM588722 | Pseudoalteromonas phage H105/1 | Pseudoalteromonas phage H105/1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30651 | 40.821 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KR262148 | Klebsiella phage KLPN1 | Klebsiella phage KLPN1 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49037 | 50.525 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KR869820 | Citrobacter phage IME-CF2 | Citrobacter phage IME-CF2 Citrobacter virus CF2 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177688 | 43.181 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| KT266805 | Roseobacter phage RDJL Phi 2 | Roseobacter phage RDJL Phi 2 Xiamenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63513 | 57.295 | Xiamenvirus | Unclassified | Siphoviridae | Roseobacter |
| KT184661 | Yersinia phage vB_YenP_ISAO8 | Yersinia phage vB_YenP_ISAO8 Yersinia virus ISAO8 Aghbyvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41449 | 53.84 | Aghbyvirus | Melnykvirinae | Autographiviridae | Yersinia |
| KR908644 | Staphylococcus phage IME-SA119 | Staphylococcus phage IME-SA119 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141028 | 30.326 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KR902361 | Staphylococcus phage IME-SA118 | Staphylococcus phage IME-SA118 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139750 | 30.323 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KR816341 | Erysipelothrix phage SE-1 | Erysipelothrix phage SE-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34997 | 33.951 | Unclassified | Unclassified | Siphoviridae | Erysipelothrix |
| KR149290 | Acinetobacter phage Fri1 | Acinetobacter phage Fri1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41805 | 39.287 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KP144388 | Geobacillus virus E3 | Geobacillus virus E3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141298 | 29.635 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| KJ452292 | Staphylococcus phage 23MRA | Staphylococcus phage 23MRA Staphylococcus virus 23MRA Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43098 | 32.902 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KJ452291 | Staphylococcus phage 3MRA | Staphylococcus phage 3MRA Staphylococcus virus 3MRA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41931 | 35.411 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KR560069 | Stenotrophomonas phage IME-SM1 | Stenotrophomonas phage IME-SM1 Stenotrophomonas virus IMESM1 Menderavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159514 | 54.142 | Menderavirus | Unclassified | Myoviridae | Stenotrophomonas |
| KR781488 | Shigella phage Ss-VASD | Shigella phage Ss-VASD Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62851 | 50.071 | Oslovirus | Sepvirinae | Podoviridae | Shigella |
| KR534323 | Pseudoalteromonas phage H101 | Pseudoalteromonas phage H101 Pseudoalteromonas virus H101 Shandongvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131903 | 37.364 | Shandongvirus | Unclassified | Myoviridae | Pseudoalteromonas |
| LC064302 | Pseudomonas phage KPP21 | Pseudomonas phage KPP21 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73420 | 53.499 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KJ000058 | Salmonella phage STP4-a | Salmonella phage STP4-a Salmonella virus STP4a Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159914 | 36.862 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| LN610590 | Pseudomonas phage vB_PaeS_PAO1_Ab30 | Pseudomonas phage vB_PaeS_PAO1_Ab30 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37238 | 64.08 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| LN610589 | Pseudomonas phage vB_PaeM_C1-14_Ab28 | Pseudomonas phage vB_PaeM_C1-14_Ab28 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66181 | 54.902 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610588 | Pseudomonas phage vB_PaeM_PAO1_Ab29 | Pseudomonas phage vB_PaeM_PAO1_Ab29 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66326 | 55.64 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610587 | Pseudomonas phage vB_PaeM_C2-10_Ab15 | Pseudomonas phage vB_PaeM_C2-10_Ab15 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93308 | 49.346 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610586 | Pseudomonas phage vB_PaeM_C2-10_Ab10 | Pseudomonas phage vB_PaeM_C2-10_Ab10 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93053 | 49.252 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610585 | Pseudomonas phage vB_PaeS_PAO1_Ab20 | Pseudomonas phage vB_PaeS_PAO1_Ab20 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57745 | 63.476 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| LN610584 | Pseudomonas phage vB_PaeS_PAO1_Ab19 | Pseudomonas phage vB_PaeS_PAO1_Ab19 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58139 | 63.303 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| LN610583 | Pseudomonas phage vB_PaeM_PAO1_Ab11 | Pseudomonas phage vB_PaeM_PAO1_Ab11 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 85783 | 54.525 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| LN610580 | Pseudomonas phage vB_PaeP_PAO1_1-15pyo | Pseudomonas phage vB_PaeP_PAO1_1-15pyo Pseudomonas virus 15pyo Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42750 | 62.18 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| LN610579 | Pseudomonas phage vB_PaeM_PAO1_Ab27 | Pseudomonas phage vB_PaeM_PAO1_Ab27 unclassified Pbunavirus Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66299 | 55.668 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610578 | Pseudomonas phage vB_PaeP_C2-10_Ab22 | Pseudomonas phage vB_PaeP_C2-10_Ab22 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45808 | 52.353 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| LN610577 | Pseudomonas phage vB_PaeS_PAO1_Ab18 | Pseudomonas phage vB_PaeS_PAO1_Ab18 Abidjanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56537 | 63.498 | Abidjanvirus | Unclassified | Siphoviridae | Pseudomonas |
| LN610575 | Pseudomonas phage vB_PaeM_C2-10_Ab08 | Pseudomonas phage vB_PaeM_C2-10_Ab08 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93503 | 49.206 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| LN610574 | Pseudomonas phage vB_PaeP_PAO1_Ab05 | Pseudomonas phage vB_PaeP_PAO1_Ab05 Pseudomonas virus Ab05 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43639 | 62.346 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| LN610572 | Pseudomonas phage vB_PaeM_C2-10_Ab02 | Pseudomonas phage vB_PaeM_C2-10_Ab02 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93848 | 49.397 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KM092515 | Escherichia phage HY02 | Escherichia phage HY02 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86252 | 38.921 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| AB971354 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4217 | 48.138 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| KJ477686 | Escherichia virus T4 | Escherichia virus T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165660 | 35.263 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KJ477685 | Escherichia virus T4 | Escherichia virus T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166816 | 35.319 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KJ477684 | Escherichia virus T4 | Escherichia virus T4 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168922 | 35.291 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| AB910393 | Pseudomonas phage KPP25 | Pseudomonas phage KPP25 Kochitakasuvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64113 | 60.287 | Kochitakasuvirus | Unclassified | Podoviridae | Pseudomonas |
| AB910392 | Pseudomonas phage KPP23 | Pseudomonas phage KPP23 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62774 | 55.999 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| AB853331 | Staphylococcus phage S25-4 | Staphylococcus phage S25-4 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132123 | 30.309 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| AB853330 | Staphylococcus phage S25-3 | Staphylococcus phage S25-3 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139738 | 30.219 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KC595518 | Brevibacillus phage Davies | Brevibacillus phage Davies Abouovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45798 | 39.1 | Abouovirus | Unclassified | Myoviridae | Brevibacillus |
| KC595517 | Brevibacillus phage Abouo | Brevibacillus phage Abouo Abouovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45552 | 39.16 | Abouovirus | Unclassified | Myoviridae | Brevibacillus |
| KC595516 | Brevibacillus phage Emery | Brevibacillus phage Emery Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58573 | 41.441 | Unclassified | Unclassified | Myoviridae | Brevibacillus |
| KC595515 | Brevibacillus phage Jimmer1 | Brevibacillus phage Jimmer1 unclassified Jimmervirus Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54312 | 38.109 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| KC595514 | Brevibacillus phage Jimmer2 | Brevibacillus phage Jimmer2 Jimmervirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54312 | 38.097 | Jimmervirus | Unclassified | Myoviridae | Brevibacillus |
| AB731695 | Helicobacter phage KHP40 | Helicobacter phage KHP40 Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26449 | 35.941 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| AB647160 | Helicobacter phage KHP30 | Helicobacter phage KHP30 Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26215 | 35.77 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| KJ133707 | Brucella phage V_141 | Brucella phage V_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.167 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133706 | Brucella phage V_19 | Brucella phage V_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.173 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133705 | Brucella phage Tb_141 | Brucella phage Tb_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.172 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133704 | Brucella phage Tbilisi | Brucella phage Tbilisi Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.167 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133703 | Brucella phage 1066_141 | Brucella phage 1066_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.171 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133702 | Brucella phage 1066_19 | Brucella phage 1066_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.169 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133701 | Brucella phage 544_141 | Brucella phage 544_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.172 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133700 | Brucella phage 544_19 | Brucella phage 544_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.179 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133699 | Brucella phage 281_141 | Brucella phage 281_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.176 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133698 | Brucella phage 281_19 | Brucella phage 281_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.173 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133697 | Brucella phage 177_141 | Brucella phage 177_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.167 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133696 | Brucella phage 177_19 | Brucella phage 177_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.167 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133695 | Brucella phage 141_141 | Brucella phage 141_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.186 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133694 | Brucella phage 141_19 | Brucella phage 141_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.183 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133693 | Brucella phage 110_141 | Brucella phage 110_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.169 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133692 | Brucella phage 110_19 | Brucella phage 110_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.162 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133691 | Brucella phage 11sa_141 | Brucella phage 11sa_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.179 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133690 | Brucella phage 11sa_19 | Brucella phage 11sa_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.166 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133689 | Brucella phage 02_141 | Brucella phage 02_141 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.166 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KJ133688 | Brucella phage 02_19 | Brucella phage 02_19 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.166 | Perisivirus | Unclassified | Podoviridae | Brucella |
| HG796220 | Burkholderia phage Bp-AMP3 | Burkholderia phage Bp-AMP3 unclassified Ampunavirus Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41882 | 61.774 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| AP014684 | Pseudomonas phage PS-1 | Pseudomonas phage PS-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48666 | 59.834 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KR014248 | Escherichia phage vB_EcoM_AYO145A | Escherichia phage vB_EcoM_AYO145A Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87372 | 39.003 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KP296796 | Paenibacillus phage Sitara | Paenibacillus phage Sitara Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43724 | 41.593 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KP296795 | Paenibacillus phage Shelly | Paenibacillus phage Shelly Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41152 | 41.534 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KP296794 | Bacteriophage Redbud | Bacteriophage Redbud unclassified Fernvirus Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37971 | 41.813 | Fernvirus | Unclassified | Siphoviridae | Unspecified |
| KP296793 | Paenibacillus phage Rani | Paenibacillus phage Rani Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37990 | 41.779 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KP296792 | Bacteriophage Lily | Bacteriophage Lily Paenibacillus virus Lily Lilyvirus Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44952 | 42.73 | Lilyvirus | Unclassified | Unclassified | Unspecified |
| KP296791 | Paenibacillus phage Diva | Paenibacillus phage Diva Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37246 | 42.072 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| KM190144 | Escherichia phage Av-05 | Escherichia phage Av-05 Escherichia virus Av05 Avunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 120938 | 40.044 | Avunavirus | Vequintavirinae | Myoviridae | Escherichia |
| KJ909655 | Escherichia Stx1-converting recombinant phage HUN/2013 | Escherichia Stx1-converting recombinant phage HUN/2013 unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61137 | 49.71 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| KR091952 | Pseudomonas phage phiPsa17 | Pseudomonas phage phiPsa17 Pseudomonas virus PhiPsa17 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40525 | 57.291 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KF831354 | Staphylococcus phage phiBU01 | Staphylococcus phage phiBU01 Staphylococcus virus BU01 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43748 | 33.016 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KF356198 | Anabaena phage A-4L | Anabaena phage A-4L Anabaena virus A4L Kozyakovvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41750 | 43.437 | Kozyakovvirus | Unclassified | Podoviridae | Anabaena |
| AB897757 | Klebsiella phage K64-1 | Klebsiella phage K64-1 Klebsiella virus K64-1 Alcyoneusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 346602 | 31.717 | Alcyoneusvirus | Unclassified | Myoviridae | Klebsiella |
| KP890823 | Proteus phage vB_PmiM_Pm5461 | Proteus phage vB_PmiM_Pm5461 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161989 | 31.082 | Unclassified | Tevenvirinae | Myoviridae | Proteus |
| KP890822 | Proteus phage vB_PmiP_Pm5460 | Proteus phage vB_PmiP_Pm5460 Proteus virus Pm5460 Acadevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44573 | 39.576 | Acadevirus | Molineuxvirinae | Autographiviridae | Proteus |
| KM589516 | Parabacteroides phage YZ-2015b | Parabacteroides phage YZ-2015b unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5001 | 40.152 | Unclassified | Unclassified | Microviridae | Parabacteroides |
| KM589515 | Parabacteroides phage YZ-2015a | Parabacteroides phage YZ-2015a unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4623 | 42.808 | Unclassified | Unclassified | Microviridae | Parabacteroides |
| KM589514 | Microviridae Fen7940_21 | Microviridae Fen7940_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4453 | 52.773 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589513 | Microviridae Fen7918_21 | Microviridae Fen7918_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4199 | 44.534 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589512 | Microviridae Fen7895_21 | Microviridae Fen7895_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4424 | 43.716 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589511 | Gokushovirinae Fen7875_21 | Gokushovirinae Fen7875_21 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4631 | 53.444 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KM589510 | Microviridae Fen7786_21 | Microviridae Fen7786_21 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4686 | 43.064 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589509 | Microviridae Fen685_11 | Microviridae Fen685_11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4654 | 45.982 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589508 | Gokushovirinae Fen672_31 | Gokushovirinae Fen672_31 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4624 | 49.632 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KM589507 | Microviridae Fen51_42 | Microviridae Fen51_42 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4451 | 42.35 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589506 | Microviridae Fen4707_41 | Microviridae Fen4707_41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4552 | 44.464 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589505 | Microviridae Fen418_41 | Microviridae Fen418_41 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4949 | 42.797 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589504 | Microviridae Fen2266_11 | Microviridae Fen2266_11 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4366 | 40.609 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589503 | Microviridae Bog9017_22 | Microviridae Bog9017_22 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4671 | 41.94 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589502 | Gokushovirinae Bog8989_22 | Gokushovirinae Bog8989_22 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4656 | 39.798 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KM589501 | Gokushovirinae Bog5712_52 | Gokushovirinae Bog5712_52 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4501 | 51.744 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KM589500 | Microviridae Bog5275_51 | Microviridae Bog5275_51 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4815 | 47.622 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589499 | Microviridae Bog1249_12 | Microviridae Bog1249_12 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4451 | 42.373 | Unclassified | Unclassified | Microviridae | Unspecified |
| KM589498 | Gokushovirinae Bog1183_53 | Gokushovirinae Bog1183_53 unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4609 | 36.537 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KP797973 | Salmonella phage Det7 | Salmonella phage Det7 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157498 | 44.598 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KR149291 | Klebsiella phage K5 | Klebsiella phage K5 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41698 | 52.492 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| KR073661 | Escherichia phage pro483 | Escherichia phage pro483 Escherichia virus pro483 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29237 | 52.981 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| KR073660 | Escherichia phage pro147 | Escherichia phage pro147 Escherichia virus pro147 Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32675 | 50.742 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| KR153873 | Delftia phage IME-DE1 | Delftia phage IME-DE1 Delftia virus IMEDE1 Piedvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38084 | 60.427 | Piedvirus | Unclassified | Autographiviridae | Delftia |
| KP972569 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6722 | 47.054 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KP972568 | Vibrio phage pre-CTX | Vibrio phage pre-CTX unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6722 | 47.054 | Unclassified | Unclassified | Inoviridae | Vibrio |
| KM677185 | Lactococcus phage WRP3 | Lactococcus phage WRP3 Lactococcus virus WRP3 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 130008 | 32.36 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| KC311669 | Acinetobacter phage AB3 | Acinetobacter phage AB3 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31185 | 39.176 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KP869113 | Escherichia coli O157 typing phage 15 | Escherichia coli O157 typing phage 15 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92632 | 37.6 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KP869112 | Escherichia coli O157 typing phage 14 | Escherichia coli O157 typing phage 14 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131952 | 43.374 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KP869111 | Escherichia coli O157 typing phage 13 | Escherichia coli O157 typing phage 13 unclassified Mosigvirus Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162417 | 37.324 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KP869110 | Escherichia coli O157 typing phage 12 | Escherichia coli O157 typing phage 12 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88632 | 38.907 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KP869109 | Escherichia coli O157 typing phage 11 | Escherichia coli O157 typing phage 11 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88771 | 38.887 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KP869108 | Escherichia coli O157 typing phage 10 | Escherichia coli O157 typing phage 10 unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39234 | 48.947 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| KP869107 | Escherichia coli O157 typing phage 9 | Escherichia coli O157 typing phage 9 unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43991 | 48.208 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| KP869106 | Escherichia coli O157 typing phage 8 | Escherichia coli O157 typing phage 8 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88998 | 38.926 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KP869105 | Escherichia coli O157 typing phage 7 | Escherichia coli O157 typing phage 7 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167747 | 35.402 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KP869104 | Escherichia coli O157 typing phage 6 | Escherichia coli O157 typing phage 6 Escherichia virus O157tp6 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160570 | 37.628 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KP869103 | Escherichia coli O157 typing phage 5 | Escherichia coli O157 typing phage 5 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137973 | 43.559 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KP869102 | Escherichia coli O157 typing phage 4 | Escherichia coli O157 typing phage 4 unclassified Vequintavirus Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141707 | 43.541 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| KP869101 | Escherichia coli O157 typing phage 3 | Escherichia coli O157 typing phage 3 Escherichia virus O157tp3 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168733 | 37.627 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KP869100 | Escherichia typing phage 1 | Escherichia typing phage 1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88531 | 38.849 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KJ410740 | Synechococcus phage S-EIVl | Synechococcus phage S-EIVl unclassified bacterial viruses Viruses | 79178 | 45.86 | Unclassified | Unclassified | Unclassified | Synechococcus |
| KF977490 | Rhizobium phage vB_RglS_P106B | Rhizobium phage vB_RglS_P106B Rhizobium virus P106B Rigallicvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56024 | 47.922 | Rigallicvirus | Unclassified | Siphoviridae | Rhizobium |
| KR131710 | Fusobacterium phage Funu1 | Fusobacterium phage Funu1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39921 | 26.986 | Unclassified | Unclassified | Myoviridae | Fusobacterium |
| KJ578792 | Propionibacterium phage PHL308M00 | Propionibacterium phage PHL308M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29442 | 54.249 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578791 | Propionibacterium phage PHL301M00 | Propionibacterium phage PHL301M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29323 | 54.408 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578790 | Propionibacterium phage PHL199M00 | Propionibacterium phage PHL199M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29806 | 54.043 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578789 | Propionibacterium phage PHL194M00 | Propionibacterium phage PHL194M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.159 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578788 | Propionibacterium phage PHL179M00 | Propionibacterium phage PHL179M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29428 | 53.928 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578787 | Propionibacterium phage PHL171M01 | Propionibacterium phage PHL171M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29327 | 53.712 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578786 | Propionibacterium phage PHL163M00 | Propionibacterium phage PHL163M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.155 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578785 | Propionibacterium phage PHL152M00 | Propionibacterium phage PHL152M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29247 | 54.153 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578784 | Propionibacterium phage PHL151N00 | Propionibacterium phage PHL151N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.99 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578783 | Propionibacterium phage PHL151M00 | Propionibacterium phage PHL151M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.993 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578782 | Propionibacterium phage PHL150M00 | Propionibacterium phage PHL150M00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29438 | 54.25 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578781 | Propionibacterium phage PHL141N00 | Propionibacterium phage PHL141N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29494 | 53.943 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578780 | Propionibacterium phage PHL132N00 | Propionibacterium phage PHL132N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29003 | 54.329 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578779 | Propionibacterium phage PHL117M01 | Propionibacterium phage PHL117M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29422 | 53.912 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578778 | Propionibacterium phage PHL117M00 | Propionibacterium phage PHL117M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29255 | 54.162 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578777 | Propionibacterium phage PHL116M10 | Propionibacterium phage PHL116M10 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29396 | 53.956 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578776 | Propionibacterium phage PHL116M00 | Propionibacterium phage PHL116M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29394 | 53.96 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578775 | Propionibacterium phage PHL114N00 | Propionibacterium phage PHL114N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29464 | 54.198 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578774 | Propionibacterium phage PHL095N00 | Propionibacterium phage PHL095N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29751 | 53.995 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578773 | Propionibacterium phage PHL092M00 | Propionibacterium phage PHL092M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29261 | 53.918 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578772 | Propionibacterium phage PHL085N00 | Propionibacterium phage PHL085N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29454 | 53.833 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578771 | Propionibacterium phage PHL082M04 | Propionibacterium phage PHL082M04 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.379 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578770 | Propionibacterium phage PHL082M03 | Propionibacterium phage PHL082M03 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.383 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578769 | Propionibacterium phage PHL082M02 | Propionibacterium phage PHL082M02 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.379 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578768 | Propionibacterium phage PHL082M00 | Propionibacterium phage PHL082M00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29491 | 54.379 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578767 | Propionibacterium phage PHL070N00 | Propionibacterium phage PHL070N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29421 | 54.04 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578766 | Propionibacterium phage PHL067M09 | Propionibacterium phage PHL067M09 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29386 | 54.278 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578765 | Propionibacterium phage PHL067M01 | Propionibacterium phage PHL067M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29377 | 54.267 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578764 | Propionibacterium phage PHL064M02 | Propionibacterium phage PHL064M02 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29407 | 53.906 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578763 | Propionibacterium phage PHL064M01 | Propionibacterium phage PHL064M01 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29424 | 53.915 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578762 | Propionibacterium phage PHL055N00 | Propionibacterium phage PHL055N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29264 | 54.159 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578761 | Propionibacterium phage PHL041M10 | Propionibacterium phage PHL041M10 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29412 | 54.27 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578760 | Propionibacterium phage PHL030N00 | Propionibacterium phage PHL030N00 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29423 | 53.91 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578759 | Propionibacterium phage PHL025M00 | Propionibacterium phage PHL025M00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29496 | 53.984 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KJ578758 | Propionibacterium phage PHL009M11 | Propionibacterium phage PHL009M11 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29503 | 53.937 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KP687432 | Staphylococcus phage IME-SA2 | Staphylococcus phage IME-SA2 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140906 | 30.328 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KP687431 | Staphylococcus phage IME-SA1 | Staphylococcus phage IME-SA1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140218 | 30.328 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KP399678 | Listeria phage vB_LmoS_293 | Listeria phage vB_LmoS_293 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40759 | 36.861 | Unclassified | Unclassified | Siphoviridae | Listeria |
| KP399677 | Listeria phage vB_LmoS_188 | Listeria phage vB_LmoS_188 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38392 | 35.872 | Unclassified | Unclassified | Siphoviridae | Listeria |
| HF937074 | Staphylococcus phage PSa3 | Staphylococcus phage PSa3 Staphylococcus virus PSa3 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17602 | 29.588 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| KP313531 | Citrobacter phage phiCFP-1 | Citrobacter phage phiCFP-1 Citrobacter virus CFP1 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38625 | 50.278 | Teetrevirus | Studiervirinae | Autographiviridae | Citrobacter |
| KP791807 | Enterobacter phage E-4 | Enterobacter phage E-4 unclassified Teetrevirus Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39142 | 50.521 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| KP791806 | Enterobacter phage E-3 | Enterobacter phage E-3 Enterobacter virus E3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31522 | 50.771 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| KP791805 | Enterobacter phage E-2 | Enterobacter phage E-2 Enterobacter virus E2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36051 | 50.639 | Teetrevirus | Studiervirinae | Autographiviridae | Enterobacter |
| KP792622 | Verrucomicrobia phage P8625 | Verrucomicrobia phage P8625 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32894 | 50.967 | Unclassified | Unclassified | Siphoviridae | Verrucomicrobia |
| KP027447 | Staphylococcus phage phiIPLA-C1C | Staphylococcus phage phiIPLA-C1C Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140961 | 27.96 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KP027446 | Staphylococcus phage phiIPLA-RODI | Staphylococcus phage phiIPLA-RODI Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142348 | 30.425 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KP308307 | Escherichia phage 172-1 | Escherichia phage 172-1 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77266 | 41.984 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| KF835987 | Pectobacterium bacteriophage PM2 | Pectobacterium bacteriophage PM2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170286 | 34.785 | Unclassified | Unclassified | Myoviridae | Pectobacterium |
| KP735928 | Staphylococcus phage IME-SA4 | Staphylococcus phage IME-SA4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41843 | 33.982 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KP861231 | Acinetobacter phage YMC11/12/R1215 | Acinetobacter phage YMC11/12/R1215 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44866 | 37.594 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KP861229 | Acinetobacter phage YMC11/12/R2315 | Acinetobacter phage YMC11/12/R2315 unclassified Obolenskvirus Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44846 | 37.6 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KM262191 | Leuconostoc phage Ln-8 | Leuconostoc phage Ln-8 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28921 | 36.14 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KM262192 | Leuconostoc phage Ln-9 | Leuconostoc phage Ln-9 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28527 | 36.439 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| HG796219 | Burkholderia phage Bp-AMP2 | Burkholderia phage Bp-AMP2 unclassified Ampunavirus Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42492 | 61.758 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| HE858210 | Escherichia virus RB43 | Escherichia virus RB43 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179836 | 43.171 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| FR775895 | Enterobacteria phage phi92 | Enterobacteria phage phi92 unclassified Vequintavirinae Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148612 | 37.434 | Unclassified | Vequintavirinae | Myoviridae | Enterobacteria |
| KP209285 | Staphylococcus phage phiNM4-gamma4 | Staphylococcus phage phiNM4-gamma4 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40365 | 35.102 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| CP004084 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| KP708986 | Klebsiella phage vB_KpnP_SU552A | Klebsiella phage vB_KpnP_SU552A Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43595 | 54.153 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KP708985 | Klebsiella phage vB_KpnP_SU503 | Klebsiella phage vB_KpnP_SU503 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43809 | 53.749 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| JX403940 | Acinetobacter phage YMC/09/02/B1251 | Acinetobacter phage YMC/09/02/B1251 Acinetobacter virus B1251 Vieuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45364 | 39.053 | Vieuvirus | Unclassified | Siphoviridae | Acinetobacter |
| KP703175 | ANMV-1 virus | ANMV-1 virus unclassified archaeal viruses Viruses | 38465 | 44.492 | Unclassified | Unclassified | Unclassified | Unspecified |
| KP010268 | Shigella phage SfMu | Shigella phage SfMu Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37146 | 51.946 | Muvirus | Unclassified | Myoviridae | Shigella |
| KM236244 | Salmonella virus Stitch | Salmonella virus Stitch Epseptimavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123475 | 40.307 | Epseptimavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| KM236248 | Bacillus phage Pookie | Bacillus phage Pookie Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40214 | 40.573 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KM236247 | Bacillus phage Pascal | Bacillus phage Pascal Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39639 | 39.87 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KJ617393 | Streptococcus phage SOCP | Streptococcus phage SOCP Cepunavirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19347 | 38.823 | Cepunavirus | Unclassified | Salasmaviridae | Streptococcus |
| KP087951 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5256 | 52.683 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087937 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4634 | 43.116 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087938 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4426 | 50.926 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087939 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4265 | 46.846 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087940 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4355 | 50.792 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087941 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4331 | 61.071 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087942 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4445 | 46.929 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087943 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6172 | 47.813 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087944 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4514 | 41.117 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087945 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4314 | 57.603 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087946 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4406 | 60.826 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087947 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5713 | 46.981 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087948 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4817 | 41.52 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087949 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4180 | 56.029 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087950 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6438 | 44.237 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087952 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5310 | 47.495 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087953 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4342 | 39.06 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087954 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4700 | 49.362 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087955 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4507 | 50.055 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087956 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4790 | 41.19 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP087957 | Eel River basin pequenovirus | Eel River basin pequenovirus unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6083 | 30.659 | Unclassified | Unclassified | Microviridae | Unspecified |
| KP658157 | Klebsiella phage 1513 | Klebsiella phage 1513 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49462 | 50.611 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| LN681542 | Clostridium phage phiMMP03 | Clostridium phage phiMMP03 Clostridioides virus phiMMP03 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52261 | 28.87 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| LN681541 | Clostridium phage phiMMP01 | Clostridium phage phiMMP01 Clostridioides virus phiMMP01 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44461 | 28.92 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| LN681540 | Clostridium phage phiCD506 | Clostridium phage phiCD506 Clostridioides virus phiCD506 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33274 | 29.612 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| LN681539 | Clostridium phage phiCD505 | Clostridium phage phiCD505 Clostridioides virus phiCD505 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49316 | 29.384 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| LN681538 | Clostridium phage phiCD481-1 | Clostridium phage phiCD481-1 Clostridioides virus phiCD481-1 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32846 | 30.25 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| LN681536 | Clostridium phage phiCD146 | Clostridium phage phiCD146 Clostridioides virus phiCD146 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41507 | 30.689 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| LN681535 | Clostridium phage phiCD111 | Clostridium phage phiCD111 Clostridioides virus phiCD111 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41560 | 30.893 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| LN681534 | Clostridium phage phiCD24-1 | Clostridium phage phiCD24-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44129 | 28.503 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| HF543949 | Pseudomonas phage vB_PaeP_p2-10_Or1 | Pseudomonas phage vB_PaeP_p2-10_Or1 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44030 | 51.994 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KP063118 | Proteus phage pPM_01 | Proteus phage pPM_01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58546 | 46.862 | Unclassified | Unclassified | Siphoviridae | Proteus |
| KJ010548 | Bacillus phage Bp8p-T | Bacillus phage Bp8p-T Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151419 | 41.154 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ010547 | Bacillus phage Bp8p-C | Bacillus phage Bp8p-C Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151417 | 41.153 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| AP014715 | Edwardsiella phage PEi26 | Edwardsiella phage PEi26 unclassified Kanagawavirus Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177215 | 40.666 | Kanagawavirus | Unclassified | Myoviridae | Edwardsiella |
| AP014714 | Edwardsiella phage PEi20 | Edwardsiella phage PEi20 Edwardsiella virus PEi20 Kanagawavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177643 | 40.637 | Kanagawavirus | Unclassified | Myoviridae | Edwardsiella |
| KM058087 | Cronobacter phage vB_CsaP_Ss1 | Cronobacter phage vB_CsaP_Ss1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42205 | 46.068 | Unclassified | Unclassified | Podoviridae | Cronobacter |
| HE978309 | Enterobacteria phage GEC-3S | Enterobacteria phage GEC-3S Escherichia virus GEC3S Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163424 | 40.549 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| KM370384 | Lelliottia phage phD2B | Lelliottia phage phD2B Lelliottia virus phD2B Tuodvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44366 | 51.003 | Tuodvirus | Molineuxvirinae | Autographiviridae | Lelliottia |
