EVAL-FARMS: Evaluating the Threat of Antimicrobial Resistance in Agricultural Manures and Slurries is led by Pro Dov Stekel at the University of Nottingham.
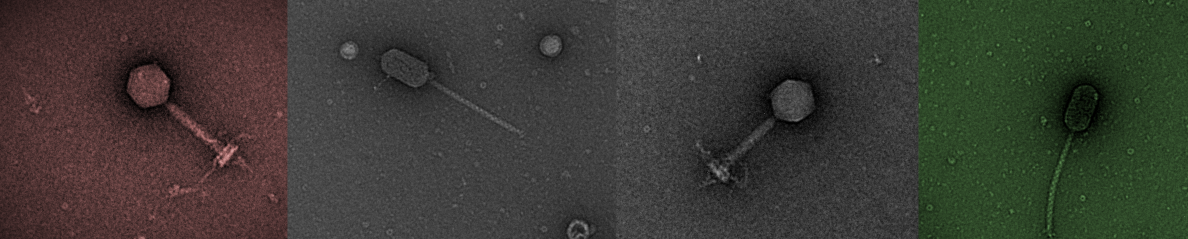
The project overview as described here is below. We are contributing to this project on the microbiology and metagenomics work package. In particular the analysis of the viral fraction of the slurry tank. We have been studying this project over several year and is a rich source of many phages used in BSc and MSc projects. Some of this phage isolation and small scale metagenomics has been published,
- Metagenomics of the Viral Community in Three Cattle Slurry Samples -PDF
- Comparative Genomics of Bacteriophage of the Genus Seuratvirus PDF
- Draft genome sequences of 14 Escherichia coli phages isolated from cattle slurry -PDF
Project overview
“Antibiotics are used extensively to fight bacterial infections and have saved millions of lives. However, the bacteria are becoming resistant to antibiotics and some antibiotics have stopped working. We refer to this as antimicrobial resistance – AMR. We don’t just use antibiotics for people; similar amounts are given to farm animals. More than 900 million farm animals are reared every year in the UK and antibiotic treatments are vital for their welfare, for farms as businesses, and for us to enjoy affordable food. However, farms may be contributing to the development of AMR. The aim of this project is to improve our understanding of how farm practice, especially the way in which manure is handled, could lead to AMR in animal and human pathogens. This understanding will help farmers and vets find new ways to reduce AMR, without harming their animals or their businesses.
For research purposes, Nottingham University maintains a typical high performance dairy farm – its 200 cows produce a lot of milk and a lot of manure. The waste is stored in a 3 million litre slurry tank, any excess goes into a 7 million litre lagoon. This slurry is applied to fields as organic fertilizer. Cow manure contains many harmless bacteria but some, e.g. E. coli O157, can cause severe infection in people. When cows get sick they are treated with antibiotics. Udder infections are treated by injection of antibiotics into the udder. Since this milk contains antibiotics, it cannot be sold but is discarded into the slurry. Foot infections are treated with an antibacterial footbath, which is also emptied into the slurry tank.
As a result, slurry tanks contain a mixture of bacteria, antibiotics and other antimicrobials that are stored for many months. The bacteria that survive in the presence of antibiotics are more likely to have antibiotic resistance. This resistance is encoded in their genes so passed to the next generation. Worse still, the genes can be passed on to other bacteria in the slurry.
Before we wrote this proposal, we investigated our own farm’s slurry tank to see if this might be happening. We tested 160 E. coli strains from the slurry; most carried antibiotic resistance. We also found antibiotics in the tank – including some that bacteria were resistant to. Our mathematical modellers showed that reducing spread of resistance genes in the tank might be more effective in preventing resistance than cutting the use of antibiotics. Conversations with the farm vets revealed that they knew about AMR and had changed some of their antibiotic prescriptions. But these analyses leave us with more questions than answers.
In this project, we want to find out if current farming methods are contributing to the development of harmful antibiotic-resistant bacteria in slurry, bacteria that may then be encountered by humans and animals. To do this, we need to integrate scientific and cultural approaches:
- What bacteria are in the slurry? How many are harmful? What resistance genes do they carry? How do these genes spread?
- How long do antibiotics remain in the tank? Do they degrade?
- What happens to the bacteria and antibiotics after they are spread on fields?
- How do farmers, vets and scientists interpret evidence about AMR? What are their hidden assumptions? Can we improve collaborative decision making on AMR risk management?
- Can we reduce resistance by avoiding mixing together bacteria and antimicrobials in slurry?
- Can we predict the risk of emergence of and exposure to resistant pathogens? Can we predict which interventions are likely to be most effective to reduce AMR, taking into account both human and scientific factors?
Through this research, we will learn what can realistically be done to reduce this risk; not just on this farm, but UK wide. We will work with farmers, vets and policy makers to ensure that our results will make a difference to reducing the risk of harmful AMR bacteria arising in agriculture.