| KP280063 | Vibrio phage phi 3 | Vibrio phage phi 3 Vibrio virus phi3 Jesfedecavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 116138 | 42.827 | Jesfedecavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| KP280062 | Vibrio phage phi 1 | Vibrio phage phi 1 Vibrio virus phi1 Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66708 | 34.503 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| KP202158 | Yersinia phage vB_YenM_TG1 | Yersinia phage vB_YenM_TG1 Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162101 | 34.56 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| KM607004 | Enterobacteria phage RB68 | Enterobacteria phage RB68 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168401 | 35.252 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM607003 | Enterobacteria phage RB59 | Enterobacteria phage RB59 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168966 | 35.293 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM607002 | Enterobacteria phage RB55 | Enterobacteria phage RB55 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168896 | 35.291 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM607001 | Enterobacteria phage RB33 | Enterobacteria phage RB33 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166007 | 35.335 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM607000 | Enterobacteria phage RB27 | Enterobacteria phage RB27 Enterobacteria virus RB27 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165179 | 35.485 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606999 | Enterobacteria phage RB10 | Enterobacteria phage RB10 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168401 | 35.373 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606998 | Enterobacteria phage RB9 | Enterobacteria phage RB9 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168395 | 35.372 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606997 | Enterobacteria phage RB7 | Enterobacteria phage RB7 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168395 | 35.372 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606996 | Enterobacteria phage RB6 | Enterobacteria phage RB6 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168394 | 35.37 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606995 | Enterobacteria phage RB5 | Enterobacteria phage RB5 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168394 | 35.37 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| KM606994 | Escherichia phage RB3 | Escherichia phage RB3 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168402 | 35.374 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JQ067085 | Pseudomonas phage PaMx73 | Pseudomonas phage PaMx73 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36570 | 64.192 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KM236246 | Bacillus phage Moonbeam | Bacillus phage Moonbeam Bacillus virus Moonbeam Moonbeamvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161239 | 40.132 | Moonbeamvirus | Bastillevirinae | Herelleviridae | Bacillus |
| KM236245 | Bacillus phage Mater | Bacillus phage Mater Bacillus virus Mater Matervirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164302 | 39.466 | Matervirus | Bastillevirinae | Herelleviridae | Bacillus |
| AB937974 | Ralstonia phage RS603 | Ralstonia phage RS603 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7679 | 59.37 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| KM272358 | Salmonella phage LSPA1 | Salmonella phage LSPA1 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41880 | 50.487 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KM236237 | Citrobacter phage Miller | Citrobacter phage Miller Citrobacter virus Miller Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178171 | 43.13 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Citrobacter |
| KM000061 | Pseudoalteromonas phage B8b | Pseudoalteromonas phage B8b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19353 | 43.942 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KJ944830 | Pseudoalteromonas phage B8b | Pseudoalteromonas phage B8b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 20209 | 43.397 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KM458633 | Salmonella virus Chi | Salmonella virus Chi Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59578 | 56.511 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| JX415536 | Escherichia phage ECBP2 | Escherichia phage ECBP2 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77315 | 42.42 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| JX415535 | Escherichia phage ECBP1 | Escherichia phage ECBP1 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69855 | 42.73 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| AB967974 | Aurantimonas phage AmM-1 | Aurantimonas phage AmM-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47800 | 66.684 | Unclassified | Unclassified | Myoviridae | Aurantimonas |
| KP202969 | Achromobacter phage JWX | Achromobacter phage JWX Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49714 | 55.413 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| FO818745 | Escherichia phage RCS47 | Escherichia phage RCS47 Escherichia virus RCS47 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115154 | 49.35 | Punavirus | Unclassified | Myoviridae | Escherichia |
| KP208803 | Streptococcus phage Str-PAP-1 | Streptococcus phage Str-PAP-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36595 | 35.289 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KP202971 | Achromobacter phage JWF | Achromobacter phage JWF Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81541 | 60.138 | Unclassified | Unclassified | Siphoviridae | Achromobacter |
| KP202970 | Achromobacter phage 83-24 | Achromobacter phage 83-24 Steinhofvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48216 | 54.853 | Steinhofvirus | Unclassified | Siphoviridae | Achromobacter |
| KP233880 | Pseudomonas phage PhiCHU | Pseudomonas phage PhiCHU Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45626 | 52.021 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KF806588 | Erwinia phage Ea9-2 | Erwinia phage Ea9-2 Johnsonvirus Erskinevirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75568 | 47.041 | Johnsonvirus | Erskinevirinae | Schitoviridae | Erwinia |
| KJ803031 | Dinoroseobacter phage vBDshPR2C | Dinoroseobacter phage vBDshPR2C unclassified Baltimorevirus Baltimorevirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74806 | 49.166 | Baltimorevirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| KM044272 | Escherichia phage vB_EcoP_SU10 | Escherichia phage vB_EcoP_SU10 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77327 | 42.063 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| KJ135004 | Escherichia phage Bp4 | Escherichia phage Bp4 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72583 | 42.883 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| KM233689 | Pseudomonas phage H70 | Pseudomonas phage H70 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37359 | 64.25 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KJ722067 | Propionibacterium phage Pacnes 2012-15 | Propionibacterium phage Pacnes 2012-15 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29741 | 54.423 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JN371769 | Synechococcus phage metaG-MbCM1 | Synechococcus phage metaG-MbCM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172879 | 39.762 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| KM983334 | Clostridium phage phiCTC2B | Clostridium phage phiCTC2B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38243 | 29.056 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KM983333 | Clostridium phage phiCTC2A | Clostridium phage phiCTC2A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47020 | 28.405 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| KM983332 | Clostridium phage phiCT19406C | Clostridium phage phiCT19406C Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35703 | 29.188 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| KM983331 | Clostridium phage phiCT19406B | Clostridium phage phiCT19406B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38243 | 29.048 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KM983330 | Clostridium phage phiCT19406A | Clostridium phage phiCT19406A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47020 | 28.396 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| KM983329 | Clostridium phage phiCT9441A | Clostridium phage phiCT9441A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46895 | 28.692 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KM983328 | Clostridium phage phiCT453B | Clostridium phage phiCT453B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36712 | 29.129 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KM983327 | Clostridium phage phiCT453A | Clostridium phage phiCT453A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42158 | 29.456 | Unclassified | Unclassified | Myoviridae | Clostridium |
| KM612265 | Vibrio phage J3 | Vibrio phage J3 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39782 | 50.545 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KM612264 | Vibrio phage J2 | Vibrio phage J2 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39530 | 50.582 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KM612263 | Vibrio phage H3 | Vibrio phage H3 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39530 | 50.539 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KM612261 | Vibrio phage H1 | Vibrio phage H1 unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39530 | 50.544 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KM612260 | Vibrio phage CJY | Vibrio phage CJY unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39542 | 50.556 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KM612259 | Vibrio phage QH | Vibrio phage QH unclassified Enhodamvirus Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39725 | 50.505 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| KJ545483 | Vibrio phage phi 2 | Vibrio phage phi 2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41476 | 46.063 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KJ572845 | Vibrio phage X29 | Vibrio phage X29 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41569 | 46.068 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KJ572844 | Vibrio phage 24 | Vibrio phage 24 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44395 | 45.39 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HG796221 | Burkholderia phage Bp-AMP4 | Burkholderia phage Bp-AMP4 unclassified Ampunavirus Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42112 | 61.785 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| KM514685 | Lactobacillus phage Ldl1 | Lactobacillus phage Ldl1 Lactobacillus virus Ldl1 Lidleunavirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74806 | 37.763 | Lidleunavirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| KP063541 | Enterobacteria phage P88 | Enterobacteria phage P88 unclassified Peduovirus Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35814 | 52.87 | Peduovirus | Peduovirinae | Myoviridae | Enterobacteria |
| DQ386162 | Streptococcus phage M102AD | Streptococcus phage M102AD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30664 | 39.62 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AY657002 | Streptococcus phage phi1207.3 | Streptococcus phage phi1207.3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52491 | 39.153 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KP007362 | Escherichia phage vB_EcoM_VR26 | Escherichia phage vB_EcoM_VR26 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171541 | 40.222 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| KP007361 | Escherichia phage vB_EcoM_VR25 | Escherichia phage vB_EcoM_VR25 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170822 | 40.418 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| KP007360 | Escherichia phage vB_EcoM_VR20 | Escherichia phage vB_EcoM_VR20 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170336 | 40.445 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| KP007359 | Escherichia phage vb_EcoM-VR5 | Escherichia phage vb_EcoM-VR5 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170473 | 39.303 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| JQ267518 | Enterobacter phage phiKDA1 | Enterobacter phage phiKDA1 Enterobacter virus KDA1 Koutsourovirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43804 | 51.678 | Koutsourovirus | Slopekvirinae | Autographiviridae | Enterobacter |
| HG793132 | Burkholderia phage Bp-AMP1 | Burkholderia phage Bp-AMP1 Burkholderia virus BpAMP1 Ampunavirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42409 | 61.749 | Ampunavirus | Okabevirinae | Autographiviridae | Burkholderia |
| KJ668714 | Escherichia phage vB_EcoM_112 | Escherichia phage vB_EcoM_112 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168470 | 35.275 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KJ725374 | Mycobacterium phage Guo1 | Mycobacterium phage Guo1 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40086 | 66.844 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KF925357 | Escherichia phage HY01 | Escherichia phage HY01 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166977 | 35.453 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| KM974184 | Pseudomonas phage YH6 | Pseudomonas phage YH6 unclassified Litunavirus Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73050 | 54.88 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| KF929199 | Staphylococcus phage vB_SepS_SEP9 | Staphylococcus phage vB_SepS_SEP9 Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92417 | 29.568 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| KF620435 | Shigella phage pSb-1 | Shigella phage pSb-1 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71629 | 42.742 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Shigella |
| KF547926 | Los Azufres archaeal virus 1 | Los Azufres archaeal virus 1 unclassified archaeal viruses Viruses | 21184 | 45.015 | Unclassified | Unclassified | Unclassified | Unspecified |
| KM253764 | Yersinia phage vB_YenP_AP5 | Yersinia phage vB_YenP_AP5 Yersinia virus AP5 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38646 | 50.696 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| KJ094032 | Enterococcus phage VD13 | Enterococcus phage VD13 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55073 | 40.003 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| KJ094031 | Listeria phage LP-124 | Listeria phage LP-124 Listeria virus LP064 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135764 | 35.874 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KP037007 | Erwinia phage phiEa2809 | Erwinia phage phiEa2809 Nezavisimistyvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162160 | 50.28 | Nezavisimistyvirus | Unclassified | Ackermannviridae | Erwinia |
| KP025626 | Pseudomonas phage Pf-10 | Pseudomonas phage Pf-10 Pseudomonas virus Pf10 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39167 | 56.474 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KM882824 | Streptococcus phage SpSL1 | Streptococcus phage SpSL1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33756 | 38.595 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KJ832078 | Enterobacteria phage Sf101 | Enterobacteria phage Sf101 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38742 | 47.445 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| KJ010489 | Enterococcus phage IME-EFm1 | Enterococcus phage IME-EFm1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42597 | 35.181 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KJ528544 | Lactococcus phage WP-2 | Lactococcus phage WP-2 Lactococcus virus WP2 Negarvirus Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18899 | 31.118 | Negarvirus | Unclassified | Rountreeviridae | Lactococcus |
| HM640230 | Clostridium phage CpV1 | Clostridium phage CpV1 Clostridium virus CpV1 Capvunavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16748 | 30.505 | Capvunavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| KM672662 | Acinetobacter phage YMC13/03/R2096 | Acinetobacter phage YMC13/03/R2096 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 98170 | 37.036 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| KJ743987 | Sinorhizobium phage phiLM21 | Sinorhizobium phage phiLM21 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50827 | 60.661 | Unclassified | Unclassified | Siphoviridae | Sinorhizobium |
| KM657822 | Escherichia phage vB_EcoM-VpaE1 | Escherichia phage vB_EcoM-VpaE1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88403 | 38.943 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KJ716335 | Dickeya phage RC-2014 | Dickeya phage RC-2014 Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155346 | 49.649 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| KJ159566 | Geobacillus phage GBK2 | Geobacillus phage GBK2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39078 | 43.093 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| KF623293 | Vibrio phage SHOU24 | Vibrio phage SHOU24 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77837 | 46.048 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KF591601 | Enterobacteria phage IME_EC2 | Enterobacteria phage IME_EC2 Escherichia virus EC2 Murrayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41510 | 59.157 | Murrayvirus | Unclassified | Siphoviridae | Escherichia |
| KF550303 | Escherichia phage 4MG | Escherichia phage 4MG Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148567 | 46.333 | Seunavirus | Vequintavirinae | Myoviridae | Escherichia |
| KF598856 | Staphylococcus phage YMC/09/04/R1988 | Staphylococcus phage YMC/09/04/R1988 Staphylococcus virus R1988 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44459 | 33.37 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| JX901189 | Mycobacterium phage FF47 | Mycobacterium phage FF47 Mapvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47724 | 58.637 | Mapvirus | Unclassified | Siphoviridae | Mycobacterium |
| FR719956 | Roseovarius Plymouth podovirus 1 | Roseovarius Plymouth podovirus 1 Plymouthvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74704 | 49.052 | Plymouthvirus | Rhodovirinae | Schitoviridae | Unspecified |
| FR682616 | Roseovarius sp. 217 phage 1 | Roseovarius sp. 217 phage 1 unclassified Plymouthvirus Plymouthvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74583 | 49.017 | Plymouthvirus | Rhodovirinae | Schitoviridae | Roseovarius |
| KM389210 | Pseudomonas phage DO4 | Pseudomonas phage DO4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49418 | 58.798 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| AB931172 | Ralstonia phage RS611 | Ralstonia phage RS611 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6386 | 62.12 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| KJ801817 | Enterococcus phage ECP3 | Enterococcus phage ECP3 Enterococcus virus ECP3 Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145518 | 35.908 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| KF692088 | Arthrobacter phage vB_ArS-ArV2 | Arthrobacter phage vB_ArS-ArV2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37372 | 62.731 | Unclassified | Unclassified | Siphoviridae | Arthrobacter |
| KM289195 | Streptococcus phage T12 | Streptococcus phage T12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37976 | 38.106 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KJ018214 | Shewanella sp. phage 3/49 | Shewanella sp. phage 3/49 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40161 | 41.966 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| KJ018213 | Shewanella sp. phage 1/44 | Shewanella sp. phage 1/44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49640 | 39.823 | Unclassified | Unclassified | Siphoviridae | Shewanella |
| KJ018212 | Shewanella sp. phage 1/41 | Shewanella sp. phage 1/41 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43510 | 42.703 | Unclassified | Unclassified | Myoviridae | Shewanella |
| KJ018211 | Shewanella sp. phage 1/40 | Shewanella sp. phage 1/40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139004 | 36.888 | Unclassified | Unclassified | Myoviridae | Shewanella |
| KJ018210 | Flavobacterium sp. phage 1/32 | Flavobacterium sp. phage 1/32 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42252 | 34.306 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| KJ018209 | Shewanella sp. phage 1/4 | Shewanella sp. phage 1/4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133824 | 36.855 | Unclassified | Unclassified | Myoviridae | Shewanella |
| KJ997912 | Enterobacteria phage MED1 | Enterobacteria phage MED1 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.801 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| KJ888149 | Staphylococcus phage MCE-2014 | Staphylococcus phage MCE-2014 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141907 | 30.385 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KF971864 | Escherichia phage phi191 | Escherichia phage phi191 Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61035 | 50.189 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| CP008753 | Burkholderia phage BEK | Burkholderia phage BEK Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37631 | 64.819 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| KM507819 | Escherichia phage 121Q | Escherichia phage 121Q Escherichia virus 121Q Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348532 | 34.114 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| KM581061 | Ruegeria phage DSS3-P1 | Ruegeria phage DSS3-P1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59601 | 64.133 | Unclassified | Unclassified | Siphoviridae | Ruegeria |
| KJ608189 | Leuconostoc phage LLC-1 | Leuconostoc phage LLC-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50014 | 39.365 | Unclassified | Unclassified | Siphoviridae | Leuconostoc |
| KJ608188 | Enterococcus phage EFC-1 | Enterococcus phage EFC-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40286 | 35.047 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| KM091444 | Lactococcus phage phi145 | Lactococcus phage phi145 Lactococcus virus phi145 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30862 | 34.901 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KM091443 | Lactococcus phage phi93 | Lactococcus phage phi93 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31841 | 34.971 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KM091442 | Lactococcus phage phi15 | Lactococcus phage phi15 Lactococcus virus phi15 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31945 | 34.575 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| AM265639 | Pseudomonas phage LKA1 | Pseudomonas phage LKA1 Stubburvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41593 | 60.902 | Stubburvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KJ173786 | Microbacterium phage vB_MoxS-ISF9 | Microbacterium phage vB_MoxS-ISF9 Microbacterium virus ISF9 Farahnazvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59254 | 62.759 | Farahnazvirus | Unclassified | Siphoviridae | Microbacterium |
| KF534715 | Pectobacterium phage PM1 | Pectobacterium phage PM1 Pectobacterium virus PM1 Suwonvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55098 | 44.925 | Suwonvirus | Cleopatravirinae | Chaseviridae | Pectobacterium |
| KC430106 | Bacillus phage BPS10C | Bacillus phage BPS10C Bacillus virus BPS10C Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159590 | 38.742 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| LK985322 | Clostridium phage phiCDHM19 | Clostridium phage phiCDHM19 Clostridioides virus phiCDHM19 Lubbockvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54295 | 28.249 | Lubbockvirus | Unclassified | Myoviridae | Clostridium |
| LK985321 | Clostridium phage phiCDHM14 | Clostridium phage phiCDHM14 Clostridioides virus phiCDHM14 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32651 | 29.54 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| HG796225 | Clostridium phage phiCDHM13 | Clostridium phage phiCDHM13 Clostridioides virus phiCDHM13 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33596 | 29.989 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| KM407600 | Shigella phage Shf125875 | Shigella phage Shf125875 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169062 | 37.558 | Mosigvirus | Tevenvirinae | Myoviridae | Shigella |
| KM373208 | Listeria phage WIL-1 | Listeria phage WIL-1 Listeria virus WIL1 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134369 | 35.988 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ847052 | Idiomarinaceae phage 1N2-2 | Idiomarinaceae phage 1N2-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34773 | 51.989 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| KM233261 | Sulfitobacter phage NYA-2014a | Sulfitobacter phage NYA-2014a Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42092 | 58.541 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| KM247287 | Escherichia phage J8-65 | Escherichia phage J8-65 Escherichia virus J8-65 Bonnellvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40981 | 55.687 | Bonnellvirus | Unclassified | Autographiviridae | Escherichia |
| KJ564038 | Lactobacillus phage Ld3 | Lactobacillus phage Ld3 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29616 | 41.646 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| KJ564037 | Lactobacillus phage Ld17 | Lactobacillus phage Ld17 Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32975 | 41.498 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| KJ603229 | Shigella phage POCJ13 | Shigella phage POCJ13 Diegovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62699 | 49.347 | Diegovirus | Sepvirinae | Podoviridae | Shigella |
| KF156339 | Synechococcus phage S-MbCM25 | Synechococcus phage S-MbCM25 unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176044 | 39.092 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| KF156338 | Synechococcus phage ACG-2014h | Synechococcus phage ACG-2014h Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 189311 | 40.491 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| JN371768 | Synechococcus phage ACG-2014c | Synechococcus phage ACG-2014c unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176043 | 39.119 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| AB983711 | Ralstonia phage RSJ5 | Ralstonia phage RSJ5 Ralstonia virus RSJ5 Risjevirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43745 | 60.661 | Risjevirus | Okabevirinae | Autographiviridae | Ralstonia |
| KM216423 | Staphylococcus phage P108 | Staphylococcus phage P108 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140807 | 30.217 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KM233151 | Escherichia phage EK99P-1 | Escherichia phage EK99P-1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44332 | 54.475 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| KM199771 | Mesorhizobium phage vB_MloP_Lo5R7ANS | Mesorhizobium phage vB_MloP_Lo5R7ANS Mesorhizobium virus Lo5R7ANS Pairvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45718 | 61.053 | Pairvirus | Unclassified | Autographiviridae | Mesorhizobium |
| KM199770 | Rhizobium phage vB_RleM_P10VF | Rhizobium phage vB_RleM_P10VF unclassified Ackermannviridae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156446 | 49.877 | Unclassified | Unclassified | Ackermannviridae | Rhizobium |
| KM224879 | Vibrio phage ICP2_2013_A_Haiti | Vibrio phage ICP2_2013_A_Haiti Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50440 | 42.797 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| KM224878 | Vibrio phage ICP2_2011_A | Vibrio phage ICP2_2011_A Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50250 | 42.476 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| KF728385 | Enterococcus phage IME_EF3 | Enterococcus phage IME_EF3 Enterococcus virus EF3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41687 | 34.598 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| CP003186 | Thermoanaerobacterium phage THSA-485A | Thermoanaerobacterium phage THSA-485A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41938 | 36.771 | Unclassified | Unclassified | Siphoviridae | Thermoanaerobacterium |
| AB920995 | Ralstonia phage RSJ2 | Ralstonia phage RSJ2 Ralstonia virus RSJ2 Risjevirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44360 | 60.911 | Risjevirus | Okabevirinae | Autographiviridae | Ralstonia |
| KJ804259 | Staphylococcus phage 6ec | Staphylococcus phage 6ec Sextaecvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93794 | 29.296 | Sextaecvirus | Unclassified | Siphoviridae | Staphylococcus |
| KJ473423 | Acinetobacter phage vB_AbaP_Acibel007 | Acinetobacter phage vB_AbaP_Acibel007 Acinetobacter virus Acibel007 Daemvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42654 | 41.166 | Daemvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KJ473422 | Acinetobacter phage vB_AbaM_Acibel004 | Acinetobacter phage vB_AbaM_Acibel004 unclassified Saclayvirus Saclayvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99730 | 37.27 | Saclayvirus | Unclassified | Myoviridae | Acinetobacter |
| AB981170 | Ralstonia phage RSMSuper | Ralstonia phage RSMSuper Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8956 | 59.669 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| AB981169 | Ralstonia phage RSY1 | Ralstonia phage RSY1 Ralstonia virus RSY1 Aresaunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40002 | 64.822 | Aresaunavirus | Peduovirinae | Myoviridae | Ralstonia |
| KJ668715 | Listeria phage LMTA-34 | Listeria phage LMTA-34 Listeria virus LMTA34 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113482 | 36.363 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ858521 | Aeromonas phage pAh6-C | Aeromonas phage pAh6-C Aeromonas virus pAh6C Pahsextavirus Nefertitivirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53744 | 52.826 | Pahsextavirus | Nefertitivirinae | Chaseviridae | Aeromonas |
| HQ317387 | Sulfitobacter phage phiCB2047-B | Sulfitobacter phage phiCB2047-B Sulfitobacter virus phiCB2047B Raunefjordenvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74485 | 42.983 | Raunefjordenvirus | Rhodovirinae | Schitoviridae | Sulfitobacter |
| KF475786 | Pseudomonas phage MP48 | Pseudomonas phage MP48 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36838 | 64.091 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KF356199 | Microcystis phage MaMV-DC | Microcystis phage MaMV-DC Microcystis virus MVDC Fukuivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169223 | 46.032 | Fukuivirus | Unclassified | Myoviridae | Microcystis |
| KJ417497 | Streptococcus phage Spn1 | Streptococcus phage Spn1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41323 | 39.939 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| HG799497 | Streptococcus phage DCC1738 | Streptococcus phage DCC1738 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38392 | 40.813 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| HG799496 | Streptococcus phage K13 | Streptococcus phage K13 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39360 | 40.615 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| HG799490 | Streptococcus phage IC1 | Streptococcus phage IC1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39965 | 41.121 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC012913 | Staphylococcus phage Team1 | Staphylococcus phage Team1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 140903 | 30.327 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX126920 | Listeria phage LP-037 | Listeria phage LP-037 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64756 | 36.559 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| JX120799 | Listeria phage LP-030-2 | Listeria phage LP-030-2 Psavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38275 | 34.751 | Psavirus | Unclassified | Siphoviridae | Listeria |
| JX126918 | Listeria phage LP-125 | Listeria phage LP-125 Listeria virus LP083-2 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135281 | 35.93 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| JX126919 | Listeria phage LP-110 | Listeria phage LP-110 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65132 | 36.342 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| KJ094020 | Listeria phage LP-026 | Listeria phage LP-026 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67150 | 36.258 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| KJ094033 | Listeria phage LP-048 | Listeria phage LP-048 Listeria virus LP048 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133048 | 35.966 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ094030 | Listeria phage LP-083-2 | Listeria phage LP-083-2 Listeria virus LP083-2 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135831 | 35.868 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ094029 | Listeria phage LP-064 | Listeria phage LP-064 Listeria virus LP064 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135279 | 35.926 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ094023 | Listeria phage LP-101 | Listeria phage LP-101 Listeria virus LP101 Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43767 | 35.451 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| KJ094022 | Listeria phage LP-030-3 | Listeria phage LP-030-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41156 | 36.554 | Unclassified | Unclassified | Siphoviridae | Listeria |
| KJ094021 | Listeria phage LP-114 | Listeria phage LP-114 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66676 | 36.434 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| AB910602 | Xanthomonas phage XacF1 | Xanthomonas phage XacF1 Coriovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7325 | 58.171 | Coriovirus | Unclassified | Inoviridae | Xanthomonas |
| KJ564036 | Lactobacillus phage Ld25A | Lactobacillus phage Ld25A Cequinquevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32799 | 41.663 | Cequinquevirus | Unclassified | Siphoviridae | Lactobacillus |
| JX976549 | Acinetobacter phage vB_AbaM-IME-AB2 | Acinetobacter phage vB_AbaM-IME-AB2 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43665 | 37.497 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KF771239 | Escherichia phage vB_EcoS_AKS96 | Escherichia phage vB_EcoS_AKS96 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45746 | 43.943 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KF771238 | Escherichia phage vB_EcoS_AHS24 | Escherichia phage vB_EcoS_AHS24 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46440 | 43.811 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KF771237 | Escherichia phage vB_EcoS_AHP42 | Escherichia phage vB_EcoS_AHP42 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46847 | 43.977 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KF771236 | Escherichia phage bV_EcoS_AHP24 | Escherichia phage bV_EcoS_AHP24 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46719 | 43.83 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KF554508 | Bacillus phage CP-51 | Bacillus phage CP-51 Siminovitchvirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138658 | 40.86 | Siminovitchvirus | Spounavirinae | Herelleviridae | Bacillus |
| KF301602 | Caulobacter phage Cr30 | Caulobacter phage Cr30 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155997 | 37.842 | Unclassified | Unclassified | Myoviridae | Caulobacter |
| KF381361 | Sinorhizobium phage phiM12 | Sinorhizobium phage phiM12 Emdodecavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194701 | 49.013 | Emdodecavirus | Unclassified | Myoviridae | Sinorhizobium |
| KC595279 | Staphylococcus phage StauST398-5 | Staphylococcus phage StauST398-5 Staphylococcus virus StauST398-5 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43301 | 35.135 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KC595278 | Staphylococcus phage StauST398-4 | Staphylococcus phage StauST398-4 Staphylococcus virus StauST398-4 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42906 | 33.084 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| JX013863 | Staphylococcus phage StauST398-1 | Staphylococcus phage StauST398-1 Staphylococcus virus StauST398-1 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45242 | 34.464 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| HQ317384 | Sulfitobacter phage pCB2047-C | Sulfitobacter phage pCB2047-C Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40931 | 58.994 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| HQ332142 | Sulfitobacter phage pCB2047-A | Sulfitobacter phage pCB2047-A Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40929 | 58.802 | Unclassified | Unclassified | Podoviridae | Sulfitobacter |
| KF669663 | Bacillus phage Staley | Bacillus phage Staley Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81656 | 35.346 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| KF669662 | Bacillus phage Spock | Bacillus phage Spock Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164297 | 37.621 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KF669661 | Bacillus phage Slash | Bacillus phage Slash Slashvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80382 | 35.232 | Slashvirus | Unclassified | Siphoviridae | Bacillus |
| KF669660 | Bacillus phage Pony | Bacillus phage Pony Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39844 | 40.699 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KF669658 | Acinetobacter phage Presley | Acinetobacter phage Presley Acinetobacter virus Presley Presleyvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77792 | 37.766 | Presleyvirus | Unclassified | Schitoviridae | Acinetobacter |
| KF669656 | Acinetobacter phage Petty | Acinetobacter phage Petty Acinetobacter virus Petty Pettyvirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40739 | 42.186 | Pettyvirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| KF669655 | Bacillus phage Page | Bacillus phage Page Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39874 | 40.706 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| KF669654 | Salmonella phage Maynard | Salmonella phage Maynard Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154701 | 45.552 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KF669653 | Salmonella phage Marshall | Salmonella phage Marshall Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156338 | 45.587 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| KF669652 | Bacillus phage Grass | Bacillus phage Grass Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156648 | 42.247 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KF669650 | Clavibacter phage CN1A | Clavibacter phage CN1A Clavibacter virus CN1A Cinunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56789 | 62.088 | Cinunavirus | Unclassified | Siphoviridae | Clavibacter |
| KF669649 | Bacillus phage CampHawk | Bacillus phage CampHawk Okubovirus Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146193 | 40.195 | Okubovirus | Spounavirinae | Herelleviridae | Bacillus |
| KF669647 | Bacillus phage BigBertha | Bacillus phage BigBertha Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165238 | 37.773 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| JX100809 | Caulobacter virus Swift | Caulobacter virus Swift Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219216 | 66.073 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX100810 | Caulobacter phage CcrColossus | Caulobacter phage CcrColossus Caulobacter virus CcrColossus Colossusvirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 279967 | 62.173 | Colossusvirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX100811 | Caulobacter virus Karma | Caulobacter virus Karma unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 221828 | 66.207 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX100812 | Caulobacter virus Magneto | Caulobacter virus Magneto unclassified Shapirovirus Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218929 | 66.15 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX100813 | Caulobacter virus phiCbK | Caulobacter virus phiCbK Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 215710 | 66.153 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX100814 | Caulobacter virus Rogue | Caulobacter virus Rogue Phicbkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 223720 | 66.105 | Phicbkvirus | Unclassified | Siphoviridae | Caulobacter |
| JN882285 | Cronobacter phage vB_CsaM_GAP32 | Cronobacter phage vB_CsaM_GAP32 Cronobacter virus GAP32 Mimasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 358663 | 35.551 | Mimasvirus | Unclassified | Myoviridae | Cronobacter |
| HQ317393 | Vibrio phage nt-1 | Vibrio phage nt-1 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 247511 | 41.327 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| KF676640 | Lactococcus phage SK1833 | Lactococcus phage SK1833 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28452 | 34.511 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KF302037 | Pseudoalteromonas phage HP1 | Pseudoalteromonas phage HP1 Pseudoalteromonas virus HP1 Melvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45035 | 44.665 | Melvirus | Unclassified | Zobellviridae | Pseudoalteromonas |
| KF302036 | Pseudoalteromonas phage HS2 | Pseudoalteromonas phage HS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38208 | 40.232 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KF302035 | Pseudoalteromonas phage HS6 | Pseudoalteromonas phage HS6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35330 | 44.874 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KF302034 | Pseudoalteromonas phage HM1 | Pseudoalteromonas phage HM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129439 | 35.733 | Unclassified | Unclassified | Myoviridae | Pseudoalteromonas |
| KF302033 | Pseudoalteromonas phage HS1 | Pseudoalteromonas phage HS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37211 | 40.534 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KF302032 | Pseudoalteromonas phage HS5 | Pseudoalteromonas phage HS5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37730 | 40.398 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KF800937 | Vibrio phage AS51 | Vibrio phage AS51 Vibrio virus AS51 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42542 | 43.47 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| KF322026 | Vibrio phage phi-A318 | Vibrio phage phi-A318 Vibrio virus A318 Kaohsiungvirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42544 | 43.47 | Kaohsiungvirus | Colwellvirinae | Autographiviridae | Vibrio |
| LK392619 | Streptococcus phage CP-7 | Streptococcus phage CP-7 Cepunavirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19741 | 38.544 | Cepunavirus | Unclassified | Salasmaviridae | Streptococcus |
| HG818824 | Citrobacter phage CR8 | Citrobacter phage CR8 unclassified Kayfunavirus Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39651 | 49.681 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| HG818823 | Citrobacter phage CR44b | Citrobacter phage CR44b Citrobacter virus CR44b Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39207 | 50.491 | Kayfunavirus | Studiervirinae | Autographiviridae | Citrobacter |
| KJ596420 | Staphylococcus phage phiSa119 | Staphylococcus phage phiSa119 unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42600 | 33.404 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KF562341 | Escherichia phage vB_EcoM_PhAPEC2 | Escherichia phage vB_EcoM_PhAPEC2 Escherichia virus PhAPEC2 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167318 | 37.725 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| KF562340 | Escherichia phage vB_EcoP_PhAPEC7 | Escherichia phage vB_EcoP_PhAPEC7 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71778 | 43.345 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| KF192075 | Escherichia phage vB_EcoP_PhAPEC5 | Escherichia phage vB_EcoP_PhAPEC5 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71248 | 43.492 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| FM164764 | Stygiolobus rod-shaped virus | Stygiolobus rod-shaped virus Azorudivirus SRV Azorudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 28096 | 29.275 | Azorudivirus | Unclassified | Rudiviridae | Stygiolobus |
| KJ817802 | Acinetobacter phage YMC-13-01-C62 | Acinetobacter phage YMC-13-01-C62 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44844 | 37.599 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| KJ591605 | Listeria phage LMTA-57 | Listeria phage LMTA-57 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136589 | 35.848 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ591604 | Listeria phage LMTA-148 | Listeria phage LMTA-148 Listeria virus LMTA148 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132541 | 36.027 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ586795 | Listeria phage LMTA-94 | Listeria phage LMTA-94 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136468 | 35.841 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ586794 | Listeria phage LMTA-34 | Listeria phage LMTA-34 Listeria virus LMTA34 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138036 | 35.87 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ451625 | Bacillus phage Bcp1 | Bacillus phage Bcp1 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152778 | 39.757 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ433974 | Mycobacterium phage Hosp | Mycobacterium phage Hosp unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70667 | 69.886 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ433976 | Mycobacterium phage Jolie1 | Mycobacterium phage Jolie1 unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71058 | 69.877 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ433975 | Mycobacterium phage 40BC | Mycobacterium phage 40BC unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71565 | 70.005 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ410133 | Mycobacterium phage Jolie2 | Mycobacterium phage Jolie2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44306 | 68.065 | Unclassified | Unclassified | Siphoviridae | Mycobacterium |
| KJ028219 | Mycobacterium phage 32HC | Mycobacterium phage 32HC Trigintaduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50781 | 65.731 | Trigintaduovirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ192399 | Vibrio phage CHOED | Vibrio phage CHOED Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66316 | 43.679 | Unclassified | Unclassified | Podoviridae | Vibrio |
| KC013029 | Leuconostoc phage phiLNTR3 | Leuconostoc phage phiLNTR3 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28015 | 36.052 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013028 | Leuconostoc phage phiLNTR2 | Leuconostoc phage phiLNTR2 unclassified Unaquatrovirus Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28336 | 36.014 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013027 | Leuconostoc phage LN34 | Leuconostoc phage LN34 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28022 | 35.972 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013026 | Leuconostoc phage LN25 | Leuconostoc phage LN25 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28427 | 36.191 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013025 | Leuconostoc phage phiLN12 | Leuconostoc phage phiLN12 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28193 | 36.637 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013024 | Leuconostoc phage phiLN6B | Leuconostoc phage phiLN6B Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25740 | 36.465 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013022 | Leuconostoc phage LN03 | Leuconostoc phage LN03 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26752 | 36.779 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KF766114 | Staphylococcus virus K | Staphylococcus virus K Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148317 | 30.386 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| HG315669 | Stenotrophomonas phage SMA6 | Stenotrophomonas phage SMA6 Scuticavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7648 | 62.631 | Scuticavirus | Unclassified | Inoviridae | Stenotrophomonas |
| HG007973 | Stenotrophomonas phage SMA7 | Stenotrophomonas phage SMA7 Subteminivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7069 | 62.343 | Subteminivirus | Unclassified | Inoviridae | Stenotrophomonas |
| KC954775 | Cronobacter phage S13 | Cronobacter phage S13 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182145 | 40.199 | Unclassified | Tevenvirinae | Myoviridae | Cronobacter |
| KC954774 | Cronobacter phage CR8 | Cronobacter phage CR8 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149162 | 50.832 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| KJ535722 | Listeria phage LMSP-25 | Listeria phage LMSP-25 Listeria virus LMSP25 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138036 | 35.87 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ535721 | Listeria phage List-36 | Listeria phage List-36 Listeria virus List36 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131952 | 36.013 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| KJ140076 | Staphylococcus phage DW2 | Staphylococcus phage DW2 unclassified Phietavirus Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41941 | 35.977 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KJ567046 | Mycobacterium phage Damien | Mycobacterium phage Damien unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68386 | 57.604 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ507100 | Pseudomonas phage phiPSA1 | Pseudomonas phage phiPSA1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51090 | 58.565 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KJ507099 | Pseudomonas phage phiPSA2 | Pseudomonas phage phiPSA2 Pseudomonas virus PhiPSA2 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40472 | 57.371 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| KF767351 | Lactobacillus phage phi jlb1 | Lactobacillus phage phi jlb1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38269 | 37.359 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| JX507079 | Acidithiobacillus phage AcaML1 | Acidithiobacillus phage AcaML1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59363 | 65.472 | Unclassified | Unclassified | Myoviridae | Acidithiobacillus |
| AB930182 | Bacillus phage SPG24 | Bacillus phage SPG24 Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152069 | 42.18 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| KJ621082 | Dinoroseobacter phage DFL12phi1 | Dinoroseobacter phage DFL12phi1 Baltimorevirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75028 | 49.26 | Baltimorevirus | Rhodovirinae | Schitoviridae | Dinoroseobacter |
| KJ668713 | Escherichia phage e4/1c | Escherichia phage e4/1c Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47112 | 44.078 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KJ025957 | Serratia phage PS2 | Serratia phage PS2 Serratia virus PS2 Muldoonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167266 | 41.698 | Muldoonvirus | Unclassified | Myoviridae | Serratia |
| KJ619459 | Vibrio virus CTXphi | Vibrio virus CTXphi Affertcholeramvirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6446 | 44.307 | Affertcholeramvirus | Unclassified | Inoviridae | Vibrio |
| KF751797 | Campylobacter phage CJIE4-5 | Campylobacter phage CJIE4-5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36991 | 28.704 | Unclassified | Unclassified | Siphoviridae | Campylobacter |
| KF751796 | Campylobacter phage CJIE4-4 | Campylobacter phage CJIE4-4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35459 | 28.921 | Unclassified | Unclassified | Siphoviridae | Campylobacter |
| KF751795 | Campylobacter phage CJIE4-3 | Campylobacter phage CJIE4-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35417 | 28.947 | Unclassified | Unclassified | Siphoviridae | Campylobacter |
| KF751794 | Campylobacter phage CJIE4-2 | Campylobacter phage CJIE4-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36734 | 28.51 | Unclassified | Unclassified | Siphoviridae | Campylobacter |
| KF751793 | Campylobacter phage CJIE4-1 | Campylobacter phage CJIE4-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37940 | 28.461 | Unclassified | Unclassified | Siphoviridae | Campylobacter |
| AP013057 | Edwardsiella phage PEi21 | Edwardsiella phage PEi21 Yokohamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43378 | 52.57 | Yokohamavirus | Unclassified | Myoviridae | Edwardsiella |
| KC465901 | Pelagibacter phage HTVC019P | Pelagibacter phage HTVC019P Pelagibacter virus HTVC019P Pelagivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42084 | 34.046 | Pelagivirus | Unclassified | Autographiviridae | Pelagibacter |
| KC465900 | Pelagibacter phage HTVC011P | Pelagibacter phage HTVC011P Pelagibacter virus HTVC011P Stopavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39921 | 31.958 | Stopavirus | Unclassified | Autographiviridae | Pelagibacter |
| KC465899 | Pelagibacter phage HTVC008M | Pelagibacter phage HTVC008M Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147284 | 33.448 | Unclassified | Unclassified | Myoviridae | Pelagibacter |
| KC465898 | Pelagibacter phage HTVC010P | Pelagibacter phage HTVC010P Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34892 | 31.967 | Unclassified | Unclassified | Siphoviridae | Pelagibacter |
| KJ534580 | Nitrincola phage 1M3-16 | Nitrincola phage 1M3-16 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 82438 | 41.306 | Unclassified | Unclassified | Unclassified | Nitrincola |
| HG803181 | Vibrio phage PV94 | Vibrio phage PV94 Vibrio virus PV94 Vulnificusvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33828 | 48.17 | Vulnificusvirus | Peduovirinae | Myoviridae | Vibrio |
| KF765493 | Klebsiella phage F19 | Klebsiella phage F19 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43766 | 53.82 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| KJ127304 | Enterococcus phage AUEF3 | Enterococcus phage AUEF3 Enterococcus virus AUEF3 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41257 | 34.452 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| KJ127303 | Enterococcus phage VD13 | Enterococcus phage VD13 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55726 | 40.008 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| HG798806 | Pseudomonas phage vB_PaeP_Tr60_Ab31 | Pseudomonas phage vB_PaeP_Tr60_Ab31 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45550 | 57.104 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| KF981875 | Pseudomonas phage SPM-1 | Pseudomonas phage SPM-1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65729 | 54.967 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| KJ489013 | Lactococcus phage P162 | Lactococcus phage P162 Lactococcus virus P162 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55584 | 33.333 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| KJ489012 | Lactococcus phage P118 | Lactococcus phage P118 Lactococcus virus P118 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54173 | 33.495 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| KJ489011 | Lactococcus phage P092 | Lactococcus phage P092 Lactococcus virus P092 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53513 | 33.528 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| KJ489010 | Lactococcus phage P078 | Lactococcus phage P078 Lactococcus virus P078 Nevevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54452 | 33.516 | Nevevirus | Unclassified | Siphoviridae | Lactococcus |
| KJ477077 | Pseudomonas phage JD024 | Pseudomonas phage JD024 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37380 | 64.152 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| KC556898 | Brucella phage Wb | Brucella phage Wb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.171 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KC556897 | Brucella phage Tb | Brucella phage Tb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.184 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KC556896 | Brucella phage S708 | Brucella phage S708 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.171 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KC556895 | Brucella phage R/C | Brucella phage R/C Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.179 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KC556894 | Brucella phage Fz | Brucella phage Fz Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41142 | 48.187 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KC556893 | Brucella phage Bk | Brucella phage Bk Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.174 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KF626668 | Xylella phage Salvo | Xylella phage Salvo Xylella virus Salvo Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55601 | 63.04 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| KF626665 | Xylella phage Sano | Xylella phage Sano Xylella virus Sano Sanovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56147 | 62.361 | Sanovirus | Unclassified | Siphoviridae | Xylella |
| HG531805 | Clostridium phage CDMH1 | Clostridium phage CDMH1 Clostridioides virus CDMH1 Yongloolinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54279 | 28.446 | Yongloolinvirus | Unclassified | Myoviridae | Clostridium |
| KJ502657 | Vibrio phage Vc1 | Vibrio phage Vc1 Vibrio virus Vc1 Gutovirus Colwellvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44541 | 44.157 | Gutovirus | Colwellvirinae | Autographiviridae | Vibrio |
| KJ419279 | Enterobacteria phage 9g | Enterobacteria phage 9g Nonagvirus Queuovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56702 | 43.894 | Nonagvirus | Queuovirinae | Siphoviridae | Enterobacteria |
| KJ021043 | Klebsiella phage Kpn112 | Klebsiella phage Kpn112 unclassified Jedunavirus Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35560 | 48.765 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| JX080304 | Staphylococcus phage MSA6 | Staphylococcus phage MSA6 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148243 | 30.243 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX080305 | Staphylococcus phage P4W | Staphylococcus phage P4W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147590 | 30.392 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX080303 | Staphylococcus phage Fi200W | Staphylococcus phage Fi200W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 148481 | 30.393 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX080300 | Staphylococcus phage Staph1N | Staphylococcus phage Staph1N Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145647 | 30.412 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KC959568 | Flavobacterium phage 6H | Flavobacterium phage 6H Flavobacterium virus 6H Unahavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46978 | 31.604 | Unahavirus | Unclassified | Siphoviridae | Flavobacterium |
| EU418428 | Staphylococcus phage A5W | Staphylococcus phage A5W Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145542 | 30.416 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KJ174317 | Salmonella phage vB-SalM-SJ2 | Salmonella phage vB-SalM-SJ2 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152460 | 44.878 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| HG428758 | Brucella phage F1 | Brucella phage F1 Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41143 | 48.176 | Perisivirus | Unclassified | Podoviridae | Brucella |
| KF787095 | Achromobacter phage JWAlpha | Achromobacter phage JWAlpha Jwalphavirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72329 | 54.412 | Jwalphavirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| KF787094 | Achromobacter phage JWDelta | Achromobacter phage JWDelta unclassified Jwalphavirus Jwalphavirus Rothmandenesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73659 | 54.251 | Jwalphavirus | Rothmandenesvirinae | Schitoviridae | Achromobacter |
| KF856712 | Pseudomonas phage phiIBB-PAA2 | Pseudomonas phage phiIBB-PAA2 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45344 | 52.327 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KC481682 | Bacillus phage vB_BanS-Tsamsa | Bacillus phage vB_BanS-Tsamsa Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168876 | 34.323 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| JQ809663 | Stenotrophomonas phage Smp131 | Stenotrophomonas phage Smp131 Stenotrophomonas virus Smp131 Simpcentumvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33525 | 65.023 | Simpcentumvirus | Peduovirinae | Myoviridae | Stenotrophomonas |
| KJ433973 | Mycobacterium phage 39HC | Mycobacterium phage 39HC unclassified Bclasvirinae Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 71565 | 70.003 | Unclassified | Bclasvirinae | Siphoviridae | Mycobacterium |
| KJ395778 | Vibrio phage PVA1 | Vibrio phage PVA1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41529 | 43.697 | Unclassified | Unclassified | Podoviridae | Vibrio |
| KF296718 | Bacillus phage phiCM3 | Bacillus phage phiCM3 Bacillus virus phiCM3 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38772 | 35.482 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| KF623294 | Erwinia phage PhiEaH1 | Erwinia phage PhiEaH1 Erwinia virus EaH1 Iapetusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218339 | 52.341 | Iapetusvirus | Unclassified | Myoviridae | Erwinia |
| KF296717 | Bacillus phage proCM3 | Bacillus phage proCM3 unclassified Lwoffvirus Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43278 | 37.361 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| KF021268 | Staphylococcus phage phiIBB-SEP1 | Staphylococcus phage phiIBB-SEP1 Sepunavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139928 | 27.93 | Sepunavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KC542353 | Pseudoalteromonas phage TW1 | Pseudoalteromonas phage TW1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39940 | 40.188 | Unclassified | Unclassified | Siphoviridae | Pseudoalteromonas |
| KC787107 | Mycobacterium phage Bo4 | Mycobacterium phage Bo4 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39318 | 66.763 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KJ101592 | Enterobacter phage PG7 | Enterobacter phage PG7 Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 173276 | 39.822 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| KF981730 | Escherichia phage KBNP1711 | Escherichia phage KBNP1711 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76184 | 42.37 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| KF811200 | Acinetobacter phage IMEAB3 | Acinetobacter phage IMEAB3 Acinetobacter virus IMEAB3 Lokivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43050 | 45.475 | Lokivirus | Unclassified | Siphoviridae | Acinetobacter |
| HQ698922 | Acinetobacter phage ZZ1 | Acinetobacter phage ZZ1 Acinetobacter virus ZZ1 Zedzedvirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166687 | 34.409 | Zedzedvirus | Twarogvirinae | Myoviridae | Acinetobacter |
| HG424323 | Streptococcus phage 20617 | Streptococcus phage 20617 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48800 | 39.975 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| CP002495 | Enterococcus phage EF62phi | Enterococcus phage EF62phi Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30505 | 32.657 | Unclassified | Unclassified | Podoviridae | Enterococcus |
| GQ866233 | Aggregatibacter phage S1249 | Aggregatibacter phage S1249 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43970 | 42.443 | Unclassified | Unclassified | Myoviridae | Aggregatibacter |
| CP000315 | Clostridium phage phiSM101 | Clostridium phage phiSM101 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38092 | 28.129 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| CP002889 | Streptococcus phage YMC-2011 | Streptococcus phage YMC-2011 unclassified Moineauvirus Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40758 | 41.239 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| KJ003983 | IAS virus | IAS virus crAss-like viruses Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 99915 | 31.948 | Unclassified | Unclassified | Podoviridae | Unspecified |
| KF669657 | Bacillus phage poppyseed | Bacillus phage poppyseed Pagevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39874 | 40.708 | Pagevirus | Unclassified | Podoviridae | Bacillus |
| Z18946 | Mycobacterium virus L5 | Mycobacterium virus L5 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52297 | 62.258 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AB903967 | Staphylococcus phage phiSA12 | Staphylococcus phage phiSA12 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142094 | 30.31 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KF986247 | Mycobacterium phage Oaker | Mycobacterium phage Oaker unclassified Konstantinevirus Konstantinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69099 | 57.546 | Konstantinevirus | Unclassified | Siphoviridae | Mycobacterium |
| KF854250 | Vibrio phage VP_EVC_2013 | Vibrio phage VP_EVC_2013 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47283 | 59.336 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KF772234 | Edwardsiella phage eiAU-183 | Edwardsiella phage eiAU-183 unclassified Eiauvirus Eiauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43017 | 55.436 | Eiauvirus | Unclassified | Siphoviridae | Edwardsiella |
| KF772233 | Edwardsiella phage eiAU | Edwardsiella phage eiAU Eiauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43015 | 55.432 | Eiauvirus | Unclassified | Siphoviridae | Edwardsiella |
| KC131025 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4062 | 34.54 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KC131024 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4200 | 50.69 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KC131023 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4334 | 41.74 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KC131022 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4070 | 44.447 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| KC131021 | Marine gokushovirus | Marine gokushovirus unclassified Gokushovirinae Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4129 | 39.574 | Unclassified | Gokushovirinae | Microviridae | Unspecified |
| HG518155 | Pseudomonas phage TL | Pseudomonas phage TL Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45696 | 52.366 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| HG322870 | Sulfolobus monocaudavirus SMV1 | Sulfolobus monocaudavirus SMV1 unclassified Bicaudaviridae Bicaudaviridae Viruses | 48775 | 38.421 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| JX483881 | Rhizobium phage RHEph10 | Rhizobium phage RHEph10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 115042 | 60.323 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| JX483880 | Rhizobium phage RHEph09 | Rhizobium phage RHEph09 Rhizobium virus RHEph09 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45962 | 49.304 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| JX483879 | Rhizobium phage RHEph08 | Rhizobium phage RHEph08 Rhizobium virus RHEph08 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43619 | 49.435 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| JX483878 | Rhizobium phage RHEph06 | Rhizobium phage RHEph06 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53721 | 56.429 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| JX483877 | Rhizobium phage RHEph05 | Rhizobium phage RHEph05 unclassified Kleczkowskavirus Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50426 | 56.443 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| JX483876 | Rhizobium phage RHEph04 | Rhizobium phage RHEph04 Kleczkowskavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53018 | 56.422 | Kleczkowskavirus | Unclassified | Myoviridae | Rhizobium |
| JX483875 | Rhizobium phage RHEph03 | Rhizobium phage RHEph03 unclassified Cuernavacavirus Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45912 | 49.342 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| JX483874 | Rhizobium phage RHEph02 | Rhizobium phage RHEph02 Rhizobium virus RHEph02 Cuernavacavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46486 | 49.35 | Cuernavacavirus | Unclassified | Autographiviridae | Rhizobium |
| JX483873 | Rhizobium phage RHEph01 | Rhizobium phage RHEph01 Rhizobium virus RHEph01 Paadamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43444 | 59.177 | Paadamvirus | Unclassified | Autographiviridae | Rhizobium |
| AB720063 | Xanthomonas virus CP1 | Xanthomonas virus CP1 Klementvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43870 | 53.31 | Klementvirus | Unclassified | Siphoviridae | Xanthomonas |
| AB720064 | Xanthomonas citri phage CP2 | Xanthomonas citri phage CP2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42963 | 67.018 | Unclassified | Unclassified | Podoviridae | Xanthomonas |
| JX649100 | Mycobacterium phage Alex | Mycobacterium phage Alex Pegunavirus Bclasvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68910 | 66.523 | Pegunavirus | Bclasvirinae | Siphoviridae | Mycobacterium |
| KC131130 | Vibrio phage VP4B | Vibrio phage VP4B Vibrio virus VP4B Tidunavirus Gorgonvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 236053 | 43.256 | Tidunavirus | Gorgonvirinae | Myoviridae | Vibrio |
| KC131129 | Vibrio phage VH7D | Vibrio phage VH7D unclassified Schizotequatrovirus Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246964 | 41.306 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| JF900176 | Salmonella phage SPN9CC | Salmonella phage SPN9CC Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40128 | 47.326 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| KF614509 | Rhizobium phage vB_RleS_L338C | Rhizobium phage vB_RleS_L338C Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 109558 | 59.003 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| KF147927 | Oenococcus phage phi9805 | Oenococcus phage phi9805 Oenococcus virus phi9805 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46145 | 38.86 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| KC801932 | Escherichia phage Lw1 | Escherichia phage Lw1 Escherichia virus Lw1 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176227 | 43.491 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| KF183314 | Oenococcus phage phiS11 | Oenococcus phage phiS11 Oenococcus virus phiS11 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46243 | 38.603 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| KF183315 | Oenococcus phage phiS13 | Oenococcus phage phiS13 Oenococcus virus phiS13 Sozzivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43454 | 38.97 | Sozzivirus | Unclassified | Siphoviridae | Oenococcus |
| KF733017 | Enterococcus phage IME-EF4 | Enterococcus phage IME-EF4 Enterococcus virus EF4 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40692 | 34.624 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| KC710998 | Shigella phage pSf-1 | Shigella phage pSf-1 Shigella virus pSf1 Hanrivervirus Tempevirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51821 | 44.023 | Hanrivervirus | Tempevirinae | Drexlerviridae | Shigella |
| KF322032 | Enterobacteria phage fiAA91-ss | Enterobacteria phage fiAA91-ss Escherichia virus fiAA91ss Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33628 | 51.906 | Peduovirus | Peduovirinae | Myoviridae | Escherichia |
| EF177456 | Salmonella phage SETP3 | Salmonella phage SETP3 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42572 | 49.852 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KC862297 | Pseudomonas phage PAK_P1 | Pseudomonas phage PAK_P1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93198 | 49.498 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KC862296 | Pseudomonas phage P3_CHA | Pseudomonas phage P3_CHA Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88097 | 54.827 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KC862301 | Pseudomonas phage PAK_P5 | Pseudomonas phage PAK_P5 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88135 | 54.73 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KC862298 | Pseudomonas phage PAK_P2 | Pseudomonas phage PAK_P2 Pseudomonas virus PAKP2 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92495 | 49.288 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| KC862295 | Pseudomonas phage CHA_P1 | Pseudomonas phage CHA_P1 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88255 | 54.628 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| KF208639 | Bacillus phage Troll | Bacillus phage Troll Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163019 | 37.825 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| JX238501 | Bacillus phage phiAGATE | Bacillus phage phiAGATE Agatevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149844 | 40.966 | Agatevirus | Bastillevirinae | Herelleviridae | Bacillus |
| AY766464 | Lactococcus phage TP712 | Lactococcus phage TP712 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42073 | 35.12 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| KC853746 | Burkholderia phage JG068 | Burkholderia phage JG068 Burkholderia virus JG068 Mguuvirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41604 | 60.689 | Mguuvirus | Okabevirinae | Autographiviridae | Burkholderia |
| KC342645 | Staphylococcus phage JS01 | Staphylococcus phage JS01 Staphylococcus virus JS01 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43458 | 33.32 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| KF192053 | Enterococcus phage IMEEF1 | Enterococcus phage IMEEF1 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57081 | 40.045 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| AB472900 | Pseudomonas phage KPP10 | Pseudomonas phage KPP10 Nankokuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88322 | 54.794 | Nankokuvirus | Unclassified | Myoviridae | Pseudomonas |
| AB863625 | Ralstonia phage RSK1 | Ralstonia phage RSK1 Ralstonia virus RSK1 Firingavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40471 | 59.591 | Firingavirus | Unclassified | Podoviridae | Ralstonia |
| JX274647 | Staphylococcus phage SP6 | Staphylococcus phage SP6 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42902 | 34.539 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| JX274646 | Staphylococcus phage SP5 | Staphylococcus phage SP5 Staphylococcus virus SP5 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43305 | 34.446 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| HG380752 | Streptococcus phage TP-778L | Streptococcus phage TP-778L unclassified Brussowvirus Brussowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41757 | 38.978 | Brussowvirus | Unclassified | Siphoviridae | Streptococcus |
| JX997978 | Pseudomonas phage MPK6 | Pseudomonas phage MPK6 Pseudomonas virus MPK6 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42957 | 62.279 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KF700371 | Erwinia phage FE44 | Erwinia phage FE44 Erwinia virus FE44 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39860 | 48.62 | Berlinvirus | Studiervirinae | Autographiviridae | Erwinia |
| KC295538 | Escherichia phage PBECO4 | Escherichia phage PBECO4 Escherichia virus PBECO4 Asteriusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 348113 | 34.087 | Asteriusvirus | Unclassified | Myoviridae | Escherichia |
| JX880072 | Vibrio phage VPMS1 | Vibrio phage VPMS1 unclassified Kafunavirus Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42313 | 44.669 | Kafunavirus | Unclassified | Podoviridae | Vibrio |
| KF598865 | Cyanophage PP | Cyanophage PP Podoviridae Caudovirales Viruses | 42480 | 46.408 | Unclassified | Unclassified | Podoviridae | Unspecified |
| KC814930 | Shigella phage SfIV | Shigella phage SfIV Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39758 | 50.297 | Unclassified | Unclassified | Myoviridae | Shigella |
| KF279414 | Mycobacterium phage Fredward | Mycobacterium phage Fredward unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52282 | 61.241 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC677663 | Staphylococcus phage SA12 | Staphylococcus phage SA12 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42902 | 34.542 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| HE956704 | Lactobacillus phage phiAQ113 | Lactobacillus phage phiAQ113 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36566 | 36.52 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| JX501340 | Pseudomonas phage MPK7 | Pseudomonas phage MPK7 Pseudomonas virus MPK7 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42874 | 62.138 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| KF562865 | Salmonella phage SETP7 | Salmonella phage SETP7 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42749 | 49.901 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KF562864 | Salmonella phage SETP13 | Salmonella phage SETP13 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42665 | 49.788 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| HE681887 | Aeropyrum coil-shaped virus | Aeropyrum coil-shaped virus Alphaspiravirus Spiraviridae Viruses | 24893 | 46.668 | Alphaspiravirus | Unclassified | Spiraviridae | Aeropyrum |
| JX094499 | Salmonella virus Chi | Salmonella virus Chi Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59407 | 56.51 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| JN790865 | Bacillus phage B4 | Bacillus phage B4 Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162596 | 37.71 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| HF563658 | Streptococcus phage phiBHN167 | Streptococcus phage phiBHN167 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37375 | 38.341 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AB854109 | Ralstonia phage RSB3 | Ralstonia phage RSB3 Ralstonia virus RSB3 Jiaoyazivirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44578 | 61.032 | Jiaoyazivirus | Unclassified | Autographiviridae | Ralstonia |
| KC139520 | Salmonella phage FSL SP-076 | Salmonella phage FSL SP-076 Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72098 | 39.535 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| KC139517 | Salmonella phage FSL SP-058 | Salmonella phage FSL SP-058 Ithacavirus Humphriesvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72394 | 39.586 | Ithacavirus | Humphriesvirinae | Schitoviridae | Salmonella |
| KF589919 | Staphylococcus phage phiRS7 | Staphylococcus phage phiRS7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43394 | 34.267 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| KF361475 | Vibrio phage VPUSM 8 | Vibrio phage VPUSM 8 unclassified Longwoodvirus Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34145 | 48.783 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| KF010834 | Paenibacillus phage phiIBB_P123 | Paenibacillus phage phiIBB_P123 Paenibacillus virus P123 Fernvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41294 | 40.936 | Fernvirus | Unclassified | Siphoviridae | Paenibacillus |
| HM072038 | Bacillus phage phi105 | Bacillus phage phi105 Bacillus virus phi105 Spizizenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39318 | 42.673 | Spizizenvirus | Unclassified | Siphoviridae | Bacillus |
| KC736978 | Shigella phage SfII | Shigella phage SfII Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41475 | 49.174 | Unclassified | Unclassified | Myoviridae | Shigella |
| KF188458 | Bacillus phage Wip1 | Bacillus phage Wip1 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14319 | 36.846 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| JX316028 | Erwinia phage phiEaH2 | Erwinia phage phiEaH2 Erskinevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 243050 | 51.29 | Erskinevirus | Unclassified | Myoviridae | Erwinia |
| KC330681 | Bacillus phage Gemini | Bacillus phage Gemini Andromedavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49362 | 41.901 | Andromedavirus | Unclassified | Siphoviridae | Bacillus |
| JX570714 | Propionibacterium phage PHL112N00 | Propionibacterium phage PHL112N00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29266 | 54.483 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570713 | Propionibacterium phage PHL113M01 | Propionibacterium phage PHL113M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29200 | 54.096 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570712 | Propionibacterium phage PHL114L00 | Propionibacterium phage PHL114L00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29464 | 54.205 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570711 | Propionibacterium phage PHL066M04 | Propionibacterium phage PHL066M04 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29512 | 53.992 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570710 | Propionibacterium phage PHL071N05 | Propionibacterium phage PHL071N05 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29467 | 53.921 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570709 | Propionibacterium phage PHL067M10 | Propionibacterium phage PHL067M10 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29377 | 54.264 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570708 | Propionibacterium phage PHL115M02 | Propionibacterium phage PHL115M02 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29453 | 53.818 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570707 | Propionibacterium phage PHL085M01 | Propionibacterium phage PHL085M01 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29451 | 53.822 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570706 | Propionibacterium phage PHL037M02 | Propionibacterium phage PHL037M02 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29443 | 53.778 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570705 | Propionibacterium phage PHL060L00 | Propionibacterium phage PHL060L00 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29514 | 54.049 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570704 | Propionibacterium phage PHL010M04 | Propionibacterium phage PHL010M04 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29511 | 53.987 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570703 | Propionibacterium phage PHL073M02 | Propionibacterium phage PHL073M02 unclassified Pahexavirus Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29503 | 53.995 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| JX570702 | Propionibacterium phage PHL111M01 | Propionibacterium phage PHL111M01 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29140 | 54.331 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| KC911857 | Salmonella phage SPC32N | Salmonella phage SPC32N Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38689 | 50.151 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| KC911856 | Salmonella phage SPC32H | Salmonella phage SPC32H Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38689 | 50.156 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| HQ634187 | Cyanophage KBS-S-2A | Cyanophage KBS-S-2A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40658 | 49.068 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| KC237729 | Clostridium phage vB_CpeS-CP51 | Clostridium phage vB_CpeS-CP51 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39108 | 28.076 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| JQ780163 | Vibrio phage VP3 | Vibrio phage VP3 unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39481 | 42.618 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| HQ634178 | Synechococcus phage S-CAM8 | Synechococcus phage S-CAM8 unclassified Acionnavirus Acionnavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171407 | 39.333 | Acionnavirus | Unclassified | Myoviridae | Synechococcus |
| HQ317292 | Synechococcus phage S-RIM2 R1_1999 | Synechococcus phage S-RIM2 R1_1999 Synechococcus virus SRIM2 Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175430 | 42.155 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| HQ317291 | Synechococcus phage S-RIM2 R9_2006 | Synechococcus phage S-RIM2 R9_2006 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175419 | 42.161 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| HQ317290 | Synechococcus phage S-RIM2 R21_2007 | Synechococcus phage S-RIM2 R21_2007 unclassified Nerrivikvirus Nerrivikvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175430 | 42.167 | Nerrivikvirus | Unclassified | Myoviridae | Synechococcus |
| JN811560 | Pseudomonas phage JBD26 | Pseudomonas phage JBD26 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37840 | 64.115 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| CP000711 | Enterobacteria phage CUS-3 | Enterobacteria phage CUS-3 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40207 | 46.86 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| HQ317390 | Colwellia phage 9A | Colwellia phage 9A Colwellia virus 9A Franklinbayvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104936 | 36.611 | Franklinbayvirus | Unclassified | Siphoviridae | Colwellia |
| KC333879 | Enterobacteria phage phiJLA23 | Enterobacteria phage phiJLA23 Escherichia virus phiJLA23 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43017 | 44.19 | Rogunavirus | Rogunavirinae | Drexlerviridae | Escherichia |
| KC206276 | Escherichia phage EC1-UPM | Escherichia phage EC1-UPM Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70912 | 42.887 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| KC690136 | Escherichia phage JES2013 | Escherichia phage JES2013 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136910 | 43.572 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| JN673056 | Escherichia phage PhaxI | Escherichia phage PhaxI Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156628 | 44.533 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| GU071107 | Cyanophage P-SSP2 | Cyanophage P-SSP2 Synechococcus virus PSSP2 Tritonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45890 | 37.932 | Tritonvirus | Unclassified | Autographiviridae | Prochlorococcus |
| JX846612 | Staphylococcus phage vB_SauM_Remus | Staphylococcus phage vB_SauM_Remus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134643 | 29.973 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| KF114880 | Leptospira phage vB_LinZ_10-LE1 | Leptospira phage vB_LinZ_10-LE1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89607 | 39.436 | Unclassified | Unclassified | Myoviridae | Leptospira |
| KF114879 | Leptospira phage vB_LbrZ_5399-LE1 | Leptospira phage vB_LbrZ_5399-LE1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87308 | 39.377 | Unclassified | Unclassified | Myoviridae | Leptospira |
| KF114878 | Leptospira phage vb_LkmZ_Bejolso9-LE1 | Leptospira phage vb_LkmZ_Bejolso9-LE1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 47417 | 34.475 | Unclassified | Unclassified | Unclassified | Leptospira |
| KF114877 | Leptospira phage vB_LnoZ_CZ214-LE1 | Leptospira phage vB_LnoZ_CZ214-LE1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87887 | 33.96 | Unclassified | Unclassified | Myoviridae | Leptospira |
| KF114876 | Leptospira phage vB_LalZ_80412-LE1 | Leptospira phage vB_LalZ_80412-LE1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86537 | 39.082 | Unclassified | Unclassified | Myoviridae | Leptospira |
| KF044310 | Enterobacteria phage FL76 Tallahassee/FL/2012 | Enterobacteria phage FL76 Tallahassee/FL/2012 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.571 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| KF044309 | Enterobacteria phage FL68 Tallahassee/FL/2012 | Enterobacteria phage FL68 Tallahassee/FL/2012 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.589 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| JX409895 | Streptococcus phage JX01 | Streptococcus phage JX01 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43028 | 36.806 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JX297445 | Salmonella phage vB_SenS_AG11 | Salmonella phage vB_SenS_AG11 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41546 | 49.909 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| JF501022 | Klebsiella phage KP36 | Klebsiella phage KP36 Webervirus Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49797 | 50.702 | Webervirus | Unclassified | Drexlerviridae | Klebsiella |
| KC677662 | Salmonella virus iEPS5 | Salmonella virus iEPS5 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59254 | 56.308 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KC751414 | Pseudoalteromonas phage RIO-1 | Pseudoalteromonas phage RIO-1 Pseudoalteromonas virus RIO1 Melvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43882 | 44.722 | Melvirus | Unclassified | Zobellviridae | Pseudoalteromonas |
| KC685370 | Bacillus phage MG-B1 | Bacillus phage MG-B1 Bacillus virus MGB1 Klosterneuburgvirus Northropvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27190 | 30.747 | Klosterneuburgvirus | Northropvirinae | Salasmaviridae | Bacillus |
| KF055347 | Escherichia phage JH2 | Escherichia phage JH2 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 87712 | 38.816 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| KF030445 | Escherichia phage 1720a-02 | Escherichia phage 1720a-02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47343 | 52.265 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| KF208315 | Yersinia phage PST | Yersinia phage PST Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167785 | 35.32 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| KC139521 | Salmonella phage FSLSP004 | Salmonella phage FSLSP004 Salmonella virus FSLSP004 Stockinghallvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29742 | 52.838 | Stockinghallvirus | Peduovirinae | Myoviridae | Salmonella |
| KC139519 | Salmonella virus FSLSP030 | Salmonella virus FSLSP030 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59746 | 56.556 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KC139518 | Salmonella phage FSL SP-031 | Salmonella phage FSL SP-031 Cornellvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42215 | 51.067 | Cornellvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KC139516 | Salmonella phage FSL SP-016 | Salmonella phage FSL SP-016 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46698 | 50.158 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| KC139515 | Salmonella phage FSL SP-124 | Salmonella phage FSL SP-124 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59245 | 56.535 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KC139514 | Salmonella phage FSL SP-039 | Salmonella phage FSL SP-039 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59815 | 56.553 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KC139512 | Salmonella virus FSLSP088 | Salmonella virus FSLSP088 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59454 | 56.442 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| KC139511 | Salmonella phage FSL SP-101 | Salmonella phage FSL SP-101 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41873 | 50.269 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| AB823818 | Thermus phage phi OH2 | Thermus phage phi OH2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38099 | 44.689 | Unclassified | Unclassified | Myoviridae | Thermus |
| KF148055 | Salmonella phage Jersey | Salmonella phage Jersey Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43447 | 49.971 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| AB828698 | Ralstonia phage RSS30 | Ralstonia phage RSS30 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8576 | 61.719 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| HE962497 | Streptococcus phage SPQS1 | Streptococcus phage SPQS1 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58305 | 39.873 | Saphexavirus | Unclassified | Siphoviridae | Streptococcus |
| AP013029 | Bacillus phage phiNIT1 | Bacillus phage phiNIT1 Nitunavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 155631 | 42.118 | Nitunavirus | Bastillevirinae | Herelleviridae | Bacillus |
| GU557055 | Puniceispirillum phage HMO-2011 | Puniceispirillum phage HMO-2011 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55282 | 43.088 | Unclassified | Unclassified | Podoviridae | Puniceispirillum |
| JX564242 | Lactococcus phage Q33 | Lactococcus phage Q33 Lactococcus virus Q33 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31106 | 36.215 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| JX567312 | Lactococcus phage BM13 | Lactococcus phage BM13 Lactococcus virus BM13 Vedamuthuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30910 | 35.988 | Vedamuthuvirus | Unclassified | Siphoviridae | Lactococcus |
| JN175269 | Pseudomonas phage phiPto-bp6g | Pseudomonas phage phiPto-bp6g Pseudomonas virus Ptobp6g Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26499 | 42.711 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| KC117378 | Halovirus HSTV-1 | Halovirus HSTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 32189 | 60.297 | Halovirus | Unclassified | Unclassified | Unspecified |
| AB259123 | Ralstonia phage RSM1 | Ralstonia phage RSM1 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 9004 | 59.951 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| KC182552 | Lactococcus phage phi7 | Lactococcus phage phi7 Lactococcus virus phi7 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32382 | 34.217 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182551 | Lactococcus phage P680 | Lactococcus phage P680 Lactococcus virus P680 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29631 | 35.115 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182550 | Lactococcus lactis phage P475 | Lactococcus lactis phage P475 Lactococcus virus P475 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30961 | 34.324 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182549 | Lactococcus lactis phage p272 | Lactococcus lactis phage p272 Lactococcus virus p272 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30778 | 34.145 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182548 | Lactococcus lactis phage P113G | Lactococcus lactis phage P113G unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30796 | 34.147 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182547 | Lactococcus phage jm3 | Lactococcus phage jm3 Lactococcus virus jm3 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28674 | 34.428 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182546 | Lactococcus phage jm2 | Lactococcus phage jm2 Lactococcus virus jm2 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31090 | 34.31 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182545 | Lactococcus phage fd13 | Lactococcus phage fd13 Lactococcus virus fd13 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30674 | 34.717 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182544 | Lactococcus phage 936 | Lactococcus phage 936 Lactococcus virus 936 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27302 | 34.518 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182543 | Lactococcus lactis phage 645 | Lactococcus lactis phage 645 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29247 | 34.978 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| KC182542 | Lactococcus phage 340 | Lactococcus phage 340 Lactococcus virus 340 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32337 | 34.487 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| AB434711 | Ralstonia phage RSM3 | Ralstonia phage RSM3 Habenivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8929 | 59.648 | Habenivirus | Unclassified | Inoviridae | Ralstonia |
| JN255163 | Mannheimia phage vB_MhM_1152AP | Mannheimia phage vB_MhM_1152AP Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34719 | 41.568 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| HQ634195 | Vibrio phage eugene 12A10 | Vibrio phage eugene 12A10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138234 | 37.954 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ632860 | Vibrio phage jenny 12G5 | Vibrio phage jenny 12G5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40557 | 42.284 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| KC581799 | Streptococcus phage phiD12 | Streptococcus phage phiD12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50470 | 39.633 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348604 | Streptococcus phage phiS10 | Streptococcus phage phiS10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35373 | 41.797 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348603 | Streptococcus phage phiD12 | Streptococcus phage phiD12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49973 | 40.826 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348602 | Streptococcus phage phi891591 | Streptococcus phage phi891591 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34367 | 41.514 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348601 | Streptococcus phage phi7917 | Streptococcus phage phi7917 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36128 | 41.345 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348600 | Streptococcus phage phi5218 | Streptococcus phage phi5218 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38539 | 40.855 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348599 | Streptococcus phage phi30c | Streptococcus phage phi30c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42703 | 40.594 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC348598 | Streptococcus phage phi20c | Streptococcus phage phi20c Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33516 | 41.717 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC413988 | Streptococcus phage phiST1 | Streptococcus phage phiST1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38940 | 39.944 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC413987 | Streptococcus phage phiSS12 | Streptococcus phage phiSS12 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35598 | 41.561 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| KC438282 | Vibrio phage JA-1 | Vibrio phage JA-1 unclassified Pacinivirus Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69278 | 34.555 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| KC438283 | Vibrio phage VCO139 | Vibrio phage VCO139 Vibrio virus VCO139 Pacinivirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68964 | 34.593 | Pacinivirus | Unclassified | Schitoviridae | Vibrio |
| JF974315 | Rhizobium phage RR1-B | Rhizobium phage RR1-B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37378 | 58.272 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| JF974314 | Rhizobium phage RR1-A | Rhizobium phage RR1-A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53102 | 57.164 | Unclassified | Unclassified | Myoviridae | Rhizobium |
| JF974313 | Vibrio phage pYD38-B | Vibrio phage pYD38-B Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37324 | 43.077 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| JF974312 | Vibrio phage pYD38-A | Vibrio phage pYD38-A unclassified Roufvirus Roufvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47552 | 47.289 | Roufvirus | Unclassified | Siphoviridae | Vibrio |
| JF974294 | Aeromonas phage pIS4-A | Aeromonas phage pIS4-A Roufvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47624 | 47.279 | Roufvirus | Unclassified | Siphoviridae | Aeromonas |
| HQ332141 | Halorubrum phage CGphi46 | Halorubrum phage CGphi46 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39784 | 65.107 | Unclassified | Unclassified | Siphoviridae | Halorubrum |
| HQ332138 | Paenibacillus phage PG1 | Paenibacillus phage PG1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37644 | 42.4 | Unclassified | Unclassified | Siphoviridae | Paenibacillus |
| HQ317383 | Synechococcus phage S-IOM18 | Synechococcus phage S-IOM18 Synechococcus virus SIOM18 Tefnutvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171797 | 40.572 | Tefnutvirus | Unclassified | Myoviridae | Synechococcus |
| JN116825 | Rhodococcus phage REQ1 | Rhodococcus phage REQ1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51342 | 66.324 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| HQ632825 | Prochlorococcus phage P-SSM5 | Prochlorococcus phage P-SSM5 unclassified Salacisavirus Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252013 | 35.525 | Salacisavirus | Unclassified | Myoviridae | Prochlorococcus |
| HQ337021 | Prochlorococcus phage P-SSM3 | Prochlorococcus phage P-SSM3 Prochlorococcus virus PSSM3 Ronodorvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179063 | 36.731 | Ronodorvirus | Unclassified | Myoviridae | Prochlorococcus |
| KF005320 | Alteromonas phage vB_AmaP_AD45-P2 | Alteromonas phage vB_AmaP_AD45-P2 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 104036 | 43.211 | Unclassified | Unclassified | Schitoviridae | Alteromonas |
| KF005319 | Alteromonas phage vB_AmaP_AD45-P4 | Alteromonas phage vB_AmaP_AD45-P4 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 100619 | 43.191 | Unclassified | Unclassified | Schitoviridae | Alteromonas |
| KF005318 | Alteromonas phage vB_AmaP_AD45-P3 | Alteromonas phage vB_AmaP_AD45-P3 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101724 | 43.153 | Unclassified | Unclassified | Schitoviridae | Alteromonas |
| KC172839 | Mycobacterium phage BTCU-1 | Mycobacterium phage BTCU-1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45942 | 64.246 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC292029 | Halovirus HCTV-1 | Halovirus HCTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 103257 | 56.958 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292028 | Halovirus HCTV-2 | Halovirus HCTV-2 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 54291 | 68.127 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292027 | Halovirus HCTV-5 | Halovirus HCTV-5 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 102105 | 57.594 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292026 | Halovirus HGTV-1 | Halovirus HGTV-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143855 | 50.363 | Halovirus | Unclassified | Myoviridae | Unspecified |
| KC292025 | Halovirus HHTV-1 | Halovirus HHTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 49107 | 56.46 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292024 | Halovirus HHTV-2 | Halovirus HHTV-2 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 52643 | 66.592 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292023 | Halovirus HRTV-4 | Halovirus HRTV-4 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 35722 | 59.482 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC292022 | Halorubrum Tailed Virus 5 | Halorubrum Tailed Virus 5 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76134 | 56.408 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| KC292021 | Halorubrum Tailed Virus 7 | Halorubrum Tailed Virus 7 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69048 | 59.64 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| KC292020 | Halorubrum Tailed Virus 8 | Halorubrum Tailed Virus 8 Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74519 | 57.097 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| AB746912 | Nitratiruptor phage NrS-1 | Nitratiruptor phage NrS-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37159 | 40.609 | Unclassified | Unclassified | Siphoviridae | Nitratiruptor |
| AB711120 | Bacillus phage PM1 | Bacillus phage PM1 Bacillus virus PM1 Pemunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50861 | 41.285 | Pemunavirus | Unclassified | Myoviridae | Bacillus |
| JX094501 | Staphylococcus phage SA13 | Staphylococcus phage SA13 Staphylococcus virus SA13 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42652 | 35.419 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| KF005317 | Alteromonas phage vB_AmaP_AD45-P1 | Alteromonas phage vB_AmaP_AD45-P1 Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103910 | 43.154 | Unclassified | Unclassified | Schitoviridae | Alteromonas |
| KC960671 | Enterobacteria phage T3 | Enterobacteria phage T3 Escherichia virus T3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38209 | 49.918 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| JX658790 | Acinetobacter phage Abp1 | Acinetobacter phage Abp1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42185 | 39.151 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| HE983845 | Pseudomonas phage vB_PaeM_C2-10_Ab1 | Pseudomonas phage vB_PaeM_C2-10_Ab1 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 92777 | 49.277 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| HE983844 | Pseudomonas phage vB_PaeP_C1-14_Or | Pseudomonas phage vB_PaeP_C1-14_Or unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45469 | 52 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KC013023 | Leuconostoc phage LN04 | Leuconostoc phage LN04 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 25865 | 36.598 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC013021 | Leuconostoc phage P793 | Leuconostoc phage P793 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26769 | 36.628 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| KC736071 | Mycobacterium phage WIVsmall | Mycobacterium phage WIVsmall unclassified Cheoctovirus Cheoctovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53359 | 61.009 | Cheoctovirus | Unclassified | Siphoviridae | Mycobacterium |
| KC701493 | Mycobacterium phage CASbig | Mycobacterium phage CASbig unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53369 | 63.612 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC699836 | Bacillus phage SIOphi | Bacillus phage SIOphi Bacillus virus SIOphi Siophivirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 146698 | 39.019 | Siophivirus | Bastillevirinae | Herelleviridae | Bacillus |
| KC462197 | Burkholderia phage ST79 | Burkholderia phage ST79 Burkholderia virus ST79 Nampongvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35430 | 62.501 | Nampongvirus | Peduovirinae | Myoviridae | Burkholderia |
| KC832325 | Salmonella phage L13 | Salmonella phage L13 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21248 | 50.08 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KC787112 | Mycobacterium phage Clark | Mycobacterium phage Clark unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38967 | 66.205 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787111 | Mycobacterium phage Sedge | Mycobacterium phage Sedge unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39869 | 66.229 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787110 | Mycobacterium phage Leo | Mycobacterium phage Leo unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39981 | 66.862 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787109 | Mycobacterium phage Legendre | Mycobacterium phage Legendre unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40733 | 66.575 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787108 | Mycobacterium phage DNAIII | Mycobacterium phage DNAIII unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39520 | 66.827 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787106 | Mycobacterium phage Chy5 | Mycobacterium phage Chy5 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51214 | 63.6 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787105 | Mycobacterium phage Chy4 | Mycobacterium phage Chy4 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46639 | 63.678 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787104 | Mycobacterium phage Chy3 | Mycobacterium phage Chy3 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40429 | 66.559 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787103 | Mycobacterium phage Chy2 | Mycobacterium phage Chy2 unclassified Liefievirus Liefievirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40017 | 66.567 | Liefievirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787102 | Mycobacterium phage Chy1 | Mycobacterium phage Chy1 unclassified Fromanvirus Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47198 | 63.405 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| KC787101 | Mycobacterium phage 33D | Mycobacterium phage 33D unclassified Timquatrovirus Timquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50036 | 67.641 | Timquatrovirus | Unclassified | Siphoviridae | Mycobacterium |
| KC618393 | Sulfolobales Mexican fusellovirus 1 | Sulfolobales Mexican fusellovirus 1 unclassified Fuselloviridae Fuselloviridae Viruses | 14847 | 45.43 | Unclassified | Unclassified | Fuselloviridae | Unspecified |
| JQ957932 | Staphylococcus phage StauST398-2 | Staphylococcus phage StauST398-2 Staphylococcus virus StauST398-2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45572 | 33.341 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| JQ973847 | Staphylococcus phage StauST398-3 | Staphylococcus phage StauST398-3 Staphylococcus virus StauST398-3 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41392 | 35.599 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| JX296113 | Providencia phage Redjac | Providencia phage Redjac Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58104 | 49.499 | Unclassified | Unclassified | Siphoviridae | Providencia |
| AB767244 | Edwardsiella phage MSW-3 | Edwardsiella phage MSW-3 Yokohamavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42746 | 53.1 | Yokohamavirus | Unclassified | Myoviridae | Edwardsiella |
| AB757801 | Edwardsiella phage IW-1 | Edwardsiella phage IW-1 Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41684 | 48.282 | Kafunavirus | Unclassified | Podoviridae | Edwardsiella |
| AB757800 | Edwardsiella phage KF-1 | Edwardsiella phage KF-1 Kafunavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41549 | 48.276 | Kafunavirus | Unclassified | Podoviridae | Edwardsiella |
| KC522412 | Lactococcus phage CaseusJM1 | Lactococcus phage CaseusJM1 Lactococcus virus CaseusJM1 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30692 | 34.836 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JX912252 | Escherichia phage ADB-2 | Escherichia phage ADB-2 Tunavirus Tunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50552 | 45.555 | Tunavirus | Tunavirinae | Drexlerviridae | Escherichia |
| JQ957925 | Yersinia phage Y | Yersinia phage Y Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37432 | 48.301 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| GU233956 | Bacillus phage 11143 | Bacillus phage 11143 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39077 | 34.962 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| HM114277 | Rhodococcus phage E3 | Rhodococcus phage E3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142563 | 67.466 | Unclassified | Unclassified | Myoviridae | Rhodococcus |
| HQ331142 | Salmonella phage vB_SenM-S16 | Salmonella phage vB_SenM-S16 Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160221 | 36.877 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| JX174275 | Staphylococcus phage LH1 | Staphylococcus phage LH1 Staphylococcus virus LH1 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46048 | 33.191 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| KC252997 | Haloarcula hispanica virus PH1 | Haloarcula hispanica virus PH1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 28072 | 67.64 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| JX173487 | Pseudomonas phage Phi-S1 | Pseudomonas phage Phi-S1 Pseudomonas virus PhiS1 Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40192 | 56.213 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| DQ832317 | Escherichia phage V5 | Escherichia phage V5 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137947 | 43.561 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| JF974308 | Rhodobacter phage RC1 | Rhodobacter phage RC1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39573 | 62.272 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| JF974307 | Rhodovulum phage RS1 | Rhodovulum phage RS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40231 | 62.119 | Unclassified | Unclassified | Siphoviridae | Rhodovulum |
| JF974293 | Synechococcus phage KBS-M-1A | Synechococcus phage KBS-M-1A unclassified Neptunevirus Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171744 | 40.637 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| HQ634188 | Cyanophage KBS-P-1A | Cyanophage KBS-P-1A Sednavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45730 | 47.354 | Sednavirus | Unclassified | Autographiviridae | Synechococcus |
| JF974296 | Pseudoalteromonas phage pYD6-A | Pseudoalteromonas phage pYD6-A Pseudoalteromonas virus pYD6A Matsuvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76802 | 38.658 | Matsuvirus | Fuhrmanvirinae | Schitoviridae | Pseudoalteromonas |
| JF974295 | Psychrobacter phage pOW20-A | Psychrobacter phage pOW20-A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40803 | 44.056 | Unclassified | Unclassified | Myoviridae | Psychrobacter |
| JF974292 | Cyanophage S-SSM2 | Cyanophage S-SSM2 unclassified Ahtivirus Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179980 | 41.096 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| HQ634196 | Vibrio phage VBP32 | Vibrio phage VBP32 unclassified Stoningtonvirus Stoningtonvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76718 | 42.483 | Stoningtonvirus | Fuhrmanvirinae | Schitoviridae | Vibrio |
| HQ634194 | Vibrio phage VBP47 | Vibrio phage VBP47 Vibrio virus VBP47 Stoningtonvirus Fuhrmanvirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76705 | 42.456 | Stoningtonvirus | Fuhrmanvirinae | Schitoviridae | Vibrio |
| HQ634193 | Cyanophage P-RSM6 | Cyanophage P-RSM6 unclassified Libanvirus Libanvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 192497 | 39.316 | Libanvirus | Unclassified | Myoviridae | Prochlorococcus |
| HQ634192 | Cellulophaga phage phiST | Cellulophaga phage phiST Cbastvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 79114 | 30.201 | Cbastvirus | Unclassified | Siphoviridae | Cellulophaga |
| HQ634191 | Cyanophage Syn10 | Cyanophage Syn10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177103 | 40.607 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| HQ634190 | Cyanophage Syn2 | Cyanophage Syn2 unclassified Pontusvirus Pontusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175596 | 41.271 | Pontusvirus | Unclassified | Myoviridae | Synechococcus |
| HQ634189 | Synechococcus phage Syn30 | Synechococcus phage Syn30 Synechococcus virus Syn30 Leucotheavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178807 | 39.948 | Leucotheavirus | Unclassified | Myoviridae | Synechococcus |
| HQ634157 | Vibrio phage 11895-B1 | Vibrio phage 11895-B1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 126434 | 35.529 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ634156 | Vibrio phage PWH3a-P1 | Vibrio phage PWH3a-P1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129155 | 35.504 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ634153 | Salicola phage CGphi29 | Salicola phage CGphi29 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40695 | 56.422 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| HQ633071 | Synechococcus phage S-SKS1 | Synechococcus phage S-SKS1 Synechococcus virus SSKS1 Llyrvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 208007 | 36.027 | Llyrvirus | Unclassified | Myoviridae | Synechococcus |
| HQ632859 | Loktanella phage pCB2051-A | Loktanella phage pCB2051-A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56958 | 54.976 | Unclassified | Unclassified | Siphoviridae | Loktanella |
| HQ632858 | Tetraselmis viridis virus SI1 | Tetraselmis viridis virus SI1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39398 | 61.224 | Unclassified | Unclassified | Siphoviridae | Tetraselmis |
| HQ632857 | Tetraselmis viridis virus S20 | Tetraselmis viridis virus S20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38987 | 61.22 | Unclassified | Unclassified | Siphoviridae | Tetraselmis |
| JF974286 | Prochlorococcus phage MED4-184 | Prochlorococcus phage MED4-184 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38327 | 37.157 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| HQ634177 | Synechococcus phage S-CAM1 | Synechococcus phage S-CAM1 Synechococcus virus SCAM1 Anaposvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 198013 | 43.025 | Anaposvirus | Unclassified | Myoviridae | Synechococcus |
| HQ634176 | Cyanophage P-RSM3 | Cyanophage P-RSM3 unclassified Ronodorvirus Ronodorvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178750 | 36.747 | Ronodorvirus | Unclassified | Myoviridae | Prochlorococcus |
| HQ634175 | Cyanophage P-RSM1 | Cyanophage P-RSM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177211 | 40.194 | Unclassified | Unclassified | Myoviridae | Prochlorococcus |
| HQ634174 | Prochlorococcus phage MED4-213 | Prochlorococcus phage MED4-213 Prochlorococcus virus MED4-213 Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 180977 | 37.764 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| JX863101 | Pseudomonas phage UFV-P2 | Pseudomonas phage UFV-P2 Pseudomonas virus UFVP2 Vicosavirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45517 | 51.471 | Vicosavirus | Unclassified | Podoviridae | Pseudomonas |
| JQ797329 | Listeria phage vB_LmoM_AG20 | Listeria phage vB_LmoM_AG20 Listeria virus AG20 Pecentumvirus Jasinkavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 133057 | 35.933 | Pecentumvirus | Jasinkavirinae | Herelleviridae | Listeria |
| JF974321 | Cyanophage MED4-117 | Cyanophage MED4-117 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38834 | 37.246 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| JF974287 | Vibrio phage pYD21-A | Vibrio phage pYD21-A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46917 | 43.897 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| HQ332136 | Dunaliella viridis virus SI2 | Dunaliella viridis virus SI2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37328 | 62.704 | Unclassified | Unclassified | Podoviridae | Dunaliella |
| HQ317392 | Cellulophaga phage phiSM | Cellulophaga phage phiSM Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44557 | 33.927 | Unclassified | Unclassified | Myoviridae | Cellulophaga |
| HQ317391 | Cyanophage S-SSM6a | Cyanophage S-SSM6a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 232883 | 39.127 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| HQ317386 | Vibrio phage VBM1 | Vibrio phage VBM1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38374 | 42.258 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ317389 | Synechococcus phage S-RIP2 | Synechococcus phage S-RIP2 Synechococcus virus SRIP2 Sednavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45728 | 47.343 | Sednavirus | Unclassified | Autographiviridae | Synechococcus |
| HQ317388 | Synechococcus phage S-RIP1 | Synechococcus phage S-RIP1 Synechococcus virus SRIP1 Kajamvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44892 | 42.93 | Kajamvirus | Unclassified | Autographiviridae | Synechococcus |
| JN116827 | Rhodococcus phage RER2 | Rhodococcus phage RER2 Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46586 | 58.571 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| JN116826 | Rhodococcus phage RGL3 | Rhodococcus phage RGL3 Rerduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48072 | 62.66 | Rerduovirus | Unclassified | Siphoviridae | Rhodococcus |
| JX094431 | Bacillus phage vB_BceM_Bc431v3 | Bacillus phage vB_BceM_Bc431v3 Caeruleovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158621 | 39.978 | Caeruleovirus | Bastillevirinae | Herelleviridae | Bacillus |
| JX846613 | Staphylococcus phage vB_SauM_Romulus | Staphylococcus phage vB_SauM_Romulus Silviavirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 131332 | 30.011 | Silviavirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX944686 | Sulfolobales Mexican rudivirus 1 | Sulfolobales Mexican rudivirus 1 unclassified Rudivirus Rudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 27431 | 46.597 | Rudivirus | Unclassified | Rudiviridae | Unspecified |
| JX879087 | Streptococcus phage phiNJ2 | Streptococcus phage phiNJ2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37282 | 39.418 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JX560968 | Escherichia phage EC6 | Escherichia phage EC6 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86231 | 38.898 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| HQ316582 | Vibrio phage henriette 12B8 | Vibrio phage henriette 12B8 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107218 | 40.98 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ316581 | Vibrio phage martha 12B12 | Vibrio phage martha 12B12 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33277 | 45.755 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ316580 | Vibrio phage douglas 12A4 | Vibrio phage douglas 12A4 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57611 | 38.956 | Unclassified | Unclassified | Podoviridae | Vibrio |
| HQ316579 | Vibrio phage helene 12B3 | Vibrio phage helene 12B3 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 135982 | 37.584 | Unclassified | Unclassified | Myoviridae | Vibrio |
| KC107834 | Cronobacter phage vB_CskP_GAP227 | Cronobacter phage vB_CskP_GAP227 Cronobacter virus GAP227 Cronosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41796 | 55.74 | Cronosvirus | Melnykvirinae | Autographiviridae | Cronobacter |
| HQ337022 | Prochlorococcus phage P-SSP10 | Prochlorococcus phage P-SSP10 Prochlorococcus virus PSSP10 Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47325 | 39.231 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| HQ332140 | Prochlorococcus phage P-GSP1 | Prochlorococcus phage P-GSP1 Prochlorococcus virus PGSP1 Lingvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44945 | 39.551 | Lingvirus | Unclassified | Autographiviridae | Prochlorococcus |
| HQ332137 | Prochlorococcus phage P-SSP3 | Prochlorococcus phage P-SSP3 Prochlorococcus virus PSSP3 Tritonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46198 | 37.941 | Tritonvirus | Unclassified | Autographiviridae | Prochlorococcus |
| HQ316584 | Cyanophage SS120-1 | Cyanophage SS120-1 Prochlorococcus virus SS120-1 Banchanvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46997 | 40.492 | Banchanvirus | Unclassified | Autographiviridae | Prochlorococcus |
| HQ316603 | Cyanophage S-SSM6b | Cyanophage S-SSM6b Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182368 | 39.403 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| HQ316583 | Synechococcus phage S-SSM4 | Synechococcus phage S-SSM4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182801 | 39.393 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| JX202565 | Salmonella phage wksl3 | Salmonella phage wksl3 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42633 | 49.797 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| KC117377 | Halovirus HVTV-1 | Halovirus HVTV-1 Haloviruses unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 102319 | 58.271 | Halovirus | Unclassified | Unclassified | Unspecified |
| KC117376 | Halovirus HSTV-2 | Halovirus HSTV-2 unclassified Haloferacalesvirus Haloferacalesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 68527 | 60.338 | Haloferacalesvirus | Unclassified | Myoviridae | Unspecified |
| JQ340389 | Vibrio phage pVp-1 | Vibrio phage pVp-1 Vibrio virus pVp1 Vipunavirus Ermolyevavirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 111506 | 39.707 | Vipunavirus | Ermolyevavirinae | Demerecviridae | Vibrio |
| JX509734 | Enterobacteria phage SfI | Enterobacteria phage SfI Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38389 | 50.116 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| JQ691611 | Cronobacter phage CR9 | Cronobacter phage CR9 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151924 | 50.556 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| JQ691610 | Salmonella phage SPN9TCW | Salmonella phage SPN9TCW unclassified Uetakevirus Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38689 | 50.151 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| HM559717 | Synechococcus phage S-CBP4 | Synechococcus phage S-CBP4 Synechococcus virus SCBP4 Poseidonvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41824 | 44.293 | Poseidonvirus | Unclassified | Autographiviridae | Synechococcus |
| HQ634152 | Prochlorococcus phage P-SSP6 | Prochlorococcus phage P-SSP6 Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47039 | 39.187 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| HQ332139 | Prochlorococcus phage P-RSP2 | Prochlorococcus phage P-RSP2 unclassified Autographiviridae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42257 | 34.049 | Unclassified | Unclassified | Autographiviridae | Prochlorococcus |
| AP012341 | Staphylococcus phage phi7401PVL | Staphylococcus phage phi7401PVL unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47252 | 33.044 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| KC294142 | Pseudomonas phage PaP4 | Pseudomonas phage PaP4 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43895 | 52.464 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| KC237308 | Enterobacteria phage ID204 Moscow/ID | Enterobacteria phage ID204 Moscow/ID Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5475 | 45.388 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| JF974289 | Synechococcus phage S-RIM8 A.HR3 | Synechococcus phage S-RIM8 A.HR3 unclassified Neptunevirus Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171211 | 40.631 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| JF974288 | Synechococcus phage S-RIM8 A.HR1 | Synechococcus phage S-RIM8 A.HR1 unclassified Neptunevirus Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171211 | 40.631 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| HQ317385 | Synechococcus phage S-RIM8 A.HR5 | Synechococcus phage S-RIM8 A.HR5 unclassified Neptunevirus Neptunevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168327 | 40.606 | Neptunevirus | Unclassified | Myoviridae | Synechococcus |
| JF966203 | Bacillus phage Bastille | Bacillus phage Bastille Bastillevirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 153962 | 38.144 | Bastillevirus | Bastillevirinae | Herelleviridae | Bacillus |
| HM144387 | Bacillus phage W.Ph. | Bacillus phage W.Ph. Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156897 | 36.447 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| AF399011 | Pseudomonas phage phiKZ | Pseudomonas phage phiKZ Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 280334 | 36.833 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| JX434033 | Pseudomonas phage JBD88a | Pseudomonas phage JBD88a unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36429 | 63.985 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JX434032 | Pseudomonas phage JBD30 | Pseudomonas phage JBD30 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36947 | 64.276 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JX434031 | Pseudomonas phage JBD24 | Pseudomonas phage JBD24 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37095 | 64.24 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JX434030 | Pseudomonas phage JBD5 | Pseudomonas phage JBD5 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37740 | 64.269 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| HE861935 | Streptococcus phage TP-J34 | Streptococcus phage TP-J34 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45606 | 38.819 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JX681814 | Burkholderia phage phiX216 | Burkholderia phage phiX216 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37637 | 64.822 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| JF461087 | Salmonella phage FO1a | Salmonella phage FO1a Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 83331 | 38.933 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX000007 | Yersinia phage R | Yersinia phage R Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37260 | 48.36 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| AF338467 | Synechococcus phage P60 | Synechococcus phage P60 Tiilvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46675 | 53.285 | Tiilvirus | Unclassified | Autographiviridae | Synechococcus |
| HQ163896 | Staphylococcus phage Sb1 | Staphylococcus phage Sb1 Staphylococcus virus Sb1 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127188 | 30.475 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX866719 | Klebsiella phage JD001 | Klebsiella phage JD001 Klebsiella virus JD001 Jedunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48814 | 48.535 | Jedunavirus | Unclassified | Myoviridae | Klebsiella |
| JX875065 | Staphylococcus phage SA5 | Staphylococcus phage SA5 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 137031 | 30.421 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX676771 | Pseudomonas phage AF | Pseudomonas phage AF Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42689 | 58.439 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| JX878496 | Serratia phage phiMAM1 | Serratia phage phiMAM1 Miltonvirus Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157834 | 51.984 | Miltonvirus | Unclassified | Ackermannviridae | Serratia |
| JQ182736 | Enterobacteria phage vB_EcoS_Rogue1 | Enterobacteria phage vB_EcoS_Rogue1 Rogunavirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45805 | 44.214 | Rogunavirus | Rogunavirinae | Drexlerviridae | Enterobacteria |
| JQ182735 | Escherichia phage HK629 | Escherichia phage HK629 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47288 | 49.638 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| JQ182734 | Enterobacteria phage mEp043 c-1 | Enterobacteria phage mEp043 c-1 Enterobacteria virus mEp043 Aguilavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42780 | 50.79 | Aguilavirus | Unclassified | Siphoviridae | Enterobacteria |
| JQ182733 | Enterobacteria phage mEp213 | Enterobacteria phage mEp213 Enterobacteria virus mEp213 Aguilavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44120 | 50.977 | Aguilavirus | Unclassified | Siphoviridae | Enterobacteria |
| JQ182732 | Escherichia phage mEp234 | Escherichia phage mEp234 Wanchaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39578 | 49.586 | Wanchaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ182731 | Enterobacteria phage mEp235 | Enterobacteria phage mEp235 Enterobacteria virus mEp235 Nochtlivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37595 | 50.015 | Nochtlivirus | Hendrixvirinae | Siphoviridae | Enterobacteria |
| JQ182730 | Enterobacteria phage mEp237 | Enterobacteria phage mEp237 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44375 | 51.432 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| JQ182729 | Enterobacterial phage mEp390 | Enterobacterial phage mEp390 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40029 | 51.678 | Unclassified | Hendrixvirinae | Siphoviridae | Enterobacteria |
| JQ182728 | Enterobacteria phage mEp460 | Enterobacteria phage mEp460 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44510 | 50.874 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| JQ182727 | Escherichia phage mEpX1 | Escherichia phage mEpX1 Cuauhtlivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41567 | 49.311 | Cuauhtlivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ182726 | Escherichia phage mEpX2 | Escherichia phage mEpX2 Shamshuipovirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38759 | 50.076 | Shamshuipovirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ086373 | Escherichia phage HK542 | Escherichia phage HK542 Wongtaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38964 | 50.914 | Wongtaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ086371 | Enterobacteria phage HK225 | Enterobacteria phage HK225 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45366 | 51.955 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| JQ086370 | Enterobacteria phage HK140 | Enterobacteria phage HK140 Enterobacteria virus HK140 Yautsimvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40710 | 49.912 | Yautsimvirus | Hendrixvirinae | Siphoviridae | Enterobacteria |
| JQ086376 | Escherichia phage HK630 | Escherichia phage HK630 Lambdavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47090 | 50.053 | Lambdavirus | Unclassified | Siphoviridae | Escherichia |
| JQ086369 | Escherichia phage HK106 | Escherichia phage HK106 Wanchaivirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41468 | 49.342 | Wanchaivirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ086372 | Escherichia phage HK446 | Escherichia phage HK446 Kwaitsingvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39026 | 50.102 | Kwaitsingvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ086374 | Escherichia phage HK544 | Escherichia phage HK544 Kwaitsingvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40155 | 49.797 | Kwaitsingvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ086375 | Escherichia phage HK578 | Escherichia phage HK578 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43741 | 54.484 | Dhillonvirus | Unclassified | Siphoviridae | Escherichia |
| JQ086377 | Escherichia phage HK633 | Escherichia phage HK633 Saikungvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41528 | 49.653 | Saikungvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JQ031132 | Escherichia phage FV3 | Escherichia phage FV3 Vequintavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 136947 | 43.699 | Vequintavirus | Vequintavirinae | Myoviridae | Escherichia |
| JX556418 | Vibrio phage vB_VpaS_MAR10 | Vibrio phage vB_VpaS_MAR10 Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78751 | 49.703 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| JX872508 | Stenotrophomonas phage IME15 | Stenotrophomonas phage IME15 Stenotrophomonas virus IME15 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38513 | 53.652 | Teseptimavirus | Studiervirinae | Autographiviridae | Stenotrophomonas |
| JX889246 | Streptomyces phage phiCAM | Streptomyces phage phiCAM Camvirus Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50348 | 65.586 | Camvirus | Arquatrovirinae | Siphoviridae | Streptomyces |
| JX878671 | Staphylococcus phage JD007 | Staphylococcus phage JD007 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 141836 | 30.371 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JX880034 | Escherichia phage IME11 | Escherichia phage IME11 Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72570 | 43.139 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| JX867715 | Escherichia phage NJ01 | Escherichia phage NJ01 Kuravirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 77448 | 42.049 | Kuravirus | Unclassified | Podoviridae | Escherichia |
| JX536493 | Escherichia phage HX01 | Escherichia phage HX01 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169158 | 37.588 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| JX403939 | Pseudomonas phage YMC/01/01/P52_PAE_BP | Pseudomonas phage YMC/01/01/P52_PAE_BP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49381 | 62.162 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| JN882287 | Cronobacter phage vB_CsaM_GAP161 | Cronobacter phage vB_CsaM_GAP161 Cronobacter virus GAP161 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178193 | 44.523 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Cronobacter |
| JN882284 | Cronobacter phage vB_CsaM_GAP31 | Cronobacter phage vB_CsaM_GAP31 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147940 | 46.298 | Seunavirus | Vequintavirinae | Myoviridae | Cronobacter |
| JX486088 | Lactobacillus phage ATCC 8014-B2 | Lactobacillus phage ATCC 8014-B2 Lactobacillus virus B2 Douglaswolinvirus Tybeckvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 80617 | 36.962 | Douglaswolinvirus | Tybeckvirinae | Siphoviridae | Lactobacillus |
| HE981739 | Escherichia virus RB43 | Escherichia virus RB43 Pseudotevenvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179839 | 43.161 | Pseudotevenvirus | Tevenvirinae | Myoviridae | Escherichia |
| JX556417 | Vibrio phage vB_VpaM_MAR | Vibrio phage vB_VpaM_MAR Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41351 | 51.336 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| JX561091 | Escherichia phage phAPEC8 | Escherichia phage phAPEC8 Escherichia virus phAPEC8 Phapecoctavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147737 | 39.148 | Phapecoctavirus | Unclassified | Myoviridae | Escherichia |
| HQ225832 | Tetrasphaera phage TJE1 | Tetrasphaera phage TJE1 Tetrasphaera virus TJE1 Tijeunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49219 | 57.742 | Tijeunavirus | Unclassified | Myoviridae | Tetrasphaera |
| JX560521 | Acinetobacter phage phiAC-1 | Acinetobacter phage phiAC-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43216 | 38.476 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| JX442241 | Listeria phage P70 | Listeria phage P70 Homburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 67170 | 36.457 | Homburgvirus | Unclassified | Siphoviridae | Listeria |
| JN986846 | Escherichia phage vB_EcoM_ACG-C40 | Escherichia phage vB_EcoM_ACG-C40 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167396 | 35.237 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JN986845 | Escherichia phage vB_EcoS_ACG-M12 | Escherichia phage vB_EcoS_ACG-M12 Guelphvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46054 | 43.512 | Guelphvirus | Braunvirinae | Drexlerviridae | Escherichia |
| JN986844 | Escherichia phage vB_EcoP_ACG-C91 | Escherichia phage vB_EcoP_ACG-C91 Escherichia virus ACGC91 Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43731 | 45.281 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| JX145342 | Clostridium phage phiMMP04 | Clostridium phage phiMMP04 Clostridioides virus phiMMP04 Sherbrookevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31674 | 29.971 | Sherbrookevirus | Unclassified | Myoviridae | Clostridium |
| JX145341 | Clostridium phage phiMMP02 | Clostridium phage phiMMP02 Clostridioides virus phiMMP02 Colneyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48396 | 29.552 | Colneyvirus | Unclassified | Myoviridae | Clostridium |
| JN871591 | Salmonella virus SPN19 | Salmonella virus SPN19 Chivirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59203 | 56.524 | Chivirus | Unclassified | Siphoviridae | Salmonella |
| AB560486 | Pseudomonas phage KPP12 | Pseudomonas phage KPP12 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64144 | 55.623 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| JN882286 | Cronobacter phage vB_CsaP_GAP52 | Cronobacter phage vB_CsaP_GAP52 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76631 | 44.221 | Unclassified | Unclassified | Podoviridae | Cronobacter |
| JX495043 | Pseudomonas phage JBD67 | Pseudomonas phage JBD67 Pseudomonas virus JBD67 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38232 | 63.018 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| JX495042 | Pseudomonas phage JBD25 | Pseudomonas phage JBD25 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39552 | 62.535 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| JX495041 | Pseudomonas phage JD18 | Pseudomonas phage JD18 Pseudomonas virus JD18 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39014 | 63.388 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| JX181829 | Salmonella phage SKML-39 | Salmonella phage SKML-39 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 159624 | 50.154 | Agtrevirus | Aglimvirinae | Ackermannviridae | Salmonella |
| JX181828 | Salmonella phage STML-13-1 | Salmonella phage STML-13-1 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157235 | 44.782 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| JX181825 | Salmonella phage STML-198 | Salmonella phage STML-198 Gelderlandvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158099 | 36.888 | Gelderlandvirus | Tevenvirinae | Myoviridae | Salmonella |
| JX181824 | Salmonella phage SSE121 | Salmonella phage SSE121 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147745 | 45.292 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| JX181823 | Salmonella phage SPT-1 | Salmonella phage SPT-1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22518 | 38.369 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX181822 | Salmonella phage SPT-1 | Salmonella phage SPT-1 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64108 | 39.224 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX181817 | Salmonella phage SBA-1781 | Salmonella phage SBA-1781 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14850 | 39.394 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX181816 | Salmonella phage SBA-1781 | Salmonella phage SBA-1781 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18103 | 39.231 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX181815 | Salmonella phage SBA-1781 | Salmonella phage SBA-1781 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21805 | 39.211 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX181814 | Salmonella phage SBA-1781 | Salmonella phage SBA-1781 unclassified Felixounavirus Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22444 | 38.42 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| JX182369 | Streptomyces phage phiHau3 | Streptomyces phage phiHau3 Arquatrovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50255 | 67.834 | Unclassified | Arquatrovirinae | Siphoviridae | Streptomyces |
| JN700520 | Staphylococcus phage StB12 | Staphylococcus phage StB12 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44714 | 34.171 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| JN700521 | Staphylococcus phage StB20 | Staphylococcus phage StB20 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40917 | 34.067 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| JN700519 | Staphylococcus phage StB27 | Staphylococcus phage StB27 Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40071 | 33.216 | Unclassified | Azeredovirinae | Siphoviridae | Staphylococcus |
| JF270478 | Psychrobacter phage Psymv2 | Psychrobacter phage Psymv2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35725 | 44.467 | Unclassified | Unclassified | Siphoviridae | Psychrobacter |
| JX306041 | Stenotrophomonas phage IME13 | Stenotrophomonas phage IME13 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162327 | 41.179 | Tulanevirus | Unclassified | Myoviridae | Stenotrophomonas |
| JQ341413 | Weissella phage phiYS61 | Weissella phage phiYS61 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33594 | 43.919 | Unclassified | Unclassified | Podoviridae | Weissella |
| JN712910 | Bacillus phage BCD7 | Bacillus phage BCD7 Bacillus virus BCD7 Becedseptimavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93839 | 38.043 | Becedseptimavirus | Unclassified | Myoviridae | Bacillus |
| JN654439 | Bacillus phage BPS13 | Bacillus phage BPS13 Wphvirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158305 | 38.751 | Wphvirus | Bastillevirinae | Herelleviridae | Bacillus |
| JN797798 | Bacillus phage BCU4 | Bacillus phage BCU4 Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 154371 | 39.857 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| JN797796 | Bacillus phage B5S | Bacillus phage B5S Bequatrovirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 162598 | 37.71 | Bequatrovirus | Bastillevirinae | Herelleviridae | Bacillus |
| JX195166 | Pectobacterium phage My1 | Pectobacterium phage My1 Pectobacterium virus My1 Myunavirus Mccorquodalevirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 122024 | 40.614 | Myunavirus | Mccorquodalevirinae | Demerecviridae | Pectobacterium |
| JN984867 | Shigella phage EP23 | Shigella phage EP23 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44077 | 54.421 | Dhillonvirus | Unclassified | Siphoviridae | Shigella |
| AB712291 | Enterococcus phage BC611 | Enterococcus phage BC611 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 53996 | 40.449 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| HM368260 | Acinetobacter phage AB1 | Acinetobacter phage AB1 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45159 | 37.689 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| JQ966307 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3525 | 50.979 | Levivirus | Unclassified | Leviviridae | Escherichia |
| HE664024 | Escherichia phage P13374 | Escherichia phage P13374 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60894 | 50.23 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| JQ965645 | Salmonella phage SSU5 | Salmonella phage SSU5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 103299 | 51.105 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| JX123262 | Aeromonas phage CC2 | Aeromonas phage CC2 Aeromonas virus CC2 Ceceduovirus Emmerichvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231743 | 38.821 | Ceceduovirus | Emmerichvirinae | Myoviridae | Aeromonas |
| JN377894 | Aeromonas phage Aes508 | Aeromonas phage Aes508 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 160646 | 41.227 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| JX409894 | Streptococcus phage LYGO9 | Streptococcus phage LYGO9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41660 | 36.726 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JQ312117 | Agrobacterium phage 7-7-1 | Agrobacterium phage 7-7-1 Agrobacterium virus 7-7-1 Schmittlotzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69391 | 52.377 | Schmittlotzvirus | Unclassified | Myoviridae | Agrobacterium |
| JX163858 | Caulobacter virus phiCbK | Caulobacter virus phiCbK Shapirovirus Dolichocephalovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 205504 | 66.105 | Shapirovirus | Dolichocephalovirinae | Siphoviridae | Caulobacter |
| JX194238 | Pseudomonas phage PA26 | Pseudomonas phage PA26 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72321 | 54.818 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| JF946695 | Mycobacterium phage SWU1 | Mycobacterium phage SWU1 Fromanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52474 | 62.406 | Fromanvirus | Unclassified | Siphoviridae | Mycobacterium |
| AY730274 | Salmonella phage SS3e | Salmonella phage SS3e Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40793 | 50.08 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| HM997020 | Escherichia phage wV7 | Escherichia phage wV7 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166452 | 35.399 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JQ801337 | Vibrio phage VvAW1 | Vibrio phage VvAW1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38682 | 49.113 | Unclassified | Unclassified | Podoviridae | Vibrio |
| AP011113 | Escherichia phage AR1 | Escherichia phage AR1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167435 | 35.331 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| HQ110084 | Erwinia phage ENT90 | Erwinia phage ENT90 Erwinia virus ENT90 Entnonagintavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29564 | 55.811 | Entnonagintavirus | Peduovirinae | Myoviridae | Erwinia |
| JN192401 | Staphylococcus virus IPLA7 | Staphylococcus virus IPLA7 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42123 | 34.755 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| JN192400 | Staphylococcus virus IPLA5 | Staphylococcus virus IPLA5 Rockefellervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43581 | 34.726 | Rockefellervirus | Unclassified | Siphoviridae | Staphylococcus |
| JF832915 | Haemophilus phage SuMu | Haemophilus phage SuMu Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37151 | 41.872 | Unclassified | Unclassified | Myoviridae | Haemophilus |
| JQ340088 | Halocynthia phage JM-2012 | Halocynthia phage JM-2012 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167292 | 35.395 | Unclassified | Unclassified | Myoviridae | Unspecified |
| JX131330 | Pseudomonas phage MP1412 | Pseudomonas phage MP1412 Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61167 | 64.283 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| JX006077 | Saccharomonospora phage PIS 136 | Saccharomonospora phage PIS 136 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94870 | 65.881 | Unclassified | Unclassified | Siphoviridae | Saccharomonospora |
| AP009390 | Enterococcus phage EF24C | Enterococcus phage EF24C Enterococcus virus EF24C Kochikohdavirus Brockvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 142072 | 35.738 | Kochikohdavirus | Brockvirinae | Herelleviridae | Enterococcus |
| JQ837901 | Pectobacterium phage PP1 | Pectobacterium phage PP1 Pectobacterium virus PP1 Axomammavirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44400 | 49.736 | Axomammavirus | Molineuxvirinae | Autographiviridae | Pectobacterium |
| JQ686190 | Staphylococcus phage G15 | Staphylococcus phage G15 Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 139806 | 30.228 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JF734911 | Helicobacter phage phiHP33 | Helicobacter phage phiHP33 unclassified Schmidvirus Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24645 | 37.338 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| HE815464 | Campylobacter phage CP21 | Campylobacter phage CP21 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182833 | 27.151 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| JQ867100 | Marinomonas phage P12026 | Marinomonas phage P12026 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31766 | 46.603 | Unclassified | Unclassified | Siphoviridae | Marinomonas |
| JQ867099 | Croceibacter phage P2559S | Croceibacter phage P2559S Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44988 | 41.853 | Unclassified | Unclassified | Siphoviridae | Croceibacter |
| JQ823123 | Nonlabens phage P12024L | Nonlabens phage P12024L Inhavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35652 | 35.875 | Inhavirus | Unclassified | Siphoviridae | Nonlabens |
| JQ823122 | Nonlabens phage P12024S | Nonlabens phage P12024S Inhavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35700 | 35.535 | Inhavirus | Unclassified | Siphoviridae | Nonlabens |
| JX233784 | Pseudomonas phage PA7 | Pseudomonas phage PA7 Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 266743 | 36.933 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| JQ617284 | Helicobacter phage 1961P | Helicobacter phage 1961P Schmidvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26836 | 35.516 | Schmidvirus | Unclassified | Podoviridae | Helicobacter |
| JN377895 | Aeromonas phage Aes012 | Aeromonas phage Aes012 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161978 | 41.3 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| JQ659259 | Leuconostoc phage Lmd1 | Leuconostoc phage Lmd1 Limdunavirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26201 | 36.403 | Limdunavirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| HE956711 | Yersinia phage phiD1 | Yersinia phage phiD1 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167063 | 35.442 | Tequatrovirus | Tevenvirinae | Myoviridae | Yersinia |
| HE956709 | Yersinia phage phiR1-RT | Yersinia phage phiR1-RT Tegunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168809 | 34.544 | Tegunavirus | Unclassified | Myoviridae | Yersinia |
| JQ764988 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JX193904 | Enterococcus phage EfaCPT1 | Enterococcus phage EfaCPT1 Enterococcus virus EfaCPT1 Efquatrovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40923 | 34.641 | Efquatrovirus | Unclassified | Siphoviridae | Enterococcus |
| JX128259 | Escherichia phage ECML-134 | Escherichia phage ECML-134 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166783 | 35.411 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| JX128258 | Escherichia phage ECML-117 | Escherichia phage ECML-117 Escherichia virus ECML-117 Wifcevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66854 | 46.198 | Wifcevirus | Unclassified | Myoviridae | Escherichia |
| JX128257 | Escherichia phage ECML-4 | Escherichia phage ECML-4 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157308 | 45.032 | Kuttervirus | Cvivirinae | Ackermannviridae | Escherichia |
| JQ762257 | Pseudomonas phage MP42 | Pseudomonas phage MP42 unclassified Casadabanvirus Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36847 | 64.198 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JQ177064 | Vibrio phage vB_VchM-138 | Vibrio phage vB_VchM-138 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44485 | 45.555 | Unclassified | Unclassified | Myoviridae | Vibrio |
| JQ177063 | Aeromonas phage vB_AsaM-56 | Aeromonas phage vB_AsaM-56 Aeromonas virus 56 Popoffvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43551 | 55.425 | Popoffvirus | Unclassified | Myoviridae | Aeromonas |
| JQ177062 | Bdellovibrio phage phi1422 | Bdellovibrio phage phi1422 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45354 | 44.294 | Unclassified | Unclassified | Myoviridae | Bdellovibrio |
| JQ177065 | Pectobacterium phage ZF40 | Pectobacterium phage ZF40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48454 | 50.204 | Unclassified | Unclassified | Myoviridae | Pectobacterium |
| JQ177061 | Vibrio phage CP-T1 | Vibrio phage CP-T1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 45.415 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HE614282 | Bacillus phage BceA1 | Bacillus phage BceA1 Bacillus virus BceA1 Beceayunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42932 | 35.663 | Beceayunavirus | Unclassified | Siphoviridae | Bacillus |
| HE614281 | Staphylococcus phage SpaA1 | Staphylococcus phage SpaA1 unclassified Beceayunavirus Beceayunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42784 | 35.635 | Beceayunavirus | Unclassified | Siphoviridae | Staphylococcus |
| HQ316604 | Vibrio phage SIO-2 | Vibrio phage SIO-2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81184 | 44.978 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| HQ396194 | Riemerella phage RAP44 | Riemerella phage RAP44 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49329 | 34.74 | Unclassified | Unclassified | Siphoviridae | Riemerella |
| GU903191 | Escherichia phage vB_EcoM_ECO1230-10 | Escherichia phage vB_EcoM_ECO1230-10 Jilinvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41666 | 53.374 | Jilinvirus | Unclassified | Myoviridae | Escherichia |
| JQ764990 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.671 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JQ764989 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.671 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JQ764987 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JQ764986 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GQ451696 | Leuconostoc phage 1-A4 | Leuconostoc phage 1-A4 Unaquatrovirus Mccleskeyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29508 | 36.146 | Unaquatrovirus | Mccleskeyvirinae | Siphoviridae | Leuconostoc |
| GU071093 | Prochlorococcus virus PSSP7 | Prochlorococcus virus PSSP7 Tiamatvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45135 | 38.564 | Tiamatvirus | Unclassified | Autographiviridae | Prochlorococcus |
| GU071091 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39778 | 48.384 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| GU071090 | Cyanophage PSS2 | Cyanophage PSS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 105532 | 52.305 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| GQ915271 | Staphylococcus phage SAP090B | Staphylococcus phage SAP090B unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26877 | 33.709 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| GQ900512 | Staphylococcus phage SAP090D | Staphylococcus phage SAP090D Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10726 | 33.787 | Unclassified | Unclassified | Unclassified | Staphylococcus |
| GQ452243 | Enterococcus phage phiEf11 | Enterococcus phage phiEf11 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42822 | 34.389 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| JQ809650 | Celeribacter phage P12053L | Celeribacter phage P12053L Celeribacter virus P12053L Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38889 | 46.1 | Siovirus | Cobavirinae | Zobellviridae | Celeribacter |
| JQ446452 | Salinivibrio phage CW02 | Salinivibrio phage CW02 Salinivibrio virus CW02 Salinovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49390 | 47.672 | Salinovirus | Unclassified | Zobellviridae | Salinivibrio |
| JN991020 | Pseudomonas phage Bf7 | Pseudomonas phage Bf7 Pseudomonas virus Bf7 Bifseptvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40058 | 58.405 | Bifseptvirus | Unclassified | Autographiviridae | Pseudomonas |
| GU071104 | Cyanophage NATL2A-133 | Cyanophage NATL2A-133 Prochlorococcus virus NATL2A133 Tangaroavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47536 | 39.871 | Tangaroavirus | Unclassified | Autographiviridae | Prochlorococcus |
| EU876853 | Iodobacter phage PhiPLPE | Iodobacter phage PhiPLPE Iodobacter virus PLPE Iodovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47453 | 46.105 | Iodovirus | Unclassified | Myoviridae | Iodobacter |
| EU734174 | Escherichia phage 13a | Escherichia phage 13a Escherichia virus 13a Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38841 | 48.39 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| EU734173 | Klebsiella phage K11 | Klebsiella phage K11 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41181 | 53.248 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| EU734172 | Escherichia phage EcoDS1 | Escherichia phage EcoDS1 Escherichia virus EcoDS1 Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39252 | 49.944 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| EU734171 | Escherichia phage BA14 | Escherichia phage BA14 Escherichia virus BA14 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39816 | 48.777 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| EU734170 | Yersinia phage Yepe2 | Yersinia phage Yepe2 Yersinia virus Yepe2 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38677 | 47.266 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| JN564907 | Burkholderia phage vB_BceS_AH2 | Burkholderia phage vB_BceS_AH2 Burkholderia virus AH2 Ahduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58065 | 61.314 | Ahduovirus | Unclassified | Siphoviridae | Burkholderia |
| JF939047 | Burkholderia phage vB_BceS_KL1 | Burkholderia phage vB_BceS_KL1 Kilunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42832 | 54.606 | Kilunavirus | Unclassified | Siphoviridae | Burkholderia |
| GU071102 | Cyanophage NATL1A-7 | Cyanophage NATL1A-7 Prochlorococcus virus NATL1A7 Cheungvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47741 | 38.732 | Cheungvirus | Unclassified | Autographiviridae | Prochlorococcus |
| GU075905 | Prochlorococcus phage P-HM2 | Prochlorococcus phage P-HM2 unclassified Eurybiavirus Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 183806 | 38.115 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071108 | Prochlorococcus phage Syn33 | Prochlorococcus phage Syn33 Prochlorococcus virus Syn33 Brizovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174285 | 39.57 | Brizovirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071106 | Synechococcus phage Syn19 | Synechococcus phage Syn19 Synechococcus virus Syn19 Pontusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175230 | 41.251 | Pontusvirus | Unclassified | Myoviridae | Synechococcus |
| GU071105 | Prochlorococcus phage Syn1 | Prochlorococcus phage Syn1 Prochlorococcus virus Syn1 Vellamovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 191195 | 40.617 | Vellamovirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071103 | Prochlorococcus phage P-SSM7 | Prochlorococcus phage P-SSM7 Prochlorococcus virus PSSM7 Palaemonvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 182180 | 37.086 | Palaemonvirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071101 | Prochlorococcus phage P-HM1 | Prochlorococcus phage P-HM1 unclassified Eurybiavirus Eurybiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 181044 | 37.849 | Eurybiavirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071099 | Prochlorococcus phage P-RSM4 | Prochlorococcus phage P-RSM4 unclassified Thaumasvirus Thaumasvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176428 | 37.644 | Thaumasvirus | Unclassified | Myoviridae | Prochlorococcus |
| GU071098 | Synechococcus phage S-SSM7 | Synechococcus phage S-SSM7 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 232878 | 39.116 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| GU071097 | Synechococcus phage S-SSM5 | Synechococcus phage S-SSM5 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 176184 | 39.962 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| GU071096 | Synechococcus phage S-ShM2 | Synechococcus phage S-ShM2 Synechococcus virus SShM2 Ahtivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 179563 | 41.107 | Ahtivirus | Unclassified | Myoviridae | Synechococcus |
| GU071095 | Synechococcus phage S-SM2 | Synechococcus phage S-SM2 unclassified Bellamyvirus Bellamyvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 190789 | 40.426 | Bellamyvirus | Unclassified | Myoviridae | Synechococcus |
| GU071094 | Synechococcus phage S-SM1 | Synechococcus phage S-SM1 Synechococcus virus SSM1 Thetisvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 174079 | 41.138 | Thetisvirus | Unclassified | Myoviridae | Synechococcus |
| JN849462 | Vibriophage phi-pp2 | Vibriophage phi-pp2 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 246421 | 42.555 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| EF437941 | Escherichia phage Phi1 | Escherichia phage Phi1 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164270 | 40.499 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| CP000622 | Burkholderia phage phiE255 | Burkholderia phage phiE255 Bcepmuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37446 | 63.051 | Bcepmuvirus | Unclassified | Myoviridae | Burkholderia |
| CP000623 | Burkholderia phage phiE202 | Burkholderia phage phiE202 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35741 | 65.429 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| JQ780327 | Cronobacter phage phiES15 | Cronobacter phage phiES15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39974 | 53.542 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| JQ692107 | Vibrio phage SSP002 | Vibrio phage SSP002 Mardecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76350 | 48.777 | Mardecavirus | Unclassified | Siphoviridae | Vibrio |
| JQ007353 | Salmonella phage SE2 | Salmonella phage SE2 Jerseyvirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43221 | 49.645 | Jerseyvirus | Guernseyvirinae | Siphoviridae | Salmonella |
| JQ806764 | Salmonella phage vB_SosS_Oslo | Salmonella phage vB_SosS_Oslo Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49116 | 48.738 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| JQ806763 | Salmonella phage vB_SemP_Emek | Salmonella phage vB_SemP_Emek Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39783 | 47.651 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AJ006589 | Streptomyces virus phiC31 | Streptomyces virus phiC31 Lomovskayavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41491 | 63.619 | Lomovskayavirus | Unclassified | Siphoviridae | Streptomyces |
| JQ965703 | Yersinia phage YpsP-G | Yersinia phage YpsP-G Yersinia virus YpsPG Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38288 | 48.221 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| JQ965702 | Yersinia phage YpP-G | Yersinia phage YpP-G unclassified Berlinvirus Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39415 | 47.238 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| JQ965701 | Yersinia phage YpP-R | Yersinia phage YpP-R Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38284 | 48.339 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| JQ965700 | Yersinia phage YpP-Y | Yersinia phage YpP-Y Yersinia virus YpPY Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37714 | 48.348 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| AJ972879 | Yersinia phage phiR1-37 | Yersinia phage phiR1-37 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 262391 | 32.474 | Unclassified | Unclassified | Myoviridae | Yersinia |
| JN191664 | Bacillus phage BtCS33 | Bacillus phage BtCS33 Bacillus virus BtCS33 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41992 | 35.223 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| GQ141189 | Bifidobacterium phage Bbif-1 | Bifidobacterium phage Bbif-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40360 | 63.29 | Unclassified | Unclassified | Siphoviridae | Bifidobacterium |
| JQ729992 | Clostridium phage phiZP2 | Clostridium phage phiZP2 Clostridium virus phiZP2 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18078 | 34.478 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| JQ729991 | Clostridium phage phiCPV4 | Clostridium phage phiCPV4 Clostridium virus phiCPV4 Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17972 | 34.487 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| JQ729990 | Clostridium phage phiCP7R | Clostridium phage phiCP7R Clostridium virus phiCP7R Brucesealvirus Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18397 | 34.81 | Brucesealvirus | Unclassified | Guelinviridae | Clostridium |
| JQ307387 | Pseudomonas phage vB_Pae-Kakheti25 | Pseudomonas phage vB_Pae-Kakheti25 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42844 | 53.802 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| JQ307386 | Pseudomonas phage vB_Pae-TbilisiM32 | Pseudomonas phage vB_Pae-TbilisiM32 unclassified Phikmvvirus Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42966 | 62.27 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| JN022534 | Xanthomonas phage vB_XveM_DIBBI | Xanthomonas phage vB_XveM_DIBBI Xanthomonas virus DIBBI Dibbivirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49981 | 52.402 | Dibbivirus | Unclassified | Myoviridae | Xanthomonas |
| JQ660954 | Clostridium phage PhiS63 | Clostridium phage PhiS63 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33609 | 27.478 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| FJ409640 | Vibrio phage N4 | Vibrio phage N4 Vibrio virus N4 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38497 | 42.803 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| JQ691612 | Cronobacter phage CR3 | Cronobacter phage CR3 Certrevirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 149273 | 50.948 | Certrevirus | Vequintavirinae | Myoviridae | Cronobacter |
| JQ768459 | Pseudomonas phage Lu11 | Pseudomonas phage Lu11 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 280538 | 50.883 | Unclassified | Unclassified | Myoviridae | Pseudomonas |
| JQ619704 | Bacillus phage PBC1 | Bacillus phage PBC1 Bacillus virus PBC1 Pebcunavirus Gutmannvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41164 | 41.684 | Pebcunavirus | Gutmannvirinae | Siphoviridae | Bacillus |
| JQ062992 | Bacillus phage phIS3501 | Bacillus phage phIS3501 Bacillus virus phIS3501 Camtrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44401 | 34.864 | Camtrevirus | Unclassified | Siphoviridae | Bacillus |
| HE806280 | Acinetobacter phage AP22 | Acinetobacter phage AP22 Obolenskvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46387 | 37.743 | Obolenskvirus | Unclassified | Myoviridae | Acinetobacter |
| JN880423 | Salisaeta icosahedral phage 1 | Salisaeta icosahedral phage 1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 43788 | 57.226 | Unclassified | Unclassified | Unclassified | Salisaeta |
| HE608841 | Bacteroides phage B124-14 | Bacteroides phage B124-14 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47159 | 38.75 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| JQ779024 | Staphylococcus phage TEM123 | Staphylococcus phage TEM123 unclassified Dubowvirus Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43786 | 34.036 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| JQ779023 | Staphylococcus phage SMSAP5 | Staphylococcus phage SMSAP5 unclassified Triavirus Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45552 | 33.415 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| JQ740814 | Lactococcus phage ASCC544 | Lactococcus phage ASCC544 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32275 | 34.538 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740812 | Lactococcus phage ASCC531 | Lactococcus phage ASCC531 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31773 | 34.646 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740811 | Lactococcus phage ASCC527 | Lactococcus phage ASCC527 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32345 | 34.546 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740810 | Lactococcus phage ASCC506 | Lactococcus phage ASCC506 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32249 | 34.482 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740809 | Lactococcus phage ASCC502 | Lactococcus phage ASCC502 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32275 | 34.541 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740808 | Lactococcus phage ASCC497 | Lactococcus phage ASCC497 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31774 | 34.635 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740806 | Lactococcus phage ASCC476 | Lactococcus phage ASCC476 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32242 | 34.452 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740803 | Lactococcus phage ASCC460 | Lactococcus phage ASCC460 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32242 | 34.461 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740802 | Lactococcus phage ASCC454 | Lactococcus phage ASCC454 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32583 | 34.527 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740801 | Lactococcus phage ASCC406 | Lactococcus phage ASCC406 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32180 | 34.484 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740800 | Lactococcus phage ASCC397 | Lactococcus phage ASCC397 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32241 | 34.496 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740799 | Lactococcus phage ASCC395 | Lactococcus phage ASCC395 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32239 | 34.499 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740798 | Lactococcus phage ASCC368 | Lactococcus phage ASCC368 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32276 | 34.524 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740797 | Lactococcus phage ASCC365 | Lactococcus phage ASCC365 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32048 | 34.779 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740796 | Lactococcus phage ASCC358 | Lactococcus phage ASCC358 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32052 | 34.769 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740795 | Lactococcus phage ASCC356 | Lactococcus phage ASCC356 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32545 | 34.73 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740794 | Lactococcus phage ASCC337 | Lactococcus phage ASCC337 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32275 | 34.528 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740793 | Lactococcus phage ASCC324 | Lactococcus phage ASCC324 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32277 | 34.517 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740792 | Lactococcus phage ASCC310 | Lactococcus phage ASCC310 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32546 | 34.754 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740791 | Lactococcus phage ASCC287 | Lactococcus phage ASCC287 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32581 | 34.523 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740790 | Lactococcus phage ASCC284 | Lactococcus phage ASCC284 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33178 | 34.659 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740807 | Lactococcus phage ASCC489 | Lactococcus phage ASCC489 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30876 | 34.856 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ740805 | Lactococcus phage ASCC473 | Lactococcus phage ASCC473 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31772 | 34.644 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ246028 | Clostridium phage phi8074-B1 | Clostridium phage phi8074-B1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47595 | 35.382 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| JF917095 | Thermococcus prieurii virus 1 | Thermococcus prieurii virus 1 unclassified archaeal viruses Viruses | 21592 | 49.463 | Unclassified | Unclassified | Unclassified | Thermococcus |
| JN254801 | Pseudomonas phage MR299-2 | Pseudomonas phage MR299-2 unclassified Bruynoghevirus Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44789 | 52.044 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| JN254800 | Pseudomonas phage NH-4 | Pseudomonas phage NH-4 Pseudomonas virus NH4 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66116 | 55.549 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| HM452126 | Aeromonas phage phiAS5 | Aeromonas phage phiAS5 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 225268 | 42.997 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| EF116926 | Streptococcus phage SMP | Streptococcus phage SMP Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36019 | 41.581 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JN968479 | Haloarcula hispanica icosahedral virus 2 | Haloarcula hispanica icosahedral virus 2 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 30578 | 66.515 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| JQ513383 | Klebsiella phage vB_KleM_RaK2 | Klebsiella phage vB_KleM_RaK2 Klebsiella virus RaK2 Alcyoneusvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 345809 | 31.702 | Alcyoneusvirus | Unclassified | Myoviridae | Klebsiella |
| JF731128 | Enterococcus phage SAP6 | Enterococcus phage SAP6 Saphexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58619 | 39.996 | Saphexavirus | Unclassified | Siphoviridae | Enterococcus |
| HQ201307 | Cronobacter phage ENT39118 | Cronobacter phage ENT39118 Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39012 | 53.063 | Unclassified | Hendrixvirinae | Siphoviridae | Cronobacter |
| HE600015 | Dickeya virus Limestone | Dickeya virus Limestone Limestonevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152427 | 49.279 | Limestonevirus | Aglimvirinae | Ackermannviridae | Dickeya |
| JQ011318 | Escherichia phage TL-2011c | Escherichia phage TL-2011c Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60523 | 50.297 | Oslovirus | Sepvirinae | Podoviridae | Escherichia |
| JQ011317 | Escherichia phage TL-2011b | Escherichia phage TL-2011b Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44784 | 47.053 | Unclassified | Unclassified | Podoviridae | Escherichia |
| JQ011316 | Escherichia phage TL-2011a | Escherichia phage TL-2011a Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 24084 | 46.375 | Unclassified | Unclassified | Podoviridae | Escherichia |
| JQ347801 | Escherichia phage JLK-2012 | Escherichia phage JLK-2012 Escherichia virus JLK2012 Marienburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57198 | 50.719 | Marienburgvirus | Unclassified | Siphoviridae | Escherichia |
| JN800508 | Clostridium phage phi24R | Clostridium phage phi24R Clostridium virus phi24R Gregsiragusavirus Denniswatsonvirinae Guelinviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18919 | 27.803 | Gregsiragusavirus | Denniswatsonvirinae | Guelinviridae | Clostridium |
| JN939332 | Brucella phage Pr | Brucella phage Pr Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38253 | 48.182 | Perisivirus | Unclassified | Podoviridae | Brucella |
| JN939331 | Brucella phage Tb | Brucella phage Tb Perisivirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41148 | 48.194 | Perisivirus | Unclassified | Podoviridae | Brucella |
| JF713456 | Vibrio phage 1 | Vibrio phage 1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 81509 | 46.875 | Unclassified | Unclassified | Siphoviridae | Vibrio |
| HQ377373 | Liberibacter phage SC2 | Liberibacter phage SC2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38997 | 39.377 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| HQ377372 | Liberibacter phage SC1 | Liberibacter phage SC1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40048 | 41.196 | Unclassified | Unclassified | Podoviridae | Liberibacter |
| JQ362498 | Sphingomonas phage PAU | Sphingomonas phage PAU Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 219372 | 31.313 | Unclassified | Unclassified | Myoviridae | Sphingomonas |
| HQ259105 | Escherichia phage vB_EcoP_G7C | Escherichia phage vB_EcoP_G7C Gamaleyavirus Enquatrovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72917 | 43.416 | Gamaleyavirus | Enquatrovirinae | Schitoviridae | Escherichia |
| AB572858 | Vibrio phage ND1-fs1 | Vibrio phage ND1-fs1 unclassified Fibrovirus Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6856 | 43.436 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| AF323673 | Lactococcus phage bIL312 | Lactococcus phage bIL312 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15179 | 33.039 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF323672 | Lactococcus phage bIL311 | Lactococcus phage bIL311 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14510 | 34.238 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF323671 | Lactococcus phage bIL310 | Lactococcus phage bIL310 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14957 | 35.876 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF323670 | Lactococcus phage bIL309 | Lactococcus phage bIL309 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36949 | 35.679 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF323669 | Lactococcus phage bIL286 | Lactococcus phage bIL286 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41834 | 35.323 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF323668 | Lactococcus phage bIL285 | Lactococcus phage bIL285 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35538 | 35.188 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| B4PORFSX | Lactococcus phage phi41 | Lactococcus phage phi41 unclassified Skunavirus Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10170 | 35.447 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| JQ408219 | Ralstonia phage RSS0 | Ralstonia phage RSS0 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7288 | 62.047 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| JQ288021 | Salmonella phage SPN3UB | Salmonella phage SPN3UB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47355 | 49.613 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| HM770079 | Salmonella phage RE2010 | Salmonella phage RE2010 Salmonella virus RE2010 Felsduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34117 | 51.021 | Felsduovirus | Peduovirinae | Myoviridae | Salmonella |
| HM452125 | Aeromonas phage phiAS4 | Aeromonas phage phiAS4 Tulanevirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 163875 | 41.297 | Tulanevirus | Unclassified | Myoviridae | Aeromonas |
| JN651747 | Aeromonas phage phiAS7 | Aeromonas phage phiAS7 Aeromonas virus AS7 Aerosvirus Melnykvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41572 | 56.918 | Aerosvirus | Melnykvirinae | Autographiviridae | Aeromonas |
| FJ965538 | Streptococcus phage 5093 | Streptococcus phage 5093 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37184 | 38.03 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| JQ066768 | Rhodobacter phage RcapNL | Rhodobacter phage RcapNL Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40489 | 65.112 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| JQ010983 | Thermococcus prieurii virus 1 | Thermococcus prieurii virus 1 unclassified archaeal viruses Viruses | 21592 | 49.537 | Unclassified | Unclassified | Unclassified | Thermococcus |
| JQ245707 | Cyanophage S-TIM5 | Cyanophage S-TIM5 Synechococcus virus STIM5 Aurunvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 161440 | 40.458 | Aurunvirus | Unclassified | Myoviridae | Synechococcus |
| JN882298 | Escherichia phage phiKT | Escherichia phage phiKT Escherichia virus PhiKT Ermolevavirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42608 | 51.589 | Ermolevavirus | Unclassified | Autographiviridae | Escherichia |
| HQ698895 | Synechococcus phage S-CBS4 | Synechococcus phage S-CBS4 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 69420 | 50.828 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| HM480106 | Synechococcus phage S-CBS1 | Synechococcus phage S-CBS1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30332 | 58.75 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| GU936715 | Synechococcus phage S-CBS3 | Synechococcus phage S-CBS3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33004 | 60.72 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| GU936714 | Synechococcus phage S-CBS2 | Synechococcus phage S-CBS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72332 | 54.474 | Unclassified | Unclassified | Siphoviridae | Synechococcus |
| JN600672 | Mycobacterium phage Fezzik | Mycobacterium phage Fezzik unclassified Bronvirus Bronvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74155 | 58.685 | Bronvirus | Unclassified | Siphoviridae | Mycobacterium |
| JN627160 | Pseudomonas phage OBP | Pseudomonas phage OBP Petsuvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 284757 | 43.455 | Petsuvirus | Unclassified | Myoviridae | Pseudomonas |
| DQ320509 | Lactobacillus phage KC5a | Lactobacillus phage KC5a Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38239 | 36.897 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| JN808773 | Pseudomonas virus FHA0480 | Pseudomonas virus FHA0480 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37374 | 64.141 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| HQ201308 | Cronobacter phage ENT47670 | Cronobacter phage ENT47670 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47611 | 51.593 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| JN585957 | Erwinia phage PEp14 | Erwinia phage PEp14 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60714 | 62.78 | Unclassified | Unclassified | Podoviridae | Erwinia |
| HQ711985 | Pseudomonas virus PMG1 | Pseudomonas virus PMG1 Detrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54024 | 57.467 | Detrevirus | Unclassified | Siphoviridae | Pseudomonas |
| HQ711984 | Pseudomonas phage phi297 | Pseudomonas phage phi297 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49135 | 62.147 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| HE584812 | Pseudomonas virus LPB1 | Pseudomonas virus LPB1 Casadabanvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36814 | 64.364 | Casadabanvirus | Unclassified | Siphoviridae | Pseudomonas |
| JN391180 | Salmonella phage SPN1S | Salmonella phage SPN1S Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38684 | 50.163 | Uetakevirus | Unclassified | Podoviridae | Salmonella |
| GQ153922 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 44.827 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| AY583236 | Mycoplasma phage phiMFV1 | Mycoplasma phage phiMFV1 unclassified bacterial viruses Viruses | 18855 | 24.826 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| AY583235 | Mycoplasma phage phiMFV1 | Mycoplasma phage phiMFV1 unclassified bacterial viruses Viruses | 23300 | 25.888 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| HQ683709 | Planktothrix phage PaV-LD | Planktothrix phage PaV-LD Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 95299 | 41.45 | Unclassified | Unclassified | Siphoviridae | Planktothrix |
| HQ259103 | Salmonella phage SFP10 | Salmonella phage SFP10 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157950 | 44.534 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| JN797797 | Bacillus phage BCP78 | Bacillus phage BCP78 Tsarbombavirus Bastillevirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 156176 | 39.859 | Tsarbombavirus | Bastillevirinae | Herelleviridae | Bacillus |
| AB626963 | Staphylococcus phage S13′ | Staphylococcus phage S13′ unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18186 | 29.215 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| AB626962 | Staphylococcus phage S24-1 | Staphylococcus phage S24-1 Staphylococcus virus S24-1 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18168 | 28.908 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| JN116828 | Nocardia phage NBR1 | Nocardia phage NBR1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46140 | 67.503 | Unclassified | Unclassified | Siphoviridae | Nocardia |
| JN116824 | Rhodococcus phage REQ3 | Rhodococcus phage REQ3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39474 | 65.95 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| JN116823 | Rhodococcus phage REQ2 | Rhodococcus phage REQ2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49330 | 65.406 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| JN116822 | Rhodococcus phage RRH1 | Rhodococcus phage RRH1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14270 | 68.381 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| AY940168 | Prochlorococcus phage P-SSM4 | Prochlorococcus phage P-SSM4 Prochlorococcus virus PSSM4 Ronodorvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178249 | 36.744 | Ronodorvirus | Unclassified | Myoviridae | Prochlorococcus |
| AY939844 | Prochlorococcus phage P-SSM2 | Prochlorococcus phage P-SSM2 Prochlorococcus virus PSSM2 Salacisavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 252401 | 35.506 | Salacisavirus | Unclassified | Myoviridae | Prochlorococcus |
| JN641803 | Salmonella phage SPN3US | Salmonella phage SPN3US Seoulvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 240413 | 48.54 | Seoulvirus | Unclassified | Myoviridae | Salmonella |
| JF314845 | Cronobacter phage ES2 | Cronobacter phage ES2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22162 | 50.799 | Unclassified | Unclassified | Siphoviridae | Cronobacter |
| JF923797 | Gordonia phage GRU1 | Gordonia phage GRU1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65766 | 65.475 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| JF923796 | Gordonia phage GTE5 | Gordonia phage GTE5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65839 | 65.053 | Unclassified | Unclassified | Siphoviridae | Gordonia |
| JN190960 | Rhodobacter phage RcapMu | Rhodobacter phage RcapMu Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39283 | 64.896 | Unclassified | Unclassified | Siphoviridae | Rhodobacter |
| JN126049 | Salmonella phage Sh19 | Salmonella phage Sh19 Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157785 | 44.685 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| JN848801 | Vibrio phage VCY-phi | Vibrio phage VCY-phi Vicialiavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7103 | 41.405 | Vicialiavirus | Unclassified | Inoviridae | Vibrio |
| JN035618 | Gordonia phage GTE7 | Gordonia phage GTE7 Gordonia virus GTE7 Getseptimavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74431 | 56.783 | Getseptimavirus | Unclassified | Siphoviridae | Gordonia |
| HM208537 | Escherichia phage HK639 | Escherichia phage HK639 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49576 | 52.451 | Unclassified | Unclassified | Siphoviridae | Escherichia |
| HM173637 | Escherichia phage HK75 | Escherichia phage HK75 Saikungvirus Hendrixvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36661 | 50.19 | Saikungvirus | Hendrixvirinae | Siphoviridae | Escherichia |
| JF719734 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.503 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| JF719733 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.581 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| JF719732 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.393 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| JF719728 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.616 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JF719727 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.634 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JF719726 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.634 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GU070616 | Salmonella phage PVPSE1 | Salmonella phage PVPSE1 Salmonella virus PVPSE1 Seunavirus Vequintavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 145964 | 45.607 | Seunavirus | Vequintavirinae | Myoviridae | Salmonella |
| JF317274 | Acinetobacter phage Abp53 | Acinetobacter phage Abp53 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10290 | 38.62 | Unclassified | Unclassified | Myoviridae | Acinetobacter |
| AP011956 | Staphylococcus phage phi7247PVL | Staphylococcus phage phi7247PVL Staphylococcus virus phi7247PVL Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42481 | 33.309 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AP011955 | Staphylococcus phage phi5967PVL | Staphylococcus phage phi5967PVL unclassified Biseptimavirus Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42461 | 33.311 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| HQ141410 | Lactobacillus phage LF1 | Lactobacillus phage LF1 Lactobacillus virus LF1 Lafunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42606 | 45.005 | Lafunavirus | Unclassified | Siphoviridae | Lactobacillus |
| HM563683 | Escherichia phage vB_EcoM_VR7 | Escherichia phage vB_EcoM_VR7 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169285 | 40.333 | Gaprivervirus | Tevenvirinae | Myoviridae | Escherichia |
| FR852584 | Staphylococcus phage ISP | Staphylococcus phage ISP Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 138339 | 30.422 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| JF770475 | Escherichia phage phiEB49 | Escherichia phage phiEB49 Escherichia virus phiEB49 Lindendrivevirus Rogunavirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47180 | 44.019 | Lindendrivevirus | Rogunavirinae | Drexlerviridae | Escherichia |
| HQ141411 | Lactobacillus phage Sha1 | Lactobacillus phage Sha1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41726 | 40.605 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| HQ829472 | Escherichia phage Bp7 | Escherichia phage Bp7 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168066 | 39.493 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| AP011617 | Thermus phage TMA | Thermus phage TMA Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 151483 | 32.625 | Unclassified | Unclassified | Myoviridae | Thermus |
| AP011616 | Thermus phage phiYS40 | Thermus phage phiYS40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152372 | 32.586 | Unclassified | Unclassified | Myoviridae | Thermus |
| HQ728266 | Erwinia phage vB_EamP-S6 | Erwinia phage vB_EamP-S6 Erwinia virus S6 Waedenswilvirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74669 | 52.09 | Waedenswilvirus | Unclassified | Schitoviridae | Erwinia |
| HQ728264 | Erwinia phage vB_EamM-Y2 | Erwinia phage vB_EamM-Y2 Erwinia virus Y2 Loessnervirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56621 | 44.229 | Loessnervirus | Cleopatravirinae | Chaseviridae | Erwinia |
| HQ728263 | Erwinia phage vB_Eam-MM7 | Erwinia phage vB_Eam-MM7 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84694 | 43.387 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| AJ536073 | Bacillus phage pGIL01 | Bacillus phage pGIL01 Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14931 | 39.729 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| FR823450 | Campylobacter virus CP81 | Campylobacter virus CP81 Fletchervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 132454 | 26.042 | Fletchervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| HM997019 | Salmonella phage 7-11 | Salmonella phage 7-11 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 89916 | 44.128 | Unclassified | Unclassified | Podoviridae | Salmonella |
| HM071924 | Escherichia phage IME08 | Escherichia phage IME08 unclassified Dhakavirus Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172253 | 39.591 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| AB605730 | Bacillus phage SP-10 | Bacillus phage SP-10 unclassified Spounavirinae Spounavirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 143986 | 40.487 | Unclassified | Spounavirinae | Herelleviridae | Bacillus |
| JN202312 | Escherichia phage ime09 | Escherichia phage ime09 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166499 | 35.675 | Tequatrovirus | Tevenvirinae | Myoviridae | Escherichia |
| HQ615693 | Synechococcus phage S-CRM01 | Synechococcus phage S-CRM01 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 178563 | 39.745 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| HQ641354 | Vibrio phage ICP1_2004_A | Vibrio phage ICP1_2004_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 128083 | 37.165 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641353 | Vibrio phage ICP1_2001_A | Vibrio phage ICP1_2001_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124826 | 37.069 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641352 | Vibrio phage ICP1_2005_A | Vibrio phage ICP1_2005_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 129373 | 37.103 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641351 | Vibrio phage ICP1_2006_A | Vibrio phage ICP1_2006_A Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123104 | 37.077 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641350 | Vibrio phage ICP1_2006_B | Vibrio phage ICP1_2006_B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 123097 | 37.074 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641349 | Vibrio phage ICP1_2006_C | Vibrio phage ICP1_2006_C Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124497 | 37.094 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641348 | Vibrio phage ICP1_2006_D | Vibrio phage ICP1_2006_D Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 124497 | 37.092 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641347 | Vibrio phage ICP1 | Vibrio phage ICP1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 125956 | 37.073 | Unclassified | Unclassified | Myoviridae | Vibrio |
| HQ641346 | Vibrio phage ICP2_2006_A | Vibrio phage ICP2_2006_A Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48626 | 42.481 | Unclassified | Unclassified | Zobellviridae | Vibrio |
| HQ641345 | Vibrio phage ICP2 | Vibrio phage ICP2 Vibrio virus ICP2 Icepovirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49675 | 42.677 | Icepovirus | Unclassified | Zobellviridae | Vibrio |
| HQ641344 | Vibrio phage ICP3_2007_A | Vibrio phage ICP3_2007_A unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39088 | 42.855 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| HQ641343 | Vibrio phage ICP3_2008_A | Vibrio phage ICP3_2008_A unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39349 | 42.807 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| HQ641341 | Vibrio phage ICP3_2009_B | Vibrio phage ICP3_2009_B unclassified Chatterjeevirus Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39042 | 42.767 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| HQ641340 | Vibrio phage ICP3 | Vibrio phage ICP3 Vibrio virus ICP3 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39162 | 42.858 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| EU409559 | Lactobacillus phage JCL1032 | Lactobacillus phage JCL1032 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49433 | 44.246 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| FR725450 | Nitrososphaera phage Pro-Nvie1 | Nitrososphaera phage Pro-Nvie1 unclassified bacterial viruses Viruses | 23750 | 52.985 | Unclassified | Unclassified | Unclassified | Nitrososphaera |
| HM753718 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.748 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753717 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.783 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753716 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.785 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753715 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753714 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753713 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753712 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753711 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.748 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753710 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753709 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753708 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753707 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.766 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753706 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.671 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753705 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.653 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753704 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.634 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753703 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.653 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753702 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.83 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753701 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.82 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753700 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.82 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753699 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.801 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753698 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.756 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753697 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.82 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753696 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.83 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753695 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.756 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753694 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.812 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753693 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.849 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753692 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.774 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753691 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.793 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753690 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.737 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753689 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753688 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753687 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.849 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753686 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.801 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753685 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.774 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753684 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.737 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753683 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.774 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753682 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.83 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753681 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.737 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753680 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.774 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753679 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753678 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.838 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753677 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.783 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753676 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753675 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753674 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753673 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753672 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753671 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.764 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753670 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753669 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753668 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753667 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753666 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753665 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753664 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753663 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753662 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753661 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753660 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753659 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM753658 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| JF767210 | Clostridium phage phiCP9O | Clostridium phage phiCP9O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39594 | 30.191 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| JF767209 | Clostridium phage phiCP34O | Clostridium phage phiCP34O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38309 | 30.528 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| JF767208 | Clostridium phage phiCP13O | Clostridium phage phiCP13O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38329 | 30.363 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| HQ264138 | Microviridae phi-CA82 | Microviridae phi-CA82 unclassified Microviridae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5514 | 39.717 | Unclassified | Unclassified | Microviridae | Unspecified |
| JF412298 | EBPR siphovirus 3 | EBPR siphovirus 3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13588 | 61.709 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| JF412296 | EBPR podovirus 3 | EBPR podovirus 3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38991 | 41.838 | Unclassified | Unclassified | Podoviridae | Unspecified |
| JF412295 | EBPR podovirus 2 | EBPR podovirus 2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40628 | 63.917 | Unclassified | Unclassified | Podoviridae | Unspecified |
| JF412294 | EBPR podovirus 1 | EBPR podovirus 1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36122 | 59.523 | Unclassified | Unclassified | Podoviridae | Unspecified |
| AY769964 | Chlamydia phage 4 | Chlamydia phage 4 unclassified Chlamydiamicrovirus Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4530 | 41.17 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| HQ403646 | Gordonia phage GTE2 | Gordonia phage GTE2 Gordonia virus GTE2 Emalynvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45530 | 60.343 | Emalynvirus | Unclassified | Siphoviridae | Gordonia |
| HM480846 | Escherichia phage K30 | Escherichia phage K30 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40940 | 51.426 | Przondovirus | Studiervirinae | Autographiviridae | Escherichia |
| GU988610 | Pseudomonas phage JG004 | Pseudomonas phage JG004 Pakpunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 93017 | 49.261 | Pakpunavirus | Unclassified | Myoviridae | Pseudomonas |
| GQ443085 | Clostridium phage phiCP26F | Clostridium phage phiCP26F Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39188 | 30.262 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| HQ630627 | Pseudomonas phage PhiPA3 | Pseudomonas phage PhiPA3 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 309208 | 47.725 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| FJ373894 | Shigella phage Ag3 | Shigella phage Ag3 Agtrevirus Aglimvirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 158006 | 50.398 | Agtrevirus | Aglimvirinae | Ackermannviridae | Shigella |
| JF344709 | Bdellovibrio phage phi1402 | Bdellovibrio phage phi1402 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23931 | 50.349 | Unclassified | Unclassified | Myoviridae | Bdellovibrio |
| HM568888 | Clostridium phage phiCD38-2 | Clostridium phage phiCD38-2 Clostridioides virus phiCD38-2 Leicestervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41090 | 30.83 | Leicestervirus | Unclassified | Siphoviridae | Clostridium |
| FR751545 | Pantoea phage LIMEzero | Pantoea phage LIMEzero Waewaevirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43032 | 55.352 | Waewaevirus | Unclassified | Autographiviridae | Pantoea |
| FR687252 | Pantoea phage LIMElight | Pantoea phage LIMElight Limelightvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44546 | 53.991 | Limelightvirus | Unclassified | Autographiviridae | Pantoea |
| HM150760 | Stenotrophomonas phage phiSHP2 | Stenotrophomonas phage phiSHP2 unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5819 | 61.505 | Unclassified | Unclassified | Inoviridae | Stenotrophomonas |
| GQ413938 | Klebsiella phage KP34 | Klebsiella phage KP34 Drulisvirus Slopekvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43809 | 54.073 | Drulisvirus | Slopekvirinae | Autographiviridae | Klebsiella |
| FJ706172 | Propionibacterium phage PAS50 | Propionibacterium phage PAS50 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29017 | 53.972 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| FJ706171 | Propionibacterium phage PAD20 | Propionibacterium phage PAD20 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29074 | 54.1 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| HM151342 | Roseobacter phage RDJL Phi 1 | Roseobacter phage RDJL Phi 1 Xiamenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62668 | 57.881 | Xiamenvirus | Unclassified | Siphoviridae | Roseobacter |
| GU583987 | Pseudomonas phage phiIBB-PF7A | Pseudomonas phage phiIBB-PF7A Pseudomonas virus IBBPF7A Pifdecavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40973 | 56.271 | Pifdecavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| HM035025 | Shigella phage Shfl2 | Shigella phage Shfl2 Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165919 | 35.567 | Tequatrovirus | Tevenvirinae | Myoviridae | Shigella |
| HQ110083 | Cronobacter phage ESSI-2 | Cronobacter phage ESSI-2 Cronobacter virus ESSI2 Seongnamvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28765 | 55.168 | Seongnamvirus | Peduovirinae | Myoviridae | Cronobacter |
| HQ127381 | Staphylococcus phage TEM126 | Staphylococcus phage TEM126 Staphylococcus virus TEM126 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41882 | 33.676 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| HQ186308 | Acinetobacter phage phiAB1 | Acinetobacter phage phiAB1 Friunavirus Beijerinckvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41526 | 39.086 | Friunavirus | Beijerinckvirinae | Autographiviridae | Acinetobacter |
| HQ906664 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA4 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA4 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21272 | 35.836 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| HQ906663 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA2 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA2 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40003 | 33.31 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| HQ906662 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA1 | Wolbachia endosymbiont wVitA of Nasonia vitripennis phage WOVitA1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66810 | 34.505 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| HQ406778 | Salmonella virus SPC35 | Salmonella virus SPC35 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 118351 | 39.385 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| GU393987 | Caulobacter phage Cd1 | Caulobacter phage Cd1 unclassified Okabevirinae Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41581 | 61.213 | Unclassified | Okabevirinae | Autographiviridae | Caulobacter |
| FR823298 | Pseudomonas phage phi15 | Pseudomonas phage phi15 Pseudomonas virus Phi15 Troedvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39562 | 58.164 | Troedvirus | Studiervirinae | Autographiviridae | Pseudomonas |
| FJ000341 | Salmonella phage g341c | Salmonella phage g341c Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40975 | 47.402 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| GU573886 | Salmonella phage ST160 | Salmonella phage ST160 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40986 | 47.06 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| HM486077 | Tsukamurella phage TPA2 | Tsukamurella phage TPA2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61440 | 69.574 | Unclassified | Unclassified | Siphoviridae | Tsukamurella |
| FR671411 | Streptococcus phage 2167 | Streptococcus phage 2167 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36217 | 40.655 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671410 | Streptococcus phage 8140 | Streptococcus phage 8140 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35890 | 40.496 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671409 | Streptococcus phage 11865 | Streptococcus phage 11865 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32602 | 40.228 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671408 | Streptococcus phage 23782 | Streptococcus phage 23782 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32031 | 40.298 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671407 | Streptococcus phage 34117 | Streptococcus phage 34117 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37636 | 40.384 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671406 | Streptococcus phage 040922 | Streptococcus phage 040922 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40104 | 39.734 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FR671405 | Streptococcus phage V22 | Streptococcus phage V22 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37159 | 40.082 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| EU652770 | Morganella phage MmP1 | Morganella phage MmP1 Morganella virus MmP1 Minipunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38457 | 46.519 | Minipunavirus | Studiervirinae | Autographiviridae | Morganella |
| EU716414 | Pseudomonas phage PB1 | Pseudomonas phage PB1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65764 | 54.928 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| AM084414 | Escherichia phage K1F | Escherichia phage K1F Escherichia virus K1F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39699 | 49.777 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| FN997652 | Streptococcus phage phi-SsUD.1 | Streptococcus phage phi-SsUD.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63748 | 39.713 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FN422399 | Pseudomonas phage LIT1 | Pseudomonas phage LIT1 Litunavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 72544 | 55.019 | Litunavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| FN422398 | Pseudomonas phage LUZ7 | Pseudomonas phage LUZ7 Luzseptimavirus Migulavirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74901 | 53.218 | Luzseptimavirus | Migulavirinae | Schitoviridae | Pseudomonas |
| FM887021 | Pseudomonas phage SN | Pseudomonas phage SN Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66390 | 55.582 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| FM897211 | Pseudomonas phage 14-1 | Pseudomonas phage 14-1 Pseudomonas virus 14-1 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66235 | 55.593 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| FM201282 | Pseudomonas phage LMA2 | Pseudomonas phage LMA2 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66530 | 55.617 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| FM201281 | Pseudomonas phage LBL3 | Pseudomonas phage LBL3 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 64427 | 55.469 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| FQ482086 | Erwinia phage phiEa100 | Erwinia phage phiEa100 unclassified Eracentumvirus Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45554 | 49.677 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| FQ482085 | Erwinia phage phiEt88 | Erwinia phage phiEt88 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47279 | 47.33 | Unclassified | Unclassified | Myoviridae | Erwinia |
| FQ482084 | Erwinia phage phiEa1H | Erwinia phage phiEa1H unclassified Eracentumvirus Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45522 | 49.681 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| FQ482083 | Erwinia phage phiEa104 | Erwinia phage phiEa104 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84565 | 43.83 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| GU580943 | Rhodococcus phage ReqiPine5 | Rhodococcus phage ReqiPine5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59231 | 67.095 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| GU580942 | Rhodococcus phage ReqiPoco6 | Rhodococcus phage ReqiPoco6 Pepyhexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 78064 | 53.265 | Pepyhexavirus | Unclassified | Siphoviridae | Rhodococcus |
| GU580941 | Rhodococcus phage ReqiPepy6 | Rhodococcus phage ReqiPepy6 Pepyhexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 76797 | 53.411 | Pepyhexavirus | Unclassified | Siphoviridae | Rhodococcus |
| GU580940 | Rhodococcus phage ReqiDocB7 | Rhodococcus phage ReqiDocB7 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 75772 | 56.658 | Unclassified | Unclassified | Siphoviridae | Rhodococcus |
| HQ641380 | Enterobacter phage EcP1 | Enterobacter phage EcP1 Enterobacter virus EcP1 Eceepunavirus Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59080 | 36.875 | Eceepunavirus | Unclassified | Schitoviridae | Enterobacter |
| FQ312032 | Salmonella virus ViI | Salmonella virus ViI Kuttervirus Cvivirinae Ackermannviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 157061 | 45.22 | Kuttervirus | Cvivirinae | Ackermannviridae | Salmonella |
| FR667955 | Salmonella phage Vi06 | Salmonella phage Vi06 Salmonella virus Vi06 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38368 | 48.926 | Teseptimavirus | Studiervirinae | Autographiviridae | Salmonella |
| FN667789 | Campylobacter virus CPt10 | Campylobacter virus CPt10 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 175720 | 27.337 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| FN667788 | Campylobacter virus CP220 | Campylobacter virus CP220 Firehammervirus Eucampyvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177534 | 27.359 | Firehammervirus | Eucampyvirinae | Myoviridae | Campylobacter |
| FN297812 | Vibrio phage VP585 | Vibrio phage VP585 Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42612 | 50.878 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| FN594518 | Pseudomonas phage phi-2 | Pseudomonas phage phi-2 Pseudomonas virus f2 Tunggulvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43144 | 58.914 | Tunggulvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| FM864213 | Streptococcus phage phi-m46.1 | Streptococcus phage phi-m46.1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58144 | 40.068 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FM207411 | Synechococcus phage S-RSM4 | Synechococcus phage S-RSM4 unclassified Cymopoleiavirus Cymopoleiavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 194454 | 41.12 | Cymopoleiavirus | Unclassified | Myoviridae | Synechococcus |
| FN391954 | Streptococcus phage PH10 | Streptococcus phage PH10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31276 | 39.49 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| FM180578 | Enterobacteria phage 2851 | Enterobacteria phage 2851 Enterobacteria virus 2851 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57248 | 51.053 | Pankowvirus | Unclassified | Siphoviridae | Enterobacteria |
| FM163528 | Streptococcus phage PH15 | Streptococcus phage PH15 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39136 | 40.507 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AM491472 | Salmonella phage Vi II-E1 | Salmonella phage Vi II-E1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45051 | 46.126 | Unclassified | Unclassified | Siphoviridae | Salmonella |
| CU468217 | Azospirillum phage Cd | Azospirillum phage Cd Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62337 | 63.932 | Unclassified | Unclassified | Siphoviridae | Azospirillum |
| AM749441 | Pseudomonas virus Yua | Pseudomonas virus Yua Yuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58663 | 64.262 | Yuavirus | Unclassified | Siphoviridae | Pseudomonas |
| AM910650 | Pseudomonas virus LUZ24 | Pseudomonas virus LUZ24 Bruynoghevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45625 | 52.248 | Bruynoghevirus | Unclassified | Podoviridae | Pseudomonas |
| AM749121 | Streptococcus phage M102 | Streptococcus phage M102 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31147 | 39.211 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AM183667 | Yersinia phage Berlin | Yersinia phage Berlin Yersinia virus Berlin Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38564 | 47.189 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| X97918 | Bacillus phage SPP1 | Bacillus phage SPP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44010 | 43.715 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| AM084415 | Escherichia virus K1E | Escherichia virus K1E Vectrevirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45251 | 45.053 | Vectrevirus | Molineuxvirinae | Autographiviridae | Escherichia |
| AJ697969 | Pseudomonas phage EL | Pseudomonas phage EL Elvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 211215 | 49.327 | Elvirus | Unclassified | Myoviridae | Pseudomonas |
| AM156909 | Escherichia virus Rtp | Escherichia virus Rtp Rtpvirus Braunvirinae Drexlerviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46219 | 44.283 | Rtpvirus | Braunvirinae | Drexlerviridae | Escherichia |
| AJ630128 | Synechococcus phage S-PM2 | Synechococcus phage S-PM2 Synechococcus virus SPM2 Nodensvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 196280 | 37.82 | Nodensvirus | Unclassified | Myoviridae | Synechococcus |
| AJ421943 | Oenococcus phage fOg44 | Oenococcus phage fOg44 unclassified bacterial viruses Viruses | 10858 | 35.458 | Unclassified | Unclassified | Unclassified | Oenococcus |
| AJ304858 | Enterobacteria phage CP-1639 | Enterobacteria phage CP-1639 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50625 | 50.37 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| AJ556162 | Enterobacteria phage BP-4795 | Enterobacteria phage BP-4795 Enterobacteria virus BP4795 Marienburgvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57930 | 50.608 | Marienburgvirus | Unclassified | Siphoviridae | Enterobacteria |
| AJ550940 | Streptomyces virus phiBT1 | Streptomyces virus phiBT1 Lomovskayavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41831 | 62.762 | Lomovskayavirus | Unclassified | Siphoviridae | Streptomyces |
| BX571876 | Leptospira phage LE1 | Leptospira phage LE1 Leptospira virus LE1 Saintgironsvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73623 | 38.527 | Saintgironsvirus | Unclassified | Myoviridae | Leptospira |
| AJ609634 | Streptococcus phage EJ-1 | Streptococcus phage EJ-1 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42935 | 39.641 | Unclassified | Unclassified | Myoviridae | Streptococcus |
| AJ604531 | Salmonella phage 5 | Salmonella phage 5 unclassified Tequintavirus Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 13751 | 40.833 | Tequintavirus | Markadamsvirinae | Demerecviridae | Salmonella |
| AJ604530 | Escherichia virus BF23 | Escherichia virus BF23 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 14504 | 40.306 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| AJ564013 | Yersinia phage PY54 | Yersinia phage PY54 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46339 | 44.574 | Unclassified | Unclassified | Siphoviridae | Yersinia |
| AJ505558 | Pseudomonas phage phiKMV | Pseudomonas phage phiKMV Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42519 | 62.297 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| AJ560763 | Haemophilus phage Aaphi23 | Haemophilus phage Aaphi23 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43033 | 42.456 | Unclassified | Unclassified | Myoviridae | Haemophilus |
| AJ312240 | Listeria phage PSA | Listeria phage PSA Psavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37618 | 34.725 | Psavirus | Unclassified | Siphoviridae | Listeria |
| AJ245616 | Lactococcus phage BK5-T | Lactococcus phage BK5-T Lactococcus virus BK5T Sandinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40003 | 34.975 | Sandinevirus | Unclassified | Siphoviridae | Lactococcus |
| AJ251789 | Lactobacillus phage A2 | Lactobacillus phage A2 Lactobacillus virus A2 Ashduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43411 | 44.89 | Ashduovirus | Unclassified | Siphoviridae | Lactobacillus |
| AJ302074 | Streptococcus phage MM1 | Streptococcus phage MM1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40248 | 38.412 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AJ298298 | Enterobacteria phage phiP27 | Enterobacteria phage phiP27 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42575 | 49.351 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| AJ318471 | Enterobacteria phage T3 | Enterobacteria phage T3 Escherichia virus T3 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38208 | 49.901 | Teetrevirus | Studiervirinae | Autographiviridae | Escherichia |
| X96987 | Bacillus phage GA1 | Bacillus phage GA1 Gaunavirus Tatarstanvirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21129 | 34.654 | Gaunavirus | Tatarstanvirinae | Salasmaviridae | Bacillus |
| AJ251805 | Yersinia phage phiYeO3-12 | Yersinia phage phiYeO3-12 Yersinia virus YeO3-12 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39600 | 50.631 | Teetrevirus | Studiervirinae | Autographiviridae | Yersinia |
| AJ242593 | Listeria phage A118 | Listeria phage A118 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40834 | 36.107 | Unclassified | Unclassified | Siphoviridae | Listeria |
| AJ131519 | Lactobacillus phage phiadh | Lactobacillus phage phiadh Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43785 | 35.572 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| Y13918 | Pseudomonas virus phiCTX | Pseudomonas virus phiCTX Citexvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35559 | 62.617 | Citexvirus | Peduovirinae | Myoviridae | Pseudomonas |
| Z47794 | Streptococcus phage Cp1 | Streptococcus phage Cp1 Cepunavirus Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19343 | 38.831 | Cepunavirus | Unclassified | Salasmaviridae | Streptococcus |
| X98106 | Lactobacillus phage phig1e | Lactobacillus phage phig1e Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42259 | 43.061 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| X99260 | Bacillus virus B103 | Bacillus virus B103 Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18630 | 37.66 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| X51522 | Enterobacteria phage P4 | Enterobacteria phage P4 Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 11624 | 49.527 | Unclassified | Unclassified | Unclassified | Enterobacteria |
| V01146 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39937 | 48.399 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| X51344 | Spiroplasma phage R8A2B | Spiroplasma phage R8A2B Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8273 | 22.894 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| EU588980 | Clostridium phage phiCP39-O | Clostridium phage phiCP39-O Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38753 | 30.421 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| AB597179 | Ralstonia phage RSB2 | Ralstonia phage RSB2 Ralstonia virus RSB2 Kelmasvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40411 | 61.751 | Kelmasvirus | Unclassified | Autographiviridae | Ralstonia |
| HQ424691 | Enterobacteria phage VT2phi_272 | Enterobacteria phage VT2phi_272 unclassified Oslovirus Oslovirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65955 | 50.104 | Oslovirus | Sepvirinae | Podoviridae | Enterobacteria |
| DQ372923 | Streptomyces phage mu1/6 | Streptomyces phage mu1/6 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38194 | 71.194 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| DQ227764 | Lactococcus virus jj50 | Lactococcus virus jj50 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27453 | 34.943 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| AY962392 | Aeromonas virus 31 | Aeromonas virus 31 unclassified Biquartavirus Biquartavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 172963 | 43.913 | Biquartavirus | Unclassified | Myoviridae | Aeromonas |
| AP008983 | Clostridium phage c-st | Clostridium phage c-st Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 185683 | 26.284 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| AP005154 | Escherichia phage Stx2 II | Escherichia phage Stx2 II Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62706 | 49.901 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| AP000400 | Enterobacteria phage VT1-Sakai | Enterobacteria phage VT1-Sakai Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56480 | 51.71 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| AP000422 | Enterobacteria phage VT2-Sakai | Enterobacteria phage VT2-Sakai unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 65510 | 50.002 | Traversvirus | Sepvirinae | Podoviridae | Enterobacteria |
| HQ333270 | Yersinia phage Yep-phi | Yersinia phage Yep-phi Yersinia virus Yepf Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38616 | 47.1 | Berlinvirus | Studiervirinae | Autographiviridae | Yersinia |
| GU323708 | Lactobacillus phage phiPYB5 | Lactobacillus phage phiPYB5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32847 | 45.468 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| GU815091 | Pseudomonas phage JG024 | Pseudomonas phage JG024 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66275 | 55.621 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| HM064452 | Ralstonia phage PE226 | Ralstonia phage PE226 Parhipatevirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5475 | 61.662 | Parhipatevirus | Unclassified | Inoviridae | Ralstonia |
| EF160123 | Erwinia phage Era103 | Erwinia phage Era103 Eracentumvirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45445 | 49.838 | Eracentumvirus | Molineuxvirinae | Autographiviridae | Erwinia |
| AY343333 | Escherichia phage RB49 | Escherichia phage RB49 Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164018 | 40.442 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| AF234173 | Escherichia phage P1 | Escherichia phage P1 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 94481 | 47.31 | Punavirus | Unclassified | Myoviridae | Escherichia |
| AY303349 | Escherichia phage RB69 | Escherichia phage RB69 Mosigvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167560 | 37.656 | Mosigvirus | Tevenvirinae | Myoviridae | Escherichia |
| AF493143 | Pseudomonas phage gh-1 | Pseudomonas phage gh-1 Ghunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37359 | 57.416 | Ghunavirus | Studiervirinae | Autographiviridae | Pseudomonas |
| AF424783 | Staphylococcus virus 13 | Staphylococcus virus 13 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42722 | 33.472 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AF176034 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176033 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5388 | 44.655 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176032 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.598 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176031 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.653 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176030 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.523 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176029 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.671 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176028 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.598 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF176027 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| PF3COMN | Pseudomonas phage Pf3 | Pseudomonas phage Pf3 Tertilicivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5833 | 45.363 | Tertilicivirus | Unclassified | Inoviridae | Pseudomonas |
| GQ241246 | Clavibacter phage CMP1 | Clavibacter phage CMP1 Clavibacter virus CMP1 Cimpunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58652 | 57.007 | Cimpunavirus | Unclassified | Siphoviridae | Clavibacter |
| FJ539135 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4273 | 50.573 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ539134 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4273 | 50.41 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ539133 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4243 | 51.614 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ539132 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4241 | 50.931 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483844 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4219 | 47.428 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483843 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4218 | 47.274 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483842 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4218 | 46.728 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483841 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4221 | 46.979 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483840 | Escherichia virus Qbeta | Escherichia virus Qbeta Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4218 | 47.866 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| FJ483839 | Escherichia virus BZ13 | Escherichia virus BZ13 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3458 | 48.178 | Levivirus | Unclassified | Leviviridae | Escherichia |
| FJ483838 | Escherichia virus BZ13 | Escherichia virus BZ13 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3393 | 46.036 | Levivirus | Unclassified | Leviviridae | Escherichia |
| FJ483837 | Escherichia virus BZ13 | Escherichia virus BZ13 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3412 | 47.098 | Levivirus | Unclassified | Leviviridae | Escherichia |
| EF204940 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3569 | 52.059 | Levivirus | Unclassified | Leviviridae | Escherichia |
| EF108465 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3569 | 51.583 | Levivirus | Unclassified | Leviviridae | Escherichia |
| EF108464 | Escherichia virus MS2 | Escherichia virus MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3569 | 51.919 | Levivirus | Unclassified | Leviviridae | Escherichia |
| EF204939 | Escherichia phage MS2 | Escherichia phage MS2 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3569 | 51.415 | Levivirus | Unclassified | Leviviridae | Escherichia |
| GU568037 | Deep-sea thermophilic phage D6E | Deep-sea thermophilic phage D6E Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49335 | 45.998 | Unclassified | Unclassified | Myoviridae | Unspecified |
| GU396103 | Aeromonas phage PX29 | Aeromonas phage PX29 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 222006 | 42.043 | Unclassified | Tevenvirinae | Myoviridae | Aeromonas |
| HM461982 | Burkholderia phage KS14 | Burkholderia phage KS14 Burkholderia virus KS14 Kisquattuordecimvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32317 | 62.28 | Kisquattuordecimvirus | Peduovirinae | Myoviridae | Burkholderia |
| GU911304 | Burkholderia phage KL3 | Burkholderia phage KL3 Burkholderia virus KL3 Tigrvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40555 | 63.225 | Tigrvirus | Peduovirinae | Myoviridae | Burkholderia |
| GU911303 | Burkholderia phage KS5 | Burkholderia phage KS5 Burkholderia virus KS5 Kisquinquevirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37236 | 63.707 | Kisquinquevirus | Peduovirinae | Myoviridae | Burkholderia |
| GU722197 | Hyperthermophilic Archaeal Virus 2 | Hyperthermophilic Archaeal Virus 2 unclassified archaeal viruses Viruses | 17666 | 52.083 | Unclassified | Unclassified | Unclassified | Unspecified |
| GU196280 | Escherichia phage K1ind2 | Escherichia phage K1ind2 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42765 | 51.348 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| GU196279 | Escherichia phage K1ind1 | Escherichia phage K1ind1 Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42292 | 51.272 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| GU196281 | Escherichia phage K1ind3 | Escherichia phage K1ind3 unclassified Kagunavirus Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43461 | 51.152 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| GU196278 | Escherichia phage K1H | Escherichia phage K1H Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41632 | 51.17 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| GU196277 | Escherichia phage K1G | Escherichia phage K1G Kagunavirus Guernseyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43587 | 51.068 | Kagunavirus | Guernseyvirinae | Siphoviridae | Escherichia |
| GQ468526 | Escherichia phage 285P | Escherichia phage 285P Escherichia virus 285P Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39270 | 48.732 | Berlinvirus | Studiervirinae | Autographiviridae | Escherichia |
| EU236438 | Pseudomonas phage PT2 | Pseudomonas phage PT2 Pseudomonas virus PT2 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42961 | 62.182 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| AM910651 | Pseudomonas phage LUZ19 | Pseudomonas phage LUZ19 Pseudomonas virus LUZ19 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43548 | 62.278 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| DQ394810 | Lactococcus phage phismq86 | Lactococcus phage phismq86 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33641 | 35.974 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| DQ394809 | Lactococcus phage ul36.t1k1 | Lactococcus phage ul36.t1k1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34897 | 35.831 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| DQ394808 | Lactococcus phage ul36.t1 | Lactococcus phage ul36.t1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35992 | 35.989 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| DQ394807 | Lactococcus phage ul36.k1t1 | Lactococcus phage ul36.k1t1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35594 | 35.734 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| DQ394806 | Lactococcus phage ul36.k1 | Lactococcus phage ul36.k1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37131 | 35.709 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AY605066 | Lactobacillus phage phiAT3 | Lactobacillus phage phiAT3 Lactobacillus virus phiAT3 Fattrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39166 | 44.592 | Fattrevirus | Unclassified | Siphoviridae | Lactobacillus |
| AF503408 | Enterobacteria phage P7 | Enterobacteria phage P7 Punavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 101660 | 47.411 | Punavirus | Unclassified | Myoviridae | Enterobacteria |
| FN582354 | Enterobacteria phage phi80 | Enterobacteria phage phi80 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 23229 | 53.498 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| HM137666 | Enterobacteria phage T4T | Enterobacteria phage T4T Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168920 | 35.291 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| HM032710 | Acinetobacter phage Ac42 | Acinetobacter phage Ac42 unclassified Tevenvirinae Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 167716 | 36.366 | Unclassified | Tevenvirinae | Myoviridae | Acinetobacter |
| GU815339 | Enterococcus phage EFRM31 | Enterococcus phage EFRM31 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16945 | 34.577 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| HM004124 | Acinetobacter phage Acj9 | Acinetobacter phage Acj9 Acinetobacter virus Acj9 Acajnonavirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 169947 | 40.028 | Acajnonavirus | Twarogvirinae | Myoviridae | Acinetobacter |
| GU911519 | Acinetobacter phage Acj61 | Acinetobacter phage Acj61 Acinetobacter virus Acj61 Lasallevirus Twarogvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 164093 | 39.006 | Lasallevirus | Twarogvirinae | Myoviridae | Acinetobacter |
| GU323318 | Enterobacter phage CC31 | Enterobacter phage CC31 Karamvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 165540 | 39.918 | Karamvirus | Tevenvirinae | Myoviridae | Enterobacter |
| HM029250 | Lactococcus phage 949 | Lactococcus phage 949 Lactococcus virus 949 Audreyjarvisvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 114768 | 32.689 | Audreyjarvisvirus | Unclassified | Siphoviridae | Lactococcus |
| GQ981382 | Shigella phage SP18 | Shigella phage SP18 Gaprivervirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170605 | 40.445 | Gaprivervirus | Tevenvirinae | Myoviridae | Shigella |
| HM242243 | Brochothrix phage A9 | Brochothrix phage A9 Brochothrix virus A9 Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127065 | 41.424 | Unclassified | Unclassified | Herelleviridae | Brochothrix |
| HM144386 | Brochothrix phage BL3 | Brochothrix phage BL3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41518 | 36.377 | Unclassified | Unclassified | Siphoviridae | Brochothrix |
| HM144385 | Brochothrix phage NF5 | Brochothrix phage NF5 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36953 | 37.296 | Unclassified | Unclassified | Siphoviridae | Brochothrix |
| FJ188381 | Stx2-converting phage 1717 | Stx2-converting phage 1717 Escherichia virus 1717 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 62147 | 50.915 | Pankowvirus | Unclassified | Siphoviridae | Unspecified |
| AY583234 | Mycoplasma phage phiMFV1 | Mycoplasma phage phiMFV1 unclassified bacterial viruses Viruses | 15757 | 24.998 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| BL2CG | Acholeplasma virus L2 | Acholeplasma virus L2 Plasmavirus Plasmaviridae Viruses | 11965 | 31.96 | Plasmavirus | Unclassified | Plasmaviridae | Acholeplasma |
| PMULEFTEN | Escherichia phage Mu | Escherichia phage Mu Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 10661 | 48.513 | Muvirus | Unclassified | Myoviridae | Escherichia |
| FJ792813 | Enterococcus phage EFAP-1 | Enterococcus phage EFAP-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 21115 | 36.718 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| HM775311 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.43 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM775310 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.598 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM775309 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.542 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM775308 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.56 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM775307 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM775306 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| HM159959 | Clostridium phage phiCTP1 | Clostridium phage phiCTP1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59199 | 31.256 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| GU949551 | Clostridium phage phiCD6356 | Clostridium phage phiCD6356 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37664 | 28.38 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| GQ357916 | Escherichia phage D108 | Escherichia phage D108 unclassified Muvirus Muvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37235 | 51.76 | Muvirus | Unclassified | Myoviridae | Escherichia |
| AY324828 | Pseudomonas virus Pf1 | Pseudomonas virus Pf1 Primolicivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 10675 | 57.255 | Primolicivirus | Unclassified | Inoviridae | Pseudomonas |
| AF320576 | Salmonella phage FelixO1 | Salmonella phage FelixO1 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 86155 | 39.006 | Felixounavirus | Ounavirinae | Myoviridae | Salmonella |
| SFU82619 | Enterobacteria phage SfV | Enterobacteria phage SfV Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37074 | 50.769 | Unclassified | Unclassified | Myoviridae | Enterobacteria |
| B2U02303 | Escherichia phage If1 | Escherichia phage If1 Infulavirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8454 | 43.707 | Infulavirus | Unclassified | Inoviridae | Escherichia |
| GQ379227 | Streptomyces phage phiSASD1 | Streptomyces phage phiSASD1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37068 | 66.254 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| AJ278322 | Corynebacterium phage BFK20 | Corynebacterium phage BFK20 Corynebacterium virus BFK20 Sasvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42972 | 56.162 | Sasvirus | Unclassified | Siphoviridae | Corynebacterium |
| GQ413937 | Klebsiella phage KP32 | Klebsiella phage KP32 Przondovirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41119 | 52.37 | Przondovirus | Studiervirinae | Autographiviridae | Klebsiella |
| FJ896200 | Vibrio phage VP93 | Vibrio phage VP93 Vibrio virus VP93 Maculvirus Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43931 | 49.109 | Maculvirus | Unclassified | Autographiviridae | Vibrio |
| EU136189 | Staphylococcus phage SAP-2 | Staphylococcus phage SAP-2 Staphylococcus virus SAP2 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 17938 | 28.944 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| EU887664 | Bacillus phage TP21-L | Bacillus phage TP21-L Lwoffvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37456 | 37.802 | Lwoffvirus | Unclassified | Siphoviridae | Bacillus |
| GU080336 | Sulfolobus turreted icosahedral virus 2 | Sulfolobus turreted icosahedral virus 2 Alphaturrivirus Turriviridae Belfryvirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 16622 | 36.734 | Alphaturrivirus | Unclassified | Turriviridae | Sulfolobus |
| GU385905 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| FJ849058 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AB370205 | Staphylococcus virus phiMR25 | Staphylococcus virus phiMR25 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44342 | 34.326 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| EU311208 | Escherichia phage Min27 | Escherichia phage Min27 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 63395 | 49.504 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| FJ870917 | Acidianus spindle-shaped virus 1 | Acidianus spindle-shaped virus 1 Betafusellovirus Fuselloviridae Viruses | 24186 | 37.695 | Betafusellovirus | Unclassified | Fuselloviridae | Acidianus |
| AF165214 | Pseudomonas virus D3 | Pseudomonas virus D3 Detrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 56426 | 57.805 | Detrevirus | Unclassified | Siphoviridae | Pseudomonas |
| AJ969242 | Spiroplasma phage SVGII3 | Spiroplasma phage SVGII3 Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7878 | 23.013 | Unclassified | Unclassified | Plectroviridae | Spiroplasma |
| DQ500118 | Rhizobium phage 16-3 | Rhizobium phage 16-3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60195 | 58.952 | Unclassified | Unclassified | Siphoviridae | Rhizobium |
| AF440571 | Sulfolobus islandicus filamentous virus | Sulfolobus islandicus filamentous virus Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 40900 | 33.352 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Sulfolobus |
| DQ530361 | Staphylococcus phage phiNM3 | Staphylococcus phage phiNM3 Staphylococcus virus phiNM3 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44061 | 32.993 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AY131267 | Lactobacillus phage Lc-Nu | Lactobacillus phage Lc-Nu Lactobacillus virus LcNu Lacnuvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36466 | 44.28 | Lacnuvirus | Unclassified | Siphoviridae | Lactobacillus |
| AY508486 | Staphylococcus virus 77 | Staphylococcus virus 77 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41708 | 33.533 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AY257527 | Bacillus phage Bam35c | Bacillus phage Bam35c Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14935 | 39.719 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| AY007505 | Streptococcus phage SM1 | Streptococcus phage SM1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34692 | 39.15 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AY247822 | Yersinia phage phiA1122 | Yersinia phage phiA1122 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37555 | 48.316 | Teseptimavirus | Studiervirinae | Autographiviridae | Yersinia |
| AY251033 | Yersinia virus L413C | Yersinia virus L413C Peduovirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 30728 | 52.106 | Peduovirus | Peduovirinae | Myoviridae | Yersinia |
| AY242528 | Vibrio phage VGJ | Vibrio phage VGJ Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7542 | 43.37 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| AF513033 | Staphylococcus phage P68 | Staphylococcus phage P68 unclassified Rosenblumvirus Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18227 | 29.325 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| AF513032 | Staphylococcus virus 44AHJD | Staphylococcus virus 44AHJD Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 16784 | 29.623 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| AF085222 | Streptococcus virus DT1 | Streptococcus virus DT1 Moineauvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34815 | 39.072 | Moineauvirus | Unclassified | Siphoviridae | Streptococcus |
| AF440695 | Natrialba phage PhiCh1 | Natrialba phage PhiCh1 Natrialba virus PhiCh1 Myohalovirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58498 | 61.888 | Myohalovirus | Unclassified | Myoviridae | Natrialba |
| AF489521 | Lactococcus phage 4268 | Lactococcus phage 4268 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36596 | 35.367 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF453500 | Vibrio phage VSK | Vibrio phage VSK Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6882 | 43.723 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| AF454431 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.616 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| AF396866 | Myxococcus phage Mx8 | Myxococcus phage Mx8 Myxococcus virus Mx8 Myxoctovirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49534 | 67.745 | Myxoctovirus | Unclassified | Podoviridae | Myxococcus |
| AF176025 | Lactococcus phage BK5-T | Lactococcus phage BK5-T Lactococcus virus BK5T Sandinevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40003 | 34.975 | Sandinevirus | Unclassified | Siphoviridae | Lactococcus |
| AY027935 | Haemophilus virus HP2 | Haemophilus virus HP2 Hpunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31508 | 39.936 | Hpunavirus | Peduovirinae | Myoviridae | Haemophilus |
| AF304433 | Lactococcus phage TP901-1 | Lactococcus phage TP901-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37667 | 35.378 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AF306496 | Bdellovibrio phage phiMH2K | Bdellovibrio phage phiMH2K Bdellomicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4594 | 46.104 | Bdellomicrovirus | Gokushovirinae | Microviridae | Bdellovibrio |
| AF301375 | Methanothermobacter phage psiM100 | Methanothermobacter phage psiM100 Psimunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 31007 | 45.745 | Psimunavirus | Unclassified | Siphoviridae | Methanothermobacter |
| GQ403788 | Lactococcus phage 1358 | Lactococcus phage 1358 Lactococcus virus 1358 Whiteheadvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36892 | 51.038 | Whiteheadvirus | Unclassified | Siphoviridae | Lactococcus |
| EU124666 | Thalassomonas phage BA3 | Thalassomonas phage BA3 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37313 | 40.889 | Unclassified | Unclassified | Podoviridae | Thalassomonas |
| AY374414 | Ralstonia phage p12J | Ralstonia phage p12J unclassified Inoviridae Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7118 | 56.954 | Unclassified | Unclassified | Inoviridae | Ralstonia |
| EF579802 | Microbacterium phage Min1 | Microbacterium phage Min1 Microbacterium virus Min1 Minunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46365 | 68.314 | Minunavirus | Unclassified | Siphoviridae | Microbacterium |
| EF432053 | Acidianus bottle-shaped virus | Acidianus bottle-shaped virus Bottigliavirus ABV Bottigliavirus Ampullaviridae Viruses | 23814 | 34.564 | Bottigliavirus | Unclassified | Ampullaviridae | Acidianus |
| EF455602 | Lactobacillus phage LL-H | Lactobacillus phage LL-H Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34659 | 47.817 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| AY722806 | Thermoproteus tenax spherical virus 1 | Thermoproteus tenax spherical virus 1 Alphaglobulovirus TTSV1 Alphaglobulovirus Globuloviridae Viruses | 20933 | 49.529 | Alphaglobulovirus | Unclassified | Globuloviridae | Thermoproteus |
| AF217253 | Salmonella phage P22 | Salmonella phage P22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41724 | 47.081 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AF227250 | Enterobacteria phage KU1 | Enterobacteria phage KU1 Levivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 3486 | 46.529 | Levivirus | Unclassified | Leviviridae | Enterobacteria |
| AF125520 | Escherichia phage 933W | Escherichia phage 933W Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61670 | 49.368 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| AF074945 | Mycoplasma phage MAV1 | Mycoplasma phage MAV1 unclassified dsDNA phages unclassified DNA viruses unclassified viruses Viruses | 15644 | 29.033 | Unclassified | Unclassified | Unclassified | Mycoplasma |
| AF065411 | Methanobacterium phage psiM2 | Methanobacterium phage psiM2 Psimunavirus psiM2 Psimunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26111 | 46.306 | Psimunavirus | Unclassified | Siphoviridae | Methanobacterium |
| AF059243 | Enterobacteria phage NL95 | Enterobacteria phage NL95 Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4248 | 50.824 | Allolevivirus | Unclassified | Leviviridae | Enterobacteria |
| AB366653 | Ralstonia phage phiRSL1 | Ralstonia phage phiRSL1 Mieseafarmvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 231255 | 58.026 | Mieseafarmvirus | Unclassified | Myoviridae | Ralstonia |
| GQ979703 | Lactococcus virus P2 | Lactococcus virus P2 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 27595 | 34.727 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| AP000363 | Enterobacteria phage VT2-Sakai | Enterobacteria phage VT2-Sakai unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60942 | 49.913 | Traversvirus | Sepvirinae | Podoviridae | Enterobacteria |
| GQ478088 | Enterococcus phage phiFL4A | Enterococcus phage phiFL4A Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37856 | 37.777 | Unclassified | Unclassified | Siphoviridae | Enterococcus |
| GQ478087 | Enterococcus phage phiFL3B | Enterococcus phage phiFL3B Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40275 | 34.498 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478086 | Enterococcus phage phiFL3A | Enterococcus phage phiFL3A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39576 | 34.546 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478085 | Enterococcus phage phiFL2B | Enterococcus phage phiFL2B Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36826 | 34.573 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478084 | Enterococcus phage phiFL2A | Enterococcus phage phiFL2A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36270 | 34.607 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478083 | Enterococcus phage phiFL1C | Enterococcus phage phiFL1C Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38721 | 34.023 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478082 | Enterococcus phage phiFL1B | Enterococcus phage phiFL1B Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38989 | 33.979 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| GQ478081 | Enterococcus phage phiFL1A | Enterococcus phage phiFL1A Phifelvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38764 | 34.014 | Phifelvirus | Unclassified | Siphoviridae | Enterococcus |
| FJ904927 | Vibrio phage VEJ | Vibrio phage VEJ Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6842 | 42.955 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| GQ153923 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.562 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| GQ153921 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.186 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| GQ153920 | Escherichia phage G4 | Escherichia phage G4 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.257 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| GQ153919 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.799 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| GQ153918 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.643 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| GQ153917 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.362 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| GQ153915 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.709 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GQ153914 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.634 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GQ153913 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.653 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GQ153912 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.597 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| GQ502199 | Sodalis phage SO1 | Sodalis phage SO1 Dhillonvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45169 | 54.582 | Dhillonvirus | Unclassified | Siphoviridae | Sodalis |
| GQ421471 | Xylella phage Xfas53 | Xylella phage Xfas53 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36674 | 56.757 | Unclassified | Unclassified | Podoviridae | Xylella |
| GQ398772 | Staphylococcus phage P954 | Staphylococcus phage P954 Staphylococcus virus P954 Biseptimavirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40761 | 33.993 | Biseptimavirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| FJ848885 | Lactococcus virus CB20 | Lactococcus virus CB20 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28625 | 35.025 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| FJ848884 | Lactococcus phage CB19 | Lactococcus phage CB19 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 28643 | 35.146 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| FJ848883 | Lactococcus phage CB14 | Lactococcus phage CB14 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29459 | 34.804 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| FJ848882 | Lactococcus phage CB13 | Lactococcus phage CB13 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32182 | 34.7 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| FJ982340 | Burkholderia phage KS9 | Burkholderia phage KS9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39896 | 60.68 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| GQ149088 | Escherichia phage St-1 | Escherichia phage St-1 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6094 | 45.208 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| FJ713816 | Staphylococcus phage phiPVL-CN125 | Staphylococcus phage phiPVL-CN125 Staphylococcus virus CN125 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44492 | 33.568 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| FJ591094 | Sulfitobacter phage EE36phi1 | Sulfitobacter phage EE36phi1 Sulfitobacter virus EE36phi1 Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 73325 | 47.04 | Aorunvirus | Rhodovirinae | Schitoviridae | Sulfitobacter |
| FJ591093 | Silicibacter phage DSS3phi2 | Silicibacter phage DSS3phi2 unclassified Aorunvirus Aorunvirus Rhodovirinae Schitoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 74611 | 47.915 | Aorunvirus | Rhodovirinae | Schitoviridae | Silicibacter |
| EU855793 | Listeria phage P40 | Listeria phage P40 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35638 | 39.259 | Unclassified | Unclassified | Siphoviridae | Listeria |
| DQ426905 | Mannheimia phage phiMhaA1-BAA410 | Mannheimia phage phiMhaA1-BAA410 Baylorvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34600 | 41.581 | Baylorvirus | Peduovirinae | Myoviridae | Mannheimia |
| DQ003642 | Listeria phage A006 | Listeria phage A006 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38124 | 35.497 | Unclassified | Unclassified | Siphoviridae | Listeria |
| DQ003641 | Listeria phage P35 | Listeria phage P35 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 35822 | 40.818 | Unclassified | Unclassified | Siphoviridae | Listeria |
| DQ003640 | Listeria phage B054 | Listeria phage B054 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 48172 | 36.212 | Unclassified | Unclassified | Myoviridae | Listeria |
| DQ003639 | Listeria phage B025 | Listeria phage B025 unclassified Slepowronvirus Slepowronvirus Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42653 | 35.1 | Slepowronvirus | Trabyvirinae | Siphoviridae | Listeria |
| DQ003637 | Listeria phage A500 | Listeria phage A500 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38867 | 36.684 | Unclassified | Unclassified | Siphoviridae | Listeria |
| DQ029335 | Vibrio phage VP4 | Vibrio phage VP4 Vibrio virus VP4 Chatterjeevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39503 | 42.612 | Chatterjeevirus | Studiervirinae | Autographiviridae | Vibrio |
| AY320035 | Streptomyces phage VWB | Streptomyces phage VWB Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49220 | 71.138 | Unclassified | Unclassified | Siphoviridae | Streptomyces |
| AY714348 | Vibrio phage KSF1 | Vibrio phage KSF1 Capistrivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7107 | 44.379 | Capistrivirus | Unclassified | Inoviridae | Vibrio |
| AF448724 | Sinorhizobium phage PBC5 | Sinorhizobium phage PBC5 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57416 | 61.486 | Unclassified | Unclassified | Podoviridae | Sinorhizobium |
| GQ334450 | Cyanophage PSS2 | Cyanophage PSS2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 107530 | 52.301 | Unclassified | Unclassified | Siphoviridae | Prochlorococcus |
| FJ867914 | Roseobacter virus SIO1 | Roseobacter virus SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38252 | 44.144 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| FJ867913 | Roseobacter virus SIO1 | Roseobacter virus SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38205 | 44.842 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| FJ867912 | Roseobacter virus SIO1 | Roseobacter virus SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38150 | 44.155 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| FJ867911 | Roseobacter virus SIO1 | Roseobacter virus SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38155 | 44.196 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| FJ867910 | Roseobacter virus SIO1 | Roseobacter virus SIO1 Siovirus Cobavirinae Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39413 | 45.257 | Siovirus | Cobavirinae | Zobellviridae | Roseobacter |
| DQ149023 | Synechococcus phage syn9 | Synechococcus phage syn9 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 177300 | 40.634 | Unclassified | Unclassified | Myoviridae | Synechococcus |
| DQ126339 | Escherichia phage phiV10 | Escherichia phage phiV10 Uetakevirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39104 | 48.972 | Uetakevirus | Unclassified | Podoviridae | Escherichia |
| DQ997624 | Thermus phage phiYS40 | Thermus phage phiYS40 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 152372 | 32.586 | Unclassified | Unclassified | Myoviridae | Thermus |
| DQ453159 | Geobacillus virus E2 | Geobacillus virus E2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40863 | 44.786 | Unclassified | Unclassified | Siphoviridae | Geobacillus |
| DQ203855 | Burkholderia phage Bcep176 | Burkholderia phage Bcep176 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44856 | 61.539 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| AY954964 | Staphylococcus virus 29 | Staphylococcus virus 29 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42802 | 35.375 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954963 | Staphylococcus virus 55 | Staphylococcus virus 55 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41902 | 35.724 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954962 | Staphylococcus virus 71 | Staphylococcus virus 71 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43114 | 35.239 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954961 | Staphylococcus phage ROSA | Staphylococcus phage ROSA Staphylococcus virus ROSA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43155 | 35.06 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954960 | Staphylococcus virus 96 | Staphylococcus virus 96 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43576 | 35.008 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954959 | Staphylococcus virus EW | Staphylococcus virus EW Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45286 | 35.991 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954958 | Staphylococcus virus 37 | Staphylococcus virus 37 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43681 | 35.114 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AY954957 | Staphylococcus virus 47 | Staphylococcus virus 47 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44777 | 33.459 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| AY954956 | Staphylococcus virus 3a | Staphylococcus virus 3a Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43095 | 33.463 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| AY954954 | Staphylococcus phage 2638A | Staphylococcus phage 2638A Staphylococcus virus 2638A Fibralongavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41318 | 36.916 | Fibralongavirus | Unclassified | Siphoviridae | Staphylococcus |
| AY954949 | Staphylococcus phage 66 | Staphylococcus phage 66 Staphylococcus virus 66 Rosenblumvirus Rakietenvirinae Rountreeviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18199 | 29.287 | Rosenblumvirus | Rakietenvirinae | Rountreeviridae | Staphylococcus |
| AY954948 | Staphylococcus phage PT1028 | Staphylococcus phage PT1028 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15603 | 31.417 | Unclassified | Unclassified | Siphoviridae | Staphylococcus |
| AY423772 | Sulfolobus virus Kamchatka 1 | Sulfolobus virus Kamchatka 1 Alphafusellovirus Fuselloviridae Viruses | 17385 | 38.746 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| AF299314 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5356 | 44.735 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299313 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5383 | 44.789 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299312 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5339 | 44.821 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299311 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5339 | 44.671 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299310 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.505 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299309 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.617 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299308 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5369 | 44.869 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299307 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5341 | 44.767 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299306 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5368 | 44.84 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299305 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5383 | 44.808 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299304 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.635 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299303 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5339 | 44.69 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299302 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.746 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299301 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5383 | 44.771 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| AF299300 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5359 | 44.635 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| CMU41758 | Chlamydia phage CPG1 | Chlamydia phage CPG1 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4529 | 40.98 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| MPU46938 | Mycobacterium phage DS6A | Mycobacterium phage DS6A Mycobacterium virus DS6A Hnatkovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15664 | 69.344 | Hnatkovirus | Unclassified | Siphoviridae | Mycobacterium |
| EU717894 | Xanthomonas phage phiL7 | Xanthomonas phage phiL7 Xanthomonas virus PhiL7 Eisenstarkvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44080 | 55.64 | Eisenstarkvirus | Unclassified | Siphoviridae | Xanthomonas |
| GU169904 | Staphylococcus phage SA1 | Staphylococcus phage SA1 unclassified Ounavirinae Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 147303 | 45.827 | Unclassified | Ounavirinae | Myoviridae | Staphylococcus |
| AY357582 | Burkholderia phage BcepNazgul | Burkholderia phage BcepNazgul Burkholderia virus BcepNazgul Nazgulvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 57455 | 60.639 | Nazgulvirus | Unclassified | Siphoviridae | Burkholderia |
| DQ289556 | Bacillus phage Gamma isolate d’Herelle | Bacillus phage Gamma isolate d’Herelle unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37373 | 35.13 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| DQ222855 | Bacillus phage Gamma | Bacillus phage Gamma unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38067 | 35.632 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| DQ222853 | Bacillus phage Gamma | Bacillus phage Gamma unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37253 | 35.221 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| DQ222851 | Bacillus phage Cherry | Bacillus phage Cherry unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36615 | 35.264 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| DQ150593 | Bacillus phage Fah | Bacillus phage Fah unclassified Wbetavirus Wbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37974 | 34.945 | Wbetavirus | Unclassified | Siphoviridae | Bacillus |
| DQ113772 | Streptococcus phage MM1 1998 | Streptococcus phage MM1 1998 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38893 | 38.11 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| DQ111067 | Escherichia phage K1F | Escherichia phage K1F Escherichia virus K1F Kayfunavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39704 | 49.776 | Kayfunavirus | Studiervirinae | Autographiviridae | Escherichia |
| DQ079909 | Enterobacteria phage WA14 | Enterobacteria phage WA14 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.078 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079908 | Enterobacteria phage ID18 | Enterobacteria phage ID18 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.224 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079906 | Enterobacteria phage NC10 | Enterobacteria phage NC10 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5529 | 46.139 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079905 | Enterobacteria phage ID12 | Enterobacteria phage ID12 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5529 | 46.319 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079904 | Enterobacteria phage WA6 | Enterobacteria phage WA6 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5529 | 46.464 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079903 | Enterobacteria phage ID41 | Enterobacteria phage ID41 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.795 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079902 | Enterobacteria phage NC19 | Enterobacteria phage NC19 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.813 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079901 | Enterobacteria phage NC13 | Enterobacteria phage NC13 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.082 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079900 | Enterobacteria phage NC2 | Enterobacteria phage NC2 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 46.279 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079899 | Enterobacteria phage WA5 | Enterobacteria phage WA5 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.957 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079898 | Enterobacteria phage ID8 | Enterobacteria phage ID8 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5540 | 46.3 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079897 | Enterobacteria phage WA3 | Enterobacteria phage WA3 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5540 | 46.029 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079896 | Enterobacteria phage WA2 | Enterobacteria phage WA2 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5540 | 46.083 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079895 | Enterobacteria phage WA11 | Enterobacteria phage WA11 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5387 | 44.552 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079894 | Enterobacteria phage WA10 | Enterobacteria phage WA10 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079893 | Enterobacteria phage WA4 | Enterobacteria phage WA4 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.541 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079892 | Enterobacteria phage NC56 | Enterobacteria phage NC56 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.374 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079891 | Enterobacteria phage NC51 | Enterobacteria phage NC51 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.393 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079890 | Enterobacteria phage NC41 | Enterobacteria phage NC41 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.579 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079889 | Enterobacteria phage NC37 | Enterobacteria phage NC37 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.504 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079888 | Enterobacteria phage NC16 | Enterobacteria phage NC16 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.449 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079887 | Enterobacteria phage NC11 | Enterobacteria phage NC11 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 45.08 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079886 | Enterobacteria phage NC7 | Enterobacteria phage NC7 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.876 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079885 | Enterobacteria phage NC5 | Enterobacteria phage NC5 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.486 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079884 | Enterobacteria phage NC1 | Enterobacteria phage NC1 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.838 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079883 | Enterobacteria phage ID45 | Enterobacteria phage ID45 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.467 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079882 | Enterobacteria phage ID34 | Enterobacteria phage ID34 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.727 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079881 | Enterobacteria phage ID22 | Enterobacteria phage ID22 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.356 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079880 | Enterobacteria phage ID1 | Enterobacteria phage ID1 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.486 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079879 | Escherichia phage NC29 | Escherichia phage NC29 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6259 | 44.352 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079878 | Enterobacteria phage NC3 | Enterobacteria phage NC3 unclassified Alphatrevirus Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6097 | 45.088 | Alphatrevirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079877 | Escherichia phage ID52 | Escherichia phage ID52 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5538 | 45.865 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| DQ079876 | Escherichia phage ID62 | Escherichia phage ID62 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6051 | 44.935 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079875 | Escherichia phage NC28 | Escherichia phage NC28 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6065 | 44.831 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079874 | Escherichia phage WA45 | Escherichia phage WA45 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 44.941 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079873 | Enterobacteria phage WA13 | Enterobacteria phage WA13 unclassified Alphatrevirus Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 44.825 | Alphatrevirus | Bullavirinae | Microviridae | Enterobacteria |
| DQ079872 | Escherichia phage NC35 | Escherichia phage NC35 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6039 | 44.726 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079871 | Escherichia phage ID32 | Escherichia phage ID32 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6071 | 45.034 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| DQ079870 | Escherichia phage ID21 | Escherichia phage ID21 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6068 | 45.27 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| AY283928 | Vibrio phage KVP40 | Vibrio phage KVP40 Schizotequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 244834 | 42.599 | Schizotequatrovirus | Tevenvirinae | Myoviridae | Vibrio |
| AY986977 | Xanthomonas phage Xp15 | Xanthomonas phage Xp15 Xanthomonas virus Xp15 Alachuavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55770 | 51.888 | Alachuavirus | Unclassified | Siphoviridae | Xanthomonas |
| AY848689 | Enterobacteria phage PRD1 | Enterobacteria phage PRD1 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14927 | 48.101 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY848688 | Enterobacteria phage PR772 | Enterobacteria phage PR772 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14942 | 48.354 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY848687 | Enterobacteria phage PR5 | Enterobacteria phage PR5 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14939 | 48.183 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY848686 | Enterobacteria phage PR4 | Enterobacteria phage PR4 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14954 | 48.322 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY848685 | Enterobacteria phage PR3 | Enterobacteria phage PR3 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14937 | 48.229 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY848684 | Enterobacteria phage L17 | Enterobacteria phage L17 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14935 | 48.256 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY751298 | Enterobacteria phage ID11 | Enterobacteria phage ID11 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5577 | 45.939 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| AY736146 | Enterobacteria phage ES18 | Enterobacteria phage ES18 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46900 | 48.593 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| AY772740 | Listonella phage phiHSIC | Listonella phage phiHSIC Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37966 | 43.963 | Unclassified | Unclassified | Siphoviridae | Listonella |
| AY701338 | Bacillus phage GIL16c | Bacillus phage GIL16c Betatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14844 | 40.07 | Betatectivirus | Unclassified | Tectiviridae | Bacillus |
| AY616446 | Bacillus phage BCASJ1c | Bacillus phage BCASJ1c Bacillus virus BCAJ1 Jarrellvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41092 | 41.74 | Jarrellvirus | Unclassified | Siphoviridae | Bacillus |
| AY576796 | Actinoplanes phage phiAsp2 | Actinoplanes phage phiAsp2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 58638 | 70.388 | Unclassified | Unclassified | Siphoviridae | Actinoplanes |
| AY575774 | Pseudomonas phage PaP2 | Pseudomonas phage PaP2 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43783 | 45.365 | Unclassified | Unclassified | Podoviridae | Pseudomonas |
| AF242738 | Lactococcus phage phiLC3 | Lactococcus phage phiLC3 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32172 | 35.509 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AY441783 | Enterobacteria phage PR772 | Enterobacteria phage PR772 Alphatectivirus Tectiviridae Kalamavirales Tectiliviricetes Preplasmiviricota Bamfordvirae Varidnaviria Viruses | 14946 | 48.341 | Alphatectivirus | Unclassified | Tectiviridae | Enterobacteria |
| AY505112 | Vibrio phage VP2 | Vibrio phage VP2 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39853 | 50.558 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| AY510084 | Vibrio phage VP5 | Vibrio phage VP5 Enhodamvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39786 | 50.485 | Enhodamvirus | Unclassified | Podoviridae | Vibrio |
| AY370673 | Salmonella phage SP6 | Salmonella phage SP6 Zindervirus Molineuxvirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43769 | 47.227 | Zindervirus | Molineuxvirinae | Autographiviridae | Salmonella |
| AY374448 | Klebsiella phage phiKO2 | Klebsiella phage phiKO2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 51601 | 51.485 | Unclassified | Unclassified | Siphoviridae | Klebsiella |
| AY388628 | Sulfolobus virus Ragged Hills | Sulfolobus virus Ragged Hills Alphafusellovirus Fuselloviridae Viruses | 16473 | 37.534 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| AY459535 | Lactobacillus prophage Lj965 | Lactobacillus prophage Lj965 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40190 | 35.141 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| AY459533 | Lactobacillus prophage Lj928 | Lactobacillus prophage Lj928 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38384 | 34.697 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| AY264774 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39938 | 48.398 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| AY176327 | Staphylococcus virus K | Staphylococcus virus K Kayvirus Twortvirinae Herelleviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 127395 | 30.602 | Kayvirus | Twortvirinae | Herelleviridae | Staphylococcus |
| AY370762 | Sulfolobus spindle-shaped virus 2 | Sulfolobus spindle-shaped virus 2 Alphafusellovirus Fuselloviridae Viruses | 14796 | 38.497 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| AF452449 | Vibrio phage VSKK | Vibrio phage VSKK Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6834 | 43.503 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| AF232233 | Pseudomonas phage B3 | Pseudomonas phage B3 Pseudomonas virus B3 Beetrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38439 | 63.16 | Beetrevirus | Unclassified | Siphoviridae | Pseudomonas |
| AY133112 | Vibrio phage VHML | Vibrio phage VHML Vhmlvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43198 | 50.618 | Vhmlvirus | Unclassified | Myoviridae | Vibrio |
| AY055382 | Salmonella phage ST64B | Salmonella phage ST64B Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40149 | 51.015 | Unclassified | Unclassified | Myoviridae | Salmonella |
| AY082070 | Clostridium phage phi3626 | Clostridium phage phi3626 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33507 | 28.382 | Unclassified | Unclassified | Siphoviridae | Clostridium |
| AF428260 | Propionibacterium phage B5 | Propionibacterium phage B5 Bifilivirus Paulinoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5804 | 64.283 | Bifilivirus | Unclassified | Paulinoviridae | Propionibacterium |
| AF335538 | Enterobacteria phage HK620 | Enterobacteria phage HK620 Escherichia virus HK620 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38297 | 46.688 | Lederbergvirus | Unclassified | Podoviridae | Escherichia |
| AF274751 | Enterobacteria phage S13 | Enterobacteria phage S13 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.319 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| AF020713 | Bacillus virus SPbeta | Bacillus virus SPbeta Spbetavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 134416 | 34.635 | Spbetavirus | Unclassified | Siphoviridae | Bacillus |
| BHU24159 | Haemophilus virus HP1 | Haemophilus virus HP1 Hpunavirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32355 | 40.006 | Hpunavirus | Peduovirinae | Myoviridae | Haemophilus |
| SPVDNA | Spiroplasma virus SpV4 | Spiroplasma virus SpV4 Spiromicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4421 | 32.052 | Spiromicrovirus | Gokushovirinae | Microviridae | Spiroplasma |
| PF3COMNY | Pseudomonas phage Pf3 | Pseudomonas phage Pf3 Tertilicivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 5833 | 45.38 | Tertilicivirus | Unclassified | Inoviridae | Pseudomonas |
| F1CCG | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.94 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| FJ870916 | Sulfolobus spindle-shaped virus 7 | Sulfolobus spindle-shaped virus 7 Alphafusellovirus Fuselloviridae Viruses | 17602 | 38.984 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| FJ870915 | Sulfolobus spindle-shaped virus 6 | Sulfolobus spindle-shaped virus 6 Betafusellovirus Fuselloviridae Viruses | 15684 | 38.192 | Betafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| FJ194439 | Kluyvera phage Kvp1 | Kluyvera phage Kvp1 Berlinvirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39472 | 48.642 | Berlinvirus | Studiervirinae | Autographiviridae | Kluyvera |
| EU622808 | Bacillus phage Nf | Bacillus phage Nf Beecentumtrevirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18753 | 37.317 | Beecentumtrevirus | Picovirinae | Salasmaviridae | Bacillus |
| EU570103 | Salmonella phage epsilon34 | Salmonella phage epsilon34 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43016 | 47.264 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AY587007 | Escherichia phage T5 | Escherichia phage T5 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 121752 | 39.326 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| AY692264 | Escherichia phage T5 | Escherichia phage T5 Tequintavirus Markadamsvirinae Demerecviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 113238 | 39.204 | Tequintavirus | Markadamsvirinae | Demerecviridae | Escherichia |
| AF527608 | Salmonella phage P22-pbi | Salmonella phage P22-pbi Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41724 | 47.09 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| ACSJ01000013 | Clostridium phage D-1873 | Clostridium phage D-1873 unclassified bacterial viruses Viruses | 10531 | 25.61 | Unclassified | Unclassified | Unclassified | Clostridium |
| ACSJ01000015 | Clostridium phage D-1873 | Clostridium phage D-1873 unclassified bacterial viruses Viruses | 17272 | 25.272 | Unclassified | Unclassified | Unclassified | Clostridium |
| ACSJ01000017 | Clostridium phage D-1873 | Clostridium phage D-1873 unclassified bacterial viruses Viruses | 74599 | 25.696 | Unclassified | Unclassified | Unclassified | Clostridium |
| ACSJ01000018 | Clostridium phage D-1873 | Clostridium phage D-1873 unclassified bacterial viruses Viruses | 14466 | 25.964 | Unclassified | Unclassified | Unclassified | Clostridium |
| EU877232 | Escherichia phage wV8 | Escherichia phage wV8 Felixounavirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 88487 | 38.894 | Felixounavirus | Ounavirinae | Myoviridae | Escherichia |
| EU863409 | Escherichia phage JS10 | Escherichia phage JS10 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 171451 | 39.515 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| EU863408 | Escherichia phage JSE | Escherichia phage JSE Krischvirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 166418 | 40.512 | Krischvirus | Tevenvirinae | Myoviridae | Escherichia |
| FJ429185 | Lactococcus phage P087 | Lactococcus phage P087 Lactococcus virus P087 Teubervirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60074 | 34.384 | Teubervirus | Unclassified | Siphoviridae | Lactococcus |
| FJ839693 | Enterobacteria phage RB51 | Enterobacteria phage RB51 unclassified Tequatrovirus Tequatrovirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 168394 | 35.253 | Tequatrovirus | Tevenvirinae | Myoviridae | Enterobacteria |
| EU221285 | Lactococcus phage bIBB29 | Lactococcus phage bIBB29 Skunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29305 | 34.724 | Skunavirus | Unclassified | Siphoviridae | Lactococcus |
| DQ079907 | Enterobacteria phage NC6 | Enterobacteria phage NC6 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5529 | 45.795 | Gequatrovirus | Bullavirinae | Microviridae | Enterobacteria |
| S13CG | Enterobacteria phage S13 | Enterobacteria phage S13 unclassified Sinsheimervirus Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.337 | Sinsheimervirus | Bullavirinae | Microviridae | Enterobacteria |
| PZACG | Bacillus phage PZA | Bacillus phage PZA Bacillus virus PZA Salasvirus Picovirinae Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 19366 | 39.662 | Salasvirus | Picovirinae | Salasmaviridae | Bacillus |
| EU710883 | Erwinia phage phiEa21-4 | Erwinia phage phiEa21-4 Kolesnikvirus Ounavirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 84576 | 43.813 | Kolesnikvirus | Ounavirinae | Myoviridae | Erwinia |
| FN263372 | Pseudomonas phage phikF77 | Pseudomonas phage phikF77 Pseudomonas virus kF77 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43152 | 62.868 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| DQ340064 | Geobacillus phage GBSV1 | Geobacillus phage GBSV1 Svunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34683 | 44.379 | Svunavirus | Unclassified | Myoviridae | Geobacillus |
| EU822883 | Burkholderia phage KS10 | Burkholderia phage KS10 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37635 | 62.87 | Unclassified | Unclassified | Myoviridae | Burkholderia |
| AB255436 | Stx2-converting phage 86 | Stx2-converting phage 86 Escherichia virus 86 Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 60238 | 49.067 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AP005153 | Escherichia Stx1 converting phage | Escherichia Stx1 converting phage unclassified Traversvirus Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 59866 | 49.694 | Traversvirus | Sepvirinae | Podoviridae | Escherichia |
| AP004402 | Stx2 converting phage I | Stx2 converting phage I Traversvirus Sepvirinae Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 61765 | 49.389 | Traversvirus | Sepvirinae | Podoviridae | Unspecified |
| AB043679 | Vibrio phage VfO4K68 | Vibrio phage VfO4K68 Versovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6891 | 47.236 | Versovirus | Unclassified | Inoviridae | Vibrio |
| AB043678 | Vibrio phage VfO3K6 | Vibrio phage VfO3K6 Versovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8784 | 45.162 | Versovirus | Unclassified | Inoviridae | Vibrio |
| AB009866 | Staphylococcus phage PVL | Staphylococcus phage PVL Staphylococcus virus PVL Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41401 | 33.555 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AB002632 | Vibrio phage fs2 | Vibrio phage fs2 Saetivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 8651 | 44.527 | Saetivirus | Unclassified | Inoviridae | Vibrio |
| AP001553 | Staphylococcus virus phiETA | Staphylococcus virus phiETA Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43081 | 35.431 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AP009624 | Pseudomonas phage PAJU2 | Pseudomonas phage PAJU2 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46872 | 56.264 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| EU871039 | Lactobacillus phage Lv-1 | Lactobacillus phage Lv-1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38934 | 37.042 | Unclassified | Unclassified | Siphoviridae | Lactobacillus |
| AB451219 | Ralstonia phage RSB1 | Ralstonia phage RSB1 Ralstonia virus RSB1 Higashivirus Okabevirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43079 | 61.738 | Higashivirus | Okabevirinae | Autographiviridae | Ralstonia |
| EU849489 | Stenotrophomonas phage S1 | Stenotrophomonas phage S1 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40287 | 63.691 | Unclassified | Unclassified | Siphoviridae | Stenotrophomonas |
| AP009146 | Sodalis phage phiSG1 | Sodalis phage phiSG1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52162 | 50.826 | Unclassified | Unclassified | Podoviridae | Sodalis |
| EU350361 | Abalone shriveling syndrome-associated virus | Abalone shriveling syndrome-associated virus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 34952 | 39.52 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| AB044554 | Staphylococcus prophage phiPV83 | Staphylococcus prophage phiPV83 Staphylococcus virus PV83 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 45636 | 33.476 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AB012574 | Vibrio phage Vf12 | Vibrio phage Vf12 Villovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7965 | 45.675 | Villovirus | Unclassified | Inoviridae | Vibrio |
| AB012573 | Vibrio virus Vf33 | Vibrio virus Vf33 Villovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7965 | 45.662 | Villovirus | Unclassified | Inoviridae | Vibrio |
| AB426868 | Salmonella phage P22 | Salmonella phage P22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41660 | 47.012 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AB370268 | Staphylococcus virus phiMR11 | Staphylococcus virus phiMR11 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43011 | 35.63 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AP008954 | Staphylococcus virus phiETA3 | Staphylococcus virus phiETA3 Phietavirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43282 | 34.899 | Phietavirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AP008953 | Staphylococcus virus phiETA2 | Staphylococcus virus phiETA2 Dubowvirus Azeredovirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43265 | 34.279 | Dubowvirus | Azeredovirinae | Siphoviridae | Staphylococcus |
| AB102868 | Enterobacteria phage ST104 | Enterobacteria phage ST104 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41391 | 47.431 | Lederbergvirus | Unclassified | Podoviridae | Enterobacteria |
| AP009363 | Staphylococcus phage phi2958PVL | Staphylococcus phage phi2958PVL Staphylococcus virus phi2958PVL Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47342 | 33.04 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| AB362338 | Salmonella phage P22 | Salmonella phage P22 Lederbergvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 41660 | 47.012 | Lederbergvirus | Unclassified | Podoviridae | Salmonella |
| AB285204 | Enterobacteria phage cdtI | Enterobacteria phage cdtI Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47021 | 49.116 | Unclassified | Unclassified | Siphoviridae | Enterobacteria |
| EU547803 | Salmonella phage phiSG-JL2 | Salmonella phage phiSG-JL2 Salmonella virus SG-JL2 Teetrevirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38815 | 50.864 | Teetrevirus | Studiervirinae | Autographiviridae | Salmonella |
| EU794049 | Bacteriophage APSE-2 | Bacteriophage APSE-2 Hamiltonella virus APSE2 Sendosyvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39867 | 42.91 | Sendosyvirus | Unclassified | Podoviridae | Unspecified |
| AJ783769 | Sulfolobus virus STSV1 | Sulfolobus virus STSV1 unclassified Bicaudaviridae Bicaudaviridae Viruses | 75294 | 35.215 | Unclassified | Unclassified | Bicaudaviridae | Sulfolobus |
| AY894696 | Bacillus thuringiensis phage MZTP02 | Bacillus thuringiensis phage MZTP02 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 15717 | 37.552 | Unclassified | Unclassified | Siphoviridae | Bacillus |
| AF133242 | Spiroplasma phage SVTS2 | Spiroplasma phage SVTS2 Suturavirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6825 | 22.652 | Suturavirus | Unclassified | Plectroviridae | Spiroplasma |
| FJ184280 | Enterobacteria phage YYZ-2008 | Enterobacteria phage YYZ-2008 Enterobacteria virus YYZ2008 Pankowvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 54896 | 51.115 | Pankowvirus | Unclassified | Siphoviridae | Enterobacteria |
| FJ008913 | Bacteroides phage B40-8 | Bacteroides phage B40-8 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44929 | 38.628 | Unclassified | Unclassified | Siphoviridae | Bacteroides |
| DQ838728 | Lactococcus phage P335 | Lactococcus phage P335 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33613 | 35.486 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| DQ499600 | Corynebacterium phage P1201 | Corynebacterium phage P1201 Corynebacterium virus P1201 Chunghsingvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 70579 | 50.919 | Chunghsingvirus | Unclassified | Siphoviridae | Corynebacterium |
| EU545650 | Acidianus filamentous virus 9 | Acidianus filamentous virus 9 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 41172 | 34.645 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| EU246945 | Lactobacillus phage Lrm1 | Lactobacillus phage Lrm1 Lactobacillus virus Lrm1 Larmunavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39989 | 45.49 | Larmunavirus | Unclassified | Siphoviridae | Lactobacillus |
| EU056923 | Pseudomonas phage PT5 | Pseudomonas phage PT5 Pseudomonas virus PT5 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42954 | 62.332 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| DQ085810 | Escherichia phage alpha3 | Escherichia phage alpha3 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6089 | 45.18 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| AB045978 | Staphylococcus virus phiSLT | Staphylococcus virus phiSLT Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42942 | 33.312 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| EU197055 | Pseudomonas phage 201phi2-1 | Pseudomonas phage 201phi2-1 unclassified Phikzvirus Phikzvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 316674 | 45.344 | Phikzvirus | Unclassified | Myoviridae | Pseudomonas |
| EU438902 | Lactococcus phage asccphi28 | Lactococcus phage asccphi28 Salasmaviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 18762 | 33.658 | Unclassified | Unclassified | Salasmaviridae | Lactococcus |
| EU081845 | Lactococcus phage 1706 | Lactococcus phage 1706 Lactococcus virus 1706 Fremauxvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 55597 | 33.689 | Fremauxvirus | Unclassified | Siphoviridae | Lactococcus |
| AM087121 | Acidianus filamentous virus 6 | Acidianus filamentous virus 6 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 39577 | 36.87 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| AM087120 | Acidianus filamentous virus 3 | Acidianus filamentous virus 3 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 40449 | 36.854 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| AM087123 | Acidianus filamentous virus 8 | Acidianus filamentous virus 8 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 38179 | 36.803 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| AM087122 | Acidianus filamentous virus 7 | Acidianus filamentous virus 7 Betalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 36895 | 37.973 | Betalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| DQ079869 | Escherichia phage ID2 Moscow/ID/2001 | Escherichia phage ID2 Moscow/ID/2001 Gequatrovirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5486 | 45.753 | Gequatrovirus | Bullavirinae | Microviridae | Escherichia |
| EF469154 | Escherichia phage JS98 | Escherichia phage JS98 Dhakavirus Tevenvirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 170523 | 39.513 | Dhakavirus | Tevenvirinae | Myoviridae | Escherichia |
| EF071488 | Pyrococcus abyssi virus 1 | Pyrococcus abyssi virus 1 unclassified archaeal dsDNA viruses unclassified DNA viruses unclassified viruses Viruses | 18098 | 47.16 | Unclassified | Unclassified | Unclassified | Pyrococcus |
| AY950802 | Haloarcula hispanica virus SH1 | Haloarcula hispanica virus SH1 Alphasphaerolipovirus Sphaerolipoviridae Halopanivirales Laserviricetes Dividoviricota Helvetiavirae Varidnaviria Viruses | 30889 | 68.384 | Alphasphaerolipovirus | Unclassified | Sphaerolipoviridae | Haloarcula |
| AF191796 | His 1 virus | His 1 virus Salterprovirus His1 Salterprovirus Halspiviridae Viruses | 14462 | 38.854 | Salterprovirus | Unclassified | Halspiviridae | Unspecified |
| L67CG | Lactococcus phage bIL67 | Lactococcus phage bIL67 Ceduovirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 22195 | 35.99 | Ceduovirus | Unclassified | Siphoviridae | Lactococcus |
| AF195902 | Lactobacillus prophage Lj771 | Lactobacillus prophage Lj771 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40881 | 36.401 | Unclassified | Unclassified | Myoviridae | Lactobacillus |
| AB374228 | Vibrio virus Kappa | Vibrio virus Kappa unclassified Longwoodvirus Longwoodvirus Peduovirinae Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 33507 | 48.835 | Longwoodvirus | Peduovirinae | Myoviridae | Vibrio |
| EF460875 | Escherichia phage phiEcoM-GJ1 | Escherichia phage phiEcoM-GJ1 Escherichia virus GJ1 Carltongylesvirus Cleopatravirinae Chaseviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52975 | 44.023 | Carltongylesvirus | Cleopatravirinae | Chaseviridae | Escherichia |
| AM076770 | Yersinia phage PY100 | Yersinia phage PY100 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 50291 | 46.841 | Unclassified | Unclassified | Myoviridae | Yersinia |
| EF583821 | Bacillus phage 0305phi8-36 | Bacillus phage 0305phi8-36 Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 218948 | 41.795 | Unclassified | Unclassified | Myoviridae | Bacillus |
| SVU28974 | Spiroplasma phage C74 | Spiroplasma phage C74 Vespertiliovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 7768 | 23.159 | Vespertiliovirus | Unclassified | Plectroviridae | Spiroplasma |
| BCP1 | Chlamydia virus Chp1 | Chlamydia virus Chp1 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4877 | 36.498 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| EF462199 | Staphylococcus phage tp310-3 | Staphylococcus phage tp310-3 unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42973 | 33.454 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| EF462198 | Staphylococcus phage tp310-2 | Staphylococcus phage tp310-2 Staphylococcus virus tp310-2 Triavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 47785 | 33.751 | Triavirus | Unclassified | Siphoviridae | Staphylococcus |
| EF462197 | Staphylococcus phage tp310-1 | Staphylococcus phage tp310-1 Staphylococcus virus tp310-1 Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42232 | 33.503 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| AB036666 | Wolbachia phage WO | Wolbachia phage WO Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 32987 | 35.008 | Unclassified | Unclassified | Myoviridae | Wolbachia |
| DQ864624 | Streptococcus phage P9 | Streptococcus phage P9 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 40539 | 39.651 | Unclassified | Unclassified | Siphoviridae | Streptococcus |
| AB259124 | Ralstonia phage RSS1 | Ralstonia phage RSS1 Restivirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6662 | 62.609 | Restivirus | Unclassified | Inoviridae | Ralstonia |
| AB334721 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.846 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| AB334720 | Enterobacteria phage f1 | Enterobacteria phage f1 Inovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6407 | 40.94 | Inovirus | Unclassified | Inoviridae | Enterobacteria |
| DQ431235 | Propionibacterium phage PA6 | Propionibacterium phage PA6 Pahexavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 29739 | 54.023 | Pahexavirus | Unclassified | Siphoviridae | Propionibacterium |
| EF523948 | Burkholderia phage BcepGomr | Burkholderia phage BcepGomr Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52414 | 56.294 | Unclassified | Unclassified | Siphoviridae | Burkholderia |
| BL5GENM | Leuconostoc phage L5 | Leuconostoc phage L5 unclassified bacterial viruses Viruses | 2435 | 33.265 | Unclassified | Unclassified | Unclassified | Leuconostoc |
| AB243556 | Staphylococcus virus 108PVL | Staphylococcus virus 108PVL unclassified Peeveelvirus Peeveelvirus Bronfenbrennervirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 44857 | 33.469 | Peeveelvirus | Bronfenbrennervirinae | Siphoviridae | Staphylococcus |
| DQ785801 | Sodalis phage phiSG1 | Sodalis phage phiSG1 Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 52162 | 50.826 | Unclassified | Unclassified | Podoviridae | Sodalis |
| EF207558 | Streptococcus phage phi3396 | Streptococcus phage phi3396 Trabyvirinae Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38528 | 37.482 | Unclassified | Trabyvirinae | Siphoviridae | Streptococcus |
| EF380032 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.579 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380031 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.56 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380030 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.022 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380029 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 43.817 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380028 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.04 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380027 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.059 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380026 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 43.91 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380025 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.207 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380024 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.281 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380023 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 43.929 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380022 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 43.892 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380021 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.356 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380020 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.3 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380019 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.244 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380018 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.152 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380017 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.337 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380016 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.3 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380015 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.281 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380014 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.207 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380013 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.152 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380012 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.003 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380011 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.133 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380010 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.263 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF380009 | Escherichia phage phiX174 | Escherichia phage phiX174 Sinsheimervirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 5386 | 44.653 | Sinsheimervirus | Bullavirinae | Microviridae | Escherichia |
| EF068134 | Escherichia virus FI | Escherichia virus FI Allolevivirus Leviviridae Levivirales Leviviricetes Lenarviricota Orthornavirae Riboviria Viruses | 4184 | 51.243 | Allolevivirus | Unclassified | Leviviridae | Escherichia |
| AJ567472 | Acidianus filamentous virus 1 | Acidianus filamentous virus 1 Gammalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 20869 | 36.94 | Gammalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| AJ842011 | Flavobacterium phage 11b | Flavobacterium phage 11b Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36012 | 30.593 | Unclassified | Unclassified | Siphoviridae | Flavobacterium |
| AM419438 | Archaeal BJ1 virus | Archaeal BJ1 virus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42271 | 64.886 | Unclassified | Unclassified | Siphoviridae | Unspecified |
| AF349457 | Lactococcus phage ul36 | Lactococcus phage ul36 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 36798 | 35.836 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| AY264778 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39085 | 47.635 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| AY264777 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39096 | 47.665 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| AY264776 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 37664 | 47.486 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| AY264775 | Escherichia phage T7 | Escherichia phage T7 Teseptimavirus Studiervirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39093 | 47.451 | Teseptimavirus | Studiervirinae | Autographiviridae | Escherichia |
| AJ854042 | Acidianus filamentous virus 2 | Acidianus filamentous virus 2 Deltalipothrixvirus Lipothrixviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 31787 | 35.656 | Deltalipothrixvirus | Unclassified | Lipothrixviridae | Acidianus |
| AM265638 | Pseudomonas phage LKD16 | Pseudomonas phage LKD16 Pseudomonas virus LKD16 Phikmvvirus Krylovirinae Autographiviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43200 | 62.317 | Phikmvvirus | Krylovirinae | Autographiviridae | Pseudomonas |
| D89074 | Vibrio virus fs1 | Vibrio virus fs1 Fibrovirus Inoviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 6340 | 43.423 | Fibrovirus | Unclassified | Inoviridae | Vibrio |
| DQ490056 | Lactococcus phage Q54 | Lactococcus phage Q54 Lactococcus virus Q54 Questintvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 26537 | 37.099 | Questintvirus | Unclassified | Siphoviridae | Lactococcus |
| AP008986 | Xanthomonas phage OP2 | Xanthomonas phage OP2 Naesvirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 46643 | 60.882 | Naesvirus | Unclassified | Myoviridae | Xanthomonas |
| AP008979 | Xanthomonas virus OP1 | Xanthomonas virus OP1 Xipdecavirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 43785 | 51.075 | Xipdecavirus | Unclassified | Siphoviridae | Xanthomonas |
| AJ888457 | Acidianus two-tailed virus | Acidianus two-tailed virus Bicaudavirus Bicaudaviridae Viruses | 62730 | 41.226 | Bicaudavirus | Unclassified | Bicaudaviridae | Acidianus |
| DQ163917 | Pseudomonas phage F8 | Pseudomonas phage F8 Pbunavirus Myoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 66015 | 54.944 | Pbunavirus | Unclassified | Myoviridae | Pseudomonas |
| DQ163915 | Pseudomonas phage PA11 | Pseudomonas phage PA11 Pseudomonas virus PA11 Paundecimvirus Zobellviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 49639 | 44.783 | Paundecimvirus | Unclassified | Zobellviridae | Pseudomonas |
| DQ163913 | Pseudomonas phage 73 | Pseudomonas phage 73 Septimatrevirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42999 | 53.559 | Septimatrevirus | Unclassified | Siphoviridae | Pseudomonas |
| DQ163912 | Pseudomonas phage F10 | Pseudomonas phage F10 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39199 | 62.083 | Unclassified | Unclassified | Siphoviridae | Pseudomonas |
| AJ875026 | Acidianus rod-shaped virus 1 | Acidianus rod-shaped virus 1 Itarudivirus ARV1 Itarudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 24655 | 39.128 | Itarudivirus | Unclassified | Rudiviridae | Acidianus |
| AJ414696 | Sulfolobus islandicus rod-shaped virus 1 | Sulfolobus islandicus rod-shaped virus 1 Icerudivirus SIRV1 Icerudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 32308 | 25.269 | Icerudivirus | Unclassified | Rudiviridae | Sulfolobus |
| AJ344259 | Sulfolobus islandicus rod-shaped virus 2 | Sulfolobus islandicus rod-shaped virus 2 Icerudivirus SIRV2 Icerudivirus Rudiviridae Ligamenvirales Tokiviricetes Taleaviricota Zilligvirae Adnaviria Viruses | 35450 | 25.199 | Icerudivirus | Unclassified | Rudiviridae | Sulfolobus |
| X07234 | Sulfolobus spindle-shaped virus 1 | Sulfolobus spindle-shaped virus 1 Alphafusellovirus Fuselloviridae Viruses | 15465 | 39.696 | Alphafusellovirus | Unclassified | Fuselloviridae | Sulfolobus |
| X60323 | Escherichia phage phiK | Escherichia phage phiK Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6089 | 44.917 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| X60322 | Escherichia phage alpha3 | Escherichia phage alpha3 Alphatrevirus Bullavirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 6087 | 45.178 | Alphatrevirus | Bullavirinae | Microviridae | Escherichia |
| AJ635161 | Pyrobaculum spherical virus | Pyrobaculum spherical virus Alphaglobulovirus PSV Alphaglobulovirus Globuloviridae Viruses | 28337 | 48.4 | Alphaglobulovirus | Unclassified | Globuloviridae | Pyrobaculum |
| AJ550635 | Chlamydia phage 3 | Chlamydia phage 3 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4554 | 41.041 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| AY526909 | Bordetella phage BIP-1 | Bordetella phage BIP-1 Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42638 | 65.402 | Rauchvirus | Unclassified | Podoviridae | Bordetella |
| AY526908 | Bordetella phage BMP-1 | Bordetella phage BMP-1 Rauchvirus Podoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 42663 | 65.438 | Rauchvirus | Unclassified | Podoviridae | Bordetella |
| AJ270057 | Chlamydia phage 2 | Chlamydia phage 2 Chlamydiamicrovirus Gokushovirinae Microviridae Petitvirales Malgrandaviricetes Phixviricota Sangervirae Monodnaviria Viruses | 4563 | 40.938 | Chlamydiamicrovirus | Gokushovirinae | Microviridae | Chlamydia |
| AB016282 | Bacillus phage phi105 | Bacillus phage phi105 Bacillus virus phi105 Spizizenvirus Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 39325 | 42.69 | Spizizenvirus | Unclassified | Siphoviridae | Bacillus |
| AF109874 | Lactococcus phage Tuc2009 | Lactococcus phage Tuc2009 Siphoviridae Caudovirales Caudoviricetes Uroviricota Heunggongvirae Duplodnaviria Viruses | 38347 | 36.222 | Unclassified | Unclassified | Siphoviridae | Lactococcus |
| X58839 | Acholeplasma phage MV-L51 | Acholeplasma phage MV-L51 Plectrovirus Plectroviridae Tubulavirales Faserviricetes Hofneiviricota Loebvirae Monodnaviria Viruses | 4491 | 33.289 | Plectrovirus | Unclassified | Plectroviridae | Acholeplasma